f73ed2523ddbffe124c2d9ccc7ec3fc2.ppt
- Количество слайдов: 23
 The use of Ontology in Organising and Managing Protein Family Resources Katy Wolstencroft, University Of Manchester
The use of Ontology in Organising and Managing Protein Family Resources Katy Wolstencroft, University Of Manchester
 Overview ¡ ¡ Research communities working on specific protein families Family Resource – central focus for the community Problems ¡ ¡ communities tend to be small Difficult to sustain resources for small number of people - funding – community input / consistency
Overview ¡ ¡ Research communities working on specific protein families Family Resource – central focus for the community Problems ¡ ¡ communities tend to be small Difficult to sustain resources for small number of people - funding – community input / consistency
 Protein Family Test Cases ¡ ¡ Protein Phosphatases: Original test family dephosphorylation, involved in control and communication. http: //www. bioinf. man. ac. uk/phosphabase ABC Transporters: (GSK Pennsylvania) transport of biological molecules through cells and organelles. Both implicated in human disease
Protein Family Test Cases ¡ ¡ Protein Phosphatases: Original test family dephosphorylation, involved in control and communication. http: //www. bioinf. man. ac. uk/phosphabase ABC Transporters: (GSK Pennsylvania) transport of biological molecules through cells and organelles. Both implicated in human disease
 Solutions From Ontology? ¡ ¡ ¡ General biological data problems * Nomenclature and free text data Sustainability Consistency and ambiguity Problems associated with both data extraction and data retrieval
Solutions From Ontology? ¡ ¡ ¡ General biological data problems * Nomenclature and free text data Sustainability Consistency and ambiguity Problems associated with both data extraction and data retrieval
 GO Accessible Resources GO – “de facto standard” – description of gene products Biological Resources – annotating data to GO terms
GO Accessible Resources GO – “de facto standard” – description of gene products Biological Resources – annotating data to GO terms
 Domain-specific Ontology GO allows efficient collection of biological data from heterogeneous sources – not rich enough to describe a whole protein family domain ¡ ¡ How should the information be stored? What information should be stored?
Domain-specific Ontology GO allows efficient collection of biological data from heterogeneous sources – not rich enough to describe a whole protein family domain ¡ ¡ How should the information be stored? What information should be stored?
 Requirements Protein Phosphatase ¡ Inhibitors - protein inhibition, transcriptional repression ¡ Activators - protein activation, transcriptional activation ¡ Domain Structure ¡ Disease Association ¡ Genetic Localisation ¡ Tissue Expression ¡ Enzyme - substrates/ products ¡ Cofactor/prosthetic group/molecule required for activation ABC Transporters ¡ Inhibitors - protein inhibition, transcriptional repression ¡ Activators - protein activation, transcriptional activation ¡ Domain Structure ¡ Disease Association ¡ Genetic Localisation ¡ Tissue Expression ¡ Transported substrates
Requirements Protein Phosphatase ¡ Inhibitors - protein inhibition, transcriptional repression ¡ Activators - protein activation, transcriptional activation ¡ Domain Structure ¡ Disease Association ¡ Genetic Localisation ¡ Tissue Expression ¡ Enzyme - substrates/ products ¡ Cofactor/prosthetic group/molecule required for activation ABC Transporters ¡ Inhibitors - protein inhibition, transcriptional repression ¡ Activators - protein activation, transcriptional activation ¡ Domain Structure ¡ Disease Association ¡ Genetic Localisation ¡ Tissue Expression ¡ Transported substrates
 System Requirements Phospha. Base ¡ ¡ Phospha. Base Doamin Ontology DAML+OIL description logic Relational Database – My. Sql User interface – Java Servlet Free access over the internet My. SQL and Java free Java platform independent ABC Transporters ¡ ¡ ABC Domain Ontology DAML+OIL description logic Relational Database Oracle User Interface – Ontology driven interface Internal Company Use Limited access
System Requirements Phospha. Base ¡ ¡ Phospha. Base Doamin Ontology DAML+OIL description logic Relational Database – My. Sql User interface – Java Servlet Free access over the internet My. SQL and Java free Java platform independent ABC Transporters ¡ ¡ ABC Domain Ontology DAML+OIL description logic Relational Database Oracle User Interface – Ontology driven interface Internal Company Use Limited access
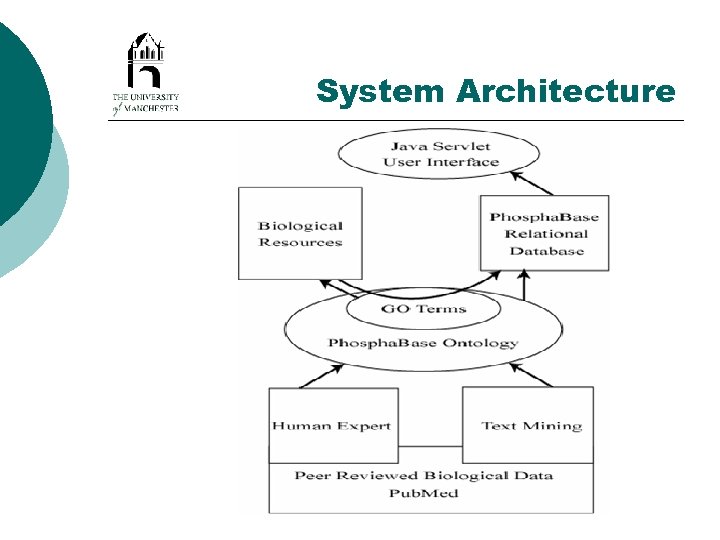 System Architecture
System Architecture
 Architecture Advantages /Disadvantages ¡ ¡ Sustainability – Data capture can be automated. Diagnostics – Classification of ‘unknown’ proteins. *Major application in annotation of new genomes* ¡ Accessibility and Portability– Free availability over the Internet. All software freely available Issues ¡ Maintenance – automation and use of ontology reduces human intervention but the ontology needs occasional maintenance ¡ Standards – DAML+OIL ontology. Need to migrate to OWL, but OWL does not currently allow qualified number restrictions
Architecture Advantages /Disadvantages ¡ ¡ Sustainability – Data capture can be automated. Diagnostics – Classification of ‘unknown’ proteins. *Major application in annotation of new genomes* ¡ Accessibility and Portability– Free availability over the Internet. All software freely available Issues ¡ Maintenance – automation and use of ontology reduces human intervention but the ontology needs occasional maintenance ¡ Standards – DAML+OIL ontology. Need to migrate to OWL, but OWL does not currently allow qualified number restrictions
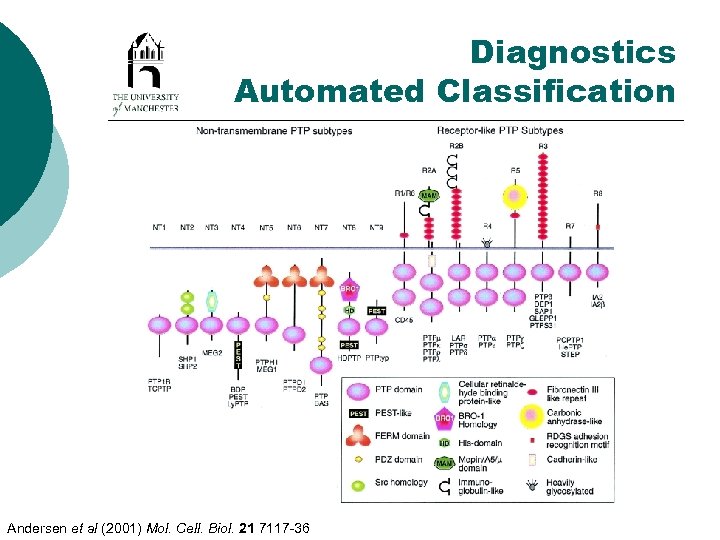 Diagnostics Automated Classification Andersen et al (2001) Mol. Cell. Biol. 21 7117 -36
Diagnostics Automated Classification Andersen et al (2001) Mol. Cell. Biol. 21 7117 -36
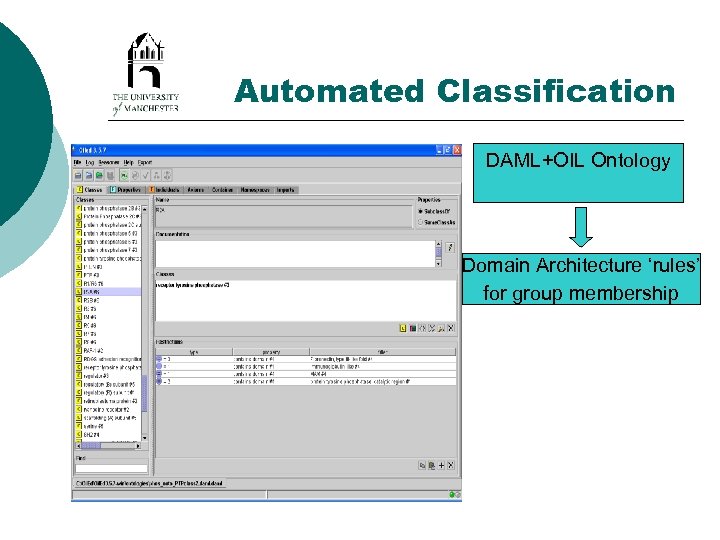 Automated Classification DAML+OIL Ontology Domain Architecture ‘rules’ for group membership
Automated Classification DAML+OIL Ontology Domain Architecture ‘rules’ for group membership
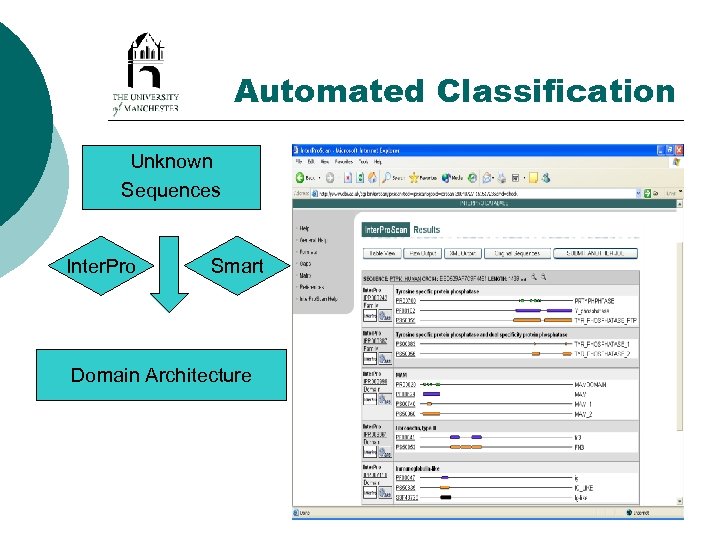 Automated Classification Unknown Sequences Inter. Pro Smart Domain Architecture
Automated Classification Unknown Sequences Inter. Pro Smart Domain Architecture
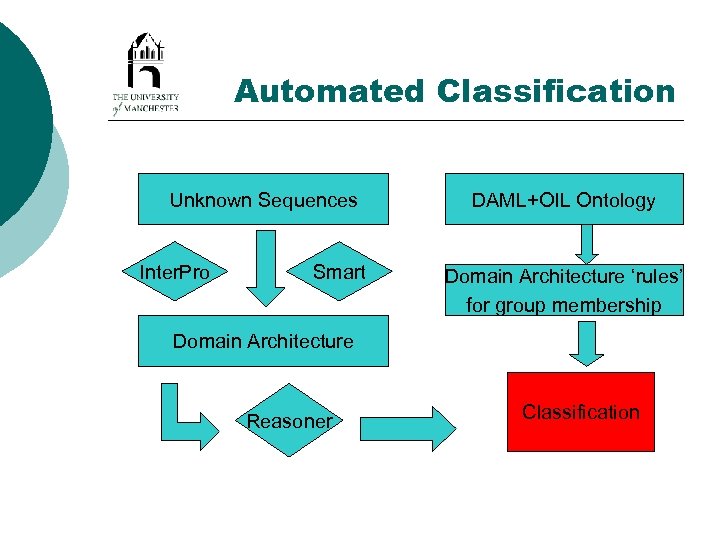 Automated Classification Unknown Sequences Inter. Pro Smart DAML+OIL Ontology Domain Architecture ‘rules’ for group membership Domain Architecture Reasoner Classification
Automated Classification Unknown Sequences Inter. Pro Smart DAML+OIL Ontology Domain Architecture ‘rules’ for group membership Domain Architecture Reasoner Classification
 Summary ¡ ¡ ¡ Two rich and useful resources for the phosphatase and ABC transporter research communities A sustainable resource with automatic classification capabilities Generic Model – A robust model could be extended to build similar resources for other protein families in the future Ontology technology – powerful tool in managing biological data
Summary ¡ ¡ ¡ Two rich and useful resources for the phosphatase and ABC transporter research communities A sustainable resource with automatic classification capabilities Generic Model – A robust model could be extended to build similar resources for other protein families in the future Ontology technology – powerful tool in managing biological data
 Next Steps Phosphorylation Ontology ¡ ¡ Control of Phosphorylation mediated by both phosphatases and kinases Collaboration - Protein Kinase Resource (UCSD) to describe whole phosphorylation events phosphatase Pi Phospho protein kinase
Next Steps Phosphorylation Ontology ¡ ¡ Control of Phosphorylation mediated by both phosphatases and kinases Collaboration - Protein Kinase Resource (UCSD) to describe whole phosphorylation events phosphatase Pi Phospho protein kinase
 Phosphorylation Ontology Goals ¡ ¡ ¡ Use ontology technology to capture whole phosphorylation events Description of phosphorylation events in the cell and the biological pathways they affect Produce phosphorylation resource useful to phosphatase and kinase community and wider.
Phosphorylation Ontology Goals ¡ ¡ ¡ Use ontology technology to capture whole phosphorylation events Description of phosphorylation events in the cell and the biological pathways they affect Produce phosphorylation resource useful to phosphatase and kinase community and wider.
 Acknowledgements Supervisors: Andy Brass, Robert Stevens Advisor and Phosphatase Biologist: Lydia Tabernero GSK: Robin Mc. Entire and the IKM group Funding: Medical Research Council
Acknowledgements Supervisors: Andy Brass, Robert Stevens Advisor and Phosphatase Biologist: Lydia Tabernero GSK: Robin Mc. Entire and the IKM group Funding: Medical Research Council
 Screen Shots
Screen Shots
 Query Result
Query Result
 Query Result - Continued
Query Result - Continued
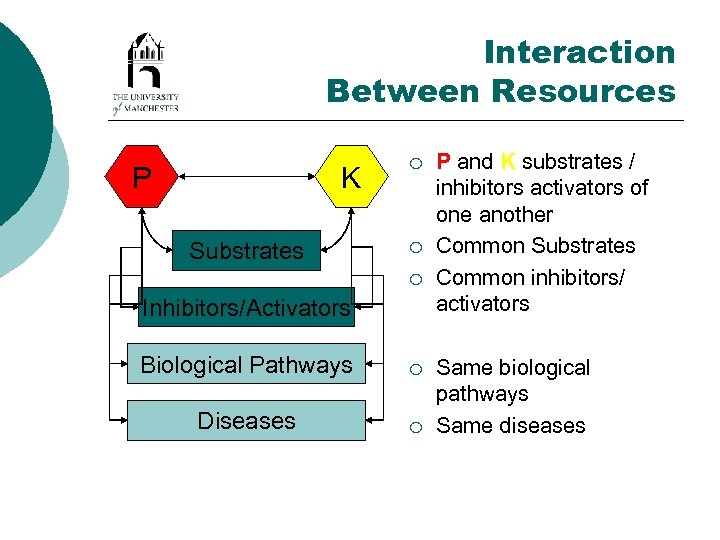 Interaction Between Resources P K Substrates ¡ ¡ ¡ Inhibitors/Activators Biological Pathways ¡ Diseases ¡ P and K substrates / inhibitors activators of one another Common Substrates Common inhibitors/ activators Same biological pathways Same diseases
Interaction Between Resources P K Substrates ¡ ¡ ¡ Inhibitors/Activators Biological Pathways ¡ Diseases ¡ P and K substrates / inhibitors activators of one another Common Substrates Common inhibitors/ activators Same biological pathways Same diseases
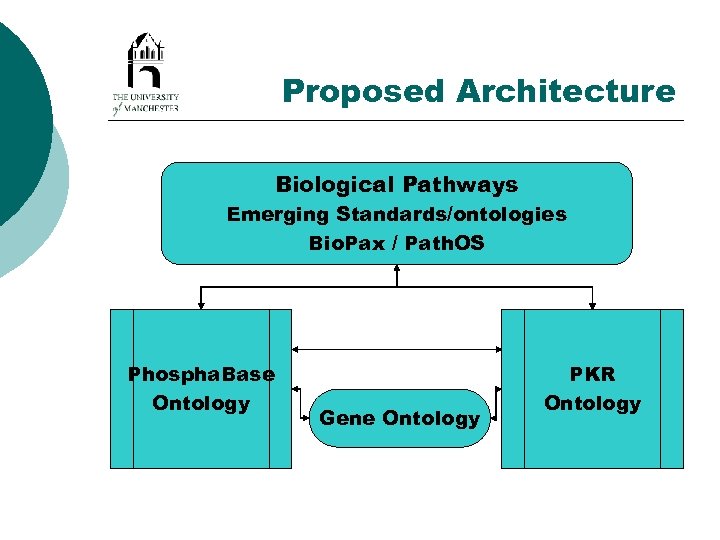 Proposed Architecture Biological Pathways Emerging Standards/ontologies Bio. Pax / Path. OS Phospha. Base Ontology Gene Ontology PKR Ontology
Proposed Architecture Biological Pathways Emerging Standards/ontologies Bio. Pax / Path. OS Phospha. Base Ontology Gene Ontology PKR Ontology


