89a17de2a017f5f5f0b1773eec9c3b07.ppt
- Количество слайдов: 25

U-BIOPRED (Unbiased BIOmarkers in PREDiction of respiratory disease outcomes) → a 5 -year European project to understand more about severe asthma Update: start of project year 4

Pre-Clinical studies are not predictive Target compounds = 5000 – 6000 Pre-Clinial Studies = 250 Phase I = 5 Approved Drug = 1

Why? • • Lack of validated biomarkers Underperforming preclinical models Inadequate and incomplete sub-phenotyping Insufficient understanding of disease mechanisms.

Hypothesis The use of biomarker profiles comprised of various types of high-dimensional data, integrated with an innovative systems biology approach into distinct phenotype handprints, will enable significantly better prediction of therapeutic efficacy than single or even clustered biomarkers of one data type, and will identify novel targets.


Despite many challenges UBIOPRED is producing.

What UBIOPRED is producing: ü Large cohort & biobank of deeply phenotyped adult and paediatric patients ü ‘Handprints’: stratification of severe asthma ü Preclinical models more reflective of clinical disease ü A GMP viral challenge exacerbation model

Cohorts
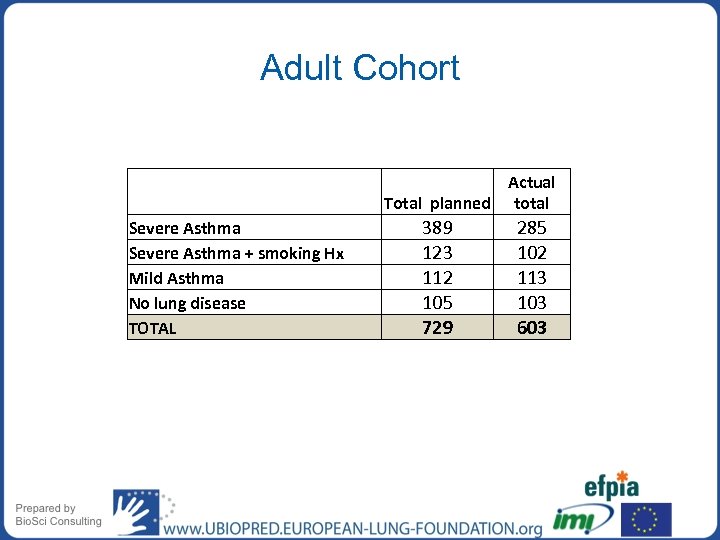
Adult Cohort Severe Asthma + smoking Hx Mild Asthma No lung disease TOTAL Actual Total planned total 389 123 112 105 729 285 102 113 103 603

Paediatric Cohort Total planned 115 50 Actual total 102 56 Pre-school severe wheeze 73 54 Pre-school mild/mod wheeze TOTAL 60 298 46 258 School aged severe asthma School aged mild asthma

Cohorts year 3 Achievements Challenges • Recruitment accelerated • Interim analysis conducted • Data cleaning – More resource to be applied • More mild asthma bronchs needed – Recall subjects for bronchs • # of exacerbation visits – More focus

Cohorts year 4 focus • • • Finishing recruitment More mild asthma bronchs Data cleaning Increase exacerbation visits Publications

Multiple data types are integrated into a ‘handprint’ to identify sub-phentoypes.
Topographical clustering on sputum and lung function produces clear clusters.

‘Handprints’ year 3 Achievements Challenges • Analysis workflows established and tested • trans. MART knowledge management platform in use – Now supported by e. TRIKS • Interim analysis shows clustering with lipidomics • Preliminary ‘Handprints’ (clusters) • Waiting for samples and clean data – Interim analyses done on what is available • Coordinating analysis efforts – Meetings every 2 months

‘Handprints’ – focus year 4 • Ongoing interim analyses • Question/publication based iterative analyses – workflows working at full speed • Interactions with other EU/IMI projects • Publications

Laboratory Models

Step 1: establish “asthmatic phenotype” AHR induction by passive sensitization ? Images before allergen exposure Images after allergen exposure Pre-incubation with serum of non-allergic donor Pre-incubation with serum of allergic donor Precision lungs are a promising model

Laboratory models year 3 Achievements Challenges • Determined that chronic House • Getting viral exacerbations Dust Mite mouse model cannot – Switch to influenza from be exacerbated by virus rhinovirus • Establihsed CFA/HDM model in • Linking with clinical data two company and one academic – Lipidomics to be done in institution light of interim analysis – Has mixed Th 1/Th 2/Th 17 results phenotype and can be exacerbated. • Explant model established • Promising work on precision cut lung slices

Laboratory models focus year 4 • • • 5 publications early 2013 Transcriptomics and analyte panels on samples Getting data into trans. MART Establish CFA/HDM in other labs Development of precision cut lung models
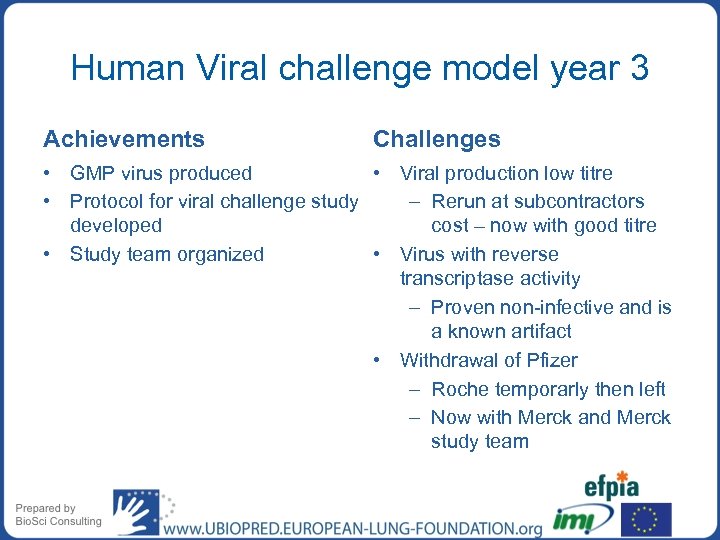

Human Viral challenge model year 3 Achievements Challenges • GMP virus produced • Viral production low titre • Protocol for viral challenge study – Rerun at subcontractors developed cost – now with good titre • Study team organized • Virus with reverse transcriptase activity – Proven non-infective and is a known artifact • Withdrawal of Pfizer – Roche temporarly then left – Now with Merck and Merck study team

Human Viral Challenge models focus year 4 • Viral challenge study • Analysis plan and linkage with ‘Handprint’ teams • Storage of virus

UBIOPRED Collaboration Successes Challenges • Two new partners • Exactly the same animal model established in 3 different laboratories • Linkage with e. TRIKS – IMI’s translational research knowledge management project • Every working group contains crucial members from both EFPIA and Academic partners • Withdrawal of Pfizer • Re-organization to enable focus on analysis • Shifting resource committments • Delays in contracting & direct financial distribution

Involvement of patients • Online forum for patients initiated • Patient representative engagement in working teleconferences • Patient representatives at yearly meeting

89a17de2a017f5f5f0b1773eec9c3b07.ppt