9281e73d162ca147d6a584a83319012a.ppt
- Количество слайдов: 58

Semantic Web Data Interoperation in Support of Enhanced Information Retrieval and Datamining in Proteomics Andrew Smith Thesis Defense 8/25/2006 Committee: Martin Schultz, Mark Gerstein (co-advisors), Drew Mc. Dermott, Steven Brenner (UC Berkeley)

Outline • Problem Description • • Enhanced information retrieval, datamining Link. Hub – supporting system • • Biological identifiers and their relationships. Semantic web RDF graph, RDF query languages Cross-database queries • Combined relational / keyword-based search • Enhanced automated information retrieval • • The web • Pub. Med (biomedical scientific literature) • Empirical performance evaluation for yeast proteins • Related Work and Conclusions

Web Information Management and Access – Opposing Paradigms Ø Search Engines over the web l l l Ø Automated People can publish in natural languages flexible Vast coverage (almost whole web) Currently, preeminent paradigm because of vast size of web and its unstructured heterogeneity Unfortunately, only gives coarse-grained topical access, no real cross-site interoperation / analysis Semantic Web l l l Very fine-grained data modeling and connection Very precise cross-resource query / question answering supported Unfortunately, requires much more manual intervention, people must change how they publish (RDF, OWL, etc. ) Thus, limited acceptance and size

Combining Search and Semantic Web Paradigms Ø Two paradigms largely independent. Ø Seem to have complementary strengths and weaknesses. Ø Key idea: These two approaches to web information management and retrieval can work together and complement one another and there are interesting, practical, and useful ways they can work with, leverage, and enhance each other.

Combining Relational and Keyword-based Search Access to Free Text Documents Ø Consider query for “all “ documents containing information for proteins which are members of the Pfam Adenylate Kinase (ADK) family” Ø Standard keyword-based search engines couldn’t support this. Ø Relational information about documents required, i. e. that they are related to particular proteins in the Pfam ADK family.

Using Semantic Web for Enhanced Automated Information Retrieval Ø Basic idea: the semantic web provides detailed information about terms and their interrelationships which can be used as additional information to improve web searches for those terms (and related terms). Ø As proof of concept, the particular, practical problem we are addressing is to find additional relevant documents for proteomics identifiers on the web or in the scientific literature.

Finding Additional Relevant Documents for Proteomics Identifiers Ø Why not just do a web search directly for the identifier, e. g. ‘P 26364’? likely not good result l l l Conflated senses of the identifier text, e. g. product catalog codes, etc. Might be synonyms of the identifier should search these too (semantic web could provide them). Many important, relevant documents might not directly mention the identifier • E. g. pages about ‘cancer pathways’ but not specifically containing cancer pathway related protein identifier • Should search for important related concepts. l Potentially much extra info from semantic web use it!

Example Cross-database Queries Ø For yeast protein interactions, find corresponding, evolutionarily related (homologous) protein interactions in Worm l Requires 4 databases: Uni. Prot, SGD, PFAM, Worm. Base Ø Explore pseudogenes for yeast essential genes; Explore pseudogenes of human homologues of yeast essential genes l Requires 4 databases: Pseudo. Gene, SGD, Uni. Prot, PFAM

Characteristics of Biological Data Biological data, especially proteomics data, is the motivation and domain of focus. Ø Vast quantities of data from high throughput experiments (genome sequencing, structural genomics, microarray, etc. ) Ø Huge and growing number of biological data resources: distributed, heterogeneous, large size variance. Ø l l Ø Practical domain to work in - need for better interoperation Challenging but not overly complex. Rich semantic relationships among data resources, but often not made explicit.

Problem Description Ø Integration and interoperation of structured, relational data (particularly proteomics data) that is: l l l Large-scale Widely distributed Independently maintained Ø In support of important applications: l l Enhanced information retrieval / web search Cross-database queries for datamining

Data Heterogeneity Lack of standard detailed description of resources Ø Data are exposed in different ways Ø l l l Ø Programmatic interfaces Web forms or pages FTP directory structures Data are presented in different ways l l l Structured text (e. g. , tab delimited format and XML format) Free text Binary (e. g. , images)

Classical Approaches to Interoperation: Data Warehousing and Federation Ø Data Warehousing l l Focuses on data translation Translate data to common format, under unified schema, cross-reference, store and query in single machine/system. Ø Federation l l Focuses on query translation Translating and distributing the parts of a query across multiple distinct, distributed databases and collating their results into single result.

General Strategy for Interoperation Ø Data is vast, distributed, independently maintained l Complete centralized data-warehousing integration is impractical Ø Must rely on federated, cooperative, loosely coupled solutions which allow partial and incremental progress l Widely used and supported standards necessary Ø Semantic Web is excellent fit to these needs

Semantic Web Resource Description Framework (RDF) Models Data as a Directed Graph with Objects and Relationships Named by URI http: //www. ncbi. nlm. nih. gov/SNP http: //purl. org/dc/elements/1. 1/creator http: //www. ncbi. nlm. nih. gov http: //purl. org/dc/elements/1. 1/language en <? xml version="1. 0"? > <rdf: RDF xmlns: rdf=“http: //www. w 3. org/1999/02/22 -rdf-syntax-ns#” xmlns: dc=“http: //purl. org/dc/elements/1. 1” xmlns: ex=“http: //www. example. org/terms”> <rdf: Description about=“http: //www. ncbi. nlm. nih. gov/SNP”> <dc: creator rdf: resource=“http: //www. ncbi. nlm. nih. gov”></dc: creator> <dc: language>en</dc: language> date> </rdf: Description> </rdf: RDF> Above RDF graph in XML
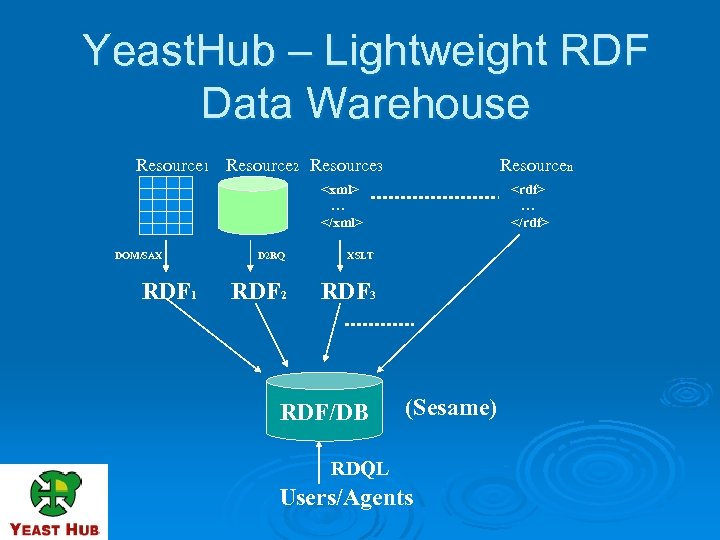
Yeast. Hub – Lightweight RDF Data Warehouse Resource 1 Resource 2 Resource 3 Resourcen <xml> … </xml> DOM/SAX RDF 1 D 2 RQ XSLT RDF 2 <rdf> … </rdf> RDF 3 RDF/DB (Sesame) RDQL Users/Agents

Name / ID proliferation problem Identifiers for biological entities are a simple but key way to identify and interrelate the entities; Important “scaffold” for biological data. Ø But often many synonyms for same entity, e. g. strange, legacy names; e. g. fly gene called “sonic hedgehog”. Ø Even simple syntactic variants can be cumbersome, e. g. GO: 0008150 vs. GO-8150, etc. Ø Not just synonyms, many kinds of relationship, one-to-many mappings. Ø Known relationships among entities not always stored, or stored in non-standard ways. Ø Implicit overall data structure: enormous, elaborate graph of relationships among biological entities. Ø
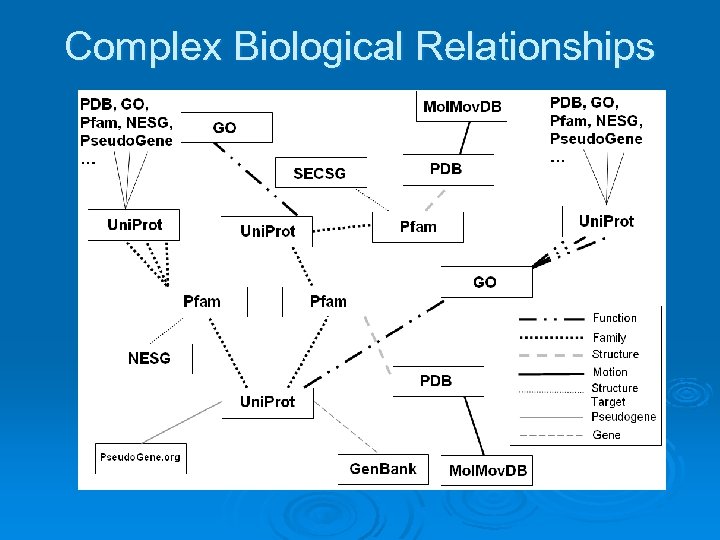
Complex Biological Relationships

Link. Hub Ø The major proteomics hub Uni. Prot performs centralized identifier mapping for large, well-known databases. l l l Large staff, resource intensive, manual curation centralization bottleneck. Not viable as complete solution Just simple mappings, no relationship types. Many data resources not covered (e. g. , small, transient, local, lab -specific, “boutique”). Need for system / toolkit to enable local, collaborative integration of data allow people to create “mini Uni. Prots” and connect them Link. Hub. Ø Practically, Link. Hub provides common “links portal” into a lab’s resources, also connecting them to larger resources. Ø l Link. Hub used this way for Gerstein Lab and NESG, connecting them to major proteomics hub Uni. Prot.
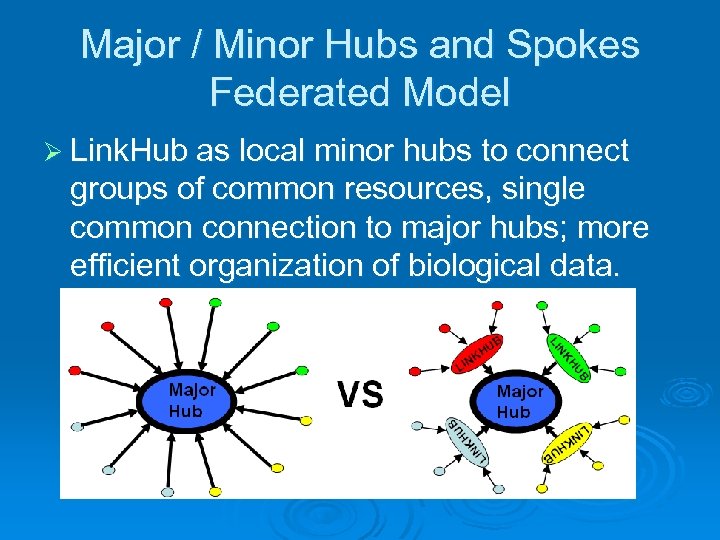
Major / Minor Hubs and Spokes Federated Model Ø Link. Hub as local minor hubs to connect groups of common resources, single common connection to major hubs; more efficient organization of biological data.
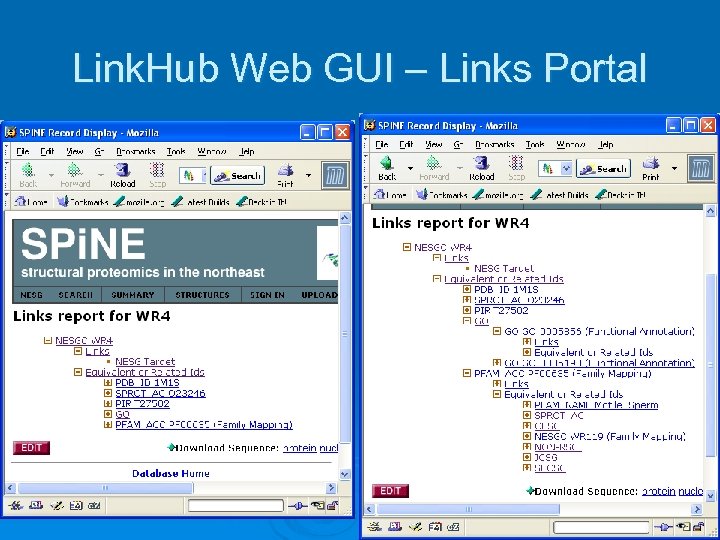
Link. Hub Web GUI – Links Portal

Example Yeast. Hub / Link. Hub queries Ø Query 1: Finding Worm ‘Interologs’ of Yeast Protein Interactions: l Ø For each yeast gene in interacting pair find corresponding Worm. Base genes: yeast gene name Uni. Prot Accession Pfam accession Uni. Prot Accession Worm. Base ID Query 2: Exploring Pseudogene Content versus Gene Essentiality in Yeast and Humans l l yeast gene name Uni. Prot Accession yeast pseudogene yeast gene name Uni. Prot Accession Pfam accession human Uni. Prot Id Uni. Prot Accession Pseudogene LSID

Combining Relational and Keyword-based Search Access to Free Text Documents Consider query for “all documents containing “ information for proteins which are members of the Pfam Adenylate Kinase (ADK) family” Ø Standard keyword-based search engines couldn’t support this. Ø Relational information about documents required, i. e. that they are related to proteins in the Pfam ADK family. Ø Link. Hub attaches documents to identifier nodes and supports such relational query access to them. Ø

Link. Hub Path Type Queries Ø View all paths in Link. Hub graph matching specific relationship types, e. g. “family views”: l l PDB ID Uni. Prot ID Pfam family Uni. Prot ID PDB ID Mol. Mov. DB Motion NESG ID Uni. Prot ID Pfam family Uni. Prot ID NESG ID Ø Practically used as secondary, orthogonal interface to other databases. l Mol. Mov. DB and NESG’s SPINE both use Link. Hub for such “family views”.

Link. Hub Subgraphs as Gold-standard Training Sets for Enhanced Automated Information Retrieval The Link. Hub subgraph emanating from a given identifier and the web pages (hyperlinks) attached to the identifiers in the subgraph is concrete, accurate, extra information about the given identifier that can be used to improve document retrieval for the given central identifier. Ø Link. Hub subgraphs and associated documents for a given identifier are used as a training set to build classifiers to rank documents obtained from the web or scientific literature. Ø
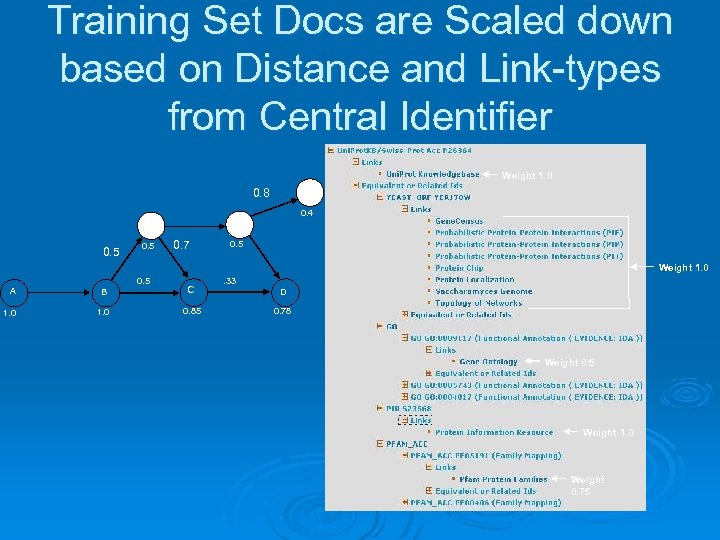

Training Set Docs are Scaled down based on Distance and Link-types from Central Identifier Weight 1. 0 0. 8 G 0. 4 F E 0. 5 0. 7 0. 5 Weight 1. 0 0. 5 A B C 1. 0 0. 85 . 33 D 0. 78 Weight 0. 5 Weight 1. 0 Weight 0. 75
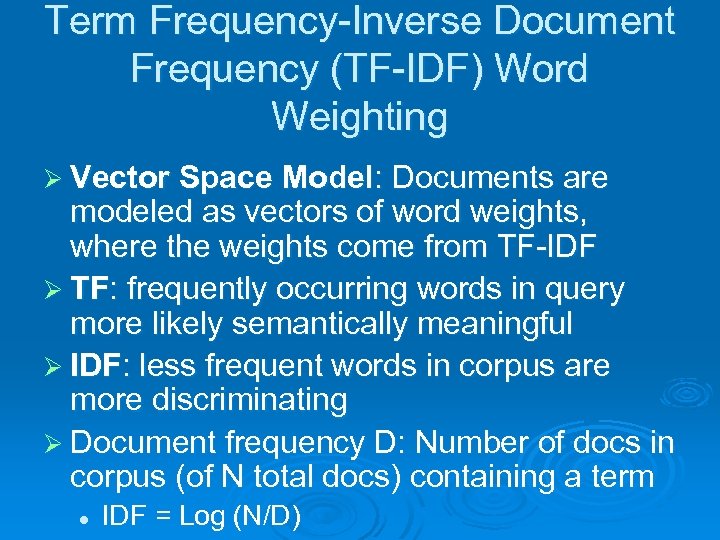

Term Frequency-Inverse Document Frequency (TF-IDF) Word Weighting Ø Vector Space Model: Documents are modeled as vectors of word weights, where the weights come from TF-IDF Ø TF: frequently occurring words in query more likely semantically meaningful Ø IDF: less frequent words in corpus are more discriminating Ø Document frequency D: Number of docs in corpus (of N total docs) containing a term l IDF = Log (N/D)

Classifier for Document Relevance Reranking Ø Use standard information retrieval techniques: tokenization, stopword filtering, word stemming, TF-IDF term weighting, and cosine similarity measures. Ø Classifier model: add weighted subgraph documents’ word vectors, TF-IDF weight it, and take top weighted 20% terms. Ø Standard cosine similarity value used to score documents against classifier.

Obtaining Documents to Rank Use major web search engines, via their web APIs. For demo purposes, we used Yahoo. Ø Perform individual, base searches Ø l l l Top 40 training set feature words Identifiers in the subgraph Combine all results into one large result set. Rerank combined result set using the constructed classifier. Ø Essentially, systematically exploring “concept space” around the identifier. Ø Searches returning most relevant docs on average could be called semantic signatures Ø l l Key concepts related to the given identifier Succinct snippets of what the identifier is “about”.
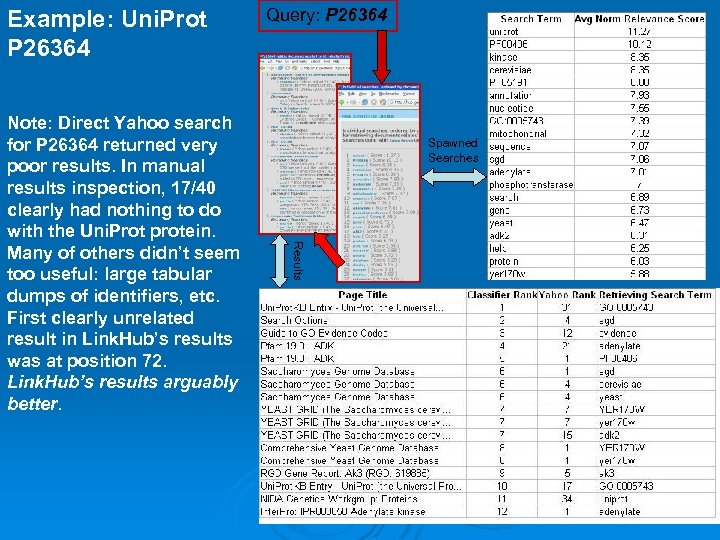
Example: Uni. Prot P 26364 Spawned Searches Results Note: Direct Yahoo search for P 26364 returned very poor results. In manual results inspection, 17/40 clearly had nothing to do with the Uni. Prot protein. Many of others didn’t seem too useful: large tabular dumps of identifiers, etc. First clearly unrelated result in Link. Hub’s results was at position 72. Link. Hub’s results arguably better. Query: P 26364

Pub. Med Application Ø Pub. Med is a database of all scientific literature citations for about the last 50 -100 years. Ø Currently, no automated information retrieval of Pub. Med for biological identifierrelated citations. Ø Built app to search for related Pub. Med abstracts, using Swish-e to index and provide base search access to Pub. Med.
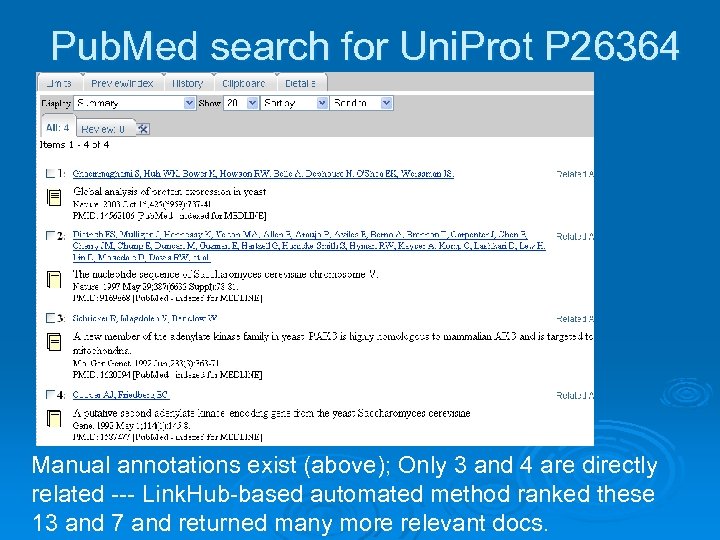
Pub. Med search for Uni. Prot P 26364 Manual annotations exist (above); Only 3 and 4 are directly related --- Link. Hub-based automated method ranked these 13 and 7 and returned many more relevant docs.

Empirical Performance Tests Ø Preceding results seemed reasonably good, but can we empirically measure performance? Ø Use gold-standard set of documents (curated bibliography) known related to particular identifiers l gene_literature. tab from yeast genome database (SGD)

Goals of Performance Tests Ø Quantify the performance level of the procedure l Performance close to optimal or lot’s of room for improvement? Ø How can we know that adding in documents (downweighted) for related identifiers actually helps? l Proof of concept: PFAM and GO are key related concepts for proteins, let’s objectively see if they help. Ø Quantify performance of a particular enhancement: pre-IDF step.

Pre-Inverse Document Frequency (pre-IDF) step Ø Idea: maximally separate all proteomics identifiers’ classifiers while making them specifically relevant and discriminating. Ø Determine document frequencies for all pages of a type, e. g. all or sample of Uni. Prot pages. Ø Perform IDF against the type’s doc freqs and then against the corpus you are searching; e. g. first Uni. Prot then Pub. Med

Pre-IDF Step is Generally Useful Ø For example, imagine wanting to find web pages highly relevant to a particular digital camera or mp 3 player. Ø Cnet has many pages about different digital cameras or mp 3 players doc freqs for these. Ø Build classifier for particular digital camera by first doing pre-IDF step against doc freqs for all Cnet digital camera pages.

Experimental Protocol Ø Pick few hundred random Yeast proteins from Tr. EMBL and Swiss-Prot separately Each with at least 20 citations and GO and PFAM relations. Ø A protein’s citations are its “in” group l Other protein’s citations are “mid” group Ø Randomly selected Pub. Med citations (not in gene_lit. tab or Uni. Prot) are an “out” group Ø Classifier should match “in” – “mid” – “out” order Ø l l But focus on “in”, most important Degree of deviation from this is the objective test
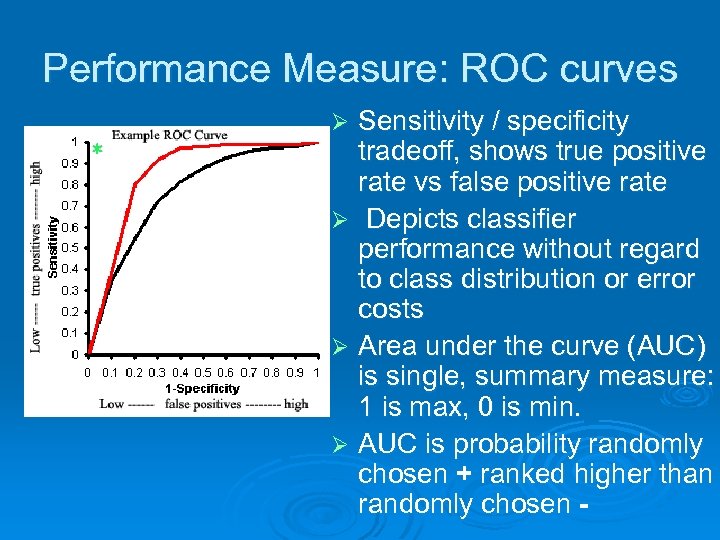
Performance Measure: ROC curves Sensitivity / specificity tradeoff, shows true positive rate vs false positive rate Ø Depicts classifier performance without regard to class distribution or error costs Ø Area under the curve (AUC) is single, summary measure: 1 is max, 0 is min. Ø AUC is probability randomly chosen + ranked higher than randomly chosen Ø

Measuring Performance Ø Full AUC and top 5% AUC (. 05 AUC) l Ø It is how well you do in the top of rankings that really matters. Uni. Prot page weight set to 1. 0. Perform optimization over parameters at 0. 1 granularity: l l l PFAM and GO weights Percentage of features kept Use/not use pre-IDF step. Compare average AUC values for various parameter values Ø Determine statistical significance with paired ttests. Ø

Results Ø Perc features kept didn’t really matter l Ø Pre-IDF gave largest performance boost l Ø GO did not help (PFAM more info rich). Tr. EMBL was helped more than Swiss-Prot l Ø Any trial with pre-IDF gave better result than one without, regardless of other parameters. Pre-IDF and PFAM at small weight increased avg AUC for all trials for both Tr. EMBL and Swiss -Prot. l Ø 0. 4 or 0. 5 for computational efficiency Swiss. Prot is curated, high quality, complete, whereas Tr. EMBL is automated, lower quality makes sense All comparisons for Tr. EMBL were statistically significant by t-tests, only pre-IDF for Swiss-Prot.
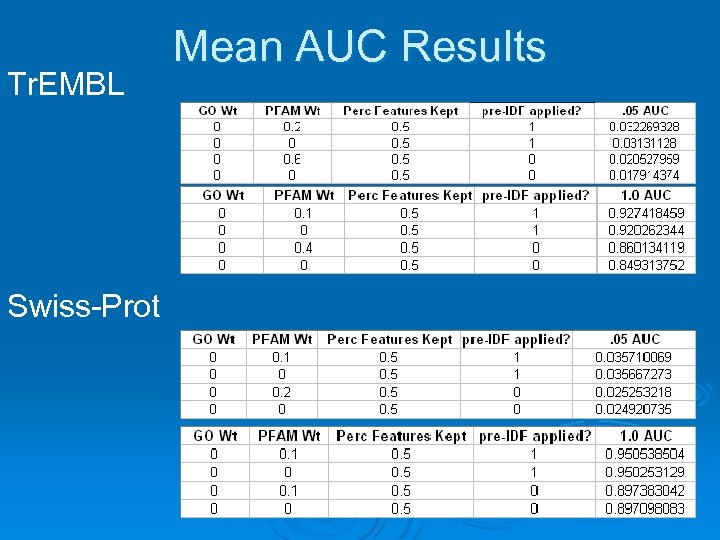
Tr. EMBL Swiss-Prot Mean AUC Results

Related Work Search Engines / Google Ø Small number of search terms entered. l l Ø So little information to go by maybe millions of result documents, hard to rank well. Good ranking was big problem for early search engines. Google (Brin and Page 1998) provided a popular solution l l Hyperlinks as “votes” for importance Rank web pages with most and best votes highest.

Alternative to Millions of Result Documents: Increased Search Precision Ø Increase search precision so fewer, more manageable number of documents returned. l More search terms Problem: users are lazy, won’t enter too many terms. Ø Solution: semantic web provides the increased precision users just select semantic web nodes for concepts, the nodes are automatically expanded to increase search precision. Ø l This is what Link. Hub does

Very Recent Related Work: Aphinyanaphongs et al 2006 Ø Argues for specialized, automated filters for finding relevant documents in huge and ever expanding scientific literature. Ø Constructed classifiers for predicting relevance of Pub. Med documents for various clinical medicine themes l l l State of the art SVM classifiers Used large, manually curated, respected bibliographies to train Used text from article title, abstract, journal name, and Me. SH terms for features

Aphinyanaphongs et al 2006 cont. Ø Link. Hub-based search by contrast: l l Fairly basic classifier model, word weight vectors compared with cosine similarity. Small training sets (Uni. Prot, GO, PFAM pgs) • Fairly noisy also: web pages vs focused text l l l Only abstract text used as features Some gene_lit. tab citations more generally relevant True performance understated Classifiers built automatically & easily at very large scale as natural byproduct of Link. Hub Ø But Link. Hub’s. 927 and. 951 AUCs better than or negligibly smaller than 0. 893, 0. 932, 0. 966 AUCs of Aphinyanaphongs et al 2006

Aphinyanaphongs et al 2006 and Citation Metrics Ø Also compared to citation-based metrics l l l Citation count Journal impact factor Indirectly to Google Page. Rank Ø SVM classifiers outperformed these, and adding citation metrics as features gave marginal improvement at best. Ø Surprising given Google’s success with Page. Rank

Aphinyanaphongs et al 2006 and Citation Metrics cont. More generally stated, the conceivable reasons for citation are so numerous that it is unrealistic to believe that citation conveys just one semantic interpretation. Instead citation metrics are a superimposition of a vast array of semantically distinct reasons to acknowledge an existing article. It follows that any specific set of criteria cannot be captured by a few general citation metrics and only focused filtering mechanisms, if attainable, would be able to identify articles satisfying the specific criteria in question. Conclusion: Aphinyanaphongs et al 2006 is consistent with and supports the general approach taken by Link. Hub of creating specialized filters (in the form of word weight vectors) for retrieval of documents specific to particular proteomics identifiers. It is arguably state of the art for focused tasks, superior to most commonly used search technology of Google; by extension, Link. Hub-based search is also.

Publications 1. 2. 3. 4. 5. 6. 7. Link. Hub: a Semantic Web System for Efficiently Handling Complex Graphs of Proteomics Identifier Relationships that Facilitates Cross-database Queries and Information Retrieval. Andrew K. Smith, Kei-Hoi Cheung, Kevin Y. Yip, Martin Schultz 1, Mark B. Gerstein. International Workshop on Semantic e-Science 3 rd September, 2006, Beijing, China. (Se. S 2006), co-located with ASWC 2006. To be published in special proceedings of BMC Bioinformatics. Yeast. Hub: a semantic web use case for integrating data in the life sciences domain. KH Cheung, KY Yip, A Smith, R Deknikker, A Masiar, M Gerstein (2005) Bioinformatics 21 Suppl 1: i 85 -96. An XML-Based Approach to Integrating Heterogeneous Yeast Genome Data. KH Cheung, D Pan, A Smith, M Seringhaus, SM Douglas, M Gerstein. 2004 International Conference on Mathematics and Engineering Techniques in Medicine and Biological Sciences (METMBS); pp 236 -242. Network security and data integrity in academia: an assessment and a proposal for large-scale archiving. A Smith, D Greenbaum, SM Douglas, M Long, M Gerstein (2005) Genome Biol 6: 119. Computer Security in academia-a potential roadblock to distributed annotation of the human genome. D Greenbaum, SM Douglas, A Smith, J Lim, M Fischer, M Schultz, M Gerstein (2004) Nat Biotechnol 22: 771 -2. Impediments to database interoperation: legal issues and security concerns. D Greenbaum, A Smith, M Gerstein (2005) Nucleic Acids Res 33: D 3 -4. Mining the structural genomics pipeline: identification of protein properties that affect high-throughput experimental analysis. CS Goh, N Lan, SM Douglas, B Wu, N Echols, A Smith, D Milburn, GT Montelione, H Zhao, M Gerstein (2004) J Mol Biol 336: 115 -30.

Acknowledgements Ø Committee: Martin Schultz, Mark Gerstein (co-advisors), Drew Mc. Dermott, Steven Brenner Ø Kei Cheung Ø Michael Krauthammer Ø Kevin Yip Ø Yale Semantic Web Interest Group Ø National Library of Medicine (NLM)
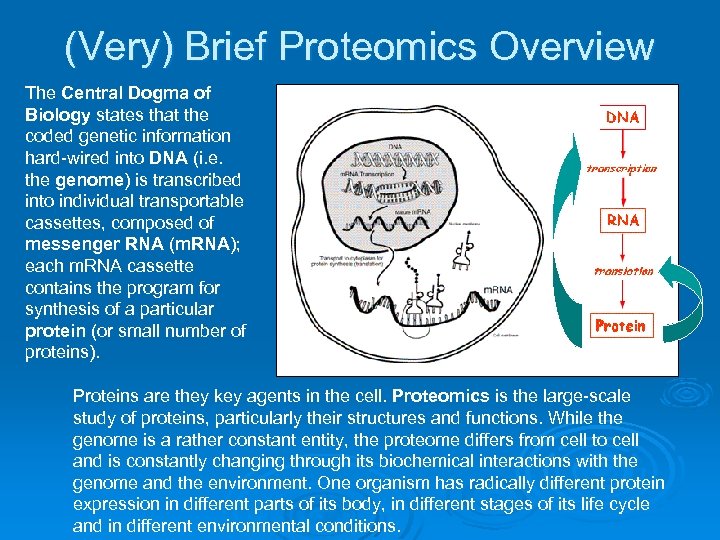
(Very) Brief Proteomics Overview The Central Dogma of Biology states that the coded genetic information hard-wired into DNA (i. e. the genome) is transcribed into individual transportable cassettes, composed of messenger RNA (m. RNA); each m. RNA cassette contains the program for synthesis of a particular protein (or small number of proteins). Proteins are they key agents in the cell. Proteomics is the large-scale study of proteins, particularly their structures and functions. While the genome is a rather constant entity, the proteome differs from cell to cell and is constantly changing through its biochemical interactions with the genome and the environment. One organism has radically different protein expression in different parts of its body, in different stages of its life cycle and in different environmental conditions.
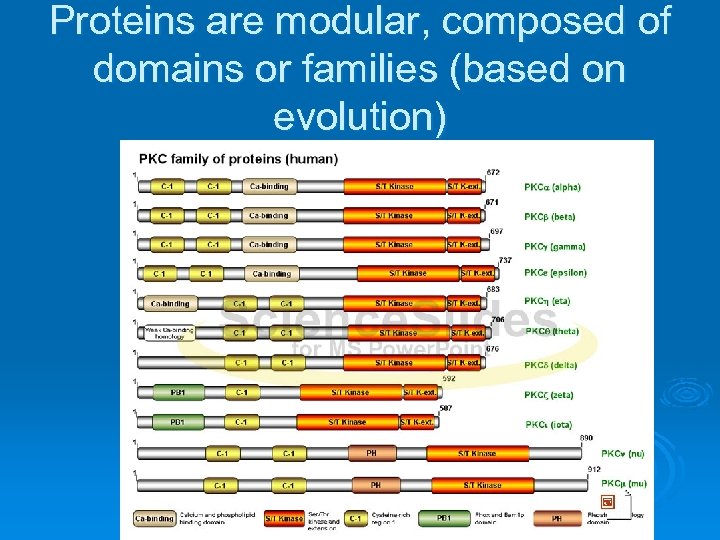
Proteins are modular, composed of domains or families (based on evolution)

Link. Hub Data Model Ø Link. Hub has identifier types, identifiers, mappings between identifiers (“mapping type” attribute gives relationship type), and resources and the identifier types they accept (and where) in their URL templates (link_exceptions gives exceptions). Ø Link. Hub stored in both RDF (Sesame) and SQL (My. SQL), translate between. My. SQL for robustness and efficiency for the GUI frontend, RDF to complement Yeast. Hub (as “glue” to make direct and indirect identifier connections).

Term Selection / Weighting using Term Associations in Corpus Lot’s of information in originating Uni. Prot, GO, PFAM, etc. pages and Pub. Med use it. Tune classifier to actual content in Pub. Med (word freqs). TF-IDF does this crudely, let’s do better. Ø Conceptually a "good" term is one that is strongly associated in the corpus (i. e. much above the background chance association in the corpus) with many other terms (which themselves have similar strong associations) from the originating documents. Ø Strength of association is ratio of doc freq in search results of a term, over the background doc freq in the corpus (Pub. Med). > 1 means association. Ø

Example associations in Pub. Med for term “browser”

Simple Algorithm for Term Selection & Weighting Ø Compute combined term freq vec for originating docs. l l For each term, do a search with it against Pub. Med and compute combined doc freq vector for result set. Filter terms which aren’t present at highly above background chance. Compute cosine similarity measure between these two vectors larger means more relevant. Ø In tests I’ve done, seems to do well, particularly for filtering noise terms (which almost always have a score close to 0).

Network analysis of Term Association Graph Can create directed graph of word interassociations; edges can be weighted by strength of association. Ø Do network analysis to aid term selection / weighting: Ø l l Find disconnected components independent concept groups. Find hubs likely central concepts in the originating documents, so good features. Find cliques groups of words that are all strongly co -associated with each other, the bigger the better. Google pagerank for term selection and weighting.

Example – Maximal Clique for Yeast Adenylate Kinase (Uni. Prot P 26364)
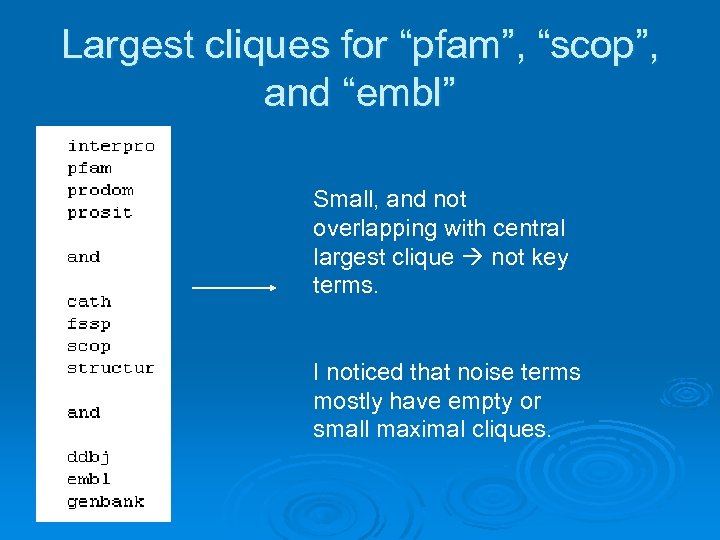

Largest cliques for “pfam”, “scop”, and “embl” Small, and not overlapping with central largest clique not key terms. I noticed that noise terms mostly have empty or small maximal cliques.

Google Page. Rank on term association graph. Actually, doesn’t work well. Seems to most highly weight relevant but more general terms (e. g. “cell”, “pfam”, etc. ) Ø In retrospect, makes sense: pagerank is trying to find nodes which are linked to by many other nodes and link out to many other nodes, and this is intuitively a signature of more general terms (i. e. they will have a larger, more diffuse set of assocations than more focused terms). Ø There are some results in the literature that Page. Rank applied to scientific citation graphs does not work well. Maybe Google really isn’t God! Ø Ø
9281e73d162ca147d6a584a83319012a.ppt