9eb6325e065c2c59f7c1f86447f6d72b.ppt
- Количество слайдов: 55

Text Mining in Biomedicine Michael Krauthammer Department of Pathology Yale University School of Medicine

Definition • Text mining is – – – the process of automatically extracting knowledge from large text collections data mining applied to text documents / knowledge discovery from text a modular process similar to reading, where facts from different articles / books are combined for novel inference (de Bruijn 2002)
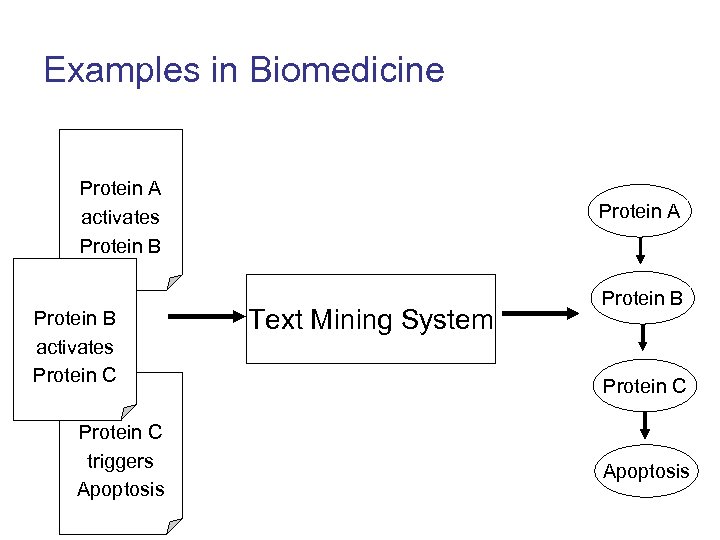
Examples in Biomedicine Protein A activates Protein B activates Protein C triggers Apoptosis Protein A Text Mining System Protein B Protein C Apoptosis
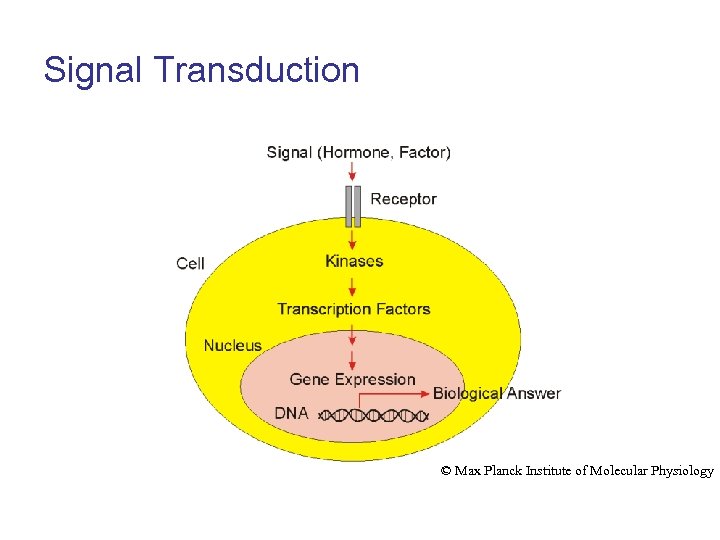
Signal Transduction © Max Planck Institute of Molecular Physiology
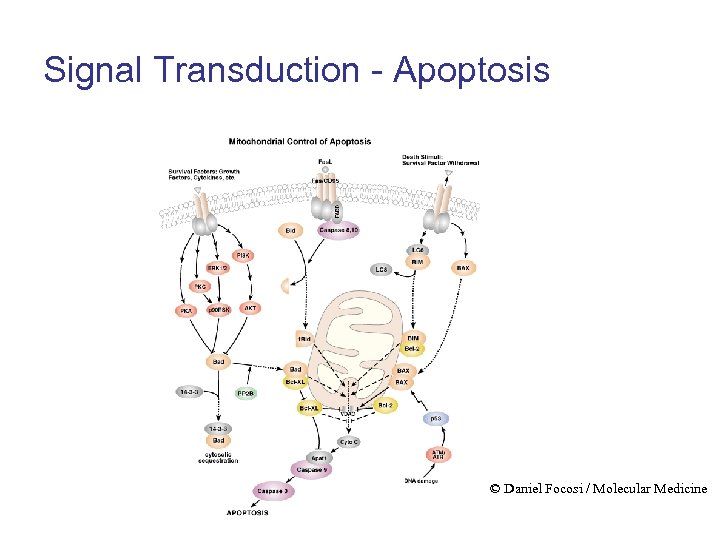
Signal Transduction - Apoptosis © Daniel Focosi / Molecular Medicine
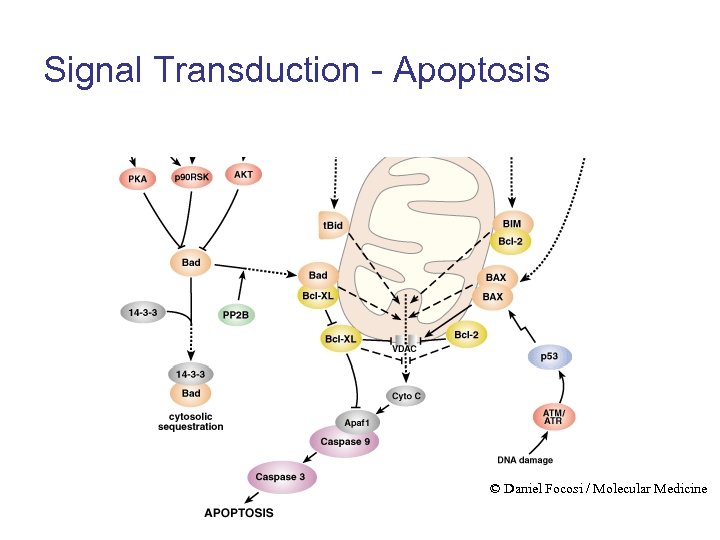
Signal Transduction - Apoptosis © Daniel Focosi / Molecular Medicine
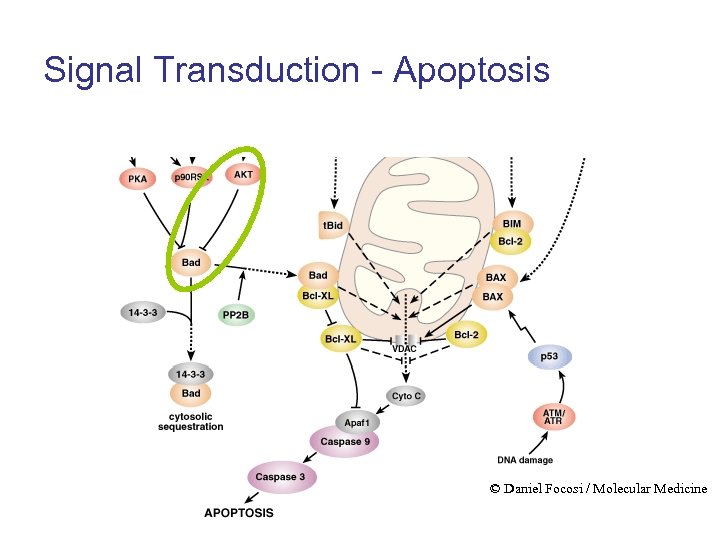
Signal Transduction - Apoptosis © Daniel Focosi / Molecular Medicine
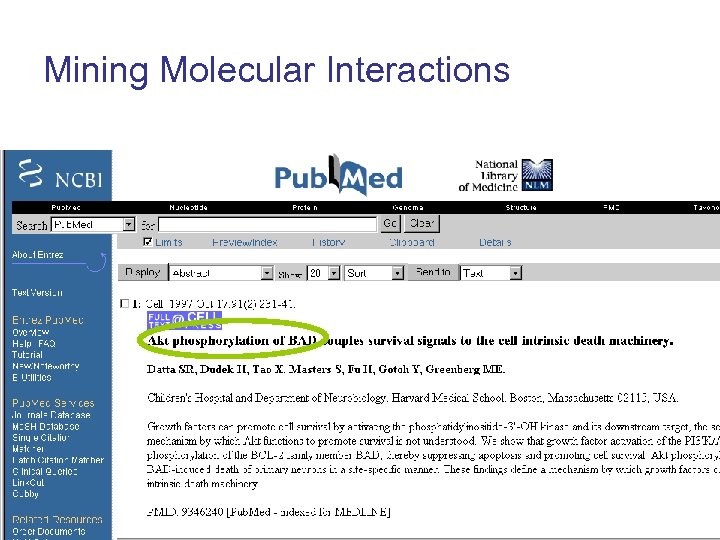

Mining Molecular Interactions

Information Explosion
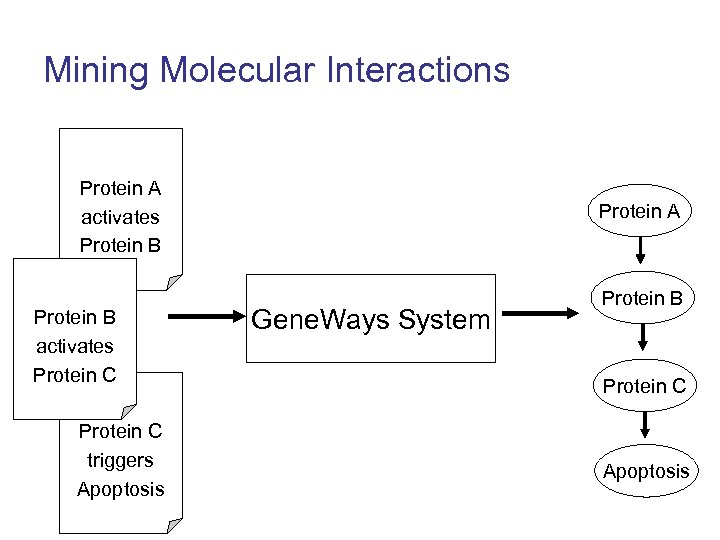
Mining Molecular Interactions Protein A activates Protein B activates Protein C triggers Apoptosis Protein A Gene. Ways System Protein B Protein C Apoptosis
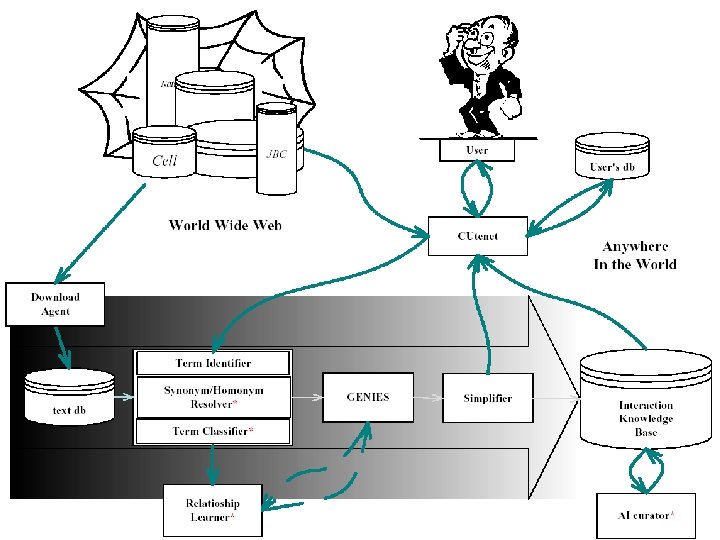
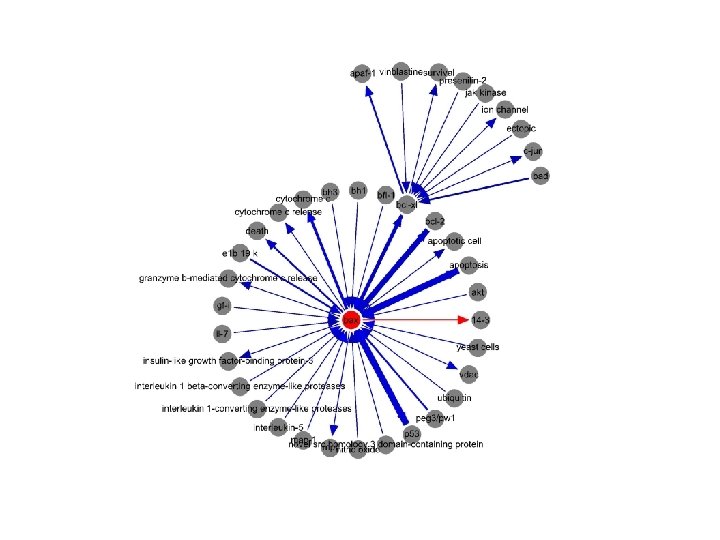

Network-based Candidate Gene Prediction
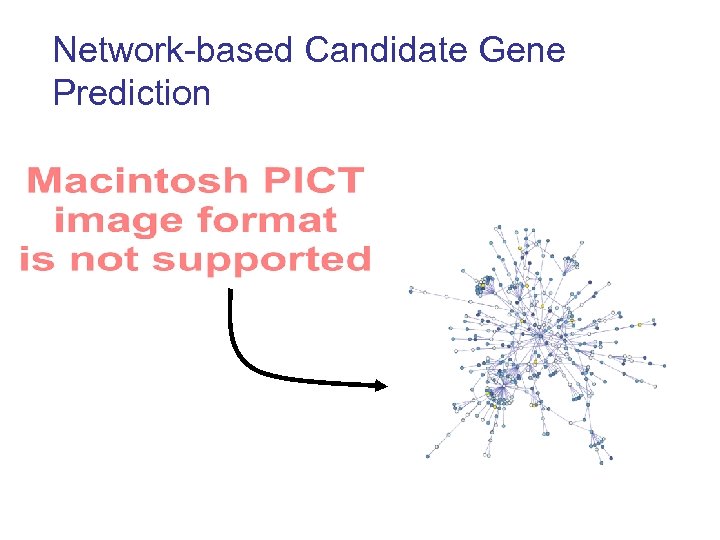
Network-based Candidate Gene Prediction
Network-based Candidate Gene Prediction
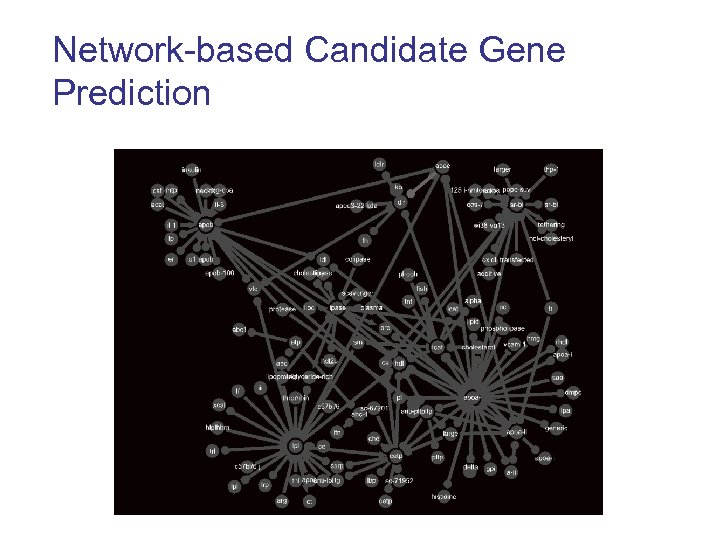
Network-based Candidate Gene Prediction
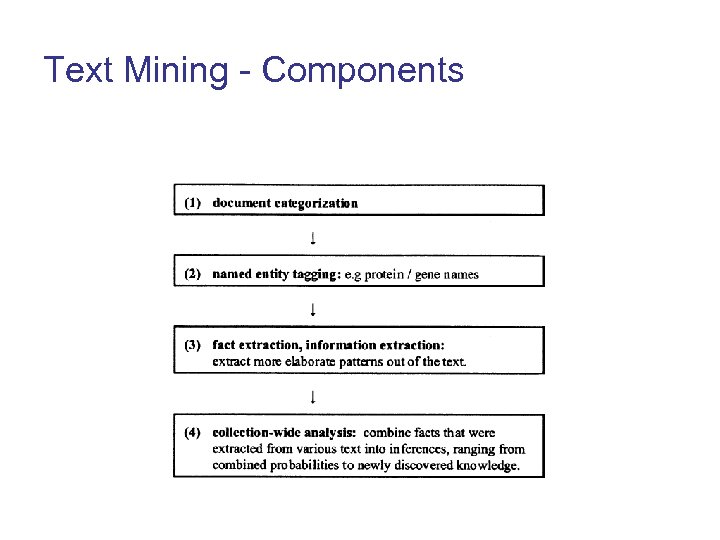

Text Mining - Components

Information Extraction • Information Extraction: “the activity of populating a structured information source (or database) from an unstructured, or free text, information source” (Gauzuskas & Wilks 1998)

Information Extraction • Many information sources are free text: • • • Law (Court Orders) Academic Research (Research Articles) Finance (Quarterly Reports) Medicine (Discharge Summaries) Biology (Molecular Interactions) • Data analysis on free text is difficult • Transformation of free text into structured data (machine-readable)
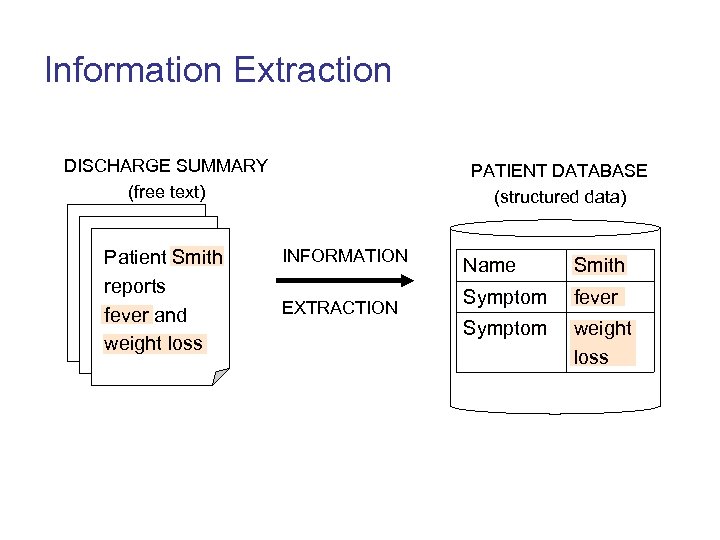
Information Extraction DISCHARGE SUMMARY (free text) Patient Smith reports fever and weight loss PATIENT DATABASE (structured data) INFORMATION EXTRACTION Name Smith Symptom fever Symptom weight loss
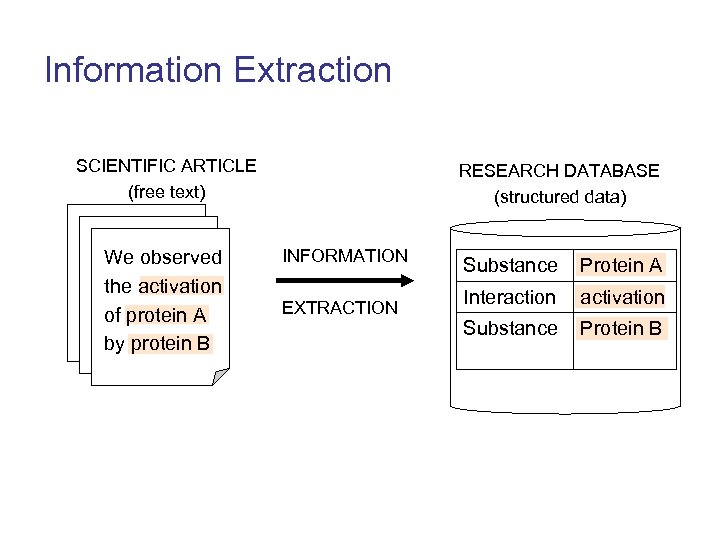
Information Extraction SCIENTIFIC ARTICLE (free text) We observed the activation of protein A by protein B RESEARCH DATABASE (structured data) INFORMATION EXTRACTION Substance Protein A Interaction activation Substance Protein B
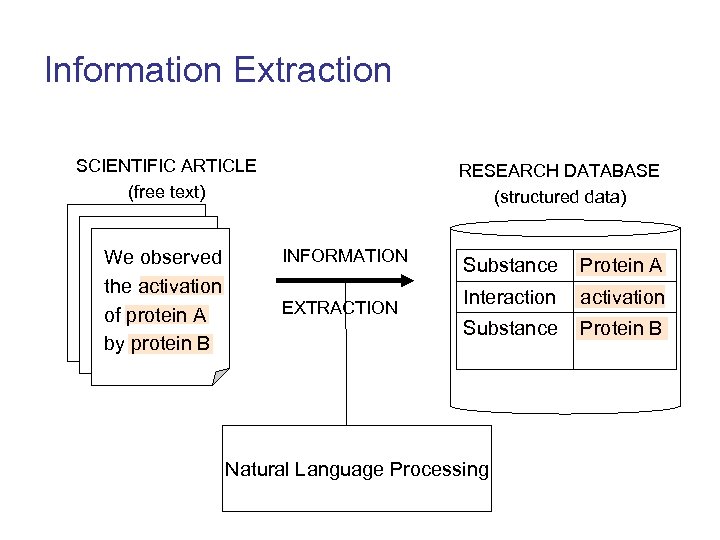
Information Extraction SCIENTIFIC ARTICLE (free text) We observed the activation of protein A by protein B RESEARCH DATABASE (structured data) INFORMATION EXTRACTION Substance Protein A Interaction activation Substance Protein B Natural Language Processing
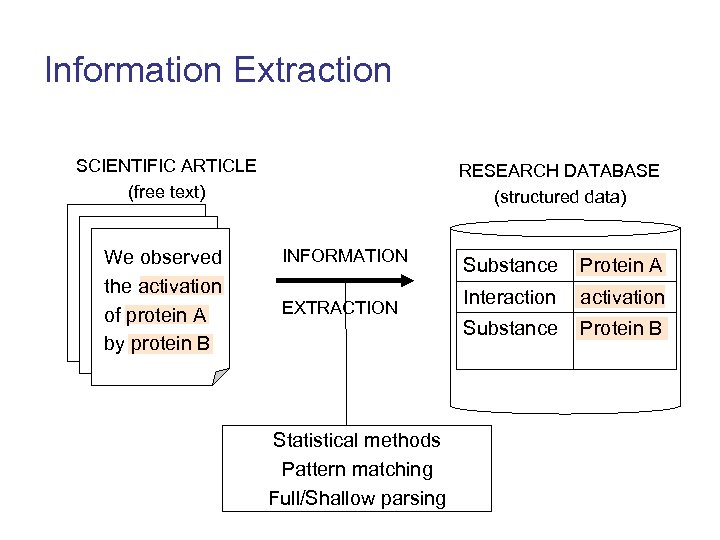
Information Extraction SCIENTIFIC ARTICLE (free text) We observed the activation of protein A by protein B RESEARCH DATABASE (structured data) INFORMATION EXTRACTION Statistical methods Pattern matching Full/Shallow parsing Substance Protein A Interaction activation Substance Protein B
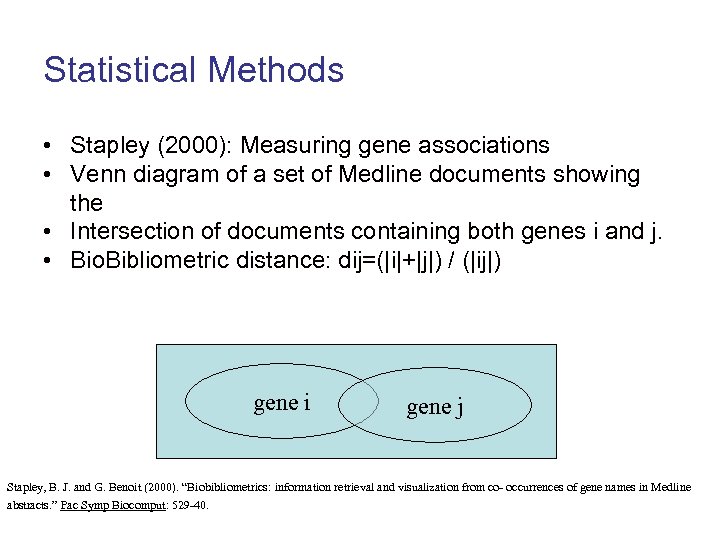

Statistical Methods • Stapley (2000): Measuring gene associations • Venn diagram of a set of Medline documents showing the • Intersection of documents containing both genes i and j. • Bio. Bibliometric distance: dij=(|i|+|j|) / (|ij|) gene i gene j Stapley, B. J. and G. Benoit (2000). “Biobibliometrics: information retrieval and visualization from co- occurrences of gene names in Medline abstracts. ” Pac Symp Biocomput: 529 -40.

Pattern Matching • Pattern matching (~regexp) to extract protein interactions • <gene> <interact with> <gene> Blaschke, C. , M. A. Andrade, et al. (1999). “Automatic extraction of biological information from scientific text: protein-protein interactions. ” Proc Int Conf Intell Syst Mol Biol: 60 -7. Ng, S. K. and M. Wong (1999). “Toward Routine Automatic Pathway Discovery from On-line Scientific Text Abstracts. ” Genome Inform Ser Workshop Genome Inform 10: 104 -112. Ono, T. , H. Hishigaki, et al. (2001). “Automated extraction of information on protein-protein interactions from the biological literature. ” Bioinformatics 17(2): 155 -61.

Full Parsing • Parsing: Detect sequence of grammar rules that describe internal structure of sentence • Grammar rule: S -> NP VP • [The house]NP [was demolished]VP. • Syntax parse tree:

Full Parsing • • Language Parsing in Biomedicine Med. LEE and GENIES semantic grammar parsers Columbia University, Dr. Carol Friedman Med. LEE: Clinical medicine parser: discharge summaries, radiology reports, pathology reports • the patient has a family history of coronary artery disease <problem v = "disease" idref = "p 64"> <bodyloc v = "coronary artery" idref = "p 60">/bodyloc> <status v=”family history”> </status> </problem>
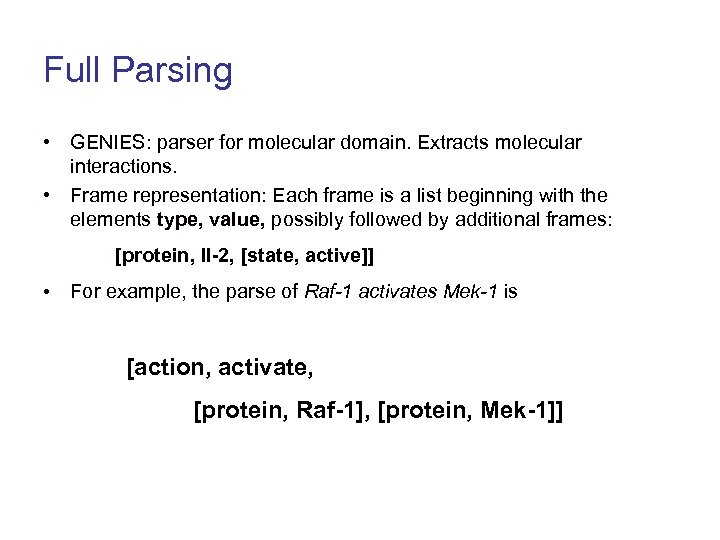

Full Parsing • GENIES: parser for molecular domain. Extracts molecular interactions. • Frame representation: Each frame is a list beginning with the elements type, value, possibly followed by additional frames: [protein, Il-2, [state, active]] • For example, the parse of Raf-1 activates Mek-1 is [action, activate, [protein, Raf-1], [protein, Mek-1]]

Full Parsing • Handles nested sentences (context free language): • mediation of sonic hedgehog-induced expression of Coup-Tfii by a protein phosphatase [action, promote, [geneorprotein, phosphatase], [action, activate, [geneorprotein, sonic hedgehog], [action, express, X, [geneorprotein, Coup-Tfii]]]]

Full Parsing Hafner, C. D. , K. Baclawski, et al. (1994). “Creating a knowledge base of biological research papers. ” Proc Int Conf Intell Syst Mol Biol 2: 147 -55. Friedman, C. , P. Kra, et al. (2001). GENIES: A Natural-Language System for the Extraction of Molecular Pathways from Complete Journal Articles. Proc Int Conf Intell Syst Mol Biol, Kopenhagen. Yakushiji A, Tateisi Y, Miyao Y, Tsujii J. Event extraction from biomedical papers using a full parser. Pac Symp Biocomput. 2001: 408 -19. Mc. Donald DM, Chen H, Su H, Marshall BB. Extracting gene pathway relations using a hybrid grammar: the Arizona relation parser. Bioinformatics. 2004 Jul 15 Leroy G, Chen H, Martinez JD. A shallow parser based on closed-class words to capture relations in biomedical text. J Biomed Inform. 2003 Jun; 36(3): 145 -58. Koike A, Niwa Y, Takagi T. Automatic extraction of gene/protein biological functions from biomedical text. Bioinformatics. 2004 Oct 27 Daraselia N, Yuryev A, Egorov S, Novichkova S, Nikitin A, Mazo I. Extracting human protein interactions from MEDLINE using a full-sentence parser. Bioinformatics. 2004 Mar 22; 20(5): 604 -11. Epub 2004 Jan 22
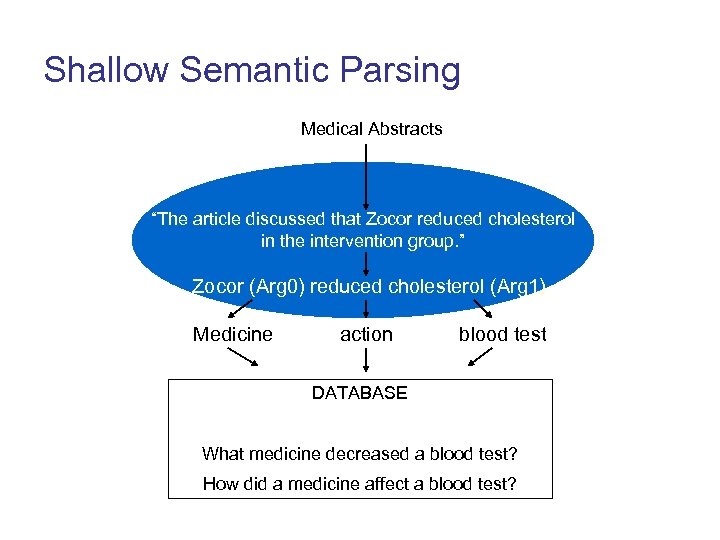

Shallow Semantic Parsing Medical Abstracts “The article discussed that Zocor reduced cholesterol in the intervention group. ” Zocor (Arg 0) reduced cholesterol (Arg 1) Medicine action blood test DATABASE What medicine decreased a blood test? How did a medicine affect a blood test?

Shallow Semantic Parsing • Shallow Semantic Parsing Technique (SSPT) – Successfully applied in non-medical domain* – “Predicate-centric” – Dissect sentences into simple WHAT did WHAT to WHOM/WHAT, and Modifiers (WHEN, WHERE, WHY and HOW) • The article discussed that Zocor (What) reduced (did What) cholesterol (to What) in the intervention group (modifiers). – Thus two core arguments, “Zocor” (Argument 0) and “cholesterol” (Argument 1), are related by the predicate “reduce(d)” – Modifier “in the intervention group” – “The article discussed that” is a null argument, i. e. it is not part of the predicate arguments. * S. Pradhan, D. Jurafsky, et al. In Proc. Of NAACL-HLT 2004.

Treebank and Propbank • Treebank contains the Wall Street Journal (WSJ) corpus annotate with syntactic information • Propbank annotates the same WSJ corpus found in Treebank with semantic information • Given the syntactic and semantic features, we can build a machine learning-based Information Extraction (IE) system, using shallow semantic parsing • Advantage of using Treebank and Propbank is its re-use of an existing corpora to do ‘free’ information extraction in the medical domain
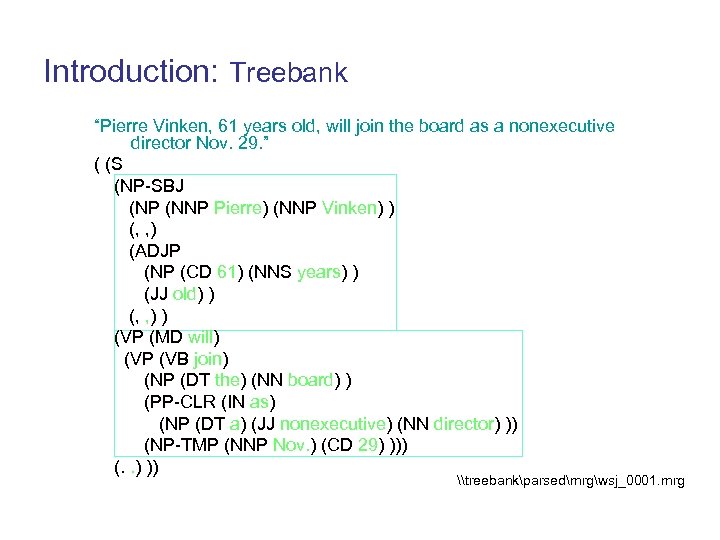

Introduction: Treebank “Pierre Vinken, 61 years old, will join the board as a nonexecutive director Nov. 29. ” ( (S (NP-SBJ (NP (NNP Pierre) (NNP Vinken) ) (, , ) (ADJP (NP (CD 61) (NNS years) ) (JJ old) ) (, , ) ) (VP (MD will) (VP (VB join) (NP (DT the) (NN board) ) (PP-CLR (IN as) (NP (DT a) (JJ nonexecutive) (NN director) )) (NP-TMP (NNP Nov. ) (CD 29) ))) (. . ) )) \treebankparsedmrgwsj_0001. mrg
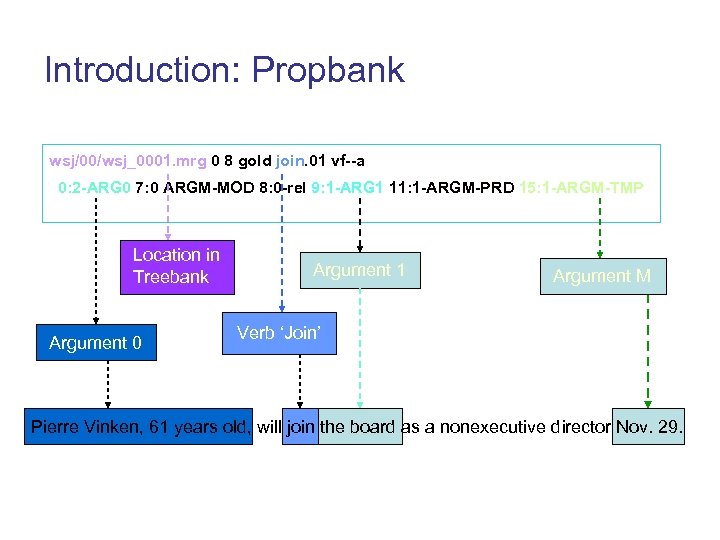
Introduction: Propbank wsj/00/wsj_0001. mrg 0 8 gold join. 01 vf--a 0: 2 -ARG 0 7: 0 ARGM-MOD 8: 0 -rel 9: 1 -ARG 1 11: 1 -ARGM-PRD 15: 1 -ARGM-TMP Location in Treebank Argument 0 Argument 1 Argument M Verb ‘Join’ Pierre Vinken, 61 years old, will join the board as a nonexecutive director Nov. 29.
Overall idea Syntax From Treebank
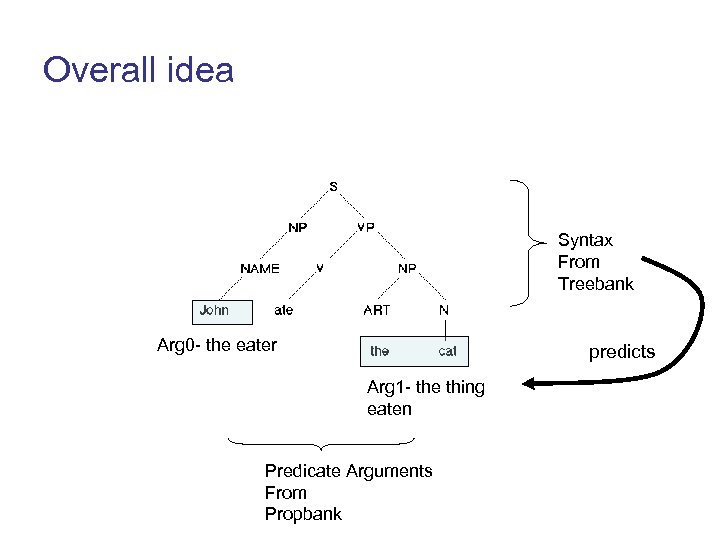
Overall idea Syntax From Treebank Arg 0 - the eater predicts Arg 1 - the thing eaten Predicate Arguments From Propbank

Problem: • WSJ corpus = business domain • In order to use WSJ, we have to make sure that the predicate distribution is “representative” for medical sentences. • We found that 99 out of top 100 predicates in medical abstracts can be found in the WSJ corpus.
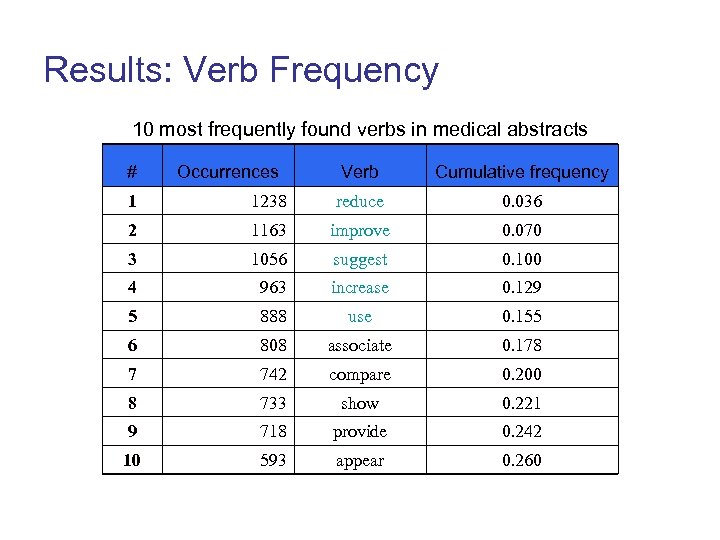
Results: Verb Frequency 10 most frequently found verbs in medical abstracts # Occurrences Verb Cumulative frequency 1 1238 reduce 0. 036 2 1163 improve 0. 070 3 1056 suggest 0. 100 4 963 increase 0. 129 5 888 use 0. 155 6 808 associate 0. 178 7 742 compare 0. 200 8 733 show 0. 221 9 718 provide 0. 242 10 593 appear 0. 260
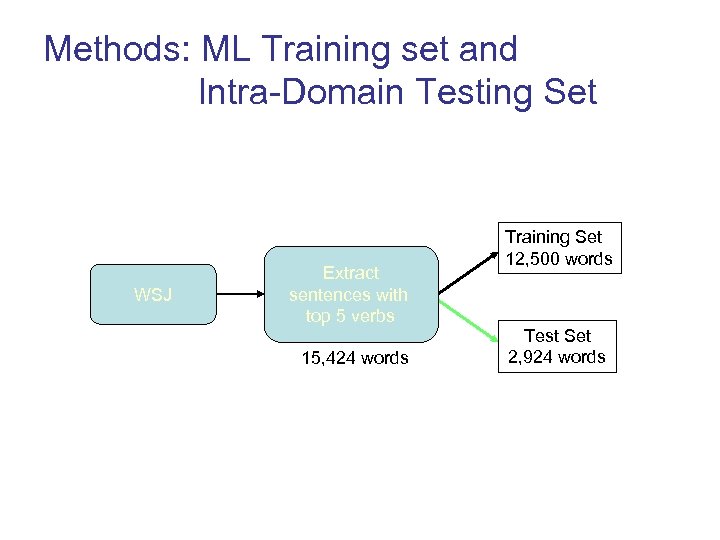
Methods: ML Training set and Intra-Domain Testing Set WSJ Extract sentences with top 5 verbs 15, 424 words Training Set 12, 500 words Test Set 2, 924 words
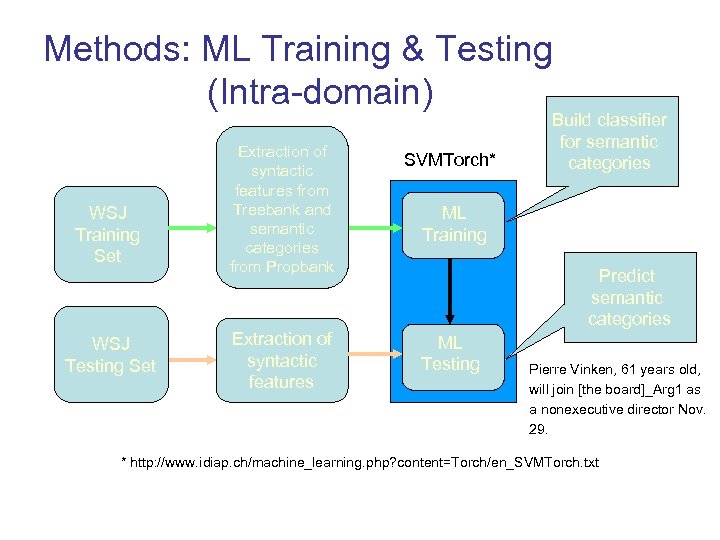
Methods: ML Training & Testing (Intra-domain) WSJ Training Set WSJ Testing Set Extraction of syntactic features from Treebank and semantic categories from Propbank SVMTorch* Extraction of syntactic features ML Testing Build classifier for semantic categories ML Training Predict semantic categories Pierre Vinken, 61 years old, will join [the board]_Arg 1 as a nonexecutive director Nov. 29. * http: //www. idiap. ch/machine_learning. php? content=Torch/en_SVMTorch. txt
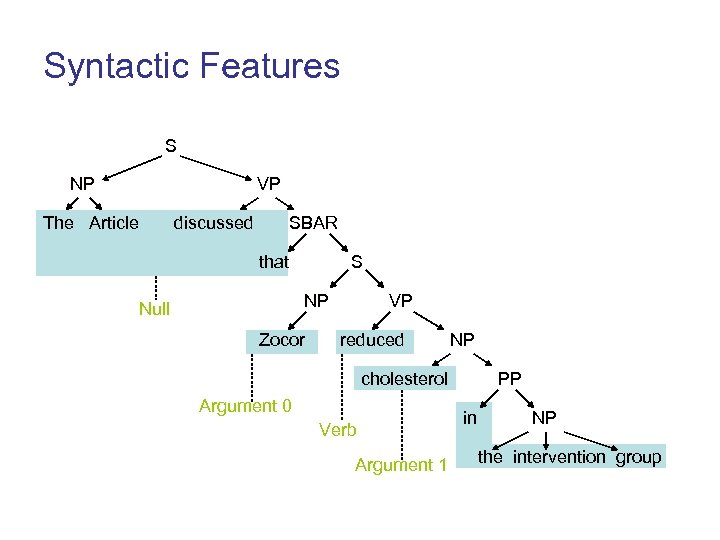
Syntactic Features S NP VP The Article discussed SBAR that S NP Null Zocor VP reduced NP cholesterol Argument 0 Verb Argument 1 PP in NP the intervention group

Syntactic Features • Predicate of the sentences • Syntactic path from a word to the sentence predicate – For the word Zocor, the paths are NP S VP VBD and S VP VBD • Phrase Type – The syntactic category of the constituent – NP and S for Zocor * S. Pradhan, D. Jurafsky, et al. In Proc. Of NAACL-HLT 2004.

Syntactic Features • Position of the word relative to the predicate • Head Word POS • The POS tag of the syntactic head of the constituent • Sub-categorization • Phrase structure expanding the predicate’s parent node in the parse tree. • VP VBD-NP for the predicate reduced
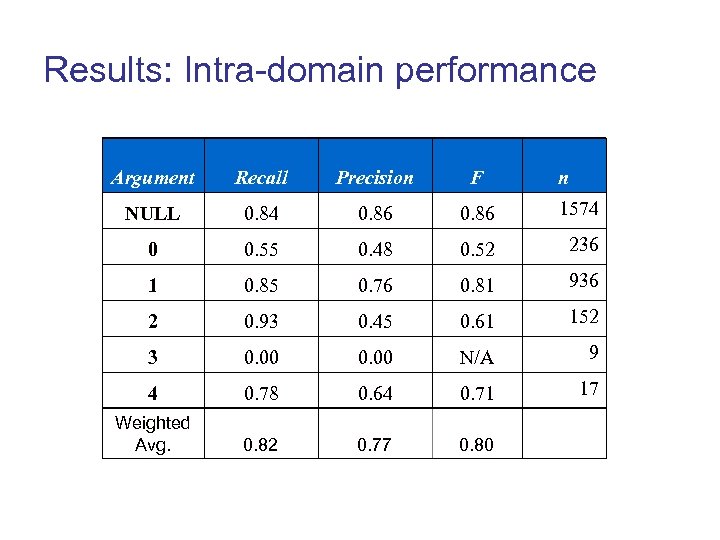
Results: Intra-domain performance Argument Recall Precision F n NULL 0. 84 0. 86 1574 0 0. 55 0. 48 0. 52 236 1 0. 85 0. 76 0. 81 936 2 0. 93 0. 45 0. 61 152 3 0. 00 N/A 9 4 0. 78 0. 64 0. 71 17 Weighted Avg. 0. 82 0. 77 0. 80
Results: Comparison with Prior Work * (Intra-domain) *Table 1: Performance on WSJ test set Arg Precision Recall F ID (null) 0. 86 0. 84 0. 86 ID + Class 0. 77 0. 82 0. 80 * S. Pradhan, D. Jurafsky, et al. In Proc. Of NAACL-HLT 2004.
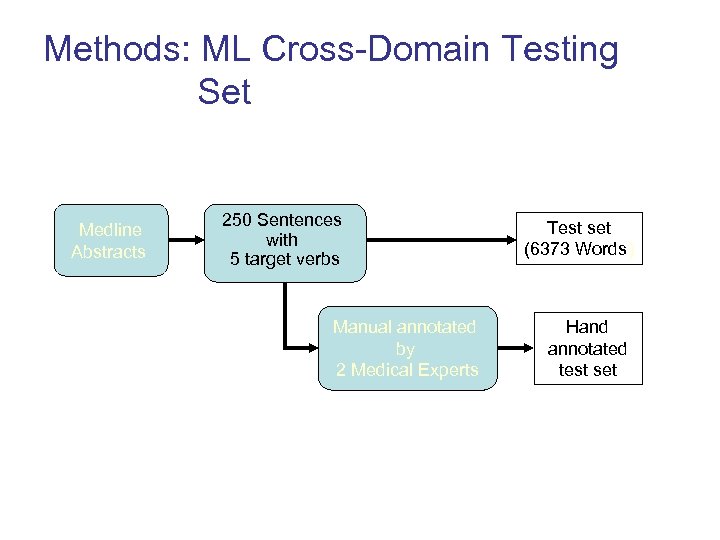
Methods: ML Cross-Domain Testing Set Medline Abstracts 250 Sentences with 5 target verbs Manual annotated by 2 Medical Experts Test set (6373 Words) Hand annotated test set
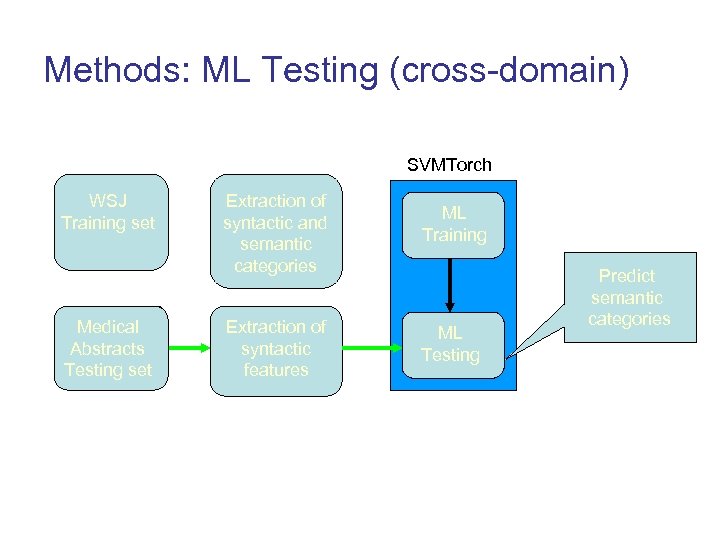
Methods: ML Testing (cross-domain) SVMTorch WSJ Propbank Training set (WSJ) Medical RCT Abstracts Testing set Extraction of syntactic and semantic categories Extraction of syntactic features ML Training ML Testing Predict semantic categories
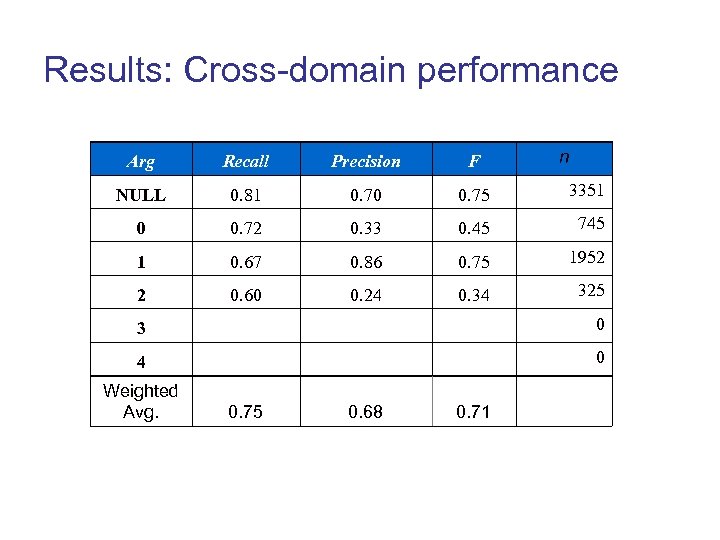
Results: Cross-domain performance n Arg Recall Precision F NULL 0. 81 0. 70 0. 75 3351 0 0. 72 0. 33 0. 45 745 1 0. 67 0. 86 0. 75 1952 2 0. 60 0. 24 0. 34 325 3 0 4 0 Weighted Avg. 0. 75 0. 68 0. 71
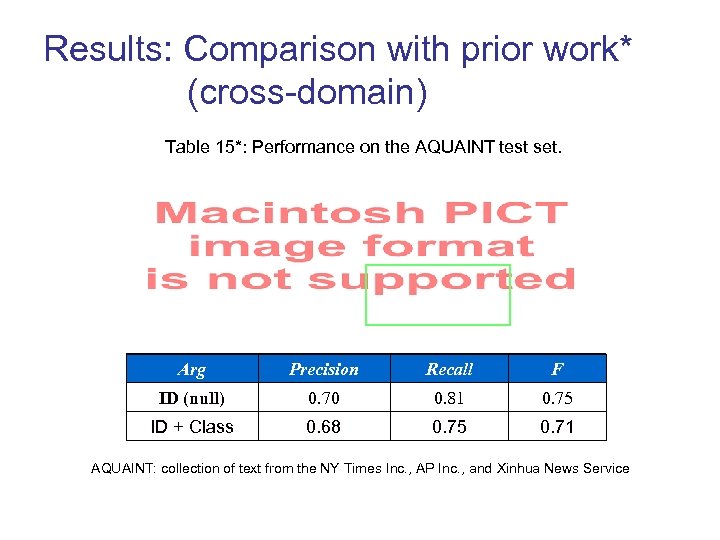

Results: Comparison with prior work* (cross-domain) Table 15*: Performance on the AQUAINT test set. Arg Precision Recall F ID (null) 0. 70 0. 81 0. 75 ID + Class 0. 68 0. 75 0. 71 AQUAINT: collection of text from the NY Times Inc. , AP Inc. , and Xinhua News Service

Discussion • Our ML classifier for null arguments – Intra-domain F = 86%, and cross-domain F = 75%, difference = 11% • Pradhan and Jurafsky article for null arguments – Intra-domain F = 92%, and cross-domain F = 81%, difference = 11% • Reuse of Propbank and Treebank information to automatically annotate medical abstract by using SSPT and ML classifier is feasible

Discussion - Limitations • Limitation – The results are based on a small medical testing set • Future directions – Improve the performance by addition of: • • Verb sense feature found in Propbank was not used Lack of lexical features Verb Clustering Temporal cue words – Test the performance using much larger medical abstract test set

Summary • Literature is an important resource for biomedical knowledge • Text mining = framework for accessing the free text in the literature, and transforming it to structured data • Machine Learning = essential element in the text mining process

Appendix: Sentence Predicate Extraction • Perl module Lingua: : EN: : Sentence -> Identified sentences • Charniak parser 1 -> Identified Parts of Speech – Based on WSJ corpus • Extracted terminals with VB* POS tags • Program morpha 2 -> Normalization of verbs 1. Charniak, E. , A Maximum-Entropy-Inspired Parser. 1999, Brown University. 2. Minning, G. , J. Carroll, and P. D. , Applied morphological processing of English. Natural Language Engineering, 2001. 7(3): p. 207 -223.
9eb6325e065c2c59f7c1f86447f6d72b.ppt