ef2b78476982c56115352e2a9cc65fda.ppt
- Количество слайдов: 38

Unorthodox mitochondrial genomes Gertraud Burger Robert-Cedergren Center for Bioinformatics and Genomics Université de Montreal QC, Canada 1
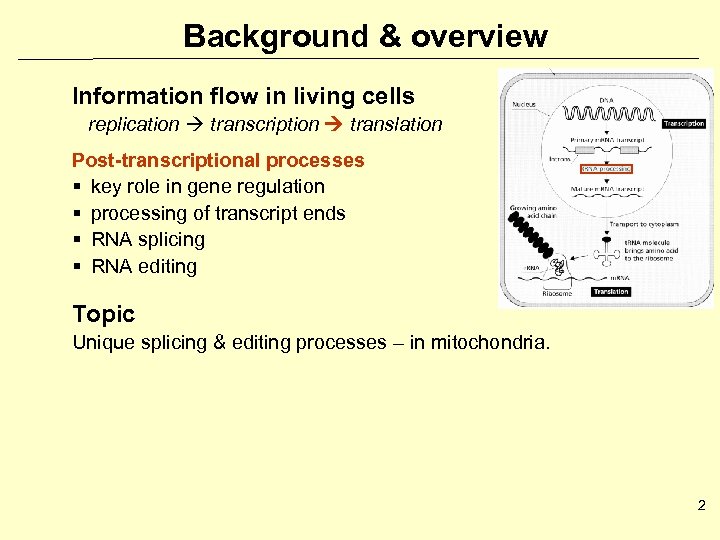
Background & overview Information flow in living cells replication transcription translation Post-transcriptional processes § key role in gene regulation § processing of transcript ends § RNA splicing § RNA editing Topic Unique splicing & editing processes – in mitochondria. 2

Outline of the talk § Current knowledge of mt. DNAs across eukaryotes § Our study in a eukaryotic microbe § Evolutionary implications. 3

Who did the work Shona Teijeiro Yifei Yan Georgette Kiethega 4
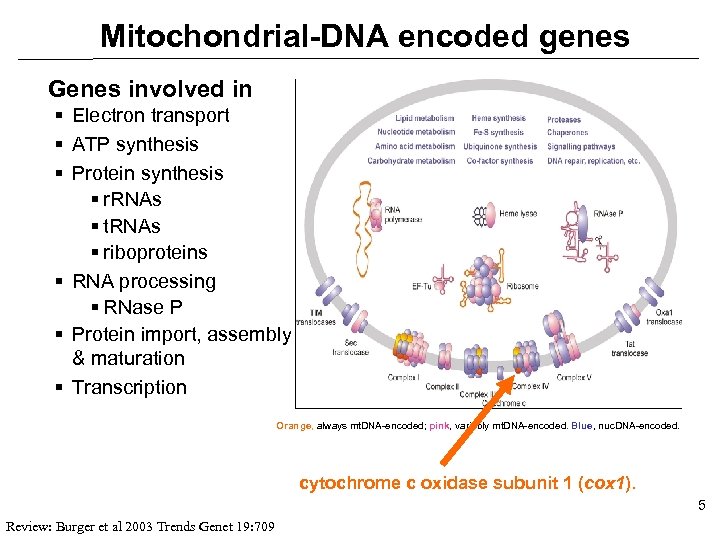
Mitochondrial-DNA encoded genes Genes involved in § Electron transport § ATP synthesis § Protein synthesis § r. RNAs § t. RNAs § riboproteins § RNA processing § RNase P § Protein import, assembly & maturation § Transcription Orange, always mt. DNA-encoded; pink, variably mt. DNA-encoded. Blue, nuc. DNA-encoded. cytochrome c oxidase subunit 1 (cox 1). 5 Review: Burger et al 2003 Trends Genet 19: 709
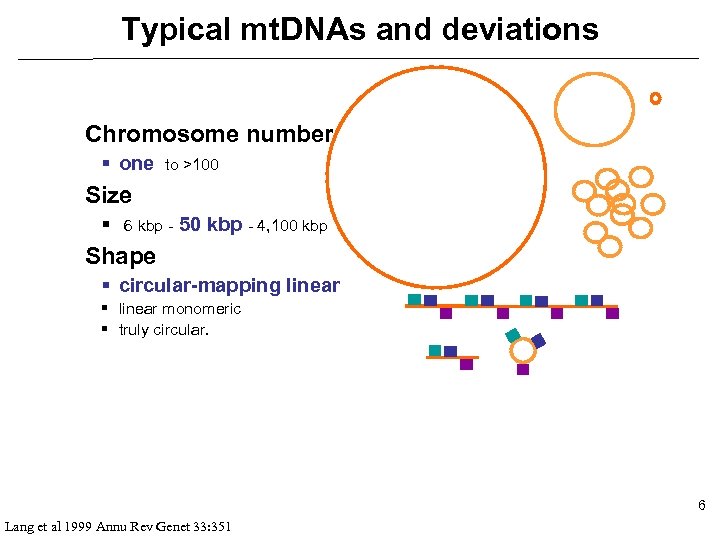
Typical mt. DNAs and deviations Chromosome number § one to >100 Size § 6 kbp - 50 kbp - 4, 100 kbp Shape § circular-mapping linear § linear monomeric § truly circular. 6 Lang et al 1999 Annu Rev Genet 33: 351
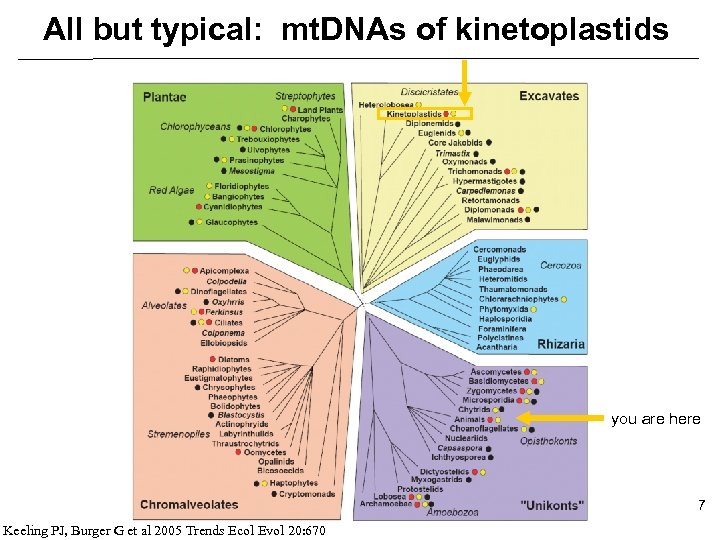
All but typical: mt. DNAs of kinetoplastids you are here 7 Keeling PJ, Burger G et al 2005 Trends Ecol Evol 20: 670
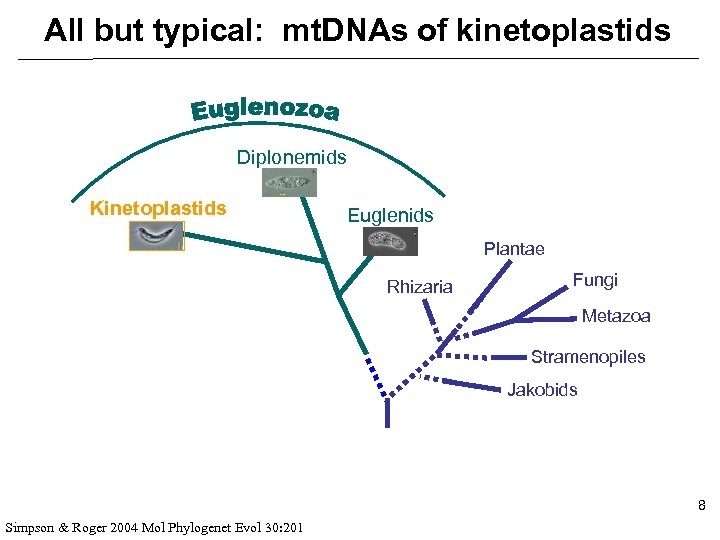
All but typical: mt. DNAs of kinetoplastids Diplonemids Kinetoplastids Euglenids Plantae Rhizaria Fungi Metazoa Stramenopiles Jakobids 8 Simpson & Roger 2004 Mol Phylogenet Evol 30: 201
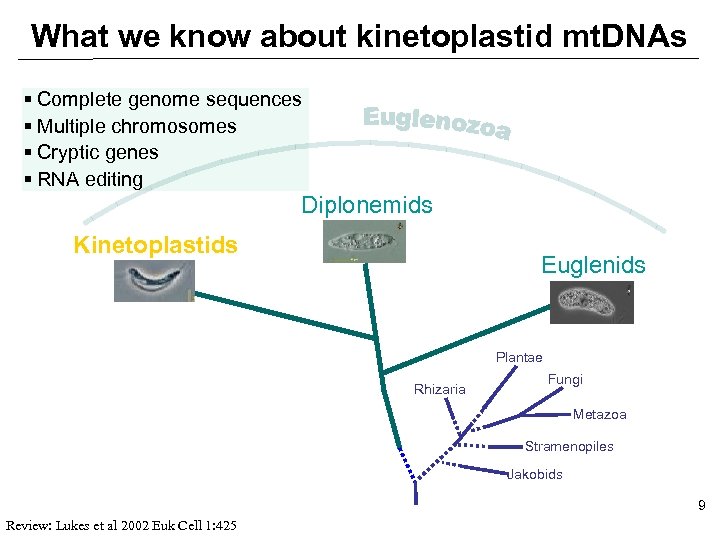
What we know about kinetoplastid mt. DNAs § Complete genome sequences § Multiple chromosomes § Cryptic genes § RNA editing Diplonemids Kinetoplastids Euglenids Plantae Rhizaria Fungi Metazoa Stramenopiles Jakobids 9 Review: Lukes et al 2002 Euk Cell 1: 425
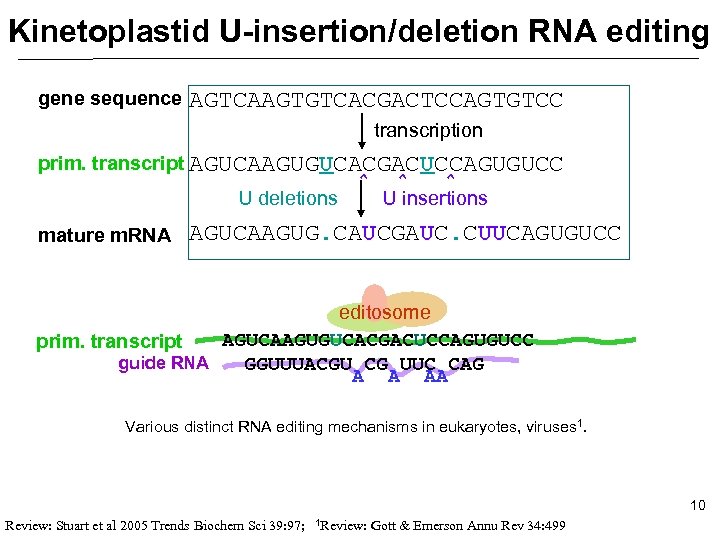
Kinetoplastid U-insertion/deletion RNA editing gene sequence AGTCAAGTGTCACGACTCCAGTGTCC transcription prim. transcript AGUCAAGUGUCACGACUCCAGUGUCC U deletions mature m. RNA ^ ^ ^ U insertions AGUCAAGUG. CAUCGAUC. CUUCAGUGUCC editosome AGUCAAGUGUCACGACUCCAGUGUCC prim. transcript guide RNA GGUUUACGU CG UUC CAG A A AA Various distinct RNA editing mechanisms in eukaryotes, viruses 1. 10 Review: Stuart et al 2005 Trends Biochem Sci 39: 97; 1 Review: Gott & Emerson Annu Rev 34: 499
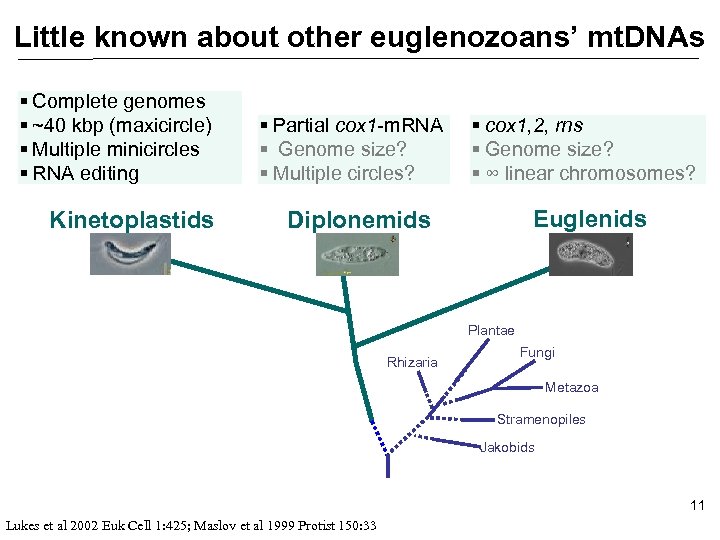
Little known about other euglenozoans’ mt. DNAs § Complete genomes § ~40 kbp (maxicircle) § Multiple minicircles § RNA editing Kinetoplastids § Partial cox 1 -m. RNA § Genome size? § Multiple circles? § cox 1, 2, rns § Genome size? § ∞ linear chromosomes? Euglenids Diplonemids Plantae Rhizaria Fungi Metazoa Stramenopiles Jakobids 11 Lukes et al 2002 Euk Cell 1: 425; Maslov et al 1999 Protist 150: 33

Our initial questions Are kinetoplastids the only Euglenozoa with unconventional mt. DNA and RNA editing? When did these unusual features emerge? . 12
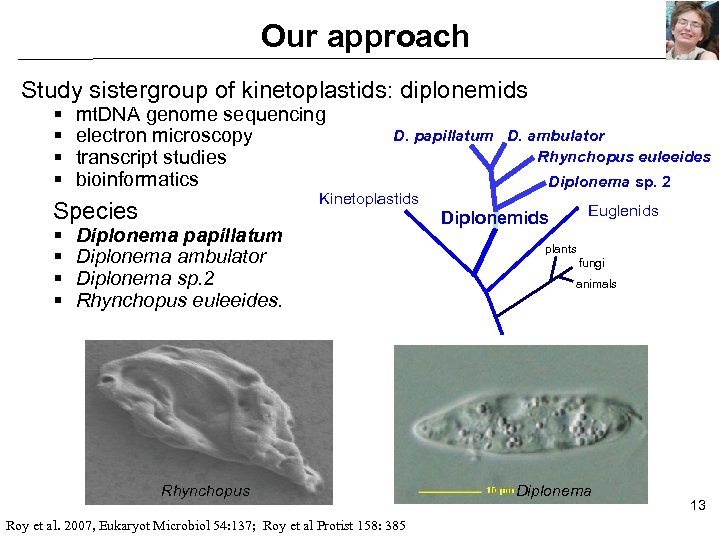
Our approach Study sistergroup of kinetoplastids: diplonemids § § mt. DNA genome sequencing electron microscopy transcript studies bioinformatics Kinetoplastids Species § § D. papillatum D. ambulator Rhynchopus euleeides Diplonema papillatum Diplonema ambulator Diplonema sp. 2 Rhynchopus euleeides. Rhynchopus Roy et al. 2007, Eukaryot Microbiol 54: 137; Roy et al Protist 158: 385 Diplonema sp. 2 Euglenids Diplonemids plants fungi animals Diplonema 13
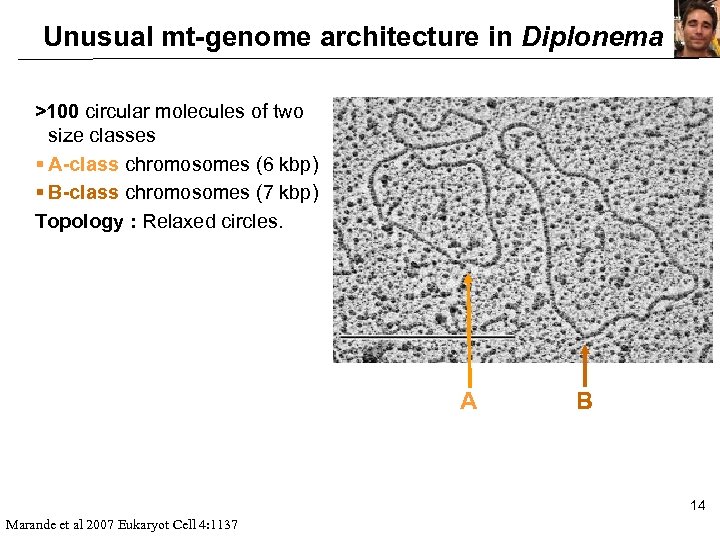
Unusual mt-genome architecture in Diplonema >100 circular molecules of two size classes § A-class chromosomes (6 kbp) § B-class chromosomes (7 kbp) Topology : Relaxed circles. A B 14 Marande et al 2007 Eukaryot Cell 4: 1137
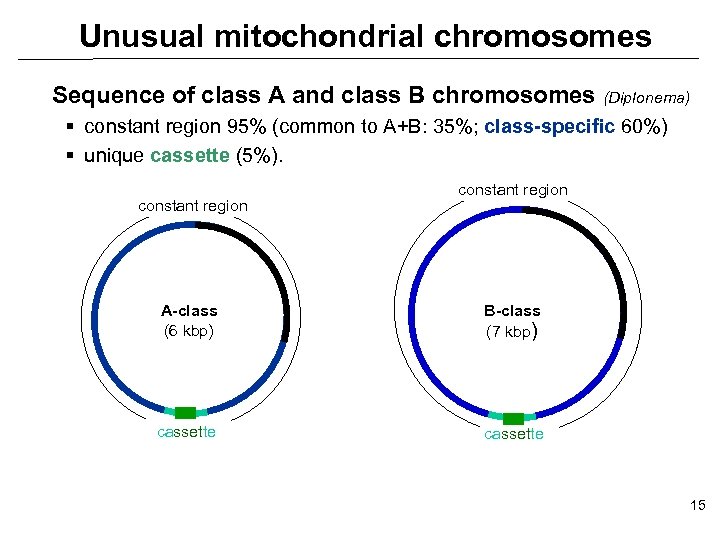
Unusual mitochondrial chromosomes Sequence of class A and class B chromosomes (Diplonema) § constant region 95% (common to A+B: 35%; class-specific 60%) § unique cassette (5%). constant region A-class (6 kbp) B-class (7 kbp) cassette 15
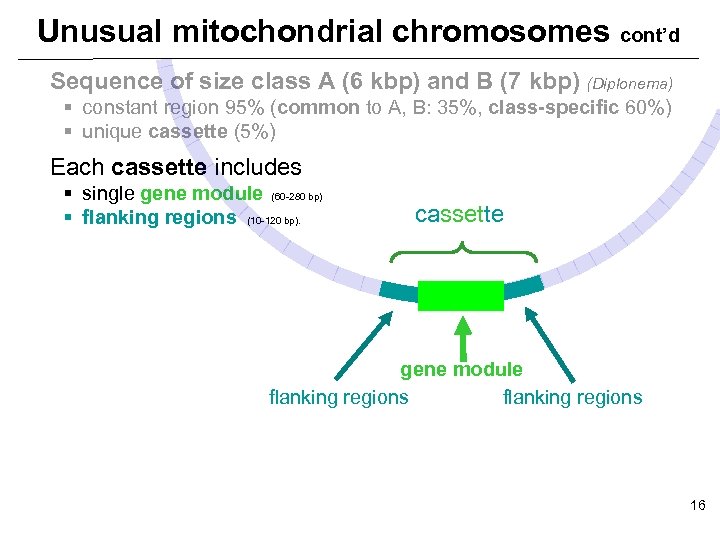
Unusual mitochondrial chromosomes cont’d Sequence of size class A (6 kbp) and B (7 kbp) (Diplonema) § constant region 95% (common to A, B: 35%, class-specific 60%) § unique cassette (5%) Each cassette includes § single gene module (60 -280 bp) § flanking regions (10 -120 bp). cassette gene module flanking regions 16

Summary 1: Diplonema mt. DNA Unusual genome architecture § ~100 chromosomes with different gene modules ~650 kbp the least compact mt. DNA known. 17
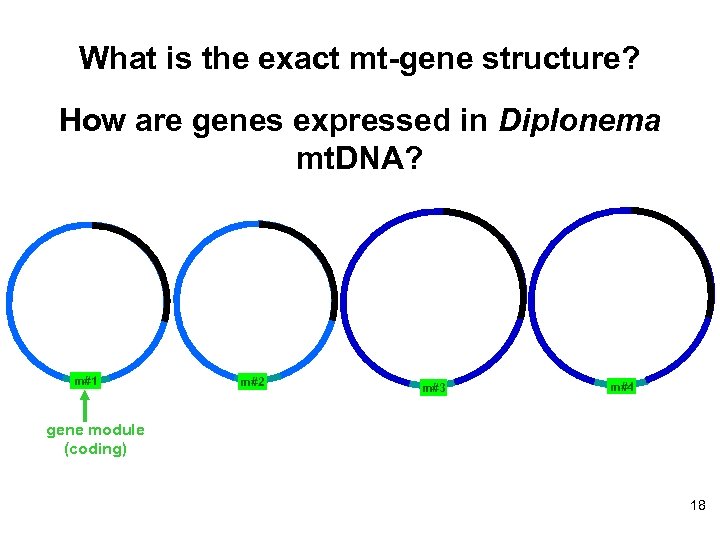
What is the exact mt-gene structure? How are genes expressed in Diplonema mt. DNA? m#1 m#2 m#3 m#4 gene module (coding) 18
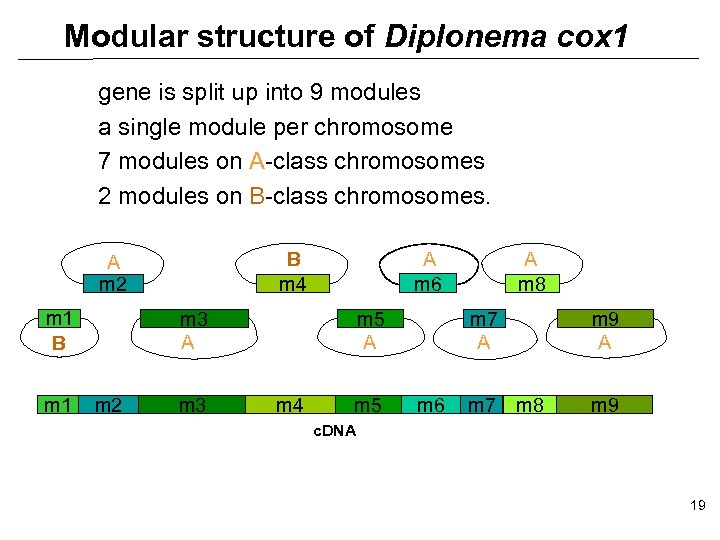
Modular structure of Diplonema cox 1 gene is split up into 9 modules a single module per chromosome 7 modules on A-class chromosomes 2 modules on B-class chromosomes. m 1 B m 1 A m 6 B m 4 A m 2 m 3 m 5 A m 4 m 5 A m 8 m 7 A m 6 m 7 m 9 A m 8 m 9 c. DNA 19
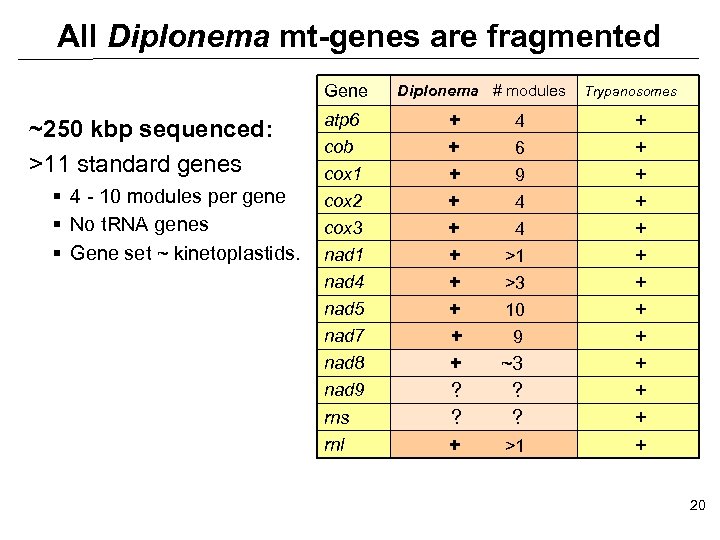
All Diplonema mt-genes are fragmented Gene ~250 kbp sequenced: >11 standard genes § 4 - 10 modules per gene § No t. RNA genes § Gene set ~ kinetoplastids. atp 6 cob cox 1 cox 2 cox 3 nad 1 nad 4 nad 5 nad 7 nad 8 nad 9 rns rnl Diplonema # modules + + + + + ? ? + 4 6 9 4 4 >1 >3 10 9 ~3 ? ? >1 Trypanosomes + + + + 20
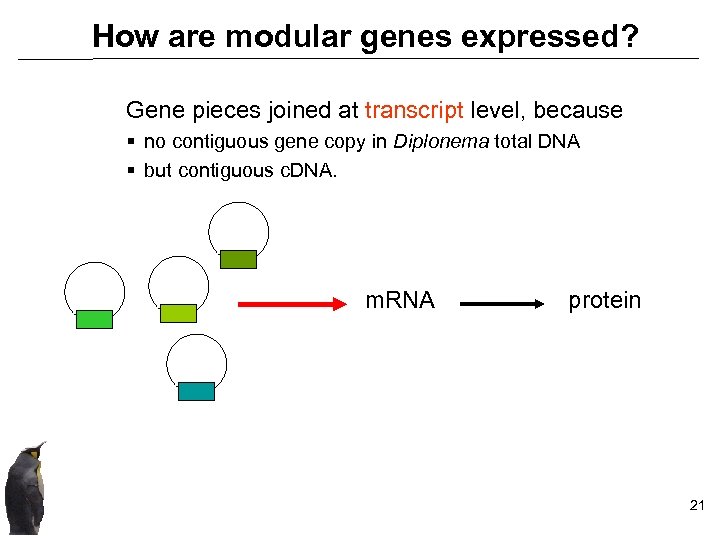
How are modular genes expressed? Gene pieces joined at transcript level, because § no contiguous gene copy in Diplonema total DNA § but contiguous c. DNA. m. RNA protein 21
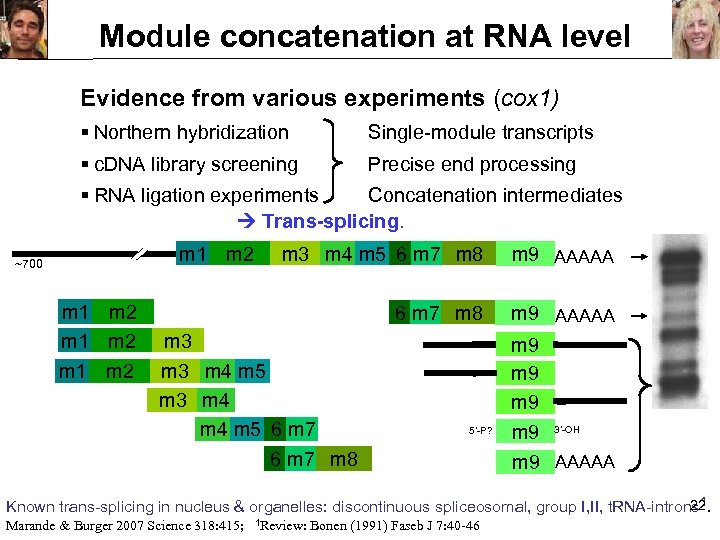
Module concatenation at RNA level Evidence from various experiments (cox 1) § Northern hybridization Single-module transcripts § c. DNA library screening Precise end processing § RNA ligation experiments Concatenation intermediates Trans-splicing. m 1 m 2 ~700 m 1 m 2 m 3 m 4 m 5 6 m 7 m 8 m 9 AAAAA m 9 m 9 3’-OH m 3 m 4 m 5 6 m 7 m 8 5’-P? m 9 AAAAA 22 Known trans-splicing in nucleus & organelles: discontinuous spliceosomal, group I, II, t. RNA-introns 1. Marande & Burger 2007 Science 318: 415; 1 Review: Bonen (1991) Faseb J 7: 40 -46
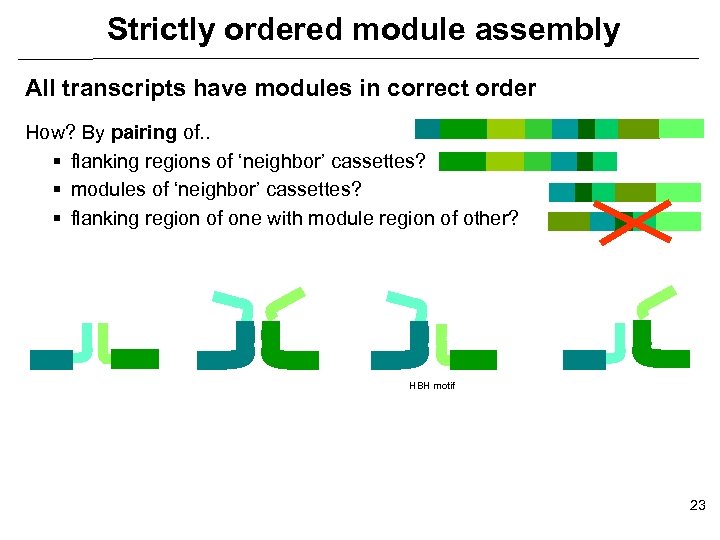
Strictly ordered module assembly All transcripts have modules in correct order How? By pairing of. . § flanking regions of ‘neighbor’ cassettes? § modules of ‘neighbor’ cassettes? § flanking region of one with module region of other? HBH motif 23
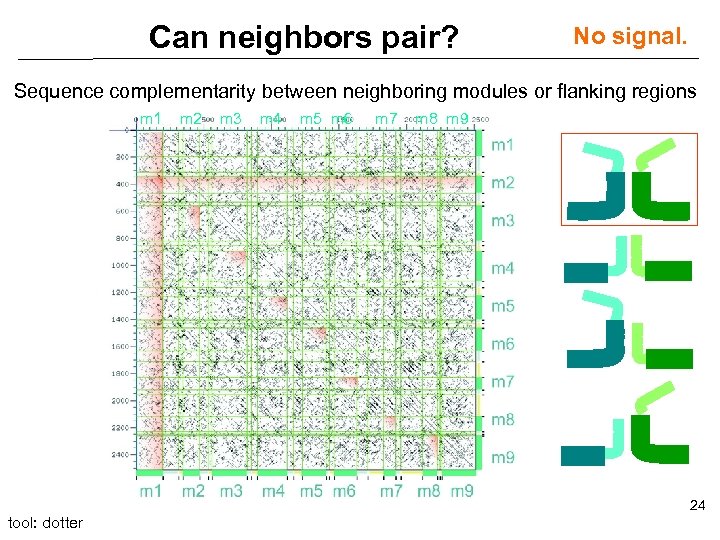
Can neighbors pair? No signal. Sequence complementarity between neighboring modules or flanking regions m 1 m 2 m 3 m 4 m 5 m 6 m 7 m 8 m 9 24 tool: dotter

Residues conserved across cassettes? No. upstream flanking cassette module downstream-flanking . . m 1. . TACGACGaggactgcgcat…. . . gatgagtagcag. CATGGCA. . m 2. . GTCGGGActcacccacgca…. . . ctgtcatcgtag. CATGGCT. . m 3. . CGTGGCAtccatac…. . . atggtatgtggt. ATCACAG. . m 4. . GGAGGACcggacactacac. . . cttggcatccac. CGCTCTA. . m 5. . GATGGTGtgtgggtgctcc…. . . taggtagcattg. GGACTTG. . m 6. . CTCTCATgcagcccgcgct…. . . atggagtcatcg. TAGGAGC. . m 7. . CCGTGTActgctacgtggt…. . . accagtgtgact. CAGGTGG. . m 8. . ACCTAGGtatctgtgctct…. . . gtacagc. TACAGTG. . m 9. . END START m 1 m 2 m 3 m 4 m 5 6 m 7 m 8 m 9 25
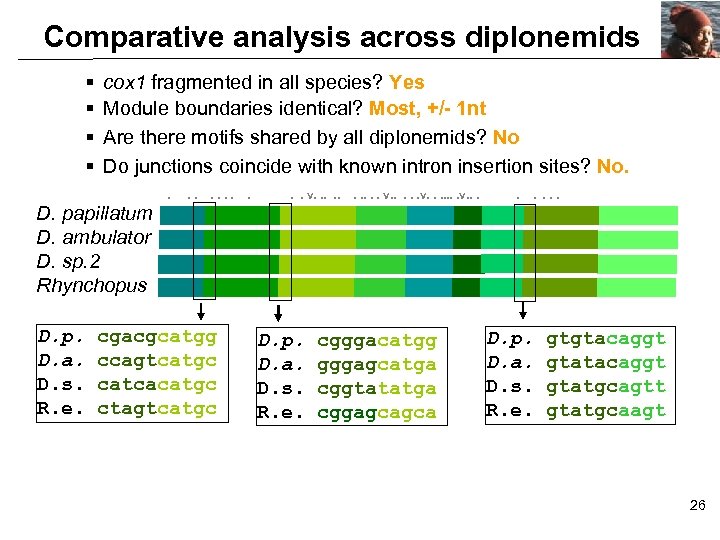

Comparative analysis across diplonemids § § cox 1 fragmented in all species? Yes Module boundaries identical? Most, +/- 1 nt Are there motifs shared by all diplonemids? No Do junctions coincide with known intron insertion sites? No. D. papillatum D. ambulator D. sp. 2 Rhynchopus D. p. D. a. D. s. R. e. . . . cgacgcatgg ccagtcatgc catcacatgc ctagtcatgc . . . v. . . D. p. D. a. D. s. R. e. . . v. . . cgggacatgg gggagcatga cggtatatga cggagcagca . . . D. p. D. a. D. s. R. e. gtgtacaggt gtatgcagtt gtatgcaagt 26
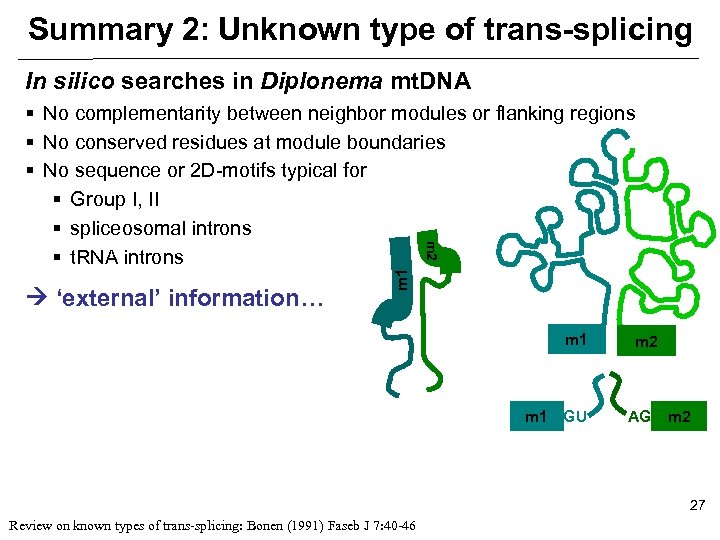
Summary 2: Unknown type of trans-splicing In silico searches in Diplonema mt. DNA ‘external’ information… m 1 m 2 § No complementarity between neighbor modules or flanking regions § No conserved residues at module boundaries § No sequence or 2 D-motifs typical for § Group I, II § spliceosomal introns § t. RNA introns m 1 GU m 2 AG m 2 27 Review on known types of trans-splicing: Bonen (1991) Faseb J 7: 40 -46
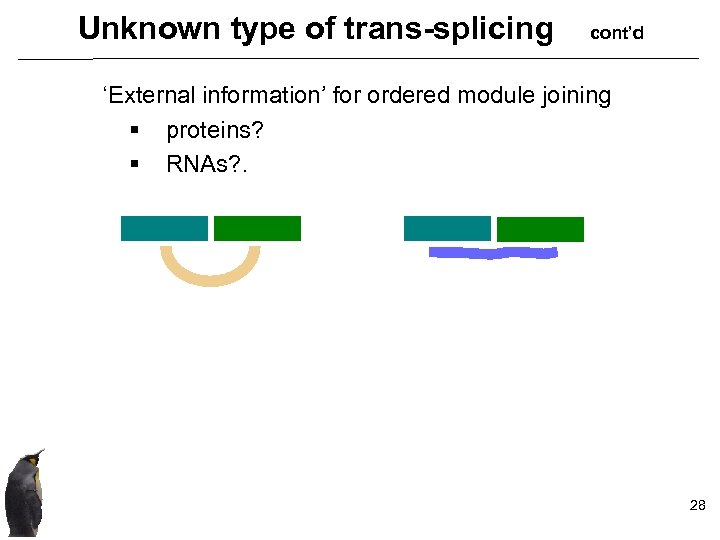
Unknown type of trans-splicing cont’d ‘External information’ for ordered module joining § proteins? § RNAs? . 28
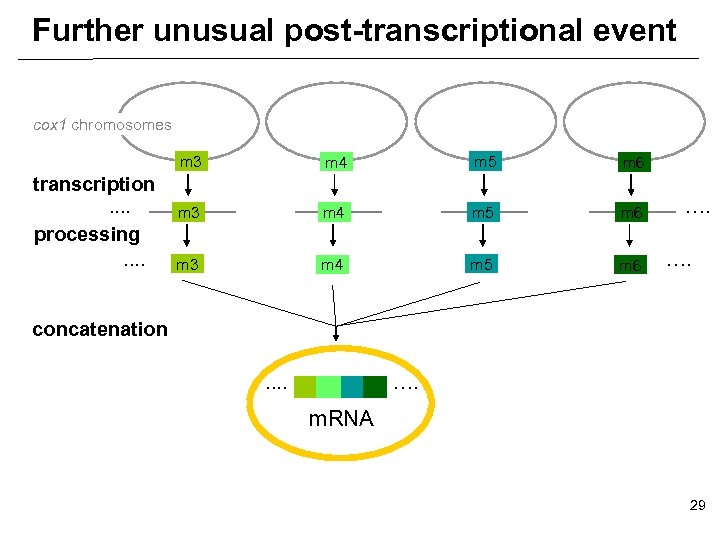
Further unusual post-transcriptional event cox 1 chromosomes m 3 transcription. . processing. . m 4 m 5 m 6 m 3 m 4 m 5 m 6 …. …. concatenation. . …. m. RNA 29
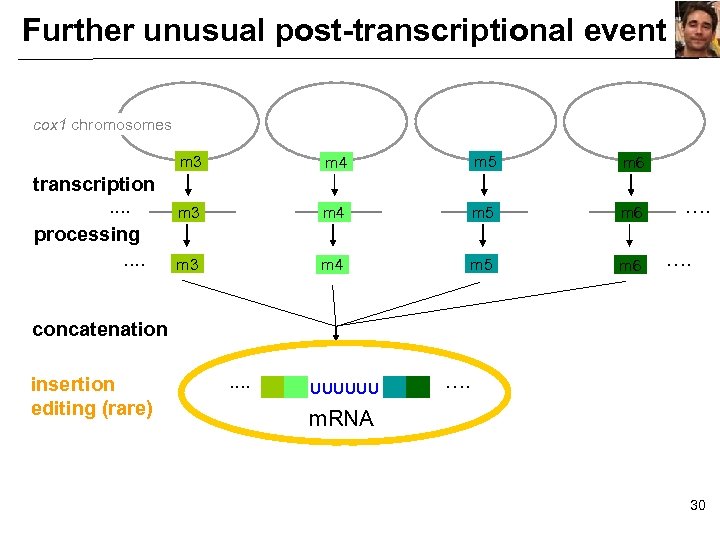
Further unusual post-transcriptional event cox 1 chromosomes m 3 transcription. . processing. . m 4 m 5 m 6 m 3 m 4 m 5 m 6 …. …. concatenation insertion editing (rare) . . UUUUUU …. m. RNA 30
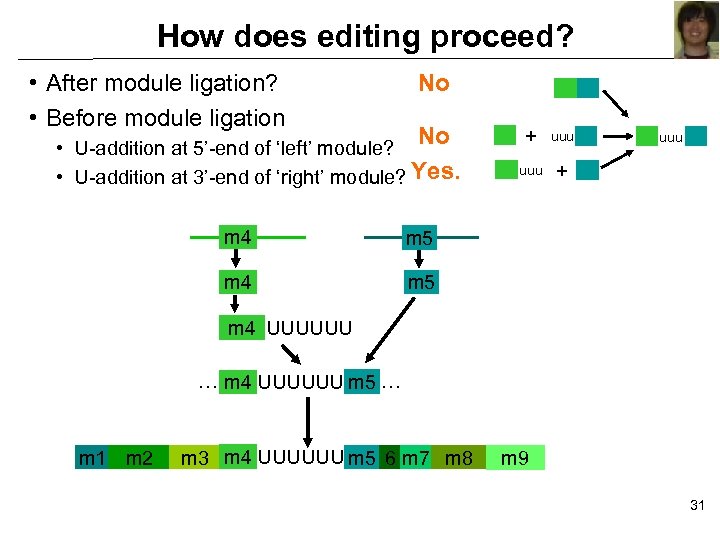
How does editing proceed? • After module ligation? • Before module ligation No No • U-addition at 5’-end of ‘left’ module? • U-addition at 3’-end of ‘right’ module? Yes. m 4 uuu + uuu m 5 m 4 + m 5 m 4 UUUUUU … m 4 UUUUUU m 5 … m 1 m 2 m 3 m 4 UUUUUU m 5 6 m 7 m 8 m 9 31
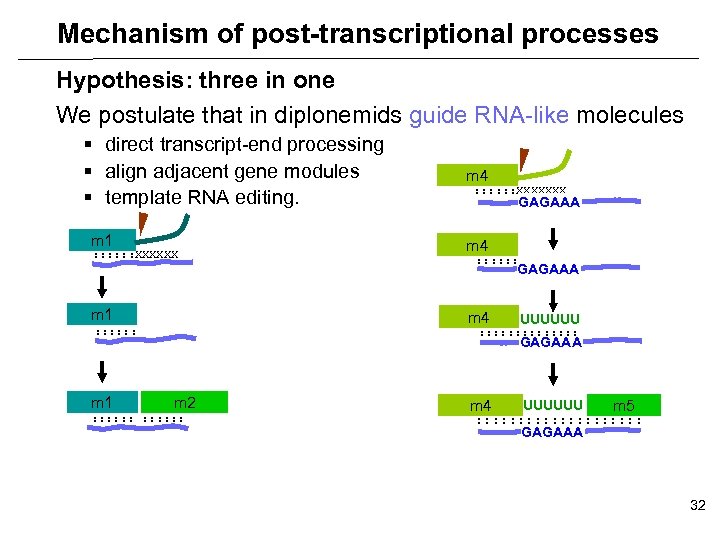
Mechanism of post-transcriptional processes Hypothesis: three in one We postulate that in diplonemids guide RNA-like molecules § direct transcript-end processing § align adjacent gene modules § template RNA editing. m 1 : : : XXXXXXX GAGAAA m 4 : : : m 1 m 4 UUUUUU : : : : GAGAAA m 2 : : : m 4 UUUUUU m 5 : : : : : GAGAAA 32
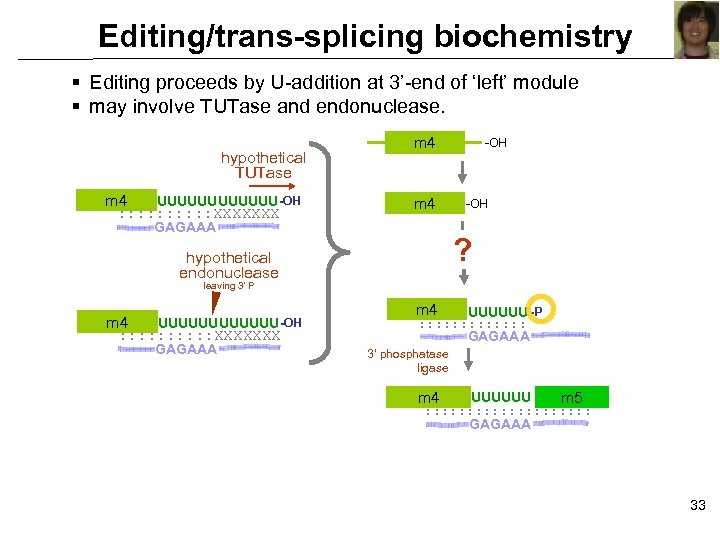
Editing/trans-splicing biochemistry § Editing proceeds by U-addition at 3’-end of ‘left’ module § may involve TUTase and endonuclease. hypothetical TUTase m 4 UUUUUU -OH : : : : : XXXXXXX m 4 GAGAAA -OH ? hypothetical endonuclease leaving 3’ P m 4 UUUUUU -OH : : : : : XXXXXXX GAGAAA m 4 UUUUUU -P : : : : GAGAAA 3’ phosphatase ligase m 4 UUUUUU m 5 : : : : : GAGAAA 33
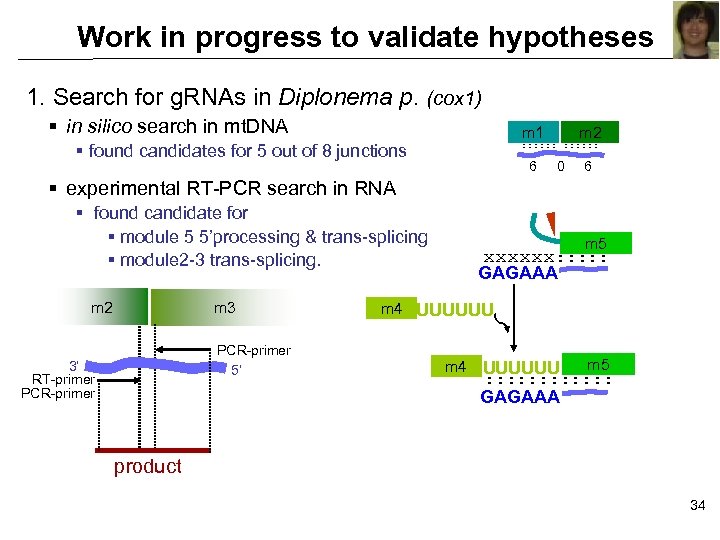
Work in progress to validate hypotheses 1. Search for g. RNAs in Diplonema p. (cox 1) § in silico search in mt. DNA m 1 m 2 : : : § found candidates for 5 out of 8 junctions 6 0 6 § experimental RT-PCR search in RNA § found candidate for § module 5 5’processing & trans-splicing § module 2 -3 trans-splicing. m 2 m 3 PCR-primer 5’ 3’ RT-primer PCR-primer m 5 xxxxxx: : : GAGAAA m 4 UUUUUU m 5 : : : GAGAAA product 34

Work in progress cont’d 1. Search for g. RNAs in Diplonema papillatum (cox 1) § in silico search in mt. DNA § experimental RT-PCR search in RNA 2. Compare cox 1 across diplonemids § search for conserved 2 D-structure (RNAshapes) 3. Develop in vitro system for validation. 35 Steffen et al 2006 Bioinformatics 22: 500
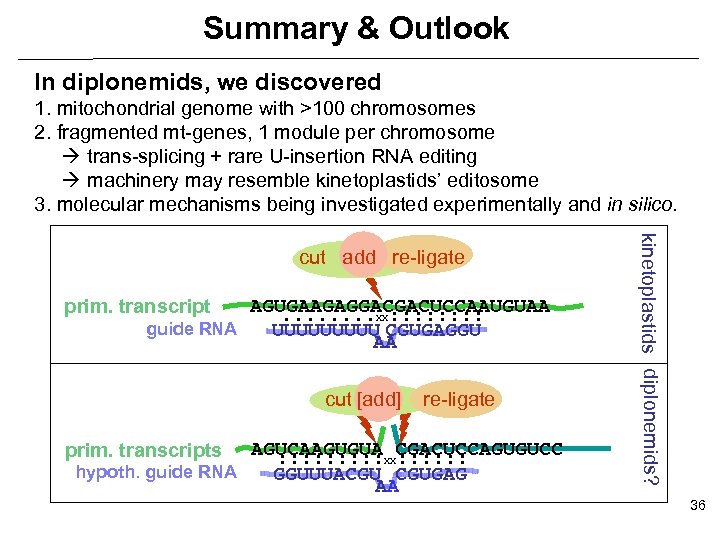
Summary & Outlook In diplonemids, we discovered 1. mitochondrial genome with >100 chromosomes 2. fragmented mt-genes, 1 module per chromosome trans-splicing + rare U-insertion RNA editing machinery may resemble kinetoplastids’ editosome 3. molecular mechanisms being investigated experimentally and in silico. prim. transcript AGUGAAGAGGACGACUCCAAUGUAA. . . . XX: : : : guide RNA UUUUU CGUGAGGU AA cut [add] prim. transcripts re-ligate AGUCAAGUGUA XXCGACUCCAGUGUCC : : : : hypoth. guide RNA GGUUUACGU CGUGAG AA kinetoplastids diplonemids? cut add re-ligate 36

Final speculations Unnecessarily complicated gene organization and expression - Advantage? - Disadvantage? - evolution under superabundant conditions. 37

Collaborations & funding Julius Lukes, University of South Bohemia, Czech Republic M. W. Gray & M. Schnare, Dalhousie University, Halifax, Canada B. Franz Lang, Université de Montréal, Canada. 38
ef2b78476982c56115352e2a9cc65fda.ppt