75750c233d0beb6c85858b158c05c4e0.ppt
- Количество слайдов: 22

Stability of the core domain of p 53 Insights from MD simulations A Madhumalar, Derek Smith, Chandra Verma Bioinformatics Institute (A-STAR) Singapore
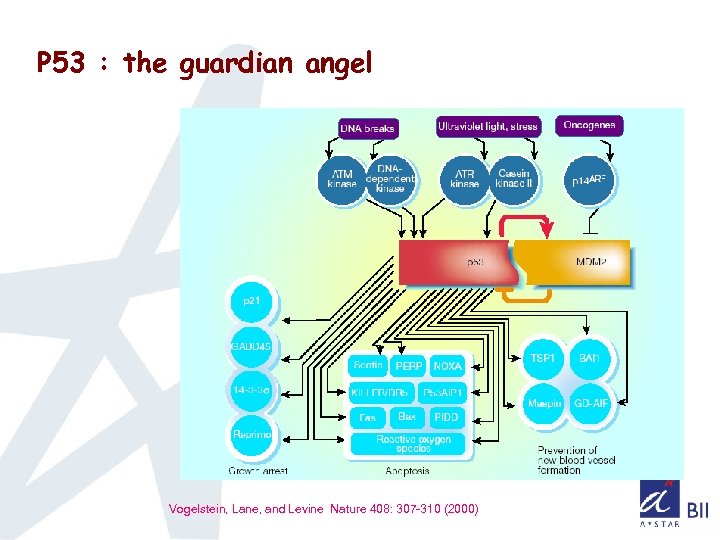
P 53 : the guardian angel Vogelstein, Lane, and Levine Nature 408: 307 -310 (2000)
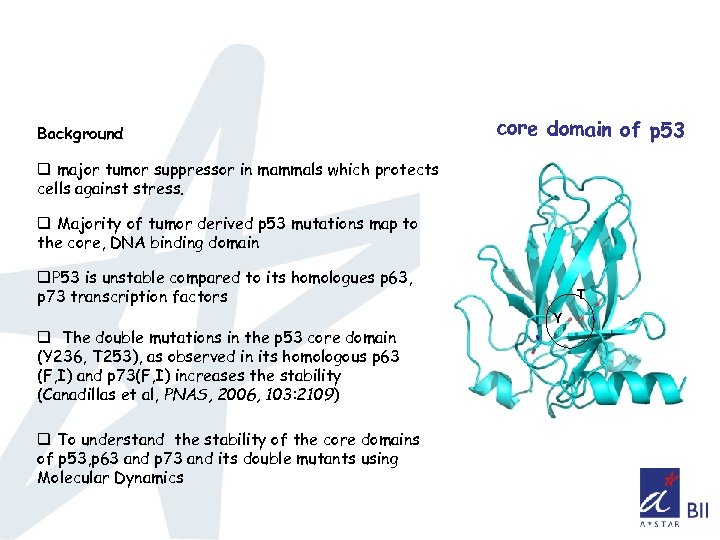
Background core domain of p 53 q major tumor suppressor in mammals which protects cells against stress. q Majority of tumor derived p 53 mutations map to the core, DNA binding domain q. P 53 is unstable compared to its homologues p 63, p 73 transcription factors q The double mutations in the p 53 core domain (Y 236, T 253), as observed in its homologous p 63 (F, I) and p 73(F, I) increases the stability (Canadillas et al, PNAS, 2006, 103: 2109) q To understand the stability of the core domains of p 53, p 63 and p 73 and its double mutants using Molecular Dynamics T Y
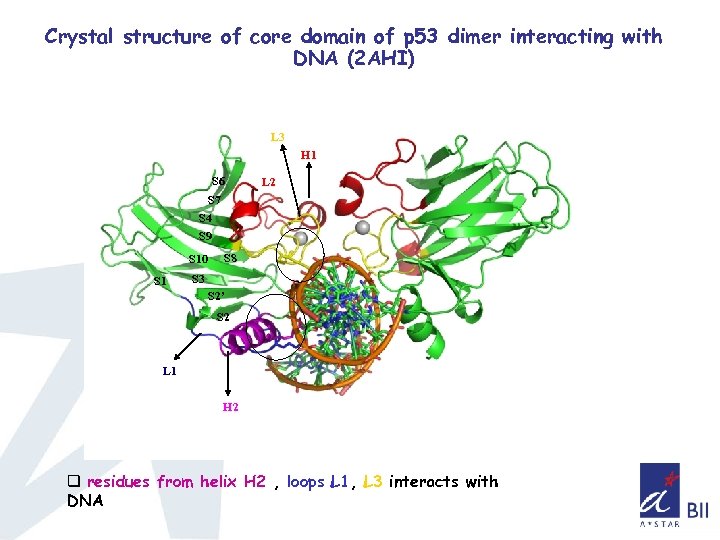
Crystal structure of core domain of p 53 dimer interacting with DNA (2 AHI) L 3 H 1 S 6 L 2 S 7 S 4 S 9 S 10 S 1 S 8 S 3 S 2’ S 2 L 1 H 2 q residues from helix H 2 , loops L 1, L 3 interacts with DNA
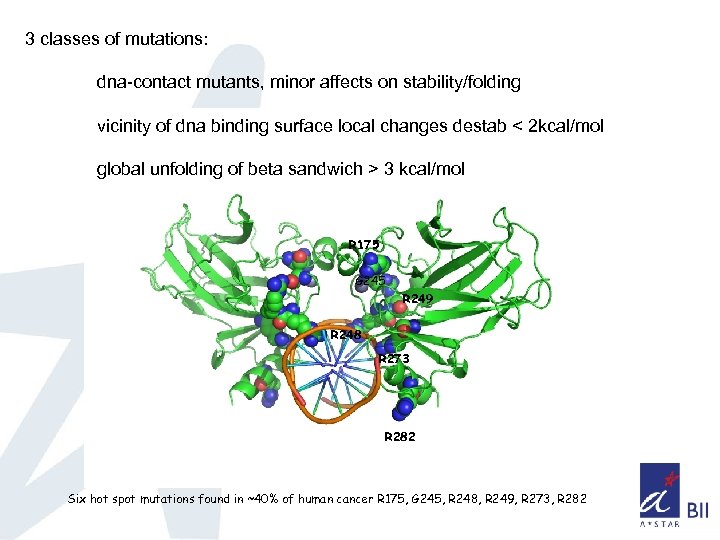
3 classes of mutations: dna-contact mutants, minor affects on stability/folding vicinity of dna binding surface local changes destab < 2 kcal/mol global unfolding of beta sandwich > 3 kcal/mol R 175 G 245 R 249 R 248 R 273 R 282 Six hot spot mutations found in ~40% of human cancer R 175, G 245, R 248, R 249, R 273, R 282
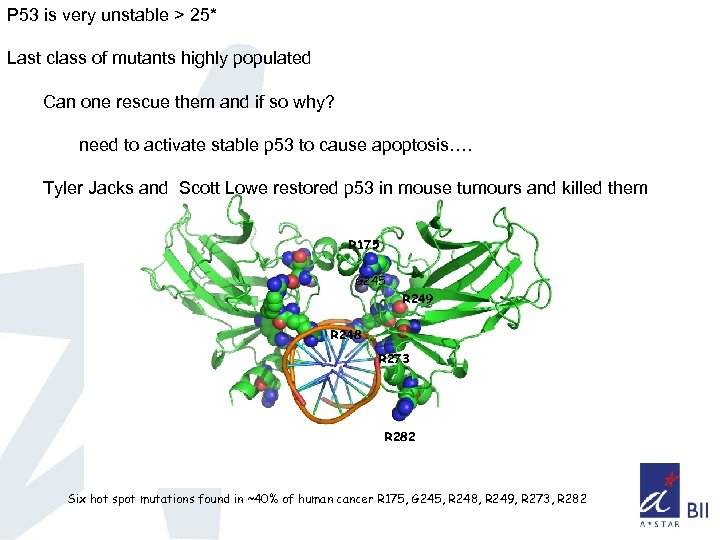
P 53 is very unstable > 25* Last class of mutants highly populated Can one rescue them and if so why? need to activate stable p 53 to cause apoptosis…. Tyler Jacks and Scott Lowe restored p 53 in mouse tumours and killed them R 175 G 245 R 249 R 248 R 273 R 282 Six hot spot mutations found in ~40% of human cancer R 175, G 245, R 248, R 249, R 273, R 282
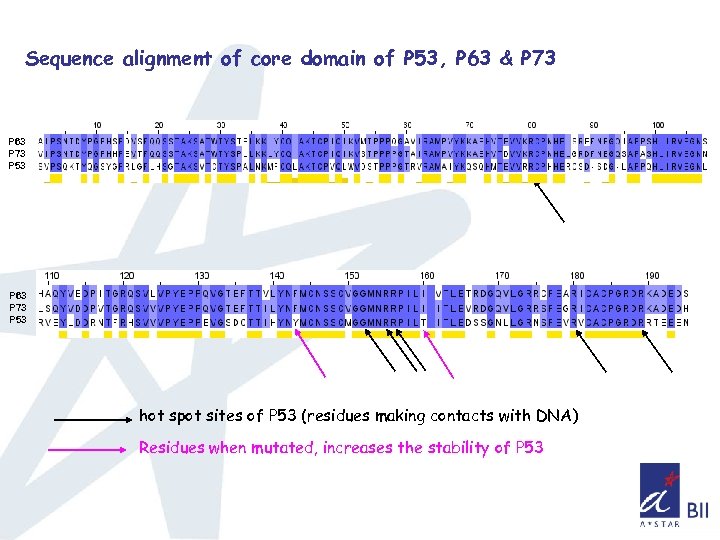
Sequence alignment of core domain of P 53, P 63 & P 73 P 63 P 73 P 53 hot spot sites of P 53 (residues making contacts with DNA) Residues when mutated, increases the stability of P 53
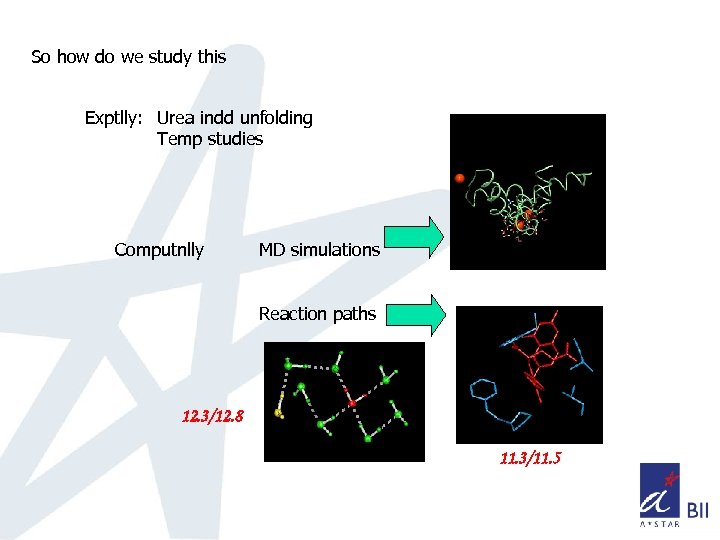
So how do we study this Exptlly: Urea indd unfolding Temp studies Computnlly MD simulations Reaction paths 12. 3/12. 8 11. 3/11. 5
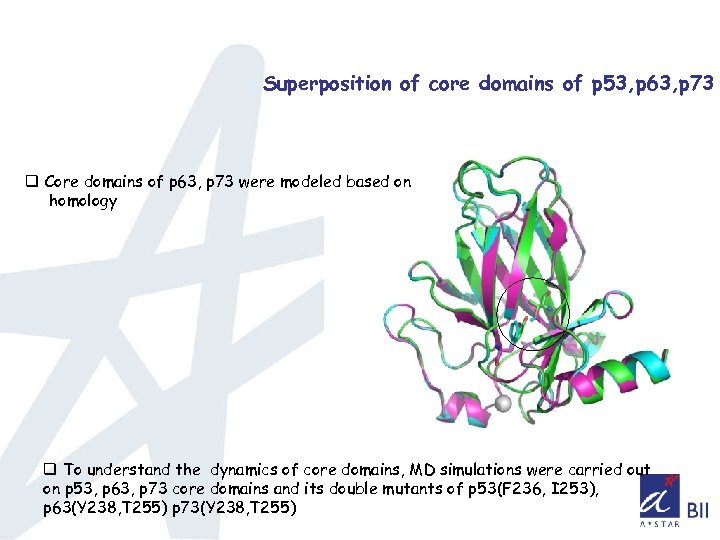
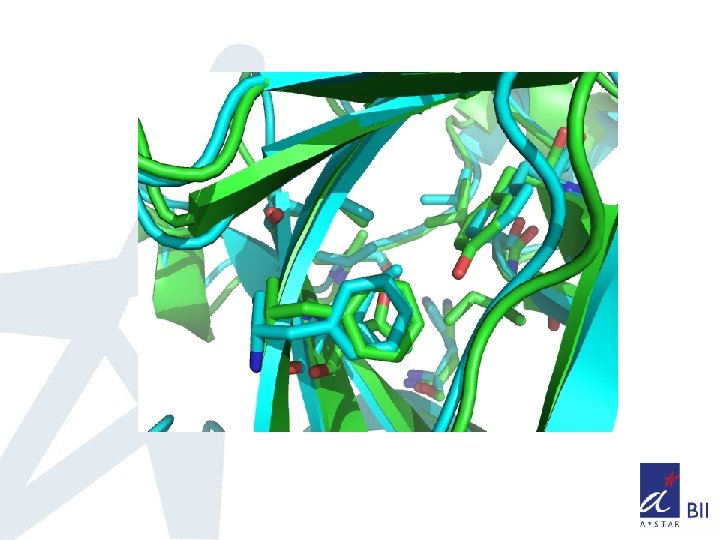
Superposition of core domains of p 53, p 63, p 73 q Core domains of p 63, p 73 were modeled based on homology q To understand the dynamics of core domains, MD simulations were carried out on p 53, p 63, p 73 core domains and its double mutants of p 53(F 236, I 253), p 63(Y 238, T 255) p 73(Y 238, T 255)
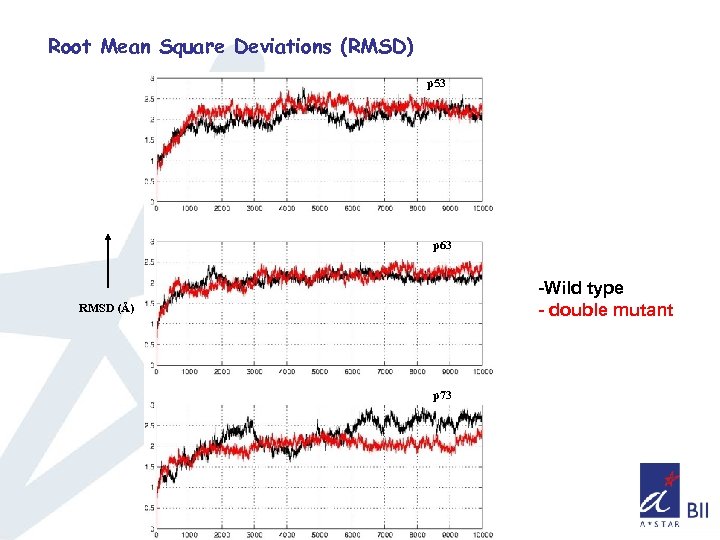
Root Mean Square Deviations (RMSD) p 53 p 63 -Wild type - double mutant RMSD (Å) p 73
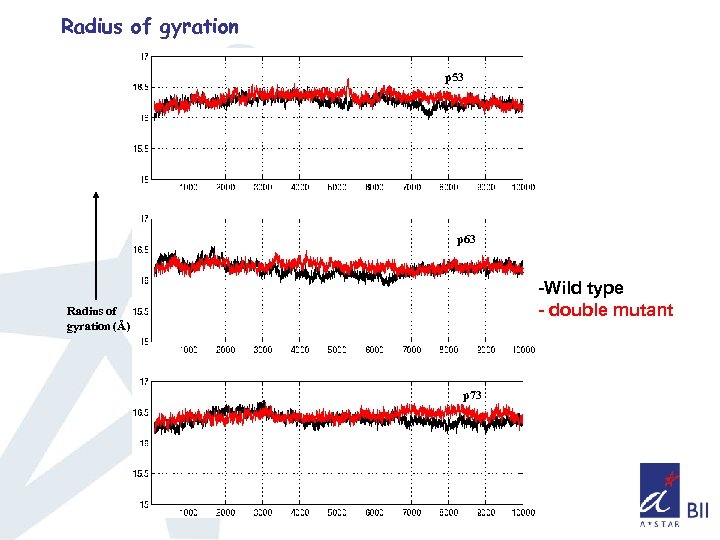
Radius of gyration p 53 p 63 -Wild type - double mutant Radius of gyration (Å) p 73
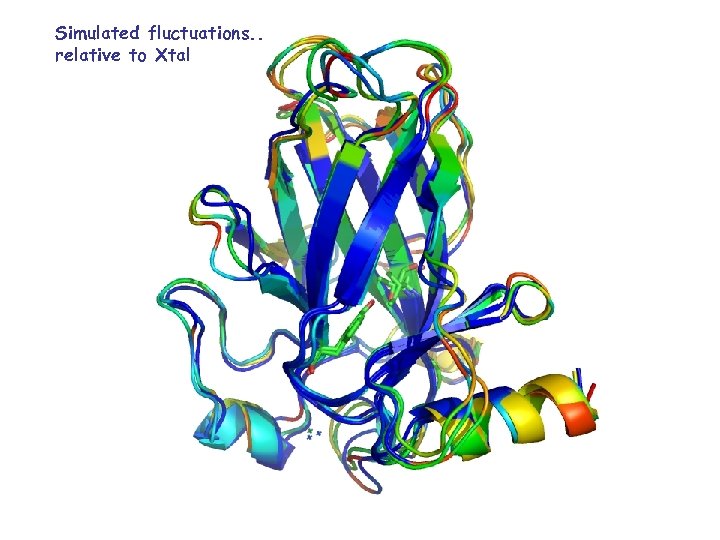
Simulated fluctuations. . relative to Xtal
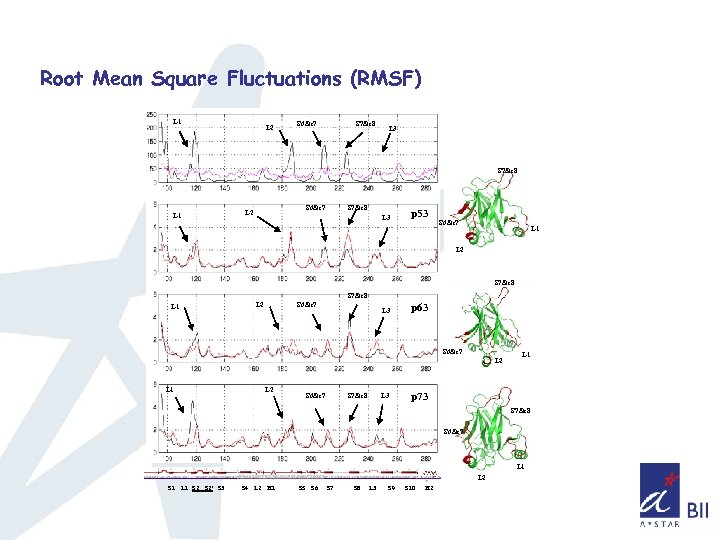
Root Mean Square Fluctuations (RMSF) L 1 L 2 S 6&s 7 S 7&s 8 L 3 S 7&s 8 S 6&s 7 L 2 L 1 S 7&s 8 L 3 p 53 S 6&s 7 L 1 L 2 S 7&s 8 S 6&s 7 L 2 L 1 L 3 p 63 S 6&s 7 L 2 L 1 L 2 S 6&s 7 S 7&s 8 L 3 L 1 p 73 S 7&s 8 S 6&s 7 L 1 L 2 S 1 L 1 S 2’ S 3 S 4 L 2 H 1 S 5 S 6 S 7 S 8 L 3 S 9 S 10 H 2
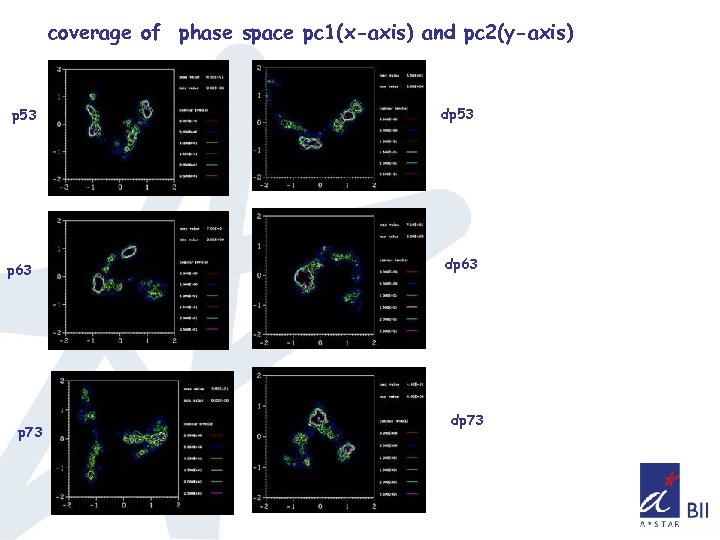
coverage of phase space pc 1(x-axis) and pc 2(y-axis) p 53 p 63 p 73 dp 53 dp 63 dp 73

Summary so far. . q. Double mutant doesn’t affect the overall structure of the core domain in all three cases as seen from the RMSD and Radius of gyration q. Consistent difference in the fluctuations of Loop L 1(observed in NMR), L 2, and loops connecting S 5 & S 6, S 7 &S 8 is observed q Sampling of phase space seems higher for p 63, p 73 compared to p 53 q Sampling of phase space also indicates that the overall mobility has been increased in the case of double mutant of p 53 and decreased in the double mutant of p 63, p 73

Structural plasticity and stability q. What is stability? q. Urea induced unfolding as a fn of temp… q. So how does urea get in? q. Can we look at certain types of motions that may give us a clue
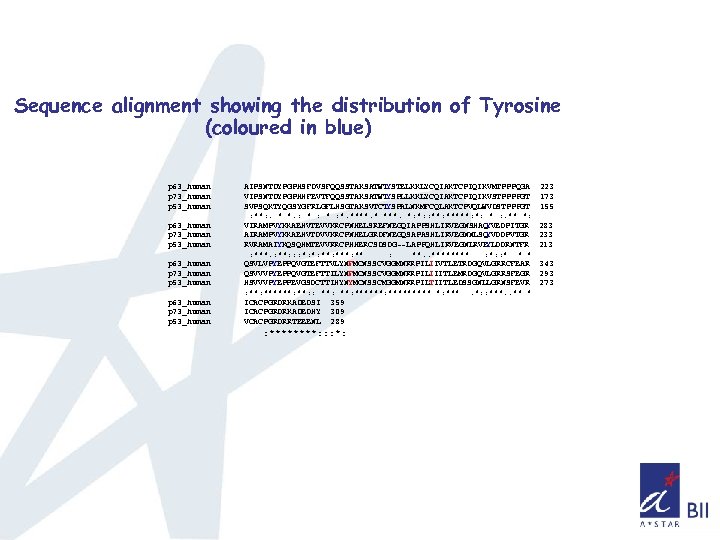

Sequence alignment showing the distribution of Tyrosine (coloured in blue) p 63_human p 73_human p 53_human AIPSNTDYPGPHSFDVSFQQSSTAKSATWT YSTELKKLYCQIAKTCPIQIKVMTPPPQGA VIPSNTDYPGPHHFEVTFQQSSTAKSATWT YSPLLKKLYCQIAKTCPIQIKVSTPPPPGT SVPSQKTYQGSYGFRLGFLHSGTAKSVTCT YSPALNKMFCQLAKTCPVQLWVDSTPPPGT : **: . * *. : * : *. ****. *: *: : *****: *: * : . ** *: VIRAMPVYKKAEHVTEVVKRCPNHELSREFNEGQIAPPSHLIRVEGNSHAQ YVEDPITGR AIRAMPVYKKAEHVTDVVKRCPNHELGRDFNEGQSAPASHLIRVEGNNLSQ YVDDPVTGR RVRAMAIYKQSQHMTEVVRRCPHHERCSDSDG--LAPPQHLIRVEGNLRVE YLDDRNTFR : ***. : **: : : *: *: ***: **. . **** : *: : * * * QSVLVPYEPPQVGTEFTTVLYNFMCNSSCVGGMNRRPILIIVTLETRDGQVLGRRCFEAR QSVVVPYEPPQVGTEFTTILYNFMCNSSCVGGMNRRPILIIITLEMRDGQVLGRRSFEGR HSVVVPYEPPEVGSDCTTIHYNYMCNSSCMGGMNRRPILTIITLEDSSGNLLGRNSFEVR : ******: : **: ********* *: ***. *: : ***. . ** * ICACPGRDRKADEDSI 359 ICACPGRDRKADEDHY 309 VCACPGRDRRTEEENL 289 : ****: : : *: 223 173 155 283 233 213 343 293 273
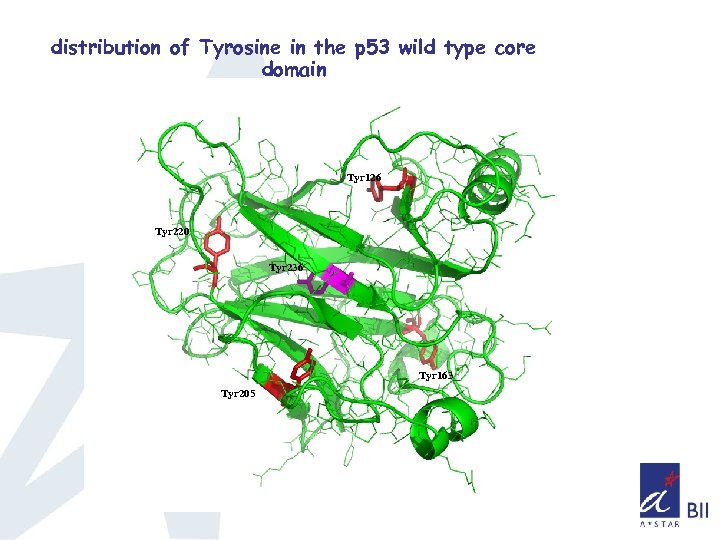
distribution of Tyrosine in the p 53 wild type core domain Tyr 126 Tyr 220 Tyr 236 Tyr 163 Tyr 205
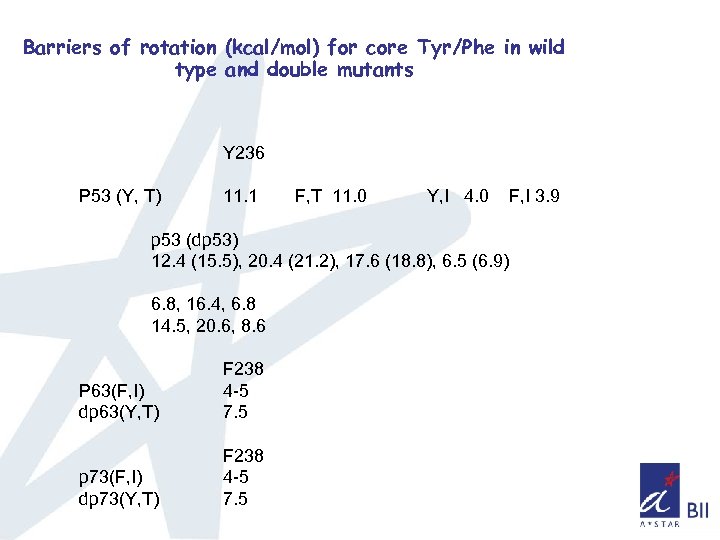

Barriers of rotation (kcal/mol) for core Tyr/Phe in wild type and double mutants Y 236 P 53 (Y, T) 11. 1 F, T 11. 0 Y, I 4. 0 F, I 3. 9 p 53 (dp 53) 12. 4 (15. 5), 20. 4 (21. 2), 17. 6 (18. 8), 6. 5 (6. 9) 6. 8, 16. 4, 6. 8 14. 5, 20. 6, 8. 6 P 63(F, I) dp 63(Y, T) F 238 4 -5 7. 5 p 73(F, I) dp 73(Y, T) F 238 4 -5 7. 5

Summary q Sampling of phase space seems higher for p 63, p 73 compared to p 53 q Sampling of phase space also indicates that the overall mobility has been increased in the case of double mutant of p 53 and decreased in the double mutant of p 63, p 73 q. The decrease in the rate of flipping of aromatic ring in the double mutant suggest that there is increase in the local packing

Thanks!
75750c233d0beb6c85858b158c05c4e0.ppt