ad210dbe1ce8ad0f3f5ff335fe91394d.ppt
- Количество слайдов: 47

Worldwide Protein Data Bank www. wwpdb. org September 7, 2007

Worldwide Protein Data Bank www. wwpdb. org Agenda § Welcome and introductions § Accomplishments § Remediation rollout summary § Toward the future Break § Matters arising – Incorrect structures § Executive session § Feedback to ww. PDB § Set next meeting date

Worldwide Protein Data Bank www. wwpdb. org ww. PDB Achievements October 2006 - September 2007 § § § § Continued growth of archive Website updates Publications and presentations Time-stamped archive Remediation rollout Annotation document One stop shop: NMR, cryo. EM
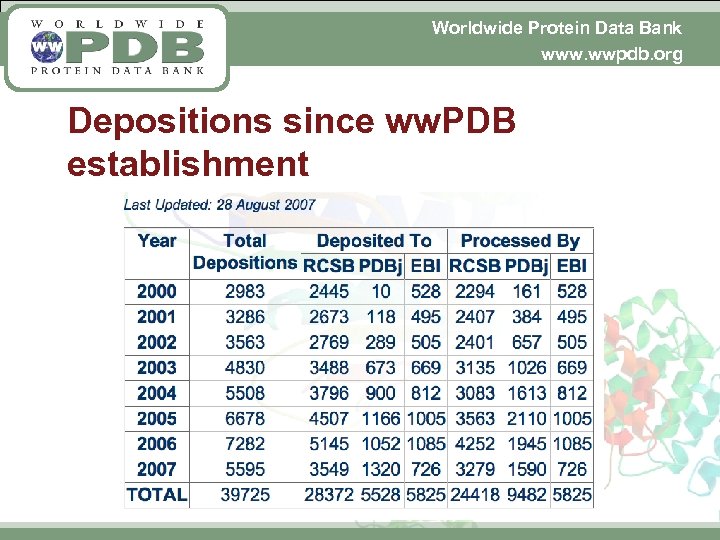
Worldwide Protein Data Bank www. wwpdb. org Depositions since ww. PDB establishment

Worldwide Protein Data Bank www. wwpdb. org PDB entry processing § 1 -1 -2000 10, 997 entries in PDB § Today 10 -Jul-2007 44, 578 entries in PDB Size now is 4 times larger than when the 3 sites started § In 1999, 2361 entries were deposited § In 2006, 7282 entries were deposited We handle more than 3 times as many entries per year with less staff – and all ww. PDB sites produce high quality annotated PDB entries No current backlog of unprocessed entries

Worldwide Protein Data Bank www. wwpdb. org Time-stamped copies of the archive § 57 Gbytes of data for 2006, released January 2, 2007 § 68 Gbytes of data for July 2007 snapshot § Both include – – – PDB format entries mm. CIF format entries PDBML format entries Experimental data Dictionary, schema, and format documentation

Worldwide Protein Data Bank www. wwpdb. org Outreach § § § ww. PDB website Discussion forums NMR Task Force Publications Professional society meetings

Worldwide Protein Data Bank www. wwpdb. org

Worldwide Protein Data Bank www. wwpdb. org Joint publications § Nucleic Acids Research, 35: D 301 (2007) – The worldwide Protein Data Bank (ww. PDB): ensuring a single, uniform archive of PDB data § Nature Structure Molecular Biology, 14: 354 (2007) – Reply to: Building meaningful models of glycoproteins § Nature Biotechnology, 25: 854 (2007) – Response to “Overhauling the PDB” § Methods in Molecular Biology, in press – Data deposition and annotation at the ww. PDB § Structural Bioinformatics 2 nd Edition, in press – The ww. PDB

Worldwide Protein Data Bank www. wwpdb. org Interactions since October 2006 § Exchange visits – – MSD/RCSB PDB (4) PDBj/RCSB PDB (1) PDBj/BMRB (2) BMRB/RCSB PDB (1) § Phone conference with site directors-twice a year § VTC’s among staff – BMRB/RCSB PDB twice a month (ADIT-NMR) – MSD/RCSB PDB twice a week (annotation procedures, remediation) – RCSB PDB/PDBj and BMRB/PDBj on necessary occasions § Email among staff – MSD/RCSB PDB ~2 per day – PDBj/RCSB PDB ~2 per day

Worldwide Protein Data Bank www. wwpdb. org New initiatives § One stop shop for NMR data and models § One stop shop for electron microscopy maps and models (NIH-funded)

Worldwide Protein Data Bank www. wwpdb. org Recommendations from 2006 ww. PDBAC report § Implement the recommendations from November 1920 2005 modeling workshop (Berman et al. Structure 14, 1211 -1217) – Models phased out October 16, 2006 § Rollout remediated data to superusers by December 31, 2006; to all users by July 1 st 2007; Provide access to PDB formatted files following the most current format. – Superusers had access to data November 2006, all users in April 2007

Worldwide Protein Data Bank www. wwpdb. org Recommendations from 2006 ww. PDBAC report § Work with SAXS community to create appropriate representation of these data, and circulate progress reports to the Committee as appropriate – Not done § Expand the four character PDB ID codes before the number of depositions reaches 400, 000 – Number of available PDB ID codes has been increased by allowing IDs to start with a character § Develop and present a formal recommendation to the ww. PDBAC regarding the purview of the PDB at our September 2007 meeting in Princeton, NJ – In process

Worldwide Protein Data Bank www. wwpdb. org Recommendations from 2006 ww. PDBAC report § Coordinate with the ww. PDBAC to obtain formal letters of support when seeking funding; establish a coordinated plan to both educate and lobby funding agency representatives; establish a charitable organization to serve as a conduit for receipt of both grant funding and gifts from pharmaceutical and biotechnology companies, involving individual Committee members as needed. – Funding Representatives Round Table Discussion

Worldwide Protein Data Bank www. wwpdb. org Remediation

Worldwide Protein Data Bank www. wwpdb. org Key drivers § § Chemistry and nomenclature Sequence and taxonomy Citations Viruses

Worldwide Protein Data Bank www. wwpdb. org IUPAC, NMR, and the PDB Atom nomenclature and NMR restraints John L. Markley

Worldwide Protein Data Bank www. wwpdb. org History of the NMR-led requested remediation of hydrogen atom nomenclature § When BMRB was established in the late 1980’s, it adopted the IUPAC atom nomenclature recommendations from Biochemistry 9, 3471 -3479, 1970 § At that time, we noted that NMR structures being deposited in the PDB did not adhere to these recommendations (particularly for H-atoms; e. g. HB 1/HB 2 instead of HB 2/HB 3), and I brought this to the attention of the director of the PDB at Brookhaven with the request that it be remedied § A group of NMR spectroscopists led by Kurt Wüthrich worked with the NMR community to develop recommendations for the deposition of NMR structures; all agreed that the prior IUPAC recommendations be maintained (Pure & Appl. Chem. , 70, 117 -142, 1998) § Over the years, ww. PDB Task Force on NMR has pushed strongly for remediation of atom nomenclature

Worldwide Protein Data Bank www. wwpdb. org Accomplished: atom nomenclature remediation § Nomenclature in PDB now matches that in BMRB § The single format will avoid confusion and errors § All discrepancies have been resolved in the remediated files, with the minor exception of atoms at the C-terminus IUPAC-IUBMB-IUPAB ww. PDB H'' HXT O' O O'' OXT – Since these atoms are not observed by NMR spectroscopists, we do not consider this to be a problem – We plan to write an addendum to the IUPAC-IUBMB-IUPAB “Recommendations” for submission to Pure & Appl. Chem. to formalize these as “accepted atom designators”

Worldwide Protein Data Bank www. wwpdb. org Remediation of NMR structure files § Required the linking of structure files and restraint files § Atom names, residue numbers and chain identifiers needed to be updated § Remediation of restraint files required the unpacking, parsing, and regularization of legacy information contained in PDB “MR” files into the “NMR Restraints Grid”

Worldwide Protein Data Bank www. wwpdb. org NMR Restraints Grid development § BMRB, University of Wisconsin-Madison, USA § MSD, European Bioinformatics Institute, Hinxton, UK § Department of Computer Sciences/Condor Project, University of Wisconsin, USA § Department of NMR Spectroscopy, Utrecht University, The Netherlands § Centre for Molecular and Biomolecular Informatics, Radboud University, The Netherlands

Worldwide Protein Data Bank www. wwpdb. org NMR Restraints Grid development § PDB MR files are converted into NMR-STAR § NMR-STAR file and the corresponding PDB coordinate file are parsed; the information is connected inside the CCPN framework; and the results are written out as NMR-STAR files; converted restraint files are filtered to remove redundant restraints § Files made available in the NMR Restraints Grid with access from links in each corresponding PDB entry § NMR restraint data files with atom nomenclature corresponding to remediated PDB data files will be available by the end of 2007

Worldwide Protein Data Bank www. wwpdb. org Current state of the NMR Restraints Grid § Grid contains 3583 entries with a total of 3, 882, 595 parsed restraints § 3583 entries out of 6508 in PDB have restraints § Database is updated continuously as new PDB entries are released that have associated NMR restraints

Worldwide Protein Data Bank www. wwpdb. org Recent agenda items considered by the ww. PDB NMR Task Force § Strongly recommend that restraints be mandatory for all NMR depositions to the PDB § Commissioned the development of procedures for representing uncertainty in NMR structures and for specifying the single model meant to be most representative of the structure § Task Force should write an article for J. Biomol. NMR on its recommendations for data representation and submission of experimental data § It was suggested that the Task Force begin to discuss validation issues

Worldwide Protein Data Bank www. wwpdb. org Most X-ray structures are supported by structure factors

Worldwide Protein Data Bank www. wwpdb. org Less than half of NMR structures are supported by restraint data
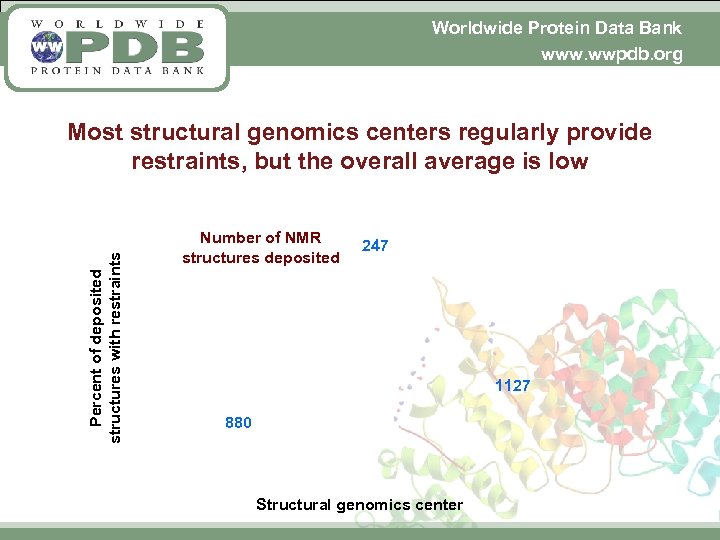
Worldwide Protein Data Bank www. wwpdb. org Percent of deposited structures with restraints Most structural genomics centers regularly provide restraints, but the overall average is low Number of NMR structures deposited 247 1127 880 Structural genomics center

Worldwide Protein Data Bank www. wwpdb. org Remediation rollout Helen M. Berman
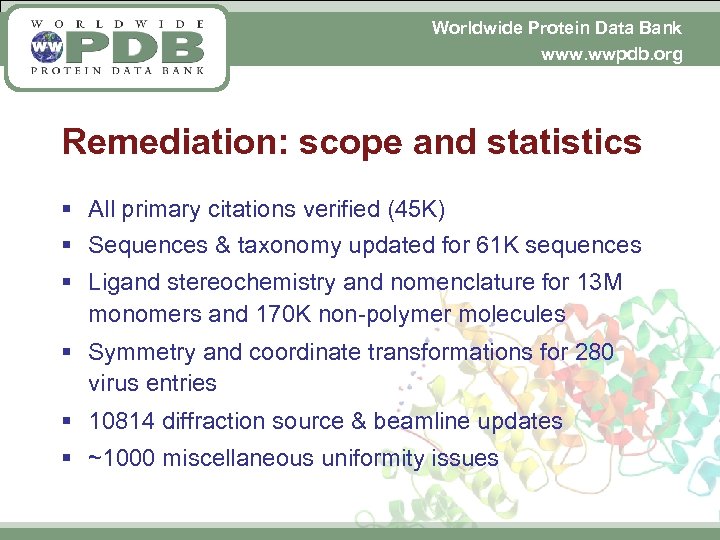

Worldwide Protein Data Bank www. wwpdb. org Remediation: scope and statistics § All primary citations verified (45 K) § Sequences & taxonomy updated for 61 K sequences § Ligand stereochemistry and nomenclature for 13 M monomers and 170 K non-polymer molecules § Symmetry and coordinate transformations for 280 virus entries § 10814 diffraction source & beamline updates § ~1000 miscellaneous uniformity issues
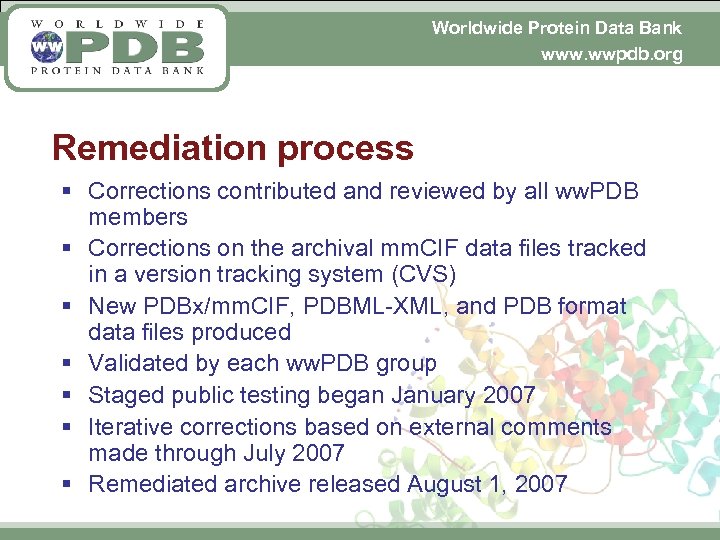

Worldwide Protein Data Bank www. wwpdb. org Remediation process § Corrections contributed and reviewed by all ww. PDB members § Corrections on the archival mm. CIF data files tracked in a version tracking system (CVS) § New PDBx/mm. CIF, PDBML-XML, and PDB format data files produced § Validated by each ww. PDB group § Staged public testing began January 2007 § Iterative corrections based on external comments made through July 2007 § Remediated archive released August 1, 2007

Worldwide Protein Data Bank www. wwpdb. org Remediation-supporting infrastructure § Internal (ww. PDB) CVS archive remediation data files § Internal (ww. PDB) rsync distribution site for remediated data files § Early tests of web, rsync, & ftp distribution sites for dictionaries, PDB, mm. CIF, and XML data files § Complete ww. PDB ftp site for remediated data and dictionaries updated with remediation corrections and weekly PDB updates § 200 K CVS remediated data file updates § 1 M+ remediated file updates to support testing and distribute from January 2007 - present

Worldwide Protein Data Bank www. wwpdb. org Checking the remediated files Haruki Nakamura

Worldwide Protein Data Bank www. wwpdb. org Different checks § § § References to external databases Data processing consistency checks PDBML/XML validation Database loads User-contributed diagnostics

Worldwide Protein Data Bank www. wwpdb. org References to external databases § Sequence and taxonomy (Uni. Prot) § Primary Citations (Pub. Med)

Worldwide Protein Data Bank www. wwpdb. org Data processing consistency checks § Covalent geometry and stereochemistry § Compliance with ww. PDB Chemical Component Dictionary – Molecular and stereochemical assignment – Atom and residue nomenclature § Compliance with PDB Exchange Dictionary – Data types, controlled vocabularies, parent-child relations § External tools such as What. IF

Worldwide Protein Data Bank www. wwpdb. org PDBML/XML schema validation § § § Version control Data type consistency Data ranges Controlled vocabularies Referential integrity XPath traversal of PDBML data hierarchy

Worldwide Protein Data Bank www. wwpdb. org Database loads § Diagnostics obtained from loading remediated data into existing database systems – Relational databases used by MSD-EBI and RCSB PDB – XML database used by PDBj

Worldwide Protein Data Bank www. wwpdb. org User-contributed diagnostics § Batch checking of remediated files by Phenix revealed consistency issues with alternate conformations - Ralf Grosse-Kunstleve § Batch checking for inconsistent linkages and missing residues by docking software - Tommy Carstensen § Nomenclature - Tom Goddard & Chimera Group § Sequence and assembly diagnostics - Roland Dunbrack § Relational data integrity diagnostics - Dan Bosler § Nomenclature and experimental details - Clemens Vonrhein § Many specific issues related to chemical assignments, disorder, and nomenclature

Worldwide Protein Data Bank www. wwpdb. org Looking toward the future Kim Henrick

Worldwide Protein Data Bank www. wwpdb. org Annotation project § Standardize annotation rules and policies among ww. PDB sites § Document annotation rules and policies § Create venue to update annotation rules and policies as necessary

Worldwide Protein Data Bank www. wwpdb. org Annotation project How did we get there? § Review and discussion of each PDB field by email and VTC § Document written and reviewed by all staff § Final review by site directors § Software compliant to new annotation procedures implemented § Tested software and trained annotators § Published document on web (January 2007)

Worldwide Protein Data Bank www. wwpdb. org Annotation document § Specification of ALL fields in PDB file § Clarification of policies – Assignment of PDB IDs – Release of files and information – Changes to entries § Clarification of data representation – Chain ID for all atoms in the file – Multi-model representation for alternate conformation or disorder – Chimeras – Microheterogenity

Worldwide Protein Data Bank www. wwpdb. org PDB IDs and DOIs § Credit for a PDB entry in CVs § Used as a reference in publications – http: //dx. doi. org/10. 2210/p db 4 hhb/pdb See also DOIs for Biological Databases Philip E. Bourne, Cross. Ref 7 th Annual Meeting, 1 November 2006 Cambridge, MA

Worldwide Protein Data Bank www. wwpdb. org Outstanding issues § Microheterogeniety § Disorder § Large structures

Worldwide Protein Data Bank www. wwpdb. org ww. PDB and software developers § ACA 24 th July 2007 meeting in Salt Lake City § “Future Challenges for the PDB: What should the PDB be doing in 2015? ” § Attended by software developers and ww. PDB staff

Worldwide Protein Data Bank www. wwpdb. org July 24 meeting § Technical discussions § TLS § Multiple models § Large structure § demand for one file per structure § Microheterogeneity § Twinning § George Sheldrick, Paul Adams and Garib Murshudov produce a draft of the PDB format to describe twinning and to represent the data in HKLF § Procedural outcomes § Yearly developer meeting § Editorial board to assist in difficult annotation problems § Ongoing electronic forum

Worldwide Protein Data Bank www. wwpdb. org Toward a single processing tool § This weekend – ww. PDB retreat with contributors from RCSB PDB Rutgers and UCSD, BMRB, PDBj, and EBI-EMBL § Task – come to agreement to pool resources to produce a single deposition tool and design of new processing pipeline
ad210dbe1ce8ad0f3f5ff335fe91394d.ppt