 Why protein structure?
Why protein structure?
 How does a protein fold?
How does a protein fold?
 The basics
The basics
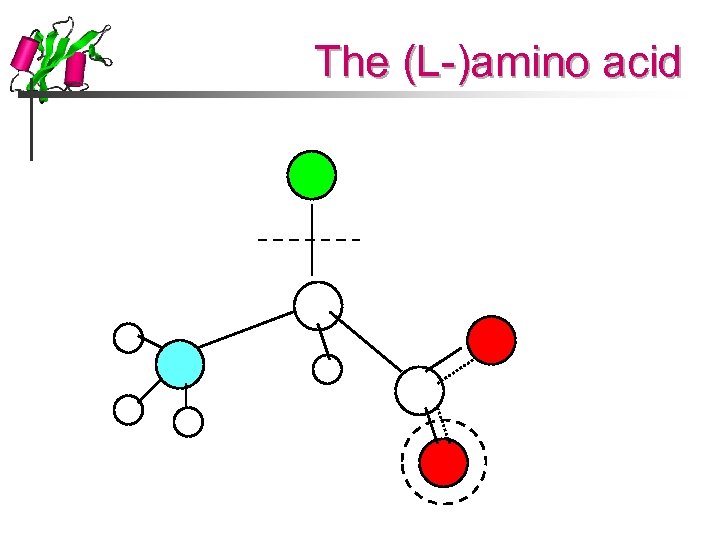 The (L-)amino acid
The (L-)amino acid
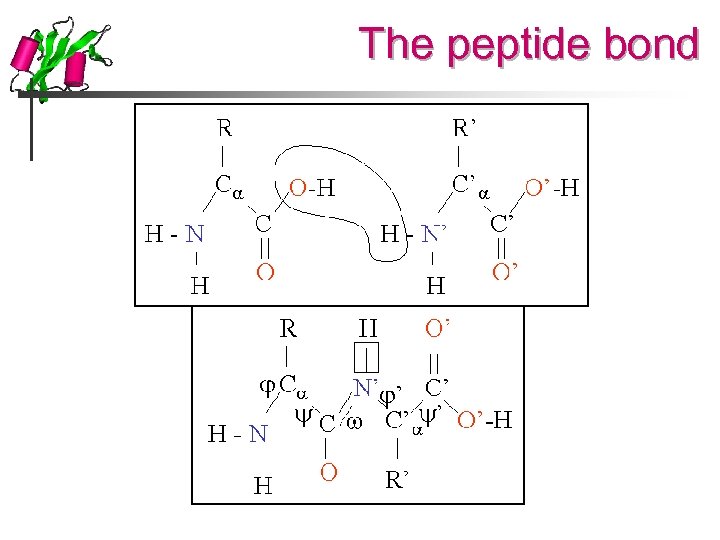 The peptide bond
The peptide bond
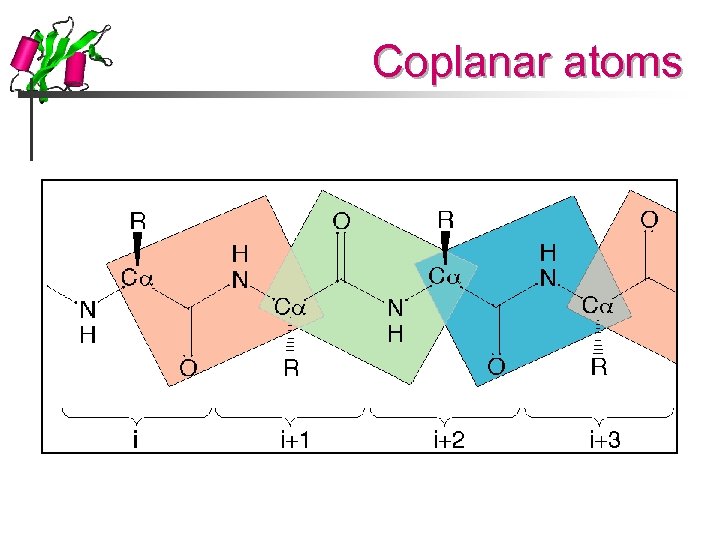 Coplanar atoms
Coplanar atoms
 Levels of protein structure
Levels of protein structure
 Levels of protein structure
Levels of protein structure
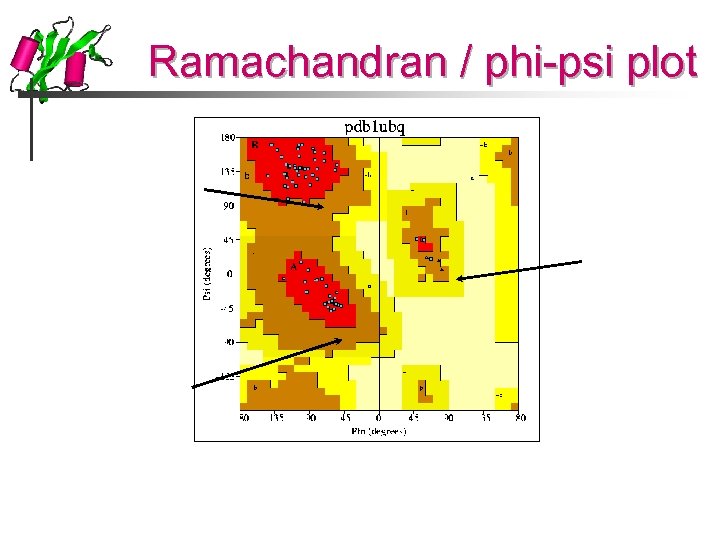 Ramachandran / phi-psi plot
Ramachandran / phi-psi plot
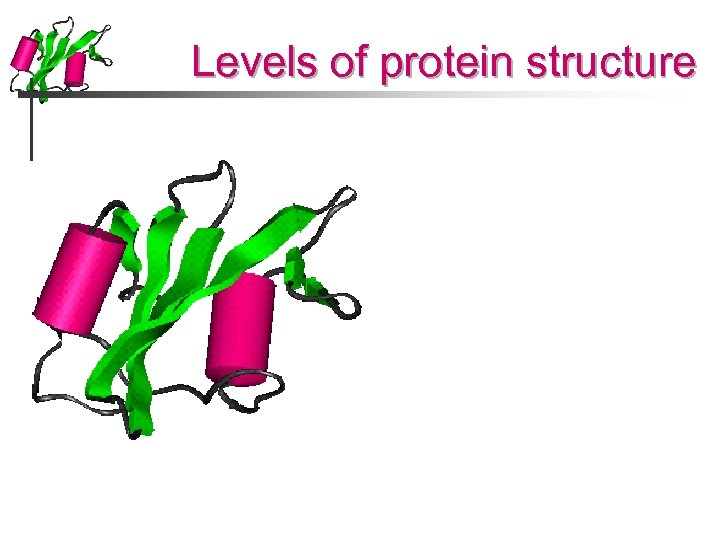 Levels of protein structure
Levels of protein structure
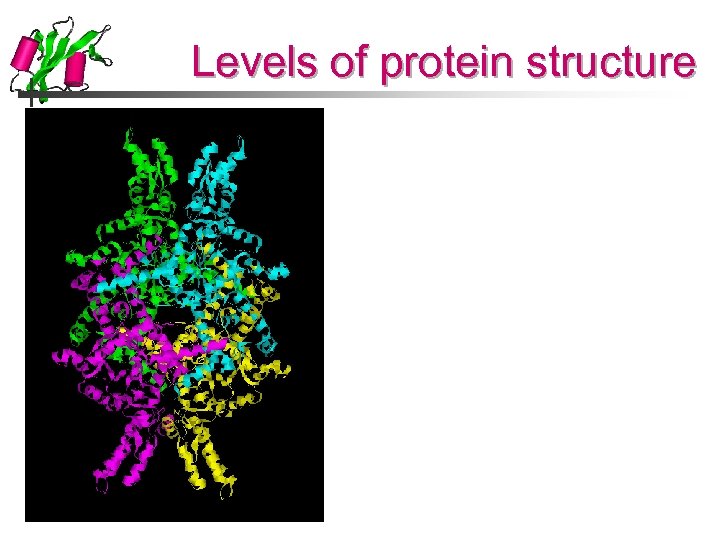 Levels of protein structure
Levels of protein structure
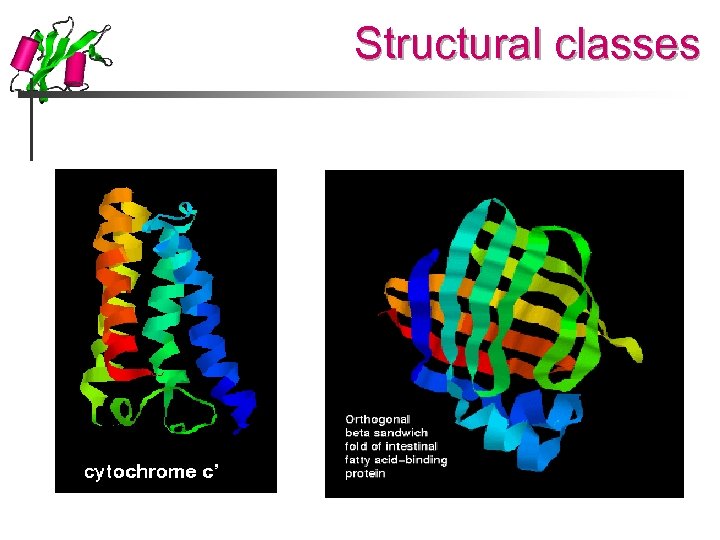 Structural classes
Structural classes
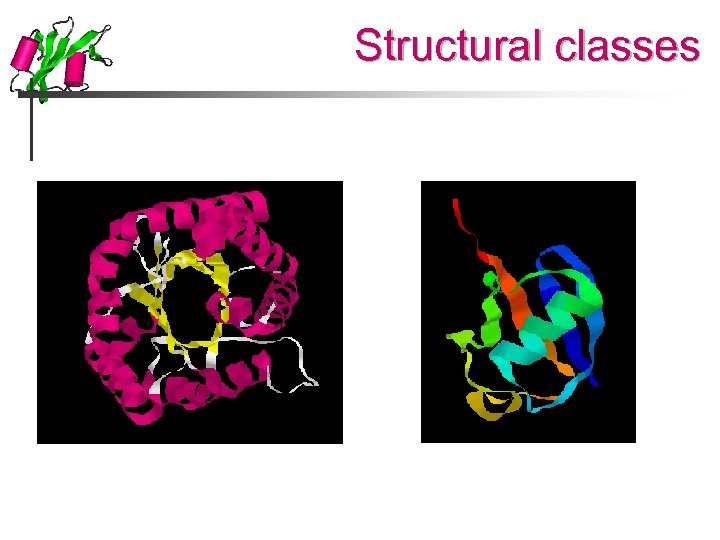 Structural classes
Structural classes
 Structural information
Structural information
 The PDB data
The PDB data
 PDB Header details
PDB Header details
 The data itself
The data itself
 Structural Families
Structural Families
 Structure comparison facts
Structure comparison facts
 Structure comparison facts
Structure comparison facts
 Visualizing PDB information
Visualizing PDB information
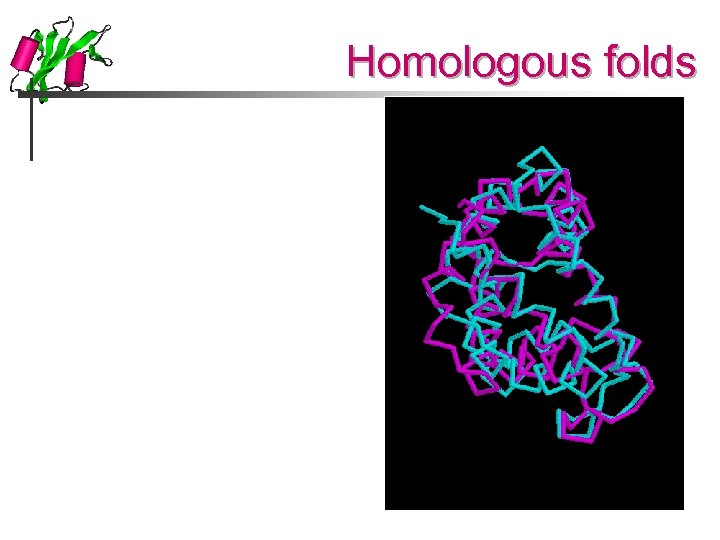 Homologous folds
Homologous folds
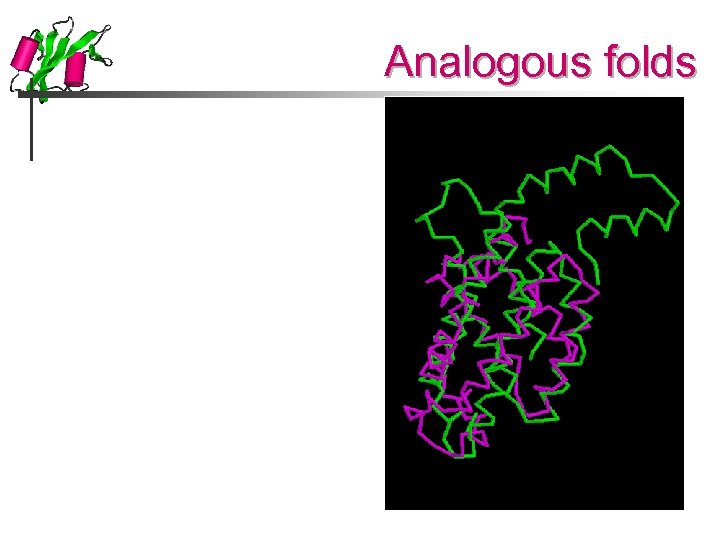 Analogous folds
Analogous folds
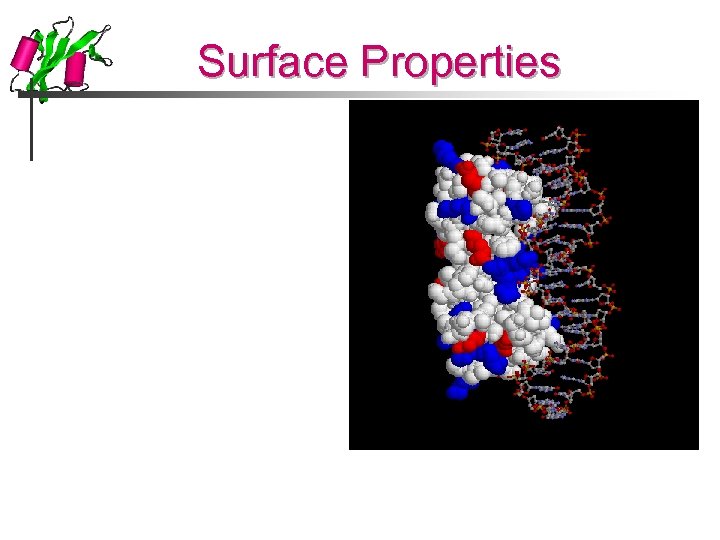 Surface Properties
Surface Properties
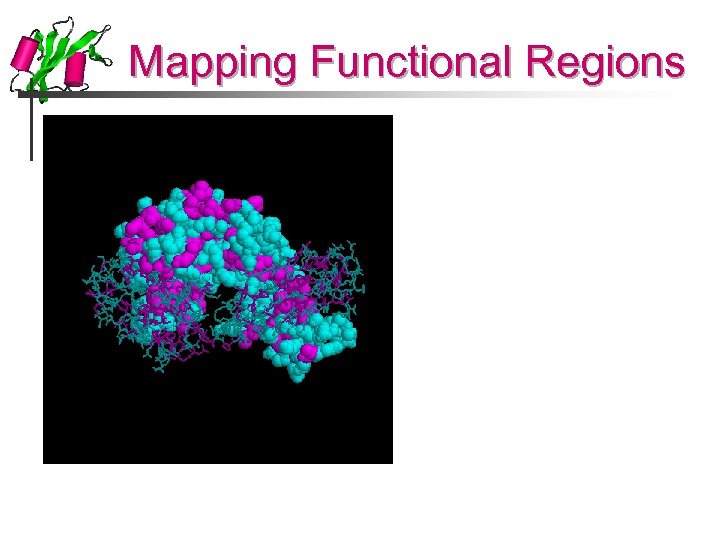 Mapping Functional Regions
Mapping Functional Regions
 Siblings and Cousins
Siblings and Cousins
 Multiple Sequence Alignment
Multiple Sequence Alignment
 MSA Methods
MSA Methods
 Alignment Checks
Alignment Checks
 Sequence Motifs & Patterns
Sequence Motifs & Patterns
 Aligned Sequence Families
Aligned Sequence Families
 Protein Domains
Protein Domains
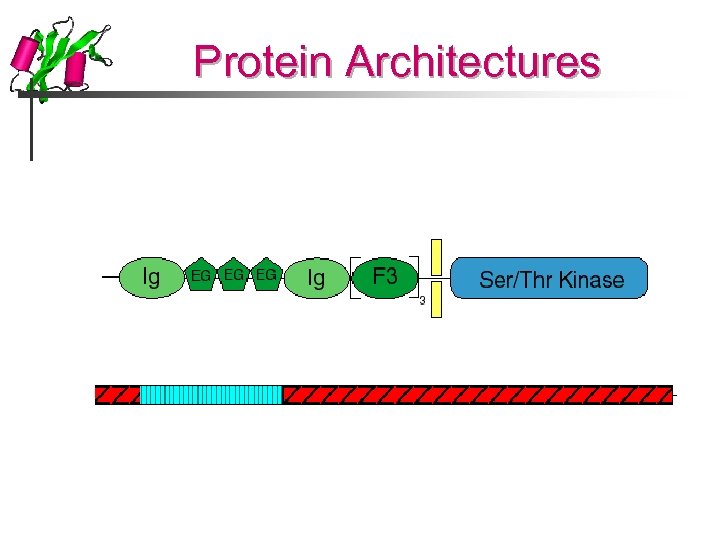 Protein Architectures
Protein Architectures
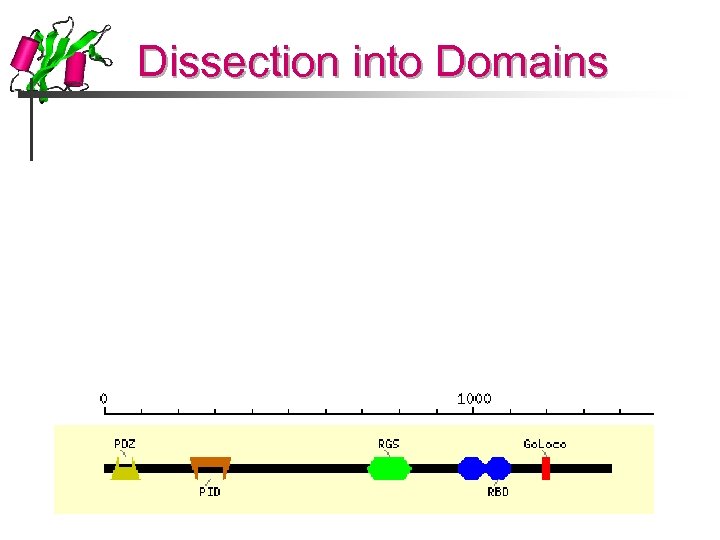 Dissection into Domains
Dissection into Domains
 Structural Homologues
Structural Homologues
 Small Proteins: Disulfide bonds
Small Proteins: Disulfide bonds
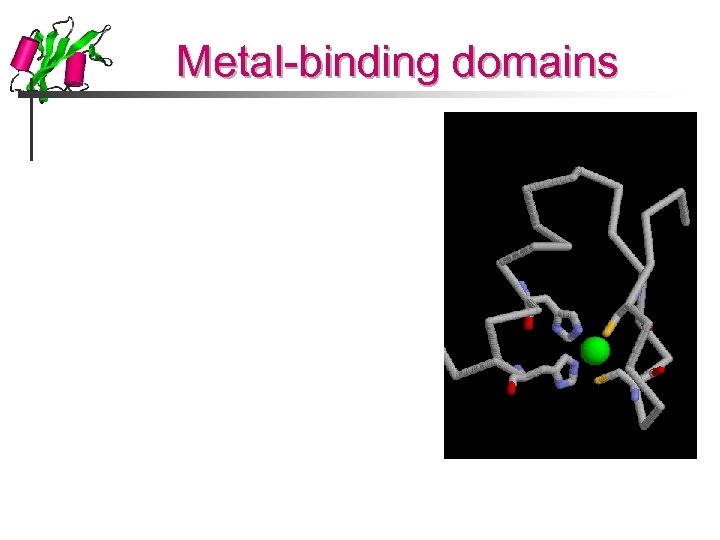 Metal-binding domains
Metal-binding domains
 Structure Prediction Methods
Structure Prediction Methods
 Tertiary Structure Prediction
Tertiary Structure Prediction
 One or many templates?
One or many templates?
 One or many templates?
One or many templates?
 Many templates
Many templates
 Query - Template Alignment
Query - Template Alignment
 QTA Checks
QTA Checks
 QTA Checks
QTA Checks
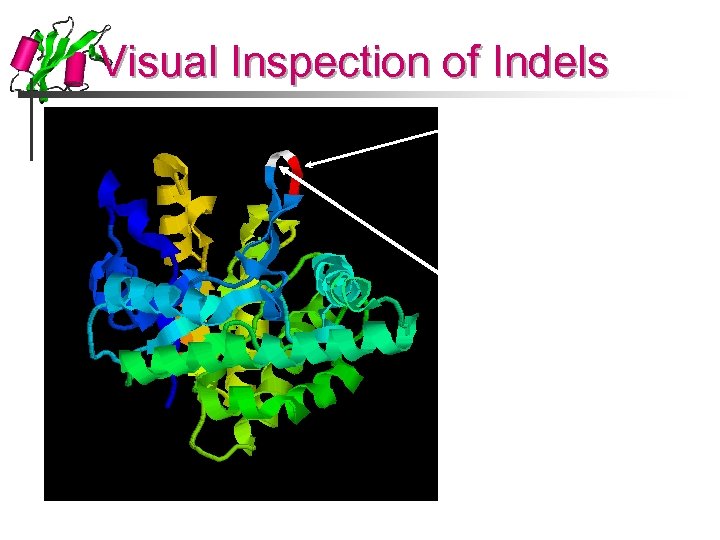 Visual Inspection of Indels
Visual Inspection of Indels
 Input for Model Building
Input for Model Building
 Methods Available
Methods Available
 Methods Available
Methods Available
 Methods Available
Methods Available
 Methods Available
Methods Available
 Methods Available
Methods Available
 Methods Available
Methods Available
 Automatic or Manual Mode?
Automatic or Manual Mode?
 How good is the model?
How good is the model?
 Improving ill-defined regions
Improving ill-defined regions
 Molecular Modeling Protocol
Molecular Modeling Protocol
 MM Protocol – Input Files
MM Protocol – Input Files
 Ex 1. High Homology Case
Ex 1. High Homology Case
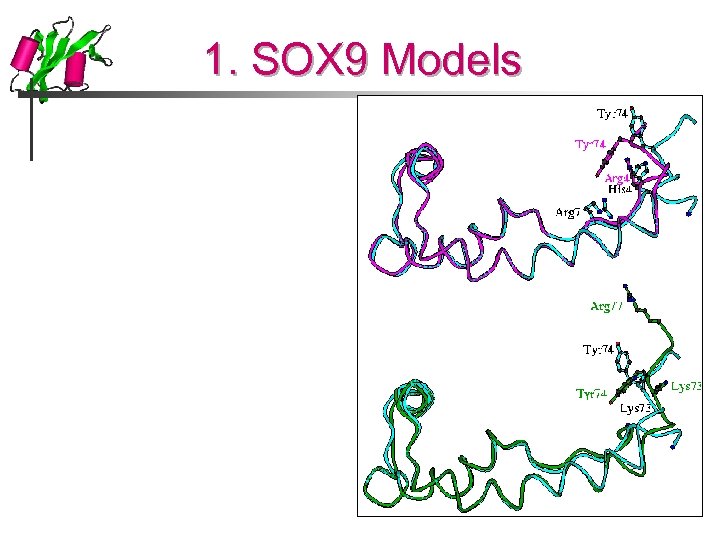 1. SOX 9 Models
1. SOX 9 Models
 Ex 2. Low Homology Situation
Ex 2. Low Homology Situation
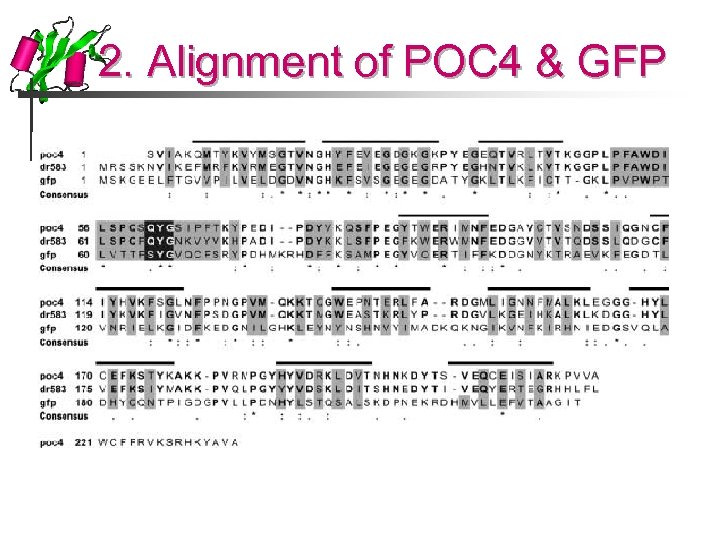 2. Alignment of POC 4 & GFP
2. Alignment of POC 4 & GFP
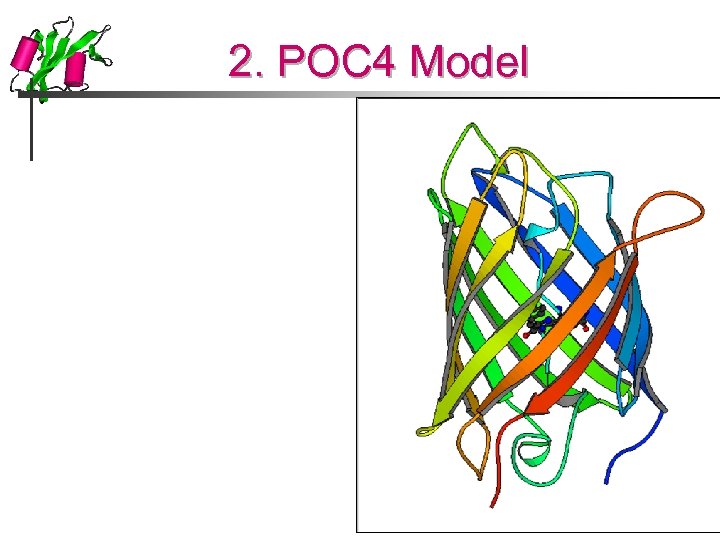 2. POC 4 Model
2. POC 4 Model
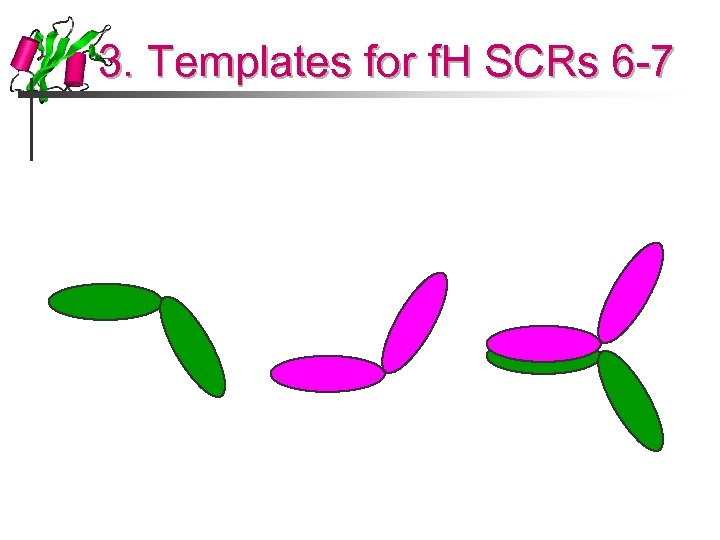 3. Templates for f. H SCRs 6 -7
3. Templates for f. H SCRs 6 -7
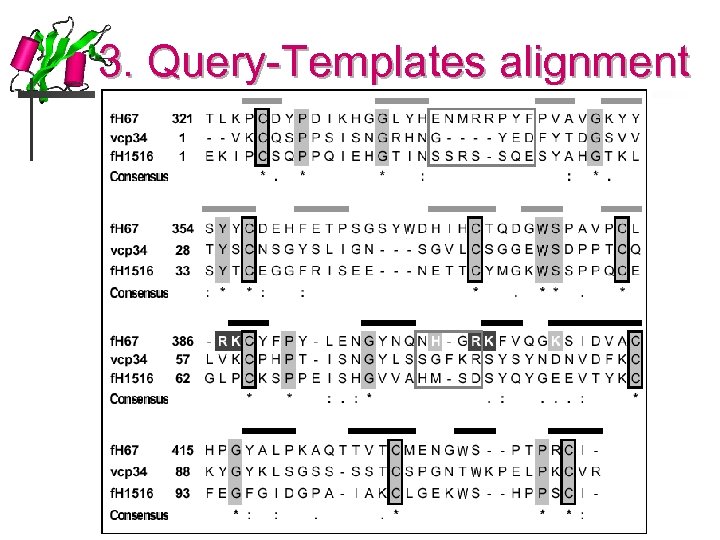 3. Query-Templates alignment
3. Query-Templates alignment
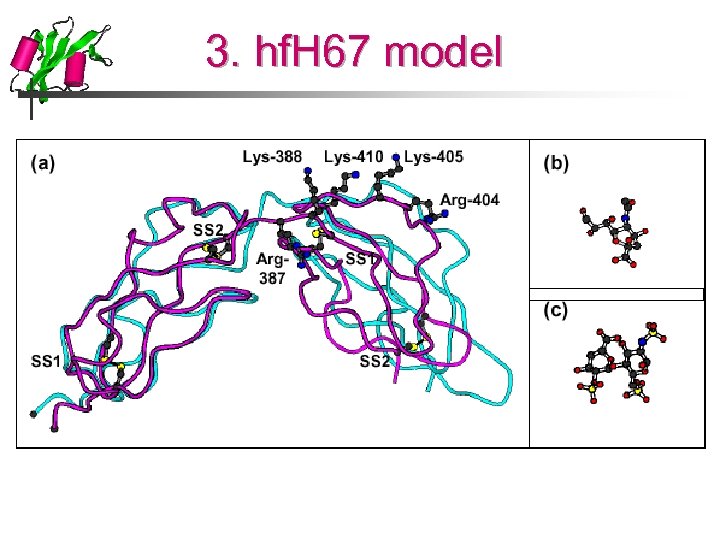 3. hf. H 67 model
3. hf. H 67 model
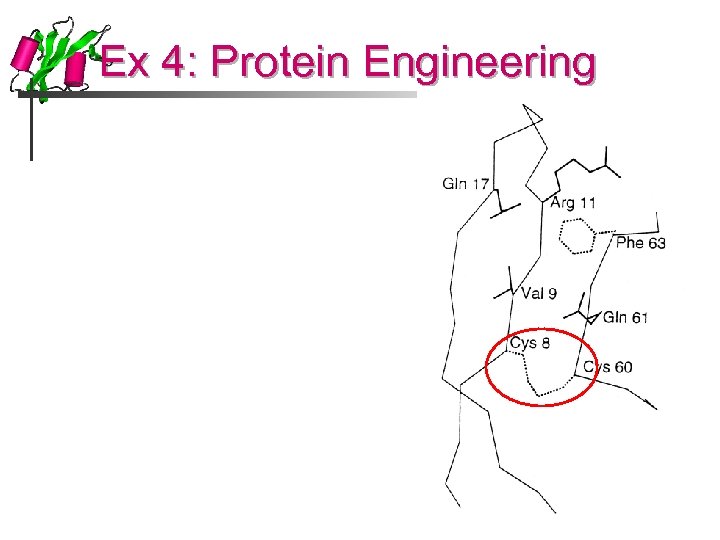 Ex 4: Protein Engineering
Ex 4: Protein Engineering
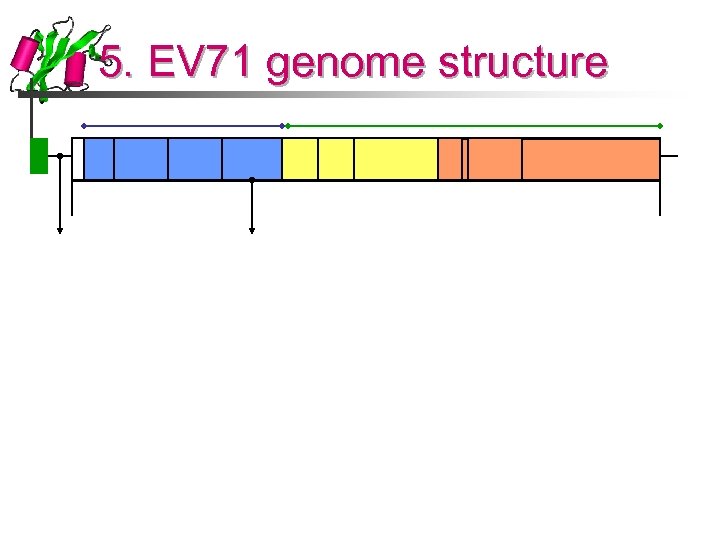 5. EV 71 genome structure
5. EV 71 genome structure
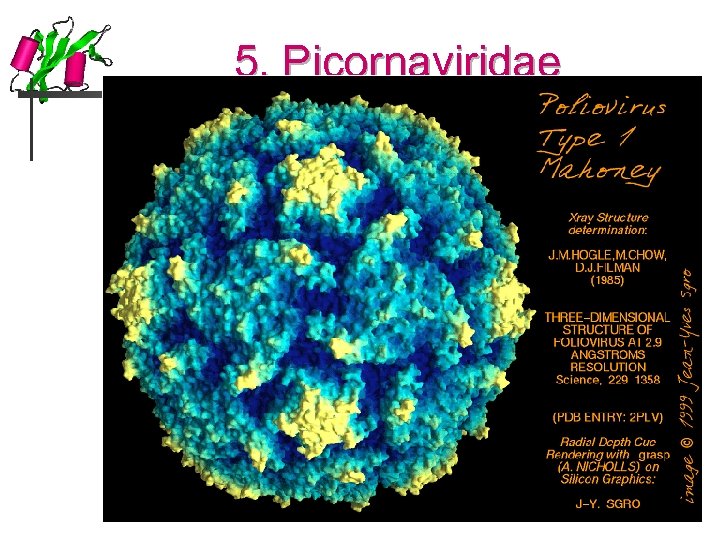 5. Picornaviridae
5. Picornaviridae
 5. Template hunt
5. Template hunt
 5. Fixing the VP 1 Alignment
5. Fixing the VP 1 Alignment
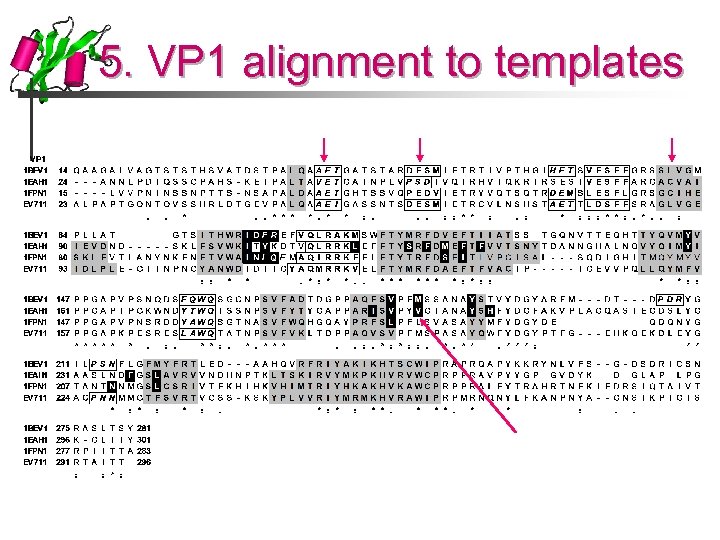 5. VP 1 alignment to templates
5. VP 1 alignment to templates
 5. Model building steps
5. Model building steps
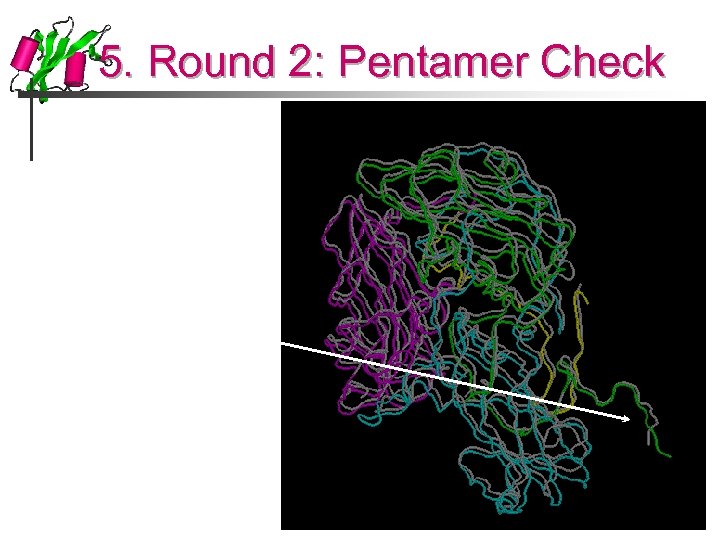 5. Round 2: Pentamer Check
5. Round 2: Pentamer Check
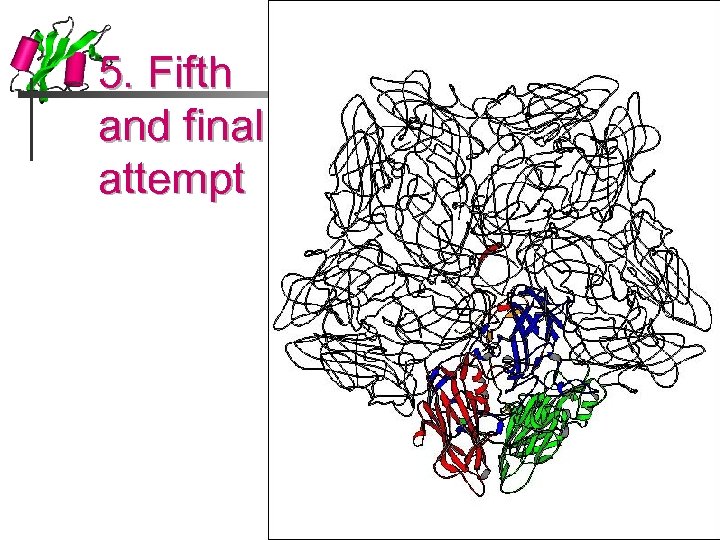 5. Fifth and final attempt
5. Fifth and final attempt
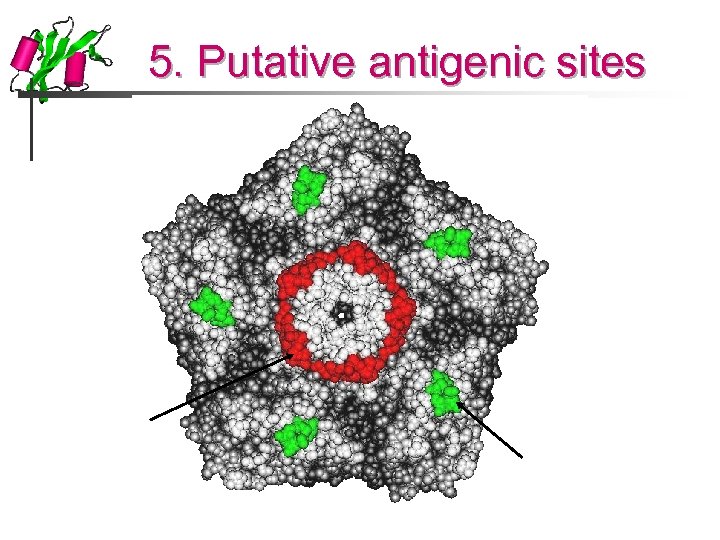 5. Putative antigenic sites
5. Putative antigenic sites