dd61bca530c70be0fe9b82d1eceb359d.ppt
- Количество слайдов: 96

Vorlesung Grundlagen der Bioinformatik http: //gobics. de/lectures/ss 07/grundlagen
Sequence alignment in molecular data analysis: Information from a Single Sequence Alone
Sequence alignment in molecular data analysis: Information from a Single Sequence Alone Multi-Organism High Quality Sequences (M. Brudno)
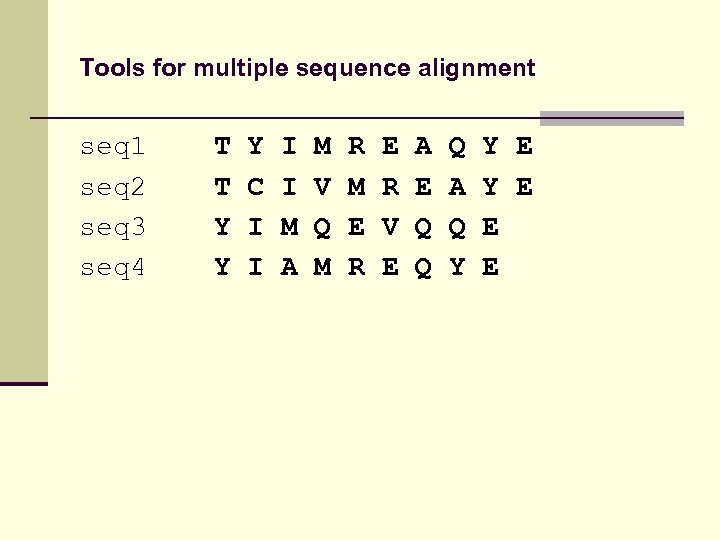
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C I I M A M V Q M R M E R V E A E Q Q Q A Q Y Y E E E
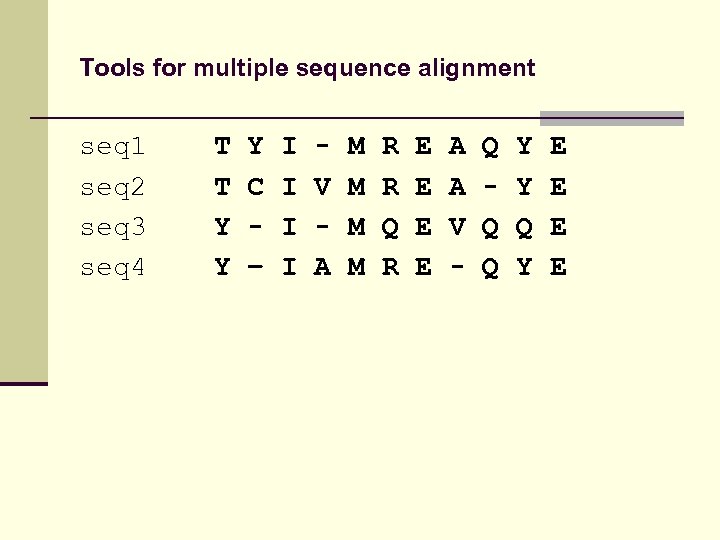
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E
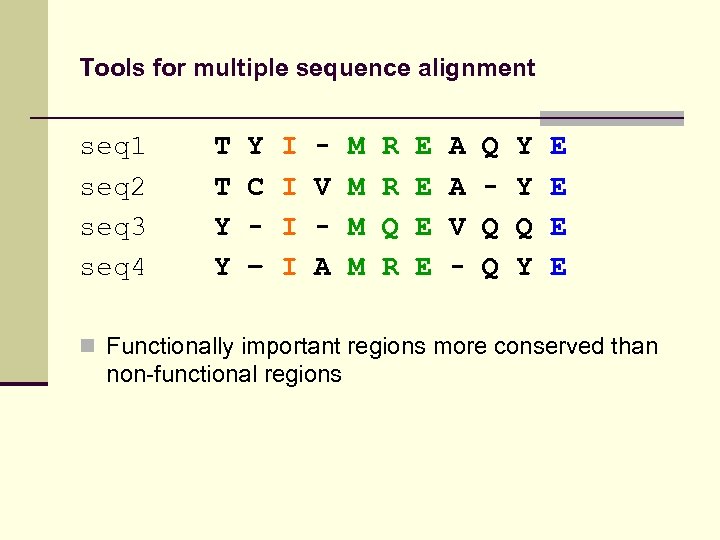
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E n Functionally important regions more conserved than non-functional regions
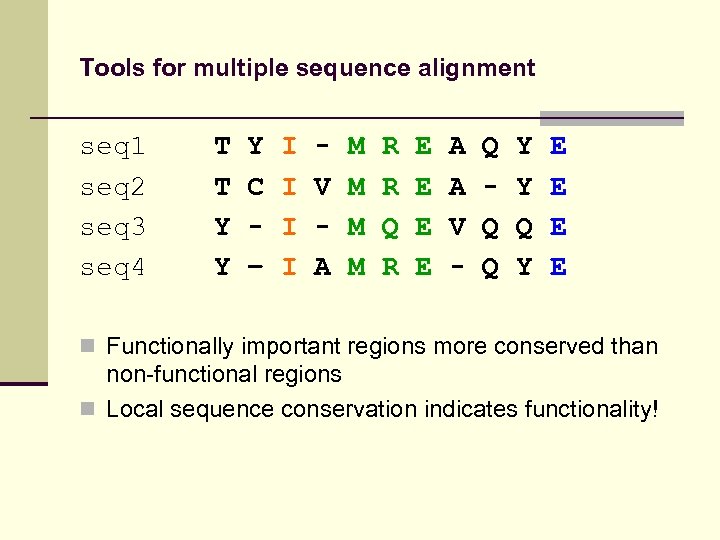
Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y Y C – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E n Functionally important regions more conserved than non-functional regions n Local sequence conservation indicates functionality!

Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y C Y – I I V A M M R R Q R E E A A V - Q Q Q Y Y Q Y E E Astronomical Number of possible alignments!

Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y C Y – I I V A M M M R M Q R E E A E V - Q A Q Q Y Y Q Y E E Astronomical Number of possible alignments!

Tools for multiple sequence alignment seq 1 seq 2 seq 3 seq 4 T T Y Y C Y – I I V A M M R R Q R E E A A V - Q Q Q Which one is the best ? ? ? Y Y Q Y E E

Tools for multiple sequence alignment Questions in development of alignment programs: (1) What is a good alignment? → objective function (`score’) (2) How to find a good alignment? → optimization algorithm First question far more important !

Tools for multiple sequence alignment Most important scoring scheme for multiple alignment: Sum-of-pairs score for global alignment.

Divide-and-Conquer Alignment (DCA) J. Stoye, A. Dress (Bielefeld) Approximate optimal global multiple alignment n n n Divide sequences into small sub-sequences Use MSA to calculate optimal alignment for subsequences Concatenate sub-alignments
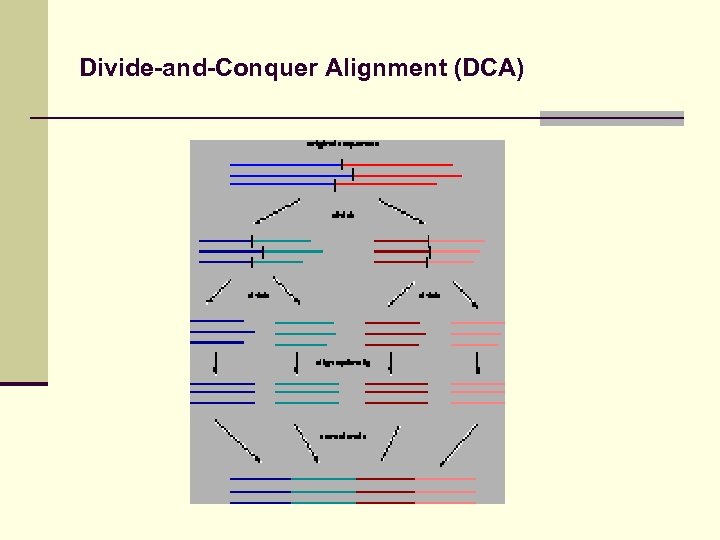
Divide-and-Conquer Alignment (DCA)
Divide-and-Conquer Alignment (DCA)

Tools for multiple sequence alignment Problems with traditional approach: n Results depend on gap penalty n Heuristic guide tree determines alignment; alignment used for phylogeny reconstruction n Algorithm produces global alignments.

First step in sequence comparison: alignment n global alignment (Needleman and Wunsch, 1970; Clustal W) atctaatagttaatactcgtccaagtat atctgtattactaaacaactggtgctacta

First step in sequence comparison: alignment n global alignment (Needleman and Wunsch, 1970; Clustal W) atc--taatagttaat--actcgtccaagtat ||| ||| || | | || atctgtattact-aaacaactggtgctacta-

First step in sequence comparison: alignment n global alignment (Needleman and Wunsch, 1970; Clustal W) atc--taatagttaat--actcgtccaagtat ||| ||| || | | || atctgtattact-aaacaactggtgctacta- n local alignment (Smith and Waterman, 1983) atctaatagttaatactcgtccaagtat gcgtgtattactaaacggttcaatctaacat

First step in sequence comparison: alignment n global alignment (Needleman and Wunsch, 1970; Clustal W) atc--taatagttaat--actcgtccaagtat ||| ||| || | | || atctgtattact-aaacaactggtgctacta- n local alignment (Smith and Waterman, 1983) atctaatagttaatactcgtccaagtat gcgtgtattactaaacggttcaatctaacat

First step in sequence comparison: alignment n global alignment (Needleman and Wunsch, 1970; Clustal W) atc--taatagttaat--actcgtccaagtat ||| ||| || | | || atctgtattact-aaacaactggtgctacta- n local alignment (Smith and Waterman, 1983) atc--taatagttaatactcgtccaagtat || || gcgtgtattact-aaacggttcaatctaacat
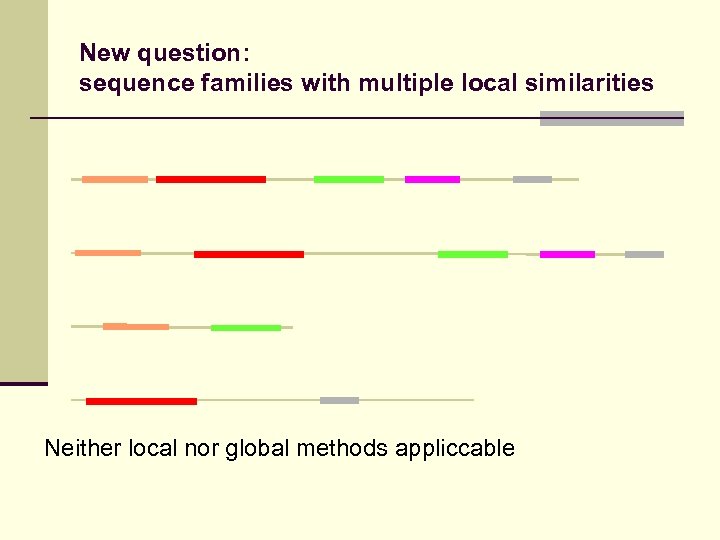

New question: sequence families with multiple local similarities Neither local nor global methods appliccable
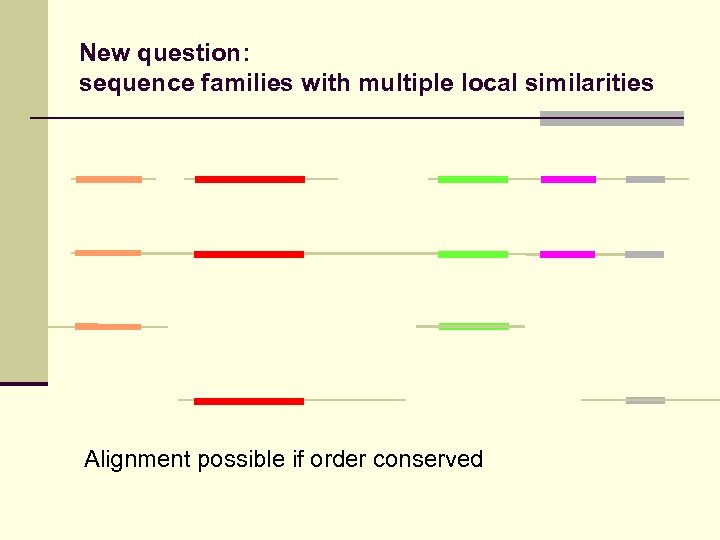
New question: sequence families with multiple local similarities Alignment possible if order conserved

The DIALIGN approach Morgenstern, Dress, Werner (1996), PNAS 93, 12098 -12103 n Combination of global and local methods n Assemble multiple alignment from gap-free local pair-wise alignments (, , fragments“)

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
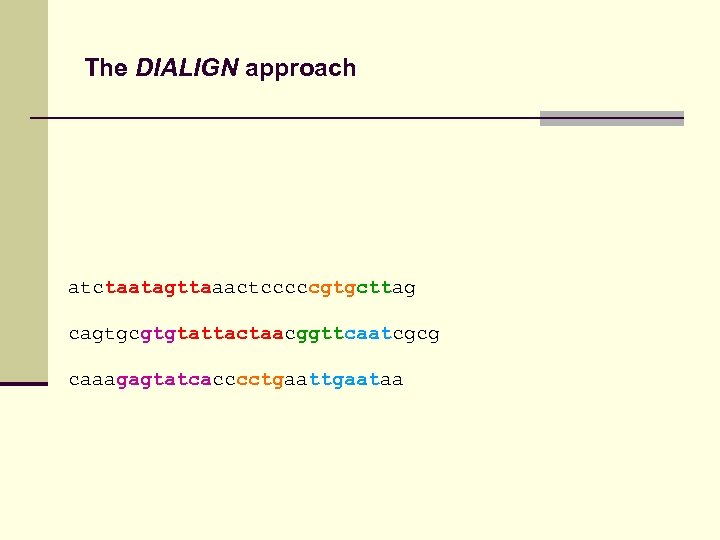
The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
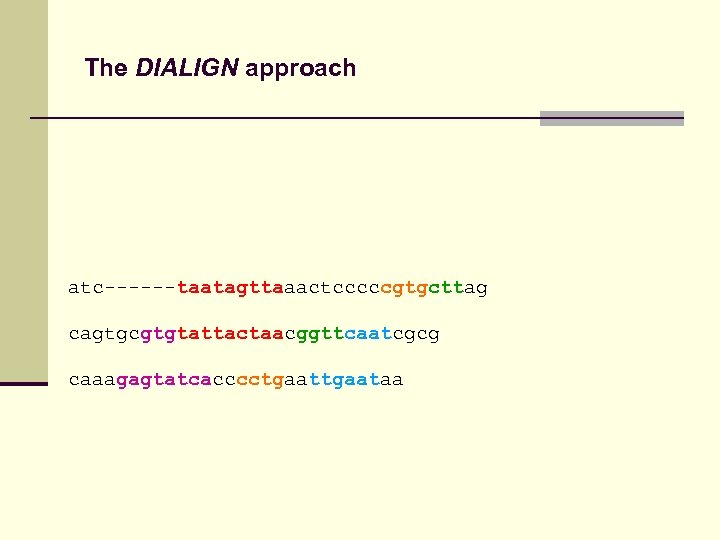
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
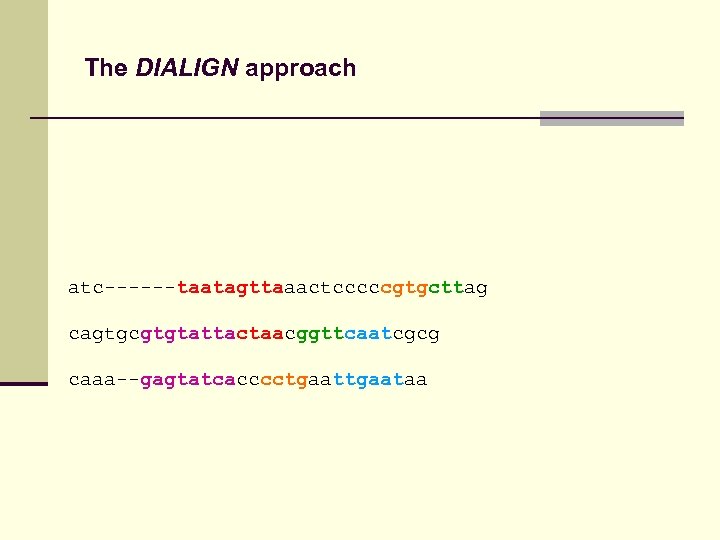
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa
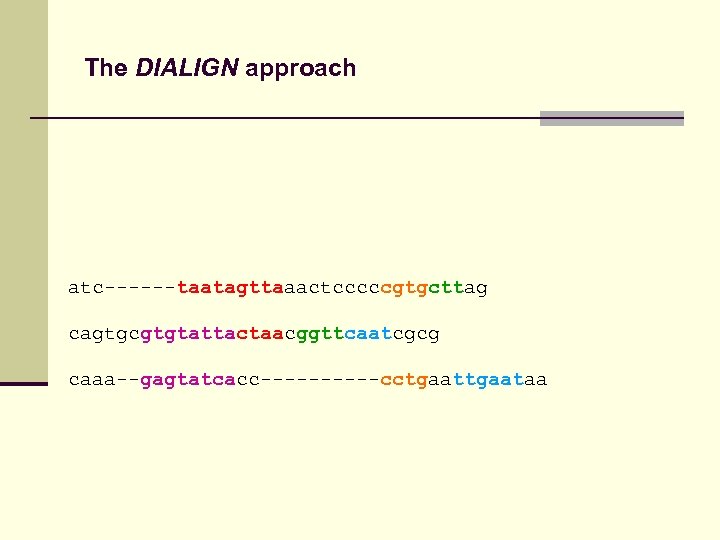
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacc-----cctgaataa
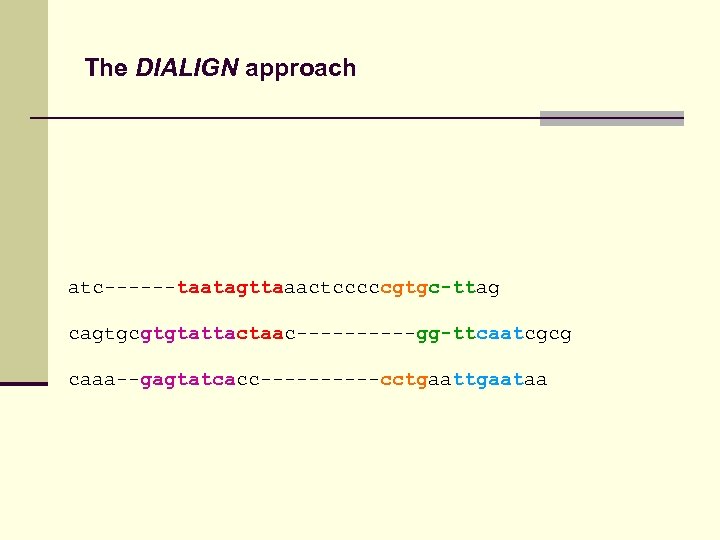
The DIALIGN approach atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa
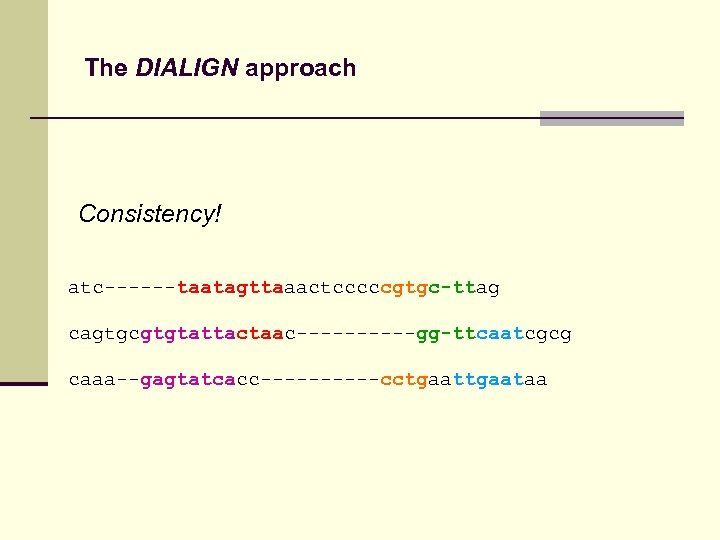

The DIALIGN approach Consistency! atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa

The DIALIGN approach atc------TAATAGTTAaactcccc. CGTGC-TTag cagtgc. GTGTATTACTAAc-----GG-TTCAATcgcg caaa--GAGTATCAcc-----CCTGaa. TTGAATaa
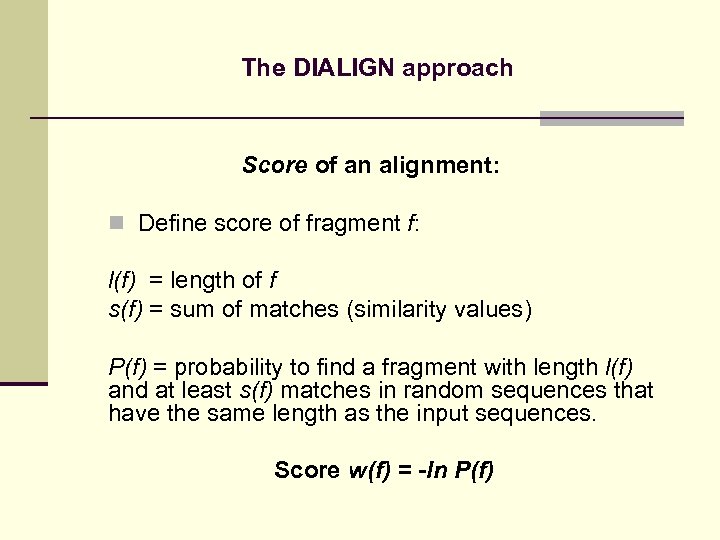

The DIALIGN approach Score of an alignment: n Define score of fragment f: l(f) = length of f s(f) = sum of matches (similarity values) P(f) = probability to find a fragment with length l(f) and at least s(f) matches in random sequences that have the same length as the input sequences. Score w(f) = -ln P(f)

The DIALIGN approach Score of an alignment: n Define score of fragment f: n Define score of alignment as sum of scores of involved fragments No gap penalty!

The DIALIGN approach Score of an alignment: Goal in fragment-based alignment approach: find Consistent collection of fragments with maximum sum of weight scores

The DIALIGN approach atctaatagttaaaccccctcgtgcttagagatccaaac cagtgcgtgtattactaacggttcaatcgcgcacatccgc Pair-wise alignment:
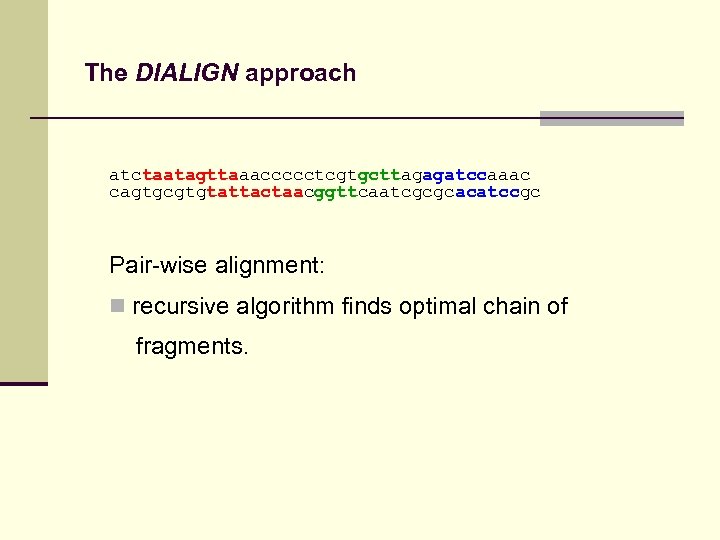

The DIALIGN approach atctaatagttaaaccccctcgtgcttagagatccaaac cagtgcgtgtattactaacggttcaatcgcgcacatccgc Pair-wise alignment: n recursive algorithm finds optimal chain of fragments.
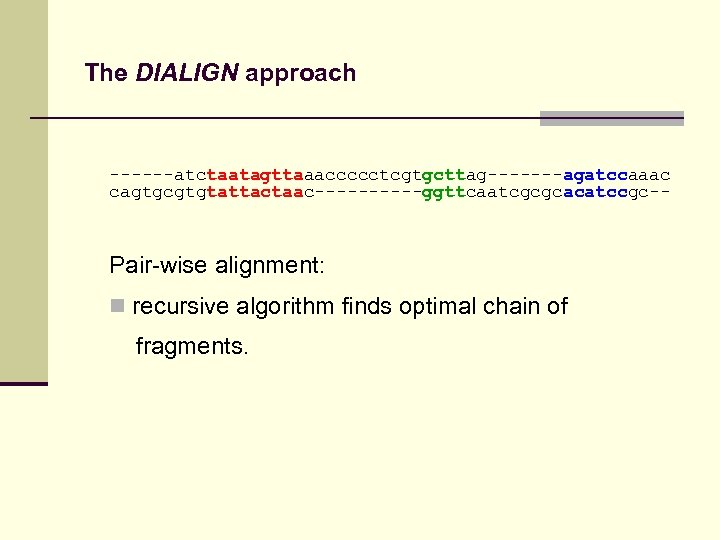
The DIALIGN approach ------atctaatagttaaaccccctcgtgcttag-------agatccaaac cagtgcgtgtattactaac-----ggttcaatcgcgcacatccgc-- Pair-wise alignment: n recursive algorithm finds optimal chain of fragments.
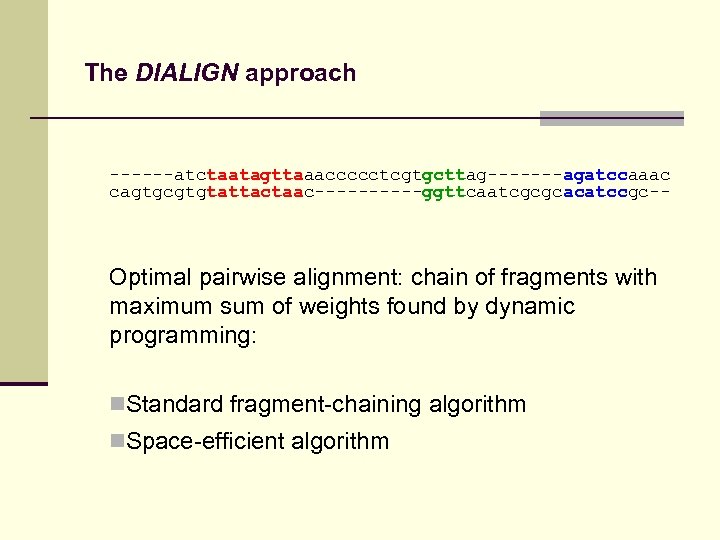

The DIALIGN approach ------atctaatagttaaaccccctcgtgcttag-------agatccaaac cagtgcgtgtattactaac-----ggttcaatcgcgcacatccgc-- Optimal pairwise alignment: chain of fragments with maximum sum of weights found by dynamic programming: n. Standard fragment-chaining algorithm n. Space-efficient algorithm
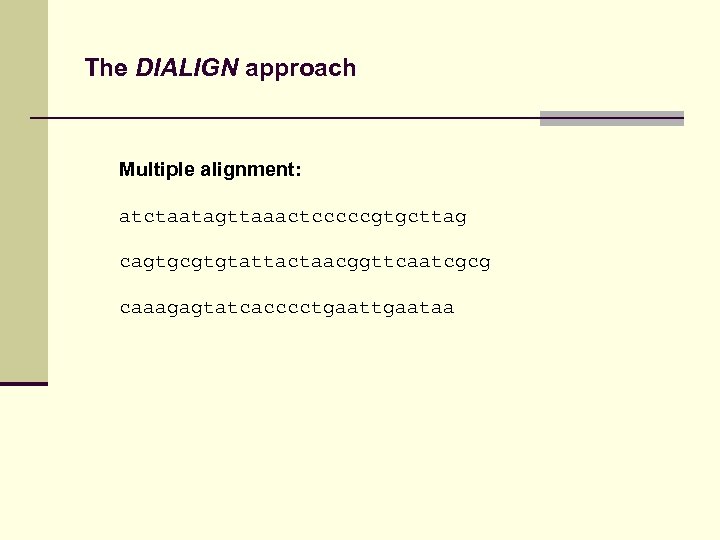

The DIALIGN approach Multiple alignment: atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
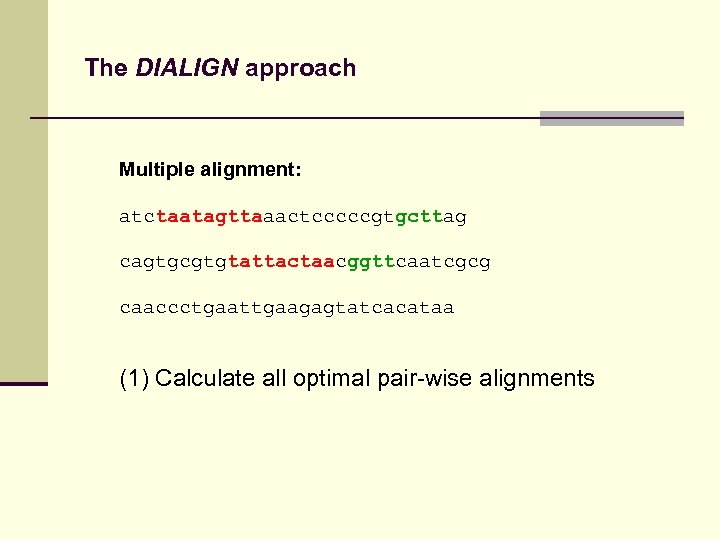

The DIALIGN approach Multiple alignment: atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaccctgaattgaagagtatcacataa (1) Calculate all optimal pair-wise alignments
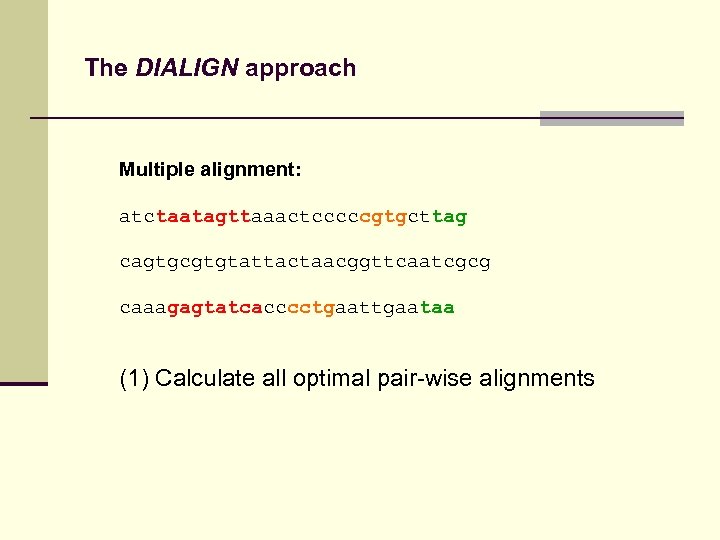

The DIALIGN approach Multiple alignment: atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa (1) Calculate all optimal pair-wise alignments
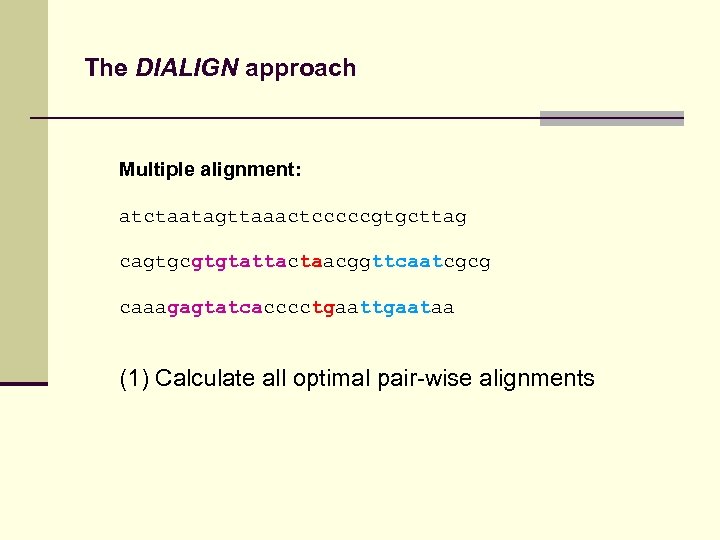

The DIALIGN approach Multiple alignment: atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa (1) Calculate all optimal pair-wise alignments

The DIALIGN approach Fragments from optimal pair-wise alignments might be inconsistent

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
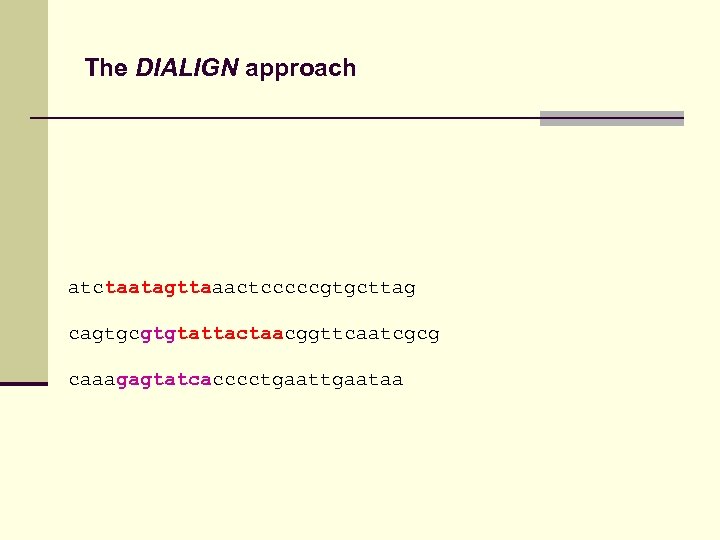

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
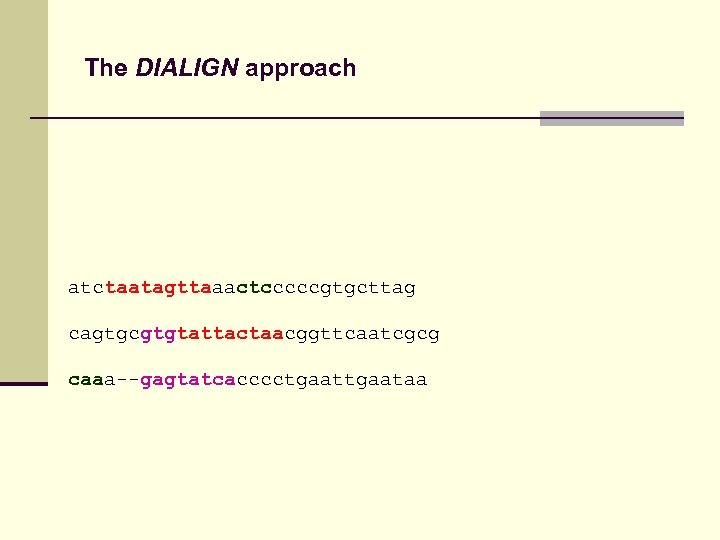

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa

The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa
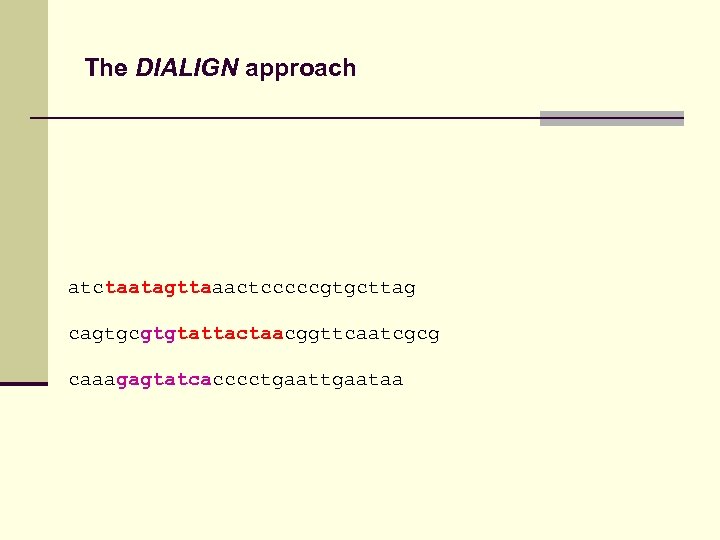

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
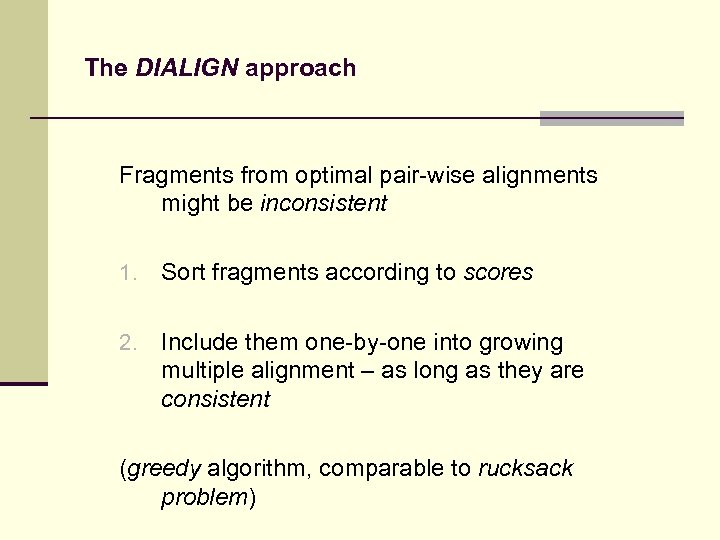

The DIALIGN approach Fragments from optimal pair-wise alignments might be inconsistent 1. Sort fragments according to scores 2. Include them one-by-one into growing multiple alignment – as long as they are consistent (greedy algorithm, comparable to rucksack problem)

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
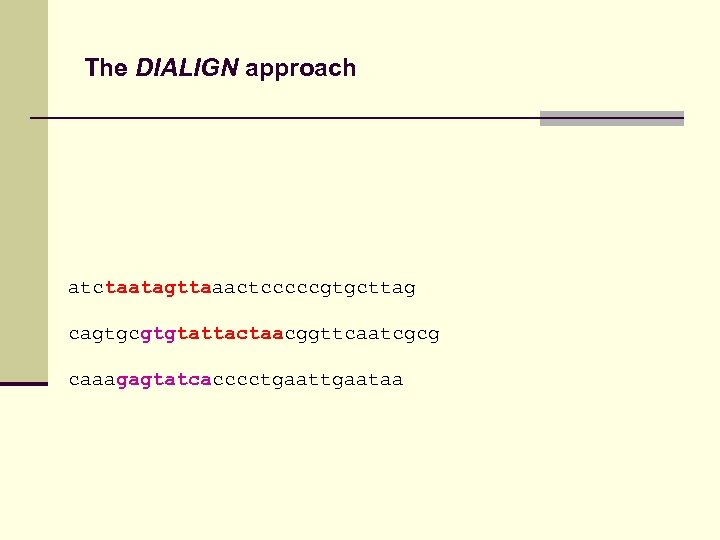

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
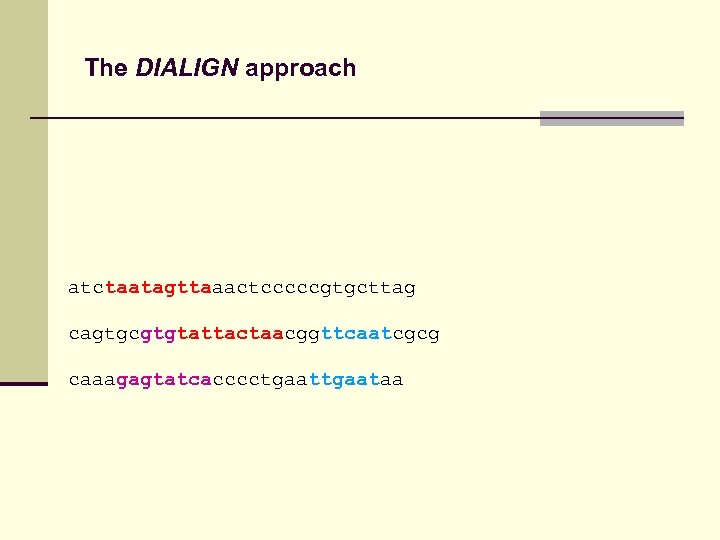

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
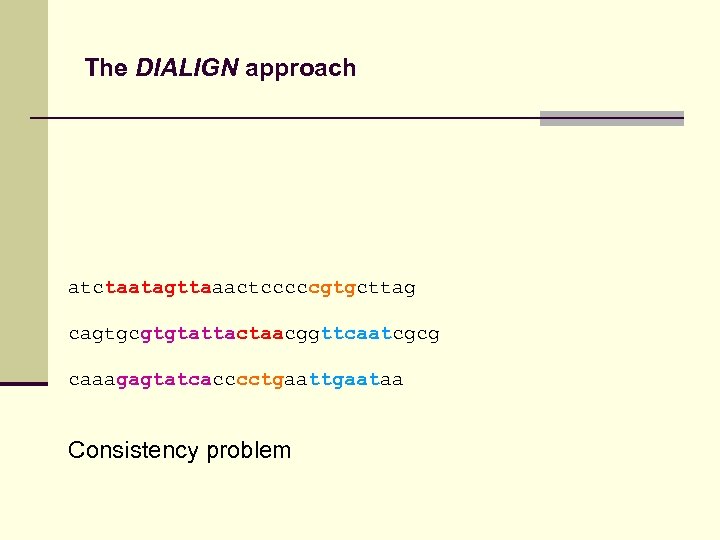

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Consistency problem
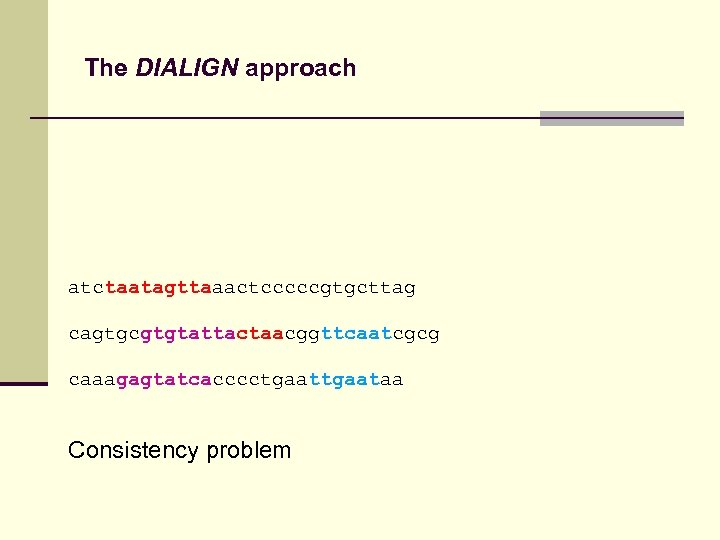

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Consistency problem
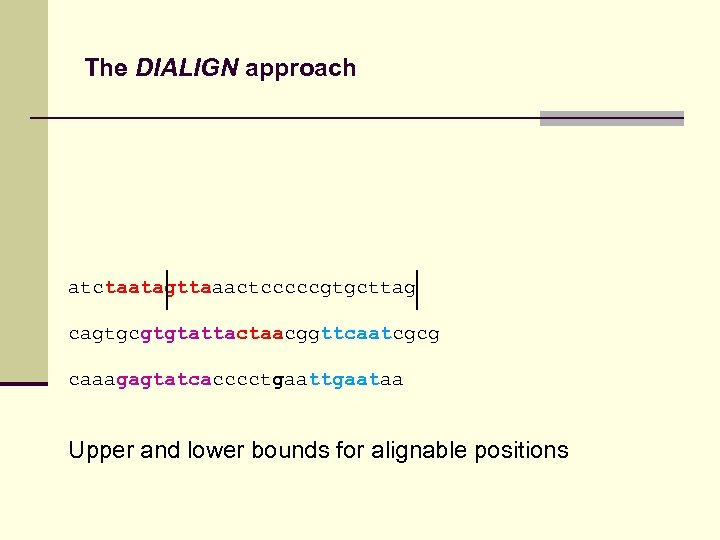

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Upper and lower bounds for alignable positions
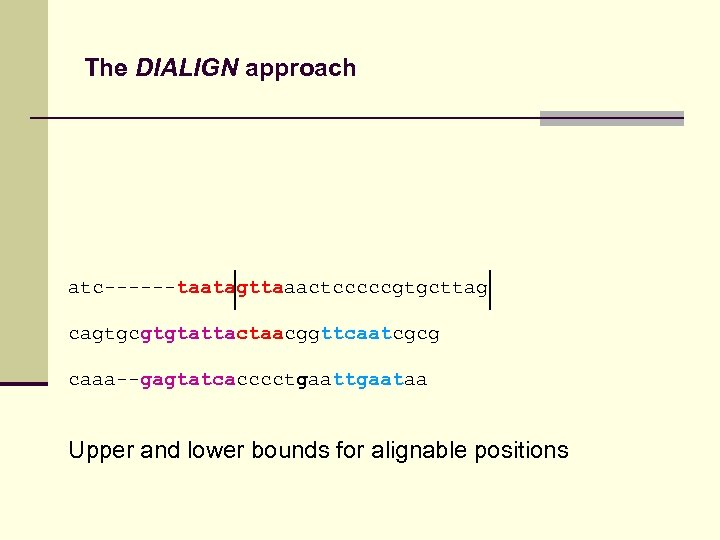

The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa Upper and lower bounds for alignable positions
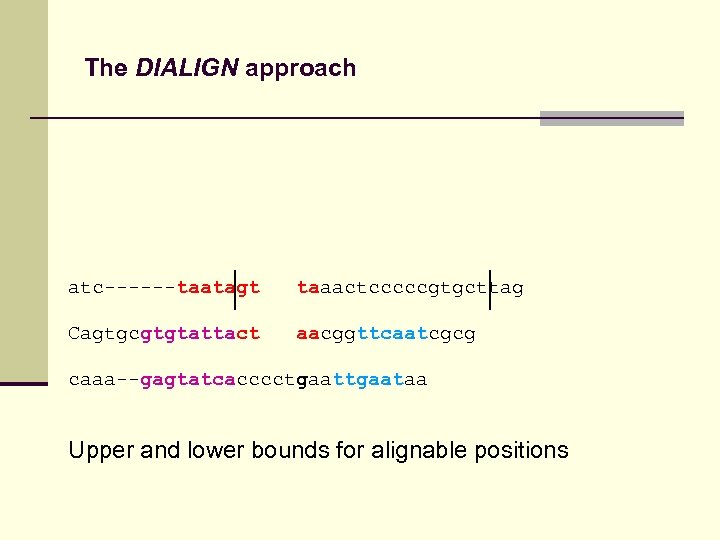
The DIALIGN approach atc------taatagt taaactcccccgtgcttag Cagtgcgtgtattact aacggttcaatcgcg caaa--gagtatcacccctgaataa Upper and lower bounds for alignable positions
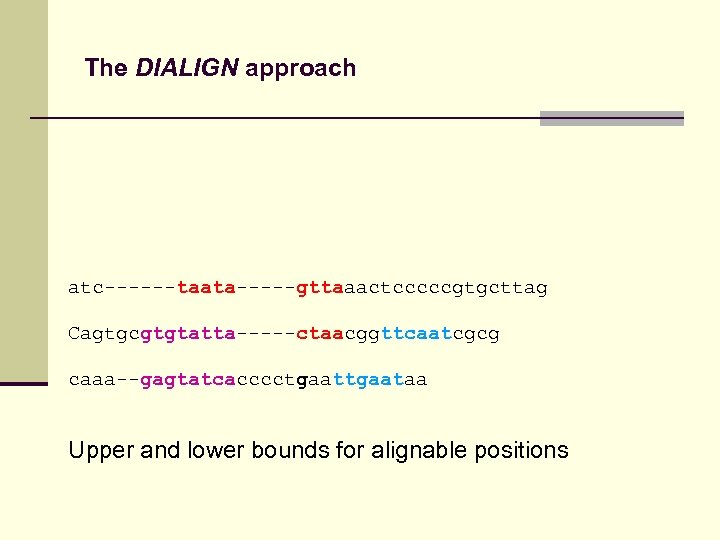

The DIALIGN approach atc------taata-----gttaaactcccccgtgcttag Cagtgcgtgtatta-----ctaacggttcaatcgcg caaa--gagtatcacccctgaataa Upper and lower bounds for alignable positions
![The DIALIGN approach site x = [i, p] (sequence i, position p) atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg The DIALIGN approach site x = [i, p] (sequence i, position p) atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg](https://present5.com/presentation/dd61bca530c70be0fe9b82d1eceb359d/image-69.jpg)
The DIALIGN approach site x = [i, p] (sequence i, position p) atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Upper and lower bounds for alignable positions
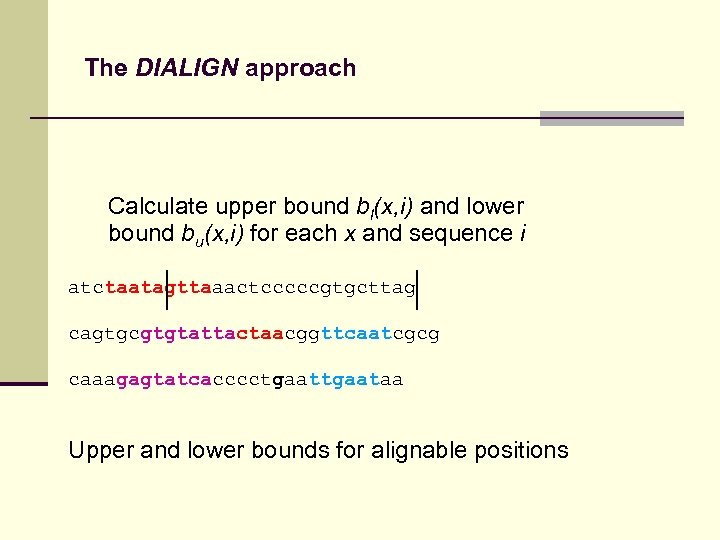

The DIALIGN approach Calculate upper bound bl(x, i) and lower bound bu(x, i) for each x and sequence i atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Upper and lower bounds for alignable positions
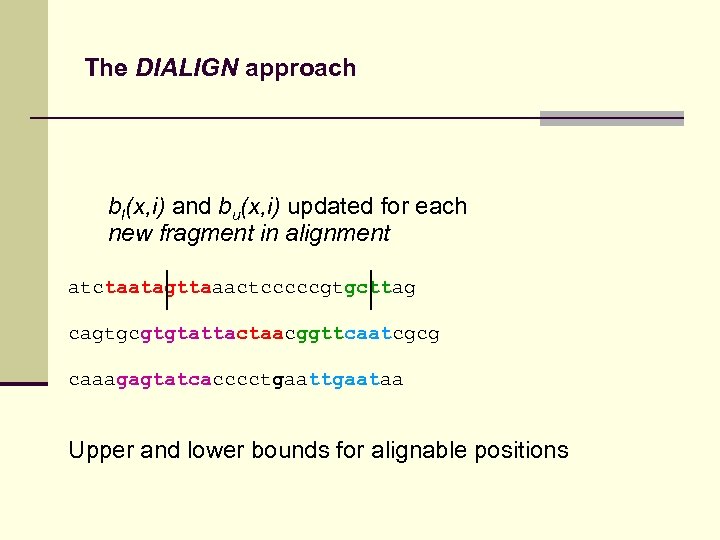

The DIALIGN approach bl(x, i) and bu(x, i) updated for each new fragment in alignment atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Upper and lower bounds for alignable positions

The DIALIGN approach Consistency bounds are to be updated for each new fragment that is included in to the growing Alignment Efficient algorithm (Abdeddaim and Morgenstern, 2002)

The DIALIGN approach Advantages of segment-based approach: n Program can produce global and local alignments! n Sequence families alignable that cannot be aligned with standard methods
![Program input Program usage: > dialign 2 -2 [options] <input_file> = multi-sequence file in Program input Program usage: > dialign 2 -2 [options] <input_file> = multi-sequence file in](https://present5.com/presentation/dd61bca530c70be0fe9b82d1eceb359d/image-74.jpg)
Program input Program usage: > dialign 2 -2 [options] <input_file> = multi-sequence file in FASTA-format

Program output DIALIGN 2. 2. 1 ******* Program code written by Burkhard Morgenstern and Said Abdeddaim e-mail contact: bmorgen@gwdg. de Published research assisted by DIALIGN 2 should cite: Burkhard Morgenstern (1999). DIALIGN 2: improvement of the segment-to-segment approach to multiple sequence alignment. Bioinformatics 15, 211 - 218. For more information, please visit the DIALIGN home page at http: //bibiserv. techfak. uni-bielefeld. de/dialign/ program call: . /dialign 2 -2 -nt -anc s Aligned sequences: ========= length: ======= 1) dog_il 4 2) bla 3) blu 300 200 Average seq. length: 233. 3 Please note that only upper-case letters are considered to be aligned.

Program output Alignment (DIALIGN format): ============== dog_il 4 bla blu 1 1 1 cagg------GTTTGA atctgataca ttgc------ctga----------GC CAAGTGGGAA ttttgatatg agaa. GTGTGA aacaagctat cctatatt. GC TAAGTGGCAG 0000000000 000011 11111 dog_il 4 bla blu 25 17 51 ----- --ATGGCACT GGGGTGAATG AGGCAG CAGAATGATC ggtgtgaata catgggtttc cagtaccttc tgaggtccag agtacc---ccctggcttt ct. ATGTGCAC AGAATGGGAG GAAAGTGCCT GCTAGTGAGC 0000000000 00000 dog_il 4 bla blu 63 63 101 GTACTGCAGC CCTGAGCTTC CACTGGCCCA TGTTGGTATC CTTGTATTTT ---------- ---TTTCCCA TGTGCTCCAT GGTGGAATGG CAGGGACTCA GAGAGAATGG AGTATAGGGG TCAGGGCat- -----0000000000 0009999999888 88888 dog_il 4 bla blu 113 90 140 TCCGCCCCTT CCCAGCACca gcattatcct ---GGGATTG GAGAAGGGGG ACCACTCCTT CTCAGCACaa caaagcccaa gaa. GGTGTTG CGTTCTAGAC ---------- ---GGGGTGG CCTTAGGCTC 8888800 00000 0007777777

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
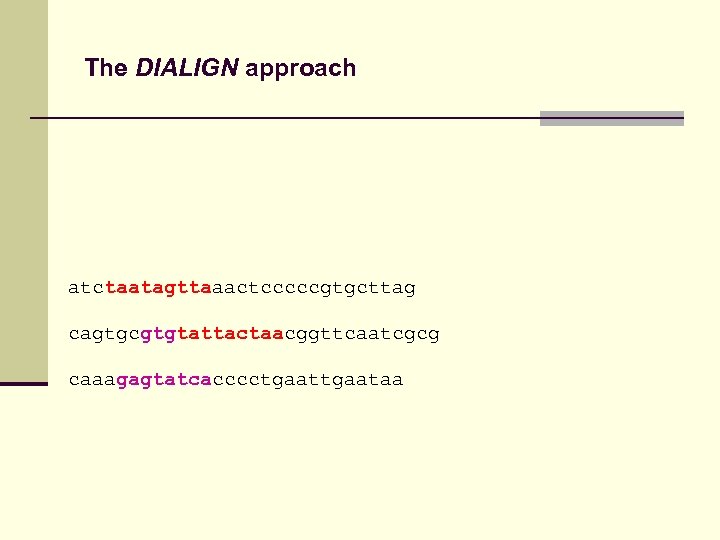

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
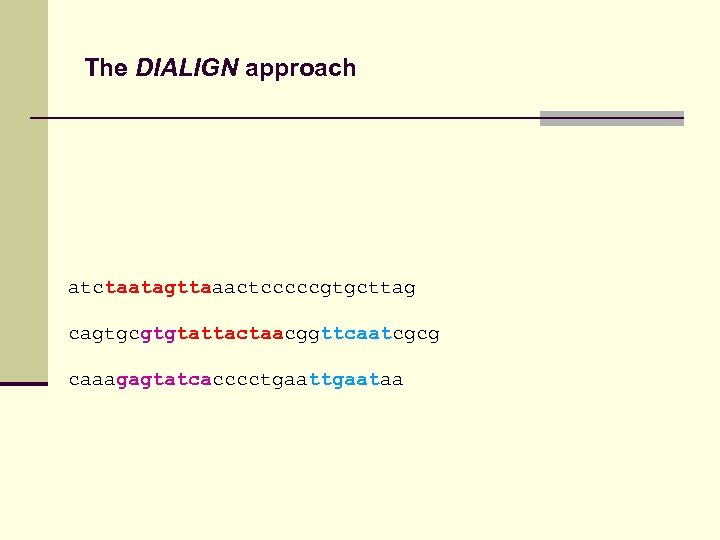

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
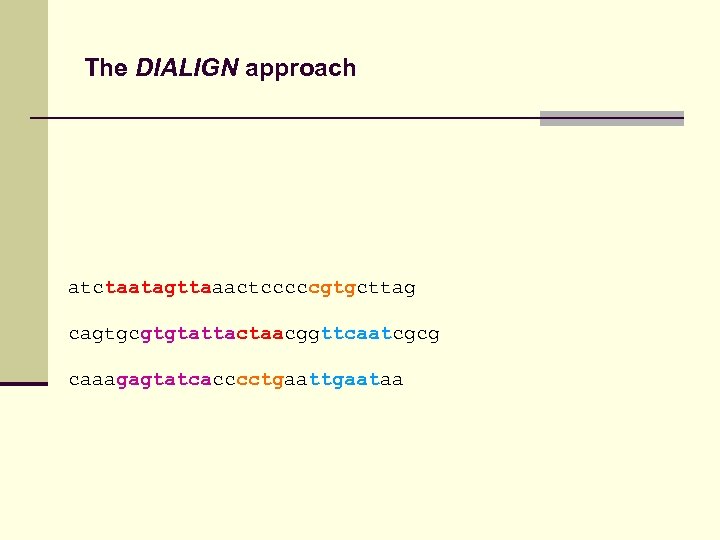

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
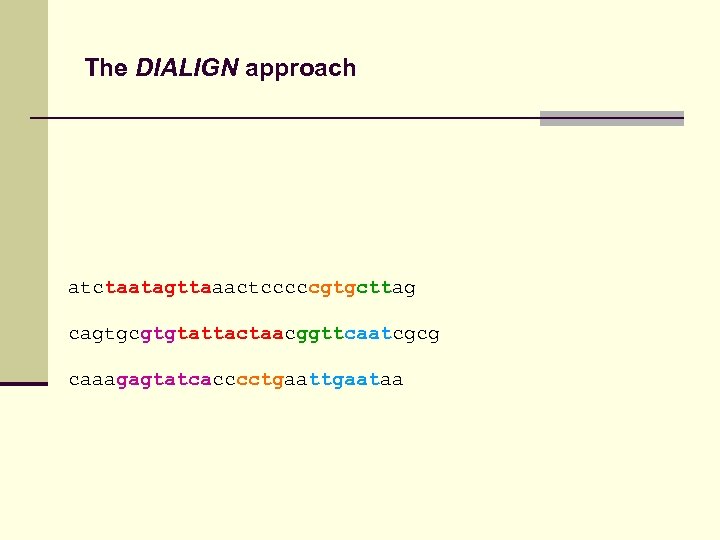
The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
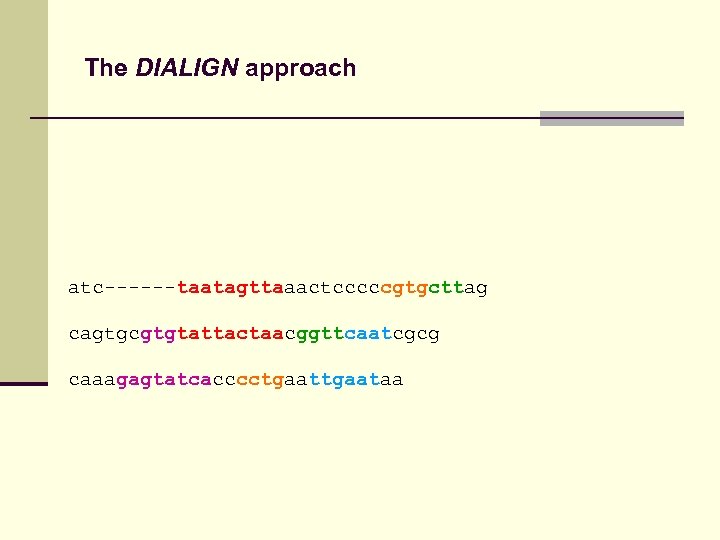
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
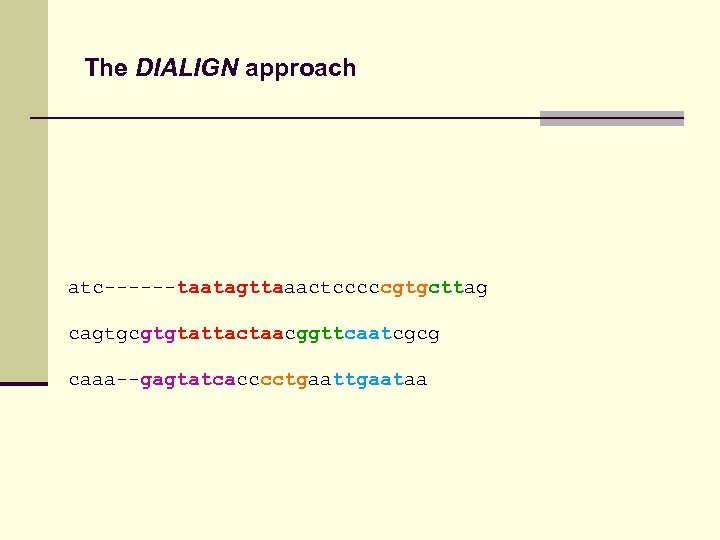
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa
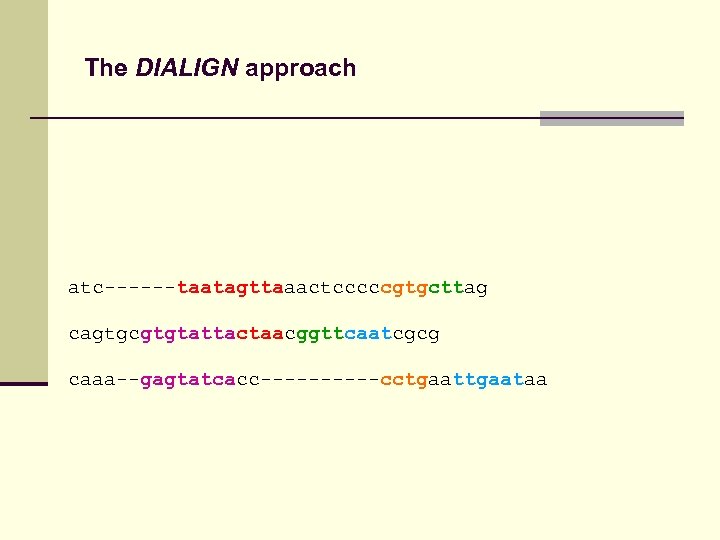
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacc-----cctgaataa
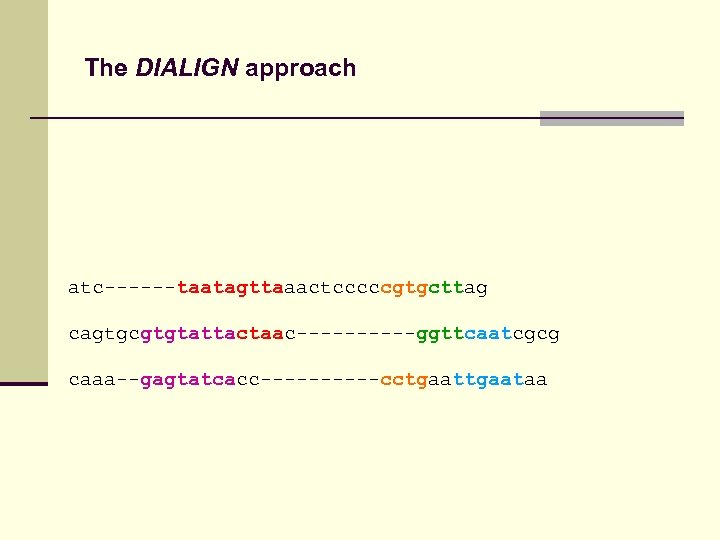
The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaac-----ggttcaatcgcg caaa--gagtatcacc-----cctgaataa
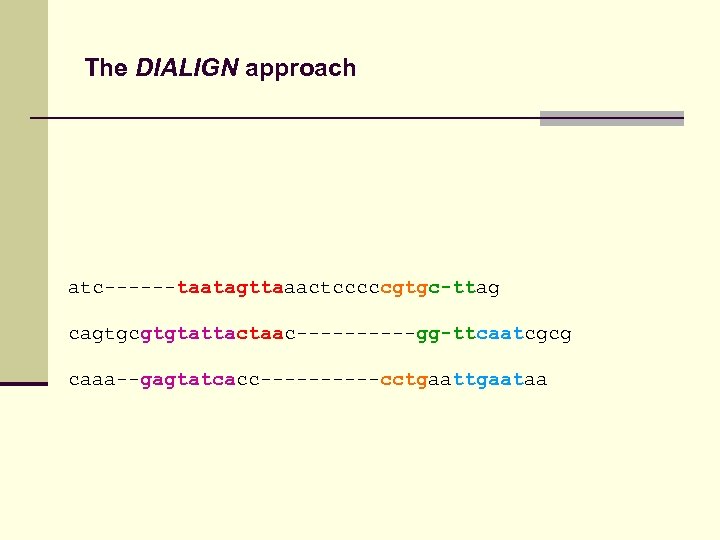
The DIALIGN approach atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa

The DIALIGN approach atc------TAATAGTTAaactcccc. CGTGC-TTag-----cagtgc. GTGTATTACTAAc-----GG-TTCAATcgcg caaa--GAGTATCAcc-----CCTGaa. TTGAATaa--
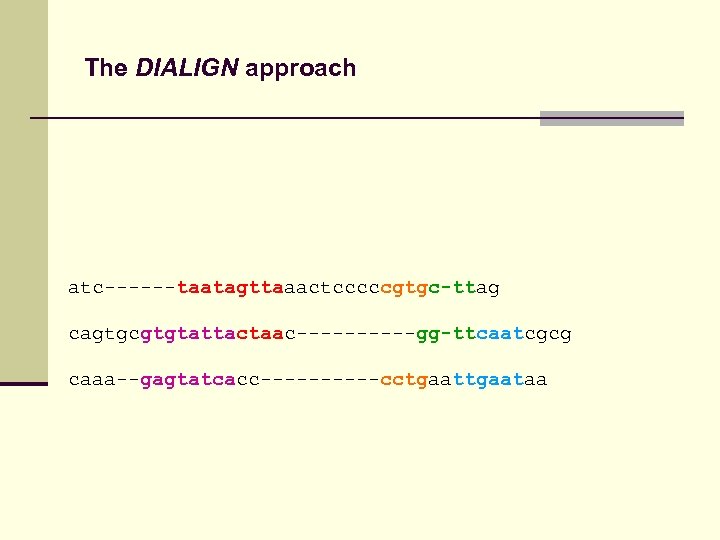
The DIALIGN approach atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa

Alignment of large genomic sequences Fragment-based alignment approach useful for alignment of genomic sequences. Possible applications: n Detection of regulatory elements n Identification of pathogenic microorganisms n Gene prediction
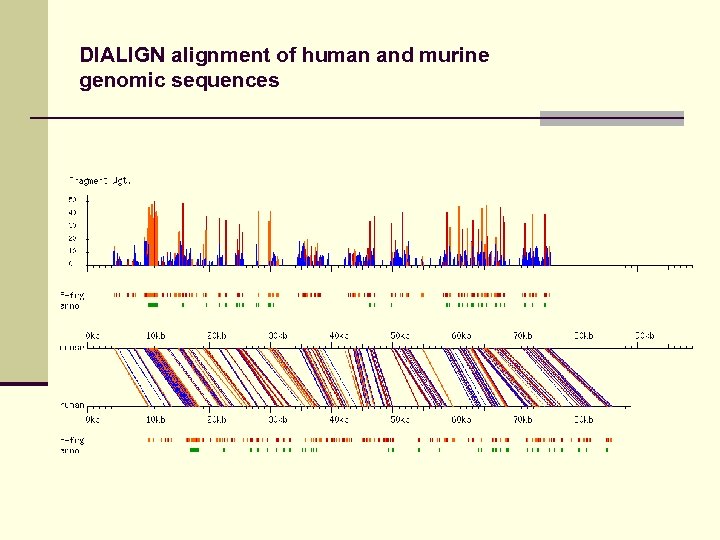
DIALIGN alignment of human and murine genomic sequences
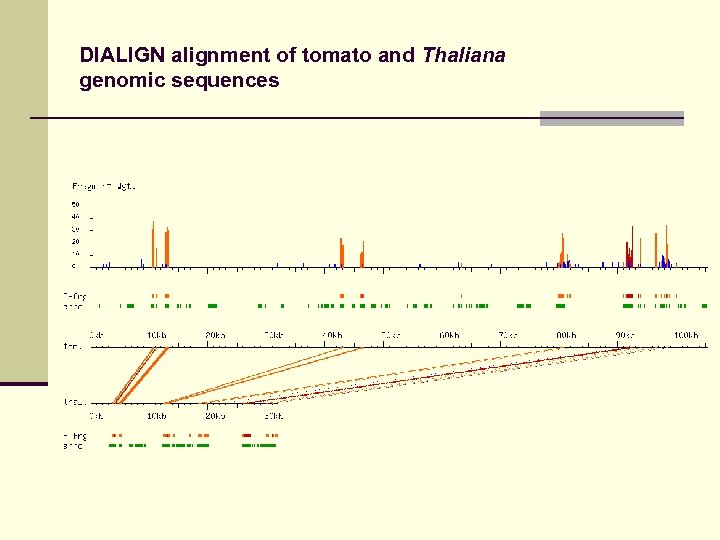
DIALIGN alignment of tomato and Thaliana genomic sequences
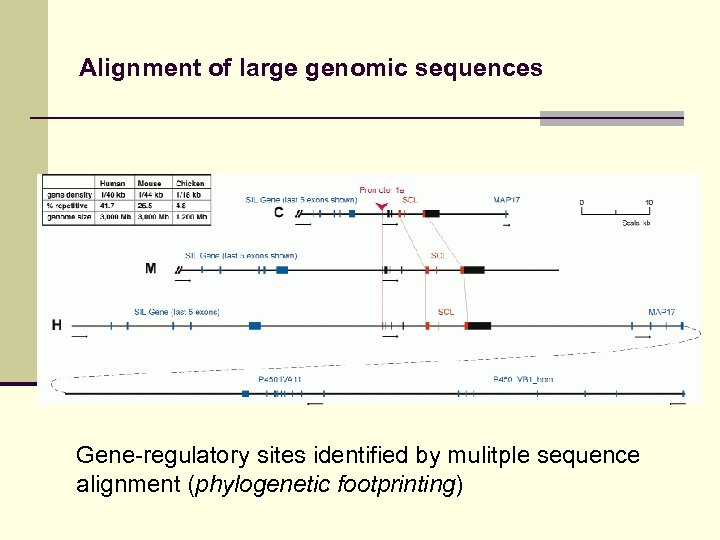
Alignment of large genomic sequences Gene-regulatory sites identified by mulitple sequence alignment (phylogenetic footprinting)
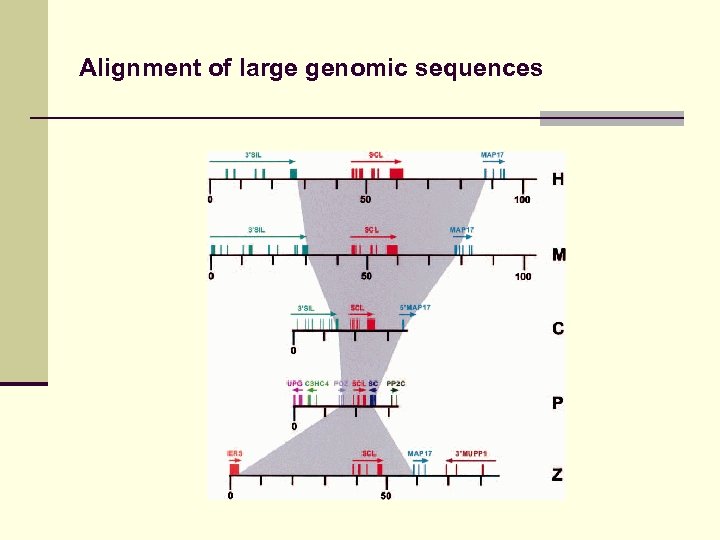
Alignment of large genomic sequences
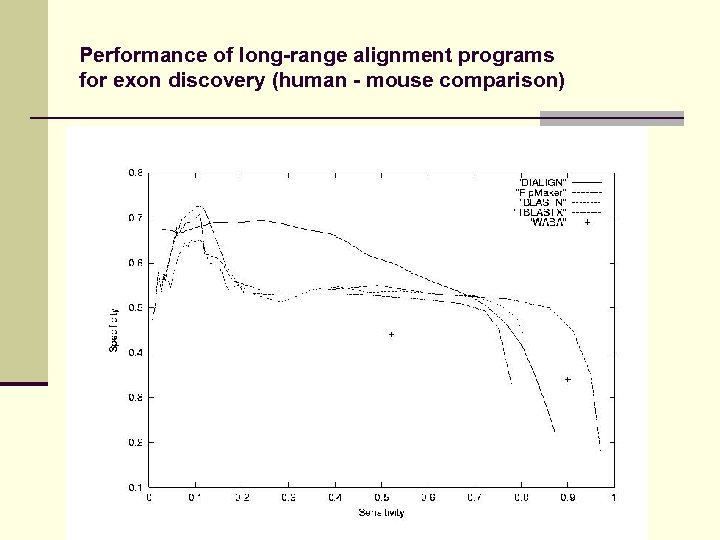
Performance of long-range alignment programs for exon discovery (human - mouse comparison)
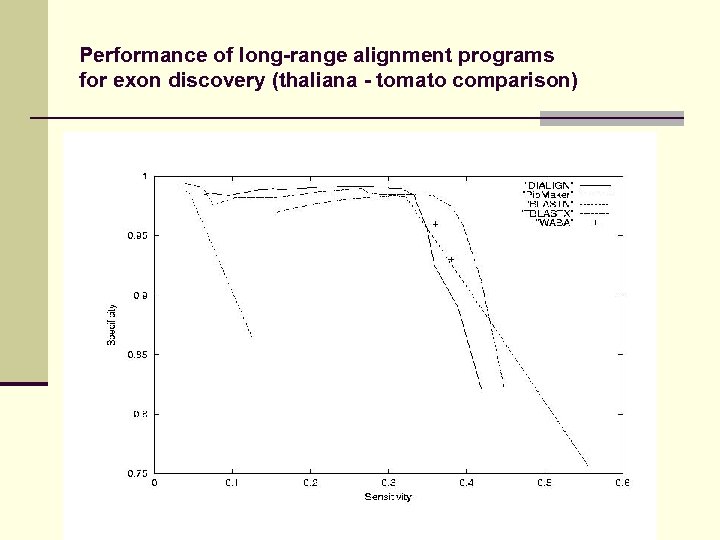
Performance of long-range alignment programs for exon discovery (thaliana - tomato comparison)
dd61bca530c70be0fe9b82d1eceb359d.ppt