90b9b2320addbf0f5603c17c99cd45bd.ppt
- Количество слайдов: 74
 Systemic Lupus Erythematosus and DNA Methylation Terrence Shin MCB 5255 Dr. Lynes Mar. 28, 2012 1
Systemic Lupus Erythematosus and DNA Methylation Terrence Shin MCB 5255 Dr. Lynes Mar. 28, 2012 1
 Systemic Lupus Erythematosus (SLE) n Chronic autoimmune disorder q Affects skin, joints, kidneys, and others n n n Joint pain and swelling Lupus nephritis Cause – Unknown Ages: 10~50 More common in women African Americans and Asians are affected more often 2
Systemic Lupus Erythematosus (SLE) n Chronic autoimmune disorder q Affects skin, joints, kidneys, and others n n n Joint pain and swelling Lupus nephritis Cause – Unknown Ages: 10~50 More common in women African Americans and Asians are affected more often 2
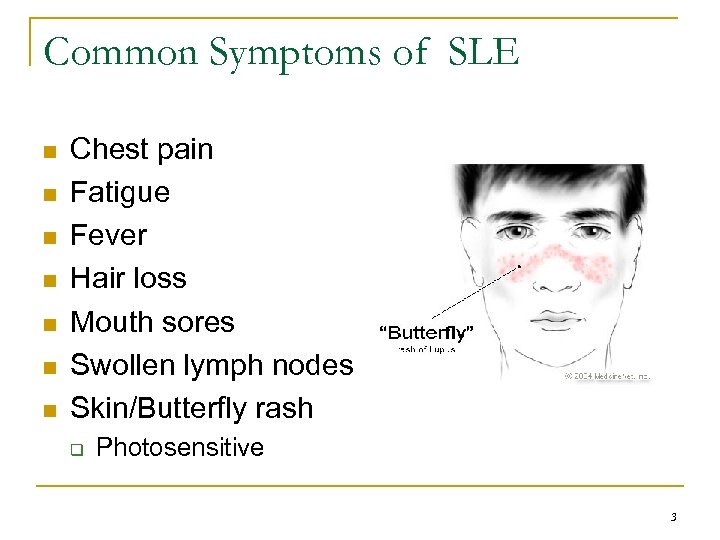 Common Symptoms of SLE n n n n Chest pain Fatigue Fever Hair loss Mouth sores Swollen lymph nodes Skin/Butterfly rash q Photosensitive 3
Common Symptoms of SLE n n n n Chest pain Fatigue Fever Hair loss Mouth sores Swollen lymph nodes Skin/Butterfly rash q Photosensitive 3
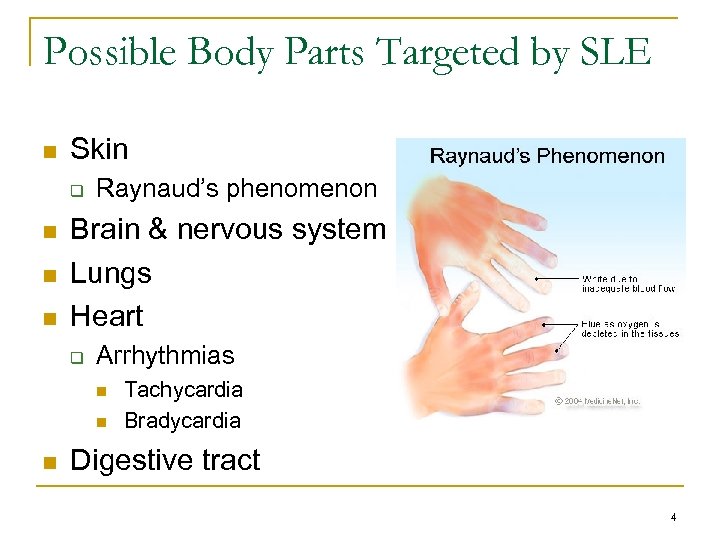 Possible Body Parts Targeted by SLE n Skin q n n n Raynaud’s phenomenon Brain & nervous system Lungs Heart q Arrhythmias n n n Tachycardia Bradycardia Digestive tract 4
Possible Body Parts Targeted by SLE n Skin q n n n Raynaud’s phenomenon Brain & nervous system Lungs Heart q Arrhythmias n n n Tachycardia Bradycardia Digestive tract 4
 Diagnosis of SLE n n n CBC ANA Chest x-ray Urinalysis Renal biopsy 5
Diagnosis of SLE n n n CBC ANA Chest x-ray Urinalysis Renal biopsy 5
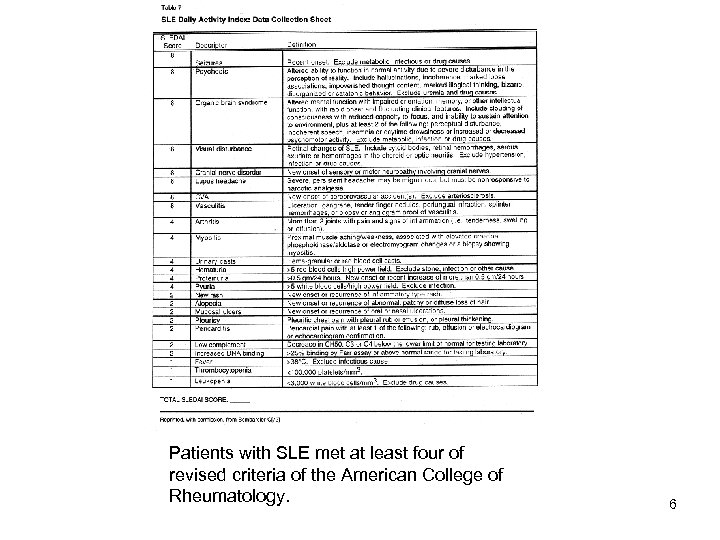 Patients with SLE met at least four of revised criteria of the American College of Rheumatology. 6
Patients with SLE met at least four of revised criteria of the American College of Rheumatology. 6
 Treatment for SLE n n No cure Control symptoms q q q Anti-inflammatory medication Corticosteroids Protection from sunlight 7
Treatment for SLE n n No cure Control symptoms q q q Anti-inflammatory medication Corticosteroids Protection from sunlight 7
 Ultraviolet B exposure of peripheral blood mononuclear cells of patients with systemic lupus erythematosus inhibits DNA methylation GS Wang, M Zhang, XP Li, H Zhang, W Chen, M Kan and YM Wang Lupus 2009 8
Ultraviolet B exposure of peripheral blood mononuclear cells of patients with systemic lupus erythematosus inhibits DNA methylation GS Wang, M Zhang, XP Li, H Zhang, W Chen, M Kan and YM Wang Lupus 2009 8
 Background n Sunlight can induce exacerbation of SLE q n SLE cells are hypomethylated q n Ultraviolet B (UVB): 290~320 nm Promoters of transcriptionally active gene DNA methylation q q q Suppressive effects on gene expression 5 th position of the cytosine ring Catalyzed by DNMT 1 9
Background n Sunlight can induce exacerbation of SLE q n SLE cells are hypomethylated q n Ultraviolet B (UVB): 290~320 nm Promoters of transcriptionally active gene DNA methylation q q q Suppressive effects on gene expression 5 th position of the cytosine ring Catalyzed by DNMT 1 9
 Specific Aims n What are the effect of UVB on DNA methylation in SLE? q What is its significance in the pathogenesis of SLE ? 10
Specific Aims n What are the effect of UVB on DNA methylation in SLE? q What is its significance in the pathogenesis of SLE ? 10
 Materials & Methods n n n Study participants Isolation of PBMCs Irradiation protocol DNA extraction and HPCE RNA isolation and RT-PCR Statistical analysis 11
Materials & Methods n n n Study participants Isolation of PBMCs Irradiation protocol DNA extraction and HPCE RNA isolation and RT-PCR Statistical analysis 11
 Study Participants n 45 patients with SLE q q q n 40 females, 5 males Mean age: 36 years; range: 18~61 years Active = SLEDAI ≥ 10 (21 out of 45 patients) 20 healthy volunteers (control) q q q Sex- and age-matched 18 females, 2 males Mean age: 31 years, range: 22~46 years 12
Study Participants n 45 patients with SLE q q q n 40 females, 5 males Mean age: 36 years; range: 18~61 years Active = SLEDAI ≥ 10 (21 out of 45 patients) 20 healthy volunteers (control) q q q Sex- and age-matched 18 females, 2 males Mean age: 31 years, range: 22~46 years 12
 Isolation of PBMCs n Total volume of 20 ml of ethylenediaminetetraacetic acid (EDTA)-K 2 preserved venous peripheral blood drawn from patients & controls q Ficoll gradient centrifugation n n PBMCs cultured in RPMI 1640 medium q Supplemented with 10% heat-inactivated FBS, 2 m. M sodium pyruvate, 100 IU/ml penicillin and 100 μg/ml streptomycin 13
Isolation of PBMCs n Total volume of 20 ml of ethylenediaminetetraacetic acid (EDTA)-K 2 preserved venous peripheral blood drawn from patients & controls q Ficoll gradient centrifugation n n PBMCs cultured in RPMI 1640 medium q Supplemented with 10% heat-inactivated FBS, 2 m. M sodium pyruvate, 100 IU/ml penicillin and 100 μg/ml streptomycin 13
 Irradiation Protocol n UVB irradiation q Waldman UV 109 B lights with TL-12 lamps (Waldman Lighting Ltd. , Germany) n n Emit within the UVB range (290– 320 nm) with an emission peak at 311 nm PBMC were irradiated in PBS with different doses of UVB (0, 50 and 100 m. J/cm 2) q PBS removed RPMI 1640 containing 10% FBS added cultured (24 h) DNA & RNA extractions 14
Irradiation Protocol n UVB irradiation q Waldman UV 109 B lights with TL-12 lamps (Waldman Lighting Ltd. , Germany) n n Emit within the UVB range (290– 320 nm) with an emission peak at 311 nm PBMC were irradiated in PBS with different doses of UVB (0, 50 and 100 m. J/cm 2) q PBS removed RPMI 1640 containing 10% FBS added cultured (24 h) DNA & RNA extractions 14
 DNA Extraction and HPCE n Five deoxynucleoside standards purchased from International Laboratory (CA, USA) q q q n d. A = 2’-deoxyadenosine d. T = 2’-deoxythymidine d. G = 2’-deoxyguanosine d. C = 2’-deoxycytidine md. C = 5 -methyl-2’-deoxycytidine All nucleosides were dissolved in dd. H 2 O water 15
DNA Extraction and HPCE n Five deoxynucleoside standards purchased from International Laboratory (CA, USA) q q q n d. A = 2’-deoxyadenosine d. T = 2’-deoxythymidine d. G = 2’-deoxyguanosine d. C = 2’-deoxycytidine md. C = 5 -methyl-2’-deoxycytidine All nucleosides were dissolved in dd. H 2 O water 15
 DNA Extraction and HPCE (cont. ) n n n DNA extraction q g. DNA extraction kit (QIAmp, DNA mini kit; Qiagen, Hilden, Germany) RNA digestion q Added 20 μg/μl RNase A (Sigma-Aldrich, St. Louis, MO) DNA hydrolysis q Heated 18 μl of DNA samples for 2 min in boiling water bath cooled rapidly in ice q Added 4. 5 μl of 10 m. M Zn. SO 4 and 7. 5 μl of nuclease P 1 (Sigma. Aldrich, St. Louis, MO) q Incubated at 37°C for 16 h q Added 7. 5 μl of Tris (0. 5 M, p. H 8. 3), and 4. 5 μl of alkaline phosphatase (Sigma-Aldrich, St. Louis, MO) and 50 U/ml in 2. 5 M (NH 4)2 SO 4 q Incubated at 37°C for 2 h 16
DNA Extraction and HPCE (cont. ) n n n DNA extraction q g. DNA extraction kit (QIAmp, DNA mini kit; Qiagen, Hilden, Germany) RNA digestion q Added 20 μg/μl RNase A (Sigma-Aldrich, St. Louis, MO) DNA hydrolysis q Heated 18 μl of DNA samples for 2 min in boiling water bath cooled rapidly in ice q Added 4. 5 μl of 10 m. M Zn. SO 4 and 7. 5 μl of nuclease P 1 (Sigma. Aldrich, St. Louis, MO) q Incubated at 37°C for 16 h q Added 7. 5 μl of Tris (0. 5 M, p. H 8. 3), and 4. 5 μl of alkaline phosphatase (Sigma-Aldrich, St. Louis, MO) and 50 U/ml in 2. 5 M (NH 4)2 SO 4 q Incubated at 37°C for 2 h 16
 DNA Extraction and HPCE (cont. ) n CE system (Beckman P/ACETM MDQ) q q q Uncoated fused-silica capillary (60 cm × 75 μm, effective length 57 cm) Connected to Millennium data-processing station Buffer: 48 m. M Na. HCO 3 (p. H 9. 6) containing 60 m. M SDS Constant voltage (20 k. V) and temperature (25°C) Pressure injected for 5 s Hydrolysed DNA or 0. 1 m. M free deoxynucleoside standard was injected into a HPCE analysis system equipped with photodiode array (PDA) detection n q Absorbance was monitored at 256 nm Capillary conditioning n Washed with 1 M Na. OH (2 min) washed with 1 m. M Na. OH (1 min) filled with the running buffer (3 min) q q Hydrolyzed samples n q Buffers and washing solutions were filtered through 0. 45 μm pores Injected hydrostatically (30 s) from 9. 8 cm above the cathode Comparison of migration time & peak area of each sample with the standards n Can calculate DNA methylation levels q Quantification of the relative methylation of each DNA sample § md. C/(d. C + md. C) x 100% 17
DNA Extraction and HPCE (cont. ) n CE system (Beckman P/ACETM MDQ) q q q Uncoated fused-silica capillary (60 cm × 75 μm, effective length 57 cm) Connected to Millennium data-processing station Buffer: 48 m. M Na. HCO 3 (p. H 9. 6) containing 60 m. M SDS Constant voltage (20 k. V) and temperature (25°C) Pressure injected for 5 s Hydrolysed DNA or 0. 1 m. M free deoxynucleoside standard was injected into a HPCE analysis system equipped with photodiode array (PDA) detection n q Absorbance was monitored at 256 nm Capillary conditioning n Washed with 1 M Na. OH (2 min) washed with 1 m. M Na. OH (1 min) filled with the running buffer (3 min) q q Hydrolyzed samples n q Buffers and washing solutions were filtered through 0. 45 μm pores Injected hydrostatically (30 s) from 9. 8 cm above the cathode Comparison of migration time & peak area of each sample with the standards n Can calculate DNA methylation levels q Quantification of the relative methylation of each DNA sample § md. C/(d. C + md. C) x 100% 17
 RNA Isolation and RT-PCR n Total RNA q Isolated from PBMCs n n Reverse transcription q n Trizol reagent (Invitrogen, Carlsbad, CA, USA) Reverse Transcription System (Promega, Madison, WI, USA) c. DNA synthesis q 1. 0 μg total RNA—single round reverse transcriptase reaction (TV = 20 μl) containing 0. 5 μg oligo (d. T) 15 primer, 2. 0 μl d. NTPs, 2. 0 μl reverse transcriptase 10 x buffer, 15 μl AMV reverse transcriptase and 0. 5 μl recombinant RNasin ribonuclease inhibitor. 18
RNA Isolation and RT-PCR n Total RNA q Isolated from PBMCs n n Reverse transcription q n Trizol reagent (Invitrogen, Carlsbad, CA, USA) Reverse Transcription System (Promega, Madison, WI, USA) c. DNA synthesis q 1. 0 μg total RNA—single round reverse transcriptase reaction (TV = 20 μl) containing 0. 5 μg oligo (d. T) 15 primer, 2. 0 μl d. NTPs, 2. 0 μl reverse transcriptase 10 x buffer, 15 μl AMV reverse transcriptase and 0. 5 μl recombinant RNasin ribonuclease inhibitor. 18
 RNA Isolation and RT-PCR (cont. ) n RT-PCR q ABI Prism 7500 Sequence Detection System n q q q Using SYBR Premix Ex Taq Kit (Ta. Ka. Ra Bio Inc. ) 10 μl of 2 x SYBR green Master Mix, 0. 2 μM primers, 5 μl 1: 5 dilution of prepared c. DNA, 20 μl of water 40 cycles at 94°C (30 s), 60°C (40 s), and 72°C (60 s) Primers used: n β-actin (forward & reverse), DNMT 1 (forward & reverse) q Purchased from Ta. Ka. Ra Bio Inc 19
RNA Isolation and RT-PCR (cont. ) n RT-PCR q ABI Prism 7500 Sequence Detection System n q q q Using SYBR Premix Ex Taq Kit (Ta. Ka. Ra Bio Inc. ) 10 μl of 2 x SYBR green Master Mix, 0. 2 μM primers, 5 μl 1: 5 dilution of prepared c. DNA, 20 μl of water 40 cycles at 94°C (30 s), 60°C (40 s), and 72°C (60 s) Primers used: n β-actin (forward & reverse), DNMT 1 (forward & reverse) q Purchased from Ta. Ka. Ra Bio Inc 19
 Statistical Analysis n Mann-Whitney U-test or one-way ANOVA q n Equality of means—to compare values Spearman’s rank correlation q Examine the relationship between two continuous variables 20
Statistical Analysis n Mann-Whitney U-test or one-way ANOVA q n Equality of means—to compare values Spearman’s rank correlation q Examine the relationship between two continuous variables 20
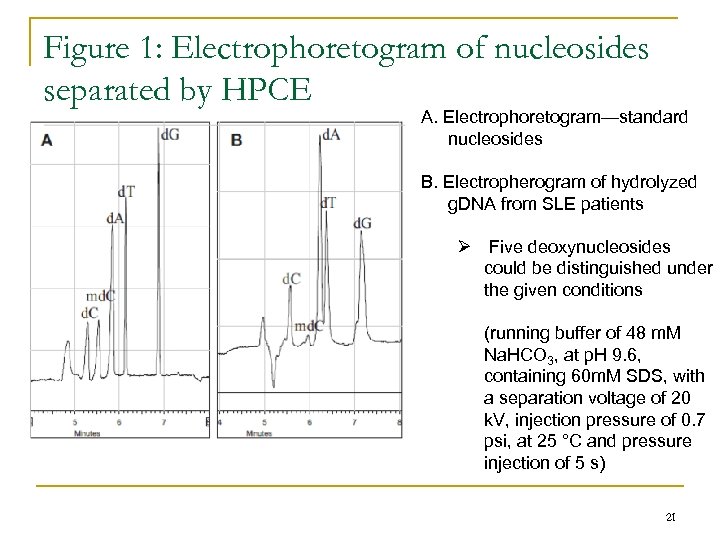 Figure 1: Electrophoretogram of nucleosides separated by HPCE A. Electrophoretogram—standard nucleosides B. Electropherogram of hydrolyzed g. DNA from SLE patients Ø Five deoxynucleosides could be distinguished under the given conditions (running buffer of 48 m. M Na. HCO 3, at p. H 9. 6, containing 60 m. M SDS, with a separation voltage of 20 k. V, injection pressure of 0. 7 psi, at 25 °C and pressure injection of 5 s) 21
Figure 1: Electrophoretogram of nucleosides separated by HPCE A. Electrophoretogram—standard nucleosides B. Electropherogram of hydrolyzed g. DNA from SLE patients Ø Five deoxynucleosides could be distinguished under the given conditions (running buffer of 48 m. M Na. HCO 3, at p. H 9. 6, containing 60 m. M SDS, with a separation voltage of 20 k. V, injection pressure of 0. 7 psi, at 25 °C and pressure injection of 5 s) 21
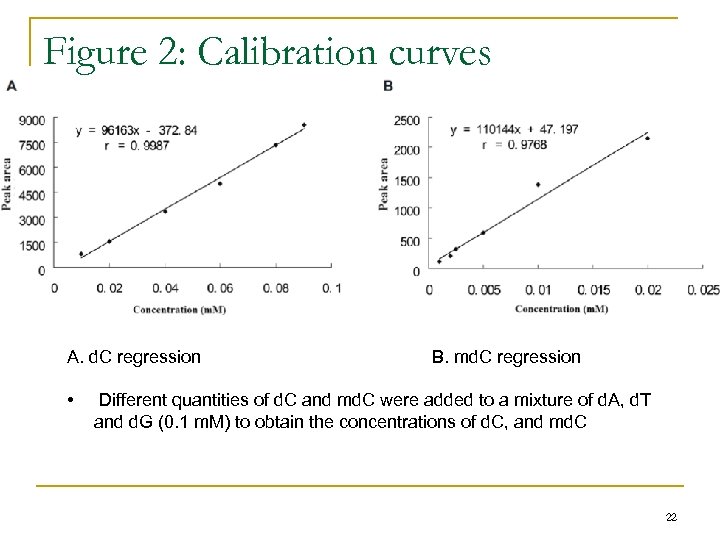 Figure 2: Calibration curves A. d. C regression • B. md. C regression Different quantities of d. C and md. C were added to a mixture of d. A, d. T and d. G (0. 1 m. M) to obtain the concentrations of d. C, and md. C 22
Figure 2: Calibration curves A. d. C regression • B. md. C regression Different quantities of d. C and md. C were added to a mixture of d. A, d. T and d. G (0. 1 m. M) to obtain the concentrations of d. C, and md. C 22
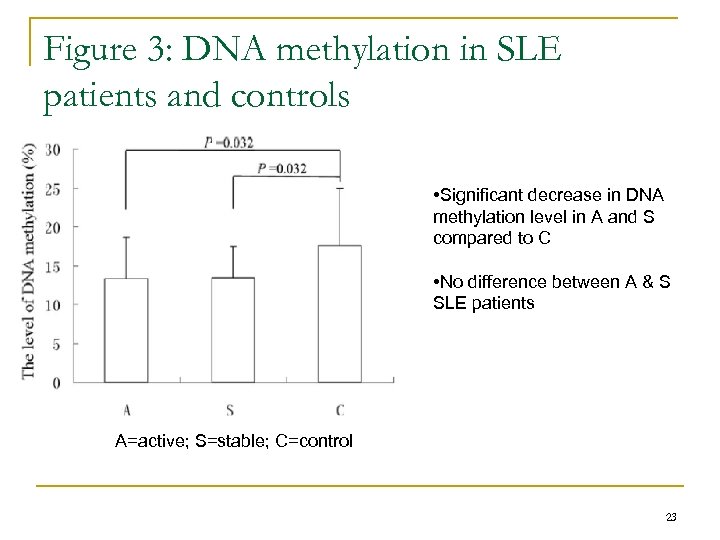 Figure 3: DNA methylation in SLE patients and controls • Significant decrease in DNA methylation level in A and S compared to C • No difference between A & S SLE patients A=active; S=stable; C=control 23
Figure 3: DNA methylation in SLE patients and controls • Significant decrease in DNA methylation level in A and S compared to C • No difference between A & S SLE patients A=active; S=stable; C=control 23
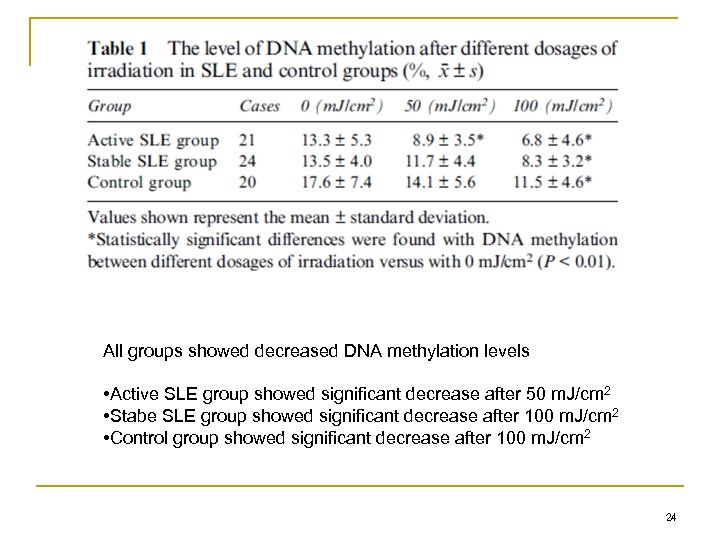 All groups showed decreased DNA methylation levels • Active SLE group showed significant decrease after 50 m. J/cm 2 • Stabe SLE group showed significant decrease after 100 m. J/cm 2 • Control group showed significant decrease after 100 m. J/cm 2 24
All groups showed decreased DNA methylation levels • Active SLE group showed significant decrease after 50 m. J/cm 2 • Stabe SLE group showed significant decrease after 100 m. J/cm 2 • Control group showed significant decrease after 100 m. J/cm 2 24
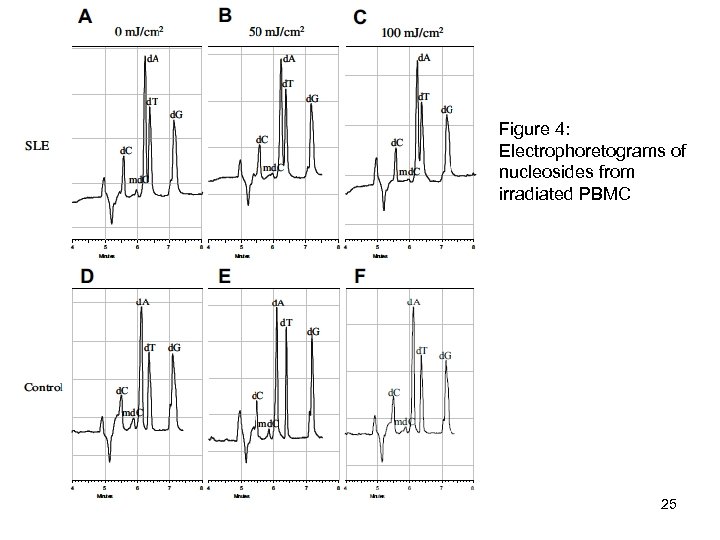 Figure 4: Electrophoretograms of nucleosides from irradiated PBMC 25
Figure 4: Electrophoretograms of nucleosides from irradiated PBMC 25
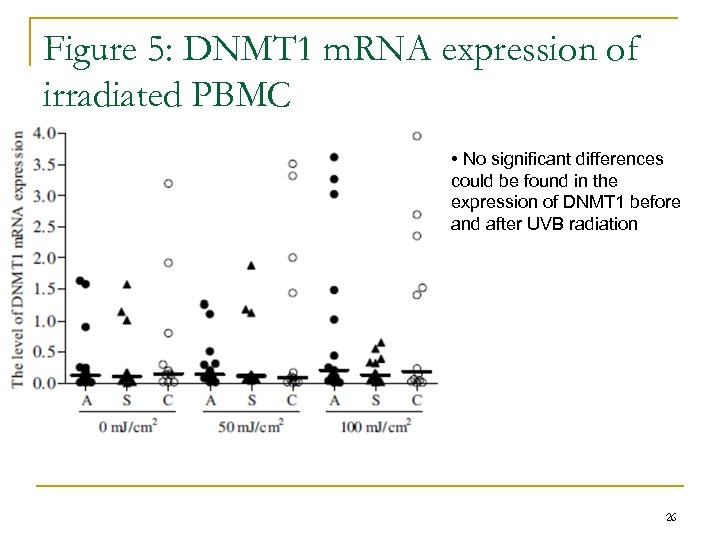 Figure 5: DNMT 1 m. RNA expression of irradiated PBMC • No significant differences could be found in the expression of DNMT 1 before and after UVB radiation 26
Figure 5: DNMT 1 m. RNA expression of irradiated PBMC • No significant differences could be found in the expression of DNMT 1 before and after UVB radiation 26
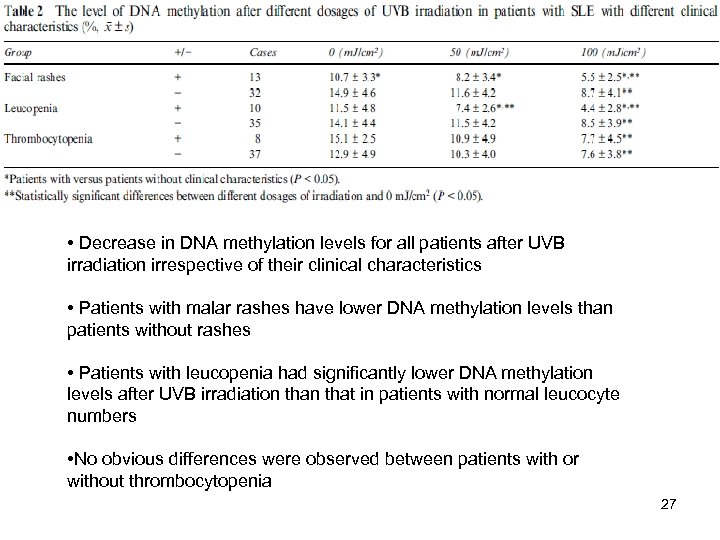 • Decrease in DNA methylation levels for all patients after UVB irradiation irrespective of their clinical characteristics • Patients with malar rashes have lower DNA methylation levels than patients without rashes • Patients with leucopenia had significantly lower DNA methylation levels after UVB irradiation that in patients with normal leucocyte numbers • No obvious differences were observed between patients with or without thrombocytopenia 27
• Decrease in DNA methylation levels for all patients after UVB irradiation irrespective of their clinical characteristics • Patients with malar rashes have lower DNA methylation levels than patients without rashes • Patients with leucopenia had significantly lower DNA methylation levels after UVB irradiation that in patients with normal leucocyte numbers • No obvious differences were observed between patients with or without thrombocytopenia 27
 Discussion & Unknowns n SLE patients have significantly lower levels of methylated DNA compared to controls (without UVB irradiation) q n Does down-regulation of DNA methylation play a role in the pathogenesis of SLE? DNA methylation levels decreased after UVB exposure q q Especially for patients with facial rash and leucopenia Does UVB play a role in pathogenesis of SLE by decreasing DNA methylation? 28
Discussion & Unknowns n SLE patients have significantly lower levels of methylated DNA compared to controls (without UVB irradiation) q n Does down-regulation of DNA methylation play a role in the pathogenesis of SLE? DNA methylation levels decreased after UVB exposure q q Especially for patients with facial rash and leucopenia Does UVB play a role in pathogenesis of SLE by decreasing DNA methylation? 28
 IL-6 Modulates CD 5 Expression in B Cells from Patients with Lupus by Regulating DNA Methylation Soizic Garaud, Christelle Le Dantec, Sandrine Jousse. Joulin, Catherine Hanrotel-Saliou, Alain Saraux, Rizgar A. Mageed, Pierre Youinou and Yves Renaudineau The Journal of Immunology, 2009 29
IL-6 Modulates CD 5 Expression in B Cells from Patients with Lupus by Regulating DNA Methylation Soizic Garaud, Christelle Le Dantec, Sandrine Jousse. Joulin, Catherine Hanrotel-Saliou, Alain Saraux, Rizgar A. Mageed, Pierre Youinou and Yves Renaudineau The Journal of Immunology, 2009 29
 Background n Features of autoimmunity in SLE q q q n B cell abnormalities in SLE q q q n B cell hyperactivity Spontaneous lymphocyte proliferation Production of antibodies to self-antigens Excess cytokine production Autoantigen presentation to T cells Modulation of the function of other immune cells SLE is considered as a B cell disease q Therapies target B cells 30
Background n Features of autoimmunity in SLE q q q n B cell abnormalities in SLE q q q n B cell hyperactivity Spontaneous lymphocyte proliferation Production of antibodies to self-antigens Excess cytokine production Autoantigen presentation to T cells Modulation of the function of other immune cells SLE is considered as a B cell disease q Therapies target B cells 30
 Background (cont. ) n CD 5 isoforms q q CD 5 -E 1 A CD 5 -E 1 B n Demethylated in B cells of SLE patients q CD 5 -E 1 B m. RNA is transcribed at the expense of CD 5 -E 1 A m. RNA transcription 31
Background (cont. ) n CD 5 isoforms q q CD 5 -E 1 A CD 5 -E 1 B n Demethylated in B cells of SLE patients q CD 5 -E 1 B m. RNA is transcribed at the expense of CD 5 -E 1 A m. RNA transcription 31
 Specific Aims n Does the engagement of the BCR with constitutive IL-6 down-regulate the level of membrane CD 5, which negatively regulates BCR signaling, in SLE B cells? q Does this signal promote the activation and expansion of autoreactive B cells in SLE patients? 32
Specific Aims n Does the engagement of the BCR with constitutive IL-6 down-regulate the level of membrane CD 5, which negatively regulates BCR signaling, in SLE B cells? q Does this signal promote the activation and expansion of autoreactive B cells in SLE patients? 32
 Materials & Methods n n n n n B lymphocyte isolation FACS analysis Cell culture m. RNA extraction and quantitative RT-PCR RACE Methylation-specific PCR Bisulfite sequencing Chromatin immunoprecipitation (Ch. IP) Computational promoter analysis Statistics 33
Materials & Methods n n n n n B lymphocyte isolation FACS analysis Cell culture m. RNA extraction and quantitative RT-PCR RACE Methylation-specific PCR Bisulfite sequencing Chromatin immunoprecipitation (Ch. IP) Computational promoter analysis Statistics 33
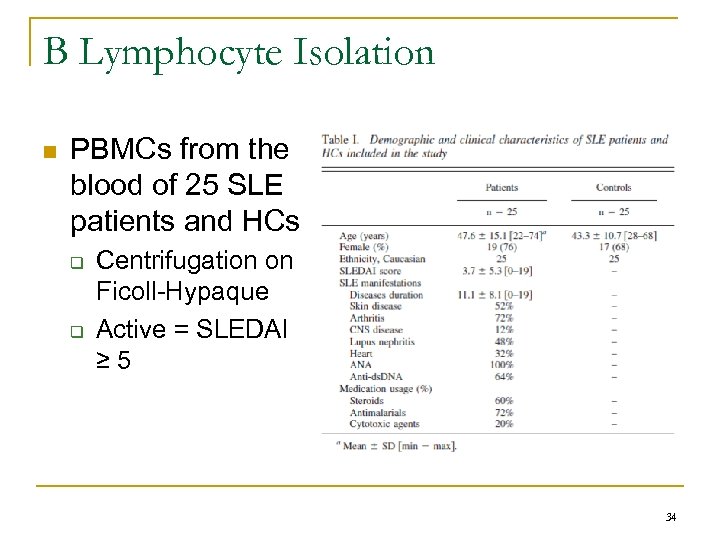 B Lymphocyte Isolation n PBMCs from the blood of 25 SLE patients and HCs q q Centrifugation on Ficoll-Hypaque Active = SLEDAI ≥ 5 34
B Lymphocyte Isolation n PBMCs from the blood of 25 SLE patients and HCs q q Centrifugation on Ficoll-Hypaque Active = SLEDAI ≥ 5 34
 B Lymphocyte Isolation (cont. ) n Cells permeabilized with 70% methanol intracellular staining q FITC-anti-CD 19 (clone J 4 -119) and PE-anti-CD 5 (clone BL 1 a) n q Anti-DNMT 1 and anti-p 27 kip 1 n n Obtained from Abcam CD 5 -CD 19+ B cells q q n Obtained from Beckman-Coulter Sorted on an Epics Elite FACS (Beckman-Coulter) 98% CD 19+ Daudi human B cell line q Purchased from American Type Culture Collection 35
B Lymphocyte Isolation (cont. ) n Cells permeabilized with 70% methanol intracellular staining q FITC-anti-CD 19 (clone J 4 -119) and PE-anti-CD 5 (clone BL 1 a) n q Anti-DNMT 1 and anti-p 27 kip 1 n n Obtained from Abcam CD 5 -CD 19+ B cells q q n Obtained from Beckman-Coulter Sorted on an Epics Elite FACS (Beckman-Coulter) 98% CD 19+ Daudi human B cell line q Purchased from American Type Culture Collection 35
 FACS Analysis n Number of CD 5 molecules per cell q Quantum Simply Cellular kit n n Amount of Ab binding to the cells (ABC) at saturating concentrations FACS analysis q 50μl calibrated microspheres stained with 20 μl of the same anti-CD 5 Ab n Mean fluorescence intensity standard ABC curve arbitrary ABC value 36
FACS Analysis n Number of CD 5 molecules per cell q Quantum Simply Cellular kit n n Amount of Ab binding to the cells (ABC) at saturating concentrations FACS analysis q 50μl calibrated microspheres stained with 20 μl of the same anti-CD 5 Ab n Mean fluorescence intensity standard ABC curve arbitrary ABC value 36
 Cell Culture n B cells (sorted via FACS) q Suspended in RPMI 1640 n q Supplemented with 10% heat-inactivated FCS, 2 m. M Lglutamine, 200 U/ml penicillin and 100 μg/ml streptomycin. B lymphocytes were seeded at 2 x 105 cells per well n Incubated with 1 μg/ml anti-Ig. M Ab-coated Sepharose beads (Bio. Rad) and 10 U/ml IL-2 q q In the presence or absence of 10– 40 ng/ml anti-IL-6 RAb (R&D Systems), or 100 ng/ml rh. IL-6 (Immuno Tools). Repression of DNMTs n Incubated with 50 μM of the ras signal blocker PD 98059 q q Detected IL-6 and IFN-ɣ in sera Detected IL-6 in the supernatant of cultured cells using ELISA kits (Beckman Coulter). 37
Cell Culture n B cells (sorted via FACS) q Suspended in RPMI 1640 n q Supplemented with 10% heat-inactivated FCS, 2 m. M Lglutamine, 200 U/ml penicillin and 100 μg/ml streptomycin. B lymphocytes were seeded at 2 x 105 cells per well n Incubated with 1 μg/ml anti-Ig. M Ab-coated Sepharose beads (Bio. Rad) and 10 U/ml IL-2 q q In the presence or absence of 10– 40 ng/ml anti-IL-6 RAb (R&D Systems), or 100 ng/ml rh. IL-6 (Immuno Tools). Repression of DNMTs n Incubated with 50 μM of the ras signal blocker PD 98059 q q Detected IL-6 and IFN-ɣ in sera Detected IL-6 in the supernatant of cultured cells using ELISA kits (Beckman Coulter). 37
 m. RNA Extraction and q. RT-PCR n n Total m. RNA q Extracted using the RNAble method (Eurobio) c. DNA synthesis q Reverse transcription n n Quantitative RT-PCR q Conducted in 20 μl mixtures n n n 20 μl volume with Superscript II RNase H-RT (Invitrogen Corporation). 50 ng template c. DNA 1 X Sybr Green PCR Master mix (Applied Biosystems) 500 n. M of each primer (Table II) All assays included a negative and positive controls CT method q Comparison of cycle thresholds q 18 S as an internal control 38
m. RNA Extraction and q. RT-PCR n n Total m. RNA q Extracted using the RNAble method (Eurobio) c. DNA synthesis q Reverse transcription n n Quantitative RT-PCR q Conducted in 20 μl mixtures n n n 20 μl volume with Superscript II RNase H-RT (Invitrogen Corporation). 50 ng template c. DNA 1 X Sybr Green PCR Master mix (Applied Biosystems) 500 n. M of each primer (Table II) All assays included a negative and positive controls CT method q Comparison of cycle thresholds q 18 S as an internal control 38
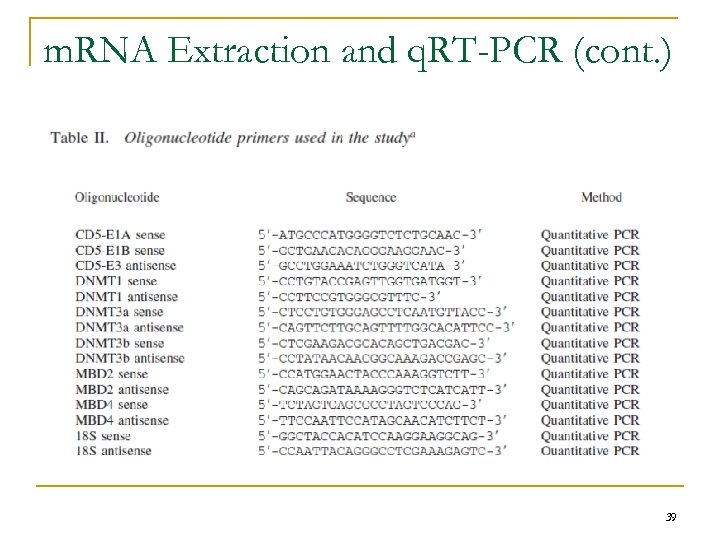 m. RNA Extraction and q. RT-PCR (cont. ) 39
m. RNA Extraction and q. RT-PCR (cont. ) 39
 RACE n m. RNA 5’ transcript ends q n c. DNA first strand synthesis q n Sense UPM primer and the gene-specific antisense primer CD 5 E 5 (Table II) PCR protocol q n Amplified with SMART-RACE kit (Clontech) Initial denaturation at 94°C for 5 min 5 touchdown-PCR cycles of denaturation at 94°C for 30 s Annealing at 72°C for 3 min Another 5 cycles at 94°C for 30 s, 70°C for 30 s, 72°C for 3 min decreasing temperature for 35 cycles of 94°C or 30 s, 68°C for 30 s, 72°C for 3 min. Nested PCR q q Sense NUP primer and the gene-specific antisense primer CD 5 E 3 Second PCR round: n 40 cycles at 94°C for 30 s, 56°C for 1 min, and 72°C for 1 min with a final extension at 72°C for 10 min. 40
RACE n m. RNA 5’ transcript ends q n c. DNA first strand synthesis q n Sense UPM primer and the gene-specific antisense primer CD 5 E 5 (Table II) PCR protocol q n Amplified with SMART-RACE kit (Clontech) Initial denaturation at 94°C for 5 min 5 touchdown-PCR cycles of denaturation at 94°C for 30 s Annealing at 72°C for 3 min Another 5 cycles at 94°C for 30 s, 70°C for 30 s, 72°C for 3 min decreasing temperature for 35 cycles of 94°C or 30 s, 68°C for 30 s, 72°C for 3 min. Nested PCR q q Sense NUP primer and the gene-specific antisense primer CD 5 E 3 Second PCR round: n 40 cycles at 94°C for 30 s, 56°C for 1 min, and 72°C for 1 min with a final extension at 72°C for 10 min. 40
 RACE (cont. ) UPM Primer NUP Primer 41
RACE (cont. ) UPM Primer NUP Primer 41
 Methylation-specific PCR n Genomic DNA q q Purification – QIAmp 96 DNA blood kit (Qiagen) Digestion – 20 U of the methylation-sensitive REs (Hpa. II, Hae. II, Fau. I, Hga. I) or the methylation insensitive restriction enzyme (Msp. I) – 37°C for 3 h n Undigested Genomic DNA q n PCR primers q Positioned upstream and downstream of E 1 A and E 1 B promoter recognition sites n n Positive control E 1 A and E 1 B of CD 5, CD 19 CD 70, Pax 5, Syk, and HRES-1 genes (Table II) PCR protocol q q Initial denaturation at 94°C for 5 min 35 cycles of denaturation at 94°C for 30 s, annealing at 56°C for 1 min, and primer extension at 72°C for 1 min final extension at 72°C for 10 min. Products separated on agarose gel n Visualized with 0. 5 g/ml ethidium bromide. 42
Methylation-specific PCR n Genomic DNA q q Purification – QIAmp 96 DNA blood kit (Qiagen) Digestion – 20 U of the methylation-sensitive REs (Hpa. II, Hae. II, Fau. I, Hga. I) or the methylation insensitive restriction enzyme (Msp. I) – 37°C for 3 h n Undigested Genomic DNA q n PCR primers q Positioned upstream and downstream of E 1 A and E 1 B promoter recognition sites n n Positive control E 1 A and E 1 B of CD 5, CD 19 CD 70, Pax 5, Syk, and HRES-1 genes (Table II) PCR protocol q q Initial denaturation at 94°C for 5 min 35 cycles of denaturation at 94°C for 30 s, annealing at 56°C for 1 min, and primer extension at 72°C for 1 min final extension at 72°C for 10 min. Products separated on agarose gel n Visualized with 0. 5 g/ml ethidium bromide. 42
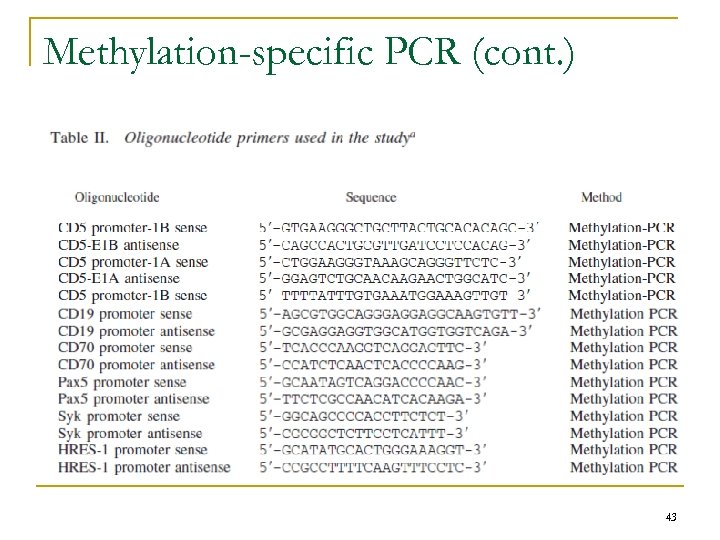 Methylation-specific PCR (cont. ) 43
Methylation-specific PCR (cont. ) 43
 Bisulfite Sequencing n Methylation status of DNA q Non-methylated cytosines were converted to uridines by bisulfite treatment n n Unmodified DNA (100 ng) q q n Amplified 40 times at 56°C using specific primers Sequenced using specific primers Bisulfite-converted DNA q n EZ-DNA methylation-Gold kit (Zymo Research) Amplified by nested PCR using two rounds of 40 cycles each at 56°C with primers specific for methylated cytosines (Table II) PCR products q q Purification – high pure PCR product purification kit (Roche) Sequencing – Internal primers n q Big. Dye Terminator Cycle Sequencing kit using an automated ABI-310 genetic analyzer (Applied Biosystems) Electrophoregram n n Quantification of T and C peaks Determination of methylation status = [peak (C)/peak (T) + peak (C)] x 100 44
Bisulfite Sequencing n Methylation status of DNA q Non-methylated cytosines were converted to uridines by bisulfite treatment n n Unmodified DNA (100 ng) q q n Amplified 40 times at 56°C using specific primers Sequenced using specific primers Bisulfite-converted DNA q n EZ-DNA methylation-Gold kit (Zymo Research) Amplified by nested PCR using two rounds of 40 cycles each at 56°C with primers specific for methylated cytosines (Table II) PCR products q q Purification – high pure PCR product purification kit (Roche) Sequencing – Internal primers n q Big. Dye Terminator Cycle Sequencing kit using an automated ABI-310 genetic analyzer (Applied Biosystems) Electrophoregram n n Quantification of T and C peaks Determination of methylation status = [peak (C)/peak (T) + peak (C)] x 100 44
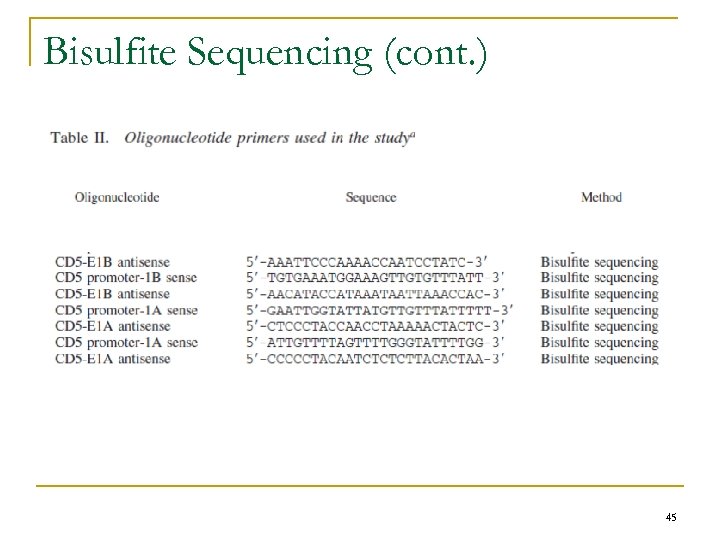 Bisulfite Sequencing (cont. ) 45
Bisulfite Sequencing (cont. ) 45
 Ch. IP n n n Epi. Quik kit (Epigentek Group) Evaluation of the activation of the CD 5 -E 1 B promoter Sonicated DNA (200– 1000 bp) q Transferred into strip wells n q q Incubated for 90 min at room temperature, then washed Precipitated DNA-protein complexes n n q Treated with 250 μg/ml proteinase K in the DNA release buffer for 15 min Left in the same buffer for 90 min at 65°C Collection of DNA samples n n Precoated with mouse anti-RNA polymerase II, or with a nonspecific mouse Ig. G, used as a negative control P-spin columns, washed with ethanol, and eluted PCR q q Template – purified DNA GAPDH & CD 5 -E 1 B specific primers (Table II) 40 cycles at 56°C Products were separated on agarose gel n Visualized with 0. 5 g/ml ethidium bromide. 46
Ch. IP n n n Epi. Quik kit (Epigentek Group) Evaluation of the activation of the CD 5 -E 1 B promoter Sonicated DNA (200– 1000 bp) q Transferred into strip wells n q q Incubated for 90 min at room temperature, then washed Precipitated DNA-protein complexes n n q Treated with 250 μg/ml proteinase K in the DNA release buffer for 15 min Left in the same buffer for 90 min at 65°C Collection of DNA samples n n Precoated with mouse anti-RNA polymerase II, or with a nonspecific mouse Ig. G, used as a negative control P-spin columns, washed with ethanol, and eluted PCR q q Template – purified DNA GAPDH & CD 5 -E 1 B specific primers (Table II) 40 cycles at 56°C Products were separated on agarose gel n Visualized with 0. 5 g/ml ethidium bromide. 46
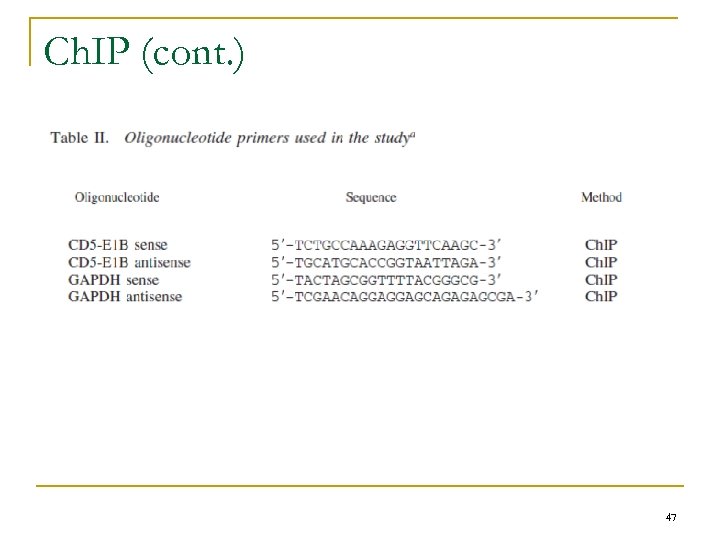 Ch. IP (cont. ) 47
Ch. IP (cont. ) 47
 Computational Promoter Analysis n Identification of putative transcription factor binding sites q q q Alibaba (v 2. 1) TESS (transcription element search system) Genomatix 48
Computational Promoter Analysis n Identification of putative transcription factor binding sites q q q Alibaba (v 2. 1) TESS (transcription element search system) Genomatix 48
 Statistical Analysis n n Results as arithmetic means with SD Mann-Whitney U test q n Wilcoxon test q n Unpaired data Paired data Spearman’s rank correlation 49
Statistical Analysis n n Results as arithmetic means with SD Mann-Whitney U test q n Wilcoxon test q n Unpaired data Paired data Spearman’s rank correlation 49
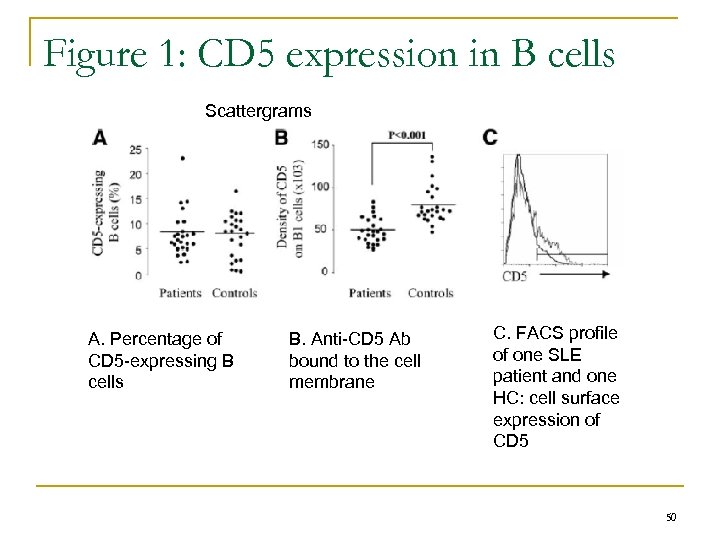 Figure 1: CD 5 expression in B cells Scattergrams A. Percentage of CD 5 -expressing B cells B. Anti-CD 5 Ab bound to the cell membrane C. FACS profile of one SLE patient and one HC: cell surface expression of CD 5 50
Figure 1: CD 5 expression in B cells Scattergrams A. Percentage of CD 5 -expressing B cells B. Anti-CD 5 Ab bound to the cell membrane C. FACS profile of one SLE patient and one HC: cell surface expression of CD 5 50
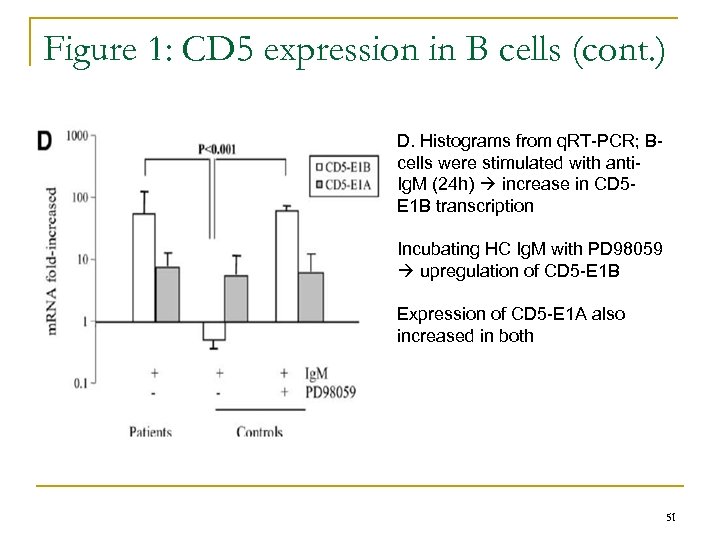 Figure 1: CD 5 expression in B cells (cont. ) D. Histograms from q. RT-PCR; Bcells were stimulated with anti. Ig. M (24 h) increase in CD 5 E 1 B transcription Incubating HC Ig. M with PD 98059 upregulation of CD 5 -E 1 B Expression of CD 5 -E 1 A also increased in both 51
Figure 1: CD 5 expression in B cells (cont. ) D. Histograms from q. RT-PCR; Bcells were stimulated with anti. Ig. M (24 h) increase in CD 5 E 1 B transcription Incubating HC Ig. M with PD 98059 upregulation of CD 5 -E 1 B Expression of CD 5 -E 1 A also increased in both 51
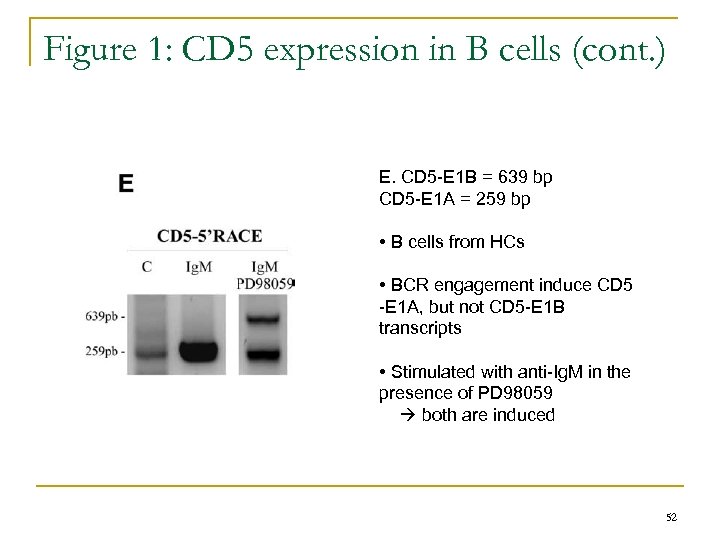 Figure 1: CD 5 expression in B cells (cont. ) E. CD 5 -E 1 B = 639 bp CD 5 -E 1 A = 259 bp • B cells from HCs • BCR engagement induce CD 5 -E 1 A, but not CD 5 -E 1 B transcripts • Stimulated with anti-Ig. M in the presence of PD 98059 both are induced 52
Figure 1: CD 5 expression in B cells (cont. ) E. CD 5 -E 1 B = 639 bp CD 5 -E 1 A = 259 bp • B cells from HCs • BCR engagement induce CD 5 -E 1 A, but not CD 5 -E 1 B transcripts • Stimulated with anti-Ig. M in the presence of PD 98059 both are induced 52
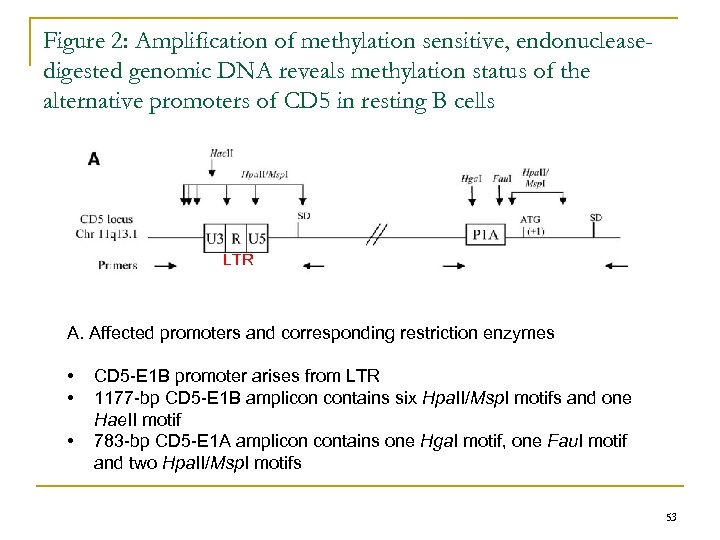 Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells LTR A. Affected promoters and corresponding restriction enzymes • • • CD 5 -E 1 B promoter arises from LTR 1177 -bp CD 5 -E 1 B amplicon contains six Hpa. II/Msp. I motifs and one Hae. II motif 783 -bp CD 5 -E 1 A amplicon contains one Hga. I motif, one Fau. I motif and two Hpa. II/Msp. I motifs 53
Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells LTR A. Affected promoters and corresponding restriction enzymes • • • CD 5 -E 1 B promoter arises from LTR 1177 -bp CD 5 -E 1 B amplicon contains six Hpa. II/Msp. I motifs and one Hae. II motif 783 -bp CD 5 -E 1 A amplicon contains one Hga. I motif, one Fau. I motif and two Hpa. II/Msp. I motifs 53
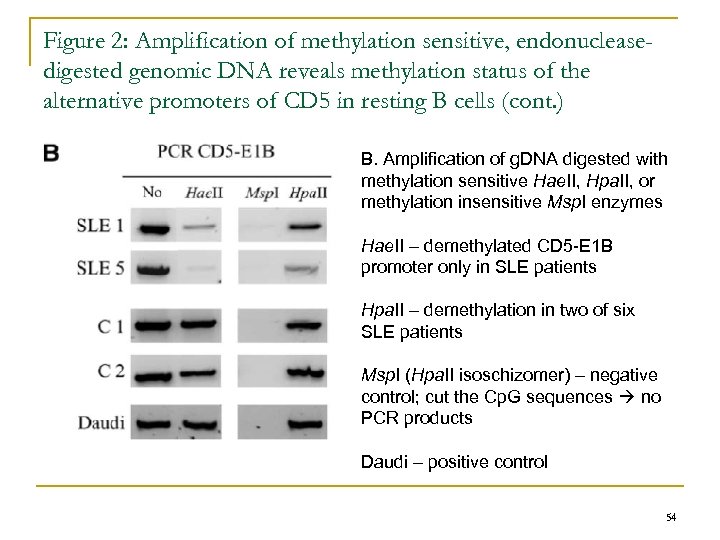 Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells (cont. ) B. Amplification of g. DNA digested with methylation sensitive Hae. II, Hpa. II, or methylation insensitive Msp. I enzymes Hae. II – demethylated CD 5 -E 1 B promoter only in SLE patients Hpa. II – demethylation in two of six SLE patients Msp. I (Hpa. II isoschizomer) – negative control; cut the Cp. G sequences no PCR products Daudi – positive control 54
Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells (cont. ) B. Amplification of g. DNA digested with methylation sensitive Hae. II, Hpa. II, or methylation insensitive Msp. I enzymes Hae. II – demethylated CD 5 -E 1 B promoter only in SLE patients Hpa. II – demethylation in two of six SLE patients Msp. I (Hpa. II isoschizomer) – negative control; cut the Cp. G sequences no PCR products Daudi – positive control 54
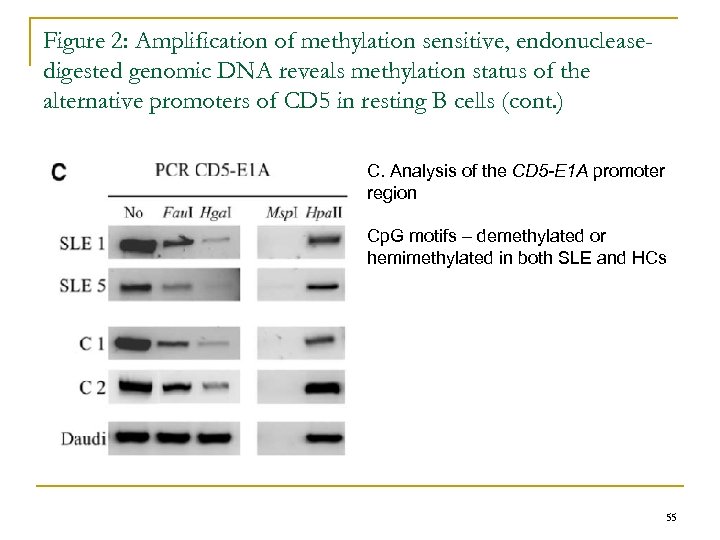 Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells (cont. ) C. Analysis of the CD 5 -E 1 A promoter region Cp. G motifs – demethylated or hemimethylated in both SLE and HCs 55
Figure 2: Amplification of methylation sensitive, endonucleasedigested genomic DNA reveals methylation status of the alternative promoters of CD 5 in resting B cells (cont. ) C. Analysis of the CD 5 -E 1 A promoter region Cp. G motifs – demethylated or hemimethylated in both SLE and HCs 55
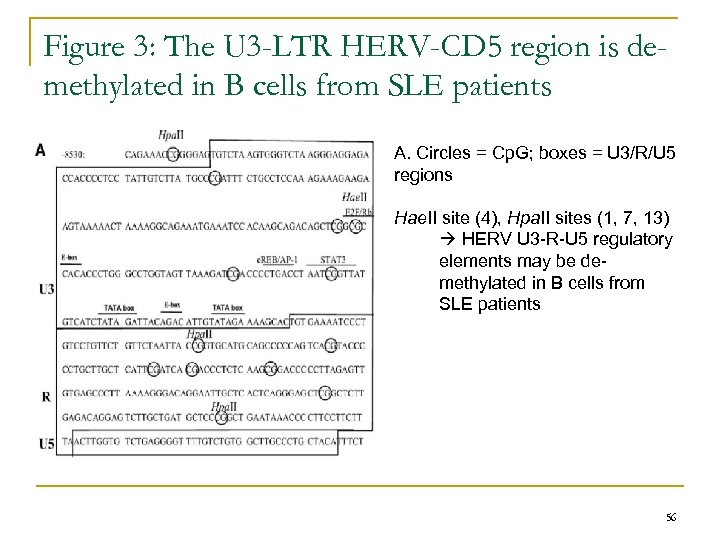 Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients A. Circles = Cp. G; boxes = U 3/R/U 5 regions Hae. II site (4), Hpa. II sites (1, 7, 13) HERV U 3 -R-U 5 regulatory elements may be demethylated in B cells from SLE patients 56
Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients A. Circles = Cp. G; boxes = U 3/R/U 5 regions Hae. II site (4), Hpa. II sites (1, 7, 13) HERV U 3 -R-U 5 regulatory elements may be demethylated in B cells from SLE patients 56
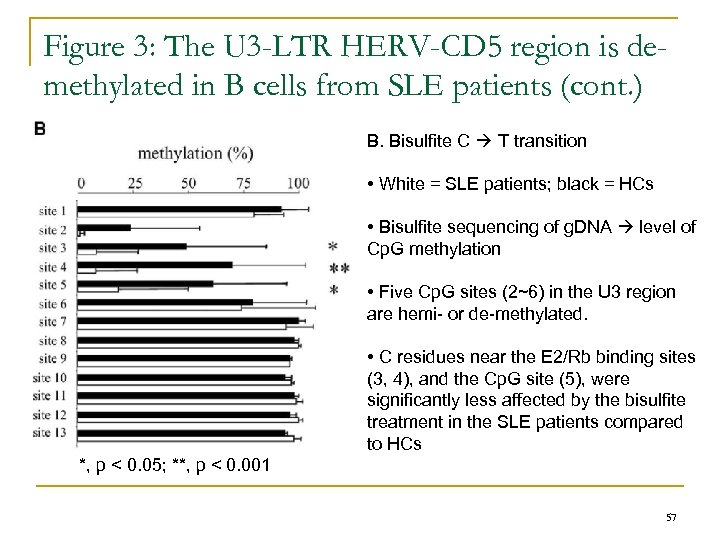 Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients (cont. ) B. Bisulfite C T transition • White = SLE patients; black = HCs • Bisulfite sequencing of g. DNA level of Cp. G methylation • Five Cp. G sites (2~6) in the U 3 region are hemi- or de-methylated. • C residues near the E 2/Rb binding sites (3, 4), and the Cp. G site (5), were significantly less affected by the bisulfite treatment in the SLE patients compared to HCs *, p < 0. 05; **, p < 0. 001 57
Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients (cont. ) B. Bisulfite C T transition • White = SLE patients; black = HCs • Bisulfite sequencing of g. DNA level of Cp. G methylation • Five Cp. G sites (2~6) in the U 3 region are hemi- or de-methylated. • C residues near the E 2/Rb binding sites (3, 4), and the Cp. G site (5), were significantly less affected by the bisulfite treatment in the SLE patients compared to HCs *, p < 0. 05; **, p < 0. 001 57
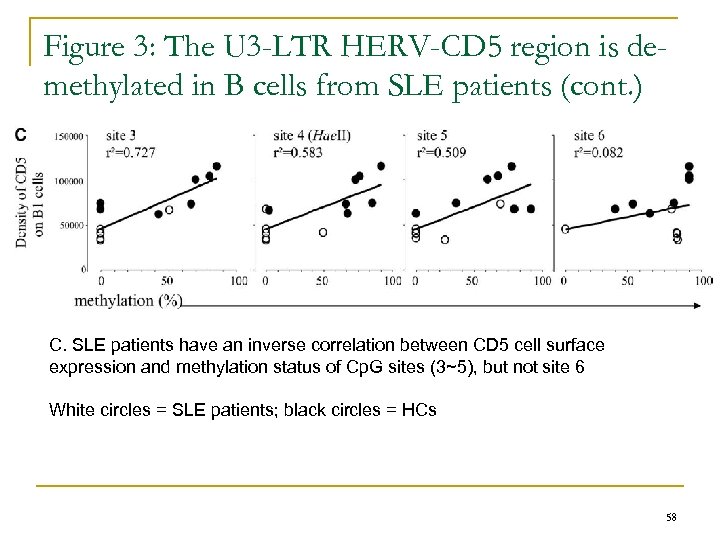 Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients (cont. ) C. SLE patients have an inverse correlation between CD 5 cell surface expression and methylation status of Cp. G sites (3~5), but not site 6 White circles = SLE patients; black circles = HCs 58
Figure 3: The U 3 -LTR HERV-CD 5 region is demethylated in B cells from SLE patients (cont. ) C. SLE patients have an inverse correlation between CD 5 cell surface expression and methylation status of Cp. G sites (3~5), but not site 6 White circles = SLE patients; black circles = HCs 58
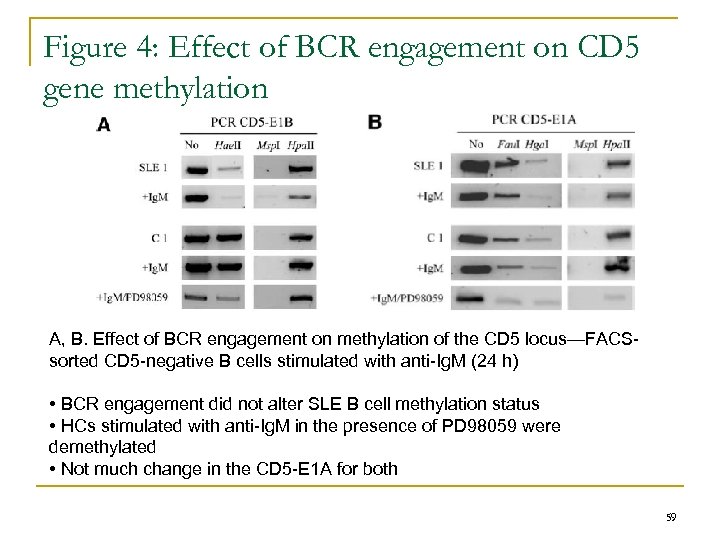 Figure 4: Effect of BCR engagement on CD 5 gene methylation A, B. Effect of BCR engagement on methylation of the CD 5 locus—FACSsorted CD 5 -negative B cells stimulated with anti-Ig. M (24 h) • BCR engagement did not alter SLE B cell methylation status • HCs stimulated with anti-Ig. M in the presence of PD 98059 were demethylated • Not much change in the CD 5 -E 1 A for both 59
Figure 4: Effect of BCR engagement on CD 5 gene methylation A, B. Effect of BCR engagement on methylation of the CD 5 locus—FACSsorted CD 5 -negative B cells stimulated with anti-Ig. M (24 h) • BCR engagement did not alter SLE B cell methylation status • HCs stimulated with anti-Ig. M in the presence of PD 98059 were demethylated • Not much change in the CD 5 -E 1 A for both 59
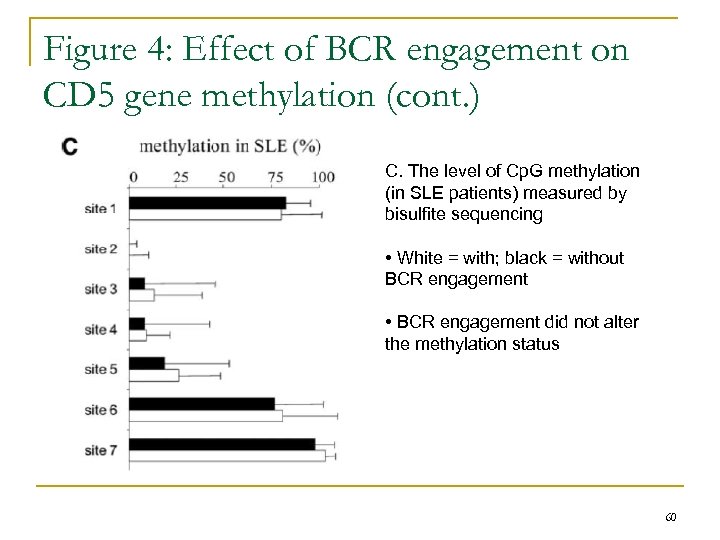 Figure 4: Effect of BCR engagement on CD 5 gene methylation (cont. ) C. The level of Cp. G methylation (in SLE patients) measured by bisulfite sequencing • White = with; black = without BCR engagement • BCR engagement did not alter the methylation status 60
Figure 4: Effect of BCR engagement on CD 5 gene methylation (cont. ) C. The level of Cp. G methylation (in SLE patients) measured by bisulfite sequencing • White = with; black = without BCR engagement • BCR engagement did not alter the methylation status 60
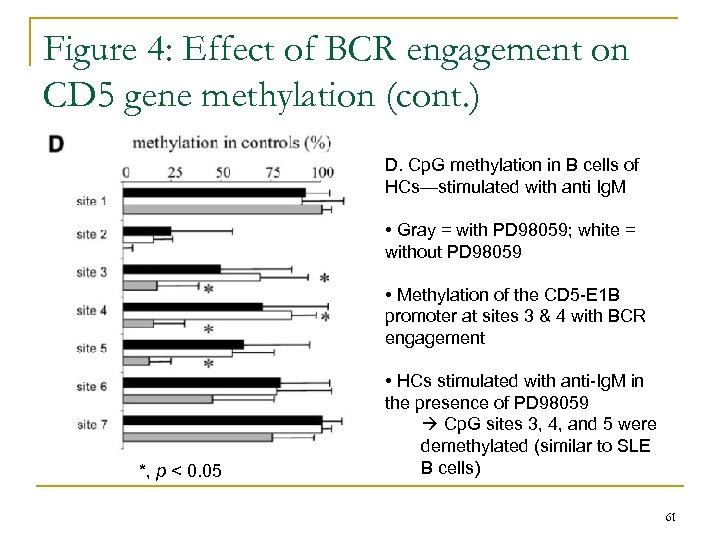 Figure 4: Effect of BCR engagement on CD 5 gene methylation (cont. ) D. Cp. G methylation in B cells of HCs—stimulated with anti Ig. M • Gray = with PD 98059; white = without PD 98059 • Methylation of the CD 5 -E 1 B promoter at sites 3 & 4 with BCR engagement *, p < 0. 05 • HCs stimulated with anti-Ig. M in the presence of PD 98059 Cp. G sites 3, 4, and 5 were demethylated (similar to SLE B cells) 61
Figure 4: Effect of BCR engagement on CD 5 gene methylation (cont. ) D. Cp. G methylation in B cells of HCs—stimulated with anti Ig. M • Gray = with PD 98059; white = without PD 98059 • Methylation of the CD 5 -E 1 B promoter at sites 3 & 4 with BCR engagement *, p < 0. 05 • HCs stimulated with anti-Ig. M in the presence of PD 98059 Cp. G sites 3, 4, and 5 were demethylated (similar to SLE B cells) 61
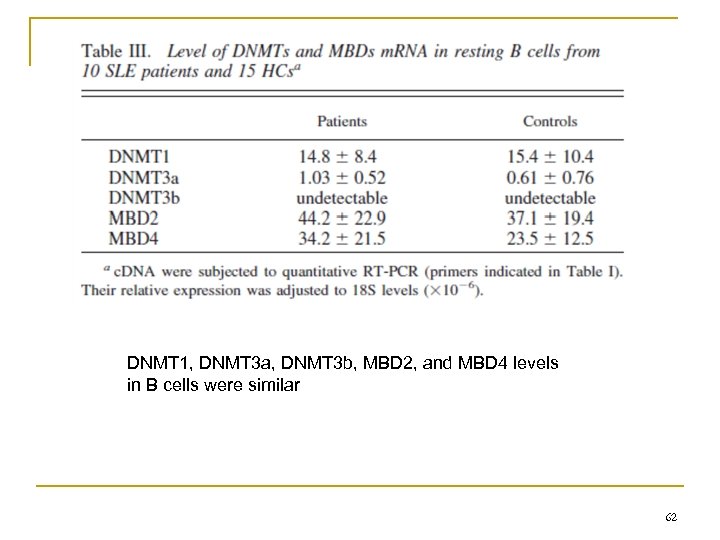 DNMT 1, DNMT 3 a, DNMT 3 b, MBD 2, and MBD 4 levels in B cells were similar 62
DNMT 1, DNMT 3 a, DNMT 3 b, MBD 2, and MBD 4 levels in B cells were similar 62
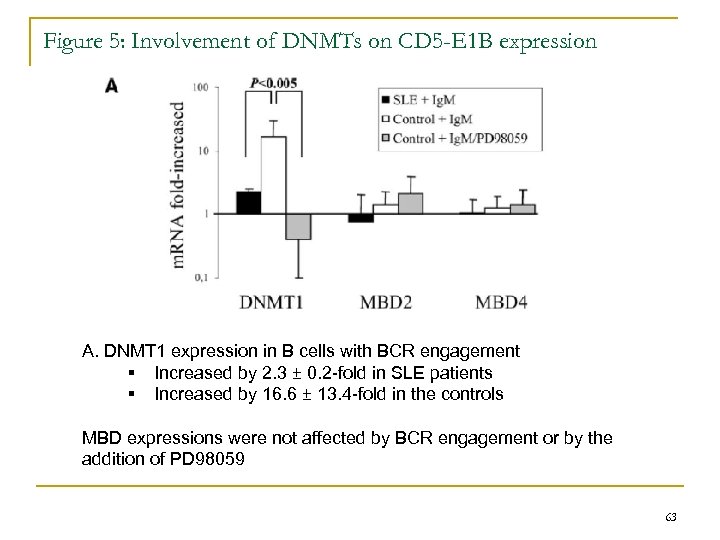 Figure 5: Involvement of DNMTs on CD 5 -E 1 B expression A. DNMT 1 expression in B cells with BCR engagement § Increased by 2. 3 ± 0. 2 -fold in SLE patients § Increased by 16. 6 ± 13. 4 -fold in the controls MBD expressions were not affected by BCR engagement or by the addition of PD 98059 63
Figure 5: Involvement of DNMTs on CD 5 -E 1 B expression A. DNMT 1 expression in B cells with BCR engagement § Increased by 2. 3 ± 0. 2 -fold in SLE patients § Increased by 16. 6 ± 13. 4 -fold in the controls MBD expressions were not affected by BCR engagement or by the addition of PD 98059 63
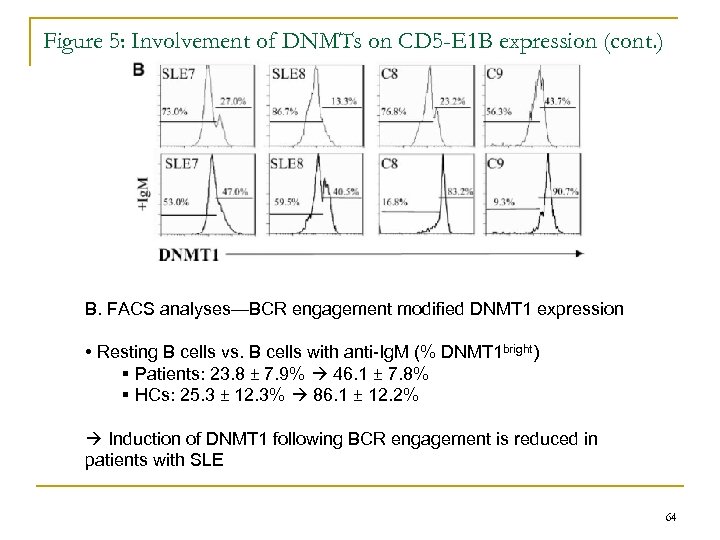 Figure 5: Involvement of DNMTs on CD 5 -E 1 B expression (cont. ) B. FACS analyses—BCR engagement modified DNMT 1 expression • Resting B cells vs. B cells with anti-Ig. M (% DNMT 1 bright) § Patients: 23. 8 ± 7. 9% 46. 1 ± 7. 8% § HCs: 25. 3 ± 12. 3% 86. 1 ± 12. 2% Induction of DNMT 1 following BCR engagement is reduced in patients with SLE 64
Figure 5: Involvement of DNMTs on CD 5 -E 1 B expression (cont. ) B. FACS analyses—BCR engagement modified DNMT 1 expression • Resting B cells vs. B cells with anti-Ig. M (% DNMT 1 bright) § Patients: 23. 8 ± 7. 9% 46. 1 ± 7. 8% § HCs: 25. 3 ± 12. 3% 86. 1 ± 12. 2% Induction of DNMT 1 following BCR engagement is reduced in patients with SLE 64
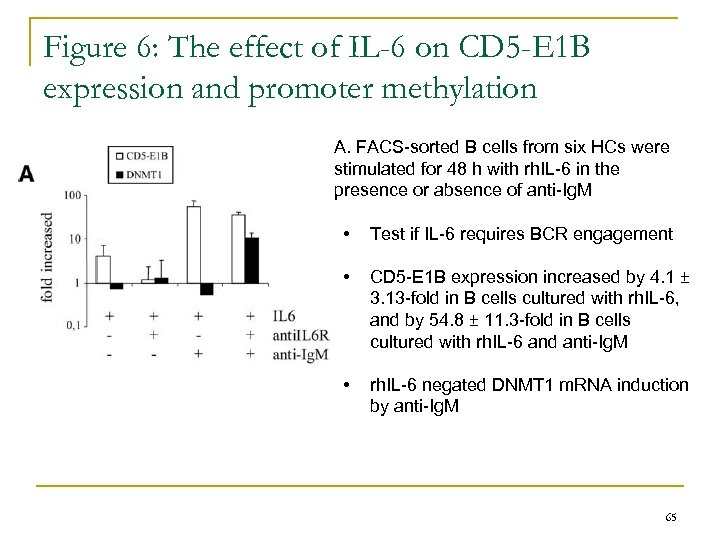 Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation A. FACS-sorted B cells from six HCs were stimulated for 48 h with rh. IL-6 in the presence or absence of anti-Ig. M • Test if IL-6 requires BCR engagement • CD 5 -E 1 B expression increased by 4. 1 ± 3. 13 -fold in B cells cultured with rh. IL-6, and by 54. 8 ± 11. 3 -fold in B cells cultured with rh. IL-6 and anti-Ig. M • rh. IL-6 negated DNMT 1 m. RNA induction by anti-Ig. M 65
Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation A. FACS-sorted B cells from six HCs were stimulated for 48 h with rh. IL-6 in the presence or absence of anti-Ig. M • Test if IL-6 requires BCR engagement • CD 5 -E 1 B expression increased by 4. 1 ± 3. 13 -fold in B cells cultured with rh. IL-6, and by 54. 8 ± 11. 3 -fold in B cells cultured with rh. IL-6 and anti-Ig. M • rh. IL-6 negated DNMT 1 m. RNA induction by anti-Ig. M 65
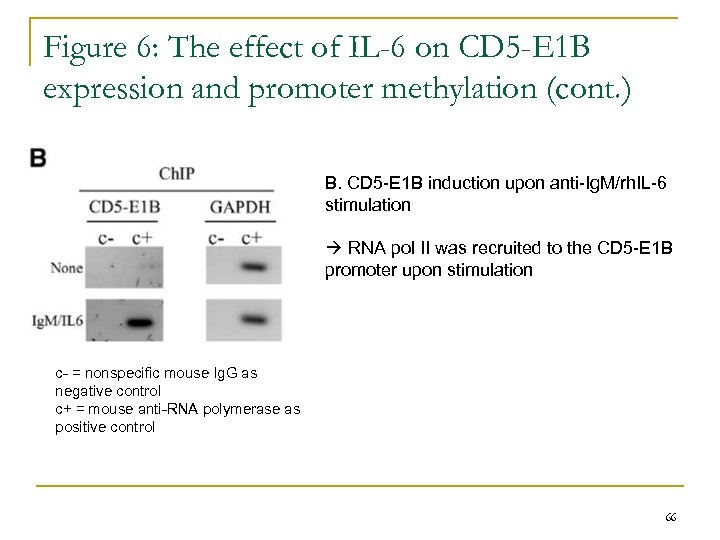 Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation (cont. ) B. CD 5 -E 1 B induction upon anti-Ig. M/rh. IL-6 stimulation RNA pol II was recruited to the CD 5 -E 1 B promoter upon stimulation c- = nonspecific mouse Ig. G as negative control c+ = mouse anti-RNA polymerase as positive control 66
Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation (cont. ) B. CD 5 -E 1 B induction upon anti-Ig. M/rh. IL-6 stimulation RNA pol II was recruited to the CD 5 -E 1 B promoter upon stimulation c- = nonspecific mouse Ig. G as negative control c+ = mouse anti-RNA polymerase as positive control 66
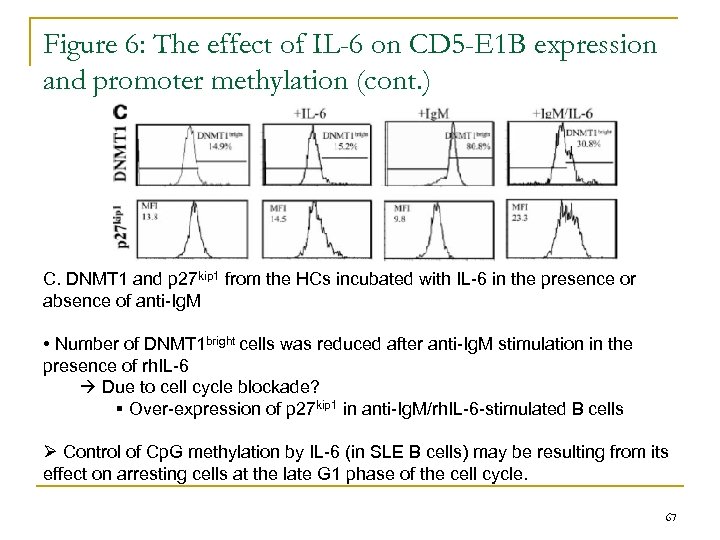 Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation (cont. ) C. DNMT 1 and p 27 kip 1 from the HCs incubated with IL-6 in the presence or absence of anti-Ig. M • Number of DNMT 1 bright cells was reduced after anti-Ig. M stimulation in the presence of rh. IL-6 Due to cell cycle blockade? § Over-expression of p 27 kip 1 in anti-Ig. M/rh. IL-6 -stimulated B cells Ø Control of Cp. G methylation by IL-6 (in SLE B cells) may be resulting from its effect on arresting cells at the late G 1 phase of the cell cycle. 67
Figure 6: The effect of IL-6 on CD 5 -E 1 B expression and promoter methylation (cont. ) C. DNMT 1 and p 27 kip 1 from the HCs incubated with IL-6 in the presence or absence of anti-Ig. M • Number of DNMT 1 bright cells was reduced after anti-Ig. M stimulation in the presence of rh. IL-6 Due to cell cycle blockade? § Over-expression of p 27 kip 1 in anti-Ig. M/rh. IL-6 -stimulated B cells Ø Control of Cp. G methylation by IL-6 (in SLE B cells) may be resulting from its effect on arresting cells at the late G 1 phase of the cell cycle. 67
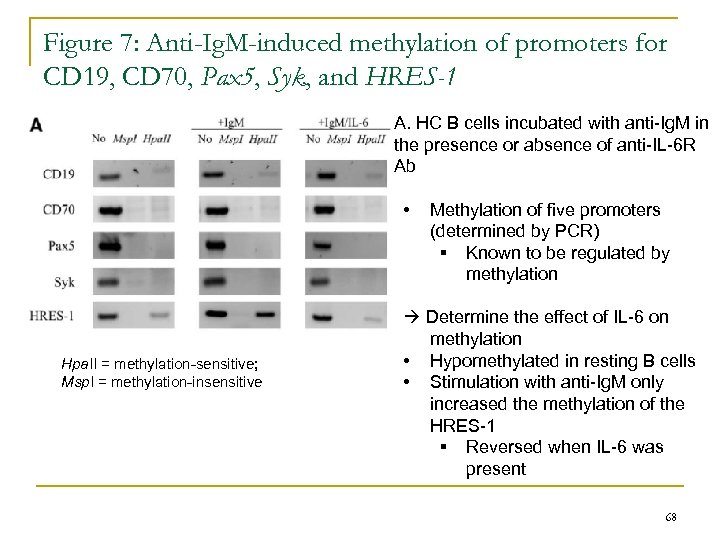 Figure 7: Anti-Ig. M-induced methylation of promoters for CD 19, CD 70, Pax 5, Syk, and HRES-1 A. HC B cells incubated with anti-Ig. M in the presence or absence of anti-IL-6 R Ab • Hpa. II = methylation-sensitive; Msp. I = methylation-insensitive Methylation of five promoters (determined by PCR) § Known to be regulated by methylation Determine the effect of IL-6 on methylation • Hypomethylated in resting B cells • Stimulation with anti-Ig. M only increased the methylation of the HRES-1 § Reversed when IL-6 was present 68
Figure 7: Anti-Ig. M-induced methylation of promoters for CD 19, CD 70, Pax 5, Syk, and HRES-1 A. HC B cells incubated with anti-Ig. M in the presence or absence of anti-IL-6 R Ab • Hpa. II = methylation-sensitive; Msp. I = methylation-insensitive Methylation of five promoters (determined by PCR) § Known to be regulated by methylation Determine the effect of IL-6 on methylation • Hypomethylated in resting B cells • Stimulation with anti-Ig. M only increased the methylation of the HRES-1 § Reversed when IL-6 was present 68
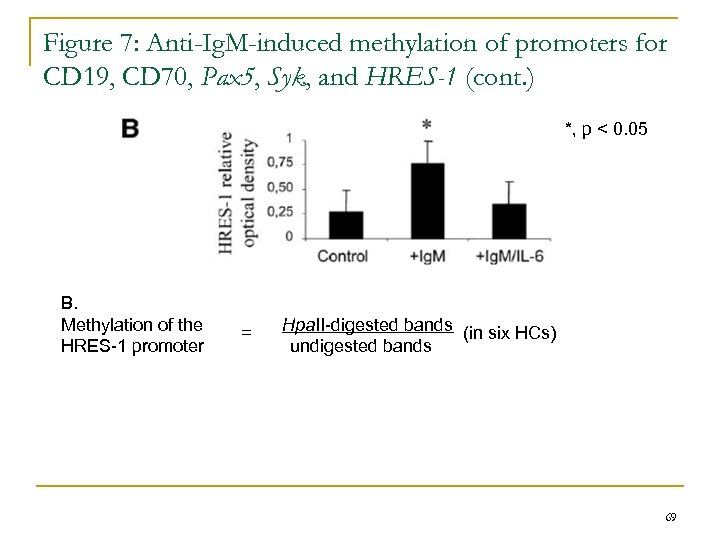 Figure 7: Anti-Ig. M-induced methylation of promoters for CD 19, CD 70, Pax 5, Syk, and HRES-1 (cont. ) *, p < 0. 05 B. Methylation of the HRES-1 promoter = Hpa. II-digested bands (in six HCs) undigested bands 69
Figure 7: Anti-Ig. M-induced methylation of promoters for CD 19, CD 70, Pax 5, Syk, and HRES-1 (cont. ) *, p < 0. 05 B. Methylation of the HRES-1 promoter = Hpa. II-digested bands (in six HCs) undigested bands 69
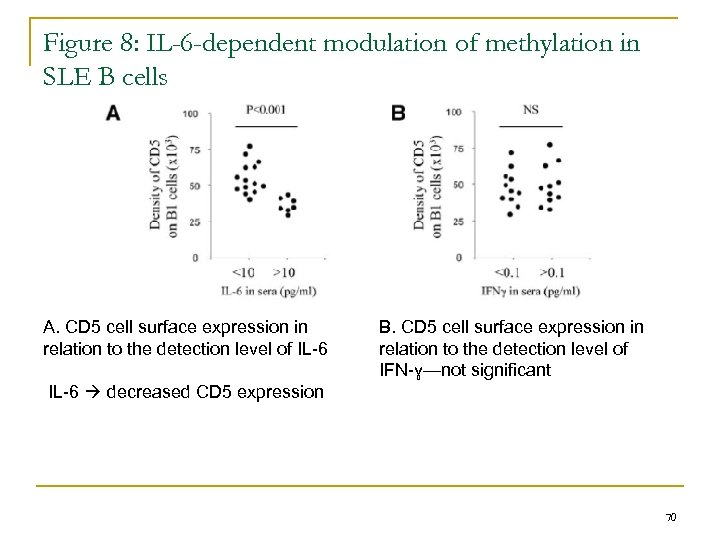 Figure 8: IL-6 -dependent modulation of methylation in SLE B cells A. CD 5 cell surface expression in relation to the detection level of IL-6 B. CD 5 cell surface expression in relation to the detection level of IFN-ɣ—not significant IL-6 decreased CD 5 expression 70
Figure 8: IL-6 -dependent modulation of methylation in SLE B cells A. CD 5 cell surface expression in relation to the detection level of IL-6 B. CD 5 cell surface expression in relation to the detection level of IFN-ɣ—not significant IL-6 decreased CD 5 expression 70
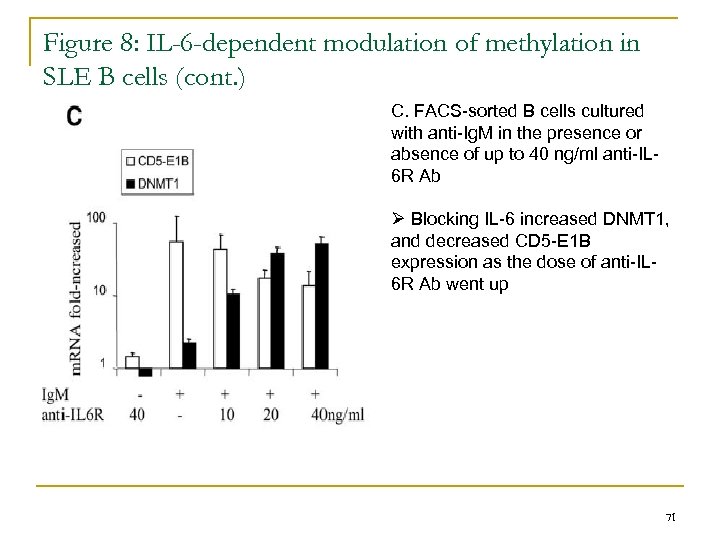 Figure 8: IL-6 -dependent modulation of methylation in SLE B cells (cont. ) C. FACS-sorted B cells cultured with anti-Ig. M in the presence or absence of up to 40 ng/ml anti-IL 6 R Ab Ø Blocking IL-6 increased DNMT 1, and decreased CD 5 -E 1 B expression as the dose of anti-IL 6 R Ab went up 71
Figure 8: IL-6 -dependent modulation of methylation in SLE B cells (cont. ) C. FACS-sorted B cells cultured with anti-Ig. M in the presence or absence of up to 40 ng/ml anti-IL 6 R Ab Ø Blocking IL-6 increased DNMT 1, and decreased CD 5 -E 1 B expression as the dose of anti-IL 6 R Ab went up 71
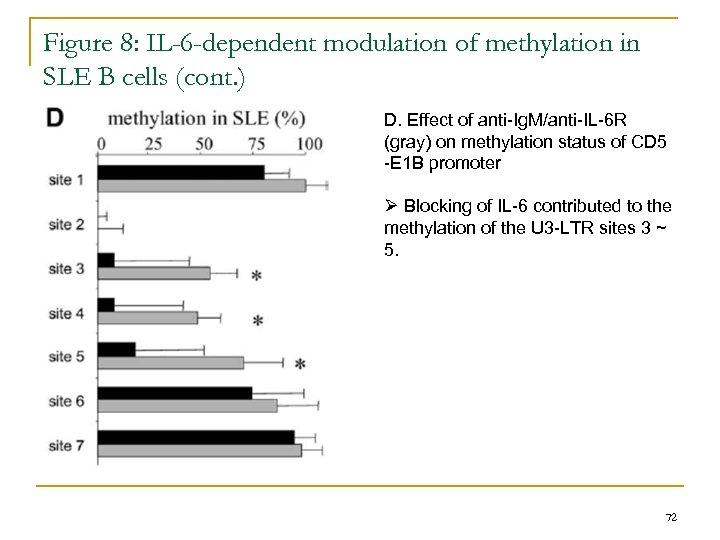 Figure 8: IL-6 -dependent modulation of methylation in SLE B cells (cont. ) D. Effect of anti-Ig. M/anti-IL-6 R (gray) on methylation status of CD 5 -E 1 B promoter Ø Blocking of IL-6 contributed to the methylation of the U 3 -LTR sites 3 ~ 5. 72
Figure 8: IL-6 -dependent modulation of methylation in SLE B cells (cont. ) D. Effect of anti-Ig. M/anti-IL-6 R (gray) on methylation status of CD 5 -E 1 B promoter Ø Blocking of IL-6 contributed to the methylation of the U 3 -LTR sites 3 ~ 5. 72
 Discussion n B cells from SLE patients have reduced membrane CD 5 levels q Due to increased expression of CD 5 -E 1 B (retained in the cytoplasm) CD 5 -E 1 B promoter (U 3 -LTR) is demethylated in SLE B cells q Differential methylation is more pronounced with BCR engagement q CD 5 -E 1 B m. RNA is transcribed at the expense of CD 5 -E 1 A m. RNA transcription B cells (of SLE patients) cannot induce DNMT 1 due to production of IL-6 q Cannot methylate DNA q Similar demethylation patterns of Cp. G islands when compared to HCs stimulated with IL-6 or PD 98059 73
Discussion n B cells from SLE patients have reduced membrane CD 5 levels q Due to increased expression of CD 5 -E 1 B (retained in the cytoplasm) CD 5 -E 1 B promoter (U 3 -LTR) is demethylated in SLE B cells q Differential methylation is more pronounced with BCR engagement q CD 5 -E 1 B m. RNA is transcribed at the expense of CD 5 -E 1 A m. RNA transcription B cells (of SLE patients) cannot induce DNMT 1 due to production of IL-6 q Cannot methylate DNA q Similar demethylation patterns of Cp. G islands when compared to HCs stimulated with IL-6 or PD 98059 73
 Therapeutic Treatment? n IL-6 activates CD 5 -E 1 B transcription q SLE patients with anti-IL-6 R m. Ab could inhibit autoreactive B cell expansion by restoring DNA methylation and cell cycle progression 74
Therapeutic Treatment? n IL-6 activates CD 5 -E 1 B transcription q SLE patients with anti-IL-6 R m. Ab could inhibit autoreactive B cell expansion by restoring DNA methylation and cell cycle progression 74


