518c44d970ca33c2d764d46a360ca282.ppt
- Количество слайдов: 27

Rosalind Elsie Franklin o Biophysicist and crystallographer o X-ray diffraction images of DNA o Tobacco mosaic and polio viruses o 1920 -1958 (source: wikipedia)

A Structural Split in the Human Genome Clara S. M. Tang and Richard J. Epstein PLo. S One (2007) 7: e 603 February 13, 2007 I. Elizabeth Cha

Introduction o PCIs Promoter-associated Cp. G islands o Mediate methylation-dependent gene silencing o Co-locate to transcriptionally active genes o 60% of human genes contains PCIs

Cp. G Islands o Genomic regions containing high frequency of CG dinucleotides o Cp. G cytidine-phosphodiester-guanosine o Formal definition n At least 200 bp n GC percentage >50% n Cp. G ratio >60%
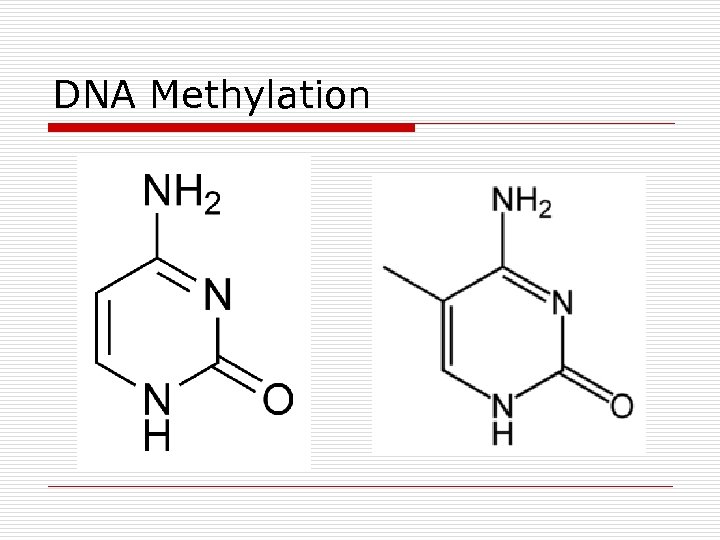
DNA Methylation

Materials and Methods o Sequence data and annotations o Determination of Cp. G island overlapping transcription start site o Housekeeping genes and paralogs of pseudogenes o Bimodal distribution of GC content o Gene expression data o Evolutionary rate determination o Principal component analysis

Sequence Data and Annotations o UCSC genomic assemblies, Ref. Seq dataset, Emsembl gene dataset n n n Human (hg 18, 3/2006) Mouse (mm 6, 3/2006) Fugu (fr 1, 8/2002) Fruit fly (dm 2, 4/2004) Worm (ce 2, 3/2004)

Data Preprocessing o Repeat. Mask – Alu o Discard sequences n Not commencing with ATG codons n Not terminating with canonical stop codons o Retain the longest genomic sequences containing identical exonic sequences

Determination of Cp. G Island Overlapping Transcription Start Site o Download Cp. G islands annotation (cpg. Island. Ext) from UCSC o Identify Cp. G islands overlapping with promoter regions o Map with Ref. Gene annotation (200 bp upstream and 500 bp downstream)

Data and Tools o 502 Housekeeping genes o 1220 pseudogene paralogs o o NOCOM program SAGEmap Homologue data XSTAT

Results – PCI+ Genes o Housekeeping gene higher GC content lower intron length/number o Pseudogene paralog lower GC content higher intron length/number o Functional distinguishable
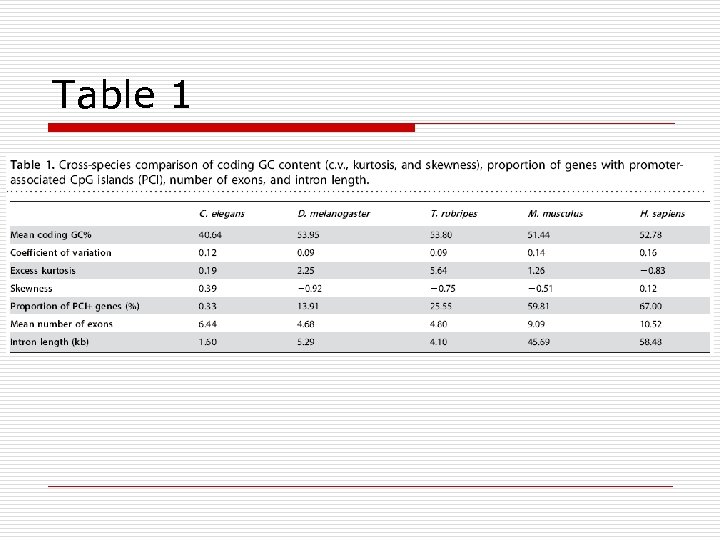
Table 1

Results – PCI- Genes o Higher evolutionary rate o Narrower expression breadth than PCI+ genes o More frequent tissue-specific inactivation
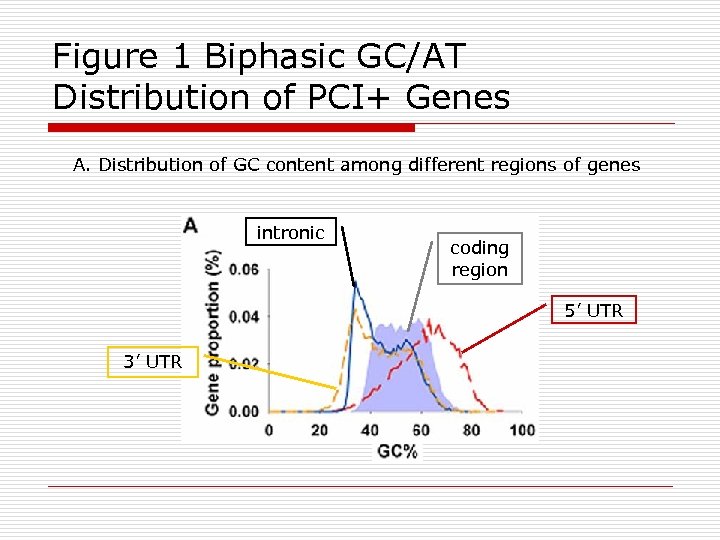
Figure 1 Biphasic GC/AT Distribution of PCI+ Genes A. Distribution of GC content among different regions of genes intronic coding region 5’ UTR 3’ UTR
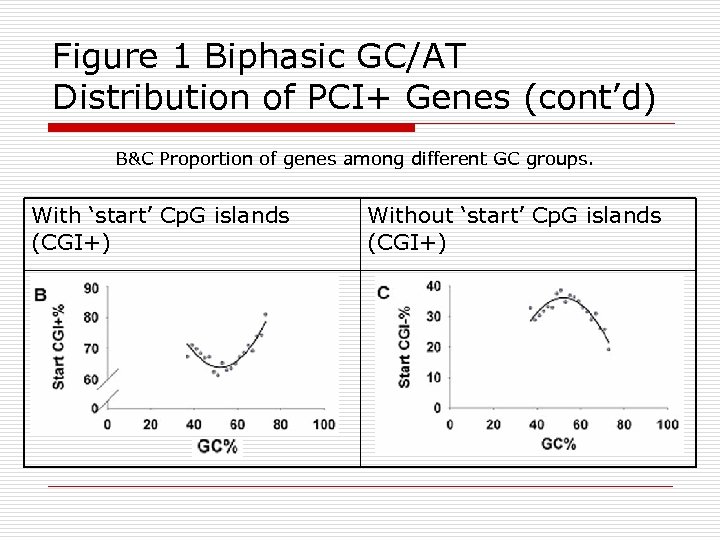
Figure 1 Biphasic GC/AT Distribution of PCI+ Genes (cont’d) B&C Proportion of genes among different GC groups. With ‘start’ Cp. G islands (CGI+) Without ‘start’ Cp. G islands (CGI+)
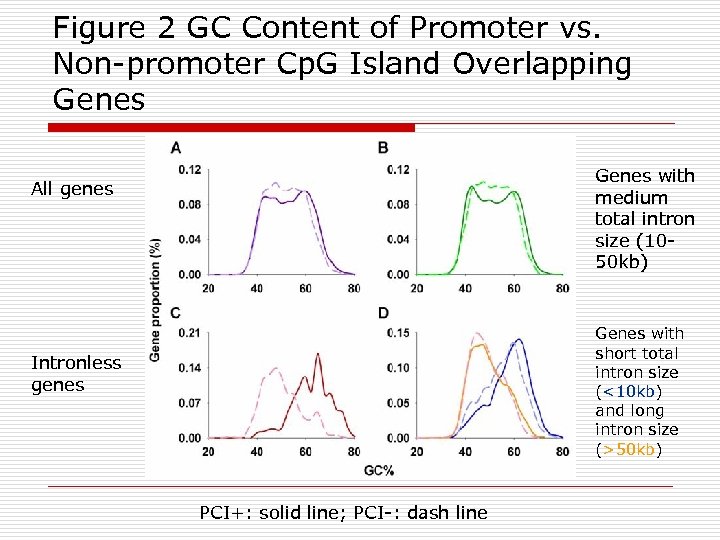
Figure 2 GC Content of Promoter vs. Non-promoter Cp. G Island Overlapping Genes with medium total intron size (1050 kb) All genes Genes with short total intron size (<10 kb) and long intron size (>50 kb) Intronless genes PCI+: solid line; PCI-: dash line
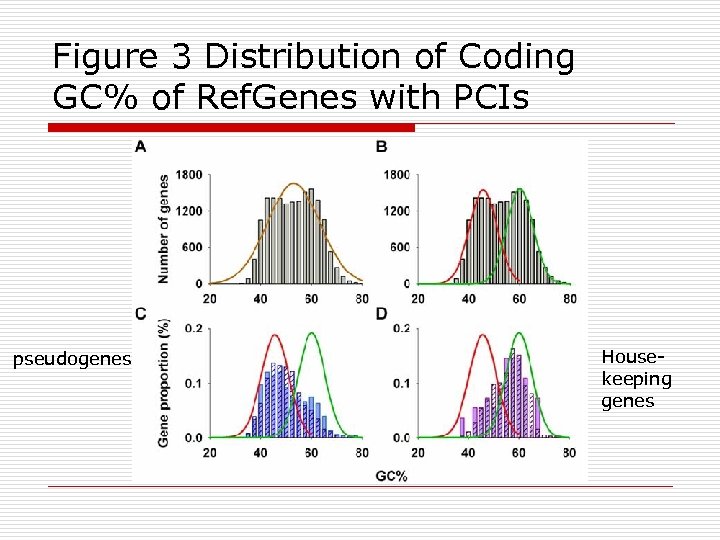
Figure 3 Distribution of Coding GC% of Ref. Genes with PCIs pseudogenes Housekeeping genes
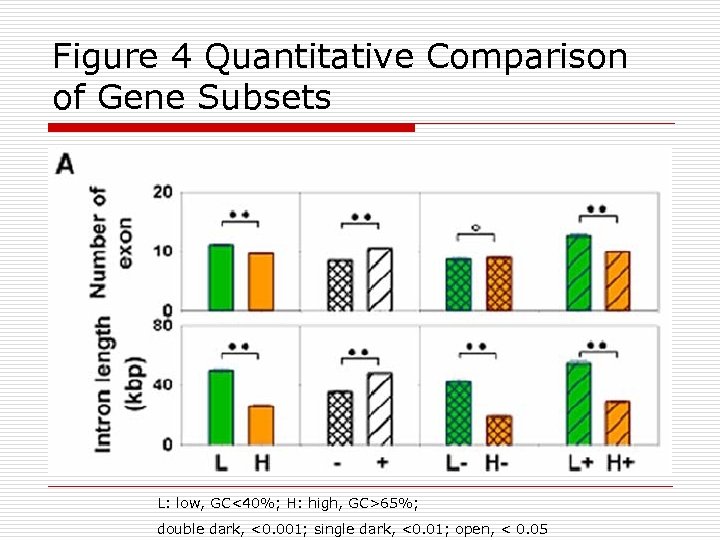
Figure 4 Quantitative Comparison of Gene Subsets L: low, GC<40%; H: high, GC>65%; double dark, <0. 001; single dark, <0. 01; open, < 0. 05
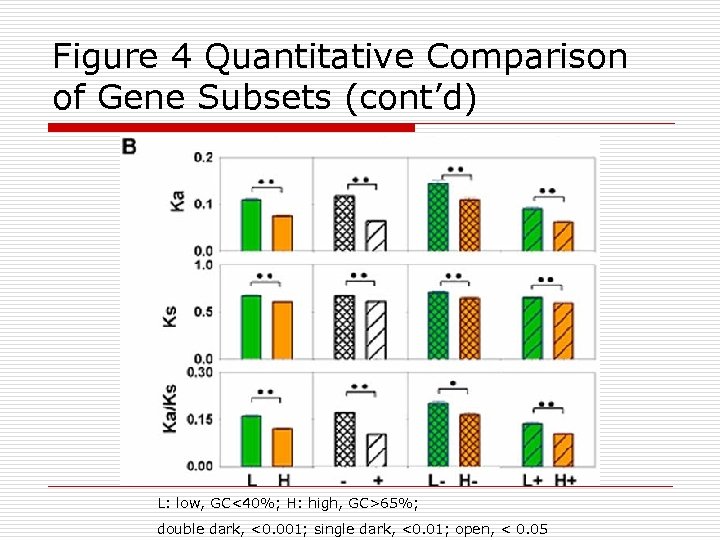
Figure 4 Quantitative Comparison of Gene Subsets (cont’d) L: low, GC<40%; H: high, GC>65%; double dark, <0. 001; single dark, <0. 01; open, < 0. 05
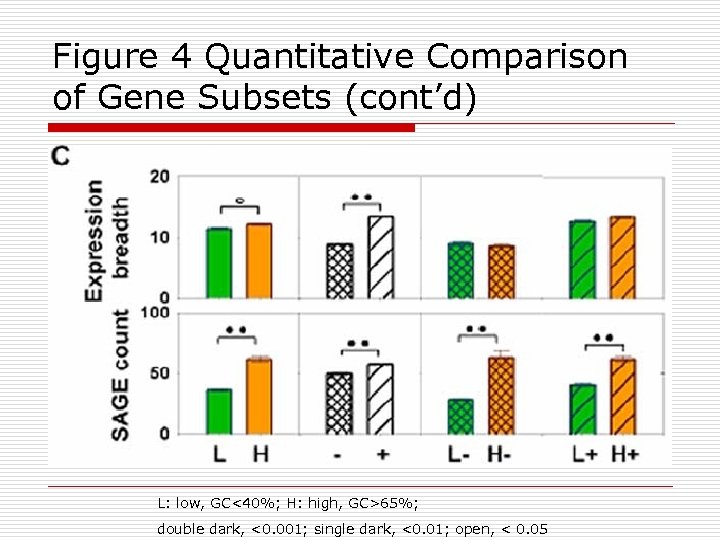
Figure 4 Quantitative Comparison of Gene Subsets (cont’d) L: low, GC<40%; H: high, GC>65%; double dark, <0. 001; single dark, <0. 01; open, < 0. 05
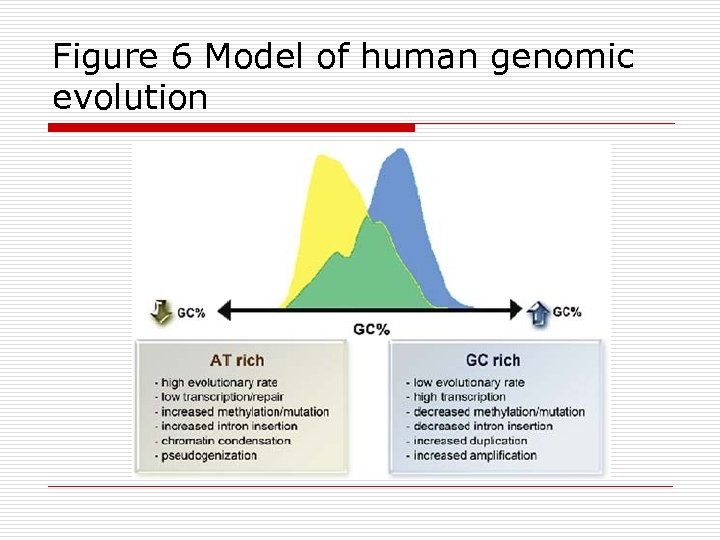
Figure 6 Model of human genomic evolution

Conclusions o PCIs n Transcriptional regulators n Evolutionary accelerators to facilitate intron insertion o Mthylated PCIs on transcription and chromatin accelerate adaptive evolution towards biological complexity

Conclusions o Adaptive evolution of human genome n Declining transcription of a subset of PCI+ genes n Predisposing to both Cp. G Tp. A mutation and intron insertion o Biological complexity model n Environmentally selected gains/losses of PCI methylation (+/-) n Polarizing PCI+ gene structures arounda genomic core of ancestral PCI- genes

Discussion o AT-rich, PCI+ gene vs. GC-rich PCI+ housekeeping gene n Lower transcriptional activity n Higher intron number n Higher evolutionary rate o Loss of negative selection pressure

Discussion (cont’d) o PCI- genes vs. PCI+ genes n Higher evolutionary rate n Lower expression breadth o Intron number relates more directly to PCI positivity
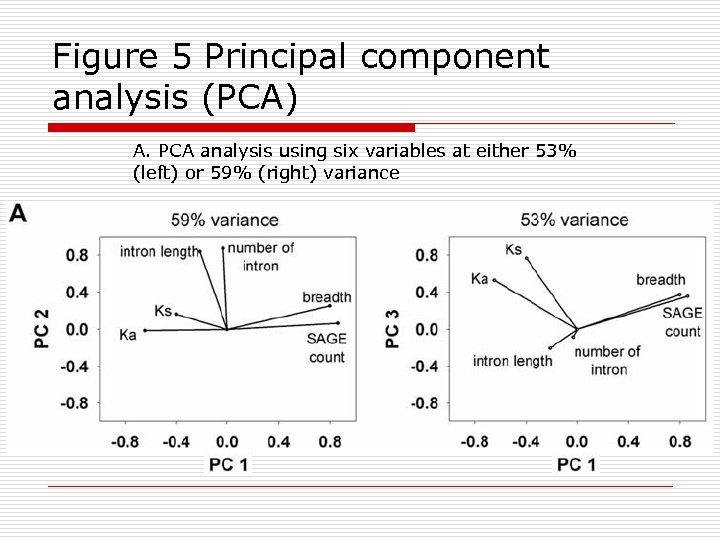
Figure 5 Principal component analysis (PCA) A. PCA analysis using six variables at either 53% (left) or 59% (right) variance
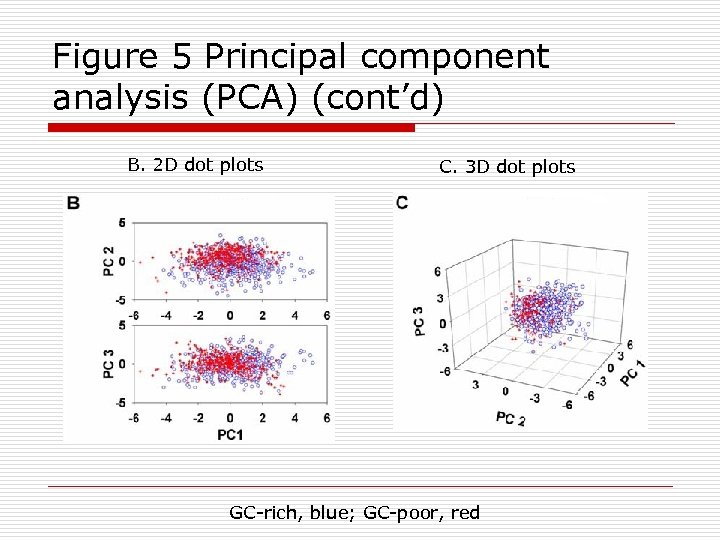
Figure 5 Principal component analysis (PCA) (cont’d) B. 2 D dot plots C. 3 D dot plots GC-rich, blue; GC-poor, red
518c44d970ca33c2d764d46a360ca282.ppt