abbb2b594306c437284811cf710fc4ec.ppt
- Количество слайдов: 26
 Site Directed Mutagenesis of Protein Pur. E Megan Silas From the University of Illinois at Urbana Champaign In Dr. Fung’s Lab in the Department of Chemistry
Site Directed Mutagenesis of Protein Pur. E Megan Silas From the University of Illinois at Urbana Champaign In Dr. Fung’s Lab in the Department of Chemistry
 Outline n Project Overview n n Bacillus anthracis Purines Pur. E Experimental Procedures and Results n n n Primer Design Polymerase Chain Reaction Transformation Sequencing Protein Purification Activity Assay
Outline n Project Overview n n Bacillus anthracis Purines Pur. E Experimental Procedures and Results n n n Primer Design Polymerase Chain Reaction Transformation Sequencing Protein Purification Activity Assay
 Bacillus anthracis – Anthrax n n n A risk to national security, biological warfare Fatal when untreated Routes of entry to the body: n n Absorption through skin Inhalation Ingestion and then absorption through the digestive tract Need a novel antibiotic to target bacteria that are resistant to current drugs n How can we exploit current knowledge to help discover alternative treatments? Hostettler, Sam. "$14 M Project to Develop Antibiotics against Biowarfare. " UIC News. University of Illinois at Chicago, 18 May 2011. Web. 06 June 2011.
Bacillus anthracis – Anthrax n n n A risk to national security, biological warfare Fatal when untreated Routes of entry to the body: n n Absorption through skin Inhalation Ingestion and then absorption through the digestive tract Need a novel antibiotic to target bacteria that are resistant to current drugs n How can we exploit current knowledge to help discover alternative treatments? Hostettler, Sam. "$14 M Project to Develop Antibiotics against Biowarfare. " UIC News. University of Illinois at Chicago, 18 May 2011. Web. 06 June 2011.
 Bacteria n n In order to survive in human plasma, bacteria must rely on de novo synthesis of many different molecules Studies show nucleotide (purine and pyrimidine) biosynthesis to be the most critical n n Limited availability of nucleotides in human blood Purines: n A major component of DNA, RNA, ATP, GTP, and more Samant, Shalaka, Hyunwoo Lee, Mahmood Ghassemi, Juan Chen, James L. Cook, Alexander S. Mankin, and Alexander A. Neyfakh. "Nucleotide Biosynthesis Is Critical for Growth of Bacteria in Human Blood. " PLo. S Pathogens 4. 2 (2008): E 37. "Purine. " Wikipedia, the Free Encyclopedia. Web. 06 June 2011. http: //en. wikipedia. org/wiki/Purine.
Bacteria n n In order to survive in human plasma, bacteria must rely on de novo synthesis of many different molecules Studies show nucleotide (purine and pyrimidine) biosynthesis to be the most critical n n Limited availability of nucleotides in human blood Purines: n A major component of DNA, RNA, ATP, GTP, and more Samant, Shalaka, Hyunwoo Lee, Mahmood Ghassemi, Juan Chen, James L. Cook, Alexander S. Mankin, and Alexander A. Neyfakh. "Nucleotide Biosynthesis Is Critical for Growth of Bacteria in Human Blood. " PLo. S Pathogens 4. 2 (2008): E 37. "Purine. " Wikipedia, the Free Encyclopedia. Web. 06 June 2011. http: //en. wikipedia. org/wiki/Purine.
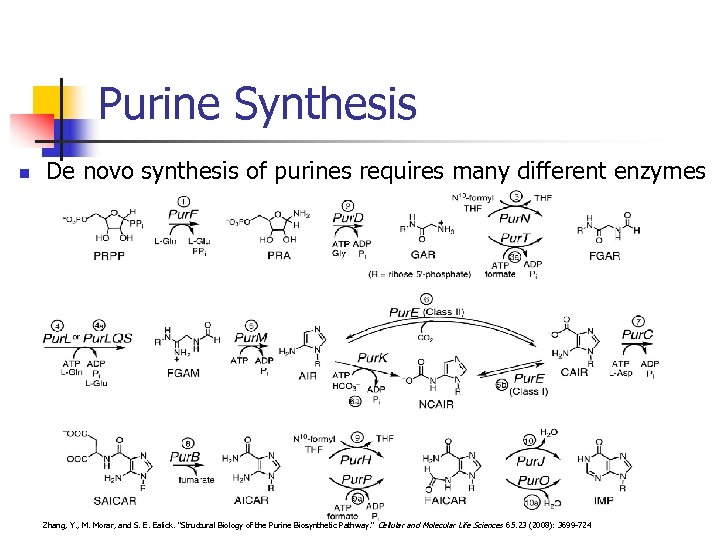 Purine Synthesis n De novo synthesis of purines requires many different enzymes Zhang, Y. , M. Morar, and S. E. Ealick. "Structural Biology of the Purine Biosynthetic Pathway. " Cellular and Molecular Life Sciences 65. 23 (2008): 3699 -724
Purine Synthesis n De novo synthesis of purines requires many different enzymes Zhang, Y. , M. Morar, and S. E. Ealick. "Structural Biology of the Purine Biosynthetic Pathway. " Cellular and Molecular Life Sciences 65. 23 (2008): 3699 -724
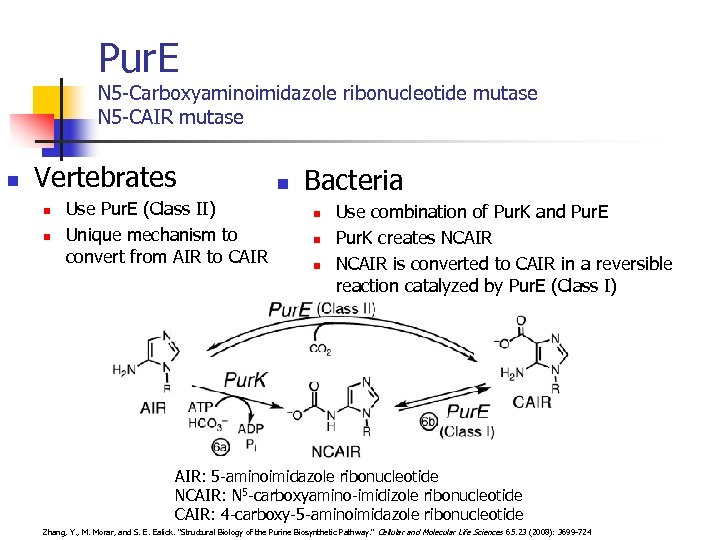 Pur. E N 5 -Carboxyaminoimidazole ribonucleotide mutase N 5 -CAIR mutase n Vertebrates n n Use Pur. E (Class II) Unique mechanism to convert from AIR to CAIR n Bacteria n n n Use combination of Pur. K and Pur. E Pur. K creates NCAIR is converted to CAIR in a reversible reaction catalyzed by Pur. E (Class I) AIR: 5 -aminoimidazole ribonucleotide NCAIR: N 5 -carboxyamino-imidizole ribonucleotide CAIR: 4 -carboxy-5 -aminoimidazole ribonucleotide Zhang, Y. , M. Morar, and S. E. Ealick. "Structural Biology of the Purine Biosynthetic Pathway. " Cellular and Molecular Life Sciences 65. 23 (2008): 3699 -724
Pur. E N 5 -Carboxyaminoimidazole ribonucleotide mutase N 5 -CAIR mutase n Vertebrates n n Use Pur. E (Class II) Unique mechanism to convert from AIR to CAIR n Bacteria n n n Use combination of Pur. K and Pur. E Pur. K creates NCAIR is converted to CAIR in a reversible reaction catalyzed by Pur. E (Class I) AIR: 5 -aminoimidazole ribonucleotide NCAIR: N 5 -carboxyamino-imidizole ribonucleotide CAIR: 4 -carboxy-5 -aminoimidazole ribonucleotide Zhang, Y. , M. Morar, and S. E. Ealick. "Structural Biology of the Purine Biosynthetic Pathway. " Cellular and Molecular Life Sciences 65. 23 (2008): 3699 -724
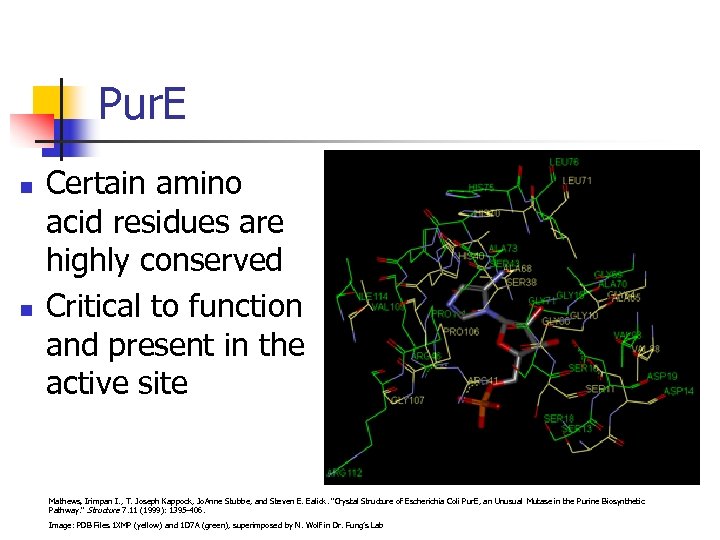 Pur. E n n Certain amino acid residues are highly conserved Critical to function and present in the active site Mathews, Irimpan I. , T. Joseph Kappock, Jo. Anne Stubbe, and Steven E. Ealick. "Crystal Structure of Escherichia Coli Pur. E, an Unusual Mutase in the Purine Biosynthetic Pathway. " Structure 7. 11 (1999): 1395 -406. Image: PDB Files 1 XMP (yellow) and 1 D 7 A (green), superimposed by N. Wolf in Dr. Fung’s Lab
Pur. E n n Certain amino acid residues are highly conserved Critical to function and present in the active site Mathews, Irimpan I. , T. Joseph Kappock, Jo. Anne Stubbe, and Steven E. Ealick. "Crystal Structure of Escherichia Coli Pur. E, an Unusual Mutase in the Purine Biosynthetic Pathway. " Structure 7. 11 (1999): 1395 -406. Image: PDB Files 1 XMP (yellow) and 1 D 7 A (green), superimposed by N. Wolf in Dr. Fung’s Lab
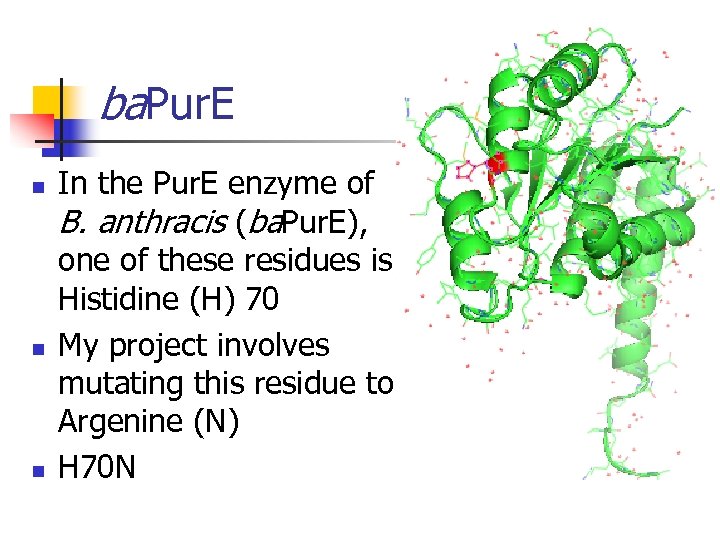 ba. Pur. E n n n In the Pur. E enzyme of B. anthracis (ba. Pur. E), one of these residues is Histidine (H) 70 My project involves mutating this residue to Argenine (N) H 70 N
ba. Pur. E n n n In the Pur. E enzyme of B. anthracis (ba. Pur. E), one of these residues is Histidine (H) 70 My project involves mutating this residue to Argenine (N) H 70 N
 Site Directed Mutagenesis n Changing an amino acid residue of interest. n n n Alter the structure of a protein Determine effect on functionality Primer: a complementary oligonucleotide (approx. 18 -27 base pairs) with a point mutation at the center such that the new codon will change the single amino acid of interest
Site Directed Mutagenesis n Changing an amino acid residue of interest. n n n Alter the structure of a protein Determine effect on functionality Primer: a complementary oligonucleotide (approx. 18 -27 base pairs) with a point mutation at the center such that the new codon will change the single amino acid of interest
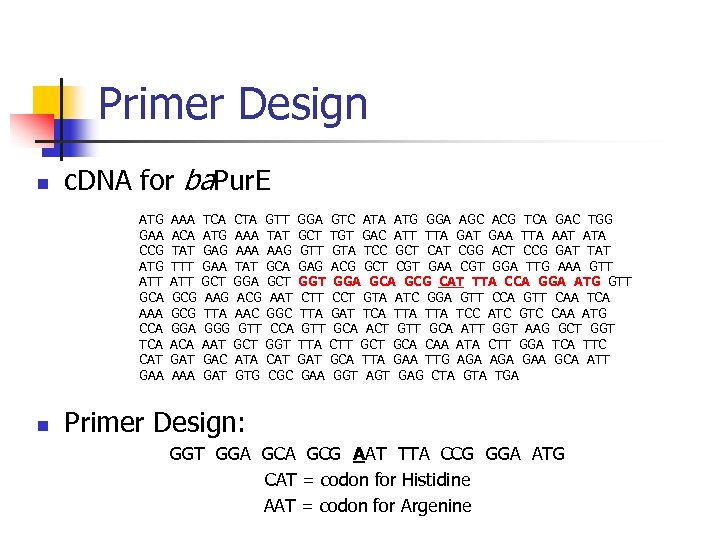 Primer Design n c. DNA for ba. Pur. E ATG GAA CCG ATT GCA AAA CCA TCA CAT GAA n AAA ACA TAT TTT ATT GCG GGA ACA GAT AAA TCA CTA GTT ATG AAA TAT GAG AAA AAG GAA TAT GCA GCT GGA GCT AAG ACG AAT TTA AAC GGG GTT CCA AAT GCT GGT GAC ATA CAT GTG CGC GGA GCT GTT GAG GGT CTT TTA GAT GAA GTC ATA ATG GGA AGC ACG TCA GAC TGG TGT GAC ATT TTA GAT GAA TTA AAT ATA GTA TCC GCT CAT CGG ACT CCG GAT TAT ACG GCT CGT GAA CGT GGA TTG AAA GTT GGA GCG CAT TTA CCA GGA ATG GTT CCT GTA ATC GGA GTT CCA GTT CAA TCA GAT TCA TTA TCC ATC GTC CAA ATG GCA ACT GTT GCA ATT GGT AAG GCT GGT CTT GCA CAA ATA CTT GGA TCA TTC GCA TTA GAA TTG AGA GAA GCA ATT GGT AGT GAG CTA GTA TGA Primer Design: GGT GGA GCG AAT TTA CCG GGA ATG CAT = codon for Histidine AAT = codon for Argenine
Primer Design n c. DNA for ba. Pur. E ATG GAA CCG ATT GCA AAA CCA TCA CAT GAA n AAA ACA TAT TTT ATT GCG GGA ACA GAT AAA TCA CTA GTT ATG AAA TAT GAG AAA AAG GAA TAT GCA GCT GGA GCT AAG ACG AAT TTA AAC GGG GTT CCA AAT GCT GGT GAC ATA CAT GTG CGC GGA GCT GTT GAG GGT CTT TTA GAT GAA GTC ATA ATG GGA AGC ACG TCA GAC TGG TGT GAC ATT TTA GAT GAA TTA AAT ATA GTA TCC GCT CAT CGG ACT CCG GAT TAT ACG GCT CGT GAA CGT GGA TTG AAA GTT GGA GCG CAT TTA CCA GGA ATG GTT CCT GTA ATC GGA GTT CCA GTT CAA TCA GAT TCA TTA TCC ATC GTC CAA ATG GCA ACT GTT GCA ATT GGT AAG GCT GGT CTT GCA CAA ATA CTT GGA TCA TTC GCA TTA GAA TTG AGA GAA GCA ATT GGT AGT GAG CTA GTA TGA Primer Design: GGT GGA GCG AAT TTA CCG GGA ATG CAT = codon for Histidine AAT = codon for Argenine
 DNA Isolation n Use DH 5α cells containing a plasmid with ba. Pur. E c. DNA
DNA Isolation n Use DH 5α cells containing a plasmid with ba. Pur. E c. DNA
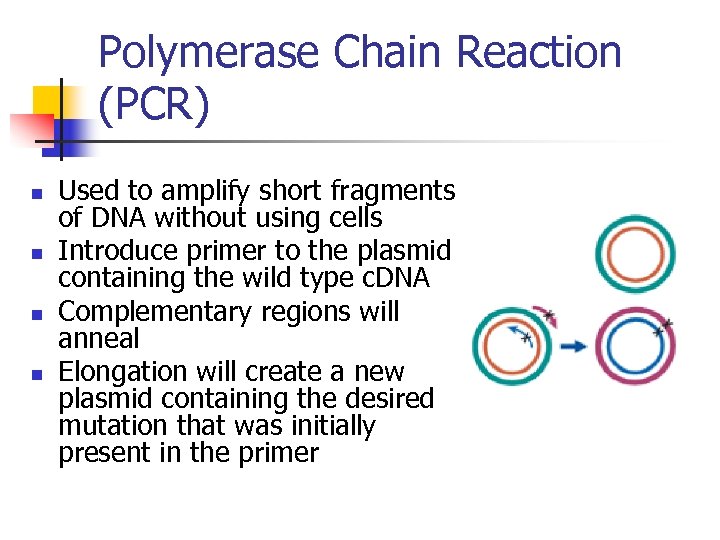 Polymerase Chain Reaction (PCR) n n Used to amplify short fragments of DNA without using cells Introduce primer to the plasmid containing the wild type c. DNA Complementary regions will anneal Elongation will create a new plasmid containing the desired mutation that was initially present in the primer
Polymerase Chain Reaction (PCR) n n Used to amplify short fragments of DNA without using cells Introduce primer to the plasmid containing the wild type c. DNA Complementary regions will anneal Elongation will create a new plasmid containing the desired mutation that was initially present in the primer
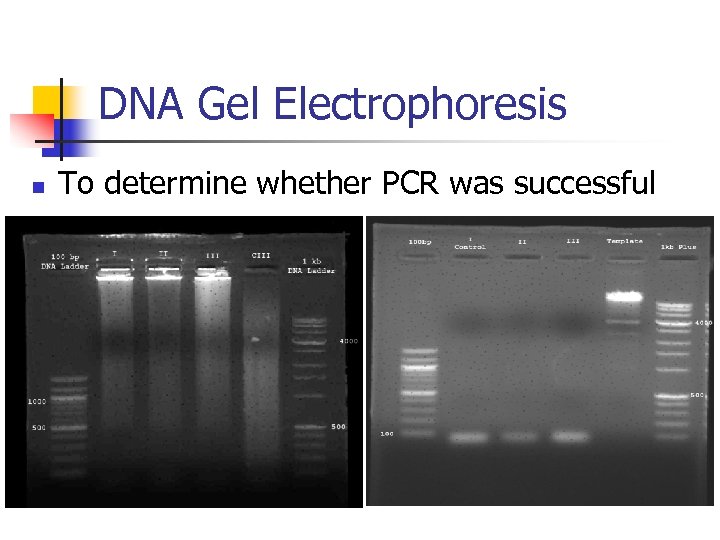 DNA Gel Electrophoresis n To determine whether PCR was successful
DNA Gel Electrophoresis n To determine whether PCR was successful
 Why is my PCR not working? n Multiple unsuccessful PCRs: n n n Varying cycling temperatures Varying concentrations for template, primers and d. NTPs Varying polymerase (“hot start” and pfu) Potential problem with the template? Re-isolate DNA from DH 5α cells
Why is my PCR not working? n Multiple unsuccessful PCRs: n n n Varying cycling temperatures Varying concentrations for template, primers and d. NTPs Varying polymerase (“hot start” and pfu) Potential problem with the template? Re-isolate DNA from DH 5α cells
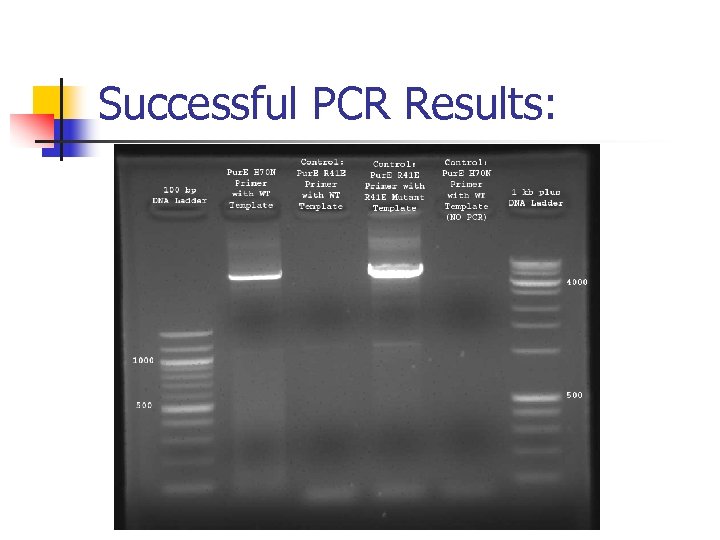 Successful PCR Results:
Successful PCR Results:
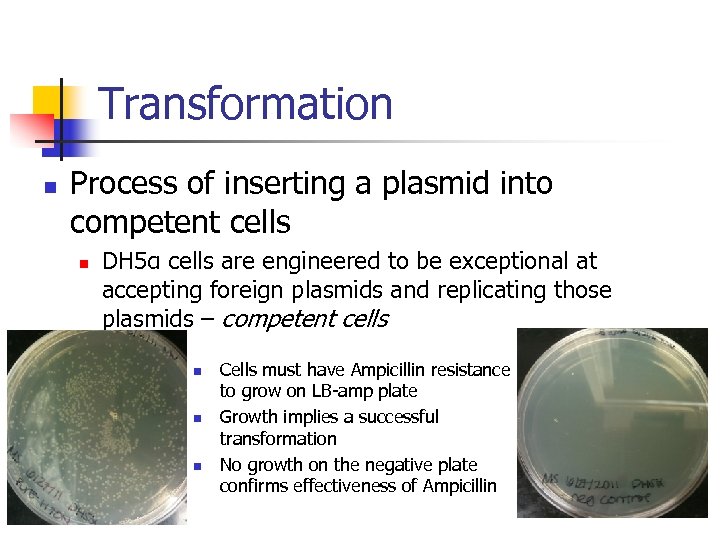 Transformation n Process of inserting a plasmid into competent cells n DH 5α cells are engineered to be exceptional at accepting foreign plasmids and replicating those plasmids – competent cells n n n Cells must have Ampicillin resistance to grow on LB-amp plate Growth implies a successful transformation No growth on the negative plate confirms effectiveness of Ampicillin
Transformation n Process of inserting a plasmid into competent cells n DH 5α cells are engineered to be exceptional at accepting foreign plasmids and replicating those plasmids – competent cells n n n Cells must have Ampicillin resistance to grow on LB-amp plate Growth implies a successful transformation No growth on the negative plate confirms effectiveness of Ampicillin
 DNA Sequencing n n Grew cultures of four distinct colonies in four different 4 m. L LB+amp liquid media Extracted DNA from all colonies and sent for sequencing at the Research Resources Center facilities available at UIC
DNA Sequencing n n Grew cultures of four distinct colonies in four different 4 m. L LB+amp liquid media Extracted DNA from all colonies and sent for sequencing at the Research Resources Center facilities available at UIC
 DNA Sequencing Results Colony 1, 2, & 4 1 2 Analyzed results using: http: //www. ebi. ac. uk/Tools/psa/emboss_needle/nucleotide. html a feature available through the European Bioinformatics Institute 4
DNA Sequencing Results Colony 1, 2, & 4 1 2 Analyzed results using: http: //www. ebi. ac. uk/Tools/psa/emboss_needle/nucleotide. html a feature available through the European Bioinformatics Institute 4
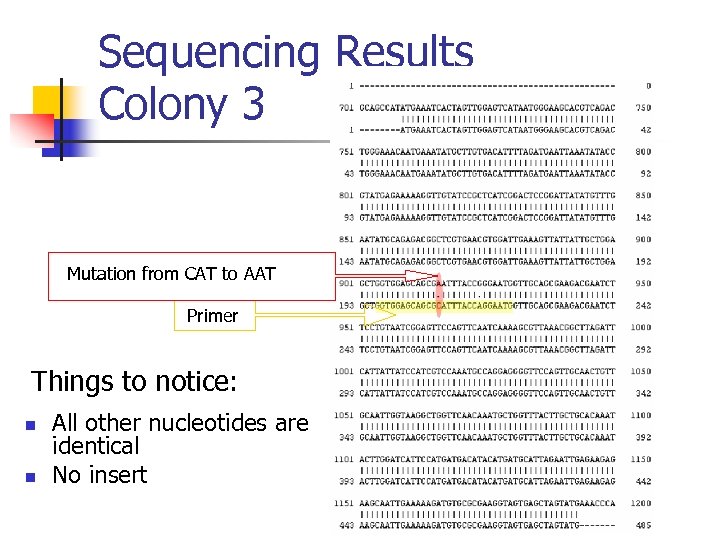 Sequencing Results Colony 3 Mutation from CAT to AAT Primer Things to notice: n n All other nucleotides are identical No insert
Sequencing Results Colony 3 Mutation from CAT to AAT Primer Things to notice: n n All other nucleotides are identical No insert
 Protein Purification n Use DNA from: n Colony 3 cells to create H 70 N protein n Protein will have an identical amino acid sequence as the wild type Pur. E, except for Arginine at position 70 Determine subsequent change in functionality Colony 1 cells to create a truncated protein n n The insert contains a stop codon Protein only has 98 amino acids instead of 161 Removing part of the active site Predict no functionality
Protein Purification n Use DNA from: n Colony 3 cells to create H 70 N protein n Protein will have an identical amino acid sequence as the wild type Pur. E, except for Arginine at position 70 Determine subsequent change in functionality Colony 1 cells to create a truncated protein n n The insert contains a stop codon Protein only has 98 amino acids instead of 161 Removing part of the active site Predict no functionality
 Creating H 70 N & Truncated Pur. E n Transform DNA into BL 21 cells n n n Grow BL 21 cells in two flasks of 2 L LB+amp liquid media Induce cells with Isopropyl β-D-1 thiogalactopyranoside (IPTG) n n BL 21 Cells are a mutated form of E. coli that over produce proteins Increases protein production and is not metabolized by cells Freeze cells overnight in -80ºC " Isopropyl β-D-1 -thiogalactopyranoside. " Wikipedia, the Free Encyclopedia. Web. 26 July 2011. http: //en. wikipedia. org/wiki/Isopropyl_%CE%B 2 -D-1 -thiogalactopyranoside.
Creating H 70 N & Truncated Pur. E n Transform DNA into BL 21 cells n n n Grow BL 21 cells in two flasks of 2 L LB+amp liquid media Induce cells with Isopropyl β-D-1 thiogalactopyranoside (IPTG) n n BL 21 Cells are a mutated form of E. coli that over produce proteins Increases protein production and is not metabolized by cells Freeze cells overnight in -80ºC " Isopropyl β-D-1 -thiogalactopyranoside. " Wikipedia, the Free Encyclopedia. Web. 26 July 2011. http: //en. wikipedia. org/wiki/Isopropyl_%CE%B 2 -D-1 -thiogalactopyranoside.
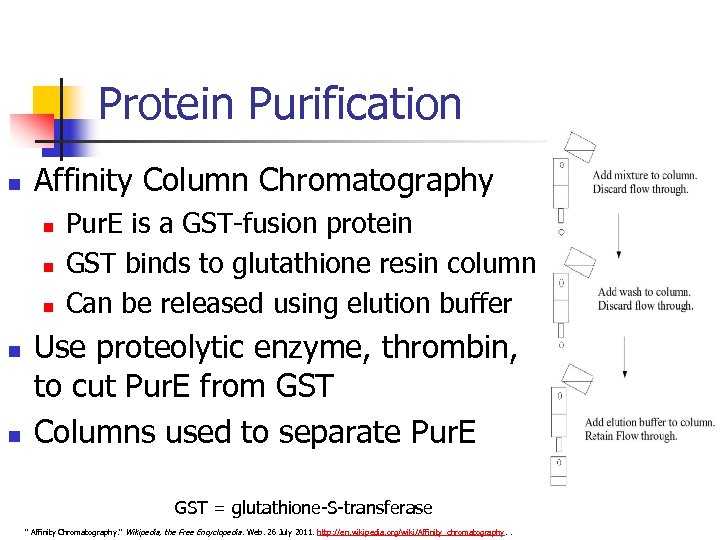 Protein Purification n Affinity Column Chromatography n n n Pur. E is a GST-fusion protein GST binds to glutathione resin column Can be released using elution buffer Use proteolytic enzyme, thrombin, to cut Pur. E from GST Columns used to separate Pur. E GST = glutathione-S-transferase " Affinity Chromatography. " Wikipedia, the Free Encyclopedia. Web. 26 July 2011. http: //en. wikipedia. org/wiki/Affinity_chromatography. .
Protein Purification n Affinity Column Chromatography n n n Pur. E is a GST-fusion protein GST binds to glutathione resin column Can be released using elution buffer Use proteolytic enzyme, thrombin, to cut Pur. E from GST Columns used to separate Pur. E GST = glutathione-S-transferase " Affinity Chromatography. " Wikipedia, the Free Encyclopedia. Web. 26 July 2011. http: //en. wikipedia. org/wiki/Affinity_chromatography. .
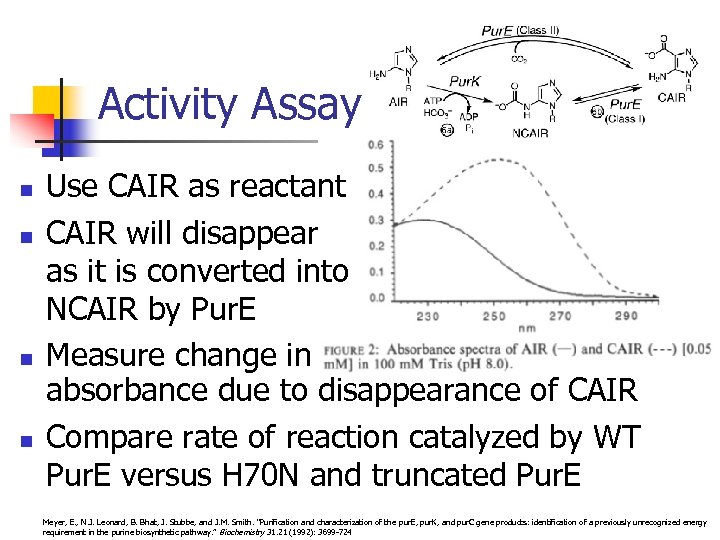 Activity Assay n n Use CAIR as reactant CAIR will disappear as it is converted into NCAIR by Pur. E Measure change in absorbance due to disappearance of CAIR Compare rate of reaction catalyzed by WT Pur. E versus H 70 N and truncated Pur. E Meyer, E. , N. J. Leonard, B. Bhat, J. Stubbe, and J. M. Smith. "Purification and characterization of the pur. E, pur. K, and pur. C gene products: identification of a previously unrecognized energy requirement in the purine biosynthetic pathway. ” Biochemistry 31. 21 (1992): 3699 -724
Activity Assay n n Use CAIR as reactant CAIR will disappear as it is converted into NCAIR by Pur. E Measure change in absorbance due to disappearance of CAIR Compare rate of reaction catalyzed by WT Pur. E versus H 70 N and truncated Pur. E Meyer, E. , N. J. Leonard, B. Bhat, J. Stubbe, and J. M. Smith. "Purification and characterization of the pur. E, pur. K, and pur. C gene products: identification of a previously unrecognized energy requirement in the purine biosynthetic pathway. ” Biochemistry 31. 21 (1992): 3699 -724
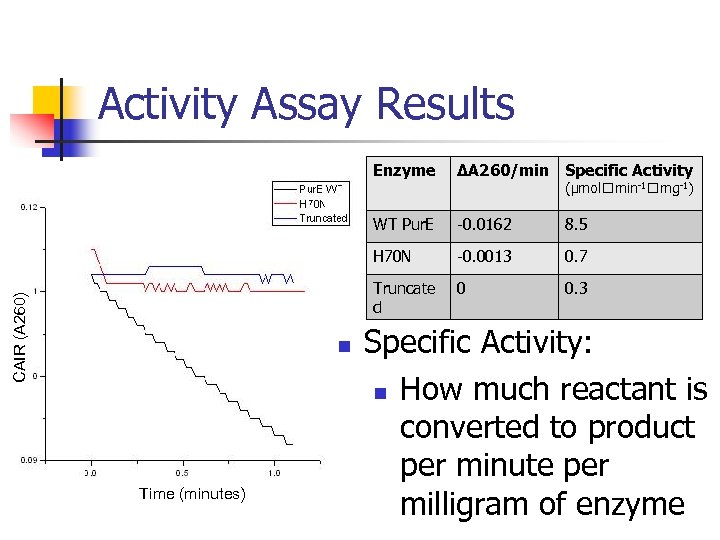 Activity Assay Results 8. 5 -0. 0013 0. 7 Truncate d Time (minutes) -0. 0162 H 70 N n ΔA 260/min Specific Activity WT Pur. E CAIR (A 260) Enzyme 0 0. 3 (μmol min-1 mg-1) Specific Activity: n How much reactant is converted to product per minute per milligram of enzyme
Activity Assay Results 8. 5 -0. 0013 0. 7 Truncate d Time (minutes) -0. 0162 H 70 N n ΔA 260/min Specific Activity WT Pur. E CAIR (A 260) Enzyme 0 0. 3 (μmol min-1 mg-1) Specific Activity: n How much reactant is converted to product per minute per milligram of enzyme
 Conclusion n Accomplishments n n n Designed an ideal primer for the H 70 N mutation Used PCR to obtain recombinant DNA with H 70 N mutation Created DNA coding for a truncated Pur. E enzyme Transformed the DNA into BL 21 cells Prepared H 70 N and truncated proteins Determined enzymatic activity of these proteins
Conclusion n Accomplishments n n n Designed an ideal primer for the H 70 N mutation Used PCR to obtain recombinant DNA with H 70 N mutation Created DNA coding for a truncated Pur. E enzyme Transformed the DNA into BL 21 cells Prepared H 70 N and truncated proteins Determined enzymatic activity of these proteins
 Acknowledgements n The financial support from the National Science Foundation EEC-NSF Grant # 1062943 n n Dr. Fung, Nina Wolf, and Esther Ng REU Program Facilitators: Dr. Takoudis, Dr. Jursich, and Arman Butt
Acknowledgements n The financial support from the National Science Foundation EEC-NSF Grant # 1062943 n n Dr. Fung, Nina Wolf, and Esther Ng REU Program Facilitators: Dr. Takoudis, Dr. Jursich, and Arman Butt


