355ea3d4e18580a39b01d4e51431cd0b.ppt
- Количество слайдов: 74
 The Use of Geometrical and Physical Models in Quantitative Medical Image Analysis James S. Duncan, Ph. D. Departments of Diagnostic Radiology and Electrical Engineering Yale University
The Use of Geometrical and Physical Models in Quantitative Medical Image Analysis James S. Duncan, Ph. D. Departments of Diagnostic Radiology and Electrical Engineering Yale University
 Collaborators • Segmentation: -Larry Staib -Amit Chakraborty -Isil Bozma -Xiaolan Zeng • Cardiac Deformation: - Xenios Papademetris - Pengcheng Shi -Albert Sinusas, M. D. -Todd Constable • Brain Shift Compensation: - Oskar Skrinjar - Dennis Spencer, M. D.
Collaborators • Segmentation: -Larry Staib -Amit Chakraborty -Isil Bozma -Xiaolan Zeng • Cardiac Deformation: - Xenios Papademetris - Pengcheng Shi -Albert Sinusas, M. D. -Todd Constable • Brain Shift Compensation: - Oskar Skrinjar - Dennis Spencer, M. D.
 Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
 Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
 Active Research Areas in Medical Image Analysis • Image Distortion Correction • Quantitative Measurement of Anatomy/ Physiology • Quantitative Characterization of Disease • Data/ Information Handling/Fusion • Modeling/ prediction of Function/Disease • Improved Visualization of All of Above • Image- Guided Intervention/Surgery
Active Research Areas in Medical Image Analysis • Image Distortion Correction • Quantitative Measurement of Anatomy/ Physiology • Quantitative Characterization of Disease • Data/ Information Handling/Fusion • Modeling/ prediction of Function/Disease • Improved Visualization of All of Above • Image- Guided Intervention/Surgery
 We’ll Look at Several of These Issues from the Viewpoint of Three Examples: a. ) Characterization of Cardiac Function from Noninvasive 4 D Imaging b. ) Study of Neuroanatomical Structure from MRI c. ) Brain Shift Compensation in Image Guided Neurosurgery
We’ll Look at Several of These Issues from the Viewpoint of Three Examples: a. ) Characterization of Cardiac Function from Noninvasive 4 D Imaging b. ) Study of Neuroanatomical Structure from MRI c. ) Brain Shift Compensation in Image Guided Neurosurgery
 a. ). Recovery of Cardiac Deformation (Problem Statement)
a. ). Recovery of Cardiac Deformation (Problem Statement)
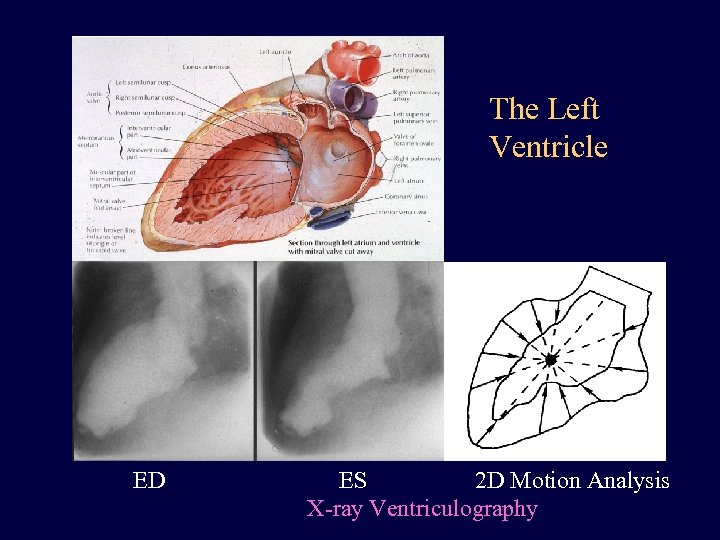 The Left Ventricle ED ES 2 D Motion Analysis X-ray Ventriculography
The Left Ventricle ED ES 2 D Motion Analysis X-ray Ventriculography
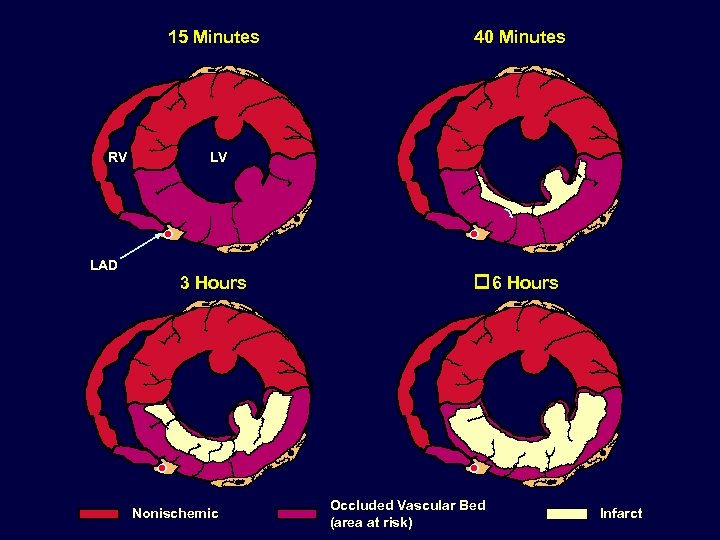 15 Minutes RV LAD 40 Minutes LV 3 Hours Nonischemic 6 Hours Occluded Vascular Bed (area at risk) Infarct
15 Minutes RV LAD 40 Minutes LV 3 Hours Nonischemic 6 Hours Occluded Vascular Bed (area at risk) Infarct
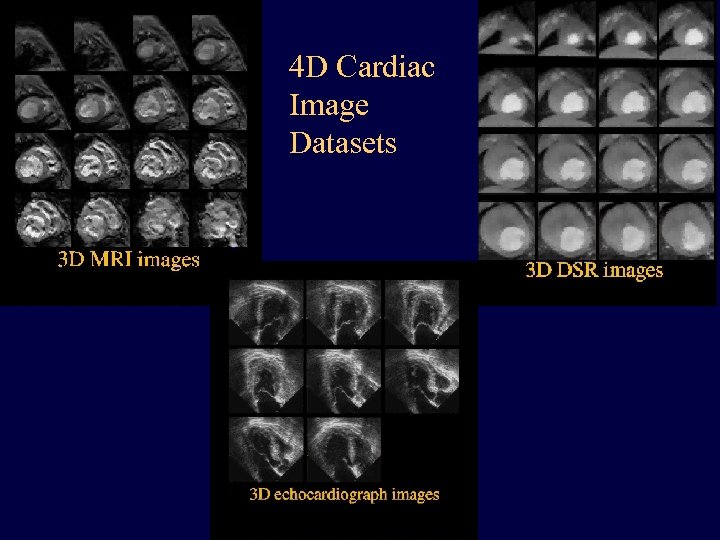 4 D Cardiac Image Datasets
4 D Cardiac Image Datasets
 Transmyocardial Revascularization (TMR) Canine Experiment
Transmyocardial Revascularization (TMR) Canine Experiment
 b. ) Measurement of Neuroanatomical Structure (Problem Statement)
b. ) Measurement of Neuroanatomical Structure (Problem Statement)
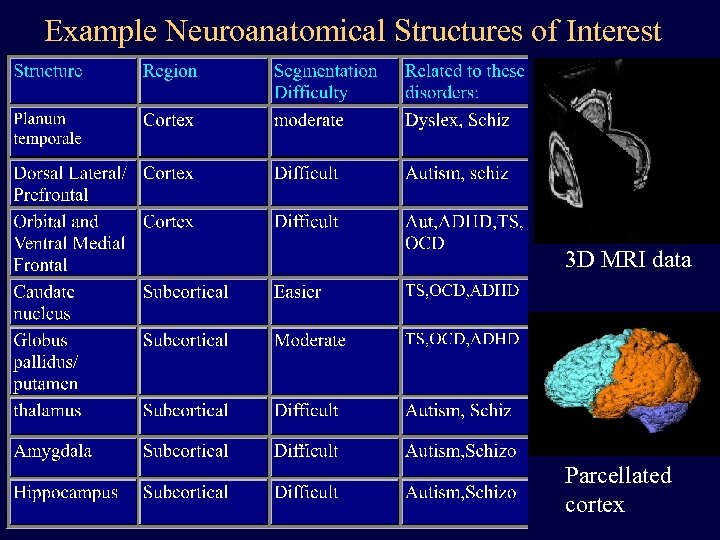 Example Neuroanatomical Structures of Interest 3 D MRI data Parcellated cortex
Example Neuroanatomical Structures of Interest 3 D MRI data Parcellated cortex
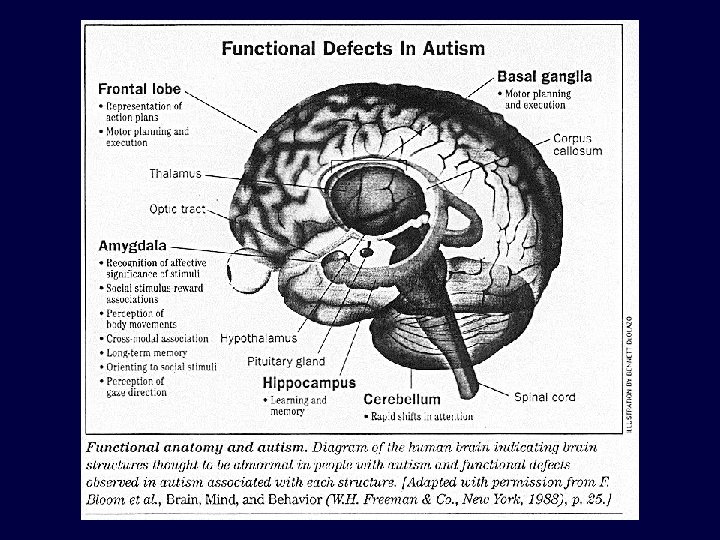
 c. ) Brain Shift Compensation in Image-Guided Neurosurgery (Problem Statement)
c. ) Brain Shift Compensation in Image-Guided Neurosurgery (Problem Statement)
 Neurosurgical Issues Related to Brain Shift • Navigation via Image-Guided Stereotactic System is Difficult/ Inaccurate: - perceived vs. actual position of structures (especially below the surface) off by many mm. - especially problematic for placing depth electrodes during epilepsy surgery • in Epilepsy Surgery: Implanted Electrode Arrays (used to gather confirmatory functional information) further deform brain: - now difficult to relate to preop estimates of seizure location (from SPECT/PET) and/or cognitive activity (f. MRI) • Tumor localization (especially below the surface) difficult in Image -Guided system both pre- and post- resection
Neurosurgical Issues Related to Brain Shift • Navigation via Image-Guided Stereotactic System is Difficult/ Inaccurate: - perceived vs. actual position of structures (especially below the surface) off by many mm. - especially problematic for placing depth electrodes during epilepsy surgery • in Epilepsy Surgery: Implanted Electrode Arrays (used to gather confirmatory functional information) further deform brain: - now difficult to relate to preop estimates of seizure location (from SPECT/PET) and/or cognitive activity (f. MRI) • Tumor localization (especially below the surface) difficult in Image -Guided system both pre- and post- resection
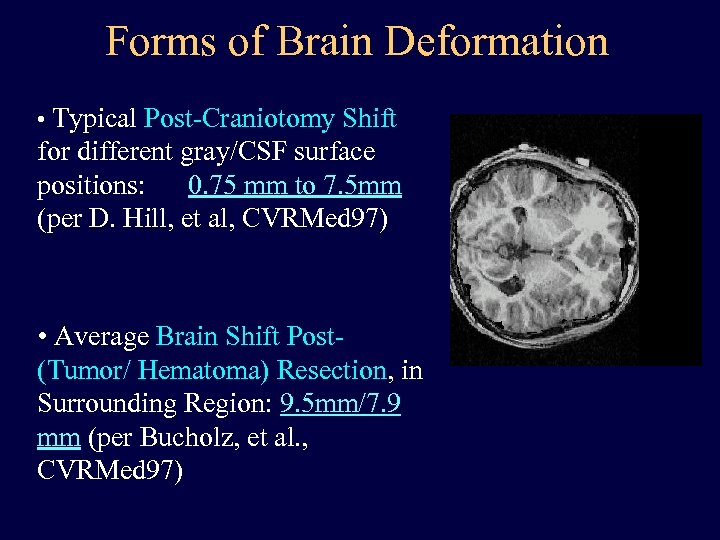 Forms of Brain Deformation • Typical Post-Craniotomy Shift for different gray/CSF surface positions: 0. 75 mm to 7. 5 mm (per D. Hill, et al, CVRMed 97) • Average Brain Shift Post(Tumor/ Hematoma) Resection, in Surrounding Region: 9. 5 mm/7. 9 mm (per Bucholz, et al. , CVRMed 97)
Forms of Brain Deformation • Typical Post-Craniotomy Shift for different gray/CSF surface positions: 0. 75 mm to 7. 5 mm (per D. Hill, et al, CVRMed 97) • Average Brain Shift Post(Tumor/ Hematoma) Resection, in Surrounding Region: 9. 5 mm/7. 9 mm (per Bucholz, et al. , CVRMed 97)
 All 3 Examples Involve the Same Basic Algorithmic Steps • Segment Structural Boundaries from 3 D/4 D image datasets: - Invoke Model for Context • Recover Quantitative Information (size/ motion/deformation) about Heart or Brain: - Models Guide this Process • Visualize Results (most helpful to relate to raw image data) Each Step Utilizes Geometrical or Biomechanical Models !
All 3 Examples Involve the Same Basic Algorithmic Steps • Segment Structural Boundaries from 3 D/4 D image datasets: - Invoke Model for Context • Recover Quantitative Information (size/ motion/deformation) about Heart or Brain: - Models Guide this Process • Visualize Results (most helpful to relate to raw image data) Each Step Utilizes Geometrical or Biomechanical Models !
 Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
 Geometrical Models: Segmentation
Geometrical Models: Segmentation
 Segmentation: Why is it useful in Medical Image analysis ? • identify anatomical structure/ landmarks/ROI’s for : • measurement of anatomy and function • to help register and then compare images • visualization of structure and function • group spatially-related functional activity
Segmentation: Why is it useful in Medical Image analysis ? • identify anatomical structure/ landmarks/ROI’s for : • measurement of anatomy and function • to help register and then compare images • visualization of structure and function • group spatially-related functional activity
 Segmentation: What’s typically used ? • Manual outlining of structure (problems: reproducibility, labor intense) • thresholding gray level histograms (no sense of spatial connectivity) • seed simple region growing algorithms that fill within isocontours (poor accuracy on real data) • feature clustering of multiecho MR images to get tissue classes (no spatial connectivity)
Segmentation: What’s typically used ? • Manual outlining of structure (problems: reproducibility, labor intense) • thresholding gray level histograms (no sense of spatial connectivity) • seed simple region growing algorithms that fill within isocontours (poor accuracy on real data) • feature clustering of multiecho MR images to get tissue classes (no spatial connectivity)
 Our Take on Deformable Model- based approaches to Segmentation • Have looked generally at one structure at a time • Desire to utilize BOTH bounding structure and region homogeneity constraints • Feel that there are two general classes of problems in the brain: 1. ) “find something that roughly looks like this global shape” (e. g. the LV, many subcortical structures) 2. ) “locate the most locally-coherent structure according to these constraints” (e. g. the outer cortex)
Our Take on Deformable Model- based approaches to Segmentation • Have looked generally at one structure at a time • Desire to utilize BOTH bounding structure and region homogeneity constraints • Feel that there are two general classes of problems in the brain: 1. ) “find something that roughly looks like this global shape” (e. g. the LV, many subcortical structures) 2. ) “locate the most locally-coherent structure according to these constraints” (e. g. the outer cortex)
 Related Work from Other Groups • Shape Priors: Cootes and Taylor (IPMI 93) ; Grenander/ Miller (atlases/templates- 1991) ; Vemuri, et al. (Med. IA 97) • Integrated Methods: 1. ) Region grow w/ edges. Pavlidis and Liow (PAMI 91); Zhu and Yuille (ICCV 95) ; Ahuja (PAMI 96) 2. ) level sets - Tek and Kimia (ICCV 95) • Segment/ Measure Cortical Gray Matter: Macdonald and Evans (SPIE 95) ; Teo and Sapiro (TMI 97) ; Xu and Prince (MICCAI 98) ; Davatzikos (TMI 95 - ribbons) • Also see Review of Deformable Models by Mc. Inerney and Terzopolous, Medical Image Analysis, Vol. 1 No. 2, 1996.
Related Work from Other Groups • Shape Priors: Cootes and Taylor (IPMI 93) ; Grenander/ Miller (atlases/templates- 1991) ; Vemuri, et al. (Med. IA 97) • Integrated Methods: 1. ) Region grow w/ edges. Pavlidis and Liow (PAMI 91); Zhu and Yuille (ICCV 95) ; Ahuja (PAMI 96) 2. ) level sets - Tek and Kimia (ICCV 95) • Segment/ Measure Cortical Gray Matter: Macdonald and Evans (SPIE 95) ; Teo and Sapiro (TMI 97) ; Xu and Prince (MICCAI 98) ; Davatzikos (TMI 95 - ribbons) • Also see Review of Deformable Models by Mc. Inerney and Terzopolous, Medical Image Analysis, Vol. 1 No. 2, 1996.
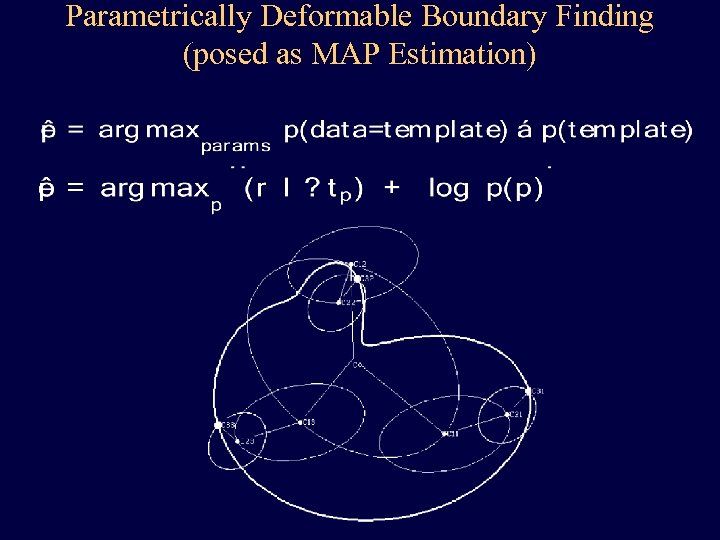 Parametrically Deformable Boundary Finding (posed as MAP Estimation)
Parametrically Deformable Boundary Finding (posed as MAP Estimation)
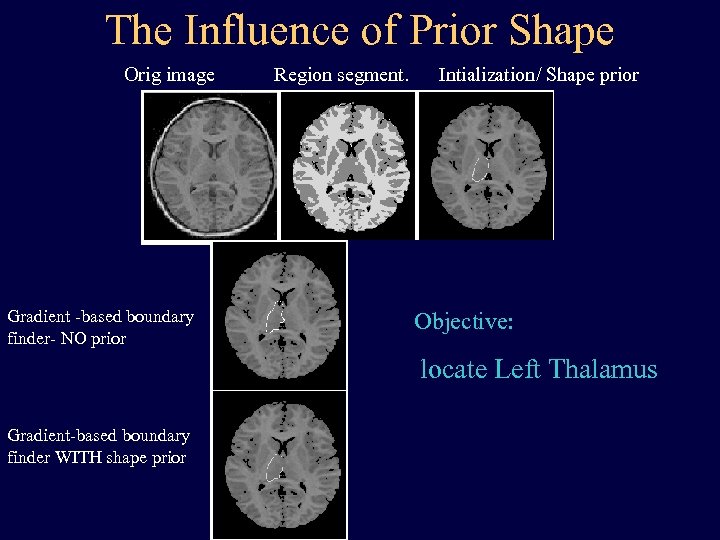 The Influence of Prior Shape Orig image Region segment. Intialization/ Shape prior Integrated (region-segment. - Gradient -based boundary finder- NO prior (Left Thalamus) boundary finderinfluenced) Objective: NO prior locate Left Thalamus Gradient-based boundary finder WITH shape prior Integrated boundary finder WITH shape prior
The Influence of Prior Shape Orig image Region segment. Intialization/ Shape prior Integrated (region-segment. - Gradient -based boundary finder- NO prior (Left Thalamus) boundary finderinfluenced) Objective: NO prior locate Left Thalamus Gradient-based boundary finder WITH shape prior Integrated boundary finder WITH shape prior
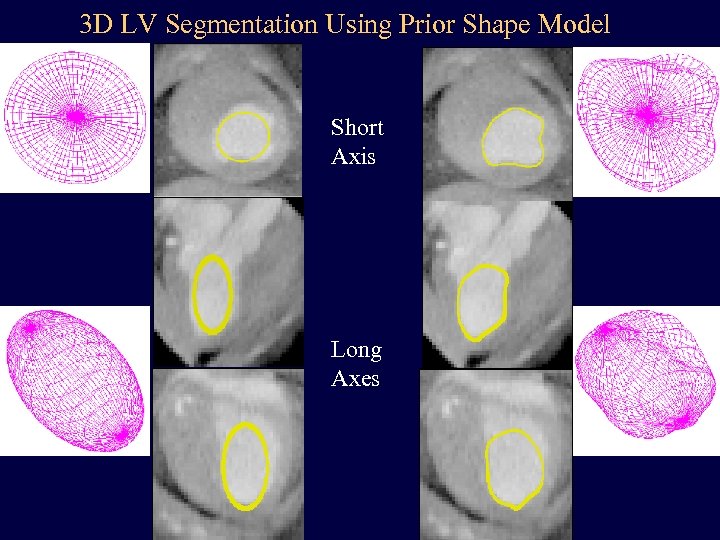 3 D LV Segmentation Using Prior Shape Model Short Axis Long Axes
3 D LV Segmentation Using Prior Shape Model Short Axis Long Axes
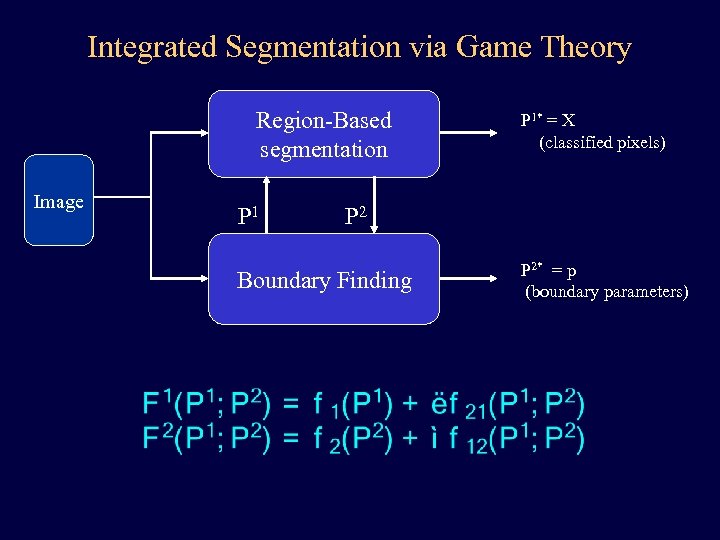 Integrated Segmentation via Game Theory Region-Based segmentation Image P 1* = X (classified pixels) P 2 Boundary Finding P 2* = p (boundary parameters)
Integrated Segmentation via Game Theory Region-Based segmentation Image P 1* = X (classified pixels) P 2 Boundary Finding P 2* = p (boundary parameters)
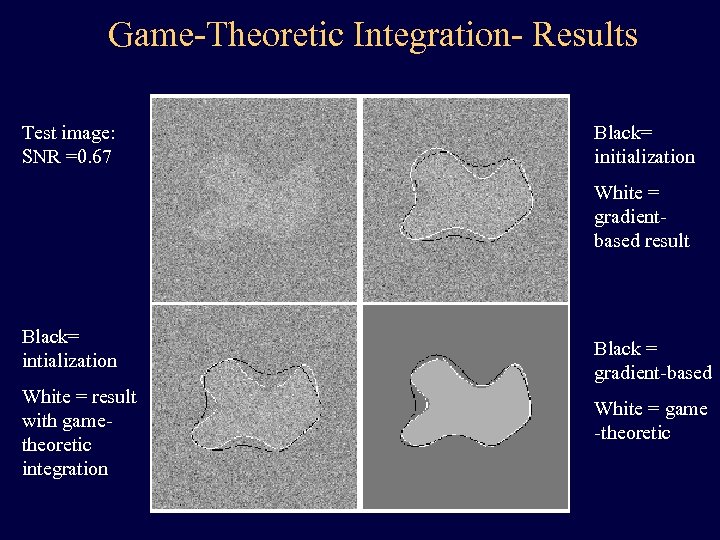 Game-Theoretic Integration- Results Test image: SNR =0. 67 Black= initialization White = gradientbased result Black= intialization White = result with gametheoretic integration Black = gradient-based White = game -theoretic
Game-Theoretic Integration- Results Test image: SNR =0. 67 Black= initialization White = gradientbased result Black= intialization White = result with gametheoretic integration Black = gradient-based White = game -theoretic
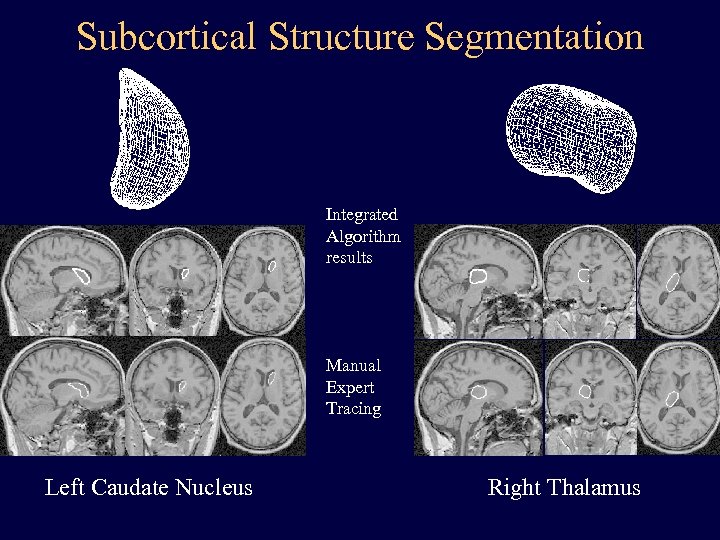 Subcortical Structure Segmentation Integrated Algorithm results Manual Expert Tracing Left Caudate Nucleus Right Thalamus
Subcortical Structure Segmentation Integrated Algorithm results Manual Expert Tracing Left Caudate Nucleus Right Thalamus
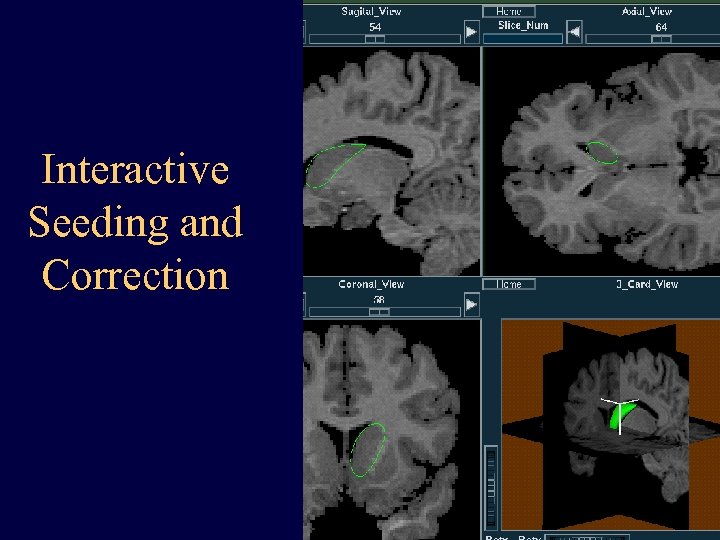 Interactive Seeding and Correction
Interactive Seeding and Correction
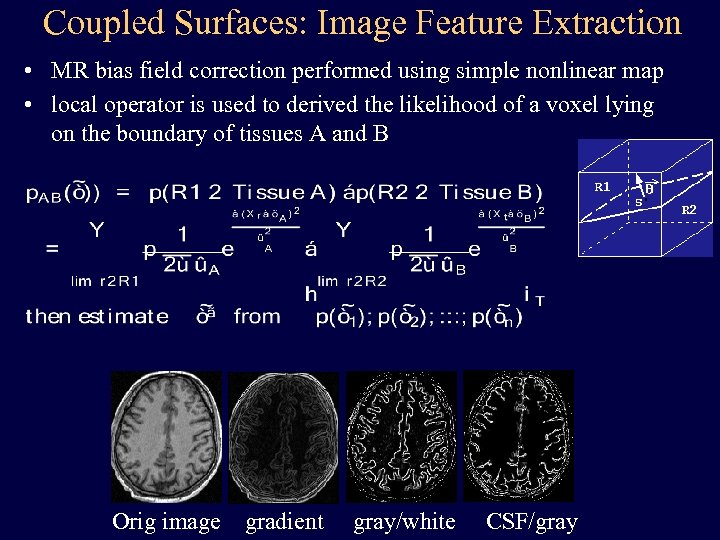 Coupled Surfaces: Image Feature Extraction • MR bias field correction performed using simple nonlinear map • local operator is used to derived the likelihood of a voxel lying on the boundary of tissues A and B Orig image gradient gray/white CSF/gray
Coupled Surfaces: Image Feature Extraction • MR bias field correction performed using simple nonlinear map • local operator is used to derived the likelihood of a voxel lying on the boundary of tissues A and B Orig image gradient gray/white CSF/gray
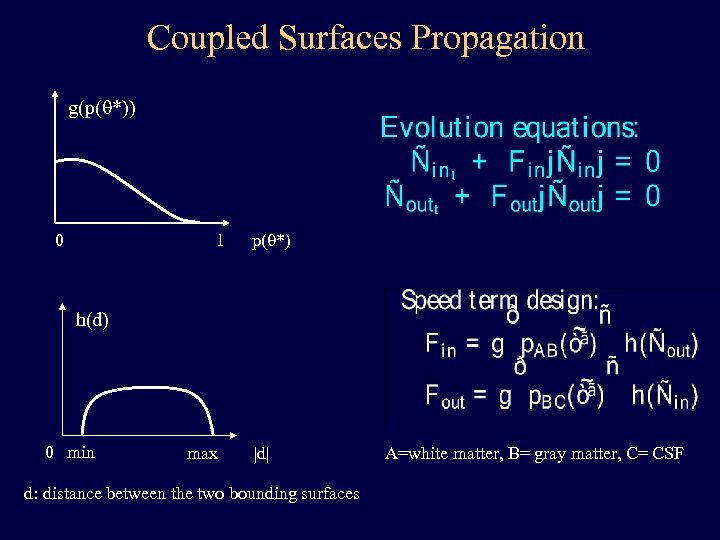 Coupled Surfaces Propagation g(p(q*)) 1 0 p(q*) h(d) 0 min max |d| d: distance between the two bounding surfaces A=white matter, B= gray matter, C= CSF
Coupled Surfaces Propagation g(p(q*)) 1 0 p(q*) h(d) 0 min max |d| d: distance between the two bounding surfaces A=white matter, B= gray matter, C= CSF
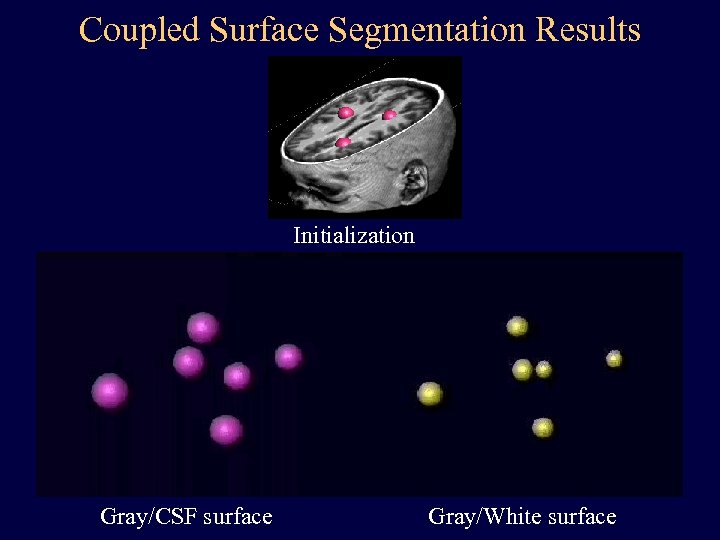 Coupled Surface Segmentation Results Initialization Gray/CSF surface Gray/White surface
Coupled Surface Segmentation Results Initialization Gray/CSF surface Gray/White surface
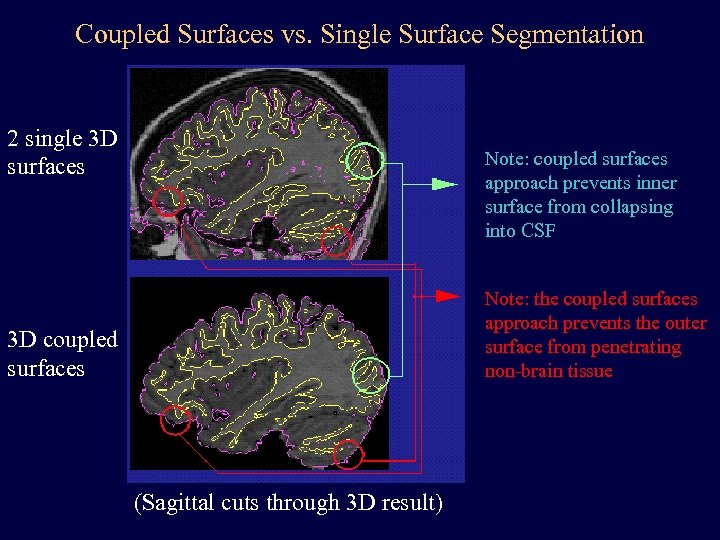 Coupled Surfaces vs. Single Surface Segmentation 2 single 3 D surfaces Note: coupled surfaces approach prevents inner surface from collapsing into CSF Note: the coupled surfaces approach prevents the outer surface from penetrating non-brain tissue 3 D coupled surfaces (Sagittal cuts through 3 D result)
Coupled Surfaces vs. Single Surface Segmentation 2 single 3 D surfaces Note: coupled surfaces approach prevents inner surface from collapsing into CSF Note: the coupled surfaces approach prevents the outer surface from penetrating non-brain tissue 3 D coupled surfaces (Sagittal cuts through 3 D result)
 Geometrical Models: Cardiac Motion Tracking
Geometrical Models: Cardiac Motion Tracking
 Cardiac Motion Tracking: Related Work from Other Groups • MR Tagging: Young and Axel, 1993 ; Park, Metaxas, and Axel, 1995 ; Denney and Prince, 1994. • MR Phase Contrast Velocities: Pelc, Herfkens and L. Pelc, 1992; Meyer, Constable, Sinusas and Duncan, 1995. • Biomechanical Model- based Tracking efforts: Guccione, Mc. Culloch and Waldman, 1991; Park and Metaxas, 1996.
Cardiac Motion Tracking: Related Work from Other Groups • MR Tagging: Young and Axel, 1993 ; Park, Metaxas, and Axel, 1995 ; Denney and Prince, 1994. • MR Phase Contrast Velocities: Pelc, Herfkens and L. Pelc, 1992; Meyer, Constable, Sinusas and Duncan, 1995. • Biomechanical Model- based Tracking efforts: Guccione, Mc. Culloch and Waldman, 1991; Park and Metaxas, 1996.
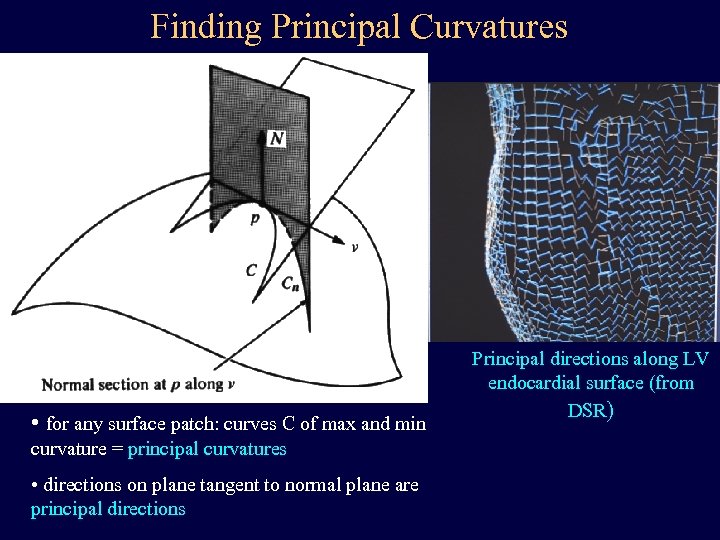 Finding Principal Curvatures • for any surface patch: curves C of max and min curvature = principal curvatures • directions on plane tangent to normal plane are principal directions Principal directions along LV endocardial surface (from DSR)
Finding Principal Curvatures • for any surface patch: curves C of max and min curvature = principal curvatures • directions on plane tangent to normal plane are principal directions Principal directions along LV endocardial surface (from DSR)
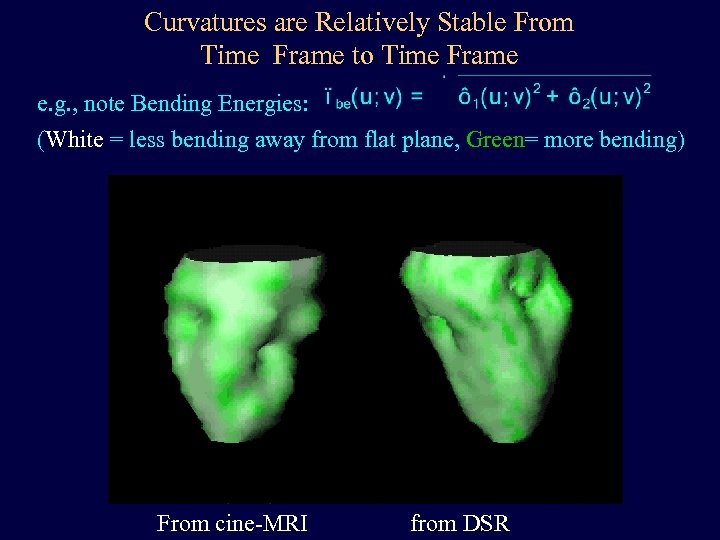 Curvatures are Relatively Stable From Time Frame to Time Frame e. g. , note Bending Energies: (White = less bending away from flat plane, Green= more bending) From cine-MRI from DSR
Curvatures are Relatively Stable From Time Frame to Time Frame e. g. , note Bending Energies: (White = less bending away from flat plane, Green= more bending) From cine-MRI from DSR
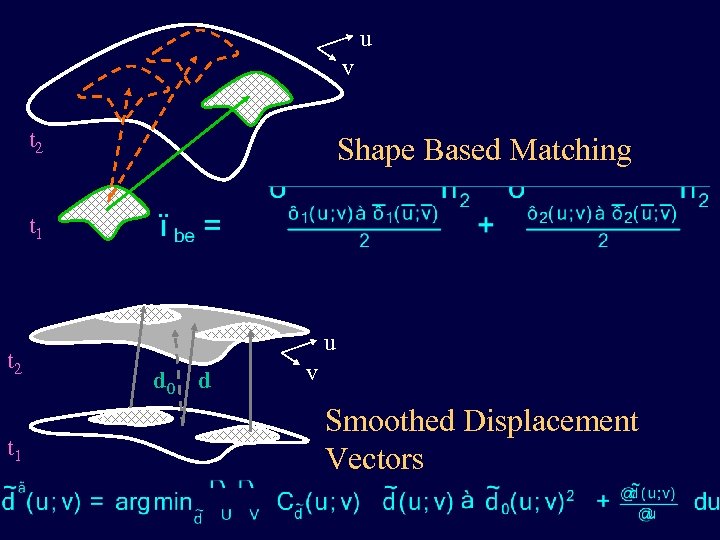 u v t 2 Shape Based Matching t 1 t 2 t 1 u d 0 d v Smoothed Displacement Vectors
u v t 2 Shape Based Matching t 1 t 2 t 1 u d 0 d v Smoothed Displacement Vectors
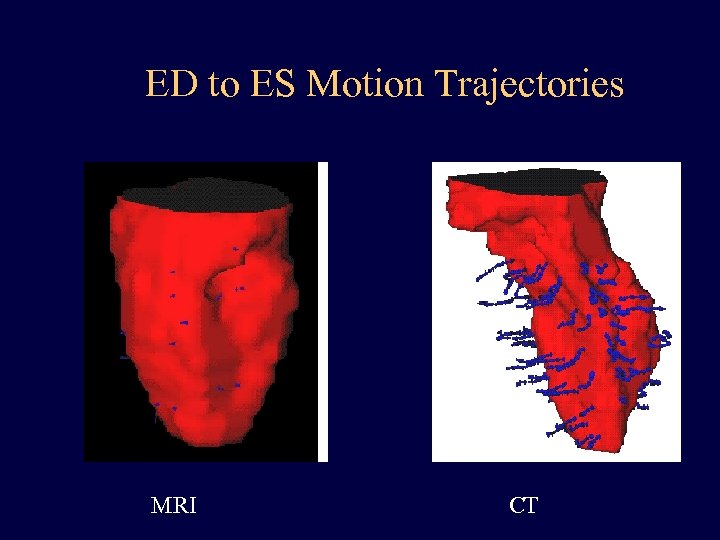 ED to ES Motion Trajectories MRI CT
ED to ES Motion Trajectories MRI CT
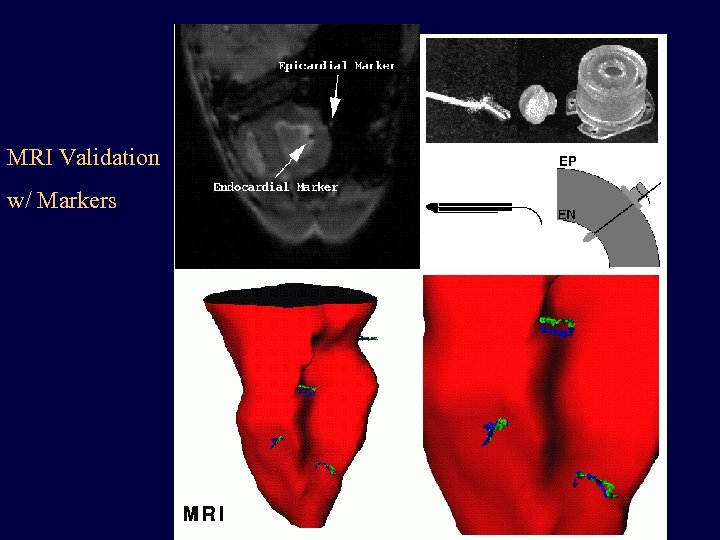 MRI Validation w/ Markers
MRI Validation w/ Markers
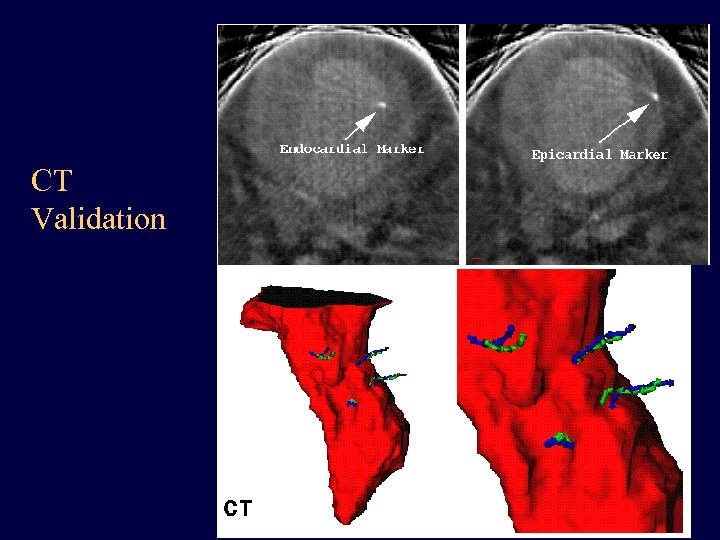 CT Validation
CT Validation
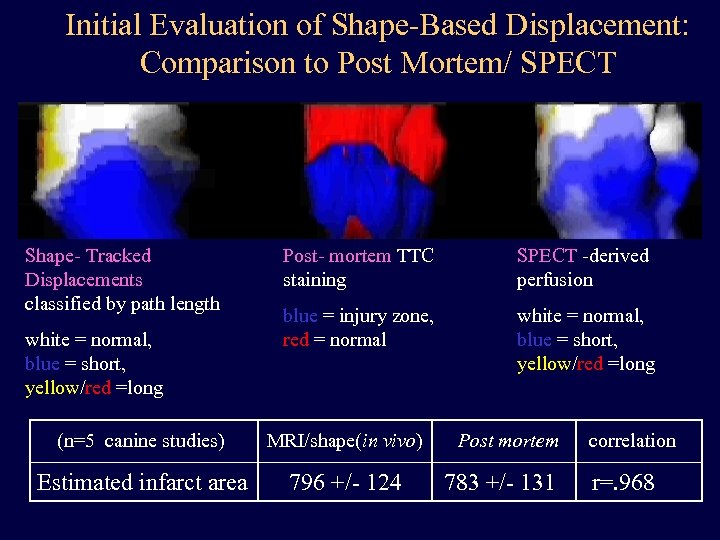 Initial Evaluation of Shape-Based Displacement: Comparison to Post Mortem/ SPECT Shape- Tracked Displacements classified by path length white = normal, blue = short, yellow/red =long Post- mortem TTC staining SPECT -derived perfusion blue = injury zone, red = normal white = normal, blue = short, yellow/red =long (n=5 canine studies) MRI/shape(in vivo) Estimated infarct area 796 +/- 124 Post mortem 783 +/- 131 correlation r=. 968
Initial Evaluation of Shape-Based Displacement: Comparison to Post Mortem/ SPECT Shape- Tracked Displacements classified by path length white = normal, blue = short, yellow/red =long Post- mortem TTC staining SPECT -derived perfusion blue = injury zone, red = normal white = normal, blue = short, yellow/red =long (n=5 canine studies) MRI/shape(in vivo) Estimated infarct area 796 +/- 124 Post mortem 783 +/- 131 correlation r=. 968
 Geometrical Models: Measurement of Neuroanatomical Structure
Geometrical Models: Measurement of Neuroanatomical Structure
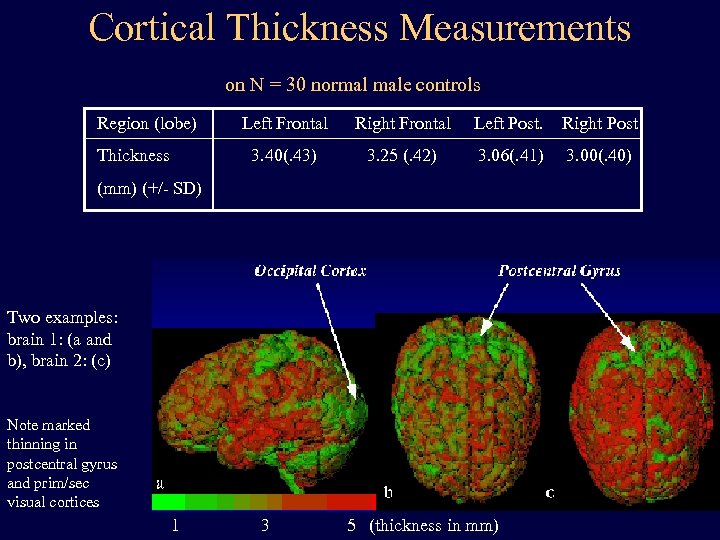 Cortical Thickness Measurements on N = 30 normal male controls: Region (lobe) Right Frontal Left Post. Right Post 3. 40(. 43) Thickness Left Frontal 3. 25 (. 42) 3. 06(. 41) 3. 00(. 40) (mm) (+/- SD) Two examples: brain 1: (a and b), brain 2: (c) Note marked thinning in postcentral gyrus and prim/sec visual cortices 1 3 5 (thickness in mm)
Cortical Thickness Measurements on N = 30 normal male controls: Region (lobe) Right Frontal Left Post. Right Post 3. 40(. 43) Thickness Left Frontal 3. 25 (. 42) 3. 06(. 41) 3. 00(. 40) (mm) (+/- SD) Two examples: brain 1: (a and b), brain 2: (c) Note marked thinning in postcentral gyrus and prim/sec visual cortices 1 3 5 (thickness in mm)
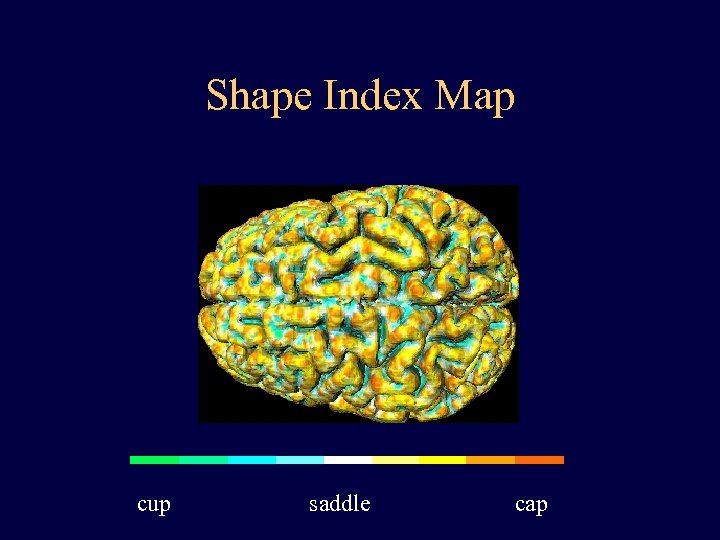 Shape Index Map cup saddle cap
Shape Index Map cup saddle cap
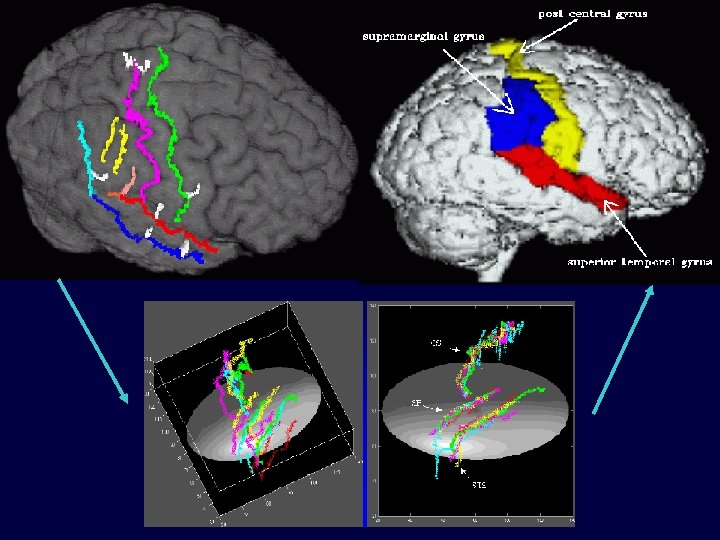
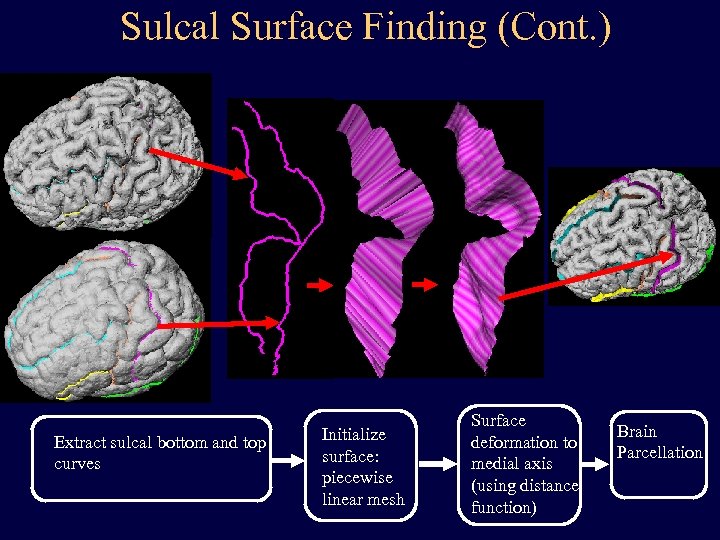 Sulcal Surface Finding (Cont. ) Extract sulcal bottom and top curves Initialize surface: piecewise linear mesh Surface deformation to medial axis (using distance function) Brain Parcellation
Sulcal Surface Finding (Cont. ) Extract sulcal bottom and top curves Initialize surface: piecewise linear mesh Surface deformation to medial axis (using distance function) Brain Parcellation
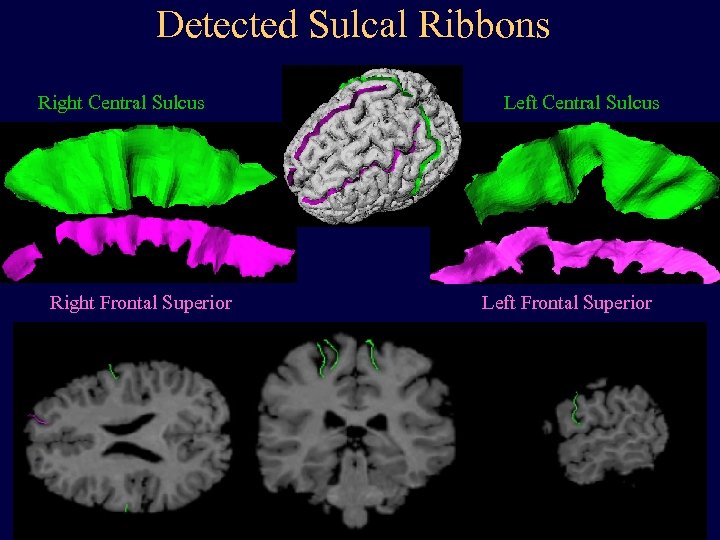 Detected Sulcal Ribbons Right Central Sulcus Right Frontal Superior Left Central Sulcus Left Frontal Superior
Detected Sulcal Ribbons Right Central Sulcus Right Frontal Superior Left Central Sulcus Left Frontal Superior
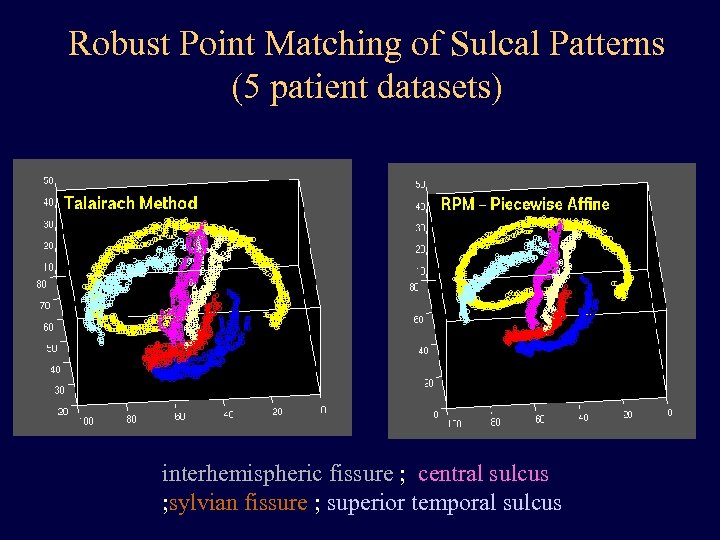 Robust Point Matching of Sulcal Patterns (5 patient datasets) interhemispheric fissure ; central sulcus ; sylvian fissure ; superior temporal sulcus
Robust Point Matching of Sulcal Patterns (5 patient datasets) interhemispheric fissure ; central sulcus ; sylvian fissure ; superior temporal sulcus
 Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
Outline I. Overview of Problem Areas - will focus on 3 heart/brain problems - types of 3 D/4 D Image Datasets II. Using Geometrically- Based Models - segmentation - motion tracking - neuroanatomical measurement III. Using Biomechanical Models - cardiac deformation analysis - brain shift compensation in neurosurgery
 Biomechanical Models: Quantifying Left Ventricular Deformation
Biomechanical Models: Quantifying Left Ventricular Deformation
 Recovery and Quantification of LV Strain • GOAL: Recover Dense 3 D Strains for Deforming Left Ventricle • DATA: Shape-based (or MR tag-based) Displacements, sometimes Phase Contrast MRI- based velocities • APPROACH: - Biomechanical models of the LV used to guide recovery and reporting of strain information - Tradeoffs as to when to use image-derived data and when to invoke model are made via data confidence measures.
Recovery and Quantification of LV Strain • GOAL: Recover Dense 3 D Strains for Deforming Left Ventricle • DATA: Shape-based (or MR tag-based) Displacements, sometimes Phase Contrast MRI- based velocities • APPROACH: - Biomechanical models of the LV used to guide recovery and reporting of strain information - Tradeoffs as to when to use image-derived data and when to invoke model are made via data confidence measures.
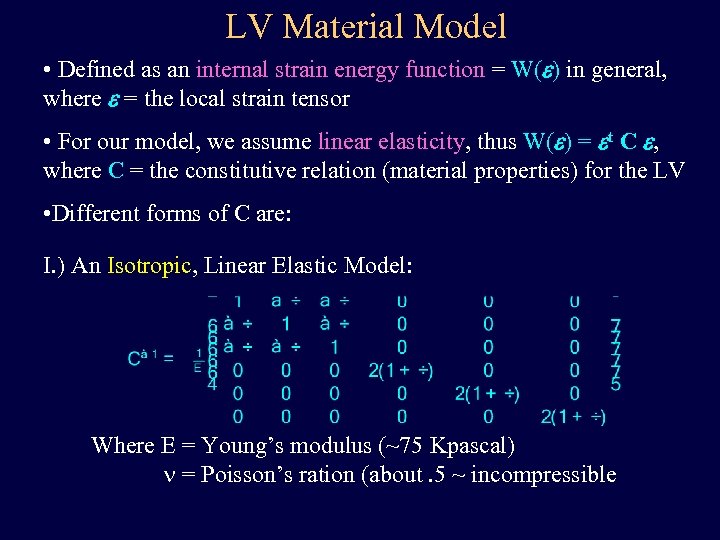 LV Material Model • Defined as an internal strain energy function = W(e) in general, where e = the local strain tensor • For our model, we assume linear elasticity, thus W(e) = et C e, where C = the constitutive relation (material properties) for the LV • Different forms of C are: I. ) An Isotropic, Linear Elastic Model: Where E = Young’s modulus (~75 Kpascal) n = Poisson’s ration (about. 5 ~ incompressible
LV Material Model • Defined as an internal strain energy function = W(e) in general, where e = the local strain tensor • For our model, we assume linear elasticity, thus W(e) = et C e, where C = the constitutive relation (material properties) for the LV • Different forms of C are: I. ) An Isotropic, Linear Elastic Model: Where E = Young’s modulus (~75 Kpascal) n = Poisson’s ration (about. 5 ~ incompressible
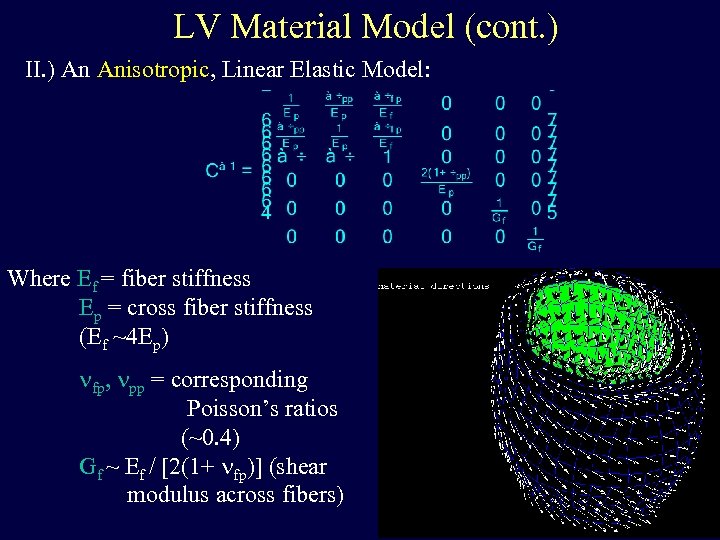 LV Material Model (cont. ) II. ) An Anisotropic, Linear Elastic Model: Where Ef = fiber stiffness Ep = cross fiber stiffness (Ef ~4 Ep) nfp, npp = corresponding Poisson’s ratios (~0. 4) Gf ~ Ef / [2(1+ nfp)] (shear modulus across fibers)
LV Material Model (cont. ) II. ) An Anisotropic, Linear Elastic Model: Where Ef = fiber stiffness Ep = cross fiber stiffness (Ef ~4 Ep) nfp, npp = corresponding Poisson’s ratios (~0. 4) Gf ~ Ef / [2(1+ nfp)] (shear modulus across fibers)
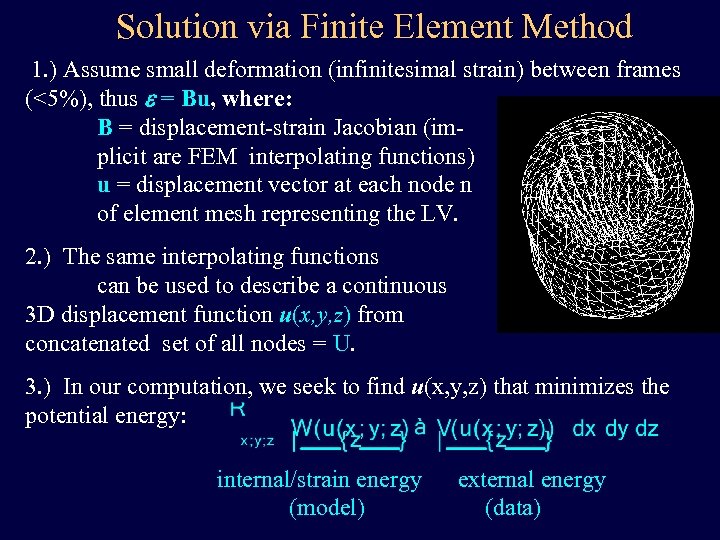 Solution via Finite Element Method 1. ) Assume small deformation (infinitesimal strain) between frames (<5%), thus e = Bu, where: B = displacement-strain Jacobian (implicit are FEM interpolating functions) u = displacement vector at each node n of element mesh representing the LV. 2. ) The same interpolating functions can be used to describe a continuous 3 D displacement function u(x, y, z) from concatenated set of all nodes = U. a 3. ) In our computation, we seek to find u(x, y, z) that minimizes the potential energy: internal/strain energy (model) external energy (data)
Solution via Finite Element Method 1. ) Assume small deformation (infinitesimal strain) between frames (<5%), thus e = Bu, where: B = displacement-strain Jacobian (implicit are FEM interpolating functions) u = displacement vector at each node n of element mesh representing the LV. 2. ) The same interpolating functions can be used to describe a continuous 3 D displacement function u(x, y, z) from concatenated set of all nodes = U. a 3. ) In our computation, we seek to find u(x, y, z) that minimizes the potential energy: internal/strain energy (model) external energy (data)
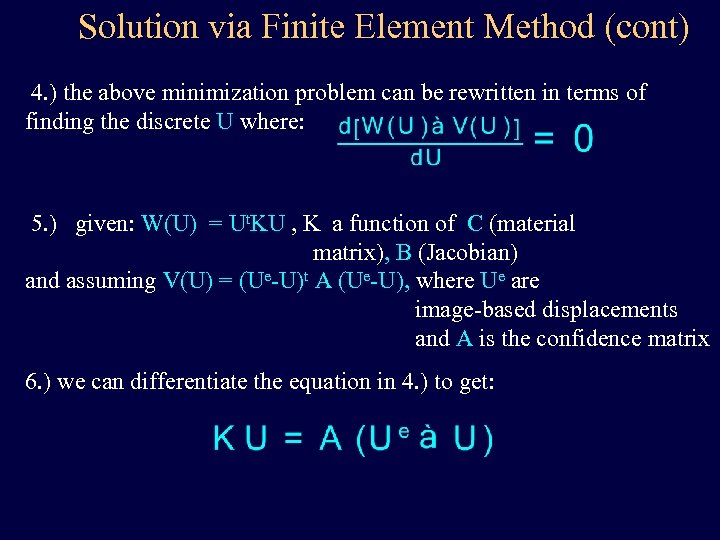 Solution via Finite Element Method (cont) 4. ) the above minimization problem can be rewritten in terms of finding the discrete U where: 5. ) given: W(U) = Ut. KU , K a function of C (material matrix), B (Jacobian) and assuming V(U) = (Ue-U)t A (Ue-U), where Ue are image-based displacements and A is the confidence matrix 6. ) we can differentiate the equation in 4. ) to get:
Solution via Finite Element Method (cont) 4. ) the above minimization problem can be rewritten in terms of finding the discrete U where: 5. ) given: W(U) = Ut. KU , K a function of C (material matrix), B (Jacobian) and assuming V(U) = (Ue-U)t A (Ue-U), where Ue are image-based displacements and A is the confidence matrix 6. ) we can differentiate the equation in 4. ) to get:
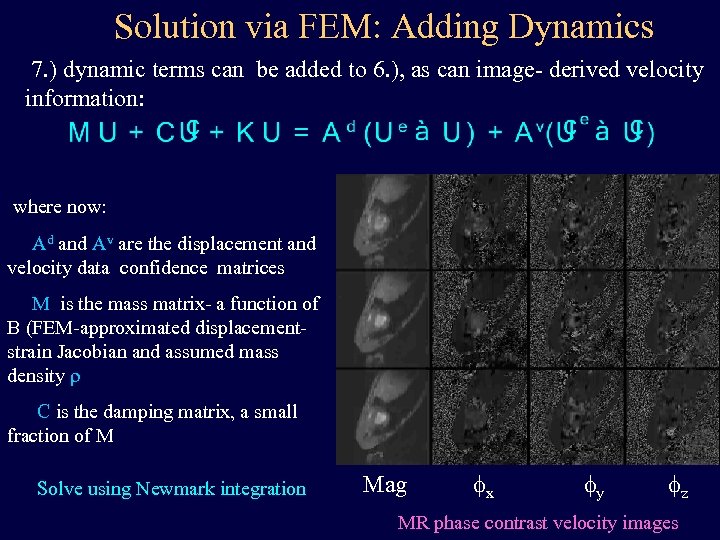 Solution via FEM: Adding Dynamics 7. ) dynamic terms can be added to 6. ), as can image- derived velocity information: where now: Ad and Av are the displacement and velocity data confidence matrices M is the mass matrix- a function of B (FEM-approximated displacementstrain Jacobian and assumed mass density r C is the damping matrix, a small fraction of M Solve using Newmark integration Mag fx fy fz MR phase contrast velocity images
Solution via FEM: Adding Dynamics 7. ) dynamic terms can be added to 6. ), as can image- derived velocity information: where now: Ad and Av are the displacement and velocity data confidence matrices M is the mass matrix- a function of B (FEM-approximated displacementstrain Jacobian and assumed mass density r C is the damping matrix, a small fraction of M Solve using Newmark integration Mag fx fy fz MR phase contrast velocity images
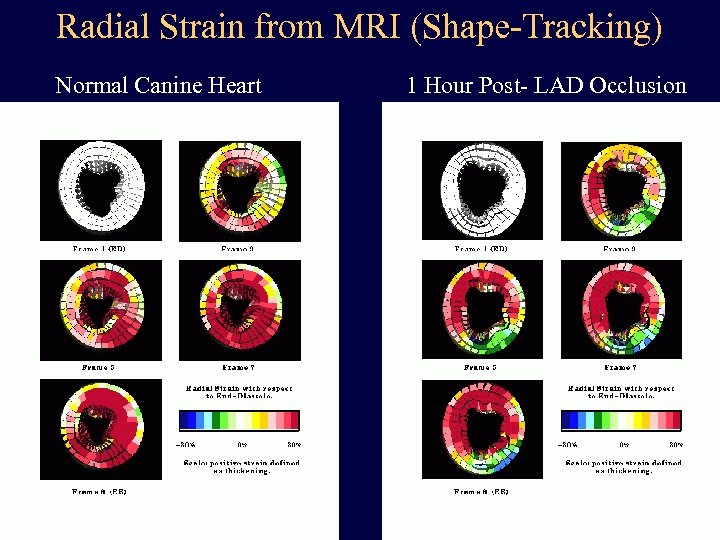 Radial Strain from MRI (Shape-Tracking) Normal Canine Heart 1 Hour Post- LAD Occlusion
Radial Strain from MRI (Shape-Tracking) Normal Canine Heart 1 Hour Post- LAD Occlusion
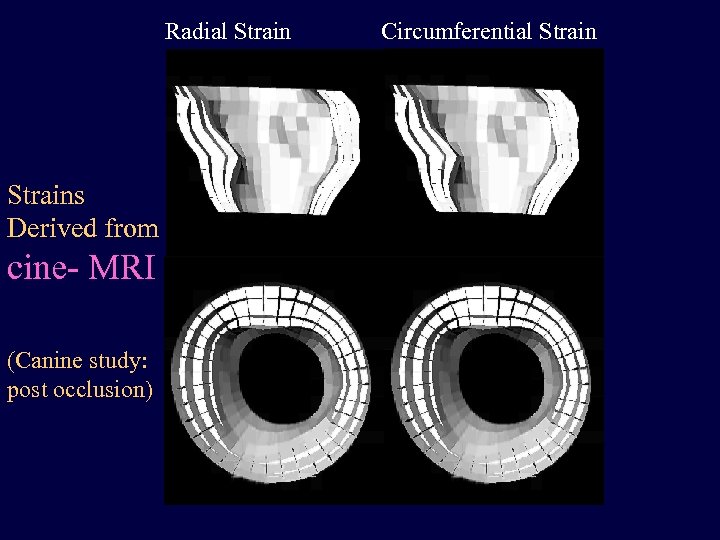 Radial Strains Derived from cine- MRI (Canine study: post occlusion) Circumferential Strain
Radial Strains Derived from cine- MRI (Canine study: post occlusion) Circumferential Strain
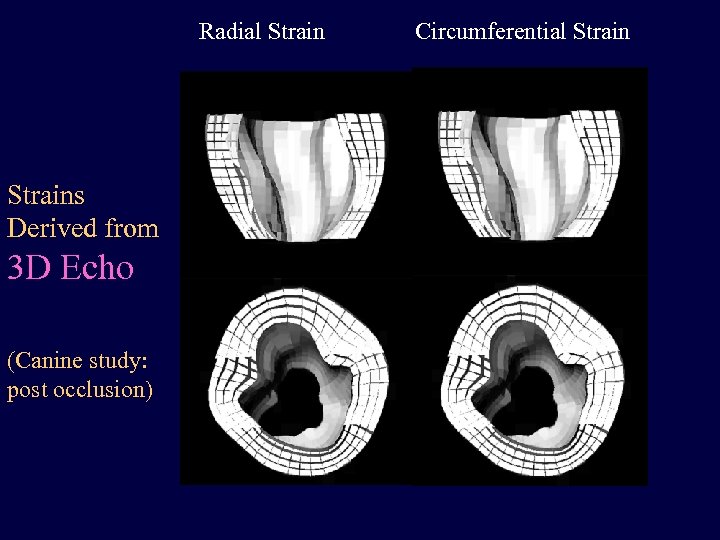 Radial Strains Derived from 3 D Echo (Canine study: post occlusion) Circumferential Strain
Radial Strains Derived from 3 D Echo (Canine study: post occlusion) Circumferential Strain
 Biomechanical Models: Brain Shift Compensation for Image- Guided Neurosurgery
Biomechanical Models: Brain Shift Compensation for Image- Guided Neurosurgery
 Previous Efforts in Brain Shift Estimation • Bucholz, et al. , (CVRMed 1997) … used Intraoperative ultrasound and 2 D spring -based mechanical model (looked at a variety of structure pre/post op on n=23 pts) … (2 D datasets only) • Hill, Fitzpatrick, et al. , (CVRMed 1997) … measured brain shifts relative to skull on 5 patients using ACUSTAR I (fiducial-based) surgical navigation system. . . (measurements only) • Edwards, Hill, Little, Hawkes, (IPMI 1997)… 3 component (rigid, deformable, fluid), physically-motivated, energy-based model; used landmark-like data. . . (2 Dran in 2 -3 hours on SPARC 20) • Kyriacou, Davatzikos, (MICCAI 1998) … 2 D Continuum model via FEMaimed at larger deformations (e. g. tumors)… (so far 2 D- primarily simulation) • Miga, Paulsen, et al. , (MICCAI 1998)… 3 D Continuum/FEM model tested/validated on Porcine brains… (nice effort- primarily forward modeling)
Previous Efforts in Brain Shift Estimation • Bucholz, et al. , (CVRMed 1997) … used Intraoperative ultrasound and 2 D spring -based mechanical model (looked at a variety of structure pre/post op on n=23 pts) … (2 D datasets only) • Hill, Fitzpatrick, et al. , (CVRMed 1997) … measured brain shifts relative to skull on 5 patients using ACUSTAR I (fiducial-based) surgical navigation system. . . (measurements only) • Edwards, Hill, Little, Hawkes, (IPMI 1997)… 3 component (rigid, deformable, fluid), physically-motivated, energy-based model; used landmark-like data. . . (2 Dran in 2 -3 hours on SPARC 20) • Kyriacou, Davatzikos, (MICCAI 1998) … 2 D Continuum model via FEMaimed at larger deformations (e. g. tumors)… (so far 2 D- primarily simulation) • Miga, Paulsen, et al. , (MICCAI 1998)… 3 D Continuum/FEM model tested/validated on Porcine brains… (nice effort- primarily forward modeling)
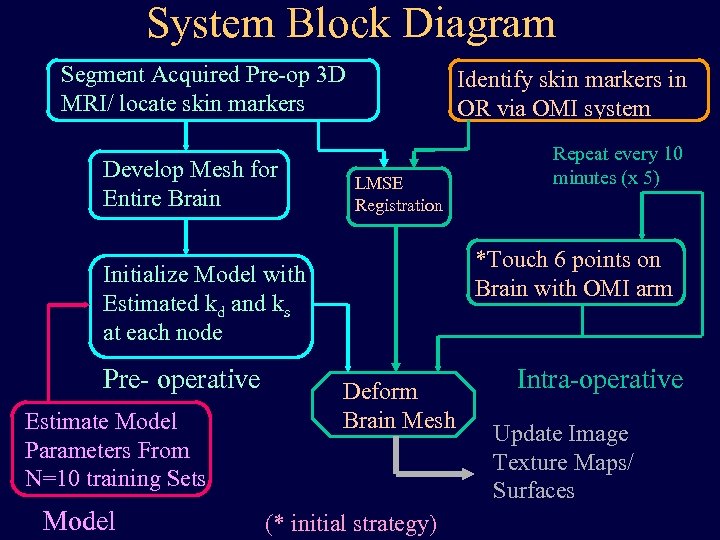 System Block Diagram Segment Acquired Pre-op 3 D MRI/ locate skin markers Develop Mesh for Entire Brain Identify skin markers in OR via OMI system LMSE Registration *Touch 6 points on Brain with OMI arm Initialize Model with Estimated kd and ks at each node Pre- operative Estimate Model Parameters From N=10 training Sets Model Repeat every 10 minutes (x 5) Deform Brain Mesh (* initial strategy) Intra-operative Update Image Texture Maps/ Surfaces
System Block Diagram Segment Acquired Pre-op 3 D MRI/ locate skin markers Develop Mesh for Entire Brain Identify skin markers in OR via OMI system LMSE Registration *Touch 6 points on Brain with OMI arm Initialize Model with Estimated kd and ks at each node Pre- operative Estimate Model Parameters From N=10 training Sets Model Repeat every 10 minutes (x 5) Deform Brain Mesh (* initial strategy) Intra-operative Update Image Texture Maps/ Surfaces
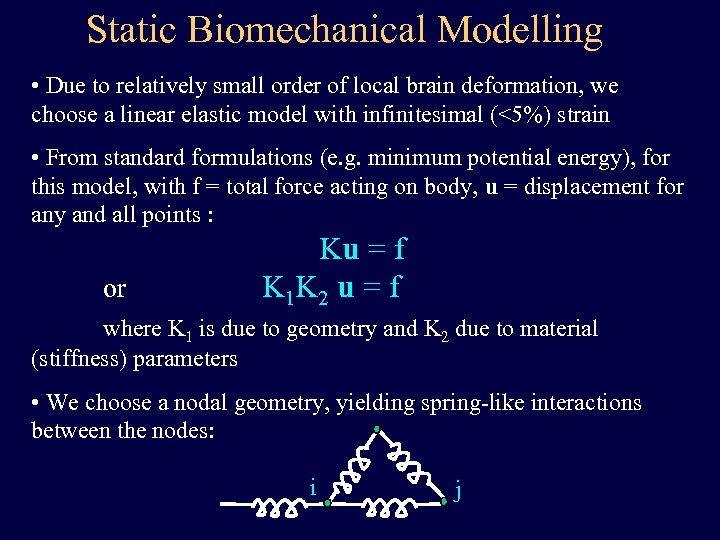 Static Biomechanical Modelling • Due to relatively small order of local brain deformation, we choose a linear elastic model with infinitesimal (<5%) strain • From standard formulations (e. g. minimum potential energy), for this model, with f = total force acting on body, u = displacement for any and all points : or Ku = f K 1 K 2 u = f where K 1 is due to geometry and K 2 due to material (stiffness) parameters • We choose a nodal geometry, yielding spring-like interactions between the nodes: i j
Static Biomechanical Modelling • Due to relatively small order of local brain deformation, we choose a linear elastic model with infinitesimal (<5%) strain • From standard formulations (e. g. minimum potential energy), for this model, with f = total force acting on body, u = displacement for any and all points : or Ku = f K 1 K 2 u = f where K 1 is due to geometry and K 2 due to material (stiffness) parameters • We choose a nodal geometry, yielding spring-like interactions between the nodes: i j
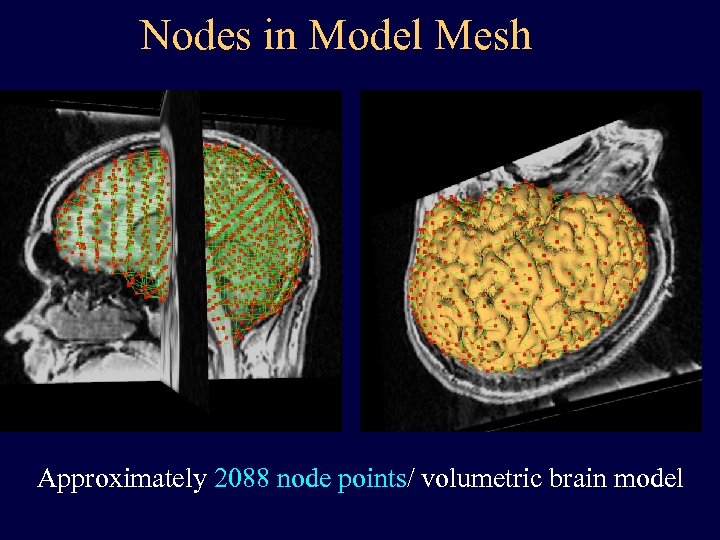 Nodes in Model Mesh Approximately 2088 node points/ volumetric brain model
Nodes in Model Mesh Approximately 2088 node points/ volumetric brain model
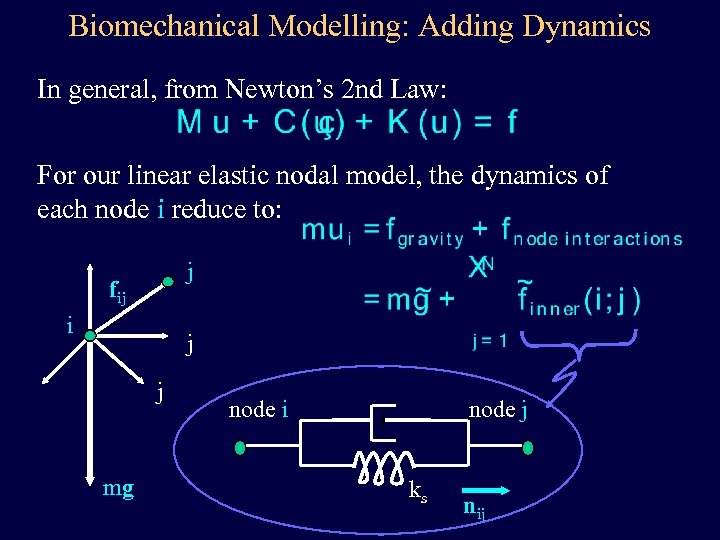 Biomechanical Modelling: Adding Dynamics In general, from Newton’s 2 nd Law: For our linear elastic nodal model, the dynamics of each node i reduce to: j fij i j j mg node i node j ks nij
Biomechanical Modelling: Adding Dynamics In general, from Newton’s 2 nd Law: For our linear elastic nodal model, the dynamics of each node i reduce to: j fij i j j mg node i node j ks nij
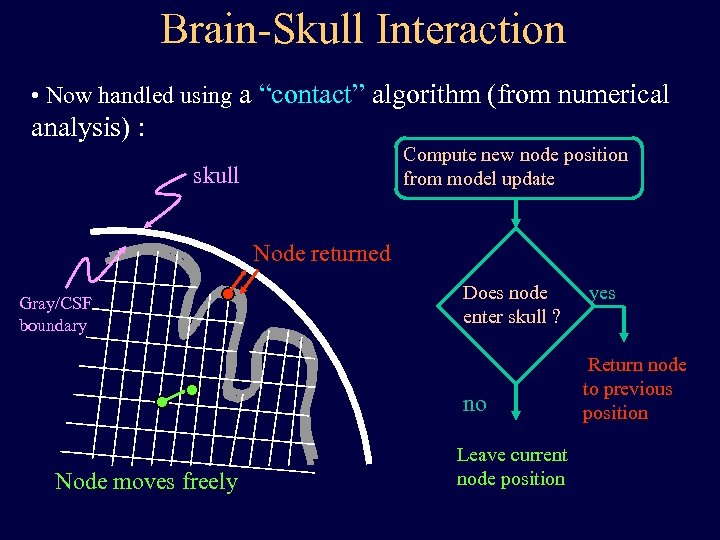 Brain-Skull Interaction • Now handled using a “contact” algorithm (from numerical analysis) : Compute new node position from model update skull Node returned Gray/CSF boundary Does node enter skull ? no Node moves freely Leave current node position yes Return node to previous position
Brain-Skull Interaction • Now handled using a “contact” algorithm (from numerical analysis) : Compute new node position from model update skull Node returned Gray/CSF boundary Does node enter skull ? no Node moves freely Leave current node position yes Return node to previous position
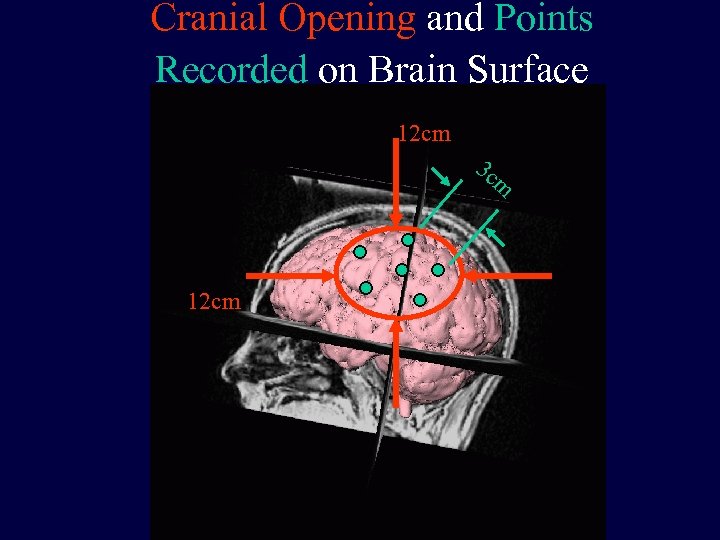 Cranial Opening and Points Recorded on Brain Surface 12 cm 3 c m 12 cm
Cranial Opening and Points Recorded on Brain Surface 12 cm 3 c m 12 cm
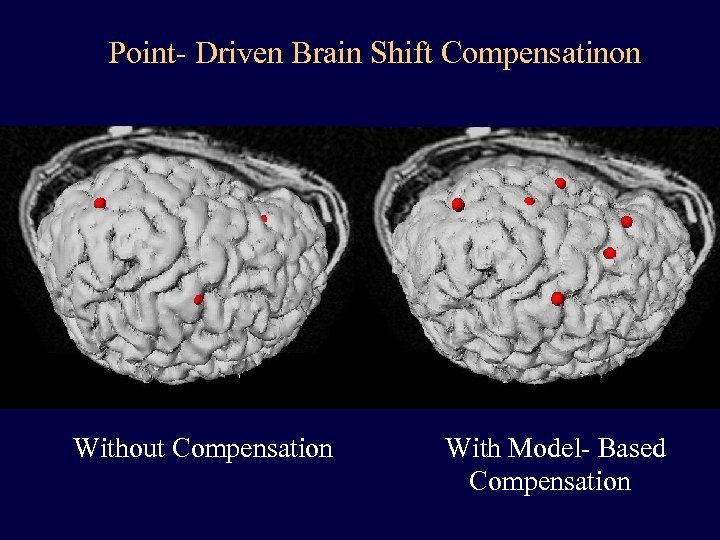 Point- Driven Brain Shift Compensatinon Without Compensation With Model- Based Compensation
Point- Driven Brain Shift Compensatinon Without Compensation With Model- Based Compensation
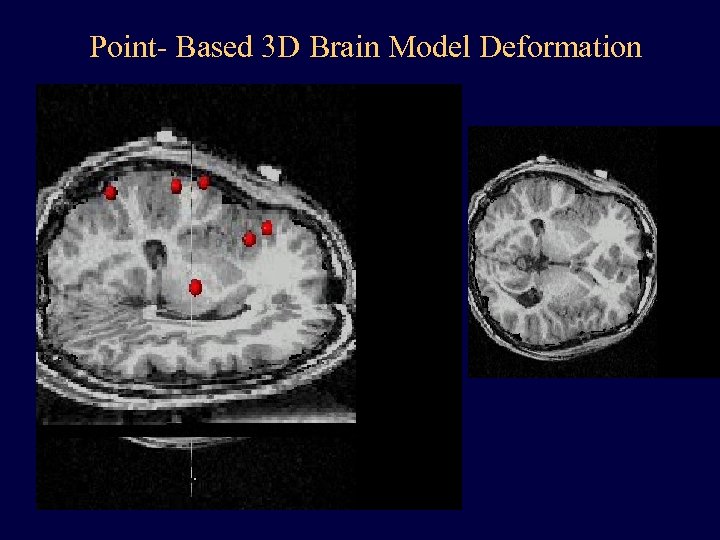 Point- Based 3 D Brain Model Deformation
Point- Based 3 D Brain Model Deformation
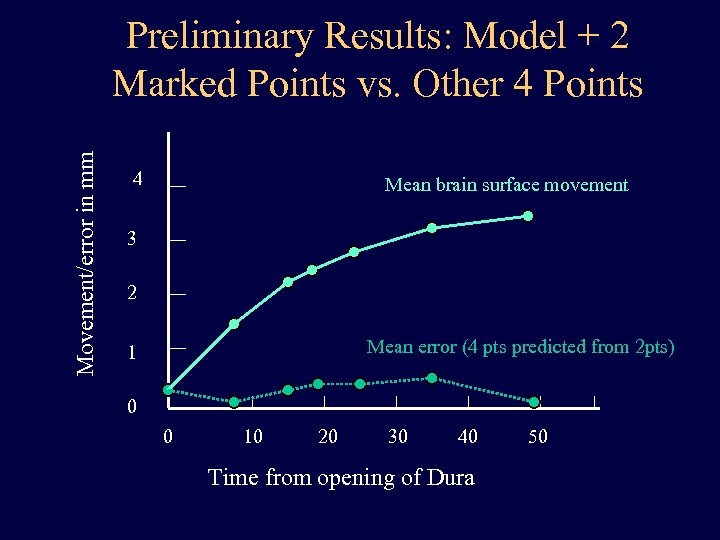 Movement/error in mm Preliminary Results: Model + 2 Marked Points vs. Other 4 Points 4 Mean brain surface movement 3 2 Mean error (4 pts predicted from 2 pts) 1 0 0 10 20 30 40 Time from opening of Dura 50
Movement/error in mm Preliminary Results: Model + 2 Marked Points vs. Other 4 Points 4 Mean brain surface movement 3 2 Mean error (4 pts predicted from 2 pts) 1 0 0 10 20 30 40 Time from opening of Dura 50
 Alternative Intra-operative Data Acquisition Strategies • Dual visible light cameras (recover surface via stereo) • Structured light (surface only) • ultrasound probe (2 D/3 D) • Intra-operative MRI • Portable CT scanner (e. g. Philips)
Alternative Intra-operative Data Acquisition Strategies • Dual visible light cameras (recover surface via stereo) • Structured light (surface only) • ultrasound probe (2 D/3 D) • Intra-operative MRI • Portable CT scanner (e. g. Philips)


