a5350967754e5523760d498bbb6ba6f6.ppt
- Количество слайдов: 38
 Vernalization Gene Architecture as a Predictor of Growth Habit in Barley • Douglas Heckart • Primary Advisor: Dr Patrick Hayes • Secondary Advisor: Dr Thomas Chastain • Project Advisor (field): Ann Corey • Project Advisor (laboratory) Dr Peter Szucs
Vernalization Gene Architecture as a Predictor of Growth Habit in Barley • Douglas Heckart • Primary Advisor: Dr Patrick Hayes • Secondary Advisor: Dr Thomas Chastain • Project Advisor (field): Ann Corey • Project Advisor (laboratory) Dr Peter Szucs
 Overview • Vernalization background • Phenotype determination methods • Genotype determination methods • Relating genotype to phenotype
Overview • Vernalization background • Phenotype determination methods • Genotype determination methods • Relating genotype to phenotype
 Purpose • Validate candidate genes for winter and facultative growth habit in barley. • Important to OSU barley breeding program for development of fall sown malting barley with cold tolerance. • Determine if growth habit can be predicted from vernalization genotype
Purpose • Validate candidate genes for winter and facultative growth habit in barley. • Important to OSU barley breeding program for development of fall sown malting barley with cold tolerance. • Determine if growth habit can be predicted from vernalization genotype
 Winter Hardiness A function of: • Low temperature tolerance • Photoperiod • Vernalization
Winter Hardiness A function of: • Low temperature tolerance • Photoperiod • Vernalization
 Vernalization Background • Vernalization: Cold temperature required for the vegetative to reproductive transition in an agronomically acceptable time frame. Required by winter barleys such as Strider
Vernalization Background • Vernalization: Cold temperature required for the vegetative to reproductive transition in an agronomically acceptable time frame. Required by winter barleys such as Strider
 Barley growth habit types as related to vernalization requirement • Winter: Vernalization required and cold tolerant • Facultative: No vernalization required and cold tolerant • Spring: No vernalization required and cold sensitive Subject of study since 1970
Barley growth habit types as related to vernalization requirement • Winter: Vernalization required and cold tolerant • Facultative: No vernalization required and cold tolerant • Spring: No vernalization required and cold sensitive Subject of study since 1970
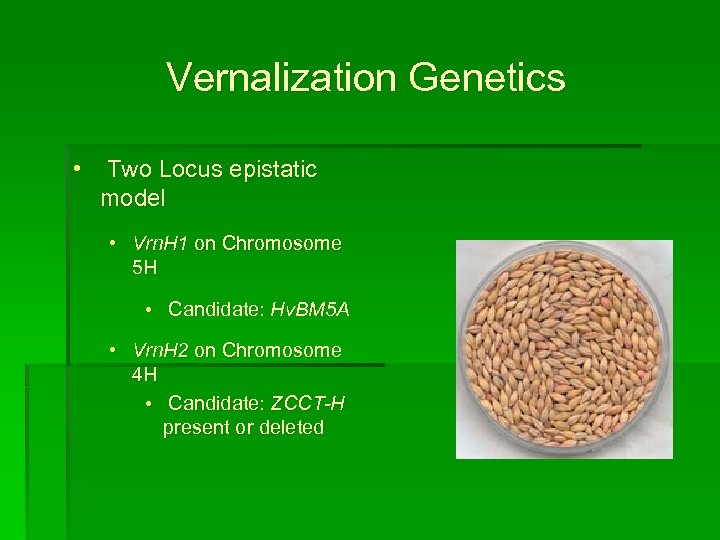 Vernalization Genetics • Two Locus epistatic model • Vrn. H 1 on Chromosome 5 H • Candidate: Hv. BM 5 A • Vrn. H 2 on Chromosome 4 H • Candidate: ZCCT-H present or deleted
Vernalization Genetics • Two Locus epistatic model • Vrn. H 1 on Chromosome 5 H • Candidate: Hv. BM 5 A • Vrn. H 2 on Chromosome 4 H • Candidate: ZCCT-H present or deleted
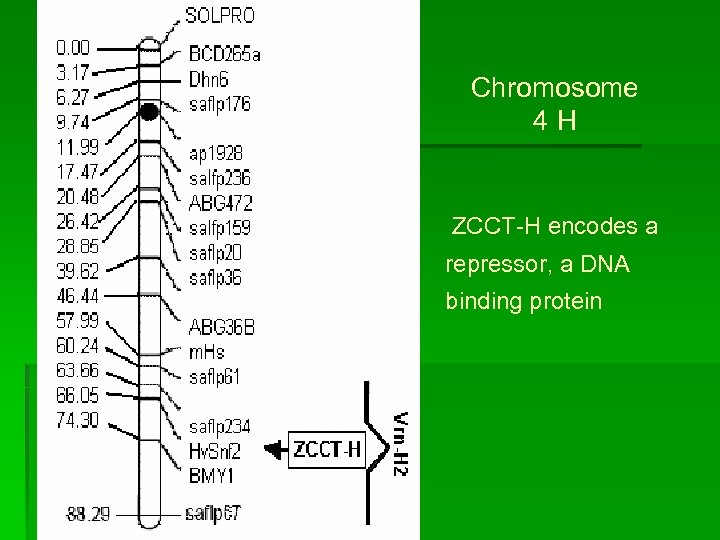 Chromosome 4 H ZCCT-H encodes a repressor, a DNA binding protein
Chromosome 4 H ZCCT-H encodes a repressor, a DNA binding protein
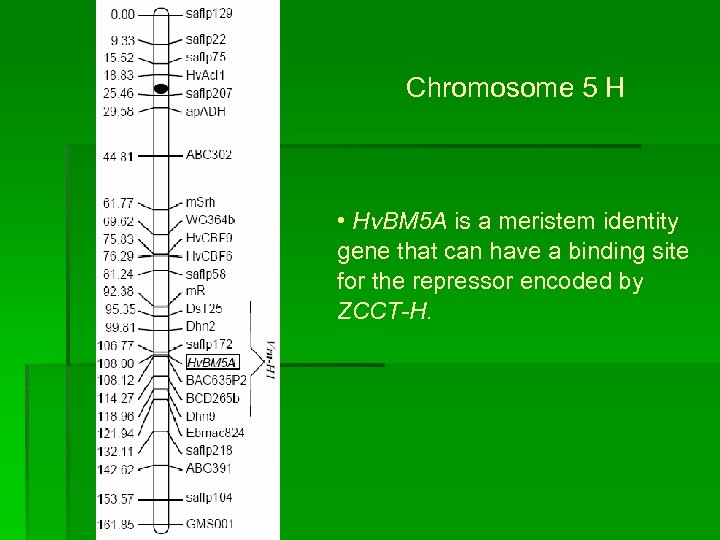 Chromosome 5 H • Hv. BM 5 A is a meristem identity gene that can have a binding site for the repressor encoded by ZCCT-H.
Chromosome 5 H • Hv. BM 5 A is a meristem identity gene that can have a binding site for the repressor encoded by ZCCT-H.
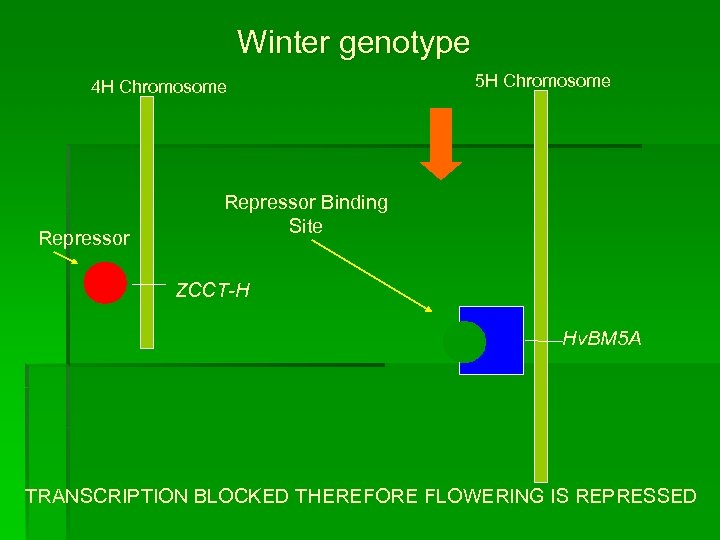 Winter genotype 5 H Chromosome 4 H Chromosome Repressor Binding Site ZCCT-H Hv. BM 5 A TRANSCRIPTION BLOCKED THEREFORE FLOWERING IS REPRESSED
Winter genotype 5 H Chromosome 4 H Chromosome Repressor Binding Site ZCCT-H Hv. BM 5 A TRANSCRIPTION BLOCKED THEREFORE FLOWERING IS REPRESSED
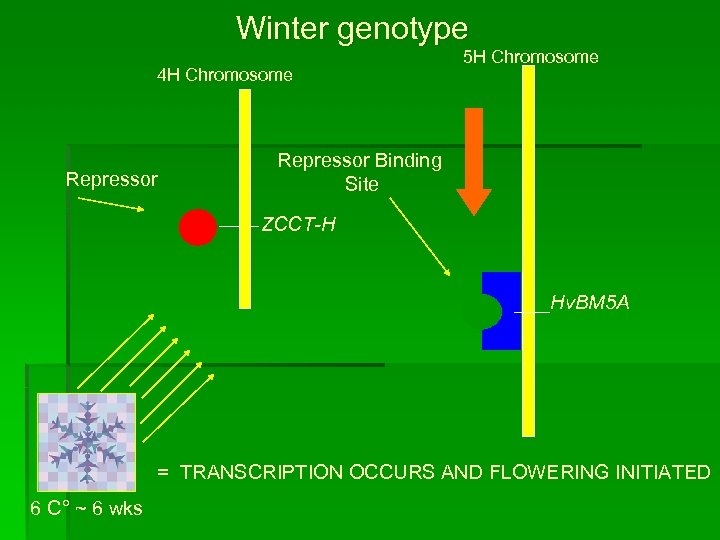 Winter genotype 4 H Chromosome Repressor 5 H Chromosome Repressor Binding Site ZCCT-H Hv. BM 5 A = TRANSCRIPTION OCCURS AND FLOWERING INITIATED 6 C° ~ 6 wks
Winter genotype 4 H Chromosome Repressor 5 H Chromosome Repressor Binding Site ZCCT-H Hv. BM 5 A = TRANSCRIPTION OCCURS AND FLOWERING INITIATED 6 C° ~ 6 wks
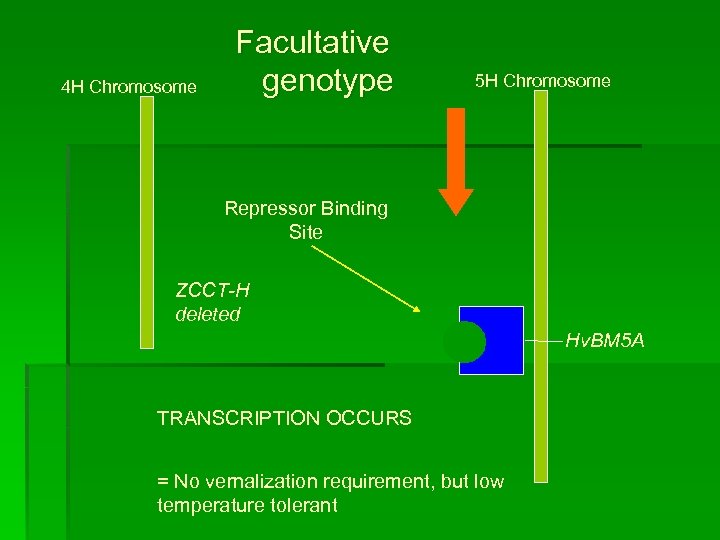 4 H Chromosome Facultative genotype 5 H Chromosome Repressor Binding Site ZCCT-H deleted TRANSCRIPTION OCCURS = No vernalization requirement, but low temperature tolerant Hv. BM 5 A
4 H Chromosome Facultative genotype 5 H Chromosome Repressor Binding Site ZCCT-H deleted TRANSCRIPTION OCCURS = No vernalization requirement, but low temperature tolerant Hv. BM 5 A
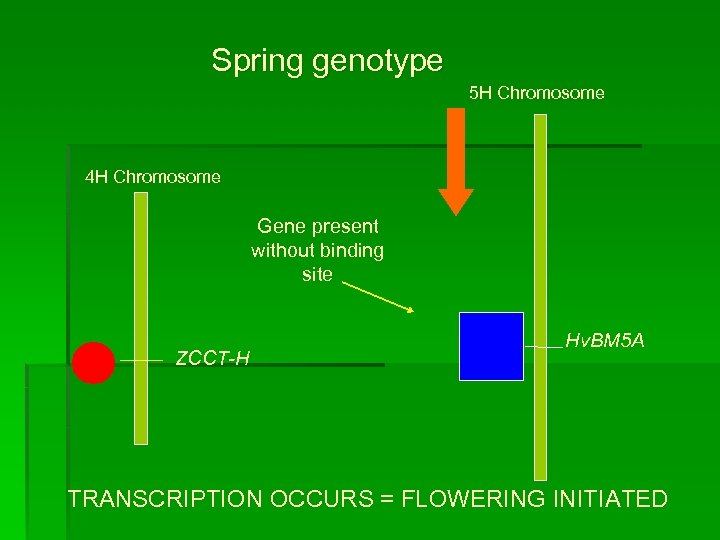 Spring genotype 5 H Chromosome 4 H Chromosome Gene present without binding site ZCCT-H Hv. BM 5 A TRANSCRIPTION OCCURS = FLOWERING INITIATED
Spring genotype 5 H Chromosome 4 H Chromosome Gene present without binding site ZCCT-H Hv. BM 5 A TRANSCRIPTION OCCURS = FLOWERING INITIATED
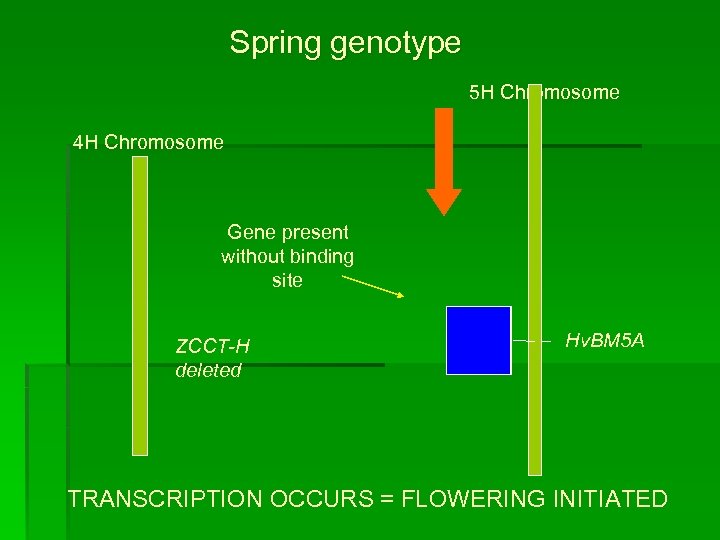 Spring genotype 5 H Chromosome 4 H Chromosome Gene present without binding site ZCCT-H deleted Hv. BM 5 A TRANSCRIPTION OCCURS = FLOWERING INITIATED
Spring genotype 5 H Chromosome 4 H Chromosome Gene present without binding site ZCCT-H deleted Hv. BM 5 A TRANSCRIPTION OCCURS = FLOWERING INITIATED
 Von Zitzewitz et al. (2005) Relates Growth Habit to Vrn Genotype • Winter types require vernalization because: • They contain the ZCCT-H (repressor) gene and the Hv. BM 5 A “winter” allele with a repressor binding site
Von Zitzewitz et al. (2005) Relates Growth Habit to Vrn Genotype • Winter types require vernalization because: • They contain the ZCCT-H (repressor) gene and the Hv. BM 5 A “winter” allele with a repressor binding site
 Von Zitzewitz et al. (cont. ) • Spring types require no vernalization because: • They contain Hv. BM 5 A allele with a deletion: no repressor binding site • ZCCT-H gene may or may not be present
Von Zitzewitz et al. (cont. ) • Spring types require no vernalization because: • They contain Hv. BM 5 A allele with a deletion: no repressor binding site • ZCCT-H gene may or may not be present
 Von Zitzewitz et al. (cont. ) • Facultative types do not require vernalization because: • • Deletion of ZCCT-H and Presence of the Hv. BM 5 A winter allele with a repressor binding site
Von Zitzewitz et al. (cont. ) • Facultative types do not require vernalization because: • • Deletion of ZCCT-H and Presence of the Hv. BM 5 A winter allele with a repressor binding site
 Methods and Materials
Methods and Materials
 Phenotype determination • Plant growth staging using Feekes growth stage scale • Each assessment was performed on 54 genotypes – fall-sown and spring-sown
Phenotype determination • Plant growth staging using Feekes growth stage scale • Each assessment was performed on 54 genotypes – fall-sown and spring-sown
 Feekes Growth Stages • Assessed every 14 days • On a plot basis • Terminated at Feekes 10. 5
Feekes Growth Stages • Assessed every 14 days • On a plot basis • Terminated at Feekes 10. 5
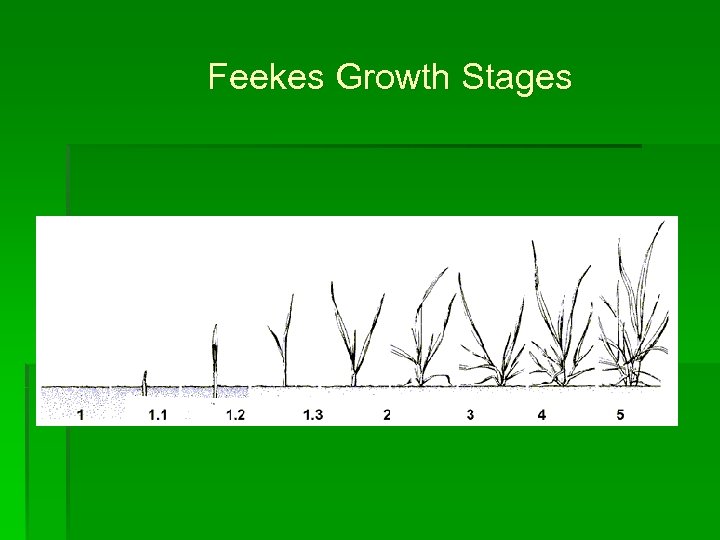 Feekes Growth Stages
Feekes Growth Stages
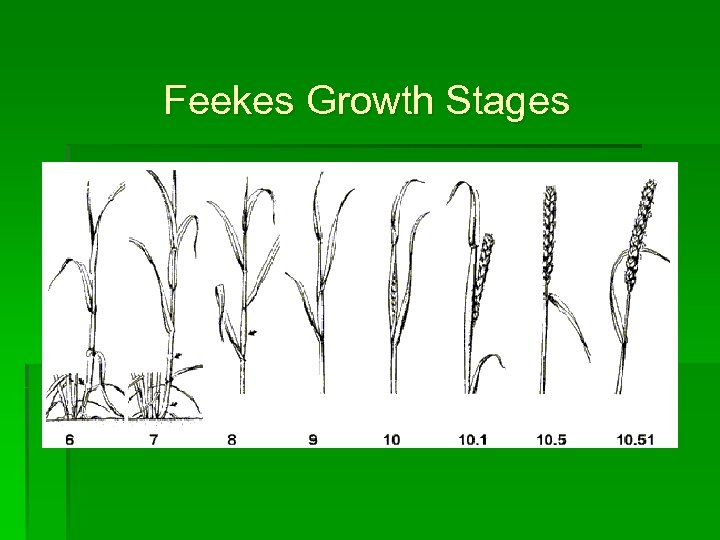 Feekes Growth Stages
Feekes Growth Stages
 Growing Degree Day Vernalization Coefficient (GDD-VC) • Growing Degree Days (GDD) from planting to Feekes 10. 5 for fall and spring sown experiments • GDD was calculated using a 10°C base • GDD-VC = spring GDD – fall GDD = 1, 000 for lines that did not flower in the spring-sown nursery
Growing Degree Day Vernalization Coefficient (GDD-VC) • Growing Degree Days (GDD) from planting to Feekes 10. 5 for fall and spring sown experiments • GDD was calculated using a 10°C base • GDD-VC = spring GDD – fall GDD = 1, 000 for lines that did not flower in the spring-sown nursery
 Genotyping procedures • 54 lines in the ORELT genotyped at two locations: USDA/ARS lab at Pullman, WA and OSU Barley project lab at Corvallis, OR • Hv. BM 5 A dominant PCRbased marker • ZCCT-H a dominant PCRbased marker
Genotyping procedures • 54 lines in the ORELT genotyped at two locations: USDA/ARS lab at Pullman, WA and OSU Barley project lab at Corvallis, OR • Hv. BM 5 A dominant PCRbased marker • ZCCT-H a dominant PCRbased marker
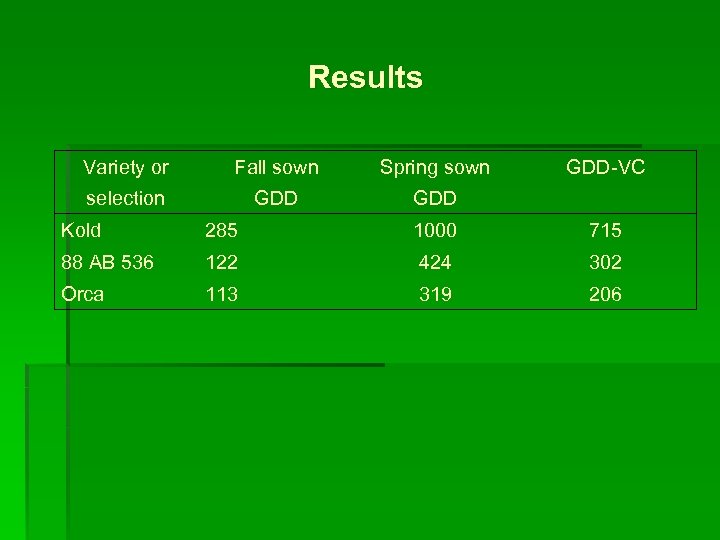 Results Variety or Fall sown Spring sown GDD-VC selection GDD Kold 285 1000 715 88 AB 536 122 424 302 Orca 113 319 206
Results Variety or Fall sown Spring sown GDD-VC selection GDD Kold 285 1000 715 88 AB 536 122 424 302 Orca 113 319 206
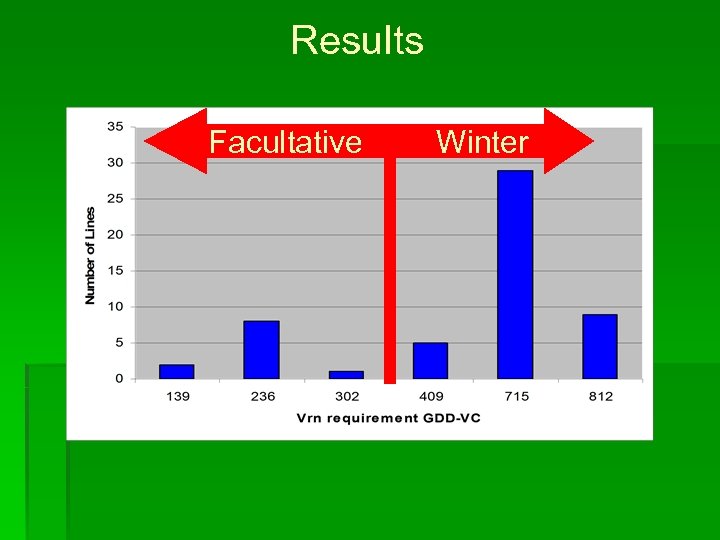 Results Facultative Winter
Results Facultative Winter
 Results § 100% agreement for Hv. BM 5 A markers § All entries are therefore winter or facultative
Results § 100% agreement for Hv. BM 5 A markers § All entries are therefore winter or facultative
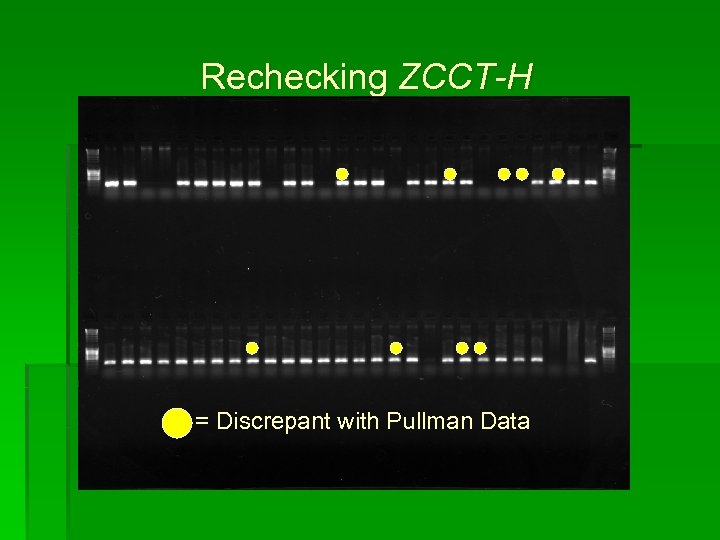 Rechecking ZCCT-H = Discrepant with Pullman Data
Rechecking ZCCT-H = Discrepant with Pullman Data
 Results ZCCT-H discrepancies between labs § Dominant marker § Considered the presence of an amplicon at one location to be evidence of gene presence
Results ZCCT-H discrepancies between labs § Dominant marker § Considered the presence of an amplicon at one location to be evidence of gene presence
 Results • 7 lines failed to amplify at USDA/ARS lab that amplified at the OSU lab • 2 lines failed to amplify at the OSU lab that amplified at the USDA/ARS lab
Results • 7 lines failed to amplify at USDA/ARS lab that amplified at the OSU lab • 2 lines failed to amplify at the OSU lab that amplified at the USDA/ARS lab
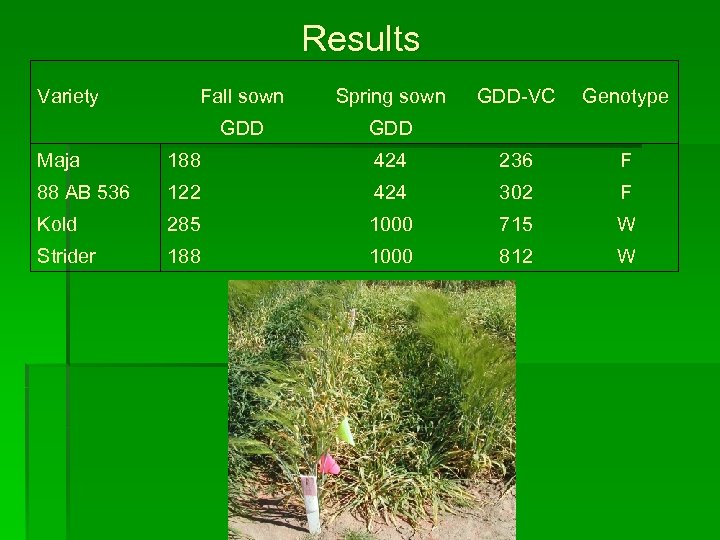 Results Variety Fall sown Spring sown GDD-VC GDD Genotype Maja 188 424 236 F 88 AB 536 122 424 302 F Kold 285 1000 715 W Strider 188 1000 812 W
Results Variety Fall sown Spring sown GDD-VC GDD Genotype Maja 188 424 236 F 88 AB 536 122 424 302 F Kold 285 1000 715 W Strider 188 1000 812 W
 3 lines violate the model ORELT Variety or Fall Spring GDD- growth habit no. Selection GDD VC classification 18 J 2 -6 -19 188 424 236 W 23 Stab. BC 50 -9 -1 188 424 236 W 24 Stab. BC 50 -7 -1 285 424 139 W
3 lines violate the model ORELT Variety or Fall Spring GDD- growth habit no. Selection GDD VC classification 18 J 2 -6 -19 188 424 236 W 23 Stab. BC 50 -9 -1 188 424 236 W 24 Stab. BC 50 -7 -1 285 424 139 W
 Conclusions • The model was validated: 51 out of 54 lines fit • Accurately predict growth habit from marker data ~ 94% of the time • The Vernalization phenotype is a suitable target for marker assisted selection (MAS)
Conclusions • The model was validated: 51 out of 54 lines fit • Accurately predict growth habit from marker data ~ 94% of the time • The Vernalization phenotype is a suitable target for marker assisted selection (MAS)
 Explanation for not having 100% fit: • Other “maturity” genes possibly play a role in determination of spring/winter phenotypes • DNA contamination • Ambiguity in rating the phenotype • Residual heterogeneity
Explanation for not having 100% fit: • Other “maturity” genes possibly play a role in determination of spring/winter phenotypes • DNA contamination • Ambiguity in rating the phenotype • Residual heterogeneity
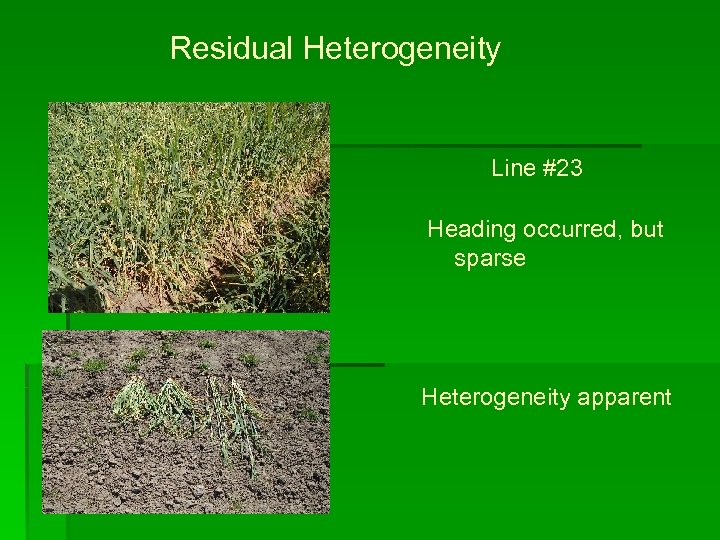 Residual Heterogeneity Line #23 Heading occurred, but sparse Heterogeneity apparent
Residual Heterogeneity Line #23 Heading occurred, but sparse Heterogeneity apparent
 Recommendations for future research • Improve genotyping • Augment the Vrn-H 2 genotyping with a codominant assay. • Hv. Snf 2 is tightly linked to ZCCT-H and present in all growth habit types
Recommendations for future research • Improve genotyping • Augment the Vrn-H 2 genotyping with a codominant assay. • Hv. Snf 2 is tightly linked to ZCCT-H and present in all growth habit types
 Recommendations for future research • Improve phenotyping • Record growth stages daily • Count Final Leaf Number on ambiguous lines
Recommendations for future research • Improve phenotyping • Record growth stages daily • Count Final Leaf Number on ambiguous lines
 Acknowledgments I would like to thank: § § Dr. Hayes for his extreme patience and help along the way Dr. Chastain for his guidance and encouragement Ann Corey for all the help with the field portion of this experiment Peter Szucs for the many hours of help in the lab. He took my project seriously and was an excellent teacher § The Hayes barley lab at Oregon State University for providing answers to my questions § The Bioresource Research program for providing an opportunity to undertake such a project as an undergraduate § The Crop and Soil Science Department at Oregon State University. I would hate to think what school would have been like without my CSS family. Thank you very much!
Acknowledgments I would like to thank: § § Dr. Hayes for his extreme patience and help along the way Dr. Chastain for his guidance and encouragement Ann Corey for all the help with the field portion of this experiment Peter Szucs for the many hours of help in the lab. He took my project seriously and was an excellent teacher § The Hayes barley lab at Oregon State University for providing answers to my questions § The Bioresource Research program for providing an opportunity to undertake such a project as an undergraduate § The Crop and Soil Science Department at Oregon State University. I would hate to think what school would have been like without my CSS family. Thank you very much!


