948f43e7617722cf79c7d909bc680d6c.ppt
- Количество слайдов: 54

SH 2 domain Nuria Bonifaci Francesc Estanyol Structural Bio. Informatics BIOINFO, 2007

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic

Why are they called SH 2? • 1911 Peyton Rous discovers Sarcoma Rous Virus • Tyrosine Kinase v-Src • 3 domains: – Src homolog 1 (Kinase) – Src homolog 2 (SH 2) – Src homolog 3 (SH 3)

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic
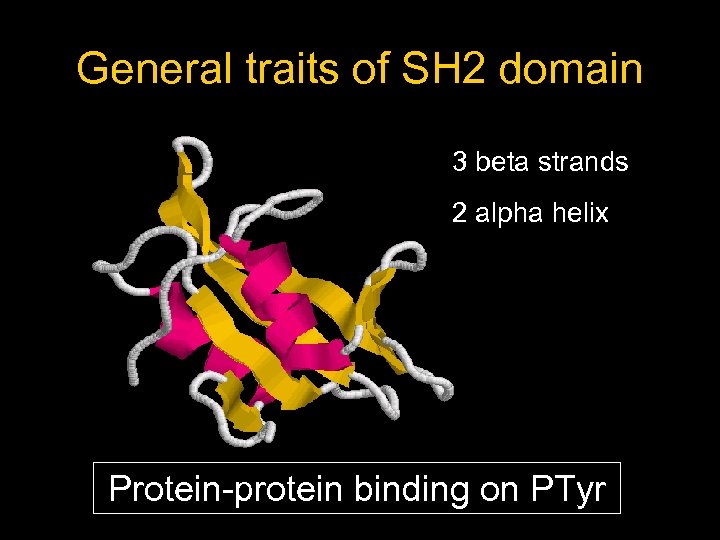
General traits of SH 2 domain 3 beta strands 2 alpha helix Protein-protein binding on PTyr

SH 2 domain
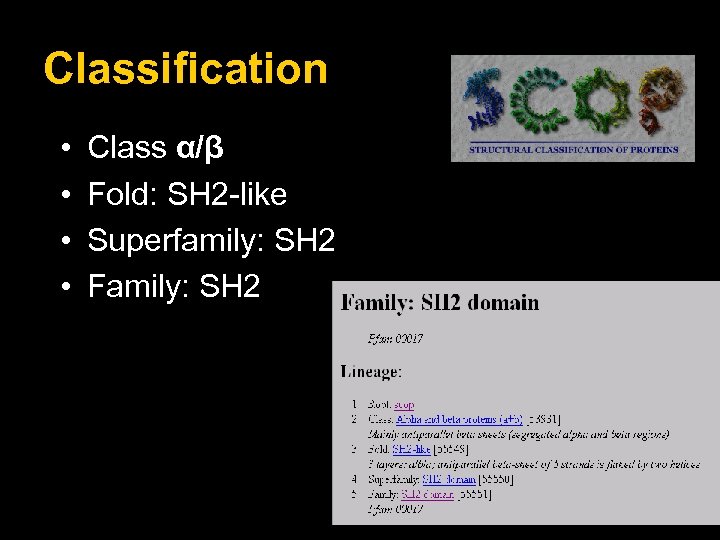

Classification • • Class α/β Fold: SH 2 -like Superfamily: SH 2 Family: SH 2

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic
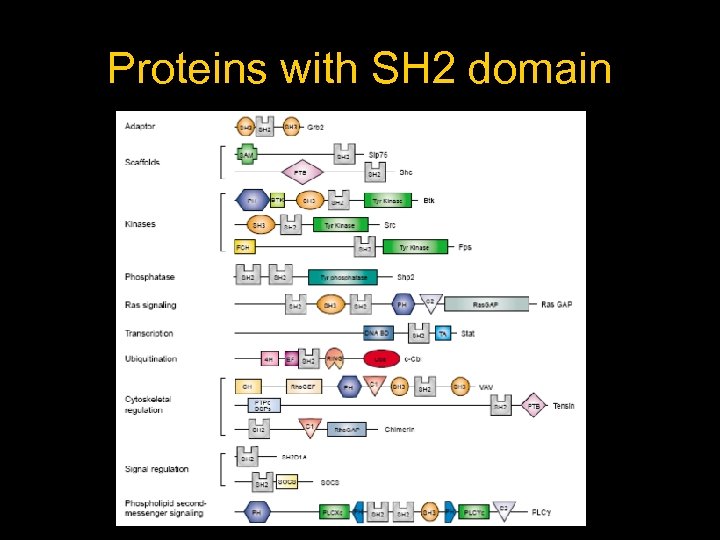
Proteins with SH 2 domain
Find the protein function:

Types of proteins with SH 2 -Kinases: Brk, Chk, Csk, Fyn, Hck, p 55 (Blk), p 56(lck), Pl 3 Kinase, Proto. oncogen. TK, Src, Syk, Zap-70 - Phosphatases: Shp 2, Syp - Adapters: Aps, Crk Proto-oncogen, Gbr 2, Gbr 7, Gbr 10, Mona - Dna Bindings: Cbl, STAT 1, STAT 3 b, STAThomolog - Others: Eat 2, PLCgamma, Sap

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic

Sequence conservation • Clustal. W Multiple sequence alignment between the kinases – Low conservation of the residues – High conservation of the characteristics
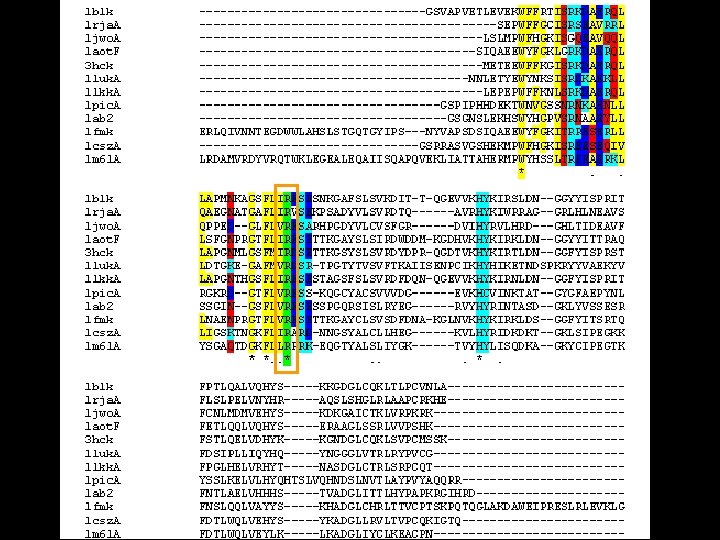
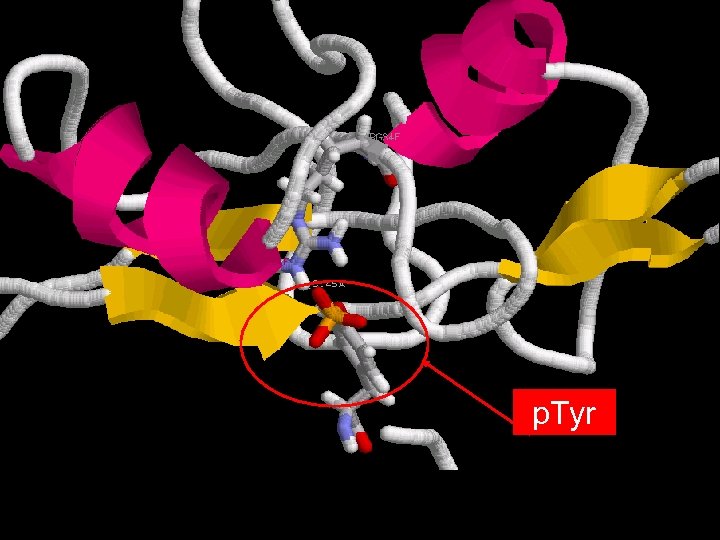
p. Tyr
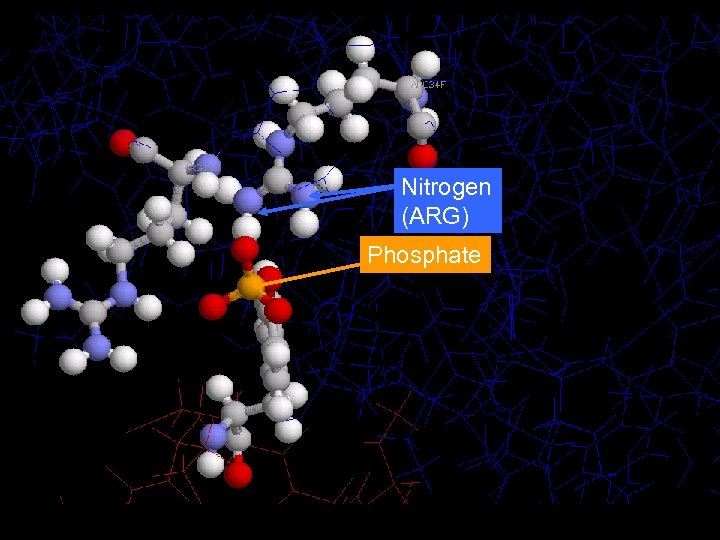
Nitrogen (ARG) Phosphate
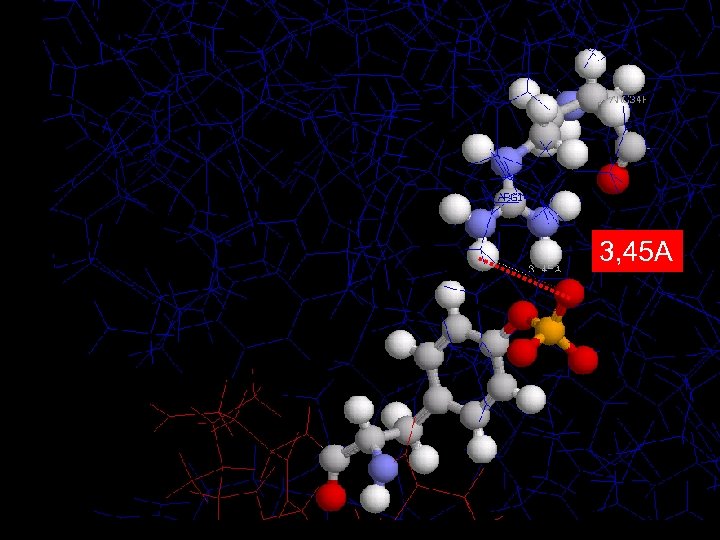
3, 45 A
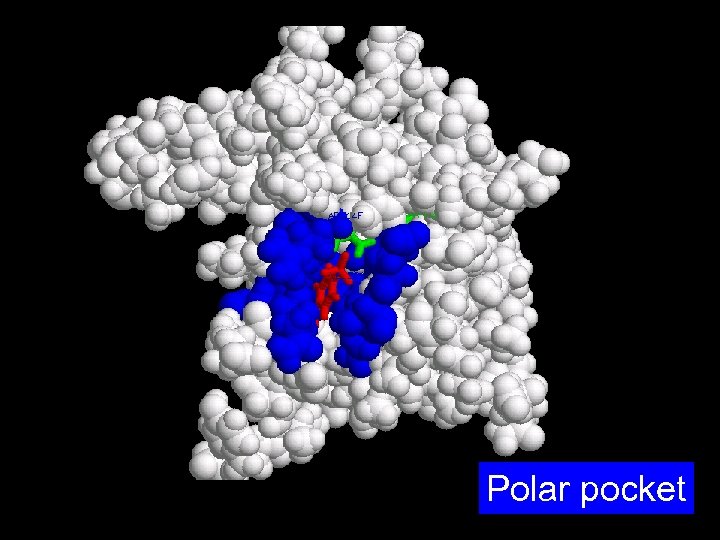
Polar pocket
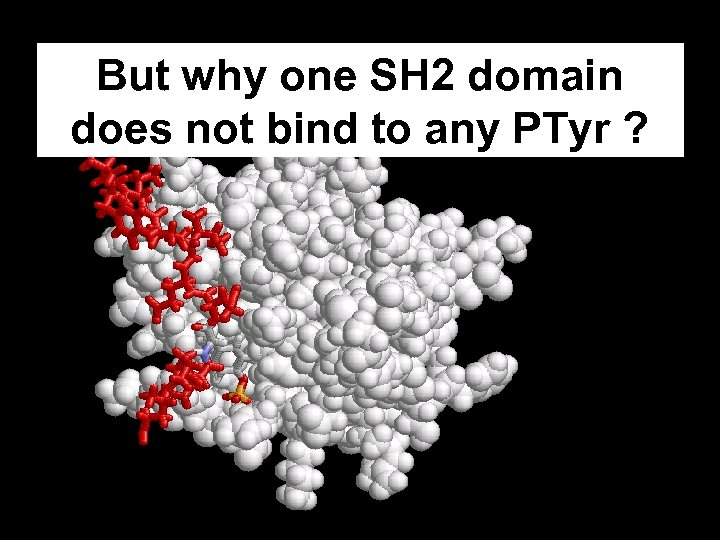
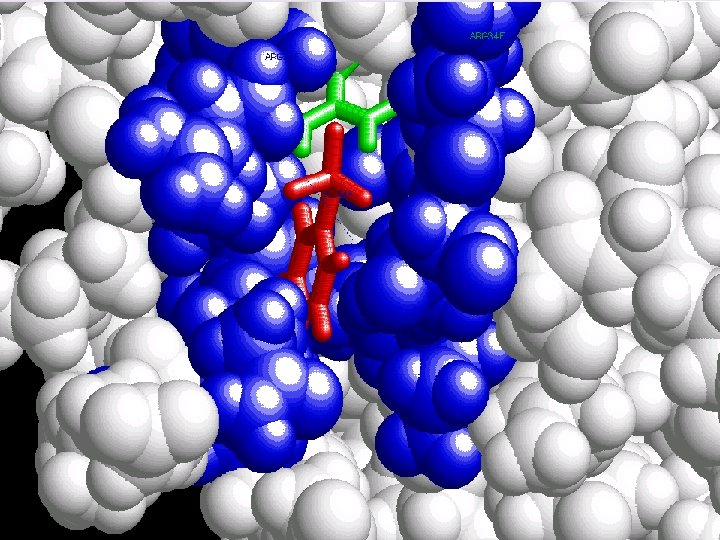

But why one SH 2 domain does not bind to any PTyr ?
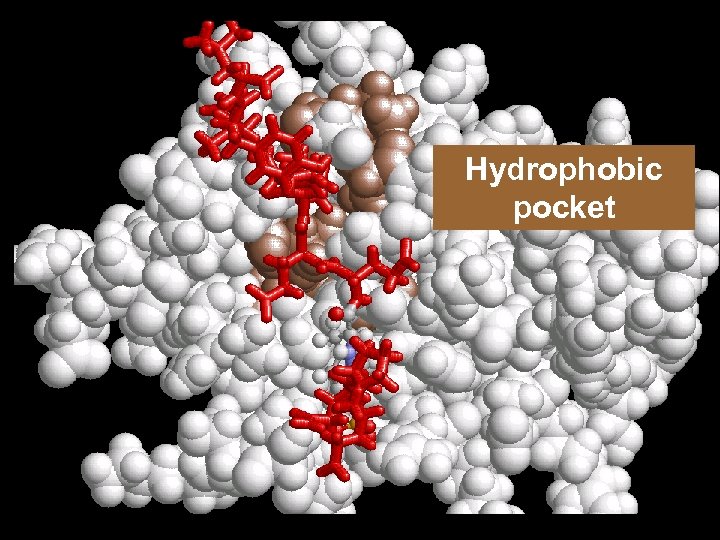
Hydrophobic pocket

Importance of the hydrophobic pocket Src Kinase PTyr-X-Asn

Grb 2 Ptyr- Val- Asn- Val

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic
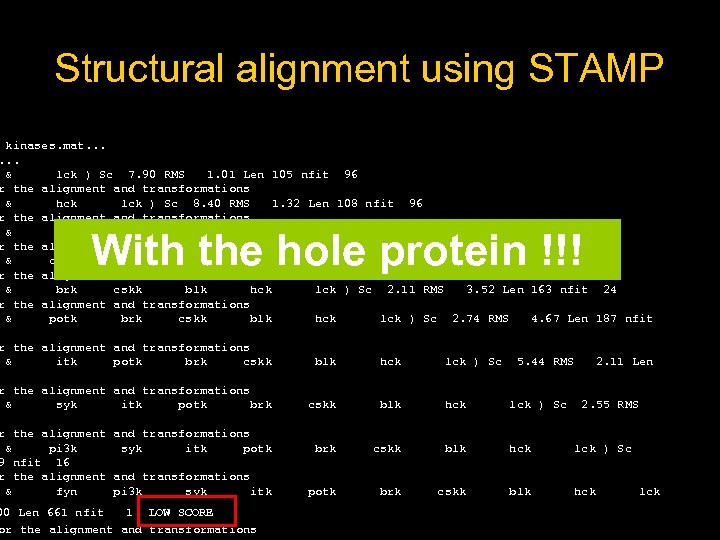

Structural alignment using STAMP kinases. mat. . . & lck ) Sc 7. 90 RMS 1. 01 Len 105 nfit 96 r the alignment and transformations & hck lck ) Sc 8. 40 RMS 1. 32 Len 108 nfit 96 r the alignment and transformations & blk hck lck ) Sc 7. 96 RMS 0. 95 Len 107 nfit 85 r the alignment and transformations & cskk blk hck lck ) Sc 7. 97 RMS 1. 31 Len 106 nfit 84 r the alignment and transformations & brk cskk blk hck lck ) Sc 2. 11 RMS 3. 52 Len 163 nfit 24 r the alignment and transformations & potk brk cskk blk hck lck ) Sc 2. 74 RMS 4. 67 Len 187 nfit With the hole protein !!! r the alignment and transformations & itk potk brk cskk blk hck lck ) Sc r the alignment and transformations & syk itk potk brk cskk blk hck lck ) Sc potk brk cskk blk hck r the alignment and transformations & pi 3 k syk itk potk 9 nfit 16 r the alignment and transformations & fyn pi 3 k syk itk 00 Len 661 nfit 1 LOW SCORE or the alignment and transformations 5. 44 RMS 2. 11 Len 2. 55 RMS lck
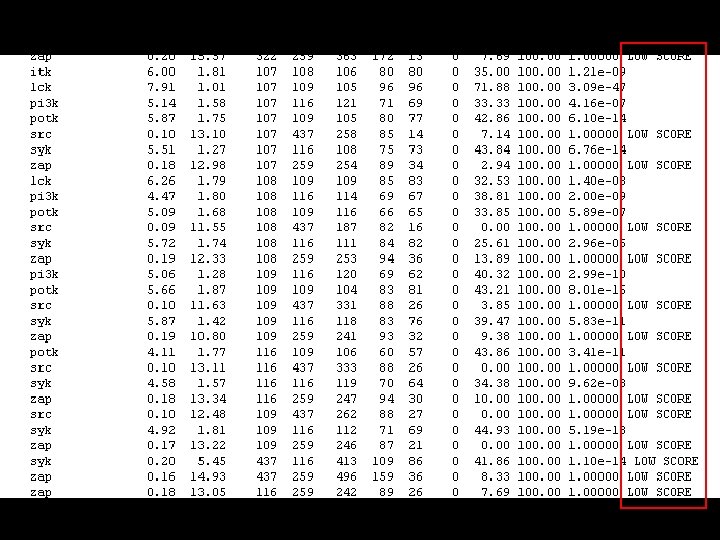

The solution • Cut manually ALL the PDB files selecting only the SH 2 domain ? ? ? A LOT OF WORK!!!

We used our bioinformatic knowledge!!! • Do a Phyton script to cut the PDB files given a starting and ending residue numbers • Get the FASTA sequences of the resulting PDBs But…where we can find the start/end residues?
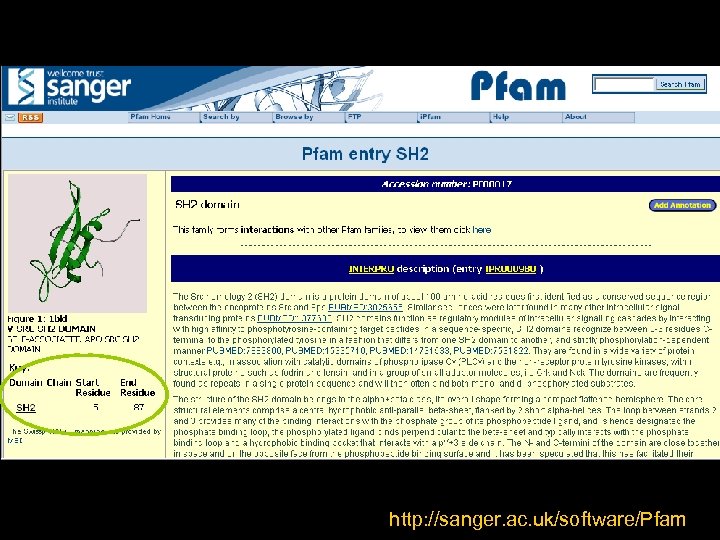
http: //sanger. ac. uk/software/Pfam

cut. PDB. py

Aminoacid 2 simbol function
NO LOW SCORES!!!
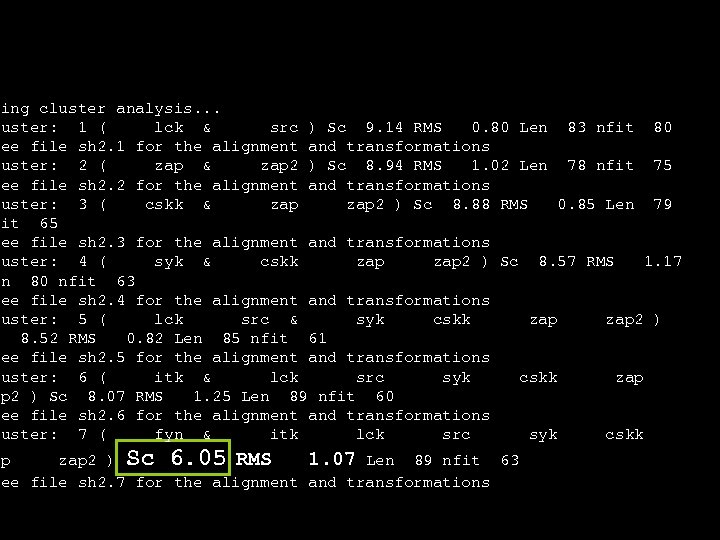
oing cluster analysis. . . luster: 1 ( lck & src ) Sc 9. 14 RMS 0. 80 Len 83 nfit 80 See file sh 2. 1 for the alignment and transformations luster: 2 ( zap & zap 2 ) Sc 8. 94 RMS 1. 02 Len 78 nfit 75 See file sh 2. 2 for the alignment and transformations luster: 3 ( cskk & zap 2 ) Sc 8. 88 RMS 0. 85 Len 79 fit 65 See file sh 2. 3 for the alignment and transformations luster: 4 ( syk & cskk zap 2 ) Sc 8. 57 RMS 1. 17 en 80 nfit 63 See file sh 2. 4 for the alignment and transformations luster: 5 ( lck src & syk cskk zap 2 ) c 8. 52 RMS 0. 82 Len 85 nfit 61 See file sh 2. 5 for the alignment and transformations luster: 6 ( itk & lck src syk cskk zap ap 2 ) Sc 8. 07 RMS 1. 25 Len 89 nfit 60 See file sh 2. 6 for the alignment and transformations luster: 7 ( fyn & itk lck src syk cskk ap zap 2 ) Sc 6. 05 RMS 1. 07 Len 89 nfit 63 See file sh 2. 7 for the alignment and transformations
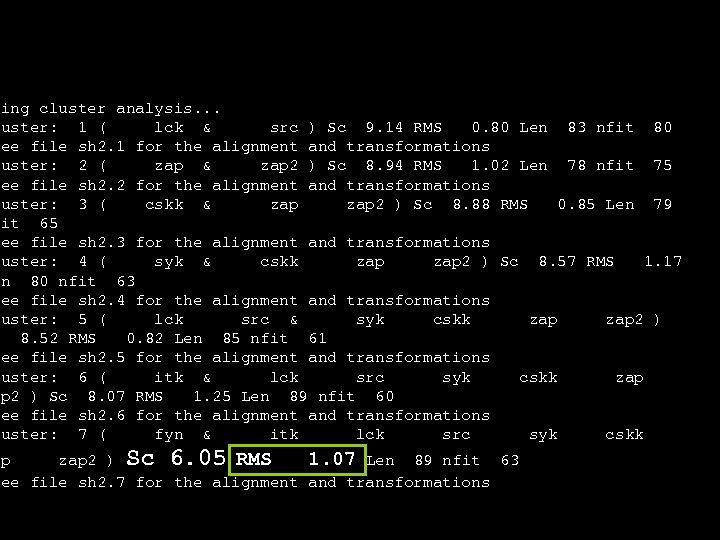
oing cluster analysis. . . luster: 1 ( lck & src ) Sc 9. 14 RMS 0. 80 Len 83 nfit 80 See file sh 2. 1 for the alignment and transformations luster: 2 ( zap & zap 2 ) Sc 8. 94 RMS 1. 02 Len 78 nfit 75 See file sh 2. 2 for the alignment and transformations luster: 3 ( cskk & zap 2 ) Sc 8. 88 RMS 0. 85 Len 79 fit 65 See file sh 2. 3 for the alignment and transformations luster: 4 ( syk & cskk zap 2 ) Sc 8. 57 RMS 1. 17 en 80 nfit 63 See file sh 2. 4 for the alignment and transformations luster: 5 ( lck src & syk cskk zap 2 ) c 8. 52 RMS 0. 82 Len 85 nfit 61 See file sh 2. 5 for the alignment and transformations luster: 6 ( itk & lck src syk cskk zap ap 2 ) Sc 8. 07 RMS 1. 25 Len 89 nfit 60 See file sh 2. 6 for the alignment and transformations luster: 7 ( fyn & itk lck src syk cskk ap zap 2 ) Sc 6. 05 RMS 1. 07 Len 89 nfit 63 See file sh 2. 7 for the alignment and transformations
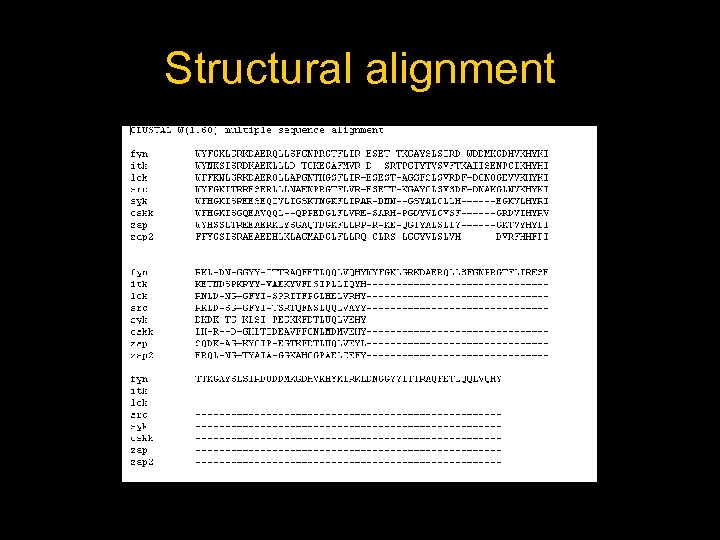

Structural alignment
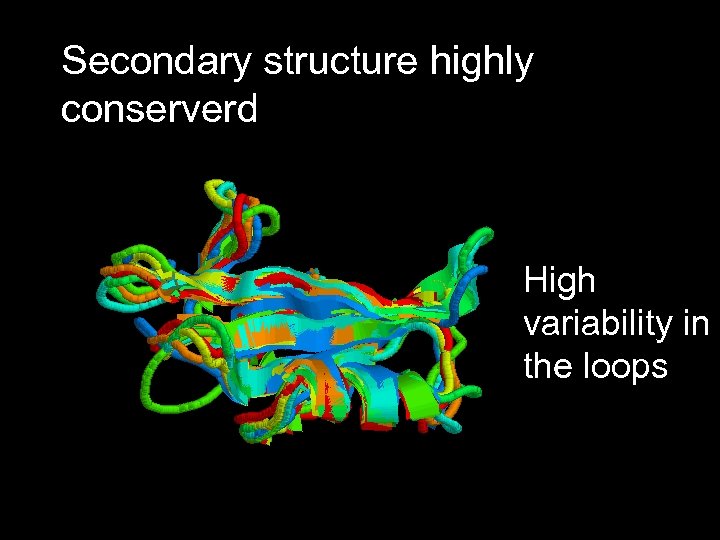
Secondary structure highly conserverd High variability in the loops
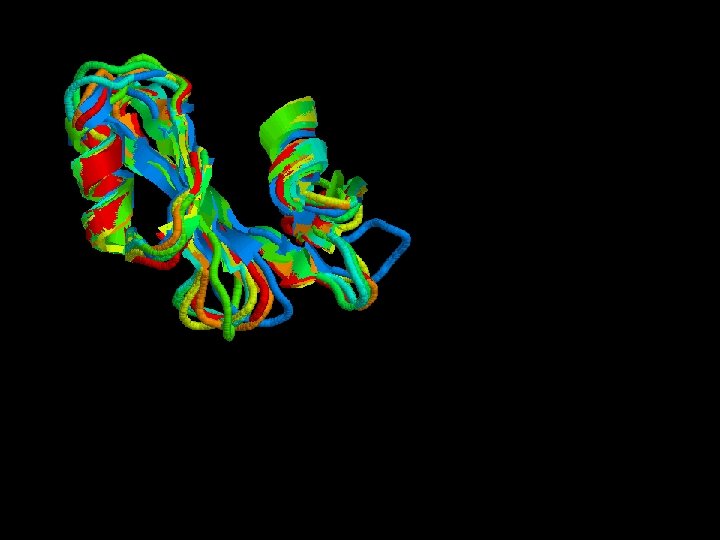
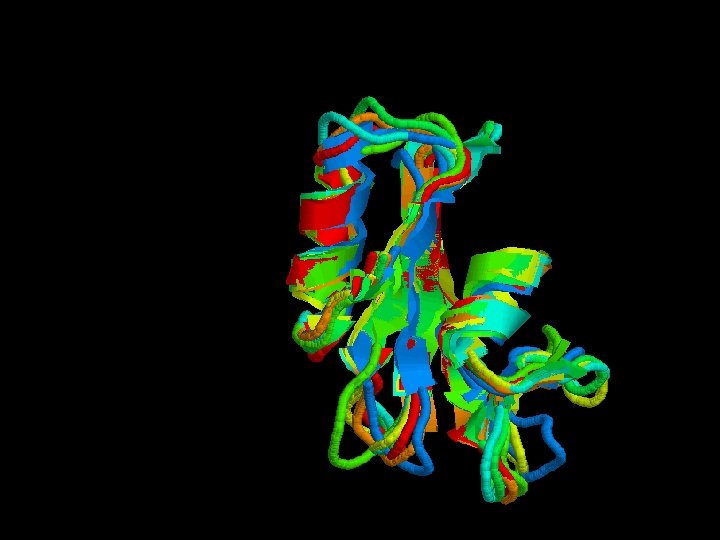
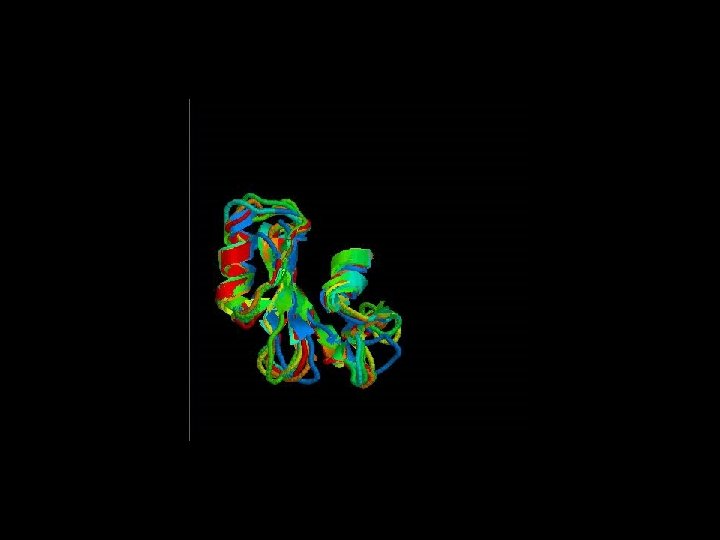
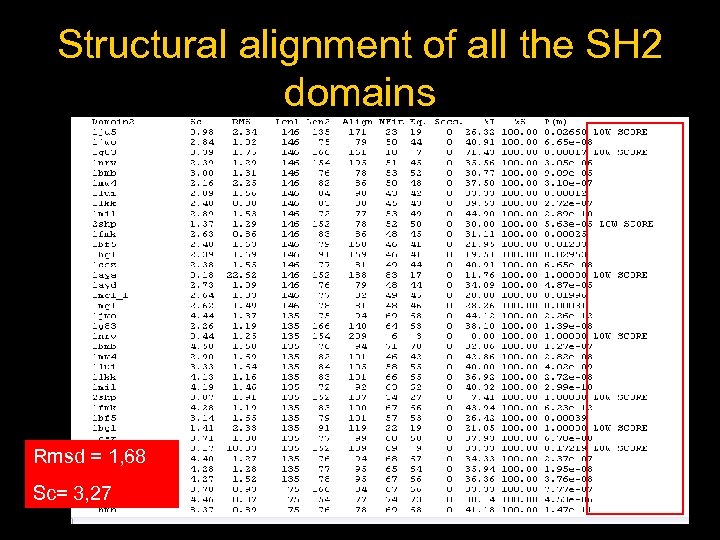
Structural alignment of all the SH 2 domains Rmsd = 1, 68 Sc= 3, 27
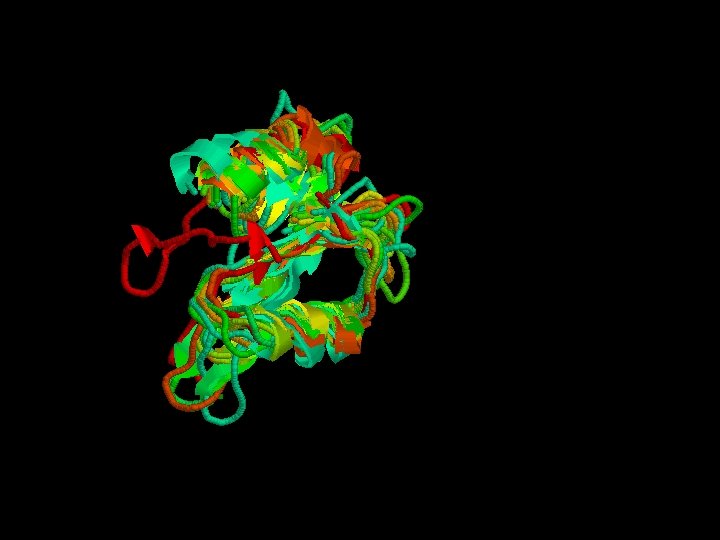
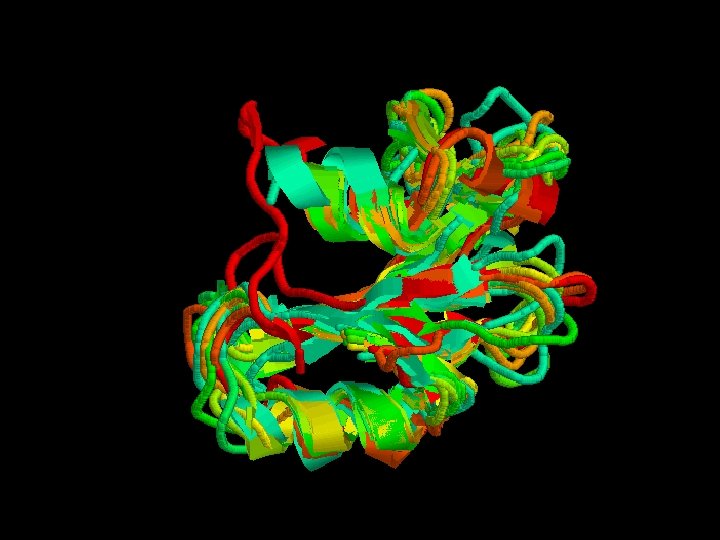

Advanced options of STAMP - Guide the program to do the superposition giving an initial alignment. – 1 -. Clustal. W of all the sequences. – 2 -. > alignfit – f sh 2. align –d sh 2. domains –out sh 2. trans > stamp –l sh 2. trans –prefix sh 2 Stamp We can’t do it : - alignfit ? - other. trans files didn’t work.

INDEX • • Introduction General traits of SH 2 domain Proteins with SH 2 domain Study of conservation – Sequence – Structure – Phylogenetic
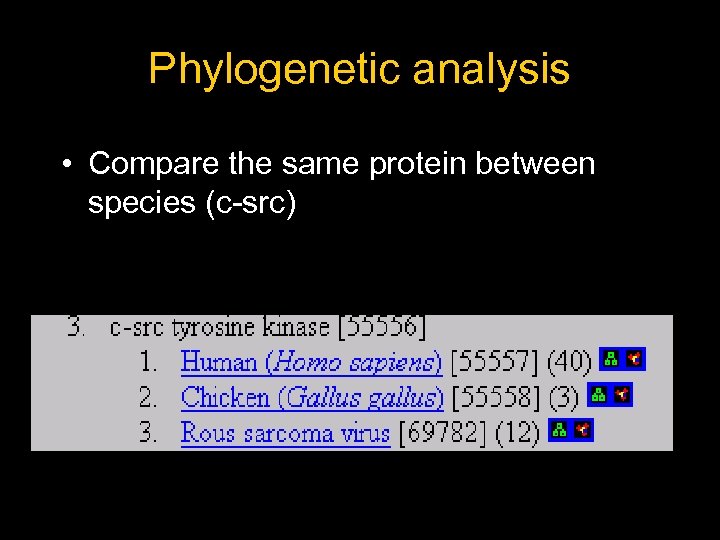

Phylogenetic analysis • Compare the same protein between species (c-src)

Good alignment !!!
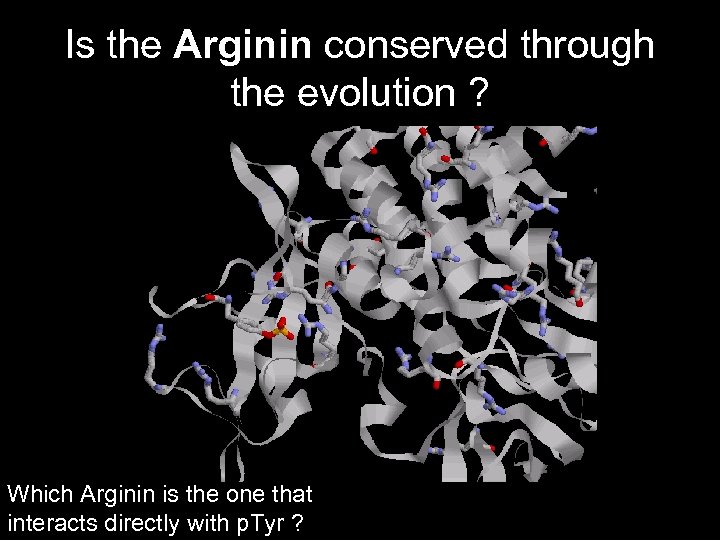
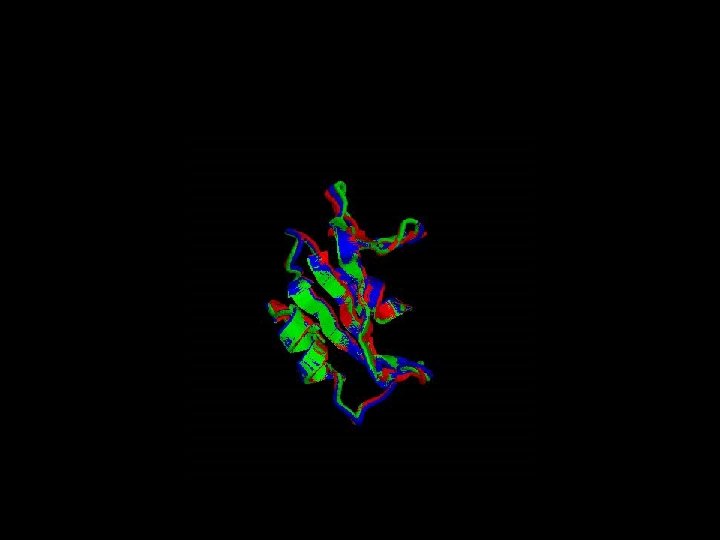

Is the Arginin conserved through the evolution ? Which Arginin is the one that interacts directly with p. Tyr ?
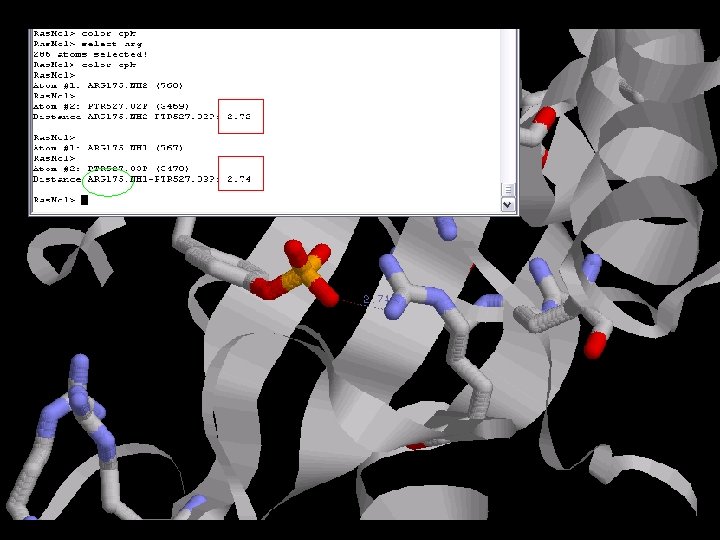
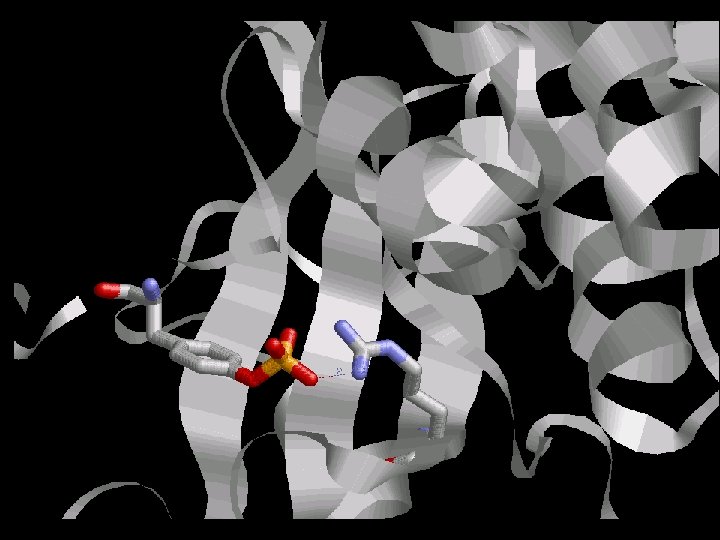

The Arginin is very conserved between the species due to its critical function

References - Bradshaw JM, Waksman G Molecular Recognition by SH 2 domains - Bhushan Nagar, Oliver Hantschel, Markus Seeliger, Jason M. Davies, William I. Weis, Giulio Superti-Furga and John Kuriyan Organization of the SH 3 -SH 2 Unit in Active and Inactive Forms of the c-Abl Tyrosine Kinase - The SRC kinases. http: //jkweb. mcb. berkeley. edu (Berkeley Univeristy)

That’s All Any questions?
948f43e7617722cf79c7d909bc680d6c.ppt