a3eed46fa3f75db5081c453a95c9b191.ppt
- Количество слайдов: 48

Scientific Workflows: Some Examples and Technical Issues Bertram Ludäscher ludaesch@sdsc. edu Data & Knowledge Systems (DAKS) San Diego Supercomputer Center University of California, San Diego GGF 9 10/07/2003, Chicago

Outline • Scientific Workflow (SWF) Examples – Genomics: Promoter Identification (DOE Sci. DAC/SDM) – Neuroscience: Functional MRI (NIH BIRN) – Ecology: Invasive Species, Climate Change (NSF SEEK) • SWFs & Analysis Pipelines … – vs Business WFs – vs Traditional Distributed Computing • Some Technical Issues GGF 9 10/07/2003, Chicago

• NSF, NIH, DOE Acknowledgements • GEOsciences Network (NSF) – www. geongrid. org • Biomedical Informatics Research Network (NIH) – www. nbirn. net • Science Environment for Ecological Knowledge (NSF) – seek. ecoinformatics. org • Scientific Data Management Center (DOE) – sdm. lbl. gov/sdmcenter/ GGF 9 10/07/2003, Chicago

Scientific Workflow Examples 1. Promoter Identification (Genomics) 2. f. MRI (Neurosciences) 3. Invasive Species (Ecology) 4. Bonus Material: Semantic Data Integration (Geology) GGF 9 10/07/2003, Chicago
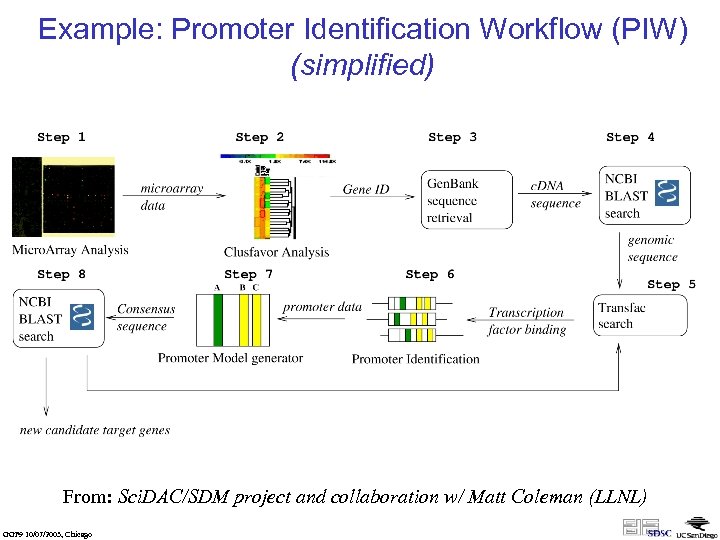
Example: Promoter Identification Workflow (PIW) (simplified) From: Sci. DAC/SDM project and collaboration w/ Matt Coleman (LLNL) GGF 9 10/07/2003, Chicago
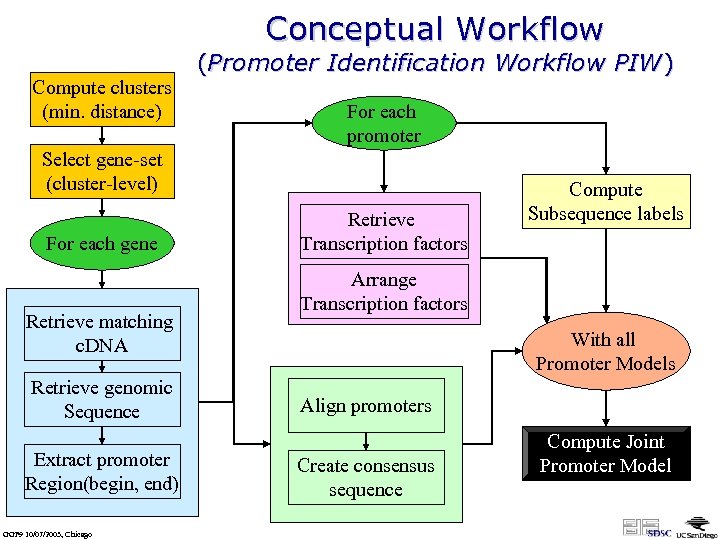
Conceptual Workflow Compute clusters (min. distance) (Promoter Identification Workflow PIW) For each promoter Select gene-set (cluster-level) For each gene Retrieve matching c. DNA Retrieve genomic Sequence Extract promoter Region(begin, end) GGF 9 10/07/2003, Chicago Retrieve Transcription factors Compute Subsequence labels Arrange Transcription factors With all Promoter Models Align promoters Create consensus sequence Compute Joint Promoter Model

SWF: Promoter Identification • More dataflow than workflow – but some branching looping, merging, … – not: documents/objects undergoing modifications – instead: dataset-out = analysis(dataset-in) • Need for “collection programming” (functional programming style) – – Iterations over lists (foreach) Filtering Functional composition Generic & higher-order operations (zip, map(f), …) • Need for abstraction and nested workflows • Need for rich user interaction / steering: – pause & resume – select & branch; e. g. , web browser capability at specific steps as part of a coordinated SWF • Need for persistence of intermediate products data provenance (virtual data concept, e. g. Gri. Phy. N) GGF 9 10/07/2003, Chicago

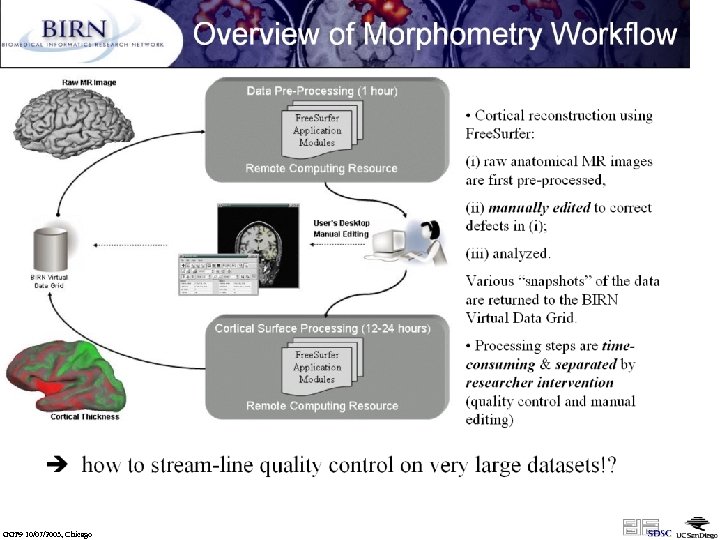
From: BIRN-CC, UCSD (Jeffrey Grethe) GGF 9 10/07/2003, Chicago
GGF 9 10/07/2003, Chicago
GGF 9 10/07/2003, Chicago

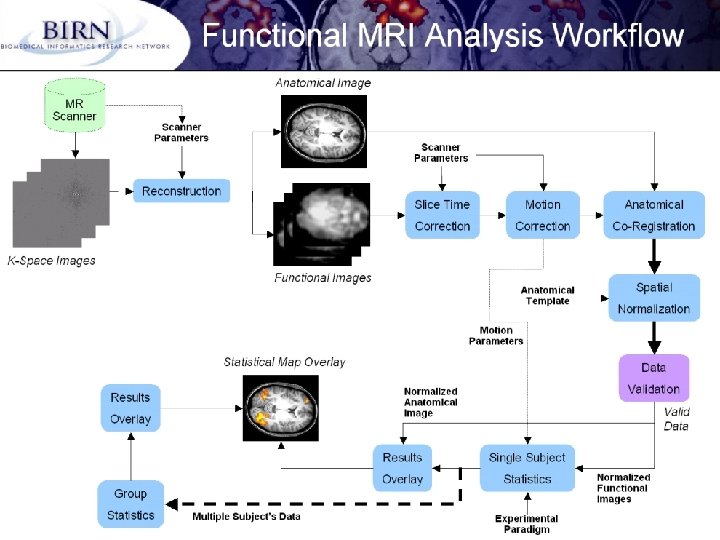
Details of the Functional MRI (Magnetic Resonance Imaging) Analysis Workflow (Jeffrey Grethe) 1. 2. 3. 4. Collect data (K-Space images in Fourier space) from MR scanner while subject performs a specific task Reconstruct K-Space data to image data (this requires scanner parameters for the reconstruction) Now have anatomical and functional data Pre-process the functional data 1. Correct for difference in slice acquisition (each slice in a volume is collected at a slightly different time). Try to correct for these differences so that all slices seem to be acquired at same time 2. Not correct for subject motion (head movement in scanner) by realigning all functional images 5. Register the functional images with the anatomical image all images are now in the same space (all aligned with one another) 6. Move all subjects into template space through non-linear spatial normalization. There exist atlas templates (made from many subjects) that one can normalize to so that all subjects are in the same space, allowing for direct comparison across subjects. 7. DATA VERIFICATION - check if all these procedures worked. If not, go back and try again (possibly tweaking some parameters for the routines or by re-doing some of it by hand). 8. Move onto statistics. First we do single subject statistics: in addition to the images, information about the experimental paradigm is required. These can be overlayed onto an anatomical to create visual displays of brain activation during a particular task. 9. Can also combine statistical data from multiple subjects and do a group/population analysis and display these results. Interactive nature of these workflows is critical (data verification) - can these steps be automated or semi-automated? need metadata from collection equipment and experimental design ! GGF 9 10/07/2003, Chicago
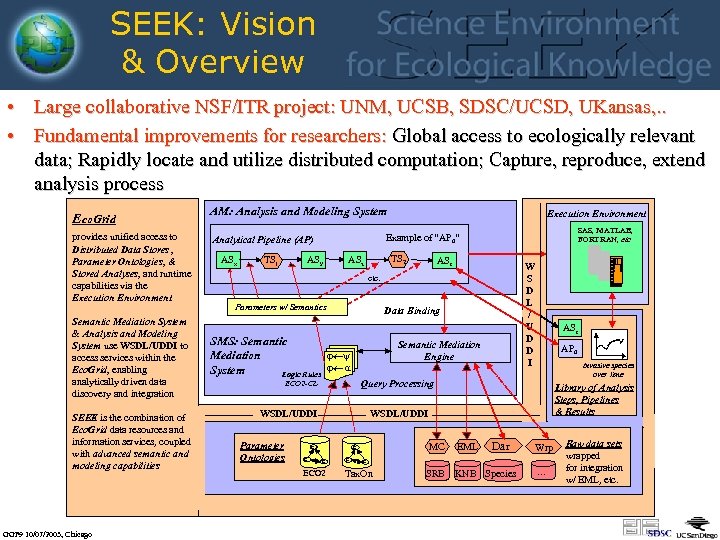
SEEK: Vision & Overview • Large collaborative NSF/ITR project: UNM, UCSB, SDSC/UCSD, UKansas, . . • Fundamental improvements for researchers: Global access to ecologically relevant data; Rapidly locate and utilize distributed computation; Capture, reproduce, extend analysis process Eco. Grid provides unified access to Distributed Data Stores , Parameter Ontologies, & Stored Analyses, and runtime capabilities via the Execution Environment Semantic Mediation System & Analysis and Modeling System use WSDL/UDDI to access services within the Eco. Grid, enabling analytically driven data discovery and integration SEEK is the combination of Eco. Grid data resources and information services, coupled with advanced semantic and modeling capabilities GGF 9 10/07/2003, Chicago AM: Analysis and Modeling System TS 1 SAS, MATLAB, FORTRAN, etc Example of “AP 0” Analytical Pipeline (AP) ASx Execution Environment ASy TS 2 ASz ASr W S D L / U D D I etc. Parameters w/ Semantics Data Binding SMS: Semantic j¬y Mediation j¬ a System Logic Rules Semantic Mediation Engine Invasive species over time Library of Analysis Steps, Pipelines & Results WSDL/UDDI ECO 2 C C AP 0 Query Processing ECO 2 -CL Parameter Ontologies ASr C ECO 2 C Dar MC C EML SRB KNB Species C Tax. On Wrp . . . Raw data sets wrapped for integration w/ EML, etc.
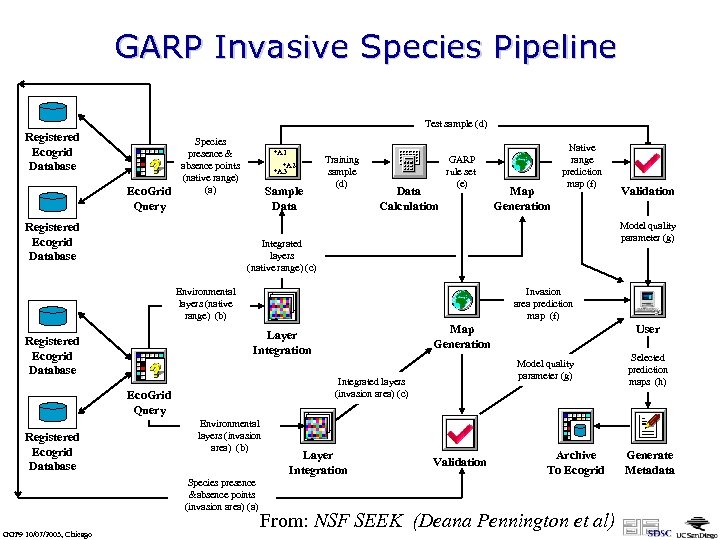
GARP Invasive Species Pipeline Test sample (d) Registered Ecogrid Database Eco. Grid Query Species presence & absence points (native range) (a) Registered Ecogrid Database +A 1 +A 2 +A 3 Sample Data Training sample (d) Data Calculation GARP rule set (e) Integrated layers (native range) (c) Invasion area prediction map (f) Environmental layers (invasion area) (b) Species presence &absence points (invasion area) (a) Layer Integration User Model quality parameter (g) Integrated layers (invasion area) (c) Eco. Grid Query GGF 9 10/07/2003, Chicago Map Generation Layer Integration Registered Ecogrid Database Validation Model quality parameter (g) Environmental layers (native range) (b) Registered Ecogrid Database Map Generation Native range prediction map (f) Validation Archive To Ecogrid From: NSF SEEK (Deana Pennington et al) Selected prediction maps (h) Generate Metadata

Details GARP Invasive Species Pipeline Modeling the distribution of a species using appropriate ecological niche modeling algorithms (e. g. , GARP—the Genetic Algorithm for Rule-set Production) in an analytical pipeline environment. a) The Eco. Grid is queried for data specifying the presence or absence of a particular species in a given area. b) Multiple environmental layers relevant to the specie’s distribution are selected with a second Eco. Grid query. c) Environmental layers, representing the current range of the species (native range) and the range of interest to the invasion study, are spatially-integrated into layer stacks. d) Samples are selected from the presence/absence data, and the corresponding values from the native range environmental layer stack are retrieved. The sample is divided into a training set and a testing set. e) The GARP algorithm is run on the training set. The GARP ruleset is then applied to both areas, creating predictive maps under current conditions (native range) and after invasion (invaded range). f) Predictive maps are sent to the scientist’s workstation for further analysis. g) A comparison is made between the ground truth occurrence data that were set aside as test data, and the corresponding location on the native range predictive map. Error measures providing an indication of model quality are sent to the scientist’s workstation. h) After multiple runs, the user may select maps for metadata generation and archiving back to the Eco. Grid. GGF 9 10/07/2003, Chicago From: NSF SEEK (Deana Pennington et al)
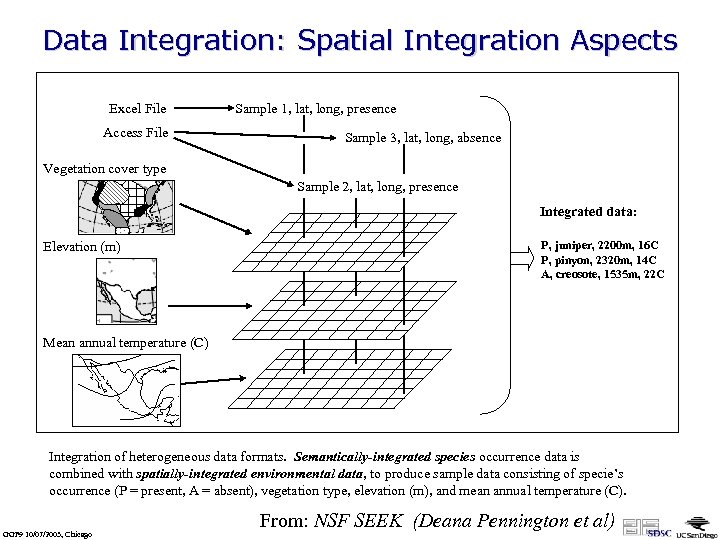
Data Integration: Spatial Integration Aspects Excel File Access File Sample 1, lat, long, presence Sample 3, lat, long, absence Vegetation cover type Sample 2, lat, long, presence Integrated data: Elevation (m) P, juniper, 2200 m, 16 C P, pinyon, 2320 m, 14 C A, creosote, 1535 m, 22 C Mean annual temperature (C) Integration of heterogeneous data formats. Semantically-integrated species occurrence data is combined with spatially-integrated environmental data, to produce sample data consisting of specie’s occurrence (P = present, A = absent), vegetation type, elevation (m), and mean annual temperature (C). GGF 9 10/07/2003, Chicago From: NSF SEEK (Deana Pennington et al)

SEEK Components • Eco. Grid • Seamless access to distributed, heterogeneous data: ecological, biodiversity, environmental data • “Semantically” mediated and metadata driven • Centralized search & management portal(s) • Analysis and Modeling System (AMS) – Capture, reproduce, and extend analysis process • Declarative means for documenting analysis • “Pipeline” system for linking generic analysis steps • Strong version control for analysis steps – Easy-to-use interface between data and analysis • Semantic Mediation System (SMS) – – “smart” data discovery, “type-correct” pipeline construction & data binding: determine whether/how to link analytic steps determine how data sets can be combined determine whether/how data sets are appropriate inputs for analysis steps GGF 9 10/07/2003, Chicago

AMS Overview • Objective – Create a semi-automated system for analyzing data and executing models that provides documentation, archiving, and versioning of the analyses, models, and their outputs (visual programming language? ) • Scope – Any type of analysis or model in ecology and biodiversity science – Massively streamline the analysis and modeling process – Archiving, rerunning analyses in SAS, Matlab, R, Sys. Stat, C(++), … – … GGF 9 10/07/2003, Chicago
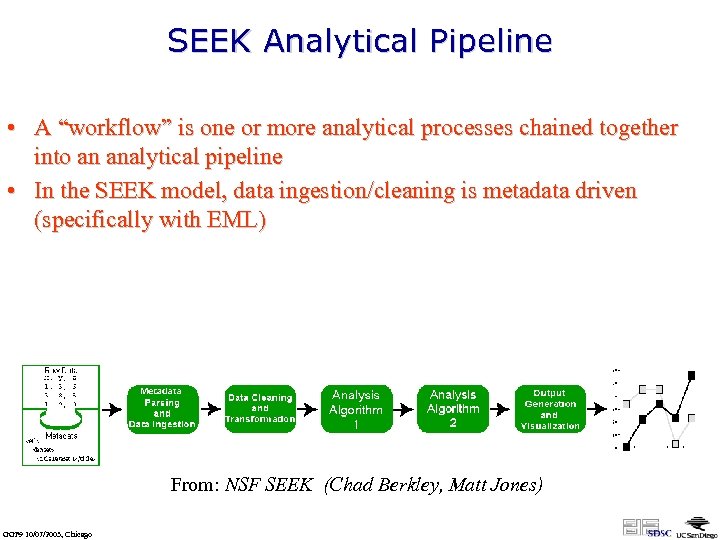
SEEK Analytical Pipeline • A “workflow” is one or more analytical processes chained together into an analytical pipeline • In the SEEK model, data ingestion/cleaning is metadata driven (specifically with EML) From: NSF SEEK (Chad Berkley, Matt Jones) GGF 9 10/07/2003, Chicago
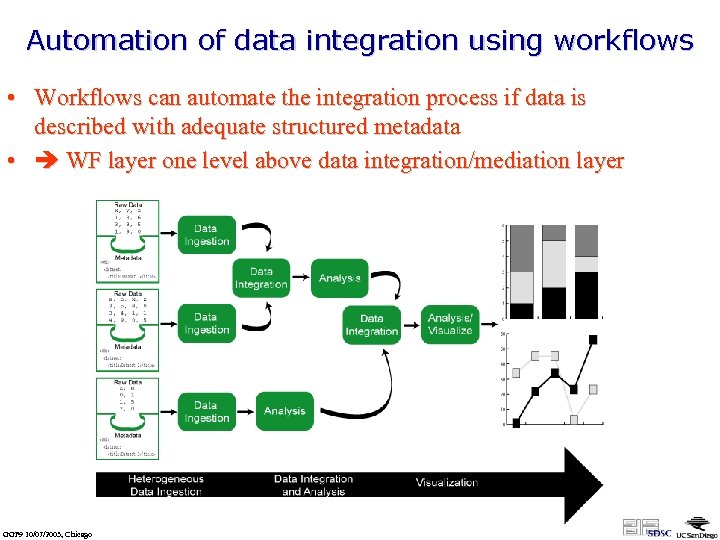
Automation of data integration using workflows • Workflows can automate the integration process if data is described with adequate structured metadata • WF layer one level above data integration/mediation layer GGF 9 10/07/2003, Chicago
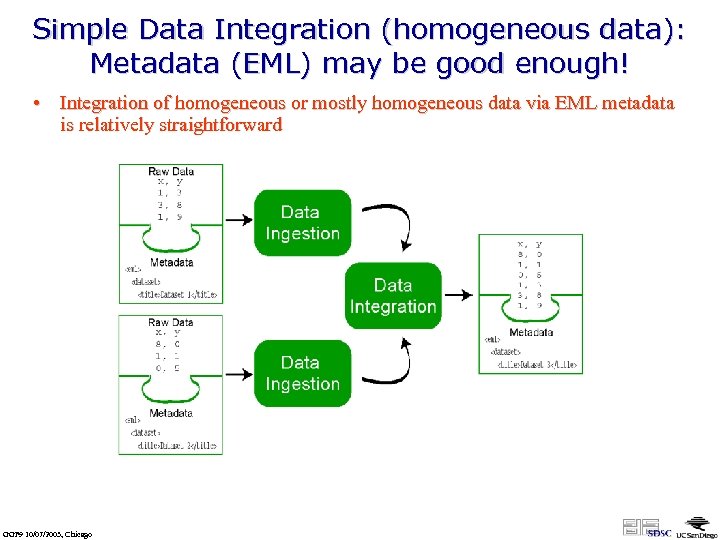
Simple Data Integration (homogeneous data): Metadata (EML) may be good enough! • Integration of homogeneous or mostly homogeneous data via EML metadata is relatively straightforward GGF 9 10/07/2003, Chicago
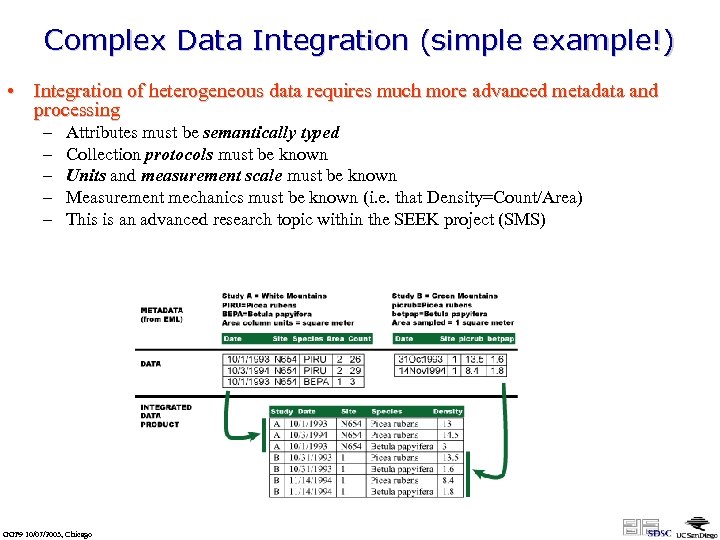

Complex Data Integration (simple example!) • Integration of heterogeneous data requires much more advanced metadata and processing – – – Attributes must be semantically typed Collection protocols must be known Units and measurement scale must be known Measurement mechanics must be known (i. e. that Density=Count/Area) This is an advanced research topic within the SEEK project (SMS) GGF 9 10/07/2003, Chicago
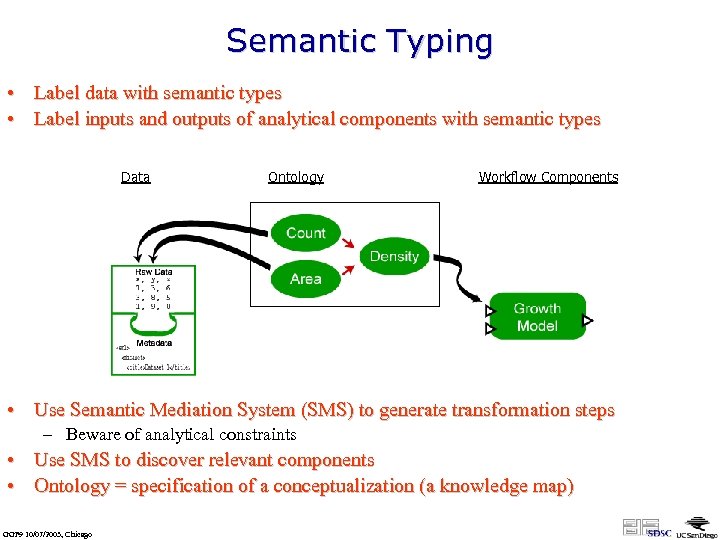
Semantic Typing • Label data with semantic types • Label inputs and outputs of analytical components with semantic types Data Ontology Workflow Components • Use Semantic Mediation System (SMS) to generate transformation steps – Beware of analytical constraints • Use SMS to discover relevant components • Ontology = specification of a conceptualization (a knowledge map) GGF 9 10/07/2003, Chicago

SWF: Ecology Examples • Similar requirements as before: – – – Rich user interaction Analysis pipelines running on an Eco. Grid Collection programming probably needed Abstraction & nested workflows Persistent intermediate steps (cf. e. g. Virtual Data concept) • Additionally: – Very heterogeneous data need for semantic typing of data and analysis steps semantic mediation support … • … for pipeline design • … for data integration at design time and at runtime GGF 9 10/07/2003, Chicago

Bonus Material: Semantic Data Integration “Geology Workbench” Kai Lin (GEON, SDSC) GGF 9 10/07/2003, Chicago
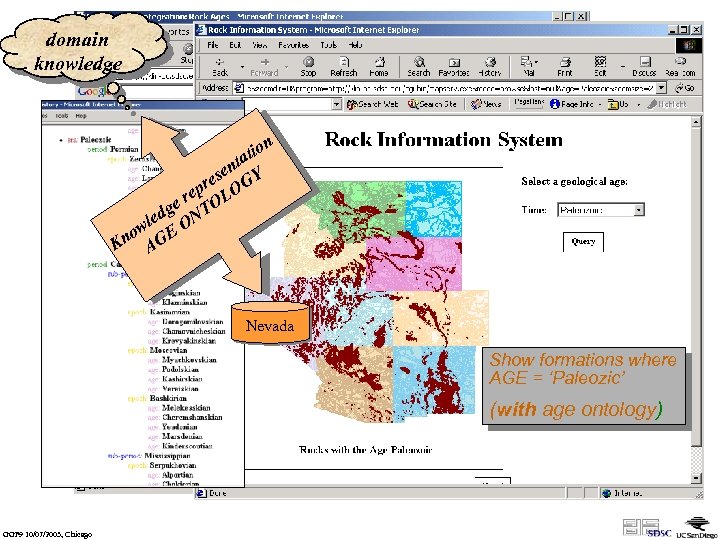
domain knowledge n tio ta en Y s pre LOG e e r TO g led ON w no GE K A Nevada Show formations where AGE = ‘Paleozic’ (without age ontology) (with GGF 9 10/07/2003, Chicago
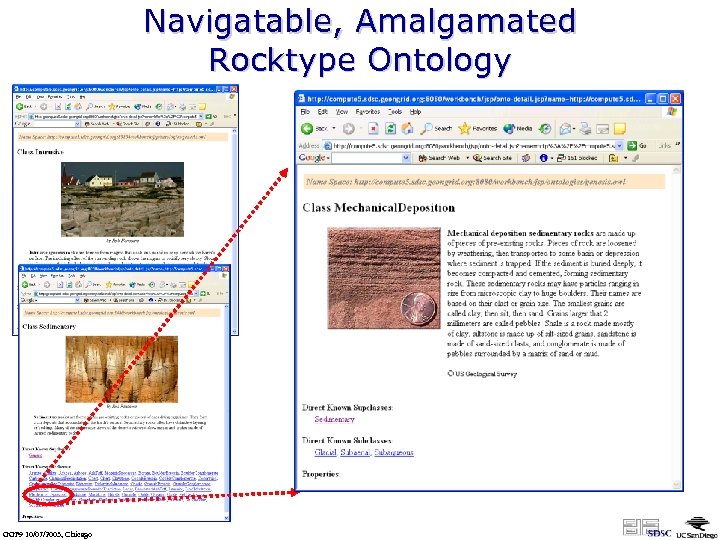
Navigatable, Amalgamated Rocktype Ontology GGF 9 10/07/2003, Chicago
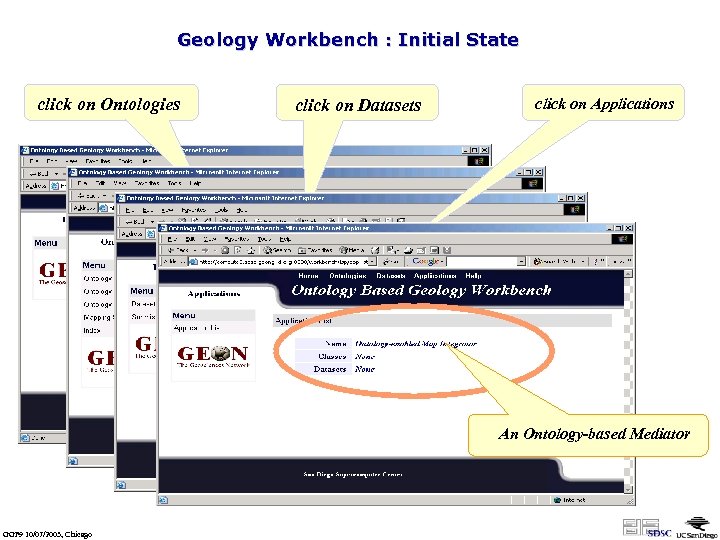
Geology Workbench : Initial State click on Ontologies click on Datasets click on Applications An Ontology-based Mediator GGF 9 10/07/2003, Chicago
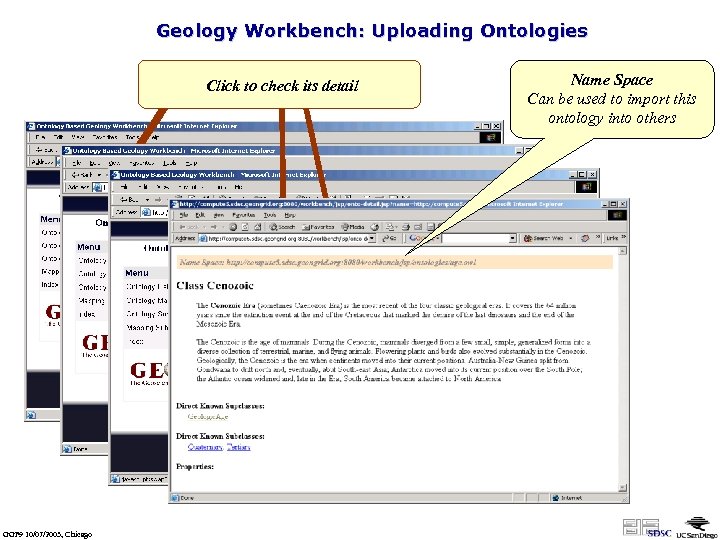
Geology Workbench: Uploading Ontologies click on Ontologyfile to upload Click to check its detail Choose an OWL Submission GGF 9 10/07/2003, Chicago Name Space Can be used to import this ontology into others
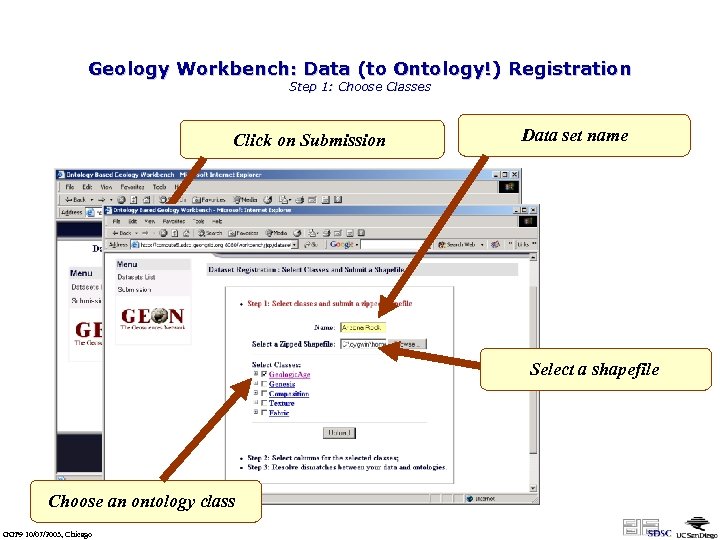
Geology Workbench: Data (to Ontology!) Registration Step 1: Choose Classes Click on Submission Data set name Select a shapefile Choose an ontology class GGF 9 10/07/2003, Chicago
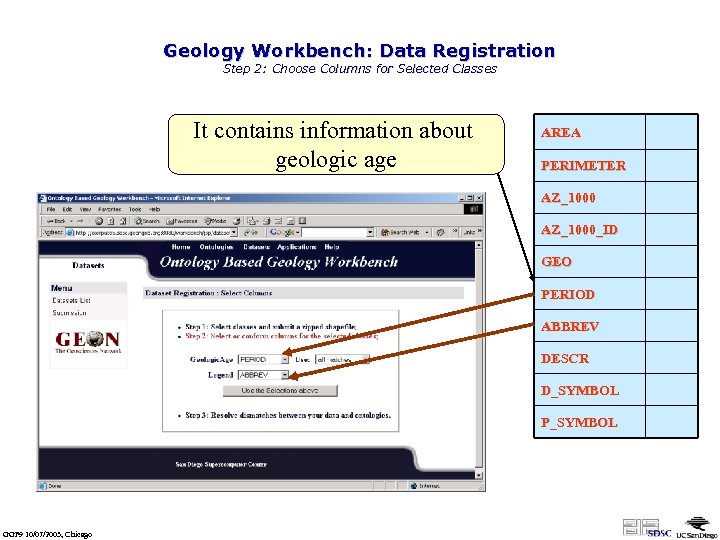
Geology Workbench: Data Registration Step 2: Choose Columns for Selected Classes It contains information about geologic age AREA PERIMETER AZ_1000_ID GEO PERIOD ABBREV DESCR D_SYMBOL P_SYMBOL GGF 9 10/07/2003, Chicago
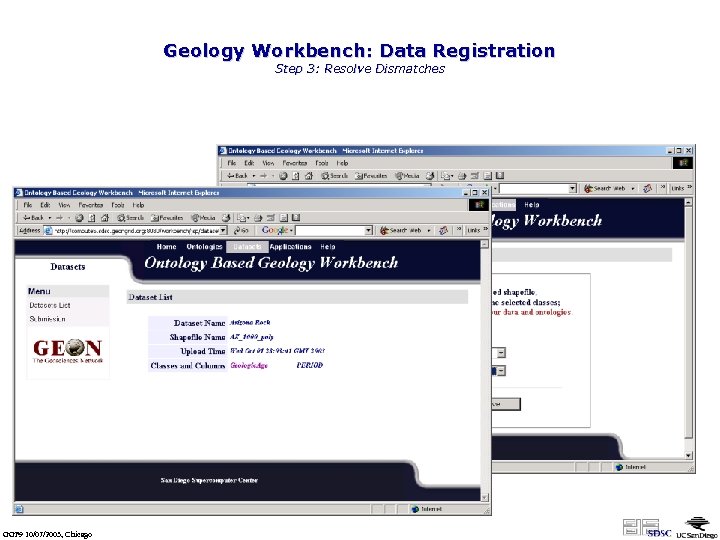
Geology Workbench: Data Registration Step 3: Resolve Dismatches Two terms are not matched any ontology terms Manually mapping algonkian into the ontology GGF 9 10/07/2003, Chicago
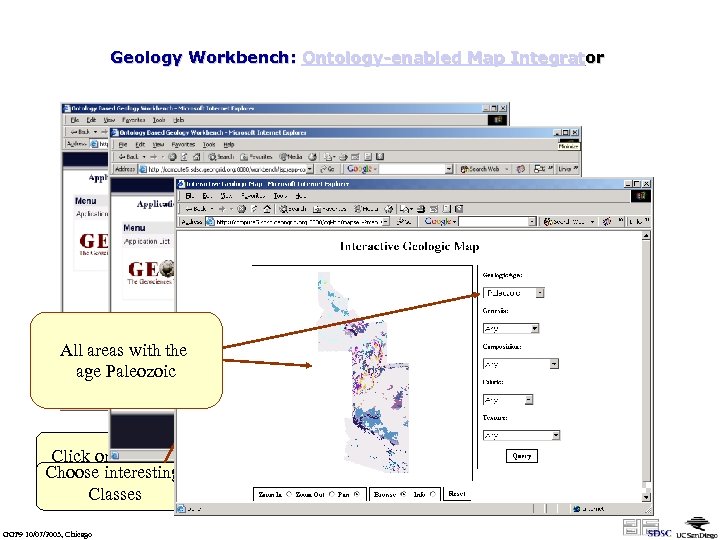
Geology Workbench: Ontology-enabled Map Integrator All areas with the age Paleozoic Click on the name Choose interesting Classes GGF 9 10/07/2003, Chicago
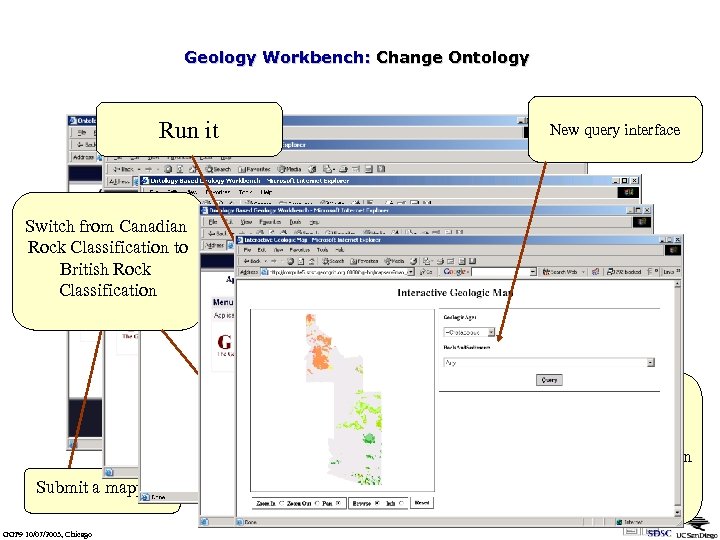
Geology Workbench: Change Ontology Run it New query interface Switch from Canadian Rock Classification to British Rock Classification Submit a mapping GGF 9 10/07/2003, Chicago Ontology mapping between British Rock Classification and Canadian Rock Classification

Scientific Workflows (SWF) vs. Business Workflows and some Technical Issues GGF 9 10/07/2003, Chicago
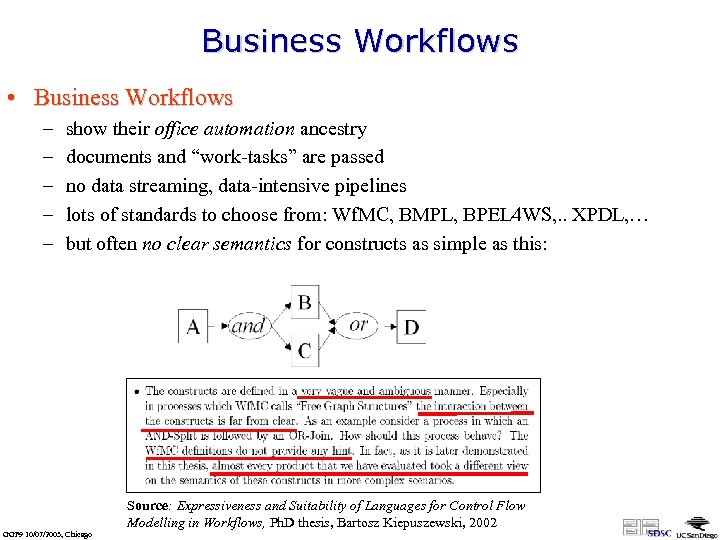

Business Workflows • Business Workflows – – – show their office automation ancestry documents and “work-tasks” are passed no data streaming, data-intensive pipelines lots of standards to choose from: Wf. MC, BMPL, BPEL 4 WS, . . XPDL, … but often no clear semantics for constructs as simple as this: Source: Expressiveness and Suitability of Languages for Control Flow Modelling in Workflows, Ph. D thesis, Bartosz Kiepuszewski, 2002 GGF 9 10/07/2003, Chicago

What is a Scientific Workflow? • A Misnomer … • … well, at least for a number of examples… • Scientific Workflows Business Workflows – Business Workflows: “control-flow-rich” – Scientific Workflows: “data-flow-rich” – … much more to say … GGF 9 10/07/2003, Chicago

More on Scientific WF vs Business WF • Business WF – Tasks, documents, etc. undergo modifications (e. g. , flight reservation from reserved to ticketed), but modified WF objects still identifiable throughout – Complex control flow, task-oriented – Transactions w/o rollback (ticket: reserved purchased) – … • SWF – data-in and data-out of an analysis step are not the same object! – dataflow, data-oriented (cf. AVS/Express, Khoros, …) – re-run automatically (a la distrib. comp. , e. g. Condor) or userdriven/interactively (based on failure type) – data integration & semantic mediation as part of SWF framework! – … GGF 9 10/07/2003, Chicago

SWF vs Distributed Computing • Distributed Computing (e. g. a la Condor-(G) ) – Batch oriented – Transparent distributed computing (“remote Unix/Java”; standard/Java universes in Condor) – HPC resource allocation & scheduling • SWF – Often highly interactive for decision making/steering of the WF and visualization (data analysis) – Transparent data access (Grid) and integration (database mediation & semantic extensions) – Desktop metaphor (“microworkflow”!? ); often (but not always!) light-weight web service invocation GGF 9 10/07/2003, Chicago

Some Technical Issues (SWFs) • Design Environment – Intuitive “visual programming” interface (ideally w/o the “programming” part!!) – “Smart Typing” extensions (for data task and task bindings) • • Structural typing (e. g. XML Schema) Semantic typing (e. g. OWL) Specialized semantic types (SI unit system, measurement scales, …) “resource typing”: token consumption/production, execution preconditions – Declarative programming extensions • Functional collection programming (e. g. , Haskell-like; cf. also Bio. Kleisli/CPL) • Also: consider what standards bring to the table (BPEL 4 WS) • Alternation of analysis and data transformation steps • Sophisticated dataflow execution models and hybrids thereof: – Ptolemy-II leads the way: Process Networks (PN), Synchronous Dataflow Networks (SDF), Continuous Time modeling (CT), Discrete Event modeling (DE) • Grid-enabling process networks and data integration: – Borrow from distributed computing technologies and tools (e. g. Globus, Condor) and distributed data access (e. g. , SRB) and integration (mediators) – Virtualize and Grid-enable everything! analysis@LOC, data@LOC, … GGF 9 10/07/2003, Chicago

Where do we go? • from Ptolemy-II to Kepler • example of what extensions are needed GGF 9 10/07/2003, Chicago
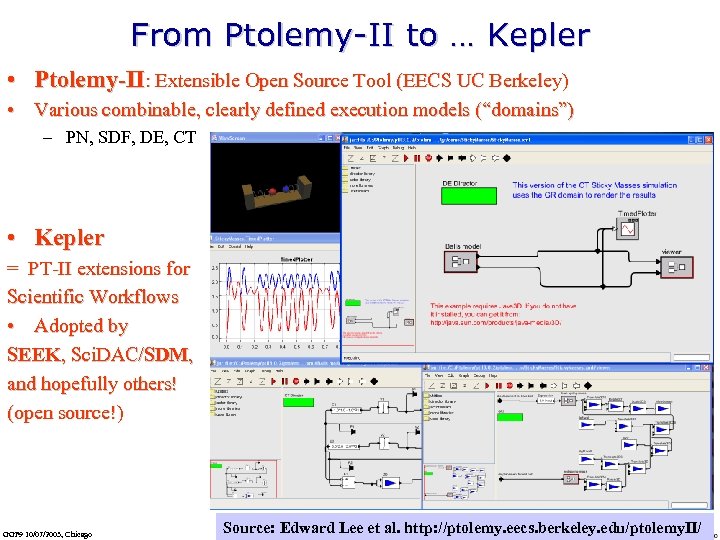
From Ptolemy-II to … Kepler • Ptolemy-II: Extensible Open Source Tool (EECS UC Berkeley) • Various combinable, clearly defined execution models (“domains”) – PN, SDF, DE, CT • Kepler = PT-II extensions for Scientific Workflows • Adopted by SEEK, Sci. DAC/SDM, and hopefully others! (open source!) GGF 9 10/07/2003, Chicago Source: Edward Lee et al. http: //ptolemy. eecs. berkeley. edu/ptolemy. II/
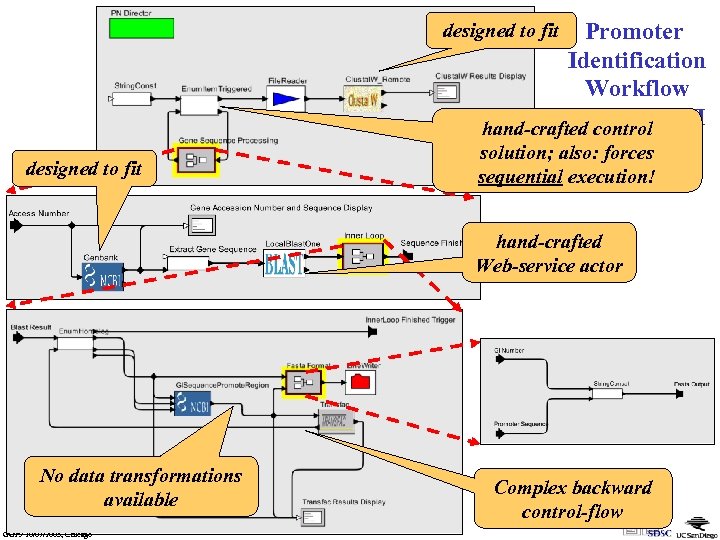
Promoter Identification Workflow in Ptolemy-II hand-crafted control (SSDBM’ 03) solution; also: forces designed to fit sequential execution! hand-crafted Web-service actor No data transformations available GGF 9 10/07/2003, Chicago Complex backward control-flow
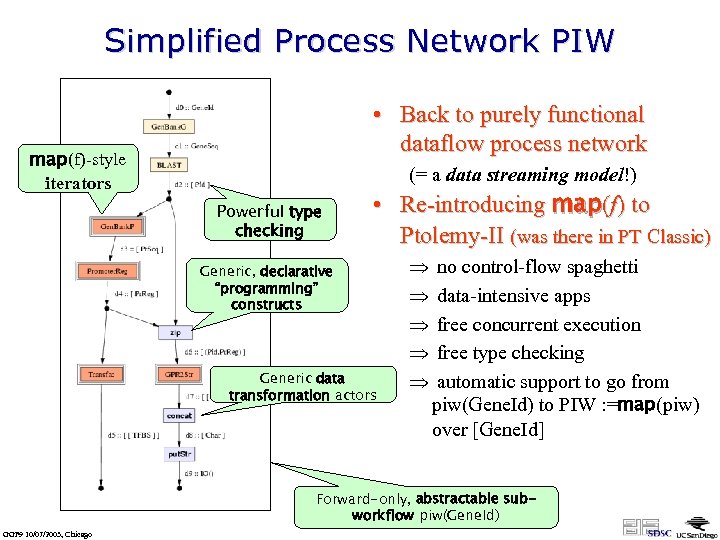
Simplified Process Network PIW • Back to purely functional dataflow process network map(f)-style iterators (= a data streaming model!) Powerful type checking • Re-introducing map(f) to Ptolemy-II (was there in PT Classic) Generic, declarative “programming” constructs Generic data transformation actors Þ no control-flow spaghetti Þ data-intensive apps Þ free concurrent execution Þ free type checking Þ automatic support to go from piw(Gene. Id) to PIW : =map(piw) over [Gene. Id] Forward-only, abstractable subworkflow piw(Gene. Id) GGF 9 10/07/2003, Chicago
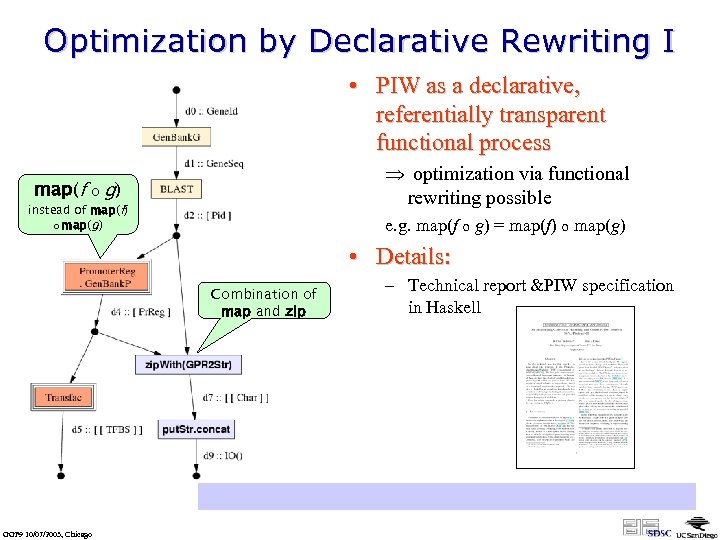
Optimization by Declarative Rewriting I • PIW as a declarative, referentially transparent functional process map(f o Þ optimization via functional rewriting possible g) instead of map(f) o map(g) e. g. map(f o g) = map(f) o map(g) • Details: Combination of map and zip – Technical report &PIW specification in Haskell http: //kbi. sdsc. edu/Sci. DAC-SDM/scidac-tn-map-constructs. pdf GGF 9 10/07/2003, Chicago
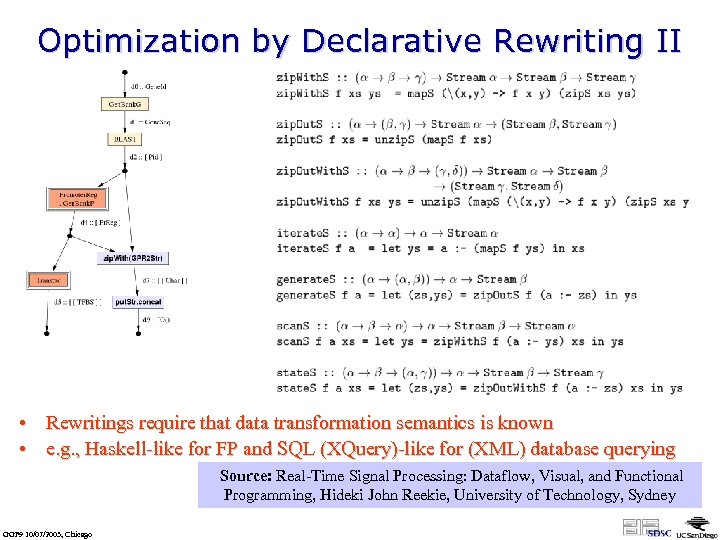

Optimization by Declarative Rewriting II • Rewritings require that data transformation semantics is known • e. g. , Haskell-like for FP and SQL (XQuery)-like for (XML) database querying Source: Real-Time Signal Processing: Dataflow, Visual, and Functional Programming, Hideki John Reekie, University of Technology, Sydney GGF 9 10/07/2003, Chicago

Summary: Scientific Workflows Everywhere • Shown bits scientific workflows in: – Sci. DAC/SDM, SEEK, BIRN, GEON, … • Many others are there: – Gri. Phy. N et al (virtual data concept): Chimera, Pegasus, DAGman, Condor. G, … Grid. ANT, … – E-Science: e. g, my. Grid: XScufl, Taverna, Discovery. Net – Pragma, i. LTER, . . – Commercial efforts: Discovery. Net (inforsense), Scitegic, IBM, Oracle, … • One size fits all? – Most likely not (Business WFs =/= Scientific WFs) – Some competition is healthy and reinventing a round wheel is OK – But some coordination & collaboration can save … • reinventing the squared wheel • “leveraging” someone else’s wheel in a squared way … – Even within SWF, quite different requirements: • exploratory and ad-hoc vs. well-designed and high throughput • interactive desktop (w/ lightweight web services/Grid) vs. distributed, batched GGF 9 10/07/2003, Chicago

Combine Everything: Die eierlegende Wollmilchsau: • Database Federation/Mediation – query rewriting under GAV/LAV – w/ binding pattern constraints – distributed query processing • Semantic Mediation – semantic integrity constraints, reasoning w/ plans, automated deduction – deductive database/logic programming technology, AI “stuff”. . . – Semantic Web technology (OWL, …) • Scientific Workflow Management – more procedural than database mediation (often the scientist is the query planner) – deployment using grid services! GGF 9 10/07/2003, Chicago

FIN GGF 9 10/07/2003, Chicago
a3eed46fa3f75db5081c453a95c9b191.ppt