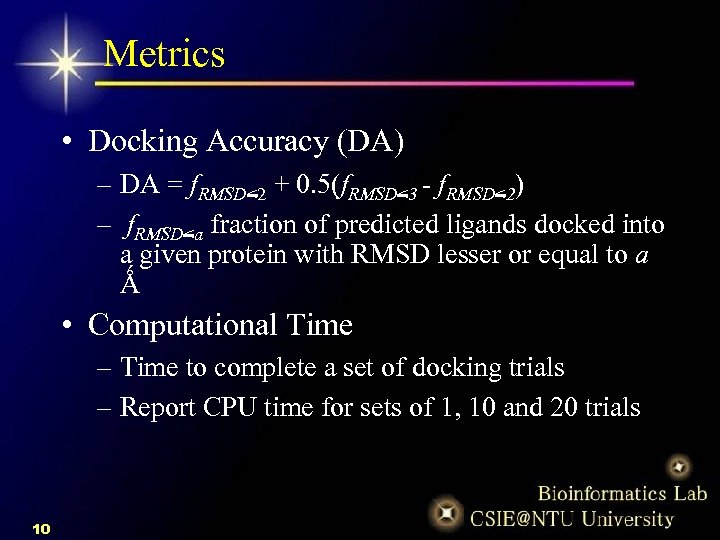
9e05faea44ee9656778602003858a30f.ppt
- Количество слайдов: 20
 Study of Highly Accurate and Fast Protein-Ligand Docking Method Based on Molecular Dynamics M. Taufer, M. Crowley, D. J. Price, A. A. Chien‡ and C. L. Brooks III∗, † Department of Molecular Biology (TPC 6), The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, U. S. A. Reporter: Yu Lun Kuo E-mail: sscc 6991@gmail. com Date: November 21, 2006 Published online 24 June 2005 in Wiley Inter. Science 1
Study of Highly Accurate and Fast Protein-Ligand Docking Method Based on Molecular Dynamics M. Taufer, M. Crowley, D. J. Price, A. A. Chien‡ and C. L. Brooks III∗, † Department of Molecular Biology (TPC 6), The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, U. S. A. Reporter: Yu Lun Kuo E-mail: sscc 6991@gmail. com Date: November 21, 2006 Published online 24 June 2005 in Wiley Inter. Science 1
 Outline • • • 2 Introduction MD-based Docking Method Algorithm Evaluation Metrics MD vs. Other Methods Conclusion & Future work
Outline • • • 2 Introduction MD-based Docking Method Algorithm Evaluation Metrics MD vs. Other Methods Conclusion & Future work
 Introduction (1/3) • Drug development • Use of small molecules (ligand) to turn on or off a protein function • Protein-ligand docking • Computational methods for the prediction of ligandprotein structure information 3
Introduction (1/3) • Drug development • Use of small molecules (ligand) to turn on or off a protein function • Protein-ligand docking • Computational methods for the prediction of ligandprotein structure information 3
 Introduction (2/3) • Exiting docking method – Current docking algorithms are fast and use simplified scoring function to direct conformational search and select the best structure – Methods based on molecular dynamics (MD) and atomically detailed force field (e. g. , CDOCKER) are more accurate but time- and resource-expensive. 4
Introduction (2/3) • Exiting docking method – Current docking algorithms are fast and use simplified scoring function to direct conformational search and select the best structure – Methods based on molecular dynamics (MD) and atomically detailed force field (e. g. , CDOCKER) are more accurate but time- and resource-expensive. 4
 Introduction (3/3) • Desktop grids – By scavenging for available and idle cycles – Provide computing power at a significant cost saving • Our algorithm – Parallel and each simulation attempt is decomposable into independent sub-jobs 5
Introduction (3/3) • Desktop grids – By scavenging for available and idle cycles – Provide computing power at a significant cost saving • Our algorithm – Parallel and each simulation attempt is decomposable into independent sub-jobs 5
 MD-based Docking Method (1/2) • Goals – Assure accuracy • Benefit from the molecular mechanics force fields – Guarantee performance • Return docking results in a short turnaround time using cost-effective platforms • Approach – Docking method based on CHARMM molecular dynamics simulations and with a highly flexible computational granularity 6
MD-based Docking Method (1/2) • Goals – Assure accuracy • Benefit from the molecular mechanics force fields – Guarantee performance • Return docking results in a short turnaround time using cost-effective platforms • Approach – Docking method based on CHARMM molecular dynamics simulations and with a highly flexible computational granularity 6
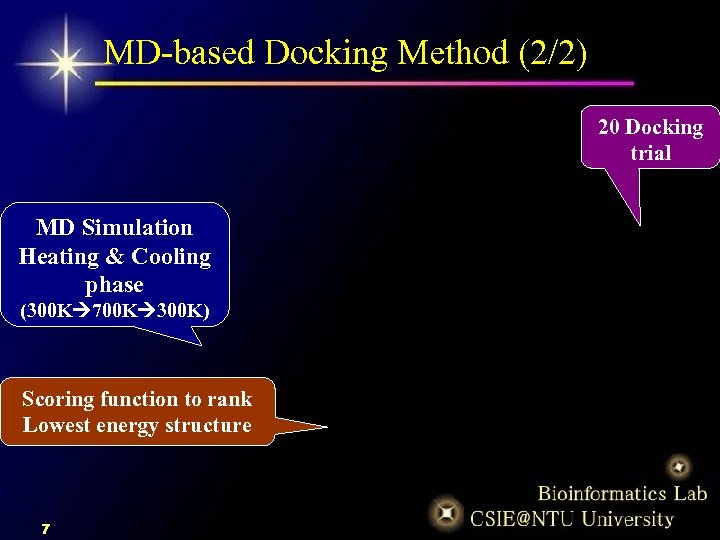 MD-based Docking Method (2/2) 20 Docking trial MD Simulation Heating & Cooling phase (300 K 700 K 300 K) Scoring function to rank Lowest energy structure 7
MD-based Docking Method (2/2) 20 Docking trial MD Simulation Heating & Cooling phase (300 K 700 K 300 K) Scoring function to rank Lowest energy structure 7
 Algorithm Evaluation (1/2) • Characterization of the docking method: – Does the MD length affect the docking accuracy? – Does the number of trials affect the docking accuracy? • Comparing algorithm with other well-known docking methods – Auto. Dock、DOCK、Flex. X、ICM、GOLD 8
Algorithm Evaluation (1/2) • Characterization of the docking method: – Does the MD length affect the docking accuracy? – Does the number of trials affect the docking accuracy? • Comparing algorithm with other well-known docking methods – Auto. Dock、DOCK、Flex. X、ICM、GOLD 8
 Algorithm Evaluation (2/2) • Experimental testbed – Platform • SGI R 10000 – Single 195 MHz IP 2 processor – 128 MB memory • A cluster of 64 dual-processor nodes at the SDSC (San Diego Supercomputer Center) – Data set: 31 protein-ligand complexes • 10 proteins • 31 ligands with different levels of complexity 9
Algorithm Evaluation (2/2) • Experimental testbed – Platform • SGI R 10000 – Single 195 MHz IP 2 processor – 128 MB memory • A cluster of 64 dual-processor nodes at the SDSC (San Diego Supercomputer Center) – Data set: 31 protein-ligand complexes • 10 proteins • 31 ligands with different levels of complexity 9
 Four Different MD Simulations 11
Four Different MD Simulations 11
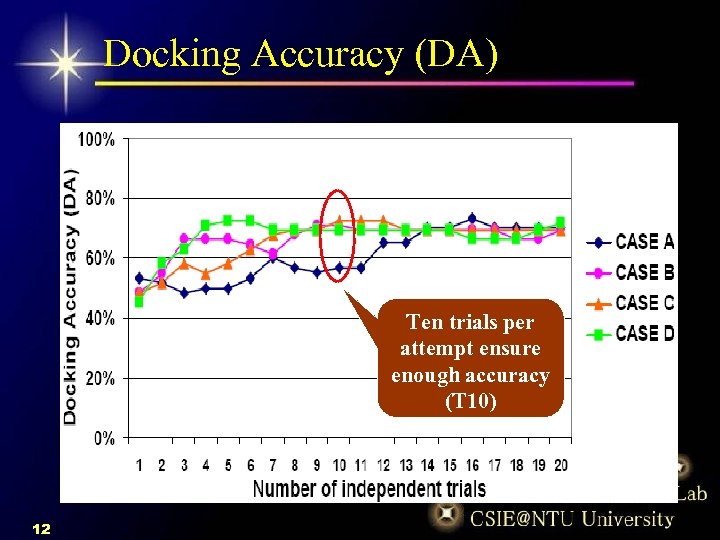 Docking Accuracy (DA) Ten trials per attempt ensure enough accuracy (T 10) 12
Docking Accuracy (DA) Ten trials per attempt ensure enough accuracy (T 10) 12
 Average Time with Different Number of MD Steps Increase of number of MD steps Almost linear increase of the simulation time 13
Average Time with Different Number of MD Steps Increase of number of MD steps Almost linear increase of the simulation time 13
 MD vs. Other Methods • Other Methods – Auto. Dock、 DOCK、Flex. X、ICM、GOLD • Comparison Metrics – Docking Accuracy (DA) – RMSD of predicted ligands – CPU time per attempt • Definition of attempt – Consider CASE B and 10 trials per attempt (T 10) 14
MD vs. Other Methods • Other Methods – Auto. Dock、 DOCK、Flex. X、ICM、GOLD • Comparison Metrics – Docking Accuracy (DA) – RMSD of predicted ligands – CPU time per attempt • Definition of attempt – Consider CASE B and 10 trials per attempt (T 10) 14
 Comparison of Docking Accuracy 15
Comparison of Docking Accuracy 15
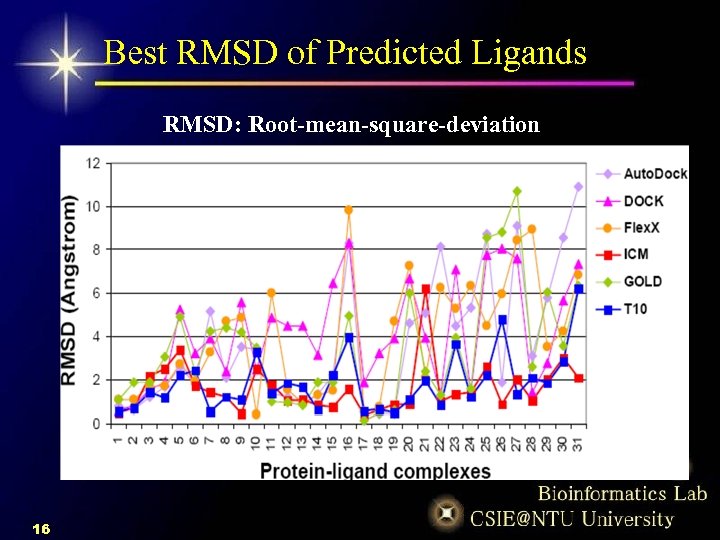 Best RMSD of Predicted Ligands RMSD: Root-mean-square-deviation 16
Best RMSD of Predicted Ligands RMSD: Root-mean-square-deviation 16
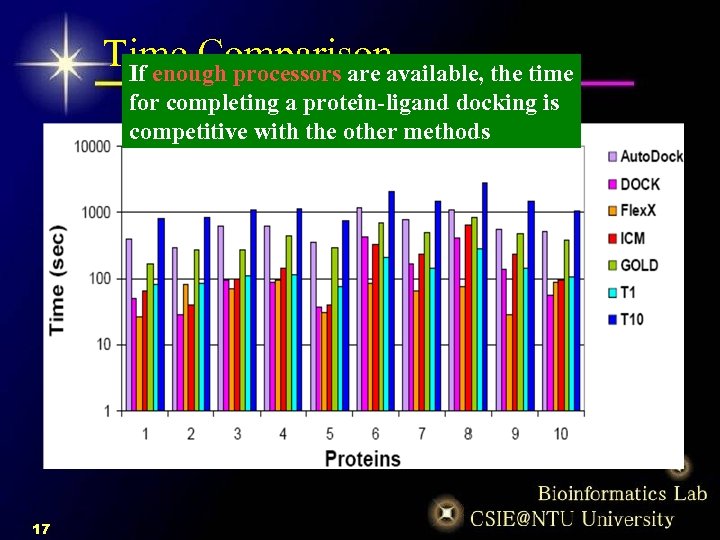 Time Comparisonavailable, the time If enough processors are for completing a protein-ligand docking is competitive with the other methods 17
Time Comparisonavailable, the time If enough processors are for completing a protein-ligand docking is competitive with the other methods 17
 Conclusion & Future work • The MD-based docking method – Reach an average accuracy of 71% • Still a lot of exciting research has to be addressed both at the application and system levels – Number of ligand orientations per trial based on resources and node reliability 18
Conclusion & Future work • The MD-based docking method – Reach an average accuracy of 71% • Still a lot of exciting research has to be addressed both at the application and system levels – Number of ligand orientations per trial based on resources and node reliability 18
 Conclusion & Future work • Future work – Plan to make a more detailed study of MD and Monte Carlo simulations for the docking process in the near future. • ICM running multiple Monte Carlo minimizations • Our docking protocol to desktop grids – Proportionally decreases the time to solution – Fine-grained parallel algorithm for docking trial 19
Conclusion & Future work • Future work – Plan to make a more detailed study of MD and Monte Carlo simulations for the docking process in the near future. • ICM running multiple Monte Carlo minimizations • Our docking protocol to desktop grids – Proportionally decreases the time to solution – Fine-grained parallel algorithm for docking trial 19
 Thanks for your attention 20
Thanks for your attention 20



