bc02b4f58025e7412b1745409044fa91.ppt
- Количество слайдов: 67
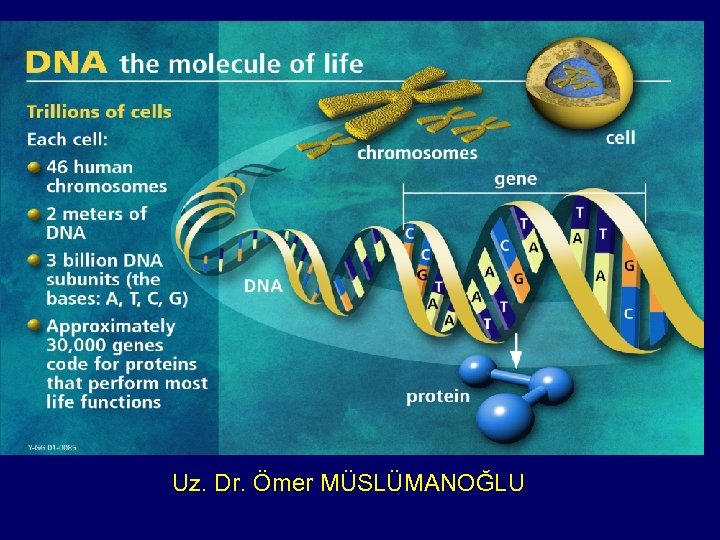 Uz. Dr. Ömer MÜSLÜMANOĞLU
Uz. Dr. Ömer MÜSLÜMANOĞLU
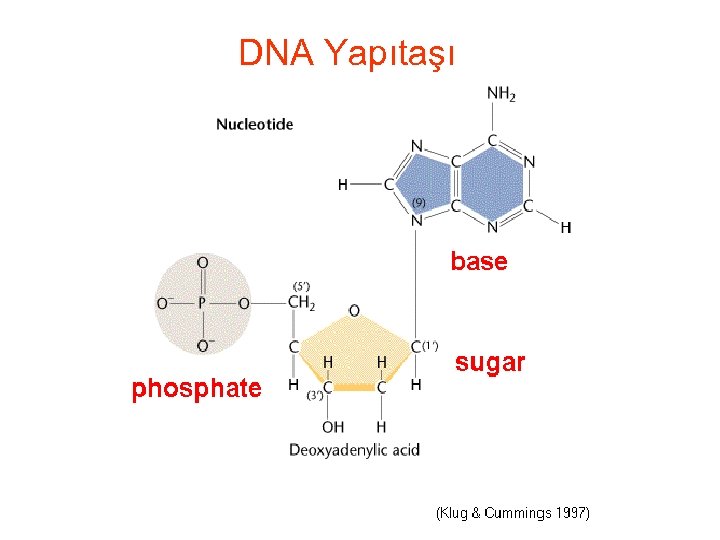 DNA Yapıtaşı
DNA Yapıtaşı
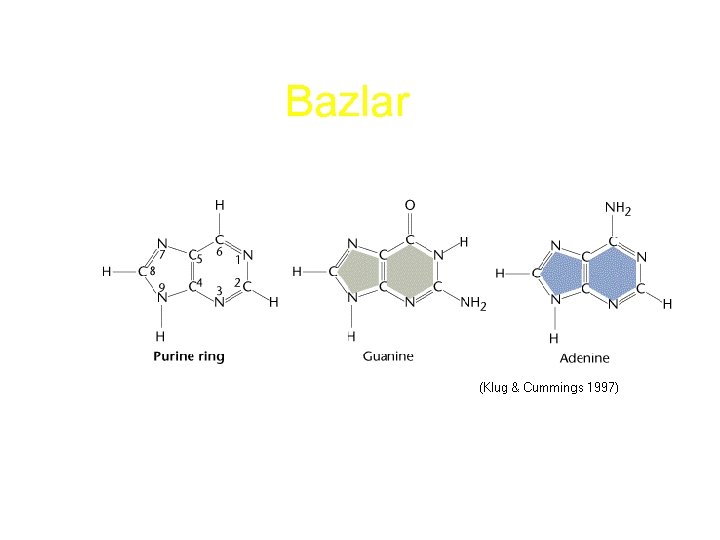 Bazlar
Bazlar
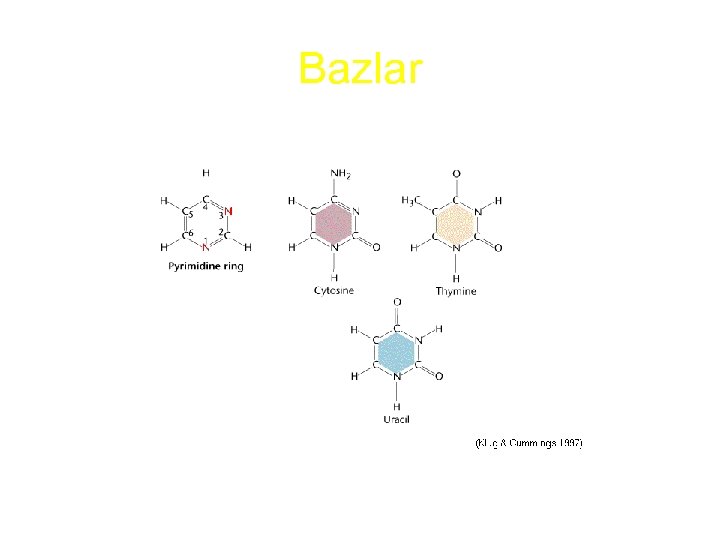 Bazlar
Bazlar
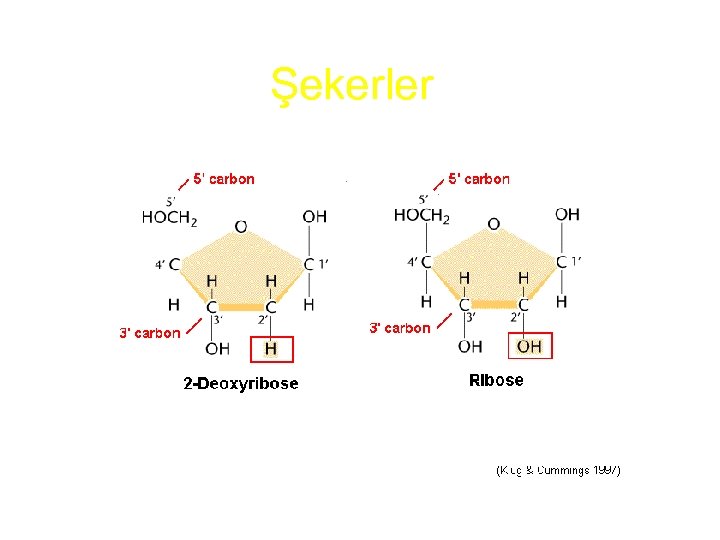 Şekerler
Şekerler
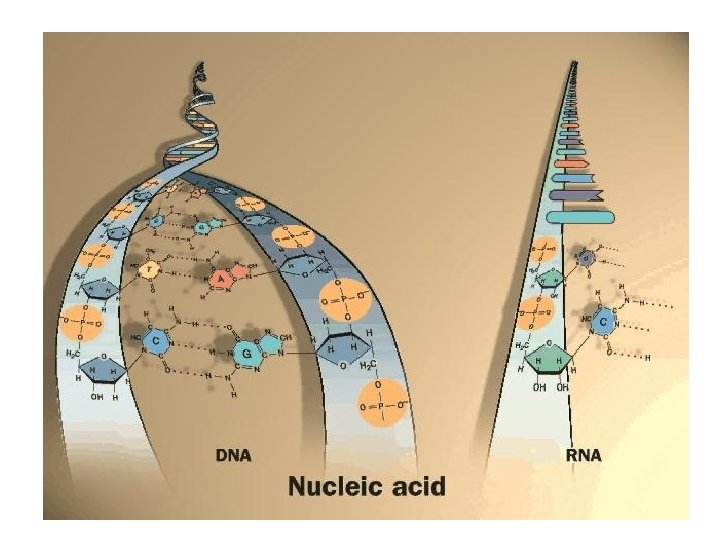
 DNA-RNA arasında temel farklar DNA RNA Şeker deoksiriboz Baz çifti Timin-Adenin Sitozin-Guanin Urasil-Adenin Sitozin-Guanin Yapısı Çift sarmal heliks Tek Sarmal Düzensiz Dayanıklılık Stabil DNAse ile yıkılır Baz hidrolizine açık RNAse ile yıkılır Fonksyonu Genetik bilgiyi nukleusta Genetik bilgiyi sitoplazmaya saklar taşır
DNA-RNA arasında temel farklar DNA RNA Şeker deoksiriboz Baz çifti Timin-Adenin Sitozin-Guanin Urasil-Adenin Sitozin-Guanin Yapısı Çift sarmal heliks Tek Sarmal Düzensiz Dayanıklılık Stabil DNAse ile yıkılır Baz hidrolizine açık RNAse ile yıkılır Fonksyonu Genetik bilgiyi nukleusta Genetik bilgiyi sitoplazmaya saklar taşır
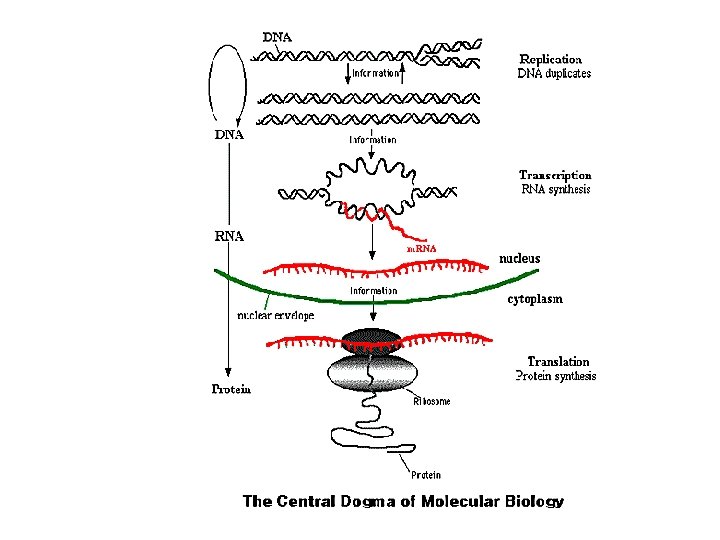
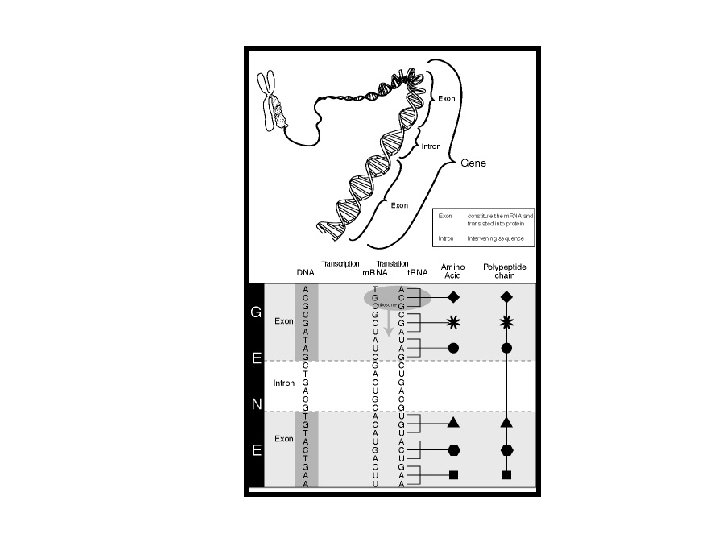
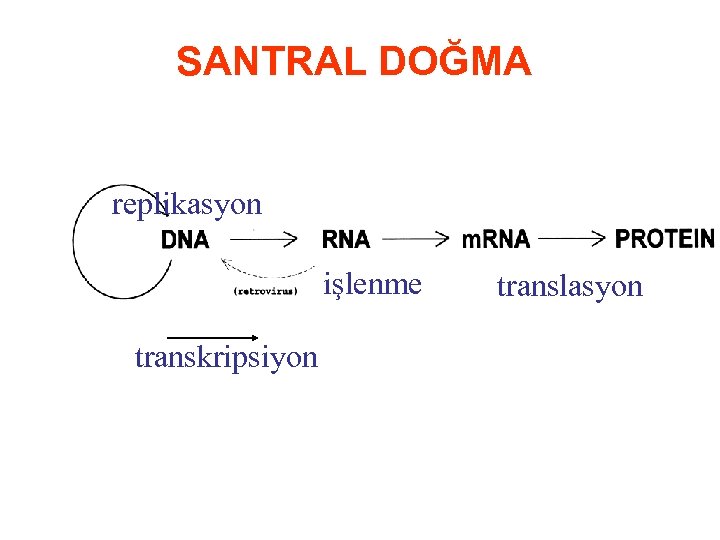 SANTRAL DOĞMA replikasyon işlenme transkripsiyon translasyon
SANTRAL DOĞMA replikasyon işlenme transkripsiyon translasyon

 DNA ISOLATION
DNA ISOLATION
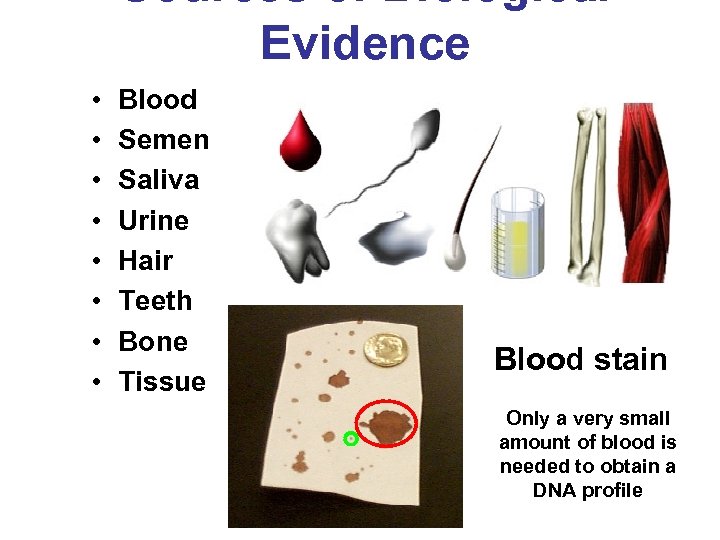 Sources of Biological Evidence • • Blood Semen Saliva Urine Hair Teeth Bone Tissue Blood stain Only a very small amount of blood is needed to obtain a DNA profile
Sources of Biological Evidence • • Blood Semen Saliva Urine Hair Teeth Bone Tissue Blood stain Only a very small amount of blood is needed to obtain a DNA profile
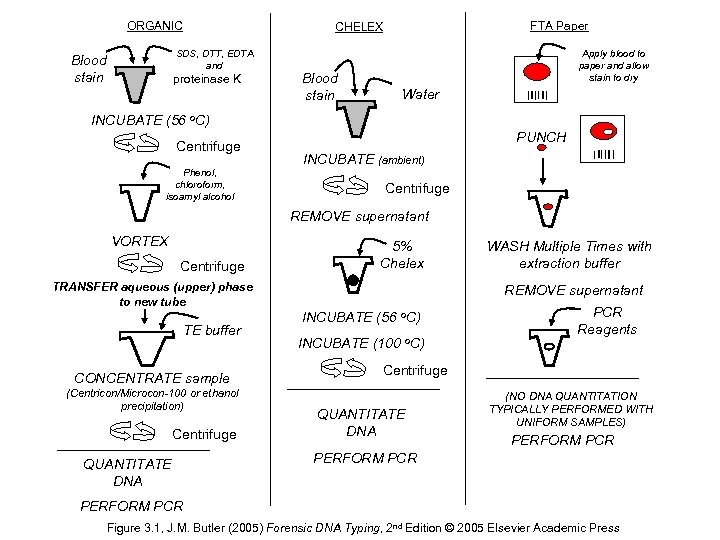 ORGANIC SDS, DTT, EDTA and Blood stain FTA Paper CHELEX proteinase K Blood stain Apply blood to paper and allow stain to dry Water INCUBATE (56 o. C) Centrifuge Phenol, chloroform, isoamyl alcohol PUNCH INCUBATE (ambient) Centrifuge REMOVE supernatant VORTEX Centrifuge 5% Chelex TRANSFER aqueous (upper) phase to new tube TE buffer CONCENTRATE sample (Centricon/Microcon-100 or ethanol precipitation) Centrifuge QUANTITATE DNA WASH Multiple Times with extraction buffer REMOVE supernatant INCUBATE (56 o. C) INCUBATE (100 o. C) PCR Reagents Centrifuge QUANTITATE DNA (NO DNA QUANTITATION TYPICALLY PERFORMED WITH UNIFORM SAMPLES) PERFORM PCR Figure 3. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
ORGANIC SDS, DTT, EDTA and Blood stain FTA Paper CHELEX proteinase K Blood stain Apply blood to paper and allow stain to dry Water INCUBATE (56 o. C) Centrifuge Phenol, chloroform, isoamyl alcohol PUNCH INCUBATE (ambient) Centrifuge REMOVE supernatant VORTEX Centrifuge 5% Chelex TRANSFER aqueous (upper) phase to new tube TE buffer CONCENTRATE sample (Centricon/Microcon-100 or ethanol precipitation) Centrifuge QUANTITATE DNA WASH Multiple Times with extraction buffer REMOVE supernatant INCUBATE (56 o. C) INCUBATE (100 o. C) PCR Reagents Centrifuge QUANTITATE DNA (NO DNA QUANTITATION TYPICALLY PERFORMED WITH UNIFORM SAMPLES) PERFORM PCR Figure 3. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
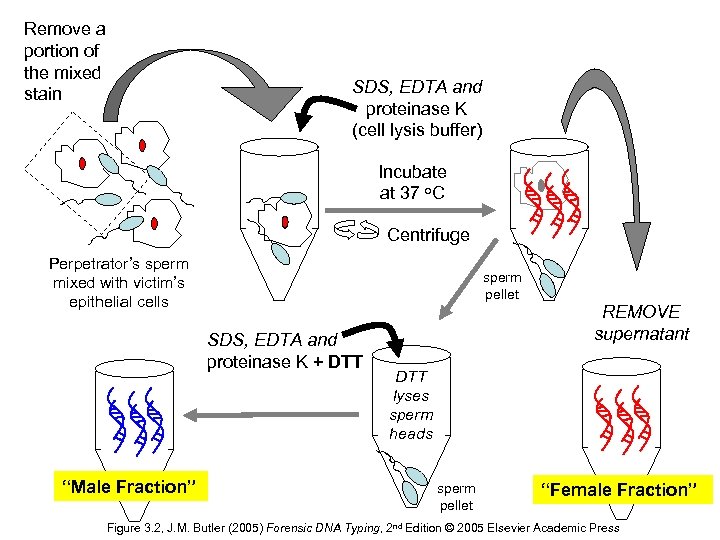 Remove a portion of the mixed stain SDS, EDTA and proteinase K (cell lysis buffer) Incubate at 37 o. C Centrifuge Perpetrator’s sperm mixed with victim’s epithelial cells sperm pellet SDS, EDTA and proteinase K + DTT “Male Fraction” REMOVE supernatant DTT lyses sperm heads sperm pellet “Female Fraction” Figure 3. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Remove a portion of the mixed stain SDS, EDTA and proteinase K (cell lysis buffer) Incubate at 37 o. C Centrifuge Perpetrator’s sperm mixed with victim’s epithelial cells sperm pellet SDS, EDTA and proteinase K + DTT “Male Fraction” REMOVE supernatant DTT lyses sperm heads sperm pellet “Female Fraction” Figure 3. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
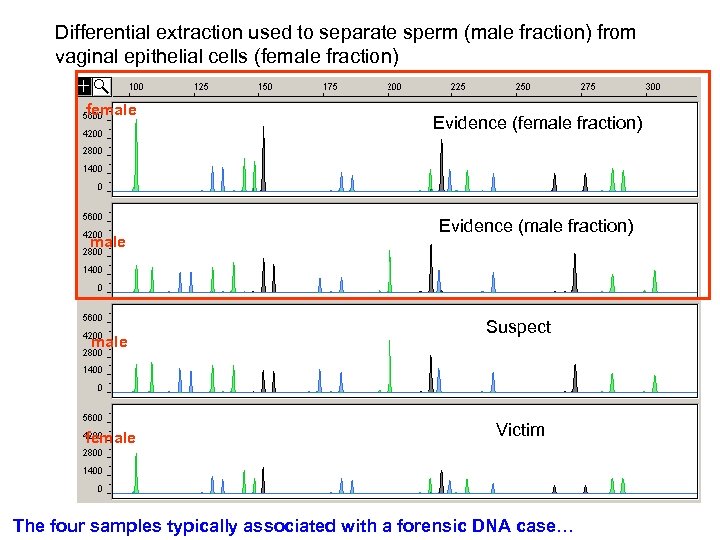 Differential extraction used to separate sperm (male fraction) from vaginal epithelial cells (female fraction) female female Evidence (female fraction) Evidence (male fraction) Suspect Victim The four samples typically associated with a forensic DNA case…
Differential extraction used to separate sperm (male fraction) from vaginal epithelial cells (female fraction) female female Evidence (female fraction) Evidence (male fraction) Suspect Victim The four samples typically associated with a forensic DNA case…
 QUANTITATION
QUANTITATION
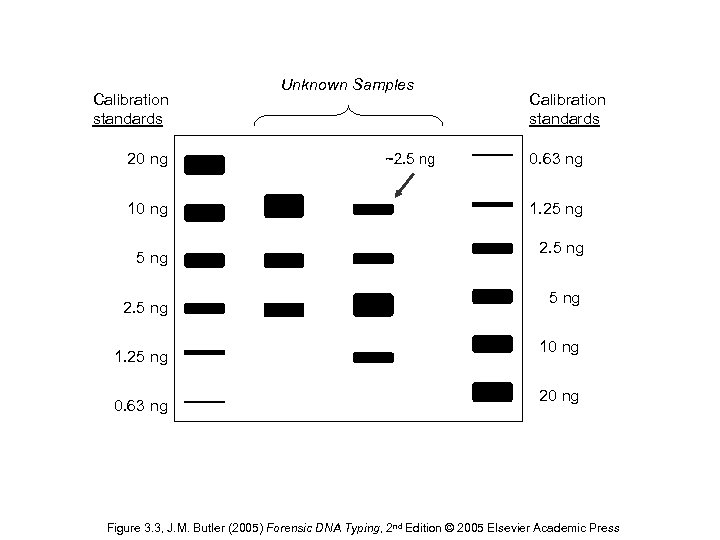 Calibration standards 20 ng 10 ng 5 ng 2. 5 ng 1. 25 ng 0. 63 ng Unknown Samples ~2. 5 ng Calibration standards 0. 63 ng 1. 25 ng 2. 5 ng 10 ng 20 ng Figure 3. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Calibration standards 20 ng 10 ng 5 ng 2. 5 ng 1. 25 ng 0. 63 ng Unknown Samples ~2. 5 ng Calibration standards 0. 63 ng 1. 25 ng 2. 5 ng 10 ng 20 ng Figure 3. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
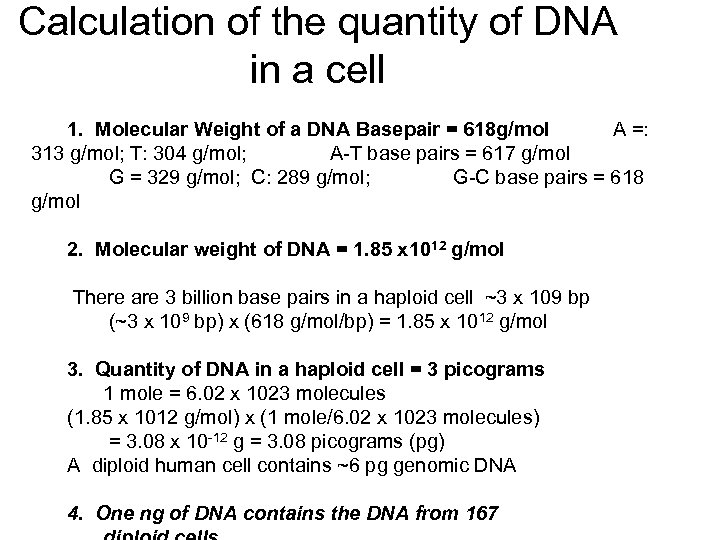 Calculation of the quantity of DNA in a cell 1. Molecular Weight of a DNA Basepair = 618 g/mol A =: 313 g/mol; T: 304 g/mol; A-T base pairs = 617 g/mol G = 329 g/mol; C: 289 g/mol; G-C base pairs = 618 g/mol 2. Molecular weight of DNA = 1. 85 x 1012 g/mol There are 3 billion base pairs in a haploid cell ~3 x 109 bp (~3 x 109 bp) x (618 g/mol/bp) = 1. 85 x 1012 g/mol 3. Quantity of DNA in a haploid cell = 3 picograms 1 mole = 6. 02 x 1023 molecules (1. 85 x 1012 g/mol) x (1 mole/6. 02 x 1023 molecules) = 3. 08 x 10 -12 g = 3. 08 picograms (pg) A diploid human cell contains ~6 pg genomic DNA 4. One ng of DNA contains the DNA from 167
Calculation of the quantity of DNA in a cell 1. Molecular Weight of a DNA Basepair = 618 g/mol A =: 313 g/mol; T: 304 g/mol; A-T base pairs = 617 g/mol G = 329 g/mol; C: 289 g/mol; G-C base pairs = 618 g/mol 2. Molecular weight of DNA = 1. 85 x 1012 g/mol There are 3 billion base pairs in a haploid cell ~3 x 109 bp (~3 x 109 bp) x (618 g/mol/bp) = 1. 85 x 1012 g/mol 3. Quantity of DNA in a haploid cell = 3 picograms 1 mole = 6. 02 x 1023 molecules (1. 85 x 1012 g/mol) x (1 mole/6. 02 x 1023 molecules) = 3. 08 x 10 -12 g = 3. 08 picograms (pg) A diploid human cell contains ~6 pg genomic DNA 4. One ng of DNA contains the DNA from 167
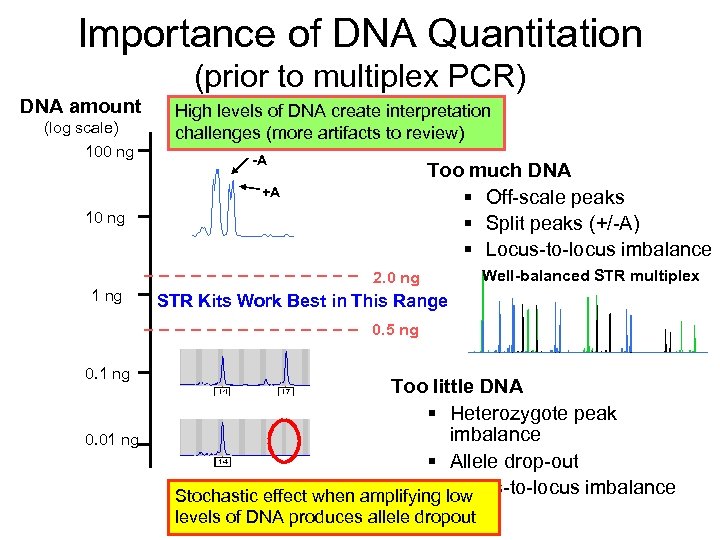 Importance of DNA Quantitation (prior to multiplex PCR) DNA amount (log scale) 100 ng High levels of DNA create interpretation challenges (more artifacts to review) -A Too much DNA § Off-scale peaks § Split peaks (+/-A) § Locus-to-locus imbalance +A 10 ng 2. 0 ng 1 ng Well-balanced STR multiplex STR Kits Work Best in This Range 0. 5 ng 0. 1 ng 0. 01 ng Too little DNA § Heterozygote peak imbalance § Allele drop-out § Locus-to-locus imbalance Stochastic effect when amplifying low levels of DNA produces allele dropout
Importance of DNA Quantitation (prior to multiplex PCR) DNA amount (log scale) 100 ng High levels of DNA create interpretation challenges (more artifacts to review) -A Too much DNA § Off-scale peaks § Split peaks (+/-A) § Locus-to-locus imbalance +A 10 ng 2. 0 ng 1 ng Well-balanced STR multiplex STR Kits Work Best in This Range 0. 5 ng 0. 1 ng 0. 01 ng Too little DNA § Heterozygote peak imbalance § Allele drop-out § Locus-to-locus imbalance Stochastic effect when amplifying low levels of DNA produces allele dropout
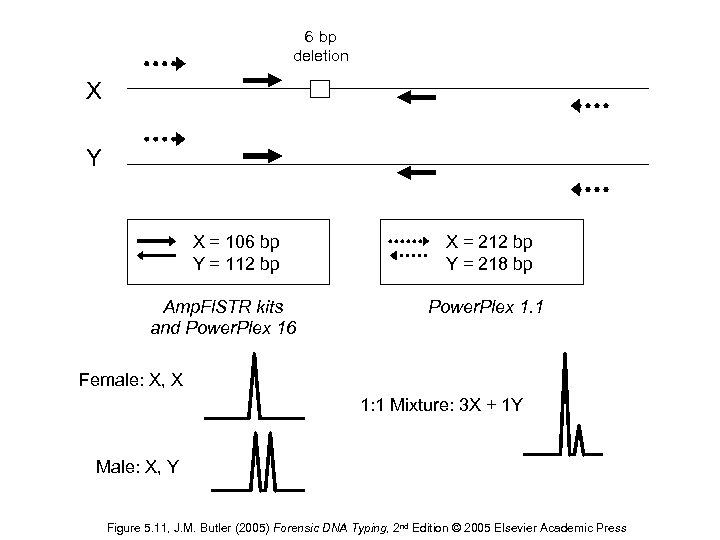 6 bp deletion X Y X = 106 bp Y = 112 bp Amp. Fl. STR kits and Power. Plex 16 X = 212 bp Y = 218 bp Power. Plex 1. 1 Female: X, X 1: 1 Mixture: 3 X + 1 Y Male: X, Y Figure 5. 11, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
6 bp deletion X Y X = 106 bp Y = 112 bp Amp. Fl. STR kits and Power. Plex 16 X = 212 bp Y = 218 bp Power. Plex 1. 1 Female: X, X 1: 1 Mixture: 3 X + 1 Y Male: X, Y Figure 5. 11, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 PCR
PCR
 POLİMORFİZM • Bir popülasyonda mevcut olan genetik çeşitliliğe polimorfizm denir • DNA Polimorfizmi, DNA üzerinde hastalığa neden olmayan, suskun nükleotid değişimleri olarak tanımlanır.
POLİMORFİZM • Bir popülasyonda mevcut olan genetik çeşitliliğe polimorfizm denir • DNA Polimorfizmi, DNA üzerinde hastalığa neden olmayan, suskun nükleotid değişimleri olarak tanımlanır.
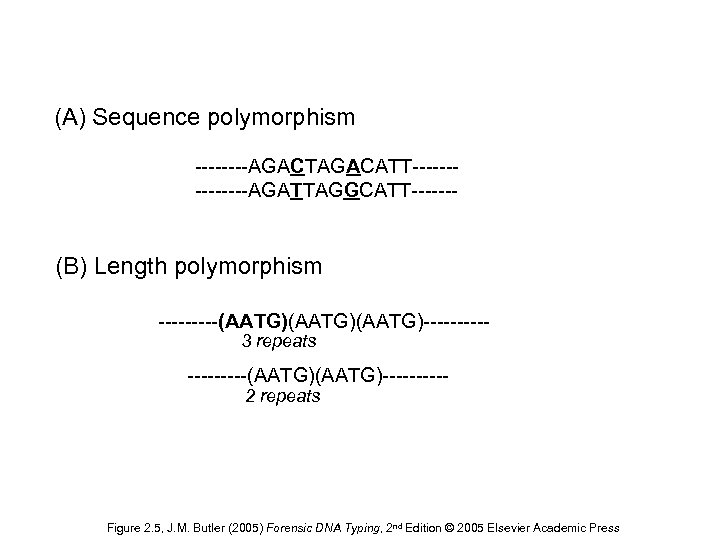 (A) Sequence polymorphism ----AGACTAGACATT-------AGATTAGGCATT------- (B) Length polymorphism -----(AATG)(AATG)-----3 repeats -----(AATG)-----2 repeats Figure 2. 5, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(A) Sequence polymorphism ----AGACTAGACATT-------AGATTAGGCATT------- (B) Length polymorphism -----(AATG)(AATG)-----3 repeats -----(AATG)-----2 repeats Figure 2. 5, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
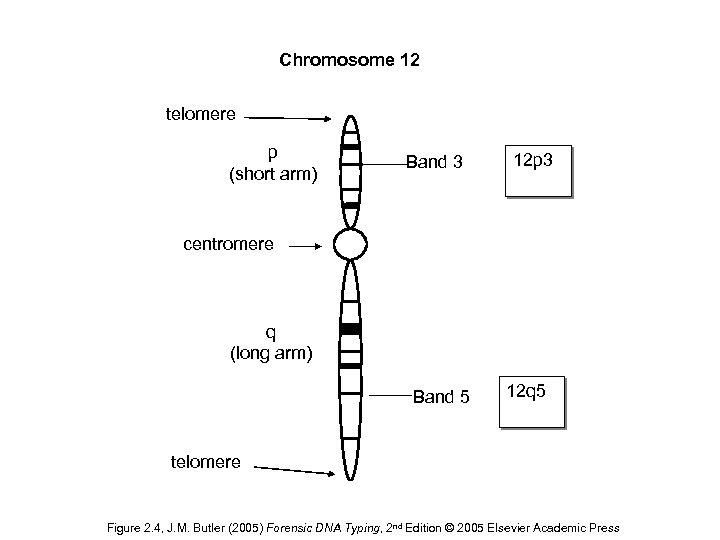 Chromosome 12 telomere p (short arm) Band 3 12 p 3 centromere q (long arm) Band 5 12 q 5 telomere Figure 2. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Chromosome 12 telomere p (short arm) Band 3 12 p 3 centromere q (long arm) Band 5 12 q 5 telomere Figure 2. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
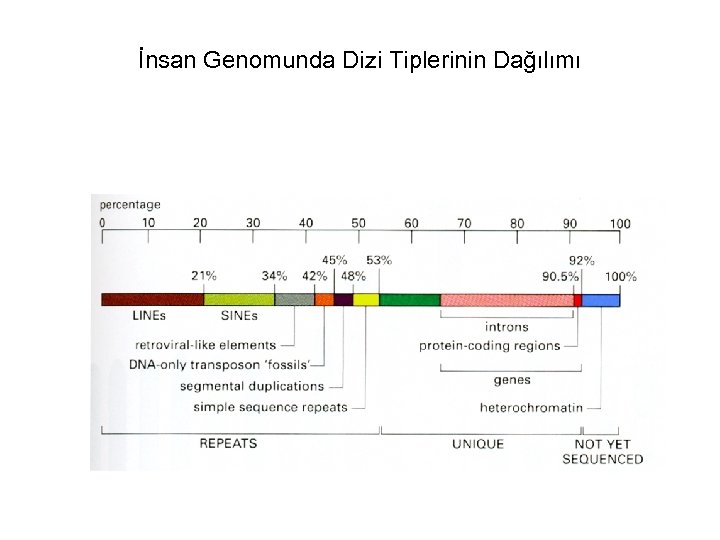 İnsan Genomunda Dizi Tiplerinin Dağılımı
İnsan Genomunda Dizi Tiplerinin Dağılımı
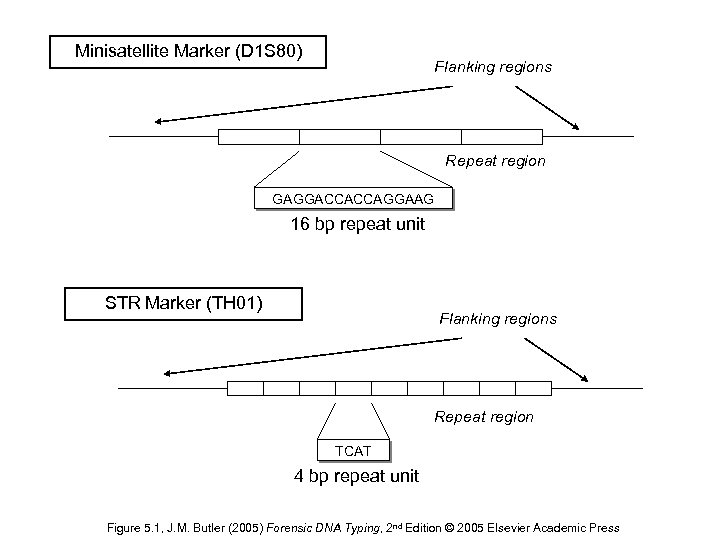 Minisatellite Marker (D 1 S 80) Flanking regions Repeat region GAGGACCACCAGGAAG 16 bp repeat unit STR Marker (TH 01) Flanking regions Repeat region TCAT 4 bp repeat unit Figure 5. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Minisatellite Marker (D 1 S 80) Flanking regions Repeat region GAGGACCACCAGGAAG 16 bp repeat unit STR Marker (TH 01) Flanking regions Repeat region TCAT 4 bp repeat unit Figure 5. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
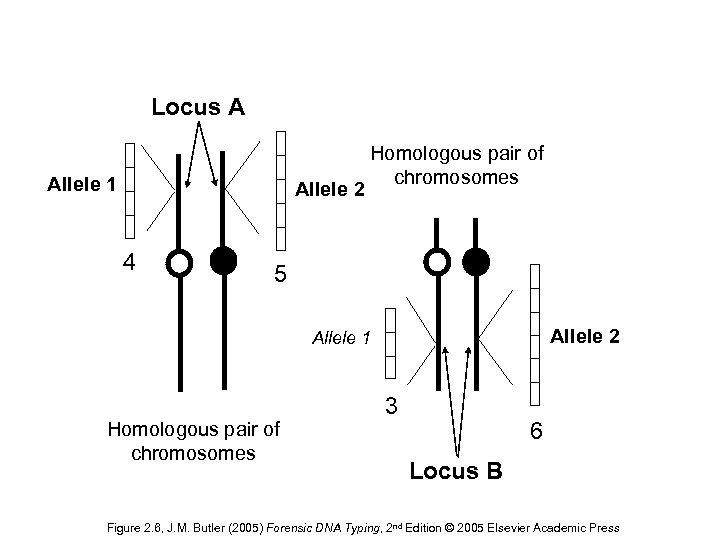 Locus A Allele 1 Allele 2 4 Homologous pair of chromosomes 5 Allele 2 Allele 1 Homologous pair of chromosomes 3 6 Locus B Figure 2. 6, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Locus A Allele 1 Allele 2 4 Homologous pair of chromosomes 5 Allele 2 Allele 1 Homologous pair of chromosomes 3 6 Locus B Figure 2. 6, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
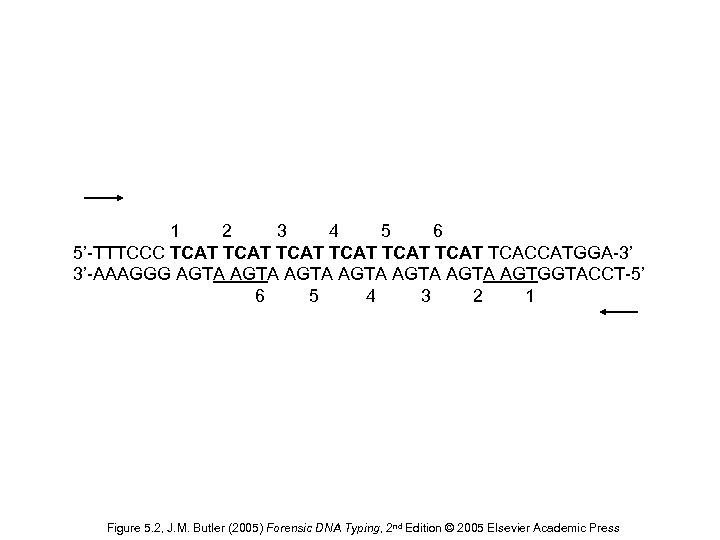 1 2 3 4 5 6 5’-TTTCCC TCAT TCAT TCACCATGGA-3’ 3’-AAAGGG AGTA AGTA AGTGGTACCT-5’ 6 5 4 3 2 1 Figure 5. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
1 2 3 4 5 6 5’-TTTCCC TCAT TCAT TCACCATGGA-3’ 3’-AAAGGG AGTA AGTA AGTGGTACCT-5’ 6 5 4 3 2 1 Figure 5. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
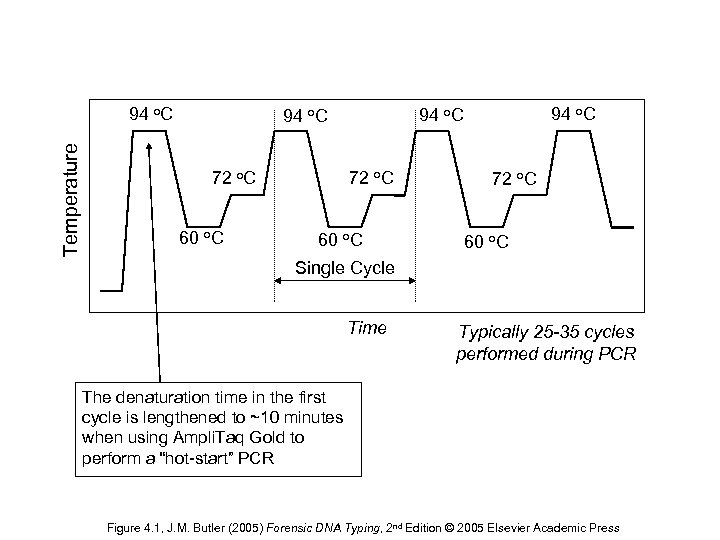 Temperature 94 o. C 72 o. C 60 o. C 94 o. C 60 o. C 72 o. C 60 o. C Single Cycle Time Typically 25 -35 cycles performed during PCR The denaturation time in the first cycle is lengthened to ~10 minutes when using Ampli. Taq Gold to perform a “hot-start” PCR Figure 4. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Temperature 94 o. C 72 o. C 60 o. C 94 o. C 60 o. C 72 o. C 60 o. C Single Cycle Time Typically 25 -35 cycles performed during PCR The denaturation time in the first cycle is lengthened to ~10 minutes when using Ampli. Taq Gold to perform a “hot-start” PCR Figure 4. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 PCR
PCR
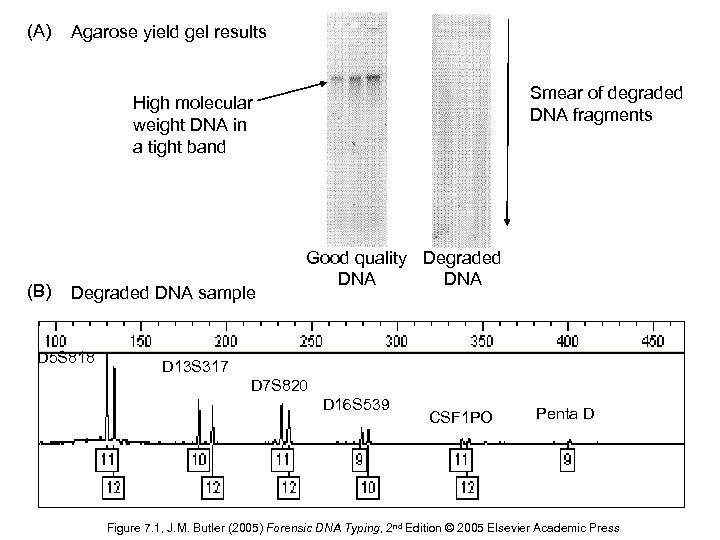 (A) Agarose yield gel results Smear of degraded DNA fragments High molecular weight DNA in a tight band (B) Degraded DNA sample D 5 S 818 Good quality Degraded DNA D 13 S 317 D 7 S 820 D 16 S 539 CSF 1 PO Penta D Figure 7. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(A) Agarose yield gel results Smear of degraded DNA fragments High molecular weight DNA in a tight band (B) Degraded DNA sample D 5 S 818 Good quality Degraded DNA D 13 S 317 D 7 S 820 D 16 S 539 CSF 1 PO Penta D Figure 7. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
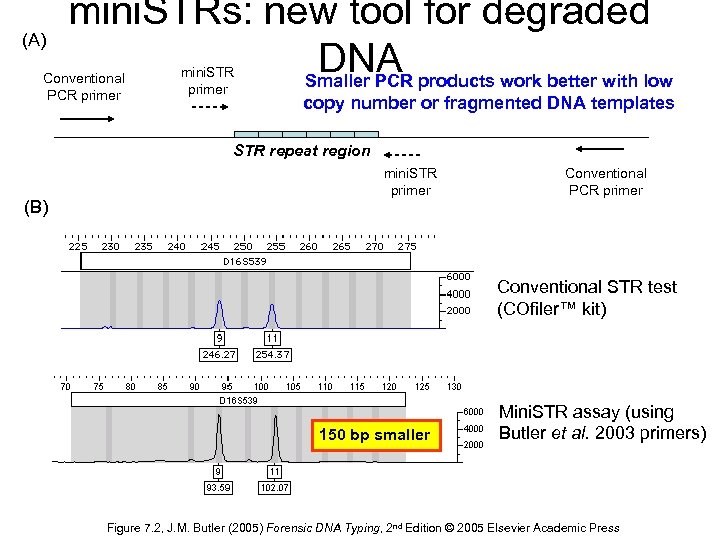 (A) mini. STRs: new tool for degraded DNA products work better with low Smaller PCR Conventional PCR primer mini. STR primer copy number or fragmented DNA templates STR repeat region (B) mini. STR primer Conventional PCR primer Conventional STR test (COfiler™ kit) 150 bp smaller Mini. STR assay (using Butler et al. 2003 primers) Figure 7. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(A) mini. STRs: new tool for degraded DNA products work better with low Smaller PCR Conventional PCR primer mini. STR primer copy number or fragmented DNA templates STR repeat region (B) mini. STR primer Conventional PCR primer Conventional STR test (COfiler™ kit) 150 bp smaller Mini. STR assay (using Butler et al. 2003 primers) Figure 7. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 Y-STR
Y-STR
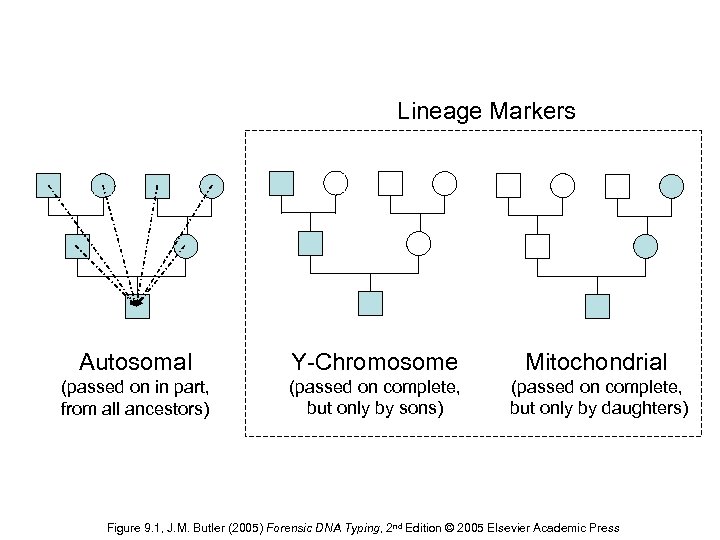 Lineage Markers Autosomal Y-Chromosome Mitochondrial (passed on in part, from all ancestors) (passed on complete, but only by sons) (passed on complete, but only by daughters) Figure 9. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Lineage Markers Autosomal Y-Chromosome Mitochondrial (passed on in part, from all ancestors) (passed on complete, but only by sons) (passed on complete, but only by daughters) Figure 9. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
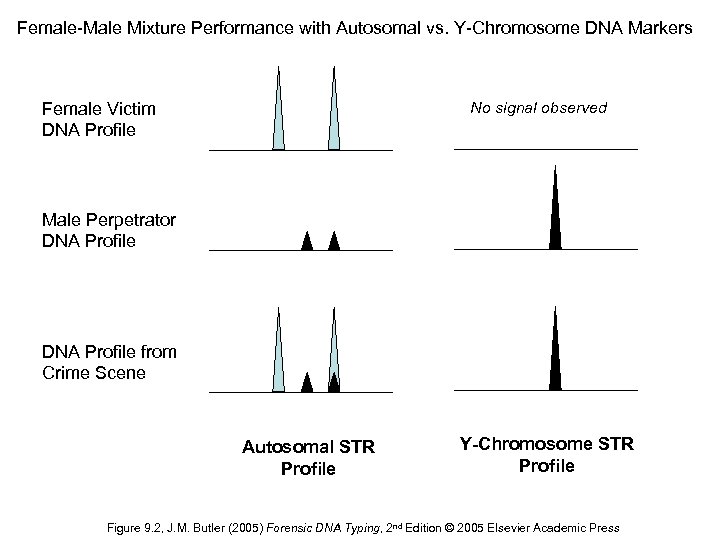 Female-Male Mixture Performance with Autosomal vs. Y-Chromosome DNA Markers No signal observed Female Victim DNA Profile Male Perpetrator DNA Profile from Crime Scene Autosomal STR Profile Y-Chromosome STR Profile Figure 9. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Female-Male Mixture Performance with Autosomal vs. Y-Chromosome DNA Markers No signal observed Female Victim DNA Profile Male Perpetrator DNA Profile from Crime Scene Autosomal STR Profile Y-Chromosome STR Profile Figure 9. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 Modern Use of Y-STR Testing Captured December 13, 2003 Matching Y-STR Haplotype Used to Confirm Identity (along with allele sharing from autosomal STRs) Uday and Qusay Hussein Is this man really Sadaam Hussein? Butler, J. M. (2005) Forensic DNA Typing, 2 nd Edition, Box 23. 1, p. 534 Killed July 22, 2003
Modern Use of Y-STR Testing Captured December 13, 2003 Matching Y-STR Haplotype Used to Confirm Identity (along with allele sharing from autosomal STRs) Uday and Qusay Hussein Is this man really Sadaam Hussein? Butler, J. M. (2005) Forensic DNA Typing, 2 nd Edition, Box 23. 1, p. 534 Killed July 22, 2003
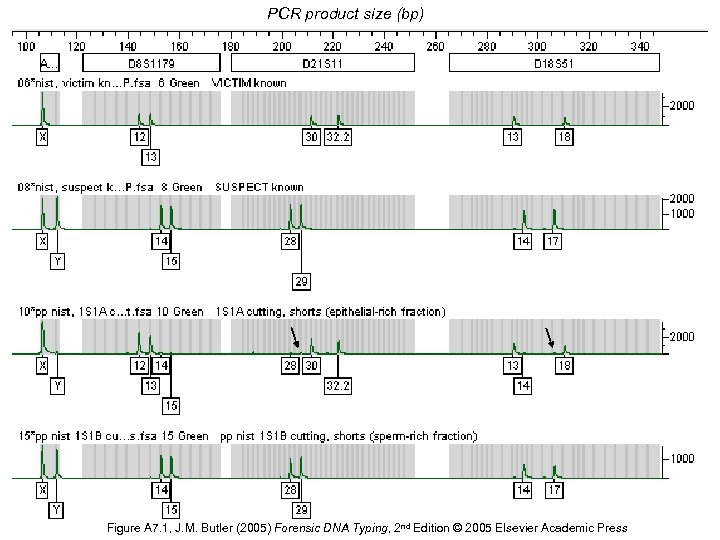 PCR product size (bp) Figure A 7. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
PCR product size (bp) Figure A 7. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
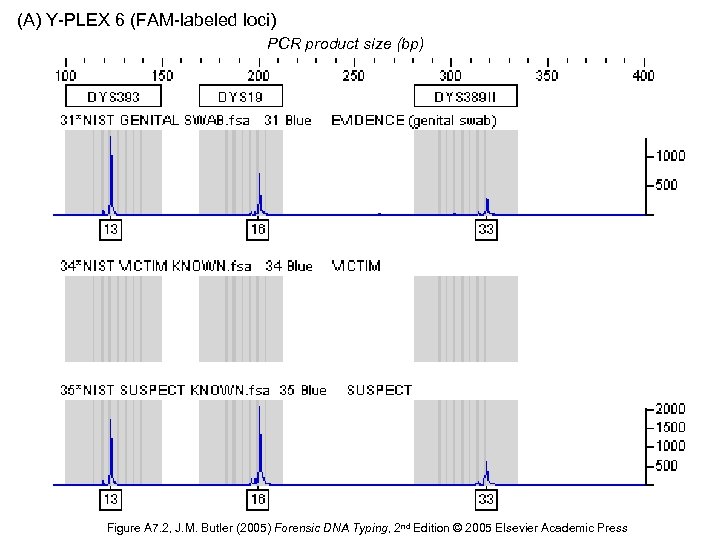 (A) Y-PLEX 6 (FAM-labeled loci) PCR product size (bp) Figure A 7. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(A) Y-PLEX 6 (FAM-labeled loci) PCR product size (bp) Figure A 7. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 ELEKTROFOREZ
ELEKTROFOREZ
 Elektroforez • Nükleik asitler (-) yüke sahitir (PO 4) • Jelde göç etmeleri büyüklükleri ve yapıları ile ilgilidir
Elektroforez • Nükleik asitler (-) yüke sahitir (PO 4) • Jelde göç etmeleri büyüklükleri ve yapıları ile ilgilidir
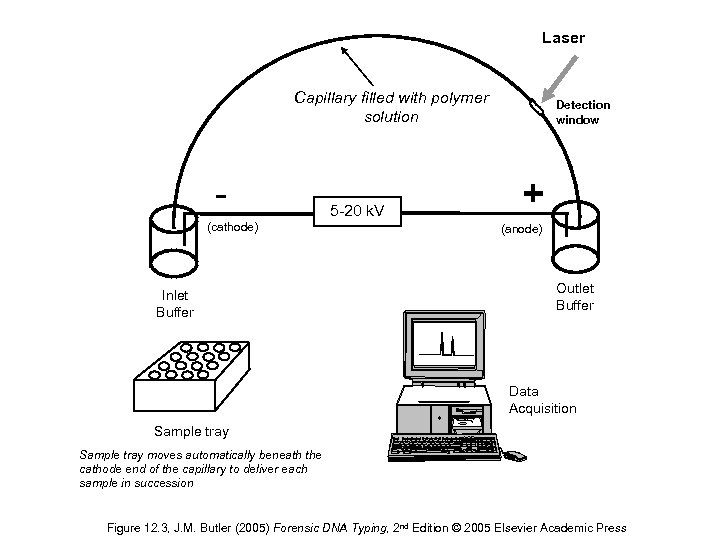 Laser Capillary filled with polymer solution (cathode) Inlet Buffer 5 -20 k. V Detection window + (anode) Outlet Buffer Data Acquisition Sample tray moves automatically beneath the cathode end of the capillary to deliver each sample in succession Figure 12. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Laser Capillary filled with polymer solution (cathode) Inlet Buffer 5 -20 k. V Detection window + (anode) Outlet Buffer Data Acquisition Sample tray moves automatically beneath the cathode end of the capillary to deliver each sample in succession Figure 12. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 CAPİLLER ELECTROPHORESİS
CAPİLLER ELECTROPHORESİS
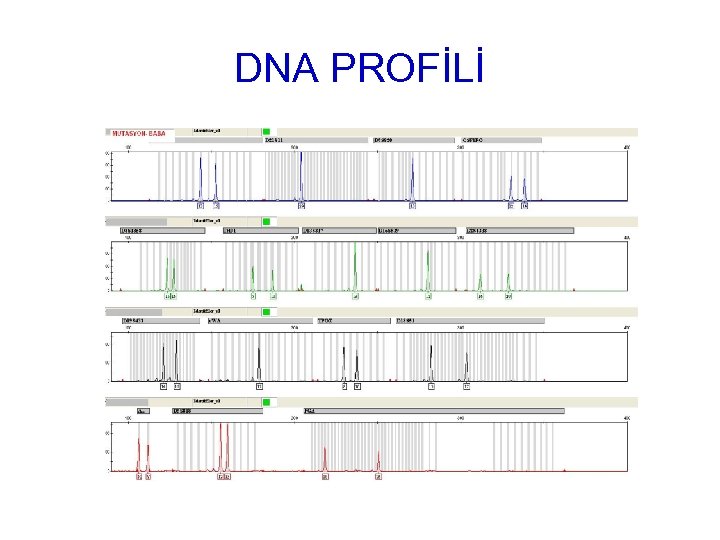 DNA PROFİLİ
DNA PROFİLİ
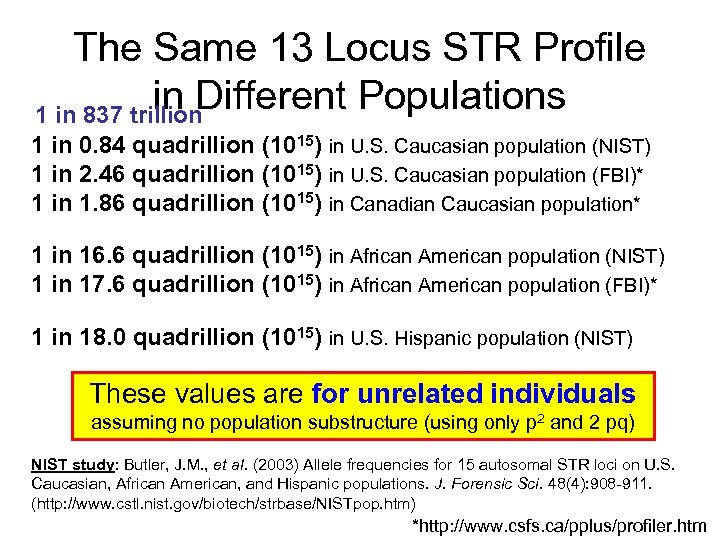 The Same 13 Locus STR Profile in Different Populations 1 in 837 trillion 1 in 0. 84 quadrillion (1015) in U. S. Caucasian population (NIST) 1 in 2. 46 quadrillion (1015) in U. S. Caucasian population (FBI)* 1 in 1. 86 quadrillion (1015) in Canadian Caucasian population* 1 in 16. 6 quadrillion (1015) in African American population (NIST) 1 in 17. 6 quadrillion (1015) in African American population (FBI)* 1 in 18. 0 quadrillion (1015) in U. S. Hispanic population (NIST) These values are for unrelated individuals assuming no population substructure (using only p 2 and 2 pq) NIST study: Butler, J. M. , et al. (2003) Allele frequencies for 15 autosomal STR loci on U. S. Caucasian, African American, and Hispanic populations. J. Forensic Sci. 48(4): 908 -911. (http: //www. cstl. nist. gov/biotech/strbase/NISTpop. htm) *http: //www. csfs. ca/pplus/profiler. htm
The Same 13 Locus STR Profile in Different Populations 1 in 837 trillion 1 in 0. 84 quadrillion (1015) in U. S. Caucasian population (NIST) 1 in 2. 46 quadrillion (1015) in U. S. Caucasian population (FBI)* 1 in 1. 86 quadrillion (1015) in Canadian Caucasian population* 1 in 16. 6 quadrillion (1015) in African American population (NIST) 1 in 17. 6 quadrillion (1015) in African American population (FBI)* 1 in 18. 0 quadrillion (1015) in U. S. Hispanic population (NIST) These values are for unrelated individuals assuming no population substructure (using only p 2 and 2 pq) NIST study: Butler, J. M. , et al. (2003) Allele frequencies for 15 autosomal STR loci on U. S. Caucasian, African American, and Hispanic populations. J. Forensic Sci. 48(4): 908 -911. (http: //www. cstl. nist. gov/biotech/strbase/NISTpop. htm) *http: //www. csfs. ca/pplus/profiler. htm
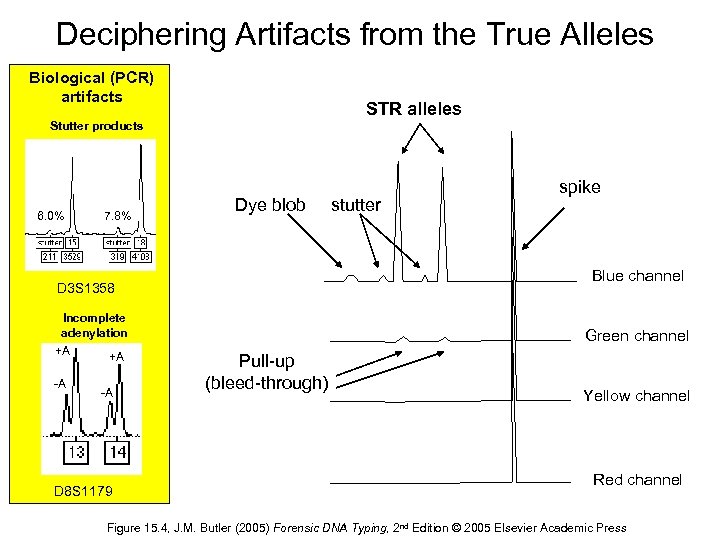 Deciphering Artifacts from the True Alleles Biological (PCR) artifacts STR alleles Stutter products 6. 0% 7. 8% Dye blob Blue channel D 3 S 1358 Incomplete adenylation +A +A -A -A D 8 S 1179 stutter spike Green channel Pull-up (bleed-through) Yellow channel Red channel Figure 15. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Deciphering Artifacts from the True Alleles Biological (PCR) artifacts STR alleles Stutter products 6. 0% 7. 8% Dye blob Blue channel D 3 S 1358 Incomplete adenylation +A +A -A -A D 8 S 1179 stutter spike Green channel Pull-up (bleed-through) Yellow channel Red channel Figure 15. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
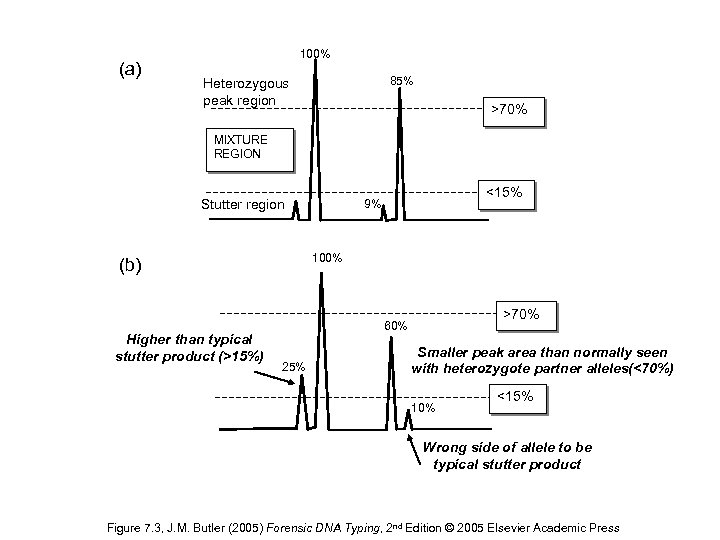 (a) 100% 85% Heterozygous peak region >70% MIXTURE REGION Stutter region 100% (b) Higher than typical stutter product (>15%) <15% 9% >70% 60% 25% Smaller peak area than normally seen with heterozygote partner alleles(<70%) 10% <15% Wrong side of allele to be typical stutter product Figure 7. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(a) 100% 85% Heterozygous peak region >70% MIXTURE REGION Stutter region 100% (b) Higher than typical stutter product (>15%) <15% 9% >70% 60% 25% Smaller peak area than normally seen with heterozygote partner alleles(<70%) 10% <15% Wrong side of allele to be typical stutter product Figure 7. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
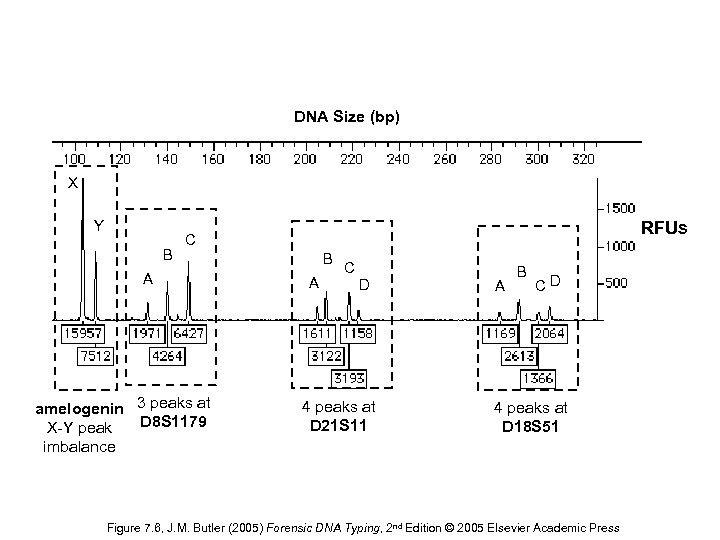 DNA Size (bp) X Y B RFUs C A amelogenin 3 peaks at X-Y peak D 8 S 1179 imbalance B A C D 4 peaks at D 21 S 11 A B CD 4 peaks at D 18 S 51 Figure 7. 6, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
DNA Size (bp) X Y B RFUs C A amelogenin 3 peaks at X-Y peak D 8 S 1179 imbalance B A C D 4 peaks at D 21 S 11 A B CD 4 peaks at D 18 S 51 Figure 7. 6, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 mt. DNA
mt. DNA
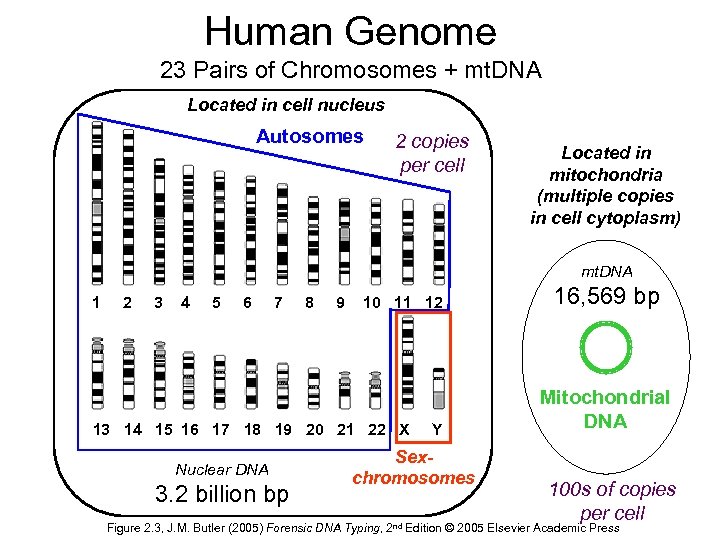 Human Genome 23 Pairs of Chromosomes + mt. DNA Located in cell nucleus http: //www. ncbi. nlm. nih. gov/genome/guide/ Autosomes 2 copies per cell Located in mitochondria (multiple copies in cell cytoplasm) mt. DNA 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 X Nuclear DNA 3. 2 billion bp Y Sexchromosomes 16, 569 bp Mitochondrial DNA 100 s of copies per cell Figure 2. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Human Genome 23 Pairs of Chromosomes + mt. DNA Located in cell nucleus http: //www. ncbi. nlm. nih. gov/genome/guide/ Autosomes 2 copies per cell Located in mitochondria (multiple copies in cell cytoplasm) mt. DNA 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 X Nuclear DNA 3. 2 billion bp Y Sexchromosomes 16, 569 bp Mitochondrial DNA 100 s of copies per cell Figure 2. 3, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
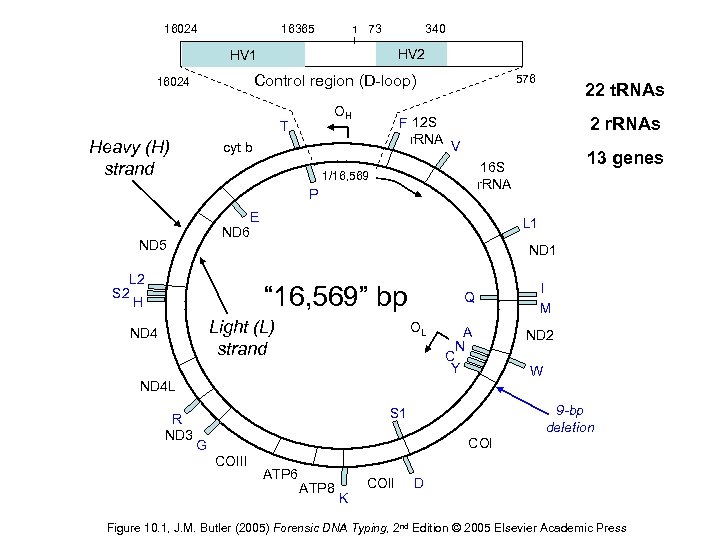 16024 16365 1 73 340 HV 2 HV 1 Control region (D-loop) 16024 OH T Heavy (H) strand F 12 S r. RNA cyt b 576 2 r. RNAs V P E ND 6 L 1 ND 1 L 2 S 2 H Q I M A N ND 2 “ 16, 569” bp Light (L) strand ND 4 OL C Y W ND 4 L R ND 3 13 genes 16 S r. RNA 1/16, 569 ND 5 22 t. RNAs 9 -bp deletion S 1 COI G COIII ATP 6 ATP 8 K COII D Figure 10. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
16024 16365 1 73 340 HV 2 HV 1 Control region (D-loop) 16024 OH T Heavy (H) strand F 12 S r. RNA cyt b 576 2 r. RNAs V P E ND 6 L 1 ND 1 L 2 S 2 H Q I M A N ND 2 “ 16, 569” bp Light (L) strand ND 4 OL C Y W ND 4 L R ND 3 13 genes 16 S r. RNA 1/16, 569 ND 5 22 t. RNAs 9 -bp deletion S 1 COI G COIII ATP 6 ATP 8 K COII D Figure 10. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 DNA DİZİ ANALİZİ
DNA DİZİ ANALİZİ
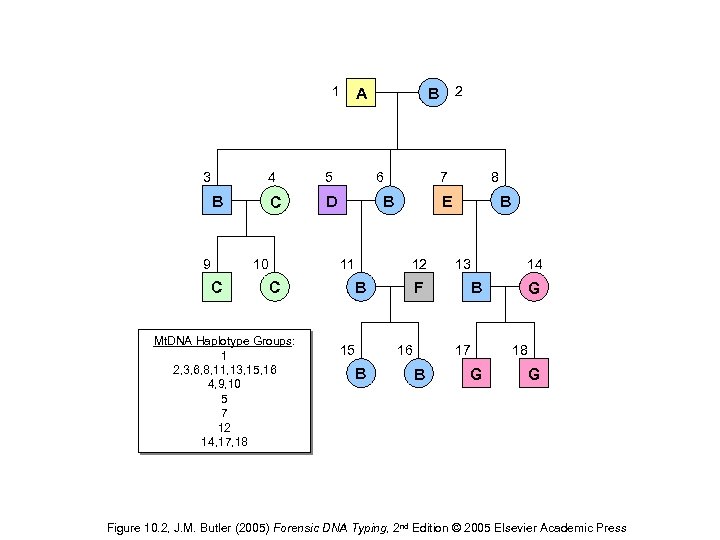 1 3 4 5 C D B 9 10 C 6 Mt. DNA Haplotype Groups: 1 2, 3, 6, 8, 11, 13, 15, 16 4, 9, 10 5 7 12 14, 17, 18 7 B 11 C B 2 A E 12 B 15 B 8 B 13 B F 16 14 17 B G 18 G G Figure 10. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
1 3 4 5 C D B 9 10 C 6 Mt. DNA Haplotype Groups: 1 2, 3, 6, 8, 11, 13, 15, 16 4, 9, 10 5 7 12 14, 17, 18 7 B 11 C B 2 A E 12 B 15 B 8 B 13 B F 16 14 17 B G 18 G G Figure 10. 2, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
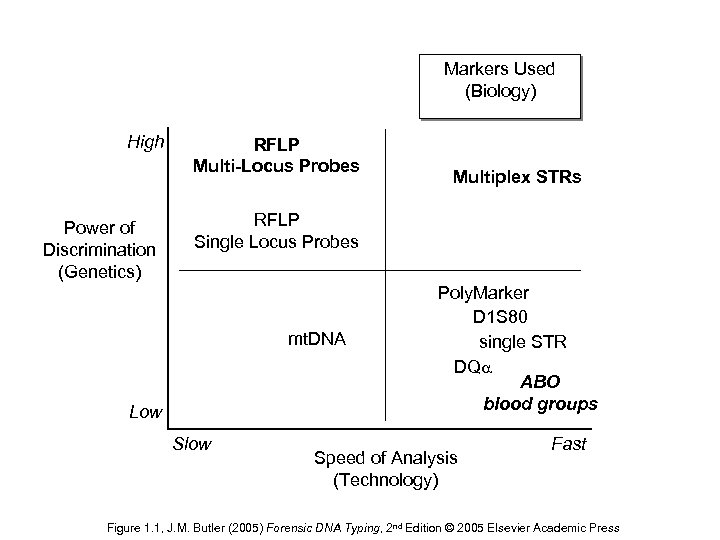 Markers Used (Biology) High Power of Discrimination (Genetics) RFLP Multi-Locus Probes Multiplex STRs RFLP Single Locus Probes mt. DNA Low Slow Poly. Marker D 1 S 80 single STR DQ ABO blood groups Speed of Analysis (Technology) Fast Figure 1. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Markers Used (Biology) High Power of Discrimination (Genetics) RFLP Multi-Locus Probes Multiplex STRs RFLP Single Locus Probes mt. DNA Low Slow Poly. Marker D 1 S 80 single STR DQ ABO blood groups Speed of Analysis (Technology) Fast Figure 1. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
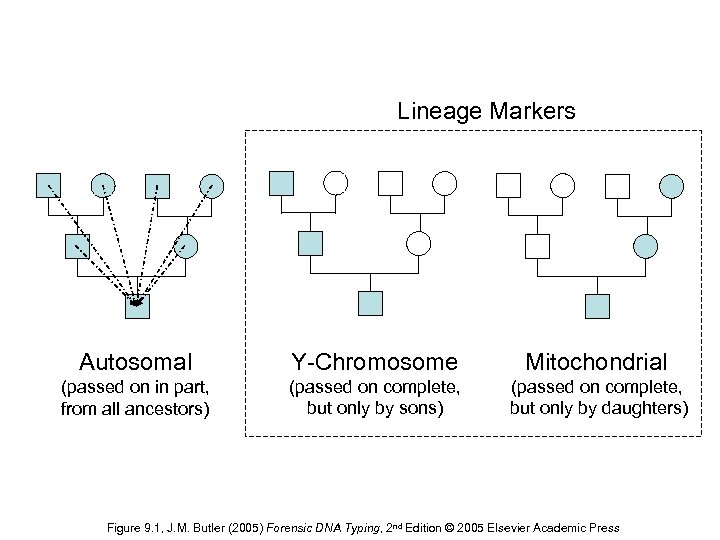 Lineage Markers Autosomal Y-Chromosome Mitochondrial (passed on in part, from all ancestors) (passed on complete, but only by sons) (passed on complete, but only by daughters) Figure 9. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Lineage Markers Autosomal Y-Chromosome Mitochondrial (passed on in part, from all ancestors) (passed on complete, but only by sons) (passed on complete, but only by daughters) Figure 9. 1, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
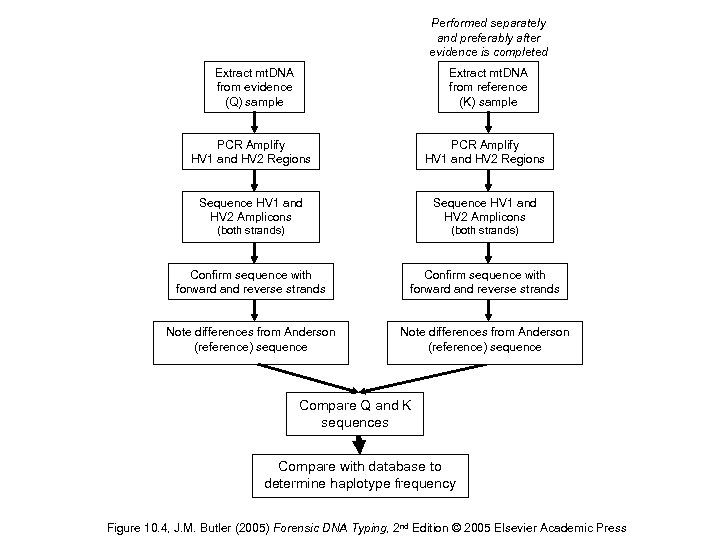 Performed separately and preferably after evidence is completed Extract mt. DNA from evidence (Q) sample Extract mt. DNA from reference (K) sample PCR Amplify HV 1 and HV 2 Regions Sequence HV 1 and HV 2 Amplicons (both strands) Confirm sequence with forward and reverse strands Note differences from Anderson (reference) sequence Compare Q and K sequences Compare with database to determine haplotype frequency Figure 10. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
Performed separately and preferably after evidence is completed Extract mt. DNA from evidence (Q) sample Extract mt. DNA from reference (K) sample PCR Amplify HV 1 and HV 2 Regions Sequence HV 1 and HV 2 Amplicons (both strands) Confirm sequence with forward and reverse strands Note differences from Anderson (reference) sequence Compare Q and K sequences Compare with database to determine haplotype frequency Figure 10. 4, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
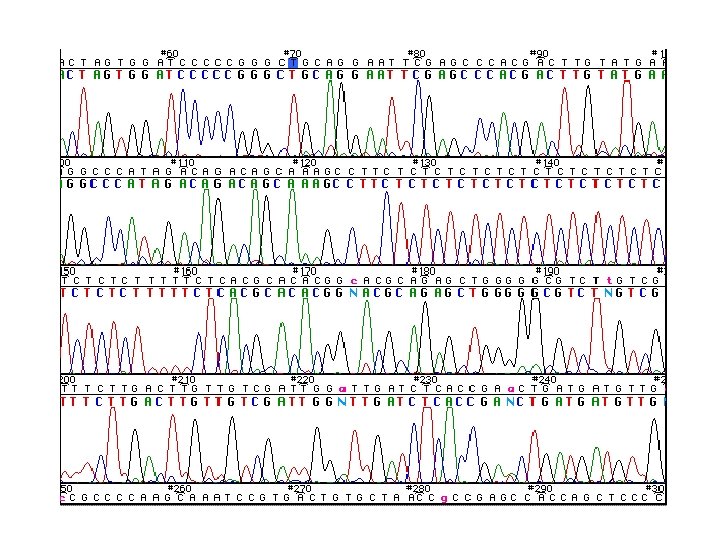
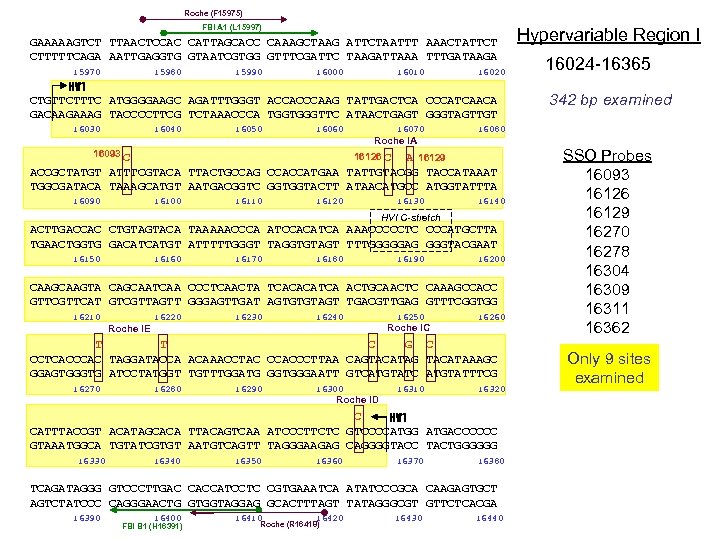 Roche (F 15975) FBI A 1 (L 15997) GAAAAAGTCT TTAACTCCAC CATTAGCACC CAAAGCTAAG ATTCTAATTT AAACTATTCT CTTTTTCAGA AATTGAGGTG GTAATCGTGG GTTTCGATTC TAAGATTAAA TTTGATAAGA 15970 15980 15990 16000 16010 16020 HV 1 CTGTTCTTTC ATGGGGAAGC AGATTTGGGT ACCACCCAAG TATTGACTCA CCCATCAACA GACAAGAAAG TACCCCTTCG TCTAAACCCA TGGTGGGTTC ATAACTGAGT GGGTAGTTGT 16030 16040 16050 16060 16070 16093 C 16126 C A 16129 ACCGCTATGT ATTTCGTACA TTACTGCCAG CCACCATGAA TATTGTACGG TACCATAAAT TGGCGATACA TAAAGCATGT AATGACGGTC GGTGGTACTT ATAACATGCC ATGGTATTTA 16100 16110 16120 16130 16140 HVI C-stretch ACTTGACCAC CTGTAGTACA TAAAAACCCA ATCCACATCA AAACCCCCTC CCCATGCTTA TGAACTGGTG GACATCATGT ATTTTTGGGT TAGGTGTAGT TTTGGGGGAG GGGTACGAAT 16150 16160 16170 16180 16190 16200 CAAGTA CAGCAATCAA CCCTCAACTA TCACACATCA ACTGCAACTC CAAAGCCACC GTTCAT GTCGTTAGTT GGGAGTTGAT AGTGTGTAGT TGACGTTGAG GTTTCGGTGG 16210 16220 16230 16240 Roche IE 16250 Roche IC 16260 T T C G C CCTCACCCAC TAGGATACCA ACAAACCTAC CCACCCTTAA CAGTACATAG TACATAAAGC GGAGTGGGTG ATCCTATGGT TGTTTGGATG GGTGGGAATT GTCATGTATC ATGTATTTCG 16270 16280 16290 16300 Roche ID 16310 16320 C HV 1 CATTTACCGT ACATAGCACA TTACAGTCAA ATCCCTTCTC GTCCCCATGG ATGACCCCCC GTAAATGGCA TGTATCGTGT AATGTCAGTT TAGGGAAGAG CAGGGGTACC TACTGGGGGG 16330 16340 16350 16360 16370 16380 TCAGATAGGG GTCCCTTGAC CACCATCCTC CGTGAAATCA ATATCCCGCA CAAGAGTGCT AGTCTATCCC CAGGGAACTG GTGGTAGGAG GCACTTTAGT TATAGGGCGT GTTCTCACGA 16390 16400 FBI B 1 (H 16391) 16410 16420 Roche (R 16418) 16430 16024 -16365 342 bp examined 16080 Roche IA 16090 Hypervariable Region I 16440 SSO Probes 16093 16126 16129 16270 16278 16304 16309 16311 16362 Only 9 sites examined
Roche (F 15975) FBI A 1 (L 15997) GAAAAAGTCT TTAACTCCAC CATTAGCACC CAAAGCTAAG ATTCTAATTT AAACTATTCT CTTTTTCAGA AATTGAGGTG GTAATCGTGG GTTTCGATTC TAAGATTAAA TTTGATAAGA 15970 15980 15990 16000 16010 16020 HV 1 CTGTTCTTTC ATGGGGAAGC AGATTTGGGT ACCACCCAAG TATTGACTCA CCCATCAACA GACAAGAAAG TACCCCTTCG TCTAAACCCA TGGTGGGTTC ATAACTGAGT GGGTAGTTGT 16030 16040 16050 16060 16070 16093 C 16126 C A 16129 ACCGCTATGT ATTTCGTACA TTACTGCCAG CCACCATGAA TATTGTACGG TACCATAAAT TGGCGATACA TAAAGCATGT AATGACGGTC GGTGGTACTT ATAACATGCC ATGGTATTTA 16100 16110 16120 16130 16140 HVI C-stretch ACTTGACCAC CTGTAGTACA TAAAAACCCA ATCCACATCA AAACCCCCTC CCCATGCTTA TGAACTGGTG GACATCATGT ATTTTTGGGT TAGGTGTAGT TTTGGGGGAG GGGTACGAAT 16150 16160 16170 16180 16190 16200 CAAGTA CAGCAATCAA CCCTCAACTA TCACACATCA ACTGCAACTC CAAAGCCACC GTTCAT GTCGTTAGTT GGGAGTTGAT AGTGTGTAGT TGACGTTGAG GTTTCGGTGG 16210 16220 16230 16240 Roche IE 16250 Roche IC 16260 T T C G C CCTCACCCAC TAGGATACCA ACAAACCTAC CCACCCTTAA CAGTACATAG TACATAAAGC GGAGTGGGTG ATCCTATGGT TGTTTGGATG GGTGGGAATT GTCATGTATC ATGTATTTCG 16270 16280 16290 16300 Roche ID 16310 16320 C HV 1 CATTTACCGT ACATAGCACA TTACAGTCAA ATCCCTTCTC GTCCCCATGG ATGACCCCCC GTAAATGGCA TGTATCGTGT AATGTCAGTT TAGGGAAGAG CAGGGGTACC TACTGGGGGG 16330 16340 16350 16360 16370 16380 TCAGATAGGG GTCCCTTGAC CACCATCCTC CGTGAAATCA ATATCCCGCA CAAGAGTGCT AGTCTATCCC CAGGGAACTG GTGGTAGGAG GCACTTTAGT TATAGGGCGT GTTCTCACGA 16390 16400 FBI B 1 (H 16391) 16410 16420 Roche (R 16418) 16430 16024 -16365 342 bp examined 16080 Roche IA 16090 Hypervariable Region I 16440 SSO Probes 16093 16126 16129 16270 16278 16304 16309 16311 16362 Only 9 sites examined
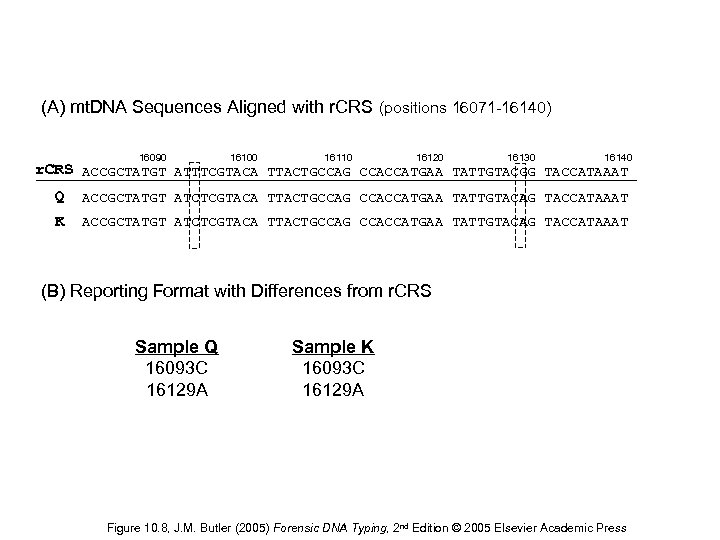 (A) mt. DNA Sequences Aligned with r. CRS (positions 16071 -16140) 16090 16100 16110 16120 16130 16140 r. CRS ACCGCTATGT ATTTCGTACA TTACTGCCAG CCACCATGAA TATTGTACGG TACCATAAAT Q ACCGCTATGT ATCTCGTACA TTACTGCCAG CCACCATGAA TATTGTACAG TACCATAAAT K ACCGCTATGT ATCTCGTACA TTACTGCCAG CCACCATGAA TATTGTACAG TACCATAAAT (B) Reporting Format with Differences from r. CRS Sample Q 16093 C 16129 A Sample K 16093 C 16129 A Figure 10. 8, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
(A) mt. DNA Sequences Aligned with r. CRS (positions 16071 -16140) 16090 16100 16110 16120 16130 16140 r. CRS ACCGCTATGT ATTTCGTACA TTACTGCCAG CCACCATGAA TATTGTACGG TACCATAAAT Q ACCGCTATGT ATCTCGTACA TTACTGCCAG CCACCATGAA TATTGTACAG TACCATAAAT K ACCGCTATGT ATCTCGTACA TTACTGCCAG CCACCATGAA TATTGTACAG TACCATAAAT (B) Reporting Format with Differences from r. CRS Sample Q 16093 C 16129 A Sample K 16093 C 16129 A Figure 10. 8, J. M. Butler (2005) Forensic DNA Typing, 2 nd Edition © 2005 Elsevier Academic Press
 • İnsan genomu 3, 164, 700, 000 nukleotidden oluşmaktadır. • Toplam gen sayısı 29, 000 -36, 000 arasındadır. • Nükleotid dizilerinin %99’u bütün insanlarda aynıdır.
• İnsan genomu 3, 164, 700, 000 nukleotidden oluşmaktadır. • Toplam gen sayısı 29, 000 -36, 000 arasındadır. • Nükleotid dizilerinin %99’u bütün insanlarda aynıdır.
 • Bu güne kadar insanda 1, 5 milyon kadar tek nukleotid değişikliği bölgesi saptanmıştır. • Tanımlanmış genlerin %50’den fazlasının işlevleri henüz bilinmemektedir. • Genomun yaklaşık %2’si proteinleri kodlamaktadır. • Proteinleri kodlamayan dizi tekrarları, genomun büyük bölümünü oluşturur.
• Bu güne kadar insanda 1, 5 milyon kadar tek nukleotid değişikliği bölgesi saptanmıştır. • Tanımlanmış genlerin %50’den fazlasının işlevleri henüz bilinmemektedir. • Genomun yaklaşık %2’si proteinleri kodlamaktadır. • Proteinleri kodlamayan dizi tekrarları, genomun büyük bölümünü oluşturur.
 İnsan Genomundan Beklentilerimiz • Moleküler Tıp -Tanı yöntemlerinin geliştirilmesi - Hastalıklara genetik yatkınlığın belirlenmesi - Genetik yapıya özgü ilaçlar geliştirilmesi - Gen tedavisi yöntemlerinin geliştirilmesi
İnsan Genomundan Beklentilerimiz • Moleküler Tıp -Tanı yöntemlerinin geliştirilmesi - Hastalıklara genetik yatkınlığın belirlenmesi - Genetik yapıya özgü ilaçlar geliştirilmesi - Gen tedavisi yöntemlerinin geliştirilmesi
 • Biyoarkeoloji, Antropoloji ve Tarih - Değişik toplumların göç yollarının ve akrabalıklarının araştırılması - Y kromozom mutasyonlarının incelenmesiyle erkek dağılımının ve göçlerin araştırılması
• Biyoarkeoloji, Antropoloji ve Tarih - Değişik toplumların göç yollarının ve akrabalıklarının araştırılması - Y kromozom mutasyonlarının incelenmesiyle erkek dağılımının ve göçlerin araştırılması
 • DNA Tanımlama - Adli tıpta suçluların belirlenmesi - Kan bağlarının saptanması - Organ nakillerinde doku uyumunun kesin şekilde saptanması - Soy ağaçlarının geliştirilmesi
• DNA Tanımlama - Adli tıpta suçluların belirlenmesi - Kan bağlarının saptanması - Organ nakillerinde doku uyumunun kesin şekilde saptanması - Soy ağaçlarının geliştirilmesi
 Kuşkular • Genetik Bilginin Özelliği ve Gizliliği - Genetik bilgiye kim sahip olacak ve kontrol edecek? - Genetik bilgilerin gizliliği tıbbi gizlilikten farklı mı? • Genetik bilginin kullanılması - Bireye ait genetik bilgilere kim ulaşabilecek ve bu bilgileri nasıl kullanacak?
Kuşkular • Genetik Bilginin Özelliği ve Gizliliği - Genetik bilgiye kim sahip olacak ve kontrol edecek? - Genetik bilgilerin gizliliği tıbbi gizlilikten farklı mı? • Genetik bilginin kullanılması - Bireye ait genetik bilgilere kim ulaşabilecek ve bu bilgileri nasıl kullanacak?
 SABRINIZ İÇİN TEŞEKKÜRLER
SABRINIZ İÇİN TEŞEKKÜRLER


