f84d5d8f8fe2c72448961d09a81092e3.ppt
- Количество слайдов: 73
 scripting inside Compu. Cell 3 D • XML gives you the ability to change simulation parameters using human-readable syntax but does not allow users to implement more complex cell behaviors, sophisticated cell type transition rules, inter-cell signaling or connecting to intracellular models • Python scripting capabilities in Compu. Cell 3 D allow users to accomplish abovementioned tasks (and much more) and are the reasons why Compu. Cell 3 D is called simulation environment, not simulation application. • Python scripting capabilities allow users to use rich set of Python modules and third party libraries and provide level flexibility comparable with packages such as Matlab or Mathematica
scripting inside Compu. Cell 3 D • XML gives you the ability to change simulation parameters using human-readable syntax but does not allow users to implement more complex cell behaviors, sophisticated cell type transition rules, inter-cell signaling or connecting to intracellular models • Python scripting capabilities in Compu. Cell 3 D allow users to accomplish abovementioned tasks (and much more) and are the reasons why Compu. Cell 3 D is called simulation environment, not simulation application. • Python scripting capabilities allow users to use rich set of Python modules and third party libraries and provide level flexibility comparable with packages such as Matlab or Mathematica
 Python Scripting Prerequisites • To make full use of Python scripting users should be familiar with Python programming language. They do not need to be experts though. • Compu. Cell 3 D comes with template and example codes that make Python scripting inside Compu. Cell 3 D quite easy even for beginners. • Python scripting in Compu. Cell 3 D typically requires users to develop a class that implements required logic. If you are unfamiliar with concept of class , think of it as a type that has several data members (such as floats, integers, other classes) and set of functions that operate on those internal members but can also take external arguments. Class is a generalization of “C” structure or “Pascal” record. • Fortunately Compu. Cell 3 D comes with plenty of examples that users can adapt to serve their needs. This does not require thorough programming knowledge. If you are unfamiliar with Python scripting, reading (and doing) “Compu. Cell 3 D Python Scripting Tutorials” should quickly get you up-to-speed.
Python Scripting Prerequisites • To make full use of Python scripting users should be familiar with Python programming language. They do not need to be experts though. • Compu. Cell 3 D comes with template and example codes that make Python scripting inside Compu. Cell 3 D quite easy even for beginners. • Python scripting in Compu. Cell 3 D typically requires users to develop a class that implements required logic. If you are unfamiliar with concept of class , think of it as a type that has several data members (such as floats, integers, other classes) and set of functions that operate on those internal members but can also take external arguments. Class is a generalization of “C” structure or “Pascal” record. • Fortunately Compu. Cell 3 D comes with plenty of examples that users can adapt to serve their needs. This does not require thorough programming knowledge. If you are unfamiliar with Python scripting, reading (and doing) “Compu. Cell 3 D Python Scripting Tutorials” should quickly get you up-to-speed.
 Typical example when Python proves to be very useful Start with a small cell that grows It reaches “doubling volume” … and divides into two cells After mitosis you want to specify types of parent and daughter cells. You may want to change target surface and target volume of daughter. And target volume is a function of a FGF concentration at the center of mass of the daughter cell. How would you do it from just XML? Python seems to be the best solution for problems like this one
Typical example when Python proves to be very useful Start with a small cell that grows It reaches “doubling volume” … and divides into two cells After mitosis you want to specify types of parent and daughter cells. You may want to change target surface and target volume of daughter. And target volume is a function of a FGF concentration at the center of mass of the daughter cell. How would you do it from just XML? Python seems to be the best solution for problems like this one
 Where Do You Begin? • Early version of Python scripting in Compu. Cell 3 D required users to provide CC 3 DML configuration file. This is no longer true. You can describe entire simulation from Python level. However, you may still use XML and Python if you want. The choice is up to you. • You will need to write Python script that implements main Compu. Cell 3 D logic i. e. reads CC 3 DML file (if you are using CC 3 DML file), initializes modules and executes calls in a loop Metropolis algorithm. This file will also call set up and initialize your modules written in Python. Compu. Cell 3 D comes with many examples of such files so in fact preparing one is reduced to minor modification of existing one. • Once you have Python script (and optionally CC 3 DML file) ready, you open them up in the Player and start running simulations.
Where Do You Begin? • Early version of Python scripting in Compu. Cell 3 D required users to provide CC 3 DML configuration file. This is no longer true. You can describe entire simulation from Python level. However, you may still use XML and Python if you want. The choice is up to you. • You will need to write Python script that implements main Compu. Cell 3 D logic i. e. reads CC 3 DML file (if you are using CC 3 DML file), initializes modules and executes calls in a loop Metropolis algorithm. This file will also call set up and initialize your modules written in Python. Compu. Cell 3 D comes with many examples of such files so in fact preparing one is reduced to minor modification of existing one. • Once you have Python script (and optionally CC 3 DML file) ready, you open them up in the Player and start running simulations.
 What Can You Do in Python? • You may implement any Compu. Cell 3 D module using Python – energy functions, lattice monitors, steppers, steppables, fixed steppers. • You need to remember that Python is an interpreted language and thus executes orders of magnitudes slower than, for example, C++. This means that although you can easily develop energy functions (remember, they are the most frequently called modules in Compu. Cell 3 D) in Python, you will probably want to avoid using them with your “production runs”. In this it makes sense to implement those functions in C++ , which is not too difficult and we provide comprehensive Developers documentation. • Since lattice monitors are called less frequently than energy functions, the performance degradation due to lattice monitor being implemented in Python is much smaller. That same is true for steppers, fixed steppers and steppables. • Notice that Compu. Cell 3 D kernel that is called from Python, as well as other core Compu. Cell 3 D modules called from Python run at native speeds, as they are implemented in C++ with only their API exposed to Python. Therefore if you run Compu. Cell 3 D through Python script but decide not to implement new Python modules, your speed of run will be essentially identical as if you ran Compu. Cell 3 D using just CC 3 DML file.
What Can You Do in Python? • You may implement any Compu. Cell 3 D module using Python – energy functions, lattice monitors, steppers, steppables, fixed steppers. • You need to remember that Python is an interpreted language and thus executes orders of magnitudes slower than, for example, C++. This means that although you can easily develop energy functions (remember, they are the most frequently called modules in Compu. Cell 3 D) in Python, you will probably want to avoid using them with your “production runs”. In this it makes sense to implement those functions in C++ , which is not too difficult and we provide comprehensive Developers documentation. • Since lattice monitors are called less frequently than energy functions, the performance degradation due to lattice monitor being implemented in Python is much smaller. That same is true for steppers, fixed steppers and steppables. • Notice that Compu. Cell 3 D kernel that is called from Python, as well as other core Compu. Cell 3 D modules called from Python run at native speeds, as they are implemented in C++ with only their API exposed to Python. Therefore if you run Compu. Cell 3 D through Python script but decide not to implement new Python modules, your speed of run will be essentially identical as if you ran Compu. Cell 3 D using just CC 3 DML file.
 What are the advantages of using Python inside Compu. Cell 3 D • Rapid development – no compilation is necessary. Write or modify your script and run • Portability – script developed by you on one machine (e. g. Mac) is ready to use under linux • Model integration - you can quite easily implement hooks to subcellular models. We have been able to set up Compu. Cell 3 D simulation that was using SBW network intracell simulators within few minutes. T • Rich set of external libraries – you may tap into rich Python library repositories that exist for essentially any task • Agile development – developing and refining your model is an iterative process. Working at the compiled language stage will force you to spend significant portion of your time waiting for the program to compile. With Python you eliminate this step thus increase productivity. Users should first prototype their models in Python and once they are ready to be used for production runs, rewrite the ones causing significant slowdown in C++.
What are the advantages of using Python inside Compu. Cell 3 D • Rapid development – no compilation is necessary. Write or modify your script and run • Portability – script developed by you on one machine (e. g. Mac) is ready to use under linux • Model integration - you can quite easily implement hooks to subcellular models. We have been able to set up Compu. Cell 3 D simulation that was using SBW network intracell simulators within few minutes. T • Rich set of external libraries – you may tap into rich Python library repositories that exist for essentially any task • Agile development – developing and refining your model is an iterative process. Working at the compiled language stage will force you to spend significant portion of your time waiting for the program to compile. With Python you eliminate this step thus increase productivity. Users should first prototype their models in Python and once they are ready to be used for production runs, rewrite the ones causing significant slowdown in C++.
 Your first Compu. Cell 3 D Python script. Make sure you have your copy of Python Scripting Tutorials Begin with template code (the file will be called cellsort_2 D. py) #import useful modules import sys from os import environ from os import getcwd import string #setup search patths sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") #add search path import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Your first Compu. Cell 3 D Python script. Make sure you have your copy of Python Scripting Tutorials Begin with template code (the file will be called cellsort_2 D. py) #import useful modules import sys from os import environ from os import getcwd import string #setup search patths sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") #add search path import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
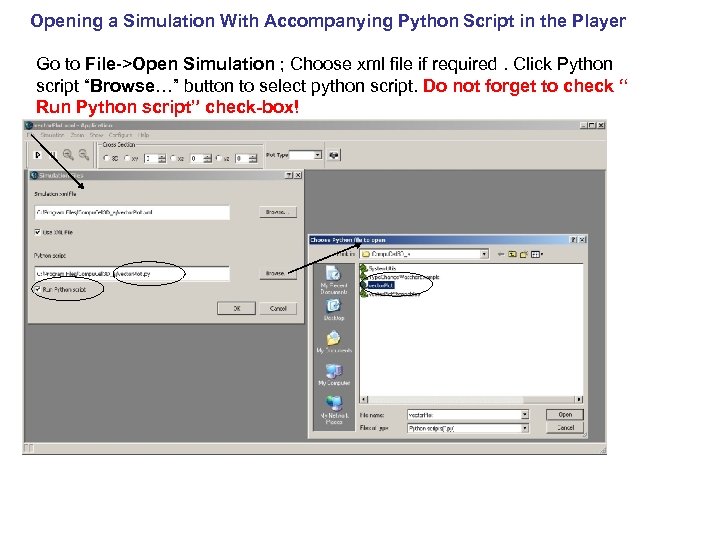 Opening a Simulation With Accompanying Python Script in the Player Go to File->Open Simulation ; Choose xml file if required. Click Python script “Browse…” button to select python script. Do not forget to check “ Run Python script” check-box!
Opening a Simulation With Accompanying Python Script in the Player Go to File->Open Simulation ; Choose xml file if required. Click Python script “Browse…” button to select python script. Do not forget to check “ Run Python script” check-box!
 Using Python to describe entire simulations • Starting with 3. 2. 0 versions you may get rid of XML file and use Python to describe entire simulation. • The advantage of doing so is that you have one less file to worry about but also you may more easily manipulate simulation parameters. For example if you want contact energy between two cell types be twice as big as between two other cell types you could easily implement it in Python. Doing the same exercise with CC 3 d. ML is a bit harder (but not impossible). • Python syntax used to describe simulation closely mimics CC 3 DML syntax. There are however certain differences and inconsistencies caused by the fact that we are using different languages to accomplish same task. Currently there is no documentation explaining in detail Python syntax that replaces CC 3 DML. It will be developed soon • The most important reason for defining entire simulation in Python is the possibility of simulation steering i. e. the ability to dynamically change simulation parameters while simulation is running (available in 3. 2. 1) • While there are certain disadvantages of this approach (mainly lack of good documentation) you get many benefits by using Python only to describe the simulation
Using Python to describe entire simulations • Starting with 3. 2. 0 versions you may get rid of XML file and use Python to describe entire simulation. • The advantage of doing so is that you have one less file to worry about but also you may more easily manipulate simulation parameters. For example if you want contact energy between two cell types be twice as big as between two other cell types you could easily implement it in Python. Doing the same exercise with CC 3 d. ML is a bit harder (but not impossible). • Python syntax used to describe simulation closely mimics CC 3 DML syntax. There are however certain differences and inconsistencies caused by the fact that we are using different languages to accomplish same task. Currently there is no documentation explaining in detail Python syntax that replaces CC 3 DML. It will be developed soon • The most important reason for defining entire simulation in Python is the possibility of simulation steering i. e. the ability to dynamically change simulation parameters while simulation is running (available in 3. 2. 1) • While there are certain disadvantages of this approach (mainly lack of good documentation) you get many benefits by using Python only to describe the simulation
 Cell -sorting simulation. Main script requires minimal modification: import sys from os import environ import string sys. path. append(environ["PYTHON_MODULE_PATH"]) import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() configure. Simulation(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) from Py. Steppables import Steppable. Registry steppable. Registry=Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Cell -sorting simulation. Main script requires minimal modification: import sys from os import environ import string sys. path. append(environ["PYTHON_MODULE_PATH"]) import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() configure. Simulation(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) from Py. Steppables import Steppable. Registry steppable. Registry=Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
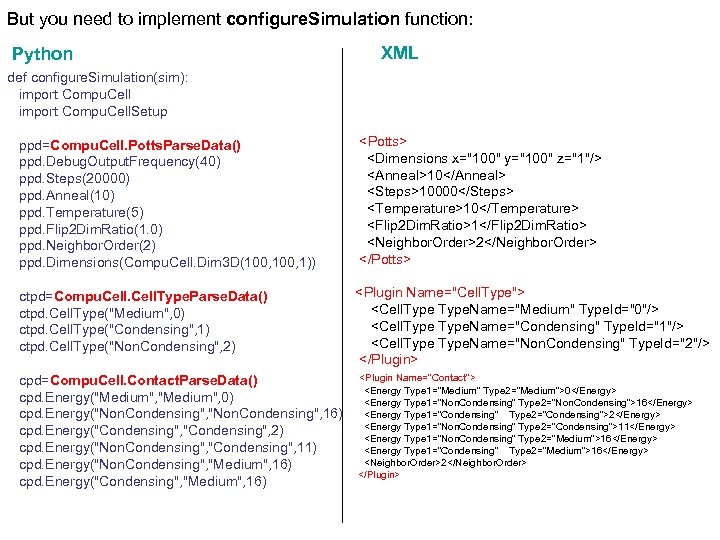 But you need to implement configure. Simulation function: Python def configure. Simulation(sim): import Compu. Cell. Setup ppd=Compu. Cell. Potts. Parse. Data() ppd. Debug. Output. Frequency(40) ppd. Steps(20000) ppd. Anneal(10) ppd. Temperature(5) ppd. Flip 2 Dim. Ratio(1. 0) ppd. Neighbor. Order(2) ppd. Dimensions(Compu. Cell. Dim 3 D(100, 1)) ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Condensing", 1) ctpd. Cell. Type("Non. Condensing", 2) cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0) cpd. Energy("Non. Condensing", 16) cpd. Energy("Condensing", 2) cpd. Energy("Non. Condensing", "Condensing", 11) cpd. Energy("Non. Condensing", "Medium", 16) cpd. Energy("Condensing", "Medium", 16) XML
But you need to implement configure. Simulation function: Python def configure. Simulation(sim): import Compu. Cell. Setup ppd=Compu. Cell. Potts. Parse. Data() ppd. Debug. Output. Frequency(40) ppd. Steps(20000) ppd. Anneal(10) ppd. Temperature(5) ppd. Flip 2 Dim. Ratio(1. 0) ppd. Neighbor. Order(2) ppd. Dimensions(Compu. Cell. Dim 3 D(100, 1)) ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Condensing", 1) ctpd. Cell. Type("Non. Condensing", 2) cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0) cpd. Energy("Non. Condensing", 16) cpd. Energy("Condensing", 2) cpd. Energy("Non. Condensing", "Condensing", 11) cpd. Energy("Non. Condensing", "Medium", 16) cpd. Energy("Condensing", "Medium", 16) XML
 Continued… Python vpd=Compu. Cell. Volume. Parse. Data() vpd. Lambda. Volume(1. 0) vpd. Target. Volume(25. 0) bipd=Compu. Cell. Blob. Initializer. Parse. Data() region=bipd. Region() region. Center(Compu. Cell. Point 3 D(50, 0)) region. Radius(40) region. Types("Condensing") region. Types("Non. Condensing") region. Width(5) #remember to register Parse. Data Compu. Cell. Setup. register. Potts(sim, ppd) Compu. Cell. Setup. register. Plugin(sim, ctpd) Compu. Cell. Setup. register. Plugin(sim, cpd) Compu. Cell. Setup. register. Plugin(sim, vpd) Compu. Cell. Setup. register. Steppable(sim, bipd) XML
Continued… Python vpd=Compu. Cell. Volume. Parse. Data() vpd. Lambda. Volume(1. 0) vpd. Target. Volume(25. 0) bipd=Compu. Cell. Blob. Initializer. Parse. Data() region=bipd. Region() region. Center(Compu. Cell. Point 3 D(50, 0)) region. Radius(40) region. Types("Condensing") region. Types("Non. Condensing") region. Width(5) #remember to register Parse. Data Compu. Cell. Setup. register. Potts(sim, ppd) Compu. Cell. Setup. register. Plugin(sim, ctpd) Compu. Cell. Setup. register. Plugin(sim, cpd) Compu. Cell. Setup. register. Plugin(sim, vpd) Compu. Cell. Setup. register. Steppable(sim, bipd) XML
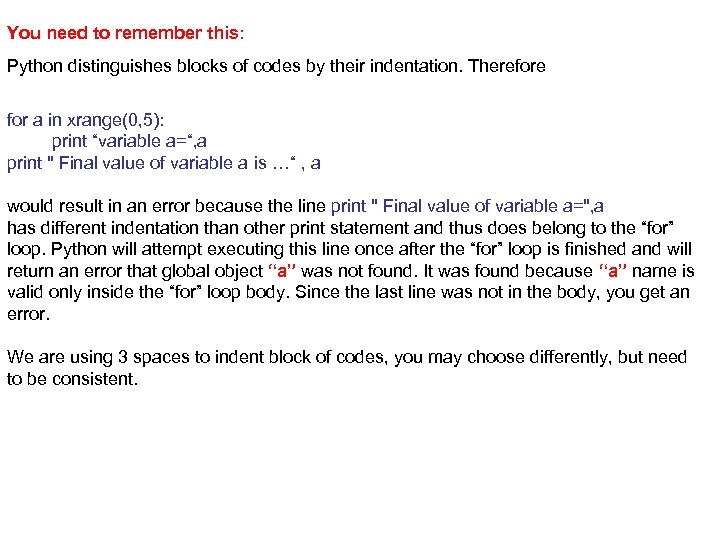 You need to remember this: Python distinguishes blocks of codes by their indentation. Therefore for a in xrange(0, 5): print “variable a=“, a print " Final value of variable a is …“ , a would result in an error because the line print " Final value of variable a=", a has different indentation than other print statement and thus does belong to the “for” loop. Python will attempt executing this line once after the “for” loop is finished and will return an error that global object “a” was not found. It was found because “a” name is valid only inside the “for” loop body. Since the last line was not in the body, you get an error. We are using 3 spaces to indent block of codes, you may choose differently, but need to be consistent.
You need to remember this: Python distinguishes blocks of codes by their indentation. Therefore for a in xrange(0, 5): print “variable a=“, a print " Final value of variable a is …“ , a would result in an error because the line print " Final value of variable a=", a has different indentation than other print statement and thus does belong to the “for” loop. Python will attempt executing this line once after the “for” loop is finished and will return an error that global object “a” was not found. It was found because “a” name is valid only inside the “for” loop body. Since the last line was not in the body, you get an error. We are using 3 spaces to indent block of codes, you may choose differently, but need to be consistent.
 Opening a Python-based simulation in the Player – no XML Go to File->Open Simulation ; Click Python script “Browse…” button to select python script. Do not forget to check “ Run Python script” checkbox!
Opening a Python-based simulation in the Player – no XML Go to File->Open Simulation ; Click Python script “Browse…” button to select python script. Do not forget to check “ Run Python script” checkbox!
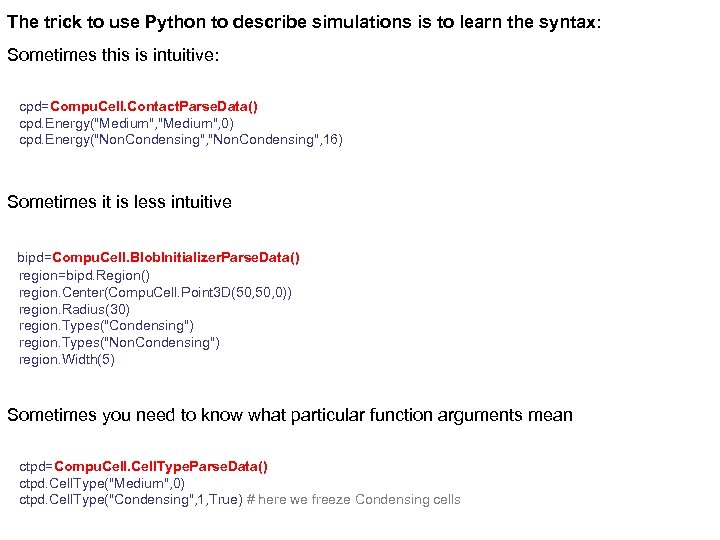 The trick to use Python to describe simulations is to learn the syntax: Sometimes this is intuitive: cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0) cpd. Energy("Non. Condensing", 16) Sometimes it is less intuitive bipd=Compu. Cell. Blob. Initializer. Parse. Data() region=bipd. Region() region. Center(Compu. Cell. Point 3 D(50, 0)) region. Radius(30) region. Types("Condensing") region. Types("Non. Condensing") region. Width(5) Sometimes you need to know what particular function arguments mean ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Condensing", 1, True) # here we freeze Condensing cells
The trick to use Python to describe simulations is to learn the syntax: Sometimes this is intuitive: cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0) cpd. Energy("Non. Condensing", 16) Sometimes it is less intuitive bipd=Compu. Cell. Blob. Initializer. Parse. Data() region=bipd. Region() region. Center(Compu. Cell. Point 3 D(50, 0)) region. Radius(30) region. Types("Condensing") region. Types("Non. Condensing") region. Width(5) Sometimes you need to know what particular function arguments mean ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Condensing", 1, True) # here we freeze Condensing cells
 Few basic rules to remember: • Plugin “My. Plugin” is initialized by creating Parse. Data type object mppd=Compu. Cell. My. Plugin. Parse. Data() Example: vpd=Compu. Cell. Volume. Parse. Data() • Initialize Parse. Data object: Example: vpd. Lambda. Volume(1. 0) vpd. Target. Volume(25. 0) • Register Parse. Data Object using either register. Potts, register. Plugin or register. Steppable functions. Example: Compu. Cell. Setup. register. Plugin(sim, vpd) See also the following examples in Compu. Cell 3 D package: *****-new-syntax. py
Few basic rules to remember: • Plugin “My. Plugin” is initialized by creating Parse. Data type object mppd=Compu. Cell. My. Plugin. Parse. Data() Example: vpd=Compu. Cell. Volume. Parse. Data() • Initialize Parse. Data object: Example: vpd. Lambda. Volume(1. 0) vpd. Target. Volume(25. 0) • Register Parse. Data Object using either register. Potts, register. Plugin or register. Steppable functions. Example: Compu. Cell. Setup. register. Plugin(sim, vpd) See also the following examples in Compu. Cell 3 D package: *****-new-syntax. py
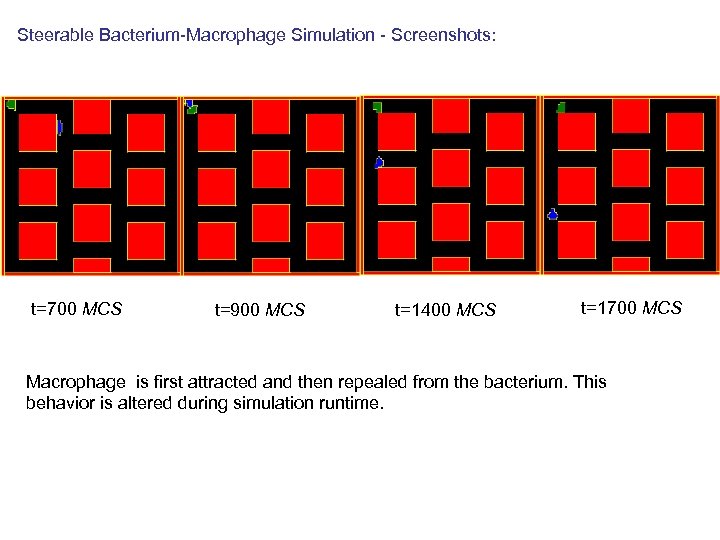
 Bacterium and macrophage simulation ppd=Compu. Cell. Potts. Parse. Data() ppd. Steps(20000) ppd. Temperature(15) ppd. Flip 2 Dim. Ratio(1. 0) ppd. Dimensions(Compu. Cell. Dim 3 D(100, 1)) ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Bacterium", 1) ctpd. Cell. Type("Macrophage", 2) Freeze ctpd. Cell. Type("Wall", 3, True)
Bacterium and macrophage simulation ppd=Compu. Cell. Potts. Parse. Data() ppd. Steps(20000) ppd. Temperature(15) ppd. Flip 2 Dim. Ratio(1. 0) ppd. Dimensions(Compu. Cell. Dim 3 D(100, 1)) ctpd=Compu. Cell. Type. Parse. Data() ctpd. Cell. Type("Medium", 0) ctpd. Cell. Type("Bacterium", 1) ctpd. Cell. Type("Macrophage", 2) Freeze ctpd. Cell. Type("Wall", 3, True)
 spd=Compu. Cell. Surface. Parse. Data() spd. Lambda. Surface(4. 0) spd. Target. Surface(20. 0)
spd=Compu. Cell. Surface. Parse. Data() spd. Lambda. Surface(4. 0) spd. Target. Surface(20. 0)
 Compu. Cell. Setup. register. Potts(sim, ppd) Compu. Cell. Setup. register. Plugin(sim, ctpd) Compu. Cell. Setup. register. Plugin(sim, vpd) Compu. Cell. Setup. register. Plugin(sim, spd) Compu. Cell. Setup. register. Plugin(sim, chpd) Compu. Cell. Setup. register. Steppable(sim, pifpd) Compu. Cell. Setup. register. Steppable(sim, fdspd) Notice that the amount of Python coding in configure. Simulation function is comparable to amount of the cc 3 DML code. Once you know CC 3 DML you should be able to pick up Python syntax easily. You get a lot more in terms of flexibility when you use Python instead of CC 3 DML
Compu. Cell. Setup. register. Potts(sim, ppd) Compu. Cell. Setup. register. Plugin(sim, ctpd) Compu. Cell. Setup. register. Plugin(sim, vpd) Compu. Cell. Setup. register. Plugin(sim, spd) Compu. Cell. Setup. register. Plugin(sim, chpd) Compu. Cell. Setup. register. Steppable(sim, pifpd) Compu. Cell. Setup. register. Steppable(sim, fdspd) Notice that the amount of Python coding in configure. Simulation function is comparable to amount of the cc 3 DML code. Once you know CC 3 DML you should be able to pick up Python syntax easily. You get a lot more in terms of flexibility when you use Python instead of CC 3 DML
 Example: Scaling contact energies – advantage of using Python to configure entire simulation energy. Scale=10 def configure. Simulation(sim): . . . cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0*energy. Scale) cpd. Energy("Macrophage", 1. 5*energy. Scale) cpd. Energy("Macrophage", "Medium", 0. 8*energy. Scale) cpd. Energy("Bacterium", 1. 5*energy. Scale) cpd. Energy("Bacterium", "Macrophage", 1. 5*energy. Scale) cpd. Energy("Bacterium", "Medium", 0. 8*energy. Scale) cpd. Energy("Wall", 0*energy. Scale) cpd. Energy("Wall", "Medium", 0*energy. Scale) cpd. Energy("Wall", "Bacterium", 5. 0*energy. Scale) cpd. Energy("Wall", "Macrophage", 5. 0*energy. Scale) It would be a bit awkward (but not impossible) to have same functionality in CC 3 DML…
Example: Scaling contact energies – advantage of using Python to configure entire simulation energy. Scale=10 def configure. Simulation(sim): . . . cpd=Compu. Cell. Contact. Parse. Data() cpd. Energy("Medium", 0*energy. Scale) cpd. Energy("Macrophage", 1. 5*energy. Scale) cpd. Energy("Macrophage", "Medium", 0. 8*energy. Scale) cpd. Energy("Bacterium", 1. 5*energy. Scale) cpd. Energy("Bacterium", "Macrophage", 1. 5*energy. Scale) cpd. Energy("Bacterium", "Medium", 0. 8*energy. Scale) cpd. Energy("Wall", 0*energy. Scale) cpd. Energy("Wall", "Medium", 0*energy. Scale) cpd. Energy("Wall", "Bacterium", 5. 0*energy. Scale) cpd. Energy("Wall", "Macrophage", 5. 0*energy. Scale) It would be a bit awkward (but not impossible) to have same functionality in CC 3 DML…
 Figuring out Parse. Data members and functions without documentation (Volume. Plugin) -Go to source code directory Compu. Cell 3 D/plugins/Volume -Open up Volume. Parse. Data. h file -Check class members and class functions and try to associate them with appropriate XML tags: class DECLSPECIFIER Volume. Parse. Data : public Parse. Data { public: Volume. Parse. Data(): Parse. Data("Volume"), target. Volume(0. ), lambda. Volume(0. ) {} double target. Volume; double lambda. Volume; void Lambda. Volume(double _lambda. Volume){lambda. Volume=_lambda. Volume; } void Target. Volume(double _target. Volume){target. Volume=_target. Volume; } };
Figuring out Parse. Data members and functions without documentation (Volume. Plugin) -Go to source code directory Compu. Cell 3 D/plugins/Volume -Open up Volume. Parse. Data. h file -Check class members and class functions and try to associate them with appropriate XML tags: class DECLSPECIFIER Volume. Parse. Data : public Parse. Data { public: Volume. Parse. Data(): Parse. Data("Volume"), target. Volume(0. ), lambda. Volume(0. ) {} double target. Volume; double lambda. Volume; void Lambda. Volume(double _lambda. Volume){lambda. Volume=_lambda. Volume; } void Target. Volume(double _target. Volume){target. Volume=_target. Volume; } };
 Plasticity. Tracker. Plugin class DECLSPECIFIER Plasticity. Tracker. Parse. Data: public Parse. Data{ public: Plasticity. Tracker. Parse. Data(): Parse. Data("Plasticity. Tracker") {} std: : set
Plasticity. Tracker. Plugin class DECLSPECIFIER Plasticity. Tracker. Parse. Data: public Parse. Data{ public: Plasticity. Tracker. Parse. Data(): Parse. Data("Plasticity. Tracker") {} std: : set
 Beyond XML - Developing Python Steppables Examples presented above showed how to run Python based simulations and how to replace XML with Python. However, the true power of Python is demonstrated in the case when you develop your own modules. We will first teach you how to develop a steppable because steppables are most likely to be developed in Python anyway. Let’s take a look at the module that prints cell id, cell type and cell volume for every cell in the simulation. Iterating over all cells is probably most frequently used task in steppables: class Info. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def start(self): print "This function is called once before simulation" def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume
Beyond XML - Developing Python Steppables Examples presented above showed how to run Python based simulations and how to replace XML with Python. However, the true power of Python is demonstrated in the case when you develop your own modules. We will first teach you how to develop a steppable because steppables are most likely to be developed in Python anyway. Let’s take a look at the module that prints cell id, cell type and cell volume for every cell in the simulation. Iterating over all cells is probably most frequently used task in steppables: class Info. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def start(self): print "This function is called once before simulation" def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume
 Python Steppable Each Python Steppable should have three functions: start() step(mcs) finish() It is OK to leave the implementation of any of above functions empty. An empty function will be then called. In addition to this, because Python steppables are implemented as classes they need to define __init__ function that act as a constructor. Steppable Template: class Your. Python. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): #your code here def start(self): #your code here def step(self, mcs): #your code here def finish(self): #your code here
Python Steppable Each Python Steppable should have three functions: start() step(mcs) finish() It is OK to leave the implementation of any of above functions empty. An empty function will be then called. In addition to this, because Python steppables are implemented as classes they need to define __init__ function that act as a constructor. Steppable Template: class Your. Python. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): #your code here def start(self): #your code here def step(self, mcs): #your code here def finish(self): #your code here
 If you are non-programmer it may looks a bit strange, but imagine how much more would be required to write do the same in C/C++. Much more. Let’s explain the code: class Info. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) First line defines our steppable class. Each class has to have __init__method that is called when object of this class is created. You can pass any arguments to this method, but the first argument must be “self”. This is required by Python language. First line in __init__ method initializes Base class Steppable. Py. Do not worry if you do not understand it. Treat it as a boiler plate code. Line self. simulator=_simulator stores a pointer or reference to simulator object as a member variable. This way, later on whenever we decide to reference simulator we would use self. simmulator instead of getting simulator some other , more complicated way. Notice, that whenever you access member variable you prepend their name with keyword “self” Subsequently we get reference to cell inventoryt (C++ object) and use it to create iterable cell. List (self. cell. List=Cell. List(self. inventory)) Notice, “self” shows up again.
If you are non-programmer it may looks a bit strange, but imagine how much more would be required to write do the same in C/C++. Much more. Let’s explain the code: class Info. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) First line defines our steppable class. Each class has to have __init__method that is called when object of this class is created. You can pass any arguments to this method, but the first argument must be “self”. This is required by Python language. First line in __init__ method initializes Base class Steppable. Py. Do not worry if you do not understand it. Treat it as a boiler plate code. Line self. simulator=_simulator stores a pointer or reference to simulator object as a member variable. This way, later on whenever we decide to reference simulator we would use self. simmulator instead of getting simulator some other , more complicated way. Notice, that whenever you access member variable you prepend their name with keyword “self” Subsequently we get reference to cell inventoryt (C++ object) and use it to create iterable cell. List (self. cell. List=Cell. List(self. inventory)) Notice, “self” shows up again.
 def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume Above function implements core functionality of our steppable. It informs that it is called every 10 MCS – see how we set frequency parameter in the __init__ function. The last two lines do actual iteration over each cell in the cell inventory Notice that it is really easy to do the iteration: for cell in self. cell. List: Now you can see how storing Cell. Type object as self. cell. List comes handy. All we need to do is to pass iterable cell list (self. cell. List) to the “for” loop. Actual printing is done in line print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume For each cell in inventory “cell” variable of the for loop will be initialized with different cell from inventory. All you need to do is to print cell. id, cell. type, and cell. volume. It is pretty simple.
def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume Above function implements core functionality of our steppable. It informs that it is called every 10 MCS – see how we set frequency parameter in the __init__ function. The last two lines do actual iteration over each cell in the cell inventory Notice that it is really easy to do the iteration: for cell in self. cell. List: Now you can see how storing Cell. Type object as self. cell. List comes handy. All we need to do is to pass iterable cell list (self. cell. List) to the “for” loop. Actual printing is done in line print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume For each cell in inventory “cell” variable of the for loop will be initialized with different cell from inventory. All you need to do is to print cell. id, cell. type, and cell. volume. It is pretty simple.
 Now save the file with the steppable as , cellsort_2 D_steppables. py. All you need to do is to provide hooks to your steppable in the main Python script: steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() ##### Steppable Registration ###### from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer= Info. Printer. Steppable(sim) steppable. Registry. register. Steppable(info. Printer) #####End of Steppable Registration ###### steppable. Registry. init(sim) Notice that registering steppable requires importing your steppable from the file: from cellsort_2 D_stepables import Info. Printer. Steppable creating steppable object: info. Printer= Info. Printer. Steppable(sim) registering it with steppable registry: steppable. Registry. register. Steppable(info. Printer)
Now save the file with the steppable as , cellsort_2 D_steppables. py. All you need to do is to provide hooks to your steppable in the main Python script: steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() ##### Steppable Registration ###### from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer= Info. Printer. Steppable(sim) steppable. Registry. register. Steppable(info. Printer) #####End of Steppable Registration ###### steppable. Registry. init(sim) Notice that registering steppable requires importing your steppable from the file: from cellsort_2 D_stepables import Info. Printer. Steppable creating steppable object: info. Printer= Info. Printer. Steppable(sim) registering it with steppable registry: steppable. Registry. register. Steppable(info. Printer)
 Full Main Script: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer. Steppable=Info. Printer. Steppable(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(info. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Full Main Script: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer. Steppable=Info. Printer. Steppable(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(info. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
 #import useful modules import sys from os import environ from os import getcwd import string #setup search patths sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") #add search path import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer. Steppable=Info. Printer. Steppable(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(info. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
#import useful modules import sys from os import environ from os import getcwd import string #setup search patths sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") #add search path import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Info. Printer. Steppable info. Printer. Steppable=Info. Printer. Steppable(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(info. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
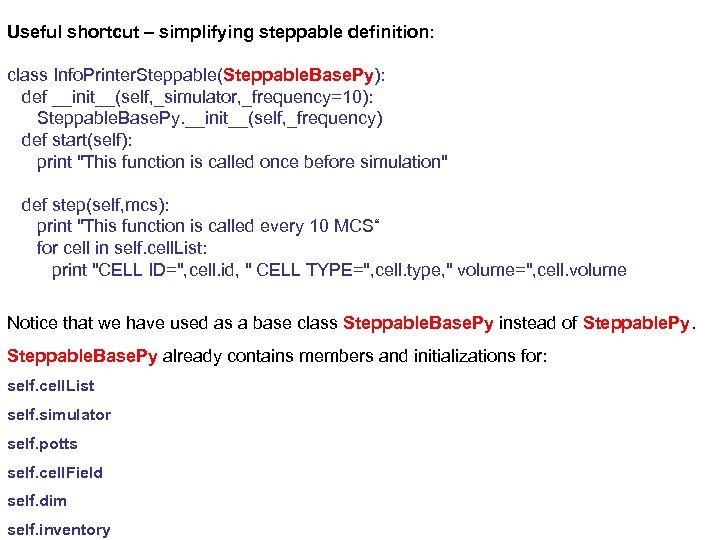 Useful shortcut – simplifying steppable definition: class Info. Printer. Steppable(Steppable. Base. Py): def __init__(self, _simulator, _frequency=10): Steppable. Base. Py. __init__(self, _frequency) def start(self): print "This function is called once before simulation" def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume Notice that we have used as a base class Steppable. Base. Py instead of Steppable. Py. Steppable. Base. Py already contains members and initializations for: self. cell. List self. simulator self. potts self. cell. Field self. dim self. inventory
Useful shortcut – simplifying steppable definition: class Info. Printer. Steppable(Steppable. Base. Py): def __init__(self, _simulator, _frequency=10): Steppable. Base. Py. __init__(self, _frequency) def start(self): print "This function is called once before simulation" def step(self, mcs): print "This function is called every 10 MCS“ for cell in self. cell. List: print "CELL ID=", cell. id, " CELL TYPE=", cell. type, " volume=", cell. volume Notice that we have used as a base class Steppable. Base. Py instead of Steppable. Py. Steppable. Base. Py already contains members and initializations for: self. cell. List self. simulator self. potts self. cell. Field self. dim self. inventory
 Steppable. Base. Py: class Steppable. Base. Py(Steppable. Py): def __init__(self, _simulator, _frequency=1): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. potts=_simulator. get. Potts() self. cell. Field=self. potts. get. Cell. Field. G() self. dim=self. cell. Field. get. Dim() self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory)
Steppable. Base. Py: class Steppable. Base. Py(Steppable. Py): def __init__(self, _simulator, _frequency=1): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. potts=_simulator. get. Potts() self. cell. Field=self. potts. get. Cell. Field. G() self. dim=self. cell. Field. get. Dim() self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory)
 Now, all you need to do is to open in the Player previous xml file (cellsort_2 D. xml) together with newly created cellsort_2 D. py and check Run Python check box. Notice that you are not loading directly cellsort_2 D_steppables. py file. The module you stored to this file will be called from cellsort_2 D. py. Try running the simulation and see if you got any performance degradation. Probably not, but by using Python you have saved yourself a lot of tedious C++ coding, not to mention that you do not need to care about dependencies, compilation, etc. . Writing your next Python steppable will require much less effort as well, as you will quickly discover that you will be using same basic code template all over again. Instead of thinking how the code you are writing fits in the overall framework you will just concentrate on it’s core functionality and leave the rest to Compu. Cell 3 D. In case you wonder how this is all possible , it is due to Object Oriented programming. Hopefully this short tutorial will encourage you to learn more of object oriented programming. It is really worth the effort.
Now, all you need to do is to open in the Player previous xml file (cellsort_2 D. xml) together with newly created cellsort_2 D. py and check Run Python check box. Notice that you are not loading directly cellsort_2 D_steppables. py file. The module you stored to this file will be called from cellsort_2 D. py. Try running the simulation and see if you got any performance degradation. Probably not, but by using Python you have saved yourself a lot of tedious C++ coding, not to mention that you do not need to care about dependencies, compilation, etc. . Writing your next Python steppable will require much less effort as well, as you will quickly discover that you will be using same basic code template all over again. Instead of thinking how the code you are writing fits in the overall framework you will just concentrate on it’s core functionality and leave the rest to Compu. Cell 3 D. In case you wonder how this is all possible , it is due to Object Oriented programming. Hopefully this short tutorial will encourage you to learn more of object oriented programming. It is really worth the effort.
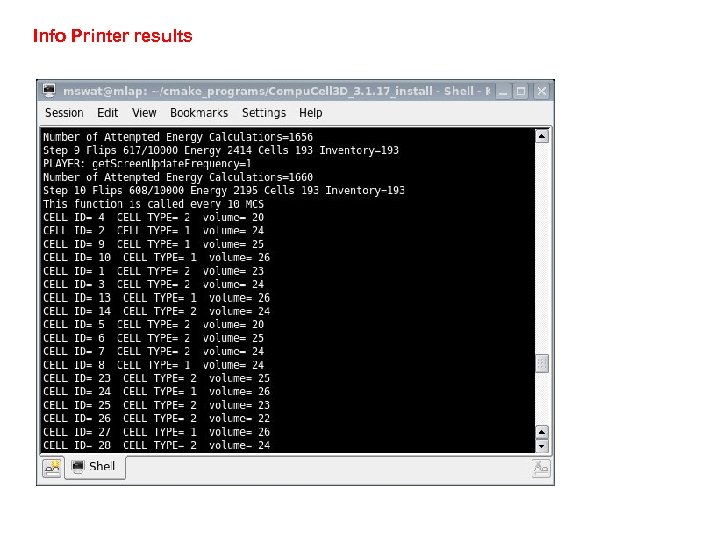 Info Printer results
Info Printer results
 Python Scripting Checklist: 1. Write main Python script (modify or reuse existing one) 2. Write Python modules in a separate file. You will import these modules from main Python script 3. Provide CC 3 DML configuration file or describe entire simulation in Python skipping CC 3 DML entirely Note: when using Python scripting your simulation may consists of many files. Make sure you keep track of them
Python Scripting Checklist: 1. Write main Python script (modify or reuse existing one) 2. Write Python modules in a separate file. You will import these modules from main Python script 3. Provide CC 3 DML configuration file or describe entire simulation in Python skipping CC 3 DML entirely Note: when using Python scripting your simulation may consists of many files. Make sure you keep track of them
 More Complicated Simulations – Adding Extra Attribute To a Cell In Compu. Cell 3 D simulations each cell by default will have several attributes such as volume, surface, centroids , target volume, cell id etc. One can write a plugin that attaches additional attributes to a cell during run time. Doing so avoids recompilation of entire Compu. Cell 3 D but requires to write and compile the C++ plugin. It is by far the easiest to attach additional cell attribute in Python. Not only there is no need to recompile anything, but the actual task takes one line of code: py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Above we told Compu. Cell 3 D to attach a Python list to each cell that will be produced by the Compu. Cell 3 D kernel. We can access this list very easily from Python level. Python list is dynamic data structure that can grow or shrink and can hold arbitrary Python objects. Therefore by attaching a list to each cell we effectively came up with a way to attach any cell attribute. We may also attach dictionary instead of the list: py. Attribute. Adder, dict. Adder=Compu. Cell. Setup. attach. Dictionary. To. Cells(sim) And everything takes place during run time…
More Complicated Simulations – Adding Extra Attribute To a Cell In Compu. Cell 3 D simulations each cell by default will have several attributes such as volume, surface, centroids , target volume, cell id etc. One can write a plugin that attaches additional attributes to a cell during run time. Doing so avoids recompilation of entire Compu. Cell 3 D but requires to write and compile the C++ plugin. It is by far the easiest to attach additional cell attribute in Python. Not only there is no need to recompile anything, but the actual task takes one line of code: py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Above we told Compu. Cell 3 D to attach a Python list to each cell that will be produced by the Compu. Cell 3 D kernel. We can access this list very easily from Python level. Python list is dynamic data structure that can grow or shrink and can hold arbitrary Python objects. Therefore by attaching a list to each cell we effectively came up with a way to attach any cell attribute. We may also attach dictionary instead of the list: py. Attribute. Adder, dict. Adder=Compu. Cell. Setup. attach. Dictionary. To. Cells(sim) And everything takes place during run time…
 Full listing of simulation where each cell gets extra attribute – a list: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() #here we will add Extra. Attribute. Cellsort steppable from cellsort_2 D_steppables import Extra. Attribute. Cellsort extra. Attribute. Cellsort=Extra. Attribute. Cellsort(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(extra. Attribute. Cellsort) from cellsort_2 D_steppables import Type. Switcher. Steppable type. Switcher. Steppable=Type. Switcher. Steppable(sim, 100) steppable. Registry. register. Steppable(type. Switcher. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Full listing of simulation where each cell gets extra attribute – a list: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() #here we will add Extra. Attribute. Cellsort steppable from cellsort_2 D_steppables import Extra. Attribute. Cellsort extra. Attribute. Cellsort=Extra. Attribute. Cellsort(_simulator=sim, _frequency=10) steppable. Registry. register. Steppable(extra. Attribute. Cellsort) from cellsort_2 D_steppables import Type. Switcher. Steppable type. Switcher. Steppable=Type. Switcher. Steppable(sim, 100) steppable. Registry. register. Steppable(type. Switcher. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
 Extra. Attribute. Cellsort class Extra. Attribute. Cellsort(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def step(self, mcs): for cell in self. cell. List: py. Attrib=Compu. Cell. get. Py. Attrib(cell) py. Attrib[0: 2]=[cell. id*mcs , cell. id*(mcs-1)] print "CELL ID modified=", py. Attrib[0], " ", py. Attrib[1] Initializing first two elements of the list Notice, you may also attach a dictionary to a cell instead of a list. See Python Scripting Tutorials for more information. Dictionaries are actually more useful then lists in the Compu. Cell 3 D context so make sure you understand them and know how to attach them to cells.
Extra. Attribute. Cellsort class Extra. Attribute. Cellsort(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def step(self, mcs): for cell in self. cell. List: py. Attrib=Compu. Cell. get. Py. Attrib(cell) py. Attrib[0: 2]=[cell. id*mcs , cell. id*(mcs-1)] print "CELL ID modified=", py. Attrib[0], " ", py. Attrib[1] Initializing first two elements of the list Notice, you may also attach a dictionary to a cell instead of a list. See Python Scripting Tutorials for more information. Dictionaries are actually more useful then lists in the Compu. Cell 3 D context so make sure you understand them and know how to attach them to cells.
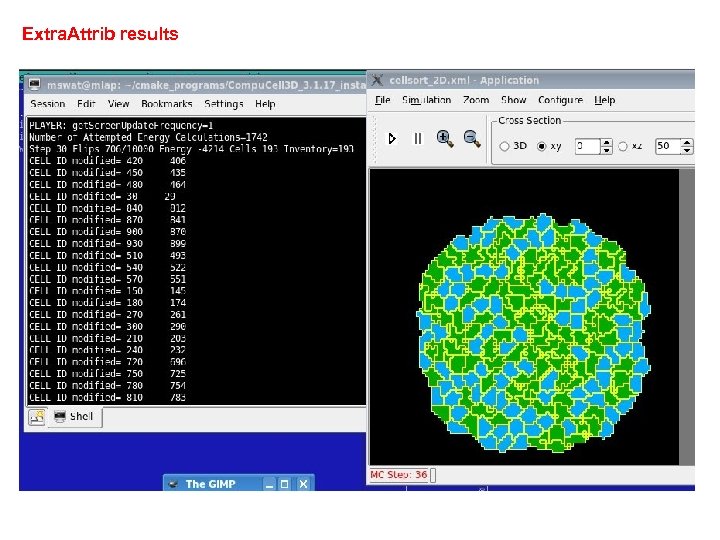 Extra. Attrib results
Extra. Attrib results
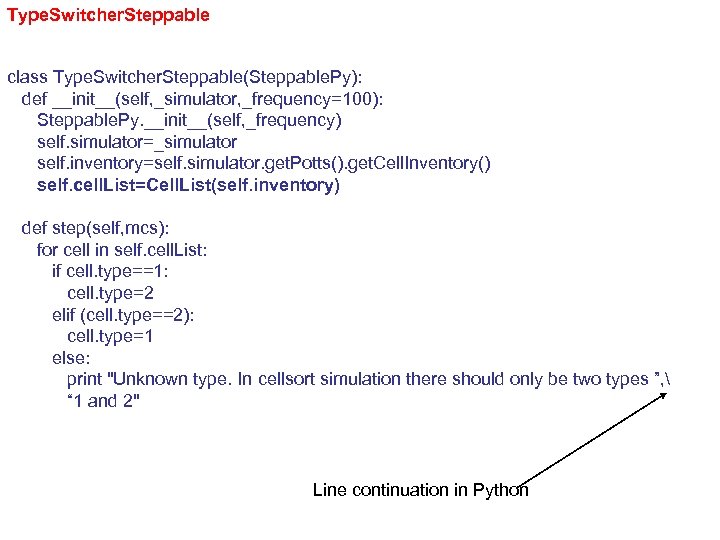 Type. Switcher. Steppable class Type. Switcher. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=100): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def step(self, mcs): for cell in self. cell. List: if cell. type==1: cell. type=2 elif (cell. type==2): cell. type=1 else: print "Unknown type. In cellsort simulation there should only be two types ”, “ 1 and 2" Line continuation in Python
Type. Switcher. Steppable class Type. Switcher. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=100): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def step(self, mcs): for cell in self. cell. List: if cell. type==1: cell. type=2 elif (cell. type==2): cell. type=1 else: print "Unknown type. In cellsort simulation there should only be two types ”, “ 1 and 2" Line continuation in Python
 Accessing Neighbor. Tracker from Python As you remember from lectures on CC 3 DML configuration files, Compu. Cell 3 D can track cell neighbors. You can access information about cell neighbors directly from Python: class Neighbor. Tracker. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=100): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. n. Tracker. Plugin=Compu. Cell. get. Neighbor. Tracker. Plugin() self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def start(self): pass def step(self, mcs): self. cell. List=Cell. List(self. inventory) for cell in self. cell. List: cell. Neighbor. List=Cell. Neighbor. List. Auto(self. n. Tracker. Plugin, cell) #create local neighbor list print "*****NEIGHBORS OF CELL WITH ID ", cell. id, " *********" for neighbor. Surface. Data in cell. Neighbor. List: #iterate over local neighbor list if neighbor. Surface. Data. neighbor. Address: #check if neighbor is non-Medium print "neighbor. id", neighbor. Surface. Data. neighbor. Address. id, " common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area #access common surface area and id else: print "Medium common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area
Accessing Neighbor. Tracker from Python As you remember from lectures on CC 3 DML configuration files, Compu. Cell 3 D can track cell neighbors. You can access information about cell neighbors directly from Python: class Neighbor. Tracker. Printer. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=100): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. n. Tracker. Plugin=Compu. Cell. get. Neighbor. Tracker. Plugin() self. inventory=self. simulator. get. Potts(). get. Cell. Inventory() self. cell. List=Cell. List(self. inventory) def start(self): pass def step(self, mcs): self. cell. List=Cell. List(self. inventory) for cell in self. cell. List: cell. Neighbor. List=Cell. Neighbor. List. Auto(self. n. Tracker. Plugin, cell) #create local neighbor list print "*****NEIGHBORS OF CELL WITH ID ", cell. id, " *********" for neighbor. Surface. Data in cell. Neighbor. List: #iterate over local neighbor list if neighbor. Surface. Data. neighbor. Address: #check if neighbor is non-Medium print "neighbor. id", neighbor. Surface. Data. neighbor. Address. id, " common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area #access common surface area and id else: print "Medium common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area
 Understanding iteration over cell neighbors def step(self, mcs): self. cell. List=Cell. List(self. inventory) for cell in self. cell. List: cell. Neighbor. List=Cell. Neighbor. List. Auto(self. n. Tracker. Plugin, cell) print "*****NEIGHBORS OF CELL WITH ID ", cell. id, " *********" for neighbor. Surface. Data in cell. Neighbor. List: if neighbor. Surface. Data. neighbor. Address: print "neighbor. id", neighbor. Surface. Data. neighbor. Address. id, " common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area else: print "Medium common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area Iterating over all cells in the simulation This function constructs list of cell neighbors neighbor. Surface. Data object has neighbor. Address and common. Surface. Area members. The first one stores a pointer to a neighbor cell, the second stores sommon surface area between neighbors
Understanding iteration over cell neighbors def step(self, mcs): self. cell. List=Cell. List(self. inventory) for cell in self. cell. List: cell. Neighbor. List=Cell. Neighbor. List. Auto(self. n. Tracker. Plugin, cell) print "*****NEIGHBORS OF CELL WITH ID ", cell. id, " *********" for neighbor. Surface. Data in cell. Neighbor. List: if neighbor. Surface. Data. neighbor. Address: print "neighbor. id", neighbor. Surface. Data. neighbor. Address. id, " common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area else: print "Medium common. Surface. Area=", neighbor. Surface. Data. common. Surface. Area Iterating over all cells in the simulation This function constructs list of cell neighbors neighbor. Surface. Data object has neighbor. Address and common. Surface. Area members. The first one stores a pointer to a neighbor cell, the second stores sommon surface area between neighbors
 import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Neighbor. Tracker. Printer. Steppable neighbor. Tracker. Printer. Steppable=Neighbor. Tracker. Printer. Steppable(sim, 100) steppable. Registry. register. Steppable(neighbor. Tracker. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_steppables import Neighbor. Tracker. Printer. Steppable neighbor. Tracker. Printer. Steppable=Neighbor. Tracker. Printer. Steppable(sim, 100) steppable. Registry. register. Steppable(neighbor. Tracker. Printer. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
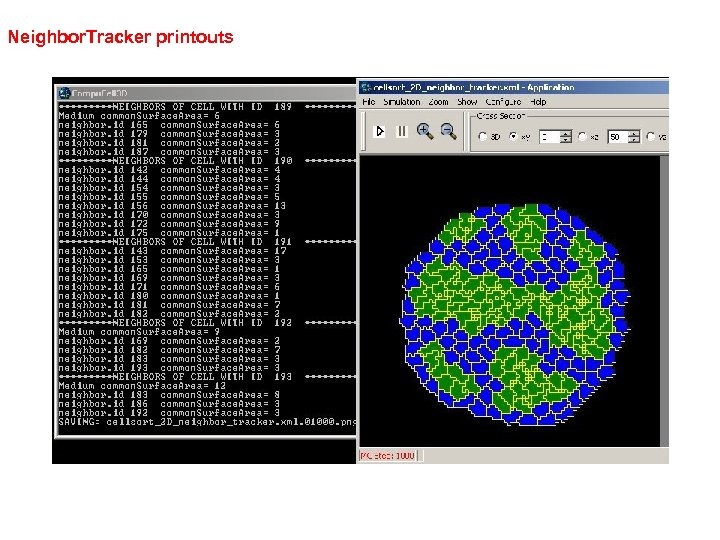 Neighbor. Tracker printouts
Neighbor. Tracker printouts
 Printing values of the concentration to a file import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here from Py. Steppables. Examples import Steppable. Registry steppable. Registry=Steppable. Registry() from cellsort_2 D_steppables import Concentration. Field. Dumper. Steppable concentration. Field. Dumper. Steppable=Concentration. Field. Dumper. Steppable(sim, _frequency=100) concentration. Field. Dumper. Steppable. set. Field. Name("FGF") steppable. Registry. register. Steppable(concentration. Field. Dumper. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Printing values of the concentration to a file import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here from Py. Steppables. Examples import Steppable. Registry steppable. Registry=Steppable. Registry() from cellsort_2 D_steppables import Concentration. Field. Dumper. Steppable concentration. Field. Dumper. Steppable=Concentration. Field. Dumper. Steppable(sim, _frequency=100) concentration. Field. Dumper. Steppable. set. Field. Name("FGF") steppable. Registry. register. Steppable(concentration. Field. Dumper. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
 class Concentration. Field. Dumper. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=1): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. dim=self. simulator. get. Potts(). get. Cell. Field. G(). get. Dim() def set. Field. Name(self, _field. Name): self. field. Name=_field. Name def step(self, mcs): file. Name=self. field. Name+"_"+str(mcs)+". dat" self. output. Field(self. field. Name, file. Name) def output. Field(self, _field. Name, _file. Name): field=Compu. Cell. get. Concentration. Field(self. simulator, _field. Name) pt=Compu. Cell. Point 3 D() if field: try: file. Handle=open(_file. Name, "w") except IOError: print "Could not open file ", _file. Name, " for writing. Check if you have necessary permissions" print "dim. x=", self. dim. x for i in xrange(self. dim. x): for j in xrange(self. dim. y): for k in xrange(self. dim. z): pt. x=i pt. y=j pt. z=k file. Handle. write("%dt%dt%fn"%(pt. x, pt. y, pt. z, field. get(pt))) #write to a file
class Concentration. Field. Dumper. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=1): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. dim=self. simulator. get. Potts(). get. Cell. Field. G(). get. Dim() def set. Field. Name(self, _field. Name): self. field. Name=_field. Name def step(self, mcs): file. Name=self. field. Name+"_"+str(mcs)+". dat" self. output. Field(self. field. Name, file. Name) def output. Field(self, _field. Name, _file. Name): field=Compu. Cell. get. Concentration. Field(self. simulator, _field. Name) pt=Compu. Cell. Point 3 D() if field: try: file. Handle=open(_file. Name, "w") except IOError: print "Could not open file ", _file. Name, " for writing. Check if you have necessary permissions" print "dim. x=", self. dim. x for i in xrange(self. dim. x): for j in xrange(self. dim. y): for k in xrange(self. dim. z): pt. x=i pt. y=j pt. z=k file. Handle. write("%dt%dt%fn"%(pt. x, pt. y, pt. z, field. get(pt))) #write to a file
 Creating, initializing and manipulating a concentration field directly from Python • Although in most cases concentration fields are created and manipulated by PDE solvers it is possible to accomplish all those tasks directly from Python. • This can be very useful if you want to develop custom visualization that is not directly supported by the Player. For example you may want to color cells according to how many neighbors they have. Player does not offer such an option but you can implement it very easily in Python in less then 5 minutes. This is not a joke. I am sure that by combining two examples from this tutorial you will accomplish this task very fast. The task of adding extra field to the Player and “managing” it consist of two steps • Creating extra field and registering it with the Player and Compu. Cell 3 D kernel • Writing steppable that manipulates values stored in the field First let’s look at the full listing:
Creating, initializing and manipulating a concentration field directly from Python • Although in most cases concentration fields are created and manipulated by PDE solvers it is possible to accomplish all those tasks directly from Python. • This can be very useful if you want to develop custom visualization that is not directly supported by the Player. For example you may want to color cells according to how many neighbors they have. Player does not offer such an option but you can implement it very easily in Python in less then 5 minutes. This is not a joke. I am sure that by combining two examples from this tutorial you will accomplish this task very fast. The task of adding extra field to the Player and “managing” it consist of two steps • Creating extra field and registering it with the Player and Compu. Cell 3 D kernel • Writing steppable that manipulates values stored in the field First let’s look at the full listing:
 import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import System. Utils. initialize. System. Resources() Creating extra field is is really easy. The location of the function call that creates the field is , however important. See the comment import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() import Compu. Cell #notice importing Compu. Cell to main script has to be done after call to get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Need to call initialize. Simulation. Objects before trying to access lattice dimension dim=sim. get. Potts(). get. Cell. Field. G(). get. Dim() extra. Player. Field=simthread. create. Float. Field. Py(dim, "Extra. Field") # initializing extra Field - this location in the #Add Python steppables here from Py. Steppables. Examples import Steppable. Registry steppable. Registry=Steppable. Registry() from cellsort_2 D_steppables import Extra. Field. Visualization. Steppable extra. Field. Visualization. Steppable=Extra. Field. Visualization. Steppable(_simulator=sim, _frequency=10) extra. Field. Visualization. Steppable. set. Scalar. Field(extra. Player. Field) steppable. Registry. register. Steppable(extra. Field. Visualization. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import System. Utils. initialize. System. Resources() Creating extra field is is really easy. The location of the function call that creates the field is , however important. See the comment import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() import Compu. Cell #notice importing Compu. Cell to main script has to be done after call to get. Core. Simulation. Objects() #Create extra player fields here or add attributes Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Need to call initialize. Simulation. Objects before trying to access lattice dimension dim=sim. get. Potts(). get. Cell. Field. G(). get. Dim() extra. Player. Field=simthread. create. Float. Field. Py(dim, "Extra. Field") # initializing extra Field - this location in the #Add Python steppables here from Py. Steppables. Examples import Steppable. Registry steppable. Registry=Steppable. Registry() from cellsort_2 D_steppables import Extra. Field. Visualization. Steppable extra. Field. Visualization. Steppable=Extra. Field. Visualization. Steppable(_simulator=sim, _frequency=10) extra. Field. Visualization. Steppable. set. Scalar. Field(extra. Player. Field) steppable. Registry. register. Steppable(extra. Field. Visualization. Steppable) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
 from Player. Python import * # necessary to manipulate Player fields from math import * # getting access to special functions from math module class Extra. Field. Visualization. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. cell. Field. G=self. simulator. get. Potts(). get. Cell. Field. G() self. dim=self. cell. Field. G. get. Dim() def set. Scalar. Field(self, _field): # getting access to newly created field self. scalar. Field=_field def start(self): pass def step(self, mcs): for x in xrange(self. dim. x): #iteration over each pixel for y in xrange(self. dim. y): for z in xrange(self. dim. z): pt=Compu. Cell. Point 3 D(x, y, z) if (not mcs%20): #filling the values of the concentration value=x*y # sometimes it is x*y fill. Scalar. Value(self. scalar. Field, x, y, z, value) else: value=sin(x*y) # sometimes sin(x*y) fill. Scalar. Value(self. scalar. Field, x, y, z, value)
from Player. Python import * # necessary to manipulate Player fields from math import * # getting access to special functions from math module class Extra. Field. Visualization. Steppable(Steppable. Py): def __init__(self, _simulator, _frequency=10): Steppable. Py. __init__(self, _frequency) self. simulator=_simulator self. cell. Field. G=self. simulator. get. Potts(). get. Cell. Field. G() self. dim=self. cell. Field. G. get. Dim() def set. Scalar. Field(self, _field): # getting access to newly created field self. scalar. Field=_field def start(self): pass def step(self, mcs): for x in xrange(self. dim. x): #iteration over each pixel for y in xrange(self. dim. y): for z in xrange(self. dim. z): pt=Compu. Cell. Point 3 D(x, y, z) if (not mcs%20): #filling the values of the concentration value=x*y # sometimes it is x*y fill. Scalar. Value(self. scalar. Field, x, y, z, value) else: value=sin(x*y) # sometimes sin(x*y) fill. Scalar. Value(self. scalar. Field, x, y, z, value)
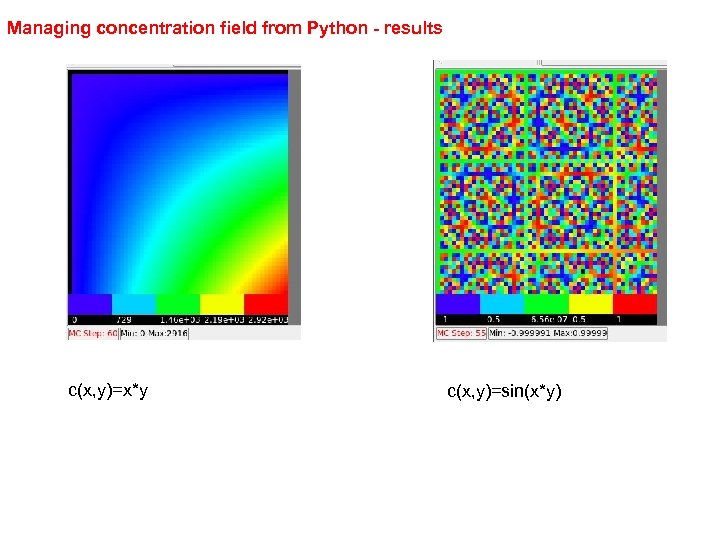 Managing concentration field from Python - results c(x, y)=x*y c(x, y)=sin(x*y)
Managing concentration field from Python - results c(x, y)=x*y c(x, y)=sin(x*y)
 Mitosis in Compu. Cell 3 D simulations Supporting cell division (mitosis) in Compu. Cell 3 D simulations is a prerequisite for building faithful biomedical simulations. You can use mitosis module (Mitosis Plugin) directly from XML however, its use will be very limited because of the following fact: After cell division you end up with two cells. What parameters should those two cells have (type, target volume etc. )? How do you modify the parameters? The best solution is to manage mitosis from Python and the example below will explain you how to do it.
Mitosis in Compu. Cell 3 D simulations Supporting cell division (mitosis) in Compu. Cell 3 D simulations is a prerequisite for building faithful biomedical simulations. You can use mitosis module (Mitosis Plugin) directly from XML however, its use will be very limited because of the following fact: After cell division you end up with two cells. What parameters should those two cells have (type, target volume etc. )? How do you modify the parameters? The best solution is to manage mitosis from Python and the example below will explain you how to do it.
 import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #add additional attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) import Compu. Cell #notice importing Compu. Cell to main script has to be done after call to get. Core. Simulation. Objects() change. Watcher. Registry=Compu. Cell. Setup. get. Change. Watcher. Registry(sim) stepper. Registry=Compu. Cell. Setup. get. Stepper. Registry(sim) from cellsort_2 D_field_modules import Cellsort. Mitosis cellsort. Mitosis=Cellsort. Mitosis(sim, change. Watcher. Registry, stepper. Registry) cellsort. Mitosis. set. Doubling. Volume(50) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_field_modules import Volume. Constraint. Steppable volume. Constraint=Volume. Constraint. Steppable(sim) steppable. Registry. register. Steppable(volume. Constraint) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #add additional attributes py. Attribute. Adder, list. Adder=Compu. Cell. Setup. attach. List. To. Cells(sim) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) import Compu. Cell #notice importing Compu. Cell to main script has to be done after call to get. Core. Simulation. Objects() change. Watcher. Registry=Compu. Cell. Setup. get. Change. Watcher. Registry(sim) stepper. Registry=Compu. Cell. Setup. get. Stepper. Registry(sim) from cellsort_2 D_field_modules import Cellsort. Mitosis cellsort. Mitosis=Cellsort. Mitosis(sim, change. Watcher. Registry, stepper. Registry) cellsort. Mitosis. set. Doubling. Volume(50) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() from cellsort_2 D_field_modules import Volume. Constraint. Steppable volume. Constraint=Volume. Constraint. Steppable(sim) steppable. Registry. register. Steppable(volume. Constraint) Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
 Mitosis function is a type of plugin that monitors lattice (field 3 DWatcher). Most of the mitosis setup is handled inside base class Mitosis. Py. Plugin. Base from random import random from Py. Plugins. Examples import Mitosis. Py. Plugin. Base class Cellsort. Mitosis(Mitosis. Py. Plugin. Base): #inherit base class def __init__(self , _simulator , _change. Watcher. Registry , _stepper. Registry): Mitosis. Py. Plugin. Base. __init__(self, _simulator, _change. Watcher. Registry, _stepper. Registry) def update. Attributes(self): #called after mitosis is done self. parent. Cell. target. Volume/=2. 0 self. child. Cell. target. Volume=self. parent. Cell. target. Volume self. child. Cell. lambda. Volume=self. parent. Cell. lambda. Volume if (random()<0. 5): self. child. Cell. type=self. parent. Cell. type else: self. child. Cell. type=3
Mitosis function is a type of plugin that monitors lattice (field 3 DWatcher). Most of the mitosis setup is handled inside base class Mitosis. Py. Plugin. Base from random import random from Py. Plugins. Examples import Mitosis. Py. Plugin. Base class Cellsort. Mitosis(Mitosis. Py. Plugin. Base): #inherit base class def __init__(self , _simulator , _change. Watcher. Registry , _stepper. Registry): Mitosis. Py. Plugin. Base. __init__(self, _simulator, _change. Watcher. Registry, _stepper. Registry) def update. Attributes(self): #called after mitosis is done self. parent. Cell. target. Volume/=2. 0 self. child. Cell. target. Volume=self. parent. Cell. target. Volume self. child. Cell. lambda. Volume=self. parent. Cell. lambda. Volume if (random()<0. 5): self. child. Cell. type=self. parent. Cell. type else: self. child. Cell. type=3
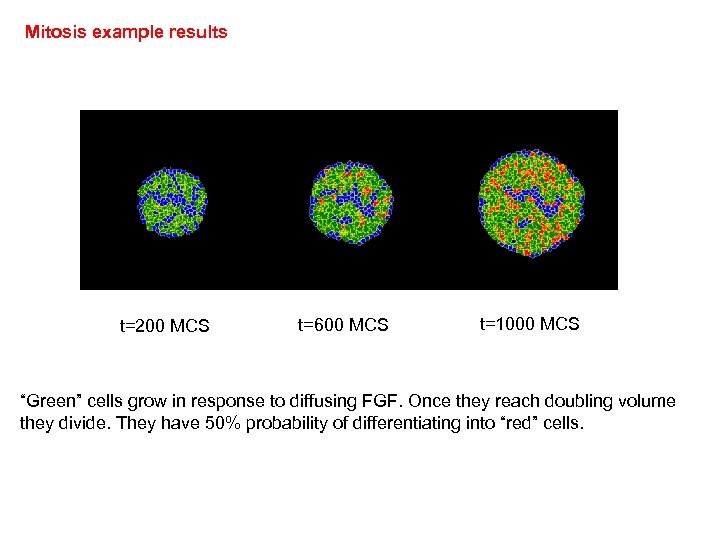 Mitosis example results t=200 MCS t=600 MCS t=1000 MCS “Green” cells grow in response to diffusing FGF. Once they reach doubling volume they divide. They have 50% probability of differentiating into “red” cells.
Mitosis example results t=200 MCS t=600 MCS t=1000 MCS “Green” cells grow in response to diffusing FGF. Once they reach doubling volume they divide. They have 50% probability of differentiating into “red” cells.
 Mitosis was our first example of a plugin implemented in Python. We can implement other plugins for example energy function in Python as well: class Volume. Energy. Function. Plugin(Energy. Function. Py): def __init__(self, _energy. Wrapper): # proper initialization Energy. Function. Py. __init__(self) self. energy. Wrapper=_energy. Wrapper self. vt=0. 0 self. lambda_v=0. 0 def set. Params(self, _lambda, _target. Volume): # configuration of the plugin self. lambda_v=_lambda; self. vt=_target. Volume def change. Energy(self): # core function of energy function plugin energy=0. 0 if(self. energy. Wrapper. new. Cell): energy+=self. lambda_v*(1+2*(self. energy. Wrapper. new. Cell. volume-self. vt)) if(self. energy. Wrapper. old. Cell): energy+=self. lambda_v*(1 -2*(self. energy. Wrapper. old. Cell. volume-self. vt)) return energy
Mitosis was our first example of a plugin implemented in Python. We can implement other plugins for example energy function in Python as well: class Volume. Energy. Function. Plugin(Energy. Function. Py): def __init__(self, _energy. Wrapper): # proper initialization Energy. Function. Py. __init__(self) self. energy. Wrapper=_energy. Wrapper self. vt=0. 0 self. lambda_v=0. 0 def set. Params(self, _lambda, _target. Volume): # configuration of the plugin self. lambda_v=_lambda; self. vt=_target. Volume def change. Energy(self): # core function of energy function plugin energy=0. 0 if(self. energy. Wrapper. new. Cell): energy+=self. lambda_v*(1+2*(self. energy. Wrapper. new. Cell. volume-self. vt)) if(self. energy. Wrapper. old. Cell): energy+=self. lambda_v*(1 -2*(self. energy. Wrapper. old. Cell. volume-self. vt)) return energy
 Full script: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes or plugins energy. Function. Registry=Compu. Cell. Setup. get. Energy. Function. Registry(sim) from cellsort_2 D_plugins import Volume. Energy. Function. Plugin volume. Energy=Volume. Energy. Function. Plugin(energy. Function. Registry) volume. Energy. set. Params(2. 0, 25. 0) energy. Function. Registry. register. Py. Energy. Function(volume. Energy) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)
Full script: import sys from os import environ from os import getcwd import string sys. path. append(environ["PYTHON_MODULE_PATH"]) sys. path. append(getcwd()+"/examples_Python. Tutorial") import Compu. Cell. Setup sim, simthread = Compu. Cell. Setup. get. Core. Simulation. Objects() #Create extra player fields here or add attributes or plugins energy. Function. Registry=Compu. Cell. Setup. get. Energy. Function. Registry(sim) from cellsort_2 D_plugins import Volume. Energy. Function. Plugin volume. Energy=Volume. Energy. Function. Plugin(energy. Function. Registry) volume. Energy. set. Params(2. 0, 25. 0) energy. Function. Registry. register. Py. Energy. Function(volume. Energy) Compu. Cell. Setup. initialize. Simulation. Objects(sim, simthread) #Add Python steppables here steppable. Registry=Compu. Cell. Setup. get. Steppable. Registry() Compu. Cell. Setup. main. Loop(sim, simthread, steppable. Registry)