c117fc7dfdf16b694260d436f809d747.ppt
- Количество слайдов: 30
 Shoshana Segal shosh@nih. gov, 301 -496 -0923 David Goldstein goldsted@ncifcrf. gov, 301 -496 -4347
Shoshana Segal shosh@nih. gov, 301 -496 -0923 David Goldstein goldsted@ncifcrf. gov, 301 -496 -4347
 CCR Technologies and Scientific Resources • • Protein-Protein Interaction Screens - Contract Antibody Production Program - Collaborative MTAs Antibody Assay Development - in-house program Multiplex Cytokine Analysis - 4 BPAs Microarray Technology - OD Subsidy Genomics/Proteomics Software Toolbox - Licenses Chemical Genomics Software - License Advanced Technologies - Research Technology Program
CCR Technologies and Scientific Resources • • Protein-Protein Interaction Screens - Contract Antibody Production Program - Collaborative MTAs Antibody Assay Development - in-house program Multiplex Cytokine Analysis - 4 BPAs Microarray Technology - OD Subsidy Genomics/Proteomics Software Toolbox - Licenses Chemical Genomics Software - License Advanced Technologies - Research Technology Program
 Myriad’s Pro. Net Yeast Two-Hybrid Platform A Program for the Discovery of Novel Protein-Protein Interactions and Networks http: //ccr. cancer. gov/research/ostp/myriad_genetics_survey_new. asp
Myriad’s Pro. Net Yeast Two-Hybrid Platform A Program for the Discovery of Novel Protein-Protein Interactions and Networks http: //ccr. cancer. gov/research/ostp/myriad_genetics_survey_new. asp
 Pro. Net Production Numbers During seven years of using the Pro. Net yeast two-hybrid platform, Myriad has accomplished the following: n >4500 proteins have been analyzed from a number of organisms n >22, 000 bait constructs have been constructed and searched n 40 different activation domain libraries have been constructed n >60, 000 yeast two-hybrid searches have been performed n >300, 000 yeast search colonies have been analyzed n >10, 000 protein-protein interactions have been identified n Extensive interaction networks in the areas of APP metabolism, glucose uptake, HIV budding, cholesterol efflux, B-cell signaling
Pro. Net Production Numbers During seven years of using the Pro. Net yeast two-hybrid platform, Myriad has accomplished the following: n >4500 proteins have been analyzed from a number of organisms n >22, 000 bait constructs have been constructed and searched n 40 different activation domain libraries have been constructed n >60, 000 yeast two-hybrid searches have been performed n >300, 000 yeast search colonies have been analyzed n >10, 000 protein-protein interactions have been identified n Extensive interaction networks in the areas of APP metabolism, glucose uptake, HIV budding, cholesterol efflux, B-cell signaling
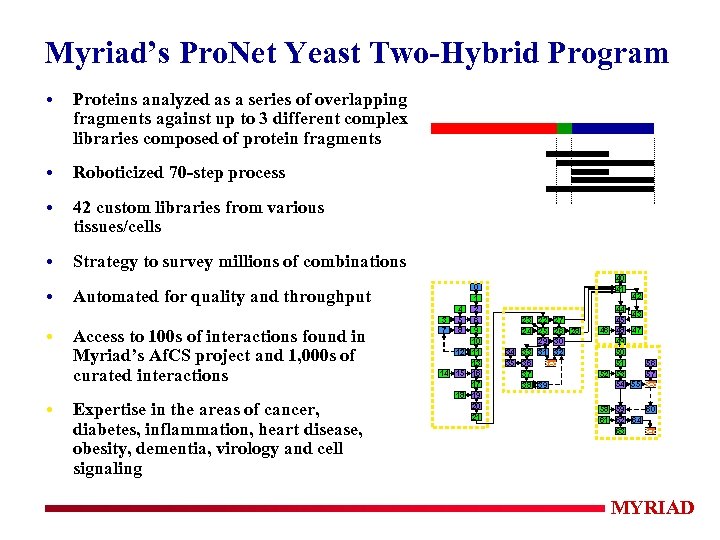 Myriad’s Pro. Net Yeast Two-Hybrid Program • Proteins analyzed as a series of overlapping fragments against up to 3 different complex libraries composed of protein fragments • Roboticized 70 -step process • 42 custom libraries from various tissues/cells • Strategy to survey millions of combinations • Automated for quality and throughput • Access to 100 s of interactions found in Myriad’s Af. CS project and 1, 000 s of curated interactions • Expertise in the areas of cancer, diabetes, inflammation, heart disease, obesity, dementia, virology and cell signaling 0 1 4 2 3 5 6 7 8 9 10 12 11 13 14 15 16 17 18 19 20 21 40 41 23 22 27 24 25 26 28 29 30 34 33 31 32 35 36 Seq 37 38 39 42 44 43 45 46 48 47 49 50 51 56 52 53 57 54 55 Seq 60 58 59 61 62 64 63 Seq MYRIAD
Myriad’s Pro. Net Yeast Two-Hybrid Program • Proteins analyzed as a series of overlapping fragments against up to 3 different complex libraries composed of protein fragments • Roboticized 70 -step process • 42 custom libraries from various tissues/cells • Strategy to survey millions of combinations • Automated for quality and throughput • Access to 100 s of interactions found in Myriad’s Af. CS project and 1, 000 s of curated interactions • Expertise in the areas of cancer, diabetes, inflammation, heart disease, obesity, dementia, virology and cell signaling 0 1 4 2 3 5 6 7 8 9 10 12 11 13 14 15 16 17 18 19 20 21 40 41 23 22 27 24 25 26 28 29 30 34 33 31 32 35 36 Seq 37 38 39 42 44 43 45 46 48 47 49 50 51 56 52 53 57 54 55 Seq 60 58 59 61 62 64 63 Seq MYRIAD
 Pro. Net Reduction of False Positives n Multiple reporter genes (3) with dissimilar promoters n Low copy number plasmids n Electronic removal of notorious false positives n Wet false positive test with heterologous protein baits
Pro. Net Reduction of False Positives n Multiple reporter genes (3) with dissimilar promoters n Low copy number plasmids n Electronic removal of notorious false positives n Wet false positive test with heterologous protein baits
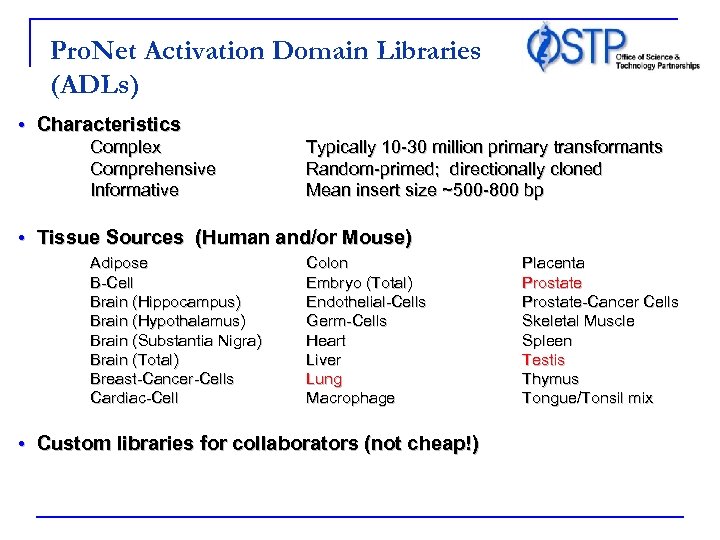 Pro. Net Activation Domain Libraries (ADLs) • Characteristics Complex Comprehensive Informative Typically 10 -30 million primary transformants Random-primed; directionally cloned Mean insert size ~500 -800 bp • Tissue Sources (Human and/or Mouse) Adipose B-Cell Brain (Hippocampus) Brain (Hypothalamus) Brain (Substantia Nigra) Brain (Total) Breast-Cancer-Cells Cardiac-Cell Colon Embryo (Total) Endothelial-Cells Germ-Cells Heart Liver Lung Macrophage • Custom libraries for collaborators (not cheap!) Placenta Prostate-Cancer Cells Skeletal Muscle Spleen Testis Thymus Tongue/Tonsil mix
Pro. Net Activation Domain Libraries (ADLs) • Characteristics Complex Comprehensive Informative Typically 10 -30 million primary transformants Random-primed; directionally cloned Mean insert size ~500 -800 bp • Tissue Sources (Human and/or Mouse) Adipose B-Cell Brain (Hippocampus) Brain (Hypothalamus) Brain (Substantia Nigra) Brain (Total) Breast-Cancer-Cells Cardiac-Cell Colon Embryo (Total) Endothelial-Cells Germ-Cells Heart Liver Lung Macrophage • Custom libraries for collaborators (not cheap!) Placenta Prostate-Cancer Cells Skeletal Muscle Spleen Testis Thymus Tongue/Tonsil mix
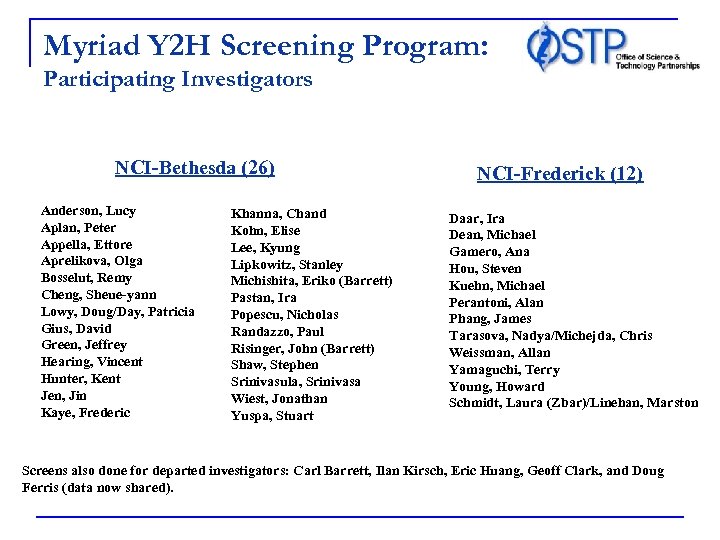 Myriad Y 2 H Screening Program: Participating Investigators NCI-Bethesda (26) Anderson, Lucy Aplan, Peter Appella, Ettore Aprelikova, Olga Bosselut, Remy Cheng, Sheue-yann Lowy, Doug/Day, Patricia Gius, David Green, Jeffrey Hearing, Vincent Hunter, Kent Jen, Jin Kaye, Frederic Khanna, Chand Kohn, Elise Lee, Kyung Lipkowitz, Stanley Michishita, Eriko (Barrett) Pastan, Ira Popescu, Nicholas Randazzo, Paul Risinger, John (Barrett) Shaw, Stephen Srinivasula, Srinivasa Wiest, Jonathan Yuspa, Stuart NCI-Frederick (12) Daar, Ira Dean, Michael Gamero, Ana Hou, Steven Kuehn, Michael Perantoni, Alan Phang, James Tarasova, Nadya/Michejda, Chris Weissman, Allan Yamaguchi, Terry Young, Howard Schmidt, Laura (Zbar)/Linehan, Marston Screens also done for departed investigators: Carl Barrett, Ilan Kirsch, Eric Huang, Geoff Clark, and Doug Ferris (data now shared).
Myriad Y 2 H Screening Program: Participating Investigators NCI-Bethesda (26) Anderson, Lucy Aplan, Peter Appella, Ettore Aprelikova, Olga Bosselut, Remy Cheng, Sheue-yann Lowy, Doug/Day, Patricia Gius, David Green, Jeffrey Hearing, Vincent Hunter, Kent Jen, Jin Kaye, Frederic Khanna, Chand Kohn, Elise Lee, Kyung Lipkowitz, Stanley Michishita, Eriko (Barrett) Pastan, Ira Popescu, Nicholas Randazzo, Paul Risinger, John (Barrett) Shaw, Stephen Srinivasula, Srinivasa Wiest, Jonathan Yuspa, Stuart NCI-Frederick (12) Daar, Ira Dean, Michael Gamero, Ana Hou, Steven Kuehn, Michael Perantoni, Alan Phang, James Tarasova, Nadya/Michejda, Chris Weissman, Allan Yamaguchi, Terry Young, Howard Schmidt, Laura (Zbar)/Linehan, Marston Screens also done for departed investigators: Carl Barrett, Ilan Kirsch, Eric Huang, Geoff Clark, and Doug Ferris (data now shared).
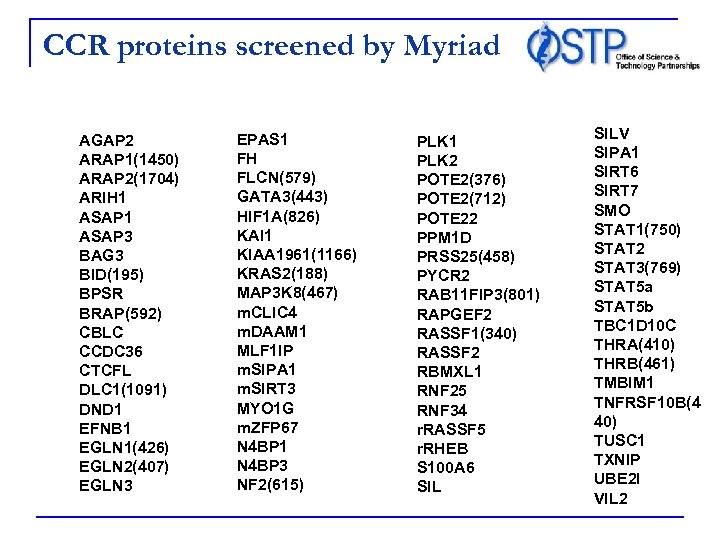 CCR proteins screened by Myriad AGAP 2 ARAP 1(1450) ARAP 2(1704) ARIH 1 ASAP 3 BAG 3 BID(195) BPSR BRAP(592) CBLC CCDC 36 CTCFL DLC 1(1091) DND 1 EFNB 1 EGLN 1(426) EGLN 2(407) EGLN 3 EPAS 1 FH FLCN(579) GATA 3(443) HIF 1 A(826) KAI 1 KIAA 1961(1166) KRAS 2(188) MAP 3 K 8(467) m. CLIC 4 m. DAAM 1 MLF 1 IP m. SIPA 1 m. SIRT 3 MYO 1 G m. ZFP 67 N 4 BP 1 N 4 BP 3 NF 2(615) PLK 1 PLK 2 POTE 2(376) POTE 2(712) POTE 22 PPM 1 D PRSS 25(458) PYCR 2 RAB 11 FIP 3(801) RAPGEF 2 RASSF 1(340) RASSF 2 RBMXL 1 RNF 25 RNF 34 r. RASSF 5 r. RHEB S 100 A 6 SILV SIPA 1 SIRT 6 SIRT 7 SMO STAT 1(750) STAT 2 STAT 3(769) STAT 5 a STAT 5 b TBC 1 D 10 C THRA(410) THRB(461) TMBIM 1 TNFRSF 10 B(4 40) TUSC 1 TXNIP UBE 2 I VIL 2
CCR proteins screened by Myriad AGAP 2 ARAP 1(1450) ARAP 2(1704) ARIH 1 ASAP 3 BAG 3 BID(195) BPSR BRAP(592) CBLC CCDC 36 CTCFL DLC 1(1091) DND 1 EFNB 1 EGLN 1(426) EGLN 2(407) EGLN 3 EPAS 1 FH FLCN(579) GATA 3(443) HIF 1 A(826) KAI 1 KIAA 1961(1166) KRAS 2(188) MAP 3 K 8(467) m. CLIC 4 m. DAAM 1 MLF 1 IP m. SIPA 1 m. SIRT 3 MYO 1 G m. ZFP 67 N 4 BP 1 N 4 BP 3 NF 2(615) PLK 1 PLK 2 POTE 2(376) POTE 2(712) POTE 22 PPM 1 D PRSS 25(458) PYCR 2 RAB 11 FIP 3(801) RAPGEF 2 RASSF 1(340) RASSF 2 RBMXL 1 RNF 25 RNF 34 r. RASSF 5 r. RHEB S 100 A 6 SILV SIPA 1 SIRT 6 SIRT 7 SMO STAT 1(750) STAT 2 STAT 3(769) STAT 5 a STAT 5 b TBC 1 D 10 C THRA(410) THRB(461) TMBIM 1 TNFRSF 10 B(4 40) TUSC 1 TXNIP UBE 2 I VIL 2
 Pro. Net Informatics n Secure, browser-based interface n Weekly updates n 70, 000 Protein Home Pages n Graphical tools n Project management tables n BLAST function to identify similar proteins or genes in Pro. Net n Curation of public two-hybrid literature
Pro. Net Informatics n Secure, browser-based interface n Weekly updates n 70, 000 Protein Home Pages n Graphical tools n Project management tables n BLAST function to identify similar proteins or genes in Pro. Net n Curation of public two-hybrid literature
 CCR Protein Interaction Data Exchange (C-Pr. IDE) Short-Term Goals n Share Y 2 H interaction data across CCR: Increase opportunity for collaboration and exchange of expertise n Upfront adoption of ca. BIG tools and guidelines: Leverage NCICB and ca. BIG Infrastructure Components n Harness potential of the ABCC computational capabilities and infrastructure n Data-driven/Dynamic architecture (i. e. , self building): Investigators will deposit data confirming interactions in mammalian systems n Take advantage of existing resources
CCR Protein Interaction Data Exchange (C-Pr. IDE) Short-Term Goals n Share Y 2 H interaction data across CCR: Increase opportunity for collaboration and exchange of expertise n Upfront adoption of ca. BIG tools and guidelines: Leverage NCICB and ca. BIG Infrastructure Components n Harness potential of the ABCC computational capabilities and infrastructure n Data-driven/Dynamic architecture (i. e. , self building): Investigators will deposit data confirming interactions in mammalian systems n Take advantage of existing resources
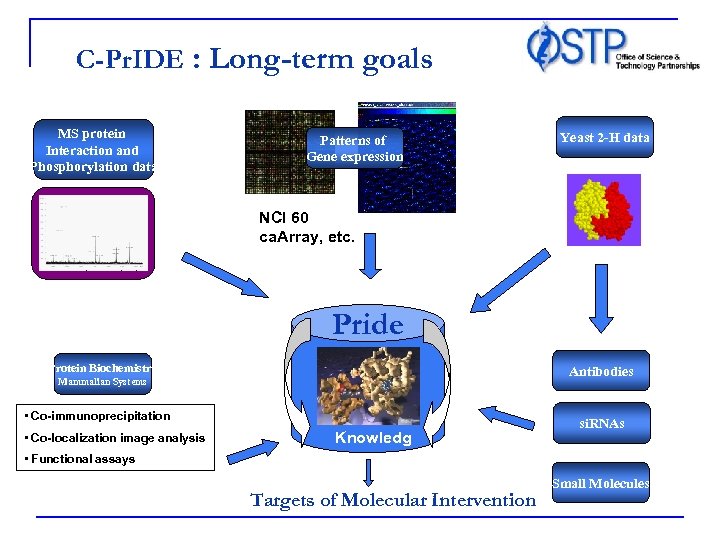 C-Pr. IDE : Long-term goals MS protein Interaction and Phosphorylation data Patterns of Gene expression Yeast 2 -H data NCI 60 ca. Array, etc. Pride Protein Biochemistry Antibodies Mammalian Systems • Co-immunoprecipitation • Co-localization image analysis • Functional assays Knowledg e Targets of Molecular Intervention si. RNAs Small Molecules
C-Pr. IDE : Long-term goals MS protein Interaction and Phosphorylation data Patterns of Gene expression Yeast 2 -H data NCI 60 ca. Array, etc. Pride Protein Biochemistry Antibodies Mammalian Systems • Co-immunoprecipitation • Co-localization image analysis • Functional assays Knowledg e Targets of Molecular Intervention si. RNAs Small Molecules
 CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals CCR has established a unique partnership with BD Pharmingen and Rockland Immunochemicals to produce mouse monoclonal and rabbit polyclonal antibodies against key cancer targets of interest to CCR - at no cost to individual investigators
CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals CCR has established a unique partnership with BD Pharmingen and Rockland Immunochemicals to produce mouse monoclonal and rabbit polyclonal antibodies against key cancer targets of interest to CCR - at no cost to individual investigators
 CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals • CCR investigators submit requests for antibodies to be produced • Peptides or recombinant proteins are synthesized and used as immunogens • BD and Rockland produce the sera, purify the antibodies, and perform preliminary analyses • Investigators receive sera and later purified antibodies for testing and validation in their cell systems • BD and Rockland commercialize the antibodies • Investigators receive 3 mg of purified antibody every year until the partnership is terminated - AT NO COST TO INVESTIGATOR • Investigators receive a discount of 30% when they purchase the antibody
CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals • CCR investigators submit requests for antibodies to be produced • Peptides or recombinant proteins are synthesized and used as immunogens • BD and Rockland produce the sera, purify the antibodies, and perform preliminary analyses • Investigators receive sera and later purified antibodies for testing and validation in their cell systems • BD and Rockland commercialize the antibodies • Investigators receive 3 mg of purified antibody every year until the partnership is terminated - AT NO COST TO INVESTIGATOR • Investigators receive a discount of 30% when they purchase the antibody
 CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals 21 mouse monoclonal antibodies delivered to investigators 50 mouse monoclonal antibodies in production 24 rabbit polyclonal antibodies delivered to investigators 100 rabbit polyclonal antibodies in production
CCR partnerships with Beckon Dickinson. Pharmingen and Rockland Immunochemicals 21 mouse monoclonal antibodies delivered to investigators 50 mouse monoclonal antibodies in production 24 rabbit polyclonal antibodies delivered to investigators 100 rabbit polyclonal antibodies in production
 Antibody Production Program Project Tracking System • Background on antigen • Peptide sequence or recombinant protein used • Project status • Links to validation data • Discussion thread for comments and figure legends • Submission of new targets
Antibody Production Program Project Tracking System • Background on antigen • Peptide sequence or recombinant protein used • Project status • Links to validation data • Discussion thread for comments and figure legends • Submission of new targets
 Antibody and Protein Purification Unit Meso Scale Discovery (MSD) Assay Development Electrochemiluminescence detection Advantages: ● Large dynamic range ● Low background signal ● Assay can be performed in as little as 2 hours As little as 750 fg protein detected ● Stable to large range of conditions: p. H, light, time, temperature ● Flexibility: wide variety of plates available (uncoated, streptavidin, avidin anti-mouse, anti-rabbit, glutathione, nickel) ● Minimal matrix effects ● Multiplex ability: Up to 10 different assays per well. Applications: ● Antibodies, peptides, proteins, cells, or membranes can be directly immobilized on carbon electrode. ● Assay cytokines, biomarkers, phosphoproteins, etc. ● High-throughput Western Blot replacement. Paul Goldsmith (paulg@nih. gov) or Michele Gunsior (gunsiorm@nih. gov)
Antibody and Protein Purification Unit Meso Scale Discovery (MSD) Assay Development Electrochemiluminescence detection Advantages: ● Large dynamic range ● Low background signal ● Assay can be performed in as little as 2 hours As little as 750 fg protein detected ● Stable to large range of conditions: p. H, light, time, temperature ● Flexibility: wide variety of plates available (uncoated, streptavidin, avidin anti-mouse, anti-rabbit, glutathione, nickel) ● Minimal matrix effects ● Multiplex ability: Up to 10 different assays per well. Applications: ● Antibodies, peptides, proteins, cells, or membranes can be directly immobilized on carbon electrode. ● Assay cytokines, biomarkers, phosphoproteins, etc. ● High-throughput Western Blot replacement. Paul Goldsmith (paulg@nih. gov) or Michele Gunsior (gunsiorm@nih. gov)
 Blanket Purchase Agreements for Multiplex Cytokine Analysis Offers the NIH community a range of platforms and a wide variety of cytokines and chemokines that can be selected for analysis 1. Linco Diagnostics (CLIA certified) BPA 00063621 - Platform: Luminex. Website: http: //www. lincodiagnostics. com 2. Pathway Diagnostics (CLIA certified) BPA 00063530 - Platforms: Mesoscale Discovery, Luminex, Pierce Searchlight, Website: http: //www. pathwaydx. com 3. Pierce Diagnostics BPA 00049380 - Platform: Searchlight, Website: http: //www. endogen. com 4. Whatman BPA 00063645 - Platform: Microspot Elisa (FAST Quant) Website: http: //www. arraying. com/Services/NIHBPAService. html User Name: nihbpa Password: whatman Cost for these assays is the responsibility of the individual labs requesting the testing. For questions contact Howard Young at youngh@ncifcrf. gov or 301846 -5700.
Blanket Purchase Agreements for Multiplex Cytokine Analysis Offers the NIH community a range of platforms and a wide variety of cytokines and chemokines that can be selected for analysis 1. Linco Diagnostics (CLIA certified) BPA 00063621 - Platform: Luminex. Website: http: //www. lincodiagnostics. com 2. Pathway Diagnostics (CLIA certified) BPA 00063530 - Platforms: Mesoscale Discovery, Luminex, Pierce Searchlight, Website: http: //www. pathwaydx. com 3. Pierce Diagnostics BPA 00049380 - Platform: Searchlight, Website: http: //www. endogen. com 4. Whatman BPA 00063645 - Platform: Microspot Elisa (FAST Quant) Website: http: //www. arraying. com/Services/NIHBPAService. html User Name: nihbpa Password: whatman Cost for these assays is the responsibility of the individual labs requesting the testing. For questions contact Howard Young at youngh@ncifcrf. gov or 301846 -5700.
 Microarray Technology Subsidy Program n n Supported platforms: q Affymetrix and LMT processing service q Agilent q Illumina q Nimble. Gen - arrays and processing service q Exon. Hit - arrays and processing service Subsidy available: q $100/microarray up to $15, 000/investigator q Up to 50% or more support for special projects (Nimble. Gen, Gen. Us, Exon. Hit, Affy HTA). Must submit brief project proposal. https: //bluethunder. nci. nih. gov/Affymetrix/ (Affy, Agilent, Illumina. etc. ) https: //bluethunder. nci. nih. gov: 443/Nimble. Gen/Order. Form. html (Nimble. Gen)
Microarray Technology Subsidy Program n n Supported platforms: q Affymetrix and LMT processing service q Agilent q Illumina q Nimble. Gen - arrays and processing service q Exon. Hit - arrays and processing service Subsidy available: q $100/microarray up to $15, 000/investigator q Up to 50% or more support for special projects (Nimble. Gen, Gen. Us, Exon. Hit, Affy HTA). Must submit brief project proposal. https: //bluethunder. nci. nih. gov/Affymetrix/ (Affy, Agilent, Illumina. etc. ) https: //bluethunder. nci. nih. gov: 443/Nimble. Gen/Order. Form. html (Nimble. Gen)
 Nimble. Gen Systems, Inc. For High Definition Genomics Identify genomic insertions and deletions and copy number changes at ultra-high resolution, genome wide or focused fine tiling, with high-definition CGH. Map regulatory protein binding sites genome wide for transcription factors, polymerases and histones, or target promoter regions with Ch. IP-chip analysis. Survey microbial genomes and identify and categorize all SNPs in a single experiment with Comparative Genome Sequencing. Determine differentially expressed genes and map transcription activity with prokaryotic or eukaryotic whole-genome expression analysis.
Nimble. Gen Systems, Inc. For High Definition Genomics Identify genomic insertions and deletions and copy number changes at ultra-high resolution, genome wide or focused fine tiling, with high-definition CGH. Map regulatory protein binding sites genome wide for transcription factors, polymerases and histones, or target promoter regions with Ch. IP-chip analysis. Survey microbial genomes and identify and categorize all SNPs in a single experiment with Comparative Genome Sequencing. Determine differentially expressed genes and map transcription activity with prokaryotic or eukaryotic whole-genome expression analysis.
 CCR/OSTP/ISCS Bioinformatics Toolbox n n Genomatix Suite: http: //ccr. cancer. gov/research/ostp/new_pathways_analysis. asp#C q El. Dorado and Gene 2 Promoter: Extended genome annotation/promoter mapping q Biblio. Sphere: Literature mining and pathway mapping q Chip. Inspector: Microarray analysis Ingenuity Pathway Analysis (IPA): http: //ccr. cancer. gov/research/ostp/new_pathways_analysis. asp#B n n n Ariadne Pathway. Expert and Pathway. Studio Partek - Statistical analysis and visualization Gene. Spring customized license that integrates: q Microarray analysis q array. CGH analytics q Ch. IP analytics q Ingenuity IPA
CCR/OSTP/ISCS Bioinformatics Toolbox n n Genomatix Suite: http: //ccr. cancer. gov/research/ostp/new_pathways_analysis. asp#C q El. Dorado and Gene 2 Promoter: Extended genome annotation/promoter mapping q Biblio. Sphere: Literature mining and pathway mapping q Chip. Inspector: Microarray analysis Ingenuity Pathway Analysis (IPA): http: //ccr. cancer. gov/research/ostp/new_pathways_analysis. asp#B n n n Ariadne Pathway. Expert and Pathway. Studio Partek - Statistical analysis and visualization Gene. Spring customized license that integrates: q Microarray analysis q array. CGH analytics q Ch. IP analytics q Ingenuity IPA
 i. Research Library (NIH-wide License Agreement) Chem. Navigator's up-to-date compilation of commercially accessible small molecular compounds for drug discovery screening from international chemistry suppliers. The database currently tracks over 26. 3 million chemical samples. Database licenses include access to regular updates, sourcing information, and Chem. Navigator's optional Chemistry Procurement Service. http: //ccr. cancer. gov/research/ostp/Chem. Navigator. asp Contact: Marc Nicklaus, Ph. D. , Computer-Aided Drug Design (CADD) Group mn 1@helix. nih. gov, 301 -846 -5903
i. Research Library (NIH-wide License Agreement) Chem. Navigator's up-to-date compilation of commercially accessible small molecular compounds for drug discovery screening from international chemistry suppliers. The database currently tracks over 26. 3 million chemical samples. Database licenses include access to regular updates, sourcing information, and Chem. Navigator's optional Chemistry Procurement Service. http: //ccr. cancer. gov/research/ostp/Chem. Navigator. asp Contact: Marc Nicklaus, Ph. D. , Computer-Aided Drug Design (CADD) Group mn 1@helix. nih. gov, 301 -846 -5903
 Mission To provide expertise in advanced technologies in the areas of genomics and gene discovery, proteomics, analytical technologies, microscopic imaging, and computation to support the scientific initiatives and programs at NCI
Mission To provide expertise in advanced technologies in the areas of genomics and gene discovery, proteomics, analytical technologies, microscopic imaging, and computation to support the scientific initiatives and programs at NCI
 What the RTP Has to Offer Access to sophisticated technological approaches and instrumentation Collaborative contribution to research projects Development of new or improved technologies through networking and strategic partnerships within or outside of NCI Innovative, integrated, and customized solutions to complex research challenges Information management & tools for extracting meaning from large data sets Research, Technology Development and Core Service
What the RTP Has to Offer Access to sophisticated technological approaches and instrumentation Collaborative contribution to research projects Development of new or improved technologies through networking and strategic partnerships within or outside of NCI Innovative, integrated, and customized solutions to complex research challenges Information management & tools for extracting meaning from large data sets Research, Technology Development and Core Service
 Research Technology Program Proteomics/MS - Timothy Veenstra and Tom Conrads Molecular Technology - David Munroe Protein Chemistry - Robert Fisher Protein Expression - James Hartley Gene Expression - Bruce Crise Image Analysis - Stephen Lockett Advanced Biomedical Computing - Stan Burt
Research Technology Program Proteomics/MS - Timothy Veenstra and Tom Conrads Molecular Technology - David Munroe Protein Chemistry - Robert Fisher Protein Expression - James Hartley Gene Expression - Bruce Crise Image Analysis - Stephen Lockett Advanced Biomedical Computing - Stan Burt
 Funding available for the development and application of new technologies: NCI-Frederick Office of Director Research Support (ODRS) RTP/ODRS Projects: Funding for RTP Projects (50%) n n Not readily categorized as Technology Development or Core Service projects - pilot, risky, uncertain outcome Represent the transition of a Technology Development project into a new research area Represent feasibility or pilot studies which will serve as a starting point for future Core Service projects Focus the direction and provide validation of Technology Development
Funding available for the development and application of new technologies: NCI-Frederick Office of Director Research Support (ODRS) RTP/ODRS Projects: Funding for RTP Projects (50%) n n Not readily categorized as Technology Development or Core Service projects - pilot, risky, uncertain outcome Represent the transition of a Technology Development project into a new research area Represent feasibility or pilot studies which will serve as a starting point for future Core Service projects Focus the direction and provide validation of Technology Development
 http: //web. ncifcrf. gov/rtp/about/access-serv. asp
http: //web. ncifcrf. gov/rtp/about/access-serv. asp
 How to find OSTP on the web http: //ccr. cancer. gov/research/ostp/default. asp
How to find OSTP on the web http: //ccr. cancer. gov/research/ostp/default. asp
 OSTP Web Page
OSTP Web Page
 Shoshana Segal shosh@nih. gov, 301 -496 -0923 David Goldstein goldsted@ncifcrf. gov, 301 -496 -4347
Shoshana Segal shosh@nih. gov, 301 -496 -0923 David Goldstein goldsted@ncifcrf. gov, 301 -496 -4347


