56630590bb155e0bc8c261895e812503.ppt
- Количество слайдов: 96

Sequence Analysis (II) Yuh-Shan Jou (周玉山) jou@ibms. sinica. edu. tw Institute of Biomedical Sciences, Academia Sinica

Other Areas to Cover • • Genomic Data Annotation Common Domains prediction WWW Other Useful Genome Browsers
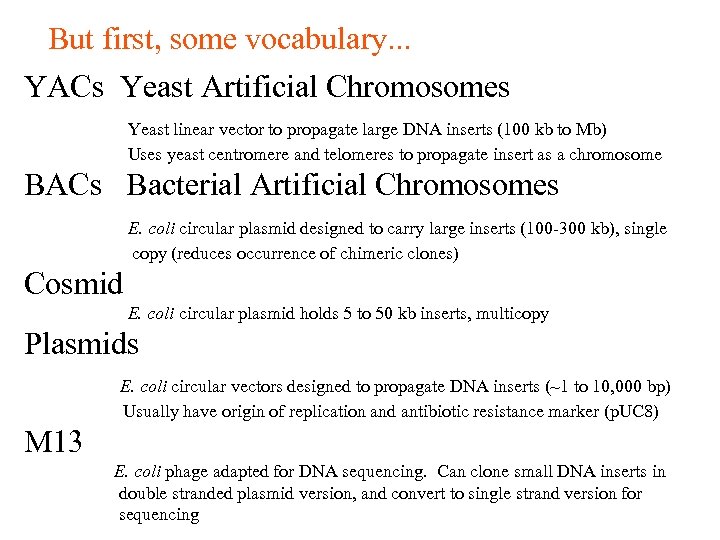


But first, some vocabulary. . . YACs Yeast Artificial Chromosomes Yeast linear vector to propagate large DNA inserts (100 kb to Mb) Uses yeast centromere and telomeres to propagate insert as a chromosome BACs Bacterial Artificial Chromosomes E. coli circular plasmid designed to carry large inserts (100 -300 kb), single copy (reduces occurrence of chimeric clones) Cosmid E. coli circular plasmid holds 5 to 50 kb inserts, multicopy Plasmids E. coli circular vectors designed to propagate DNA inserts (~1 to 10, 000 bp) Usually have origin of replication and antibiotic resistance marker (p. UC 8) M 13 E. coli phage adapted for DNA sequencing. Can clone small DNA inserts in double stranded plasmid version, and convert to single strand version for sequencing

c. DNA complementary DNA synthesized from RNA using reverse transcriptase (an RNAdependent DNA polymerase) EST Expressed Sequence Tag Single pass DNA sequence run of a c. DNA insert in a plasmid from one end ORF Open Reading Frame A region of at least 100 codons that is uninterrupted by stop codons and thus potentially encodes a protein SNP Single Nucleotide Polymorphism A single base that differs among members of a population. Can be detected by “genotyping”by PCR. Responsible for much trait diversity in populations (physical appearance, diseases, drug response). Satellite Marker Short tandem repeat (CACACACACAC, eg. ) with length polymorphisms in a population (10 CA’s vs 25, eg. ). Can be detected by genotyping. Often used for screening affected populations for disease genes(LOD scores). STS Sequenced Tag Site Short (~500 bp) segment of DNA of known sequence mapped to location

Ensembl Database and Web Browser Erin Pleasance Canada’s Michael Smith Genome Sciences Centre, Vancouver
www. ensembl. org Lecture 7. 1 7

What is Ensembl? • • • Joint project of EBI and Sanger Automated annotation of eukaryotic genomes Open source software Relational database system Web interface “The main aim of this campaign is to encourage scientists across the world - in academia, pharmaceutical companies, and the biotechnology and computer industries - to use this free information. ” - Dr. Mike Dexter, Director of the Wellcome Trust
TPMD: http: //tpmd. nhri. org. tw Nucleic Acids Research, 2005, Vol. 33, Database issue D 174 -D 177
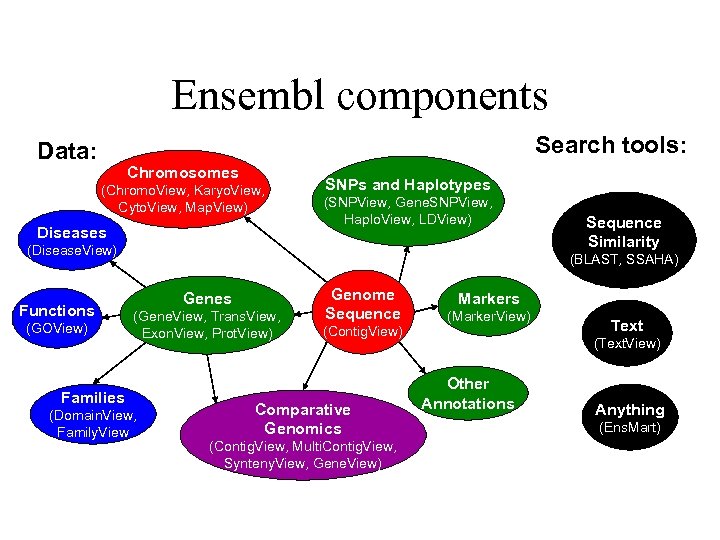
Ensembl components Search tools: Data: Chromosomes (Chromo. View, Karyo. View, Cyto. View, Map. View) Diseases SNPs and Haplotypes (SNPView, Gene. SNPView, Haplo. View, LDView) (Disease. View) Functions (GOView) Sequence Similarity (BLAST, SSAHA) Genes (Gene. View, Trans. View, Exon. View, Prot. View) Families (Domain. View, Family. View Genome Sequence (Contig. View) Comparative Genomics (Contig. View, Multi. Contig. View, Synteny. View, Gene. View) Markers (Marker. View) Text (Text. View) Other Annotations Anything (Ens. Mart)

Example 1: Exploring Caspase-3 • Aim to demonstrate basic browsing and views • Caspase-3 is a gene involved in apoptosis (cell suicide) • We will look at: – – Gene annotation SNPs Orthologs and genome alignments Alternative transcripts and EST genes

Example 1: Exploring Caspase-3 http: //www. ensembl. org Go to human homepage

Species-specific homepage Statistics of current release Site map
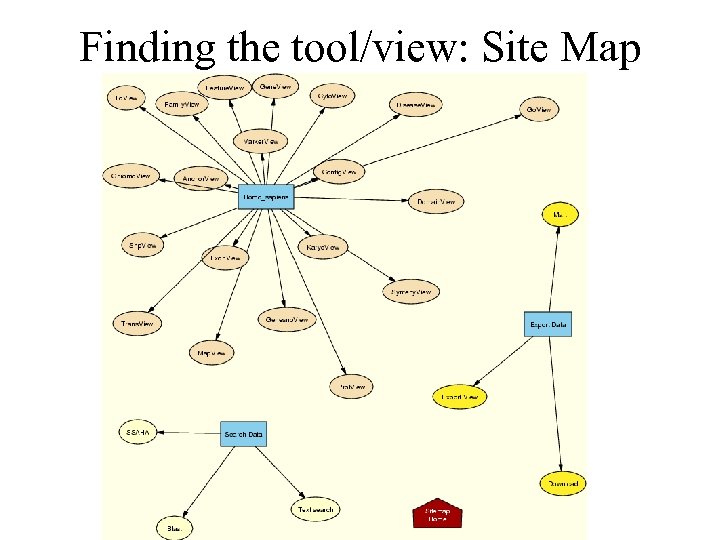
Finding the tool/view: Site Map
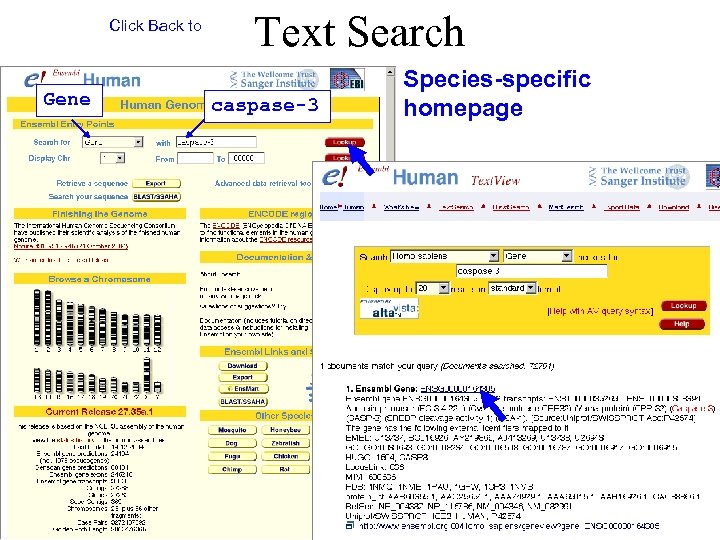
Click Back to Gene Text Search caspase-3 Species-specific homepage
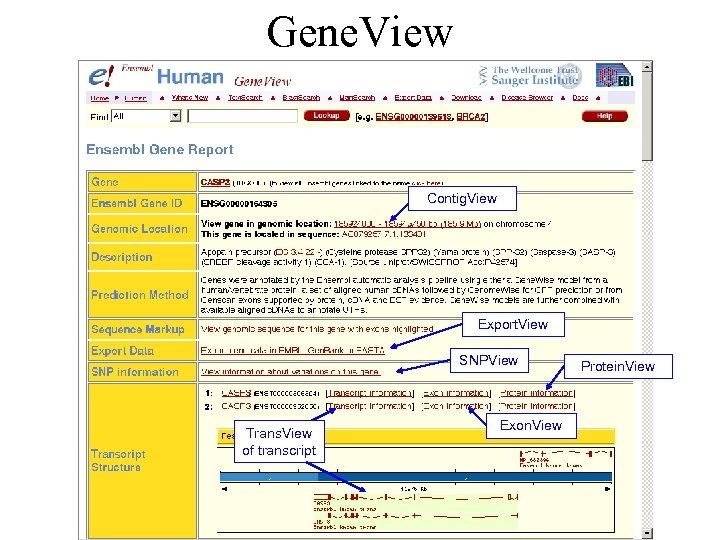
Gene. View Contig. View Export. View SNPView Trans. View of transcript Exon. View Protein. View
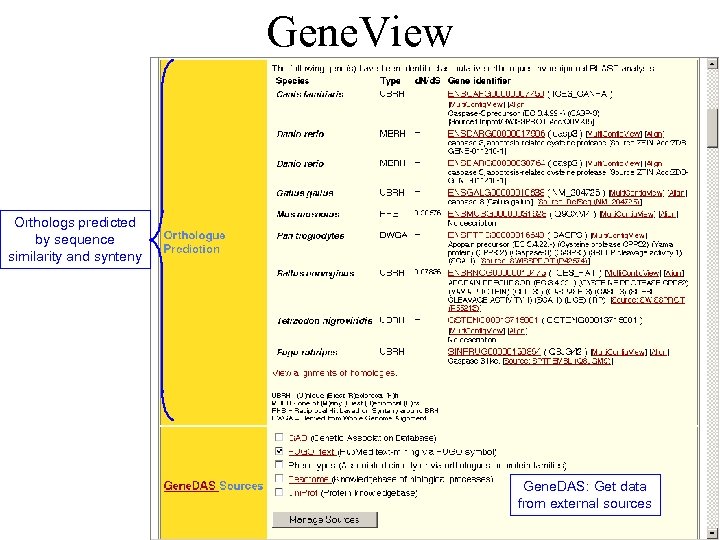

Gene. View Orthologs predicted by sequence similarity and synteny Gene. DAS: Get data from external sources
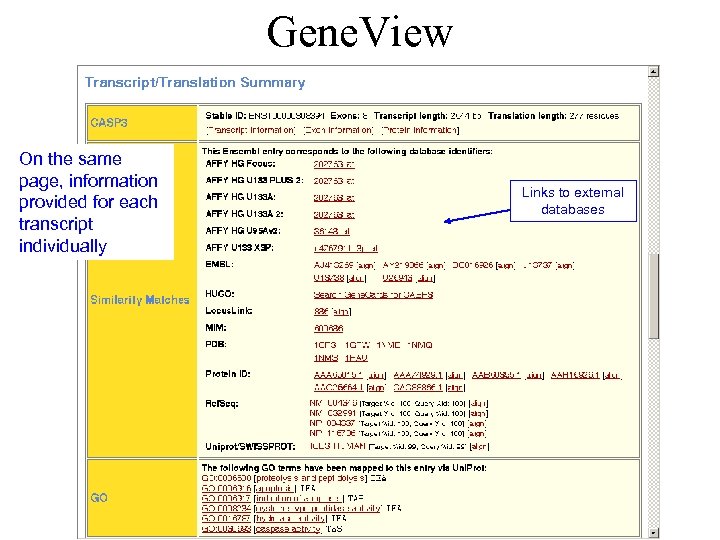
Gene. View On the same page, information provided for each transcript individually Links to external databases
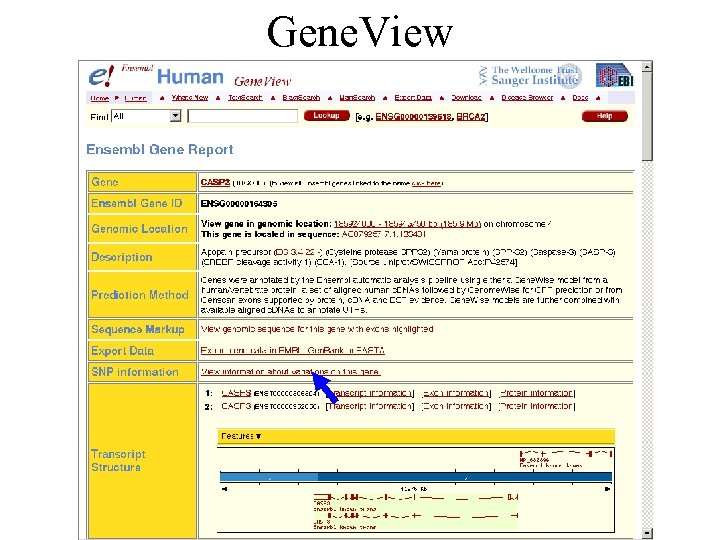
Gene. View
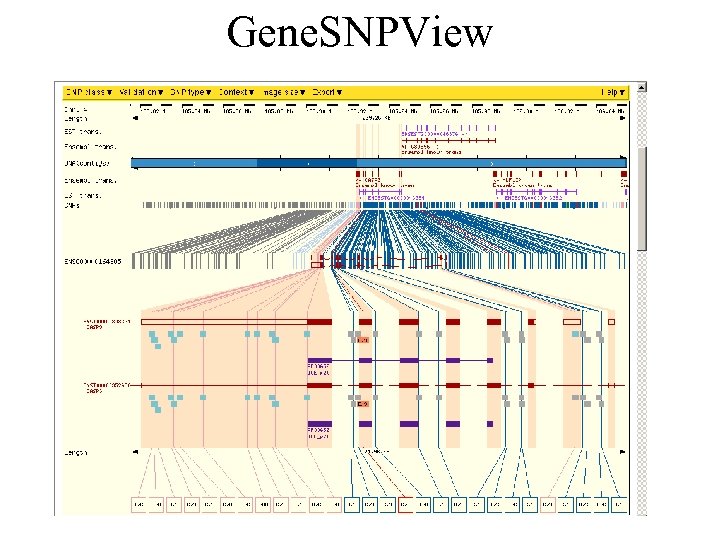
Gene. SNPView
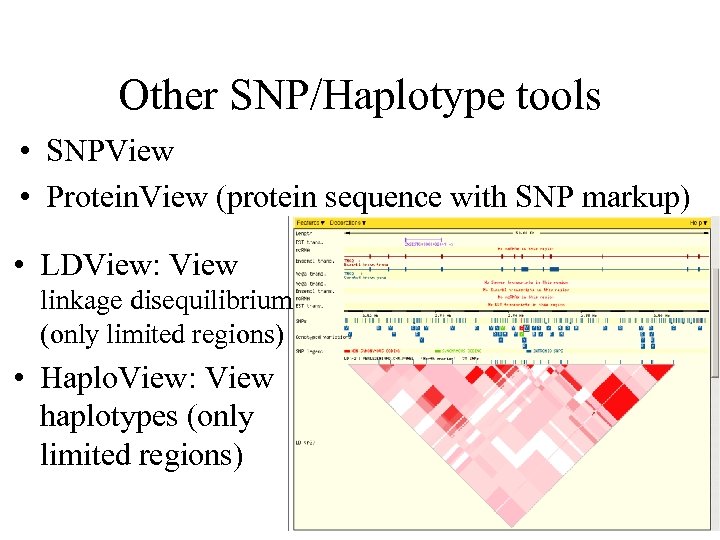

Other SNP/Haplotype tools • SNPView • Protein. View (protein sequence with SNP markup) • LDView: View linkage disequilibrium (only limited regions) • Haplo. View: View haplotypes (only limited regions)
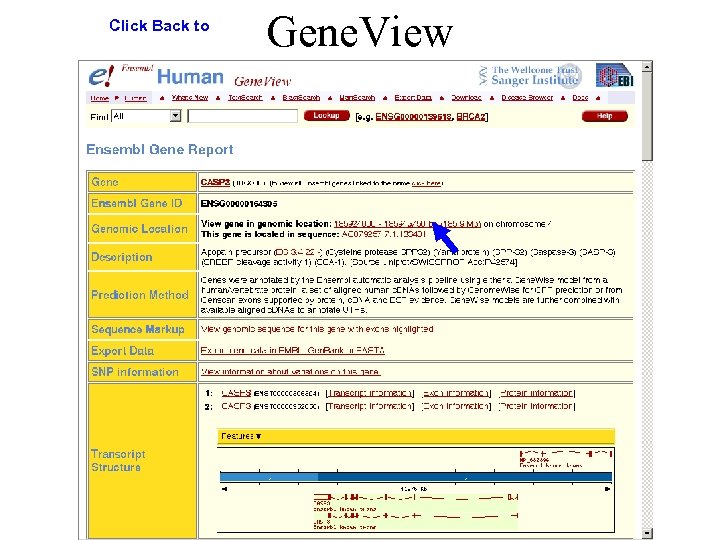
Click Back to Gene. View
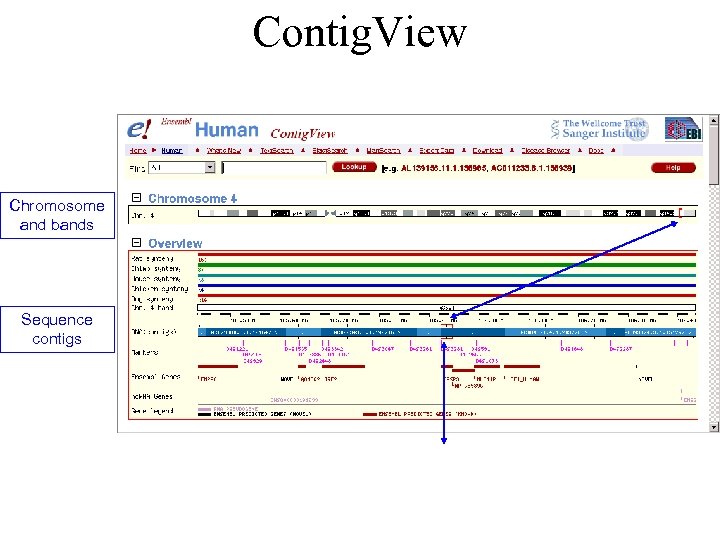
Contig. View Chromosome and bands Sequence contigs
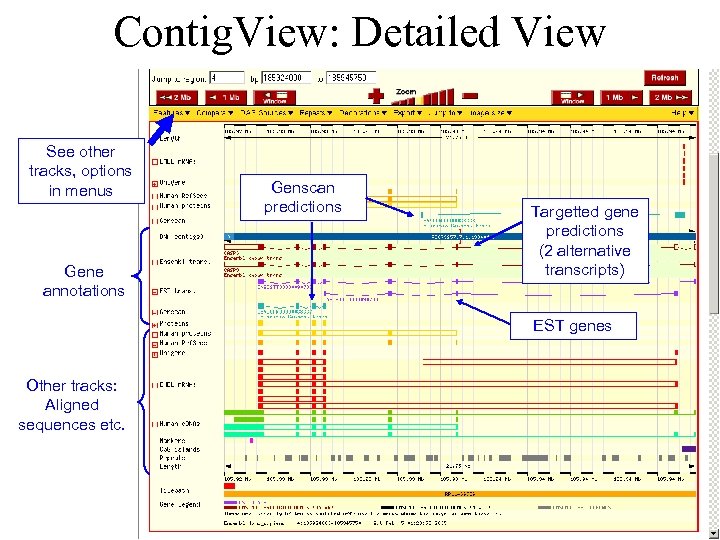
Contig. View: Detailed View See other tracks, options in menus Gene annotations Genscan predictions Targetted gene predictions (2 alternative transcripts) EST genes Other tracks: Aligned sequences etc.
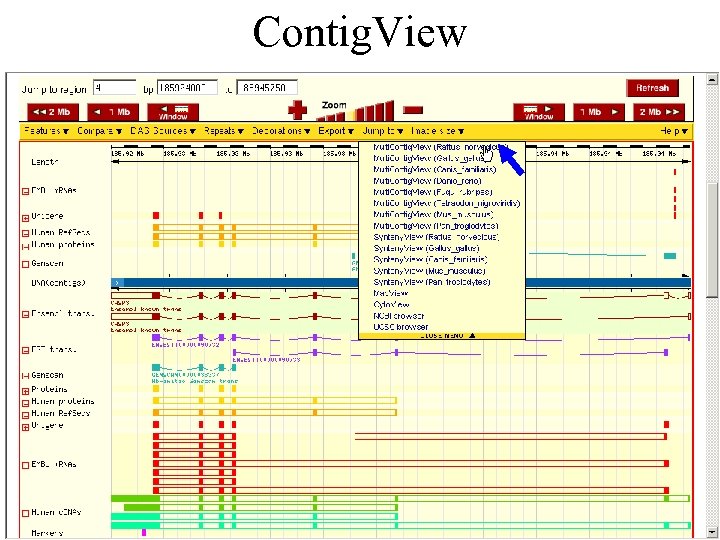
Contig. View
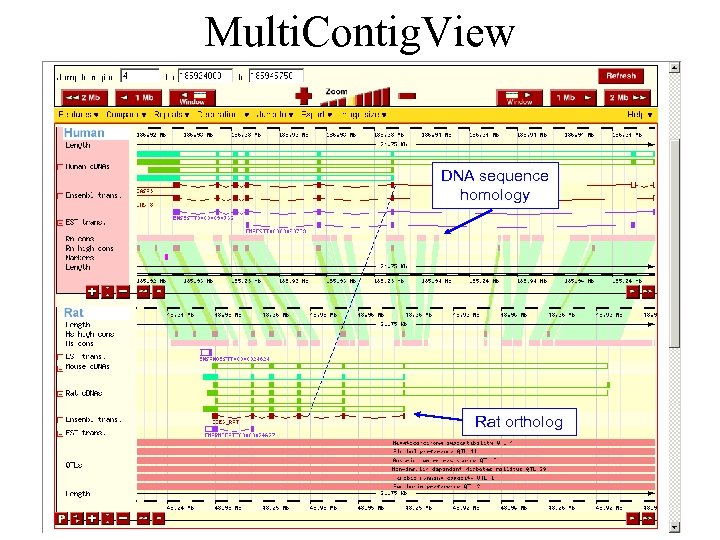
Multi. Contig. View DNA sequence homology Rat ortholog
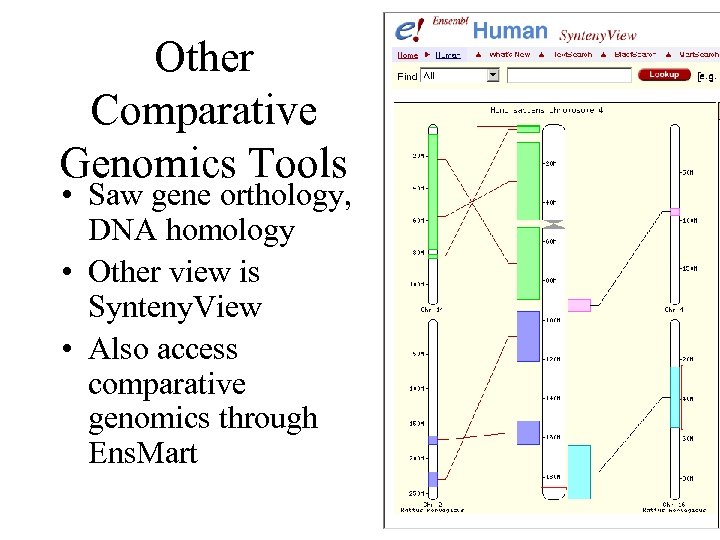

Other Comparative Genomics Tools • Saw gene orthology, DNA homology • Other view is Synteny. View • Also access comparative genomics through Ens. Mart

Data Mining with Ens. Mart • Allows very fast, cross-data source querying • Search for genes (features, sequences, etc. ) or SNPs based on – Position; function; domains; similarity; expression; etc. • Accessible from Ensembl website (Mart. View) as well as stand-alone • Extremely powerful for data mining

Example 2: Ens. Mart • A new disease locus has been mapped between markers D 21 S 1991 and D 21 S 171. It may be that the gene involved has already been identified as having a role in another disease. What candidates are in this region?

Example 2: Ens. Mart • Ens. Mart is based on Bio. Mart • http: //www. ensembl. org/Multi/martview OR • http: //www. ebi. ac. uk/Bio. Mart/martview
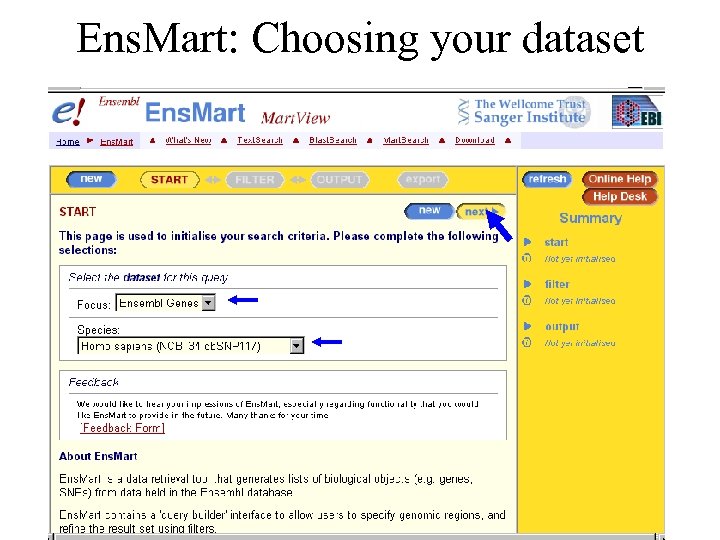
Ens. Mart: Choosing your dataset
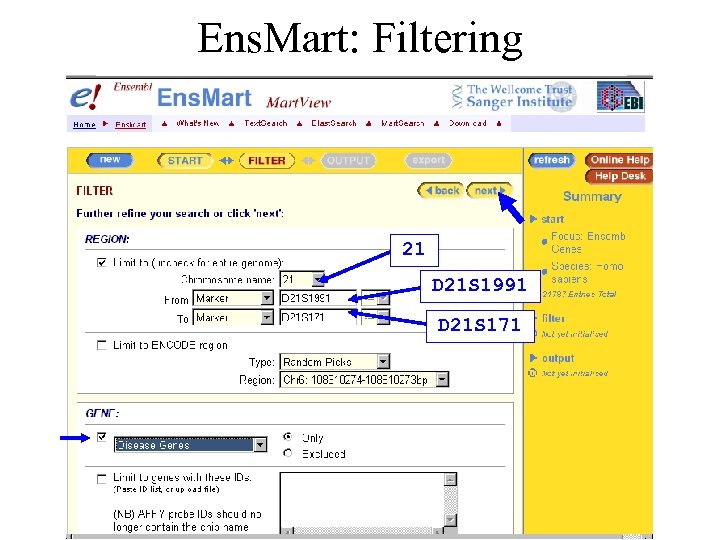
Ens. Mart: Filtering 21 D 21 S 1991 D 21 S 171
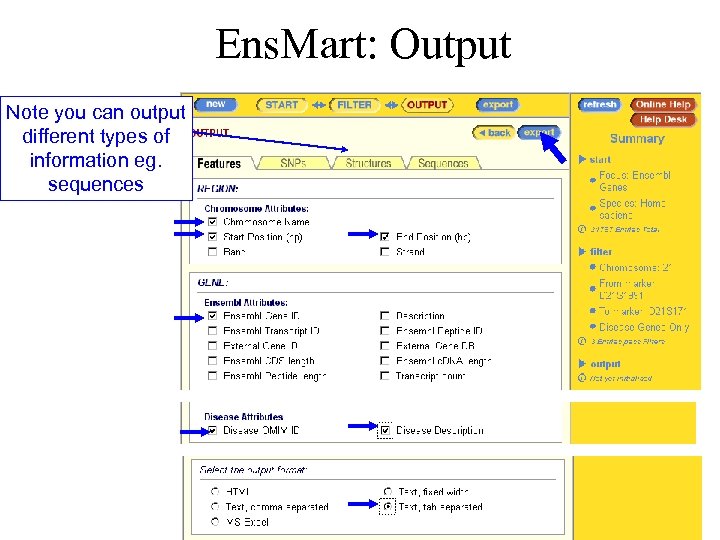
Ens. Mart: Output Note you can output different types of information eg. sequences
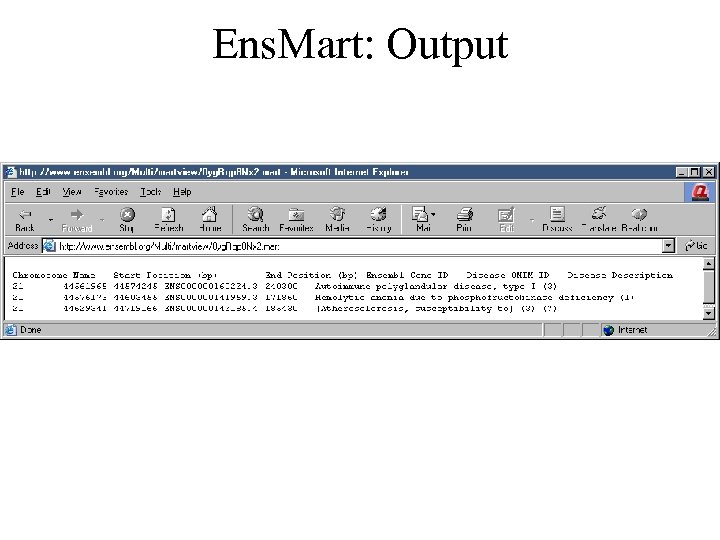
Ens. Mart: Output
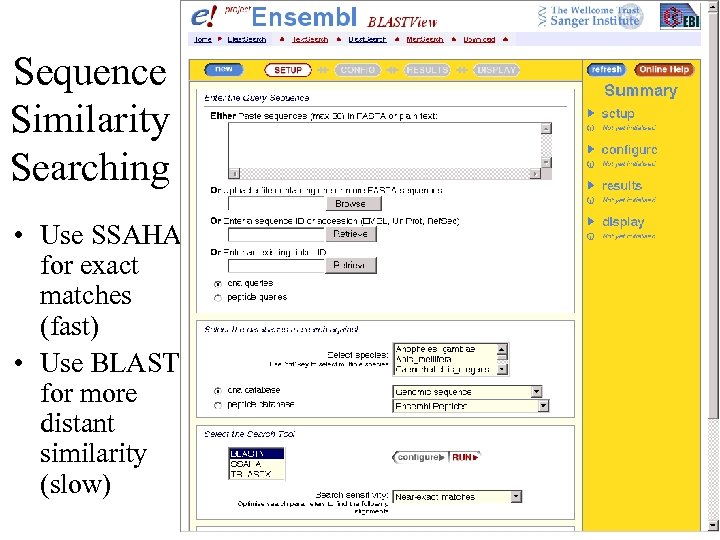
Sequence Similarity Searching • Use SSAHA for exact matches (fast) • Use BLAST for more distant similarity (slow)
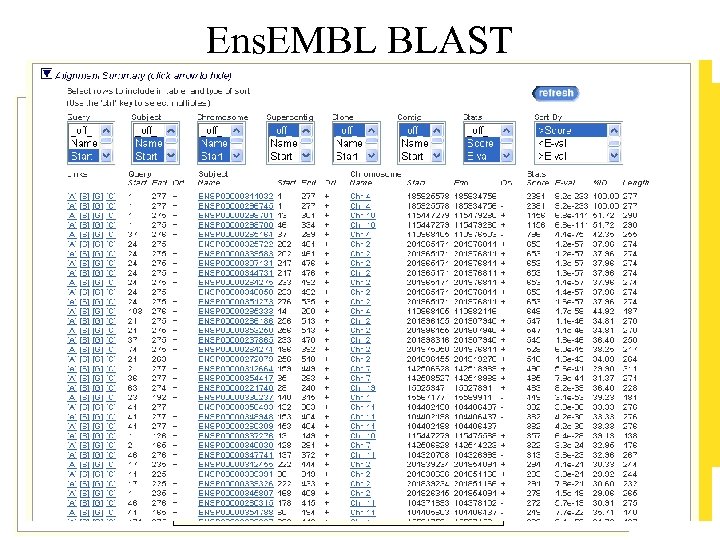
Ens. EMBL BLAST
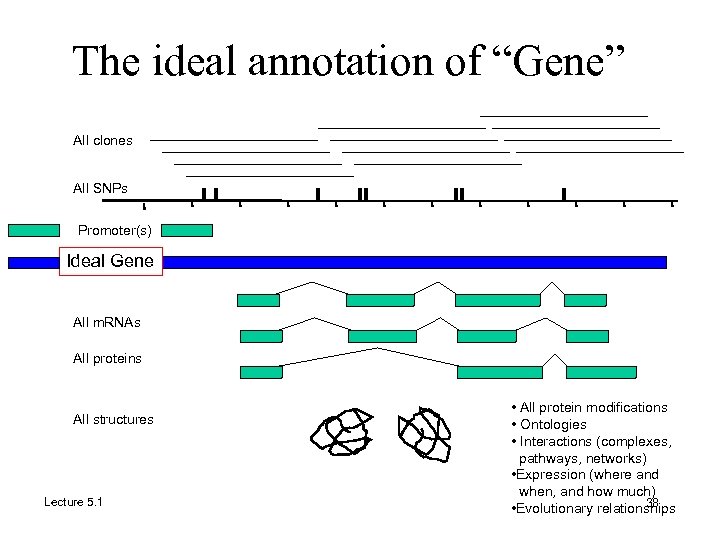
The ideal annotation of “Gene” All clones All SNPs Promoter(s) Ideal Gene All m. RNAs All proteins All structures Lecture 5. 1 • All protein modifications • Ontologies • Interactions (complexes, pathways, networks) • Expression (where and when, and how much) 38 • Evolutionary relationships
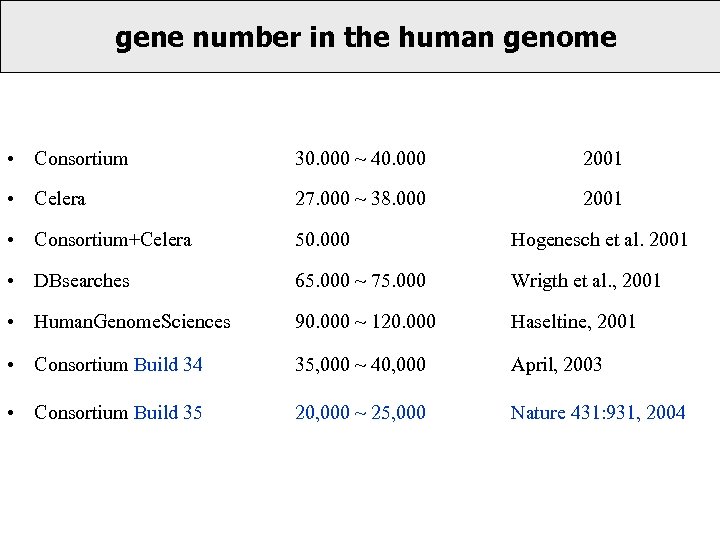

gene number in the human genome • Consortium 30. 000 ~ 40. 000 2001 • Celera 27. 000 ~ 38. 000 2001 • Consortium+Celera 50. 000 Hogenesch et al. 2001 • DBsearches 65. 000 ~ 75. 000 Wrigth et al. , 2001 • Human. Genome. Sciences 90. 000 ~ 120. 000 Haseltine, 2001 • Consortium Build 34 35, 000 ~ 40, 000 April, 2003 • Consortium Build 35 20, 000 ~ 25, 000 Nature 431: 931, 2004
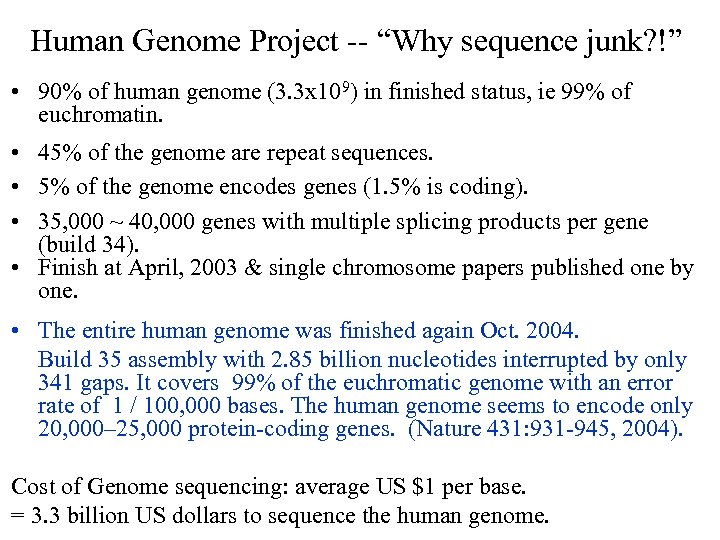

Human Genome Project -- “Why sequence junk? !” • 90% of human genome (3. 3 x 109) in finished status, ie 99% of euchromatin. • 45% of the genome are repeat sequences. • 5% of the genome encodes genes (1. 5% is coding). • 35, 000 ~ 40, 000 genes with multiple splicing products per gene (build 34). • Finish at April, 2003 & single chromosome papers published one by one. • The entire human genome was finished again Oct. 2004. Build 35 assembly with 2. 85 billion nucleotides interrupted by only 341 gaps. It covers 99% of the euchromatic genome with an error rate of 1 / 100, 000 bases. The human genome seems to encode only 20, 000– 25, 000 protein-coding genes. (Nature 431: 931 -945, 2004). Cost of Genome sequencing: average US $1 per base. = 3. 3 billion US dollars to sequence the human genome.

Ab initio gene identification • Goals – Identify coding exons – Seek gene structure information – Get a protein sequence for further analysis • Relevance – Characterization of anonymous DNA genomic sequences – Works on all DNA sequences

Gene Finding on the Web GRAIL: Oak Ridge Natl. Lab, Oak Ridge, TN – http: //compbio. ornl. gov/grailexp ORFfinder: NCBI – http: //www. ncbi. nlm. nih. gov/gorf. html DNA translation: Univ. of Minnesota Med. School – http: //alces. med. umn. edu/webtrans. html Gen. Lang – http: //cbil. humgen. upenn. edu/~sdong/genlang. html BCM Gene. Finder: Baylor College of Medicine, Houston, TX – http: //dot. imgen. bcm. tmc. edu: 9331/seq-search/gene-search. html – http: //dot. imgen. bcm. tmc. edu: 9331/gene-finder/gf. html
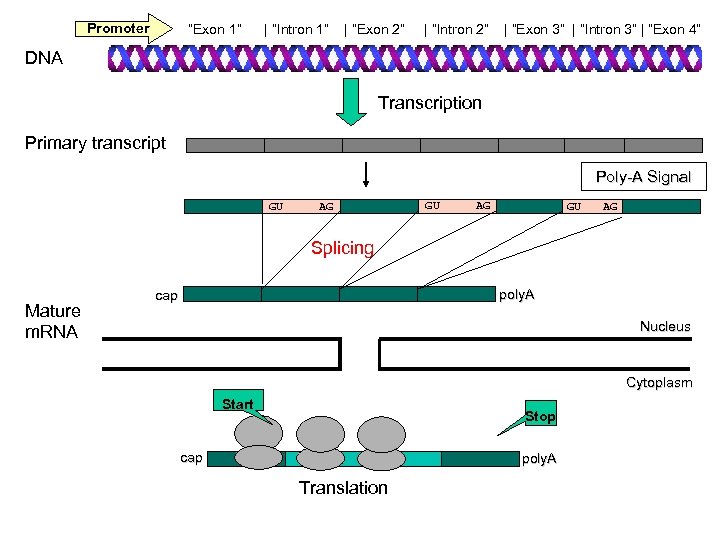
Promoter “Exon 1” | “Intron 1” | “Exon 2” | “Intron 2” | “Exon 3” | “Intron 3” | “Exon 4” DNA Transcription Primary transcript Poly-A Signal GU AG Splicing Mature m. RNA poly. A cap Nucleus Cytoplasm Start Stop cap poly. A Translation
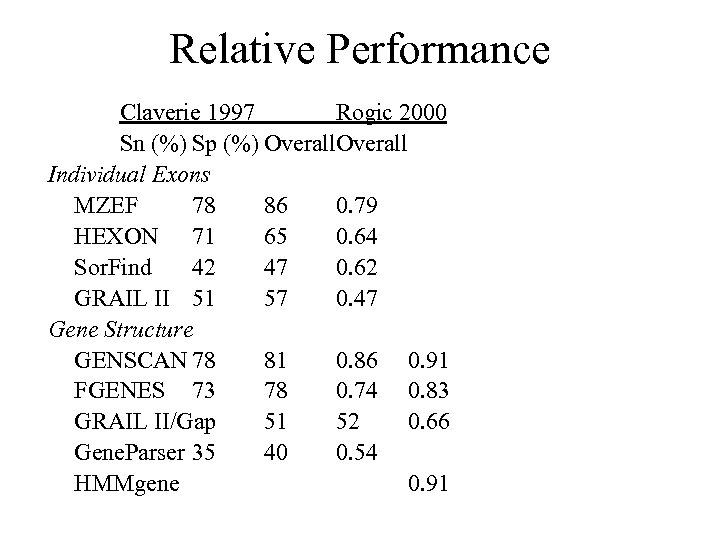
Relative Performance Claverie 1997 Rogic 2000 Sn (%) Sp (%) Overall Individual Exons MZEF 78 86 0. 79 HEXON 71 65 0. 64 Sor. Find 42 47 0. 62 GRAIL II 51 57 0. 47 Gene Structure GENSCAN 78 81 0. 86 0. 91 FGENES 73 78 0. 74 0. 83 GRAIL II/Gap 51 52 0. 66 Gene. Parser 35 40 0. 54 HMMgene 0. 91

What works best when? • Genome survey (draft) data: expect only a single exon in any given stretch of contiguous sequence – BLASTN vs. db. EST (3’ UTR) – BLASTX vs. nr (protein CDS) • Finished data: large contigs are available, providing context – GENSCAN – HMMgene

Things we are looking to annotate? • • • CDS m. RNA Alternative RNA Promoter and Poly-A Signal Pseudogenes nc. RNA Lecture 5. 1 46

Pseudogenes • Could be as high as 20 -30% of all genomic sequence predictions could be pseudogene • Non-functional copy of a gene – Processed pseudogene • • Retro-transposon derived No 5’ promoters No introns Often includes poly. A tail – Non-processed pseudogene • Gene duplication derived – Both include events that make the gene non-funtional • Frameshift • Stop codons • We assume pseudogenes have no function, but we really don’t know!

Noncoding RNA (nc. RNA) • nc. RNA represent 98% of all transcripts in a mammalian cell • nc. RNA have not been taken into account in gene counts • c. DNA • ORF computational prediction • Comparative genomics looking at ORF • nc. RNA can be: – Structural – Catalytic – Regulatory
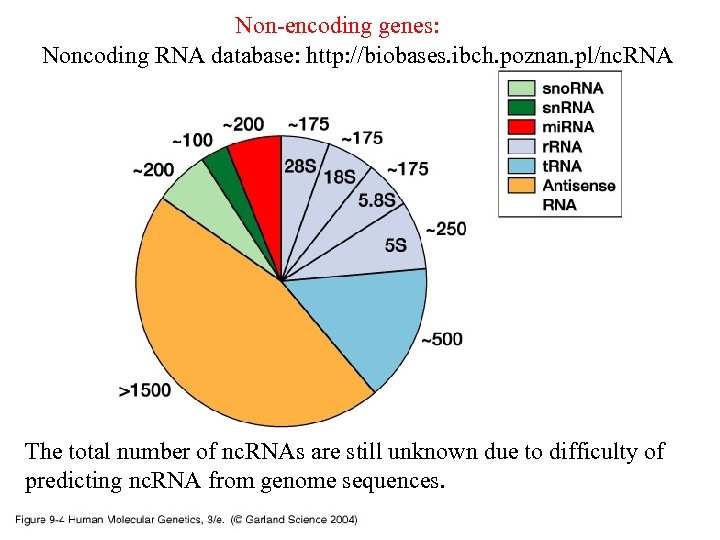
Non-encoding genes: Noncoding RNA database: http: //biobases. ibch. poznan. pl/nc. RNA 09_04. jpg The total number of nc. RNAs are still unknown due to difficulty of predicting nc. RNA from genome sequences.

Noncoding RNA (nc. RNA) • t. RNA – transfer RNA: involved in translation • r. RNA – ribosomal RNA: structural component of ribosome, where translation takes place • sno. RNA – small nucleolar RNA: functional/catalytic in RNA maturation • Antisense RNA: gene silencing
http: //rfam. wustl. edu
http: //rna. tbi. univie. ac. at/
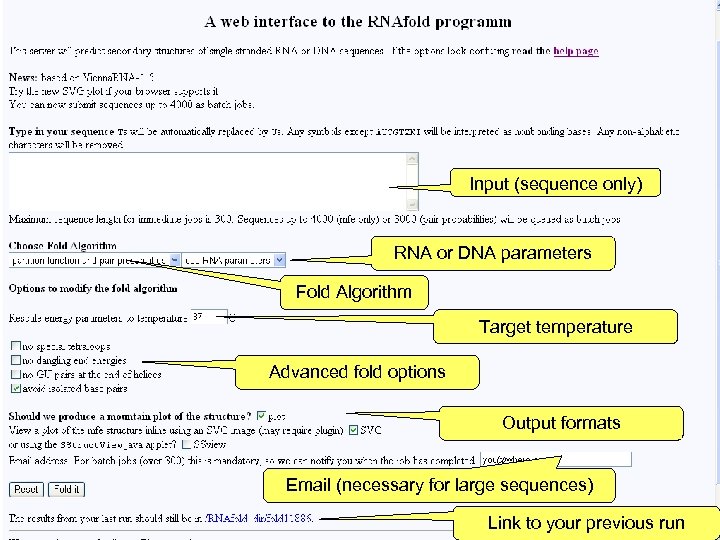
Input (sequence only) RNA or DNA parameters Fold Algorithm Target temperature Advanced fold options Output formats Email (necessary for large sequences) Link to your previous run
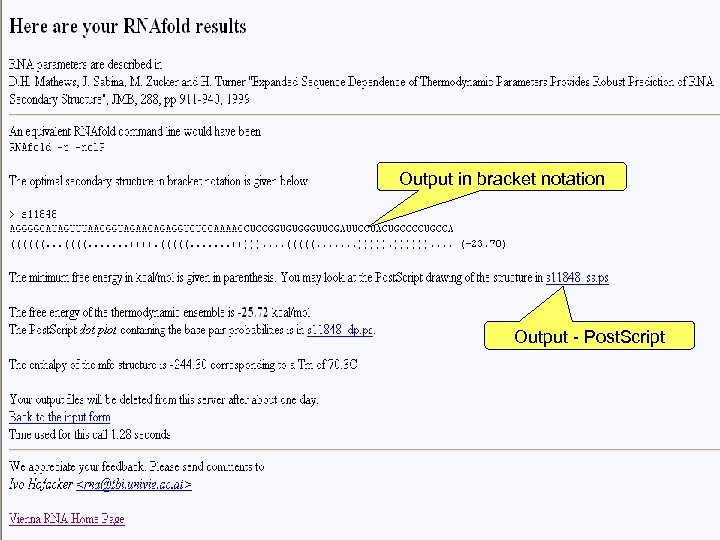
Output in bracket notation Output - Post. Script
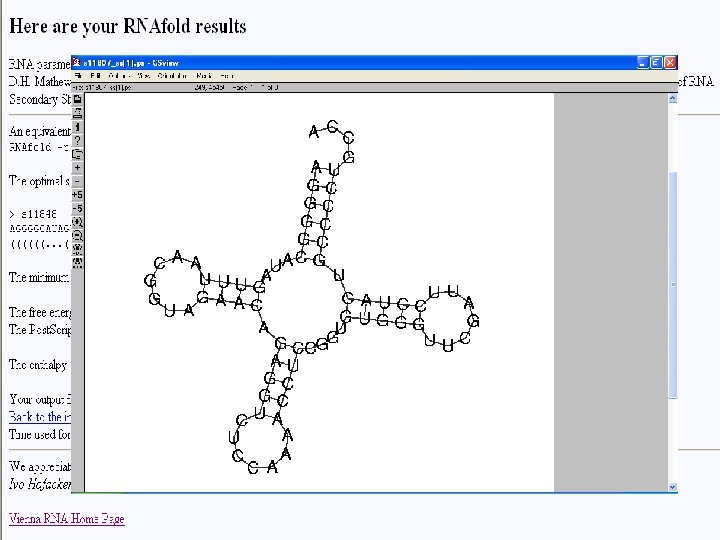
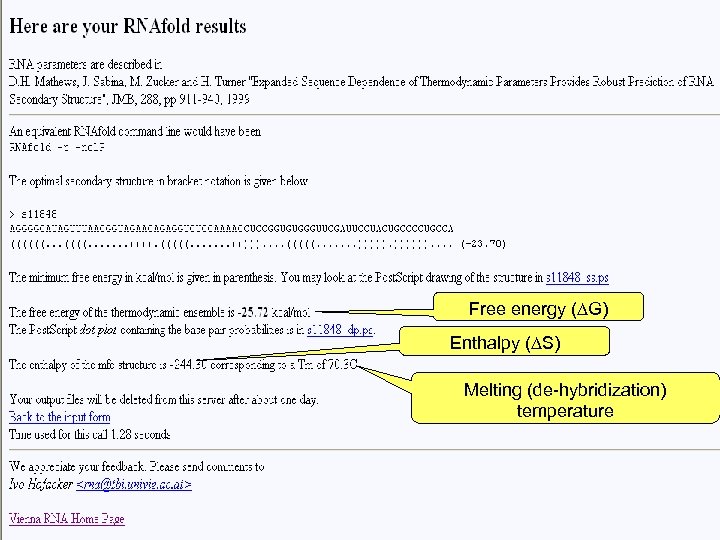
Free energy (∆G) Enthalpy (∆S) Melting (de-hybridization) temperature
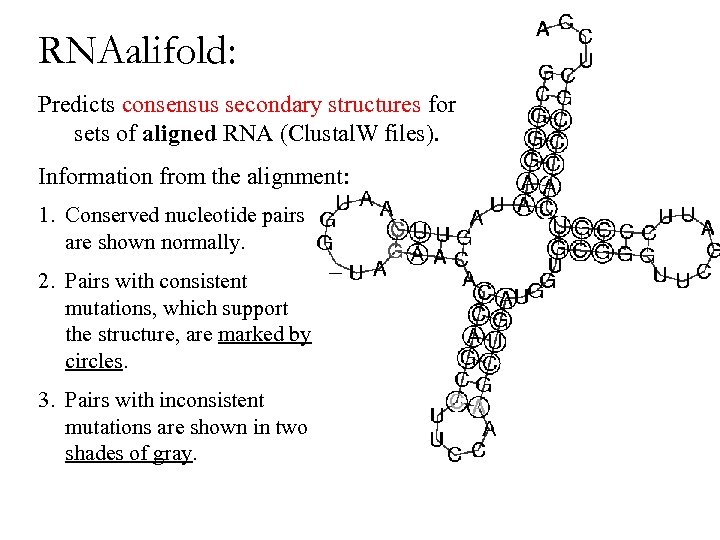
RNAalifold: Predicts consensus secondary structures for sets of aligned RNA (Clustal. W files). Information from the alignment: 1. Conserved nucleotide pairs are shown normally. 2. Pairs with consistent mutations, which support the structure, are marked by circles. 3. Pairs with inconsistent mutations are shown in two shades of gray.
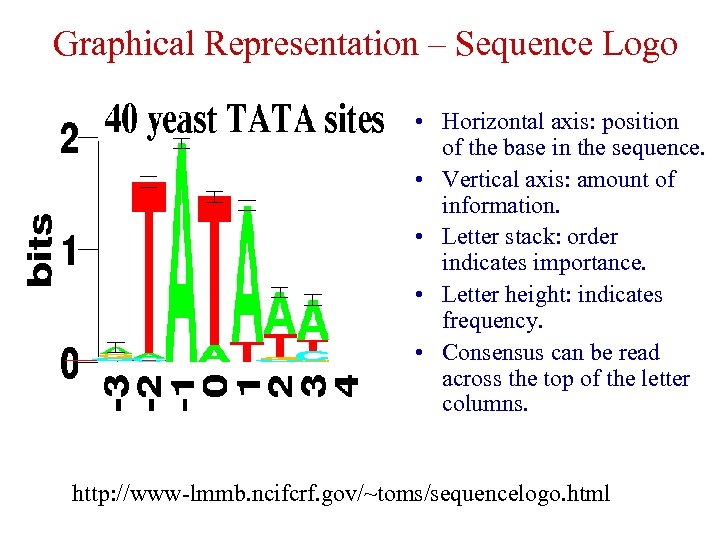
Graphical Representation – Sequence Logo • Horizontal axis: position of the base in the sequence. • Vertical axis: amount of information. • Letter stack: order indicates importance. • Letter height: indicates frequency. • Consensus can be read across the top of the letter columns. http: //www-lmmb. ncifcrf. gov/~toms/sequencelogo. html

Tools on the Web for motifs • MEME – Multiple EM for Motif Elicitation. http: //meme. sdsc. edu/meme/website/ • meta. MEME- Uses HMM method http: //meme. sdsc. edu/meme • MAST-Motif Alignment and Search Tool http: //meme. sdsc. edu/meme • TRANSFAC - database of eukaryotic cis-acting regulatory DNA elements and trans-acting factors. http: //transfac. gbf. de/TRANSFAC/ • e. Motif - allows to scan, make and search for motifs in the protein level. http: //motif. stanford. edu/emotif/

Websites for Promoter finding Promoter Scan: NIH Bioinformatics (BIMAS) http: //bimas. dcrt. nih. gov/molbio/proscan/ Promoter Scan II: Univ. of Minnesota & Axyx Pharmaceuticals http: //biosci. cbs. umn. edu/software/proscan/promoterscan. htm Signal Scan: NIH Bioinformatics (BIMAS) http: //bimas. dcrt. nih. gov: 80/molbio/signal/index. html Transcription Element Search (TESS): Center for Bioinformatics, Univ. of Pennsylvania http: //www. cbil. upenn. edu/tess/ Search Trans. Fac at GBF with Mat. Inspector, Pat. Search, and Fun. Site. P http: //transfac. gbf-braunschweig. de/TRANSFAC/programs. html Target. Finder: Telethon Inst. of Genetics and Medicine, Milan, Italy http: //hercules. tigem. it/Target. Finder. html

Transcriptional regulatory region TFs play a significant role in differentiation in a number of cell types The fact that ~ 5% of the genes are predicted to encode transcription factors underscores the importance of transcriptional regulation in gene expression (Tupler et al. 2001 Nature. 409: 832 -833) The combinatorial nature of transcriptional regulation and practically unlimited number of cellular conditions significantly complicate the experimental identification of TF binding sites on a genome scale Understanding the transcriptional regulation is a major challenge Computational approaches to identify potential regulatory elements and modules, and derive new, biologically relevant and testable hypothesis
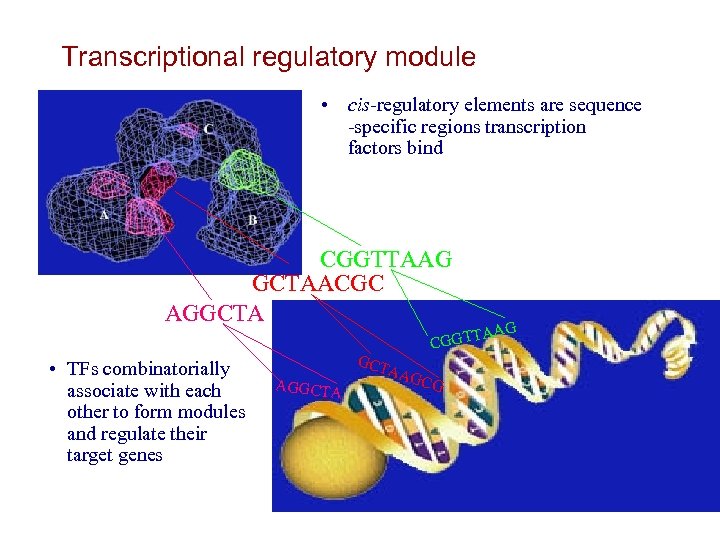
Transcriptional regulatory module • cis-regulatory elements are sequence -specific regions transcription factors bind CGGTTAAG GCTAACGC AGGCTA • TFs combinatorially associate with each other to form modules and regulate their target genes GCT AGGCTA AG TA CGGT AAG CG
Mammalian Promoter Dadabase (MProm. Db) (http: //bioinformatics. med. ohio-state. edu)
MProm. Db 1. 0 (Mammalian Promoter Database) (http: //bioinformatics. med. ohio-state. edu) Human, mouse & rat Search by gene symbol; Genbank Acc. Num; Unigene/Locus. Link ID; TF binding site name Click here to search the database
MProm. Db 1. 0 (Mammalian Promoter Database) (http: //bioinformatics. med. ohio-state. edu)
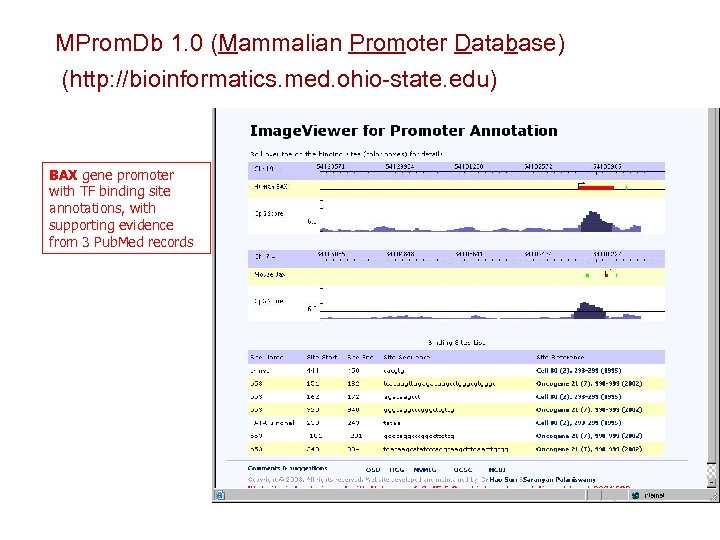

MProm. Db 1. 0 (Mammalian Promoter Database) (http: //bioinformatics. med. ohio-state. edu) BAX gene promoter with TF binding site annotations, with supporting evidence from 3 Pub. Med records

MProm. Db (Mammalian Promoter Database) Promoter sequences with annotations of experimentally supported TF binding sites Promoter sequences with annotations of computationally predicted TF binding sites A platform for statistical analysis & pattern recognition, to predict TF binding sites in uncharacterized promoters, and model combinatorial association of TF binding sites A platform for comparative genomics, to reveal conserved regions across genomes of different species Identification of core-promoters Identification of all the human-mouse-rat homologues pairs Modeling and identification of TF binding sites & modules

Specific databases of protein sequences and structures § Swissprot § PIR § TREMBL (translated from DNA) § PDB (Three Dimensional Structures)
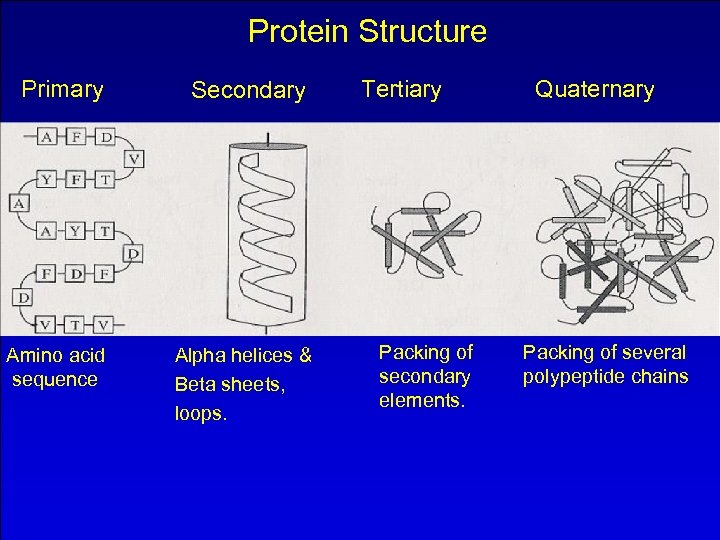
Protein Structure Primary Amino acid sequence Secondary Alpha helices & Beta sheets, loops. Tertiary Packing of secondary elements. Quaternary Packing of several polypeptide chains

Structure Prediction: Motivation • Hundreds of thousands of gene sequences translated to proteins (genbanbk, SW, PIR) • Only about 28000 solved structures (PDB) • Goal: Predict protein structure based on sequence information

Structure Prediction: Motivation • Understand protein function – Locate binding sites • Broaden homology – Detect similar function where sequence differs • Explain disease – See effect of amino acid changes – Design suitable compensatory drugs

Prediction Approaches • Primary (sequence) to secondary structure – Sequence characteristics • Secondary to tertiary structure – Fold recognition – Threading against known structures • Primary to tertiary structure – Ab initio modelling

Secondary structure prediction • • • • AGADIR - An algorithm to predict the helical content of peptides APSSP - Advanced Protein Secondary Structure Prediction Server GOR - Garnier et al, 1996 HNN - Hierarchical Neural Network method (Guermeur, 1997) Jpred - A consensus method for protein secondary structure prediction at University of Dundee JUFO - Protein secondary structure prediction from sequence (neural network) nn. Predict - University of California at San Francisco (UCSF) Predict. Protein - PHDsec, PHDacc, PHDhtm, PHDtopology, PHDthreader, Max. Hom, Eval. Sec from Columbia University Prof - Cascaded Multiple Classifiers for Secondary Structure Prediction PSA - Bio. Molecular Engineering Research Center (BMERC) / Boston PSIpred - Various protein structure prediction methods at Brunel University SOPMA - Geourjon and Del י age, 1995 SSpro - Secondary structure prediction using bidirectional recurrent neural networks at University of California DLP - Domain linker prediction at RIKEN
http: //searchlauncher. bcm. tmc. edu/

Multiple Sequence Alignment

Clustal. W Algorithm Progressive Sequences Alignment (Higgins and Sharp 1988) • Compute pairwise alignment for all the pairs of sequences. • Use the alignment scores to build a phylogenetic tree such that • similar sequences are neighbors in the tree • distant sequences are distant from each other in the tree. • The sequences are progressively aligned according to the branching order in the guide tree. • http: //www. ebi. ac. uk/clustalw/
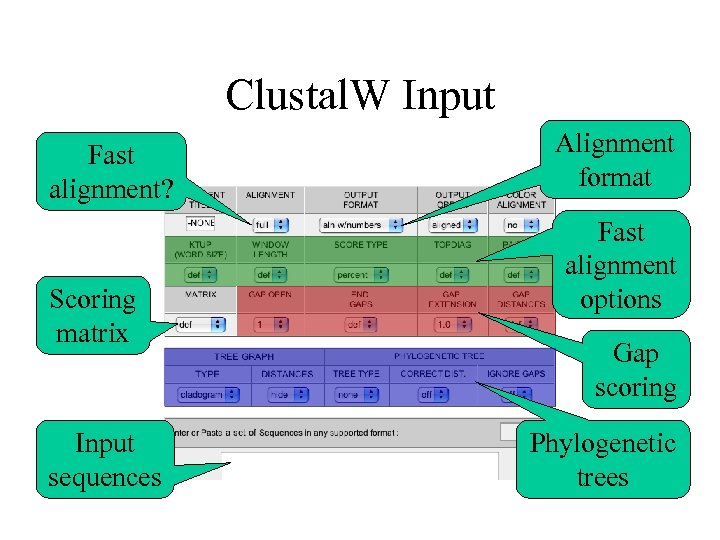
Clustal. W Input Fast alignment? Scoring matrix Input sequences Alignment format Fast alignment options Gap scoring Phylogenetic trees
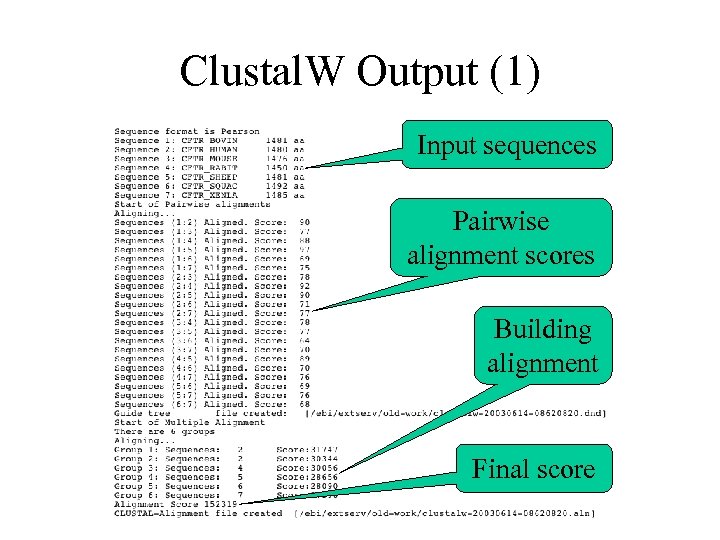
Clustal. W Output (1) Input sequences Pairwise alignment scores Building alignment Final score
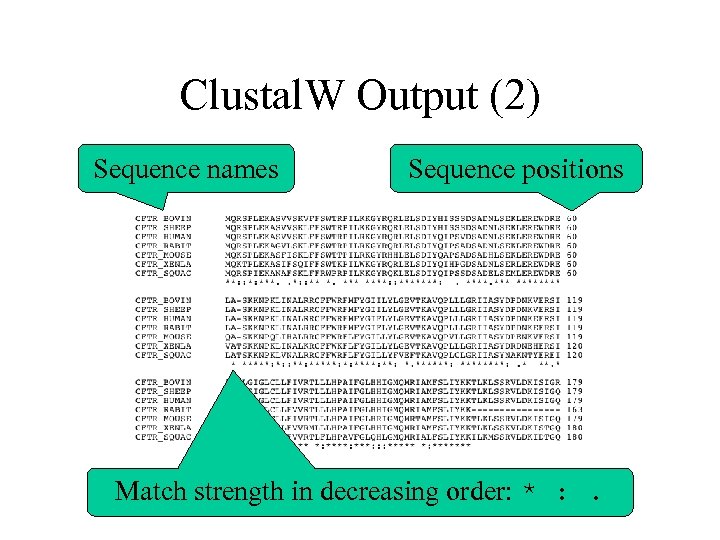
Clustal. W Output (2) Sequence names Sequence positions Match strength in decreasing order: * : .
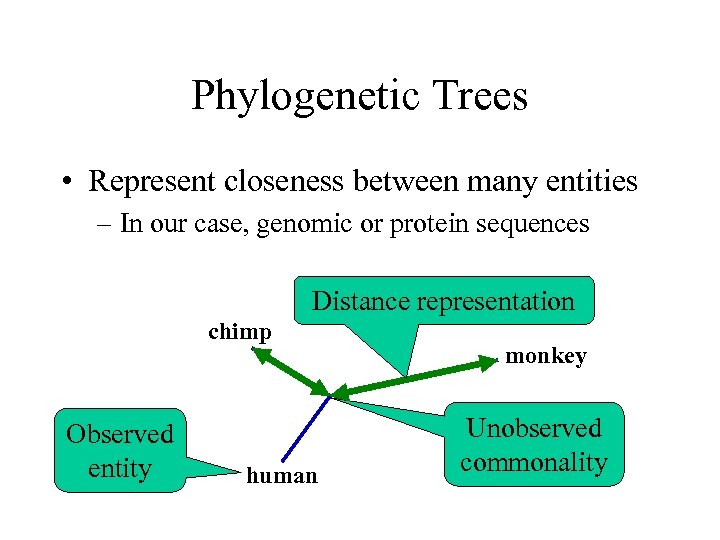
Phylogenetic Trees • Represent closeness between many entities – In our case, genomic or protein sequences Distance representation chimp Observed entity human monkey Unobserved commonality
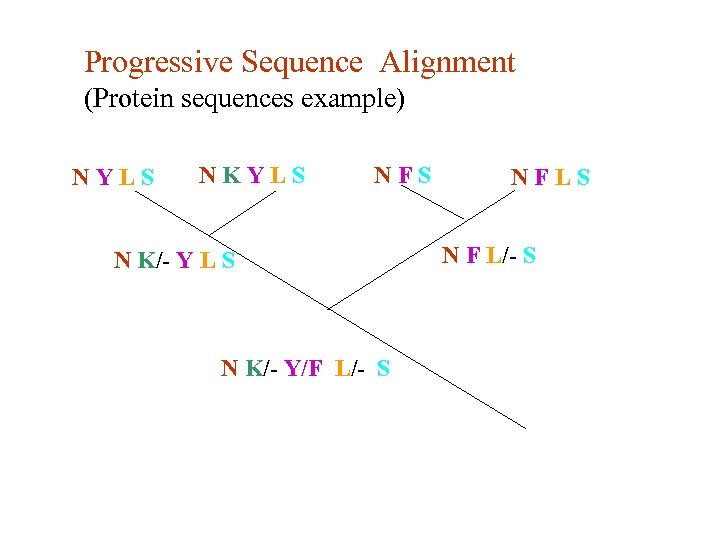
Progressive Sequence Alignment (Protein sequences example) NYLS NKYLS NFS N K/- Y L S N K/- Y/F L/- S NFLS N F L/- S

MSA Approaches • Progressive approach CLUSTALW (CLUSTALX) PILEUP T-COFFEE • Iterative approach: Repeatedly realign subsets of sequences. Mult. Alin, Di. Align. • Statistical Methods: Hidden Markov Models SAM 2 K • Genetic algorithm SAGA

Multiple Alignment tools on the Web (Some URLs) u. EMBL-EBI http: //www. ebi. ac. uk/clustalw/ u. BCM Search Launcher: Multiple Alignment http: //dot. imgen. bcm. tmc. edu: 9331/multi-align/multialign. html u. Multiple Sequence Alignment for Proteins (Wash. U. St. Louis) http: //www. ibc. wustl. edu/service/msa/

Editing Multiple Alignments • There a variety of tools that can be used to modify a multiple alignment. • These programs can be very useful in formatting and annotating an alignment for publication. • An editor can also be used to make modifications by hand to improve biologically significant regions in a multiple alignment created by one of the automated alignment programs.


GCG alignment editors • Alignments produced with PILEUP (or CLUSTAL) can be adjusted with LINEUP. • Nicely shaded printouts can be produced with PRETTYBOX • GCG's Seq. Lab X-Windows interface has a superb multiple sequence editor - the best editor of any kind.
Seq. Vu

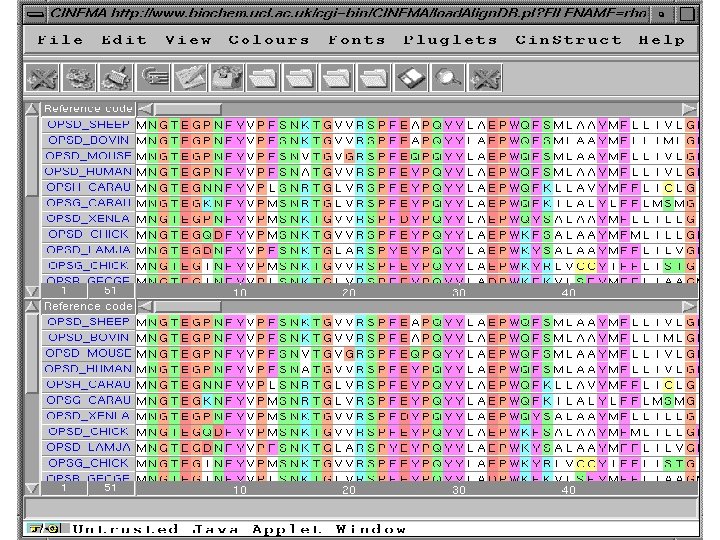
Editors on the Web • Check out CINEMA (Colour INteractive Editor for Multiple Alignments) – It is an editor created completely in JAVA (old browsers beware) – It includes a fully functional version of CLUSTAL, BLAST, and a Dot. Plot module http: //www. bioinf. man. ac. uk/dbbrowser/CINEMA 2. 1/

Questions: 1. Download protein seq and predict domain, secondary structure and post-translational modifications. 2. Download all SARS virus genome and perform MSA.
56630590bb155e0bc8c261895e812503.ppt