2a12d90518e7b3d19e0ac0c4575a6fcc.ppt
- Количество слайдов: 30
 Results of donor screening with nucleic acid amplification tests (NAT) and implications for HIV research, diagnosis and surveillance Michael P. Busch, MD, Ph. D Professor of Laboratory Medicine, UCSF Director, Blood Systems Research Institute
Results of donor screening with nucleic acid amplification tests (NAT) and implications for HIV research, diagnosis and surveillance Michael P. Busch, MD, Ph. D Professor of Laboratory Medicine, UCSF Director, Blood Systems Research Institute
 Overview of presentation 1. RNA dynamics in primary HIV infection and projected yield of NAT in low and high incidence populations 2. Design of NAT assays for blood screening 3. Yield and cost effectiveness of NAT screening of blood donors 4. Applications of donor NAT assays in research settings
Overview of presentation 1. RNA dynamics in primary HIV infection and projected yield of NAT in low and high incidence populations 2. Design of NAT assays for blood screening 3. Yield and cost effectiveness of NAT screening of blood donors 4. Applications of donor NAT assays in research settings
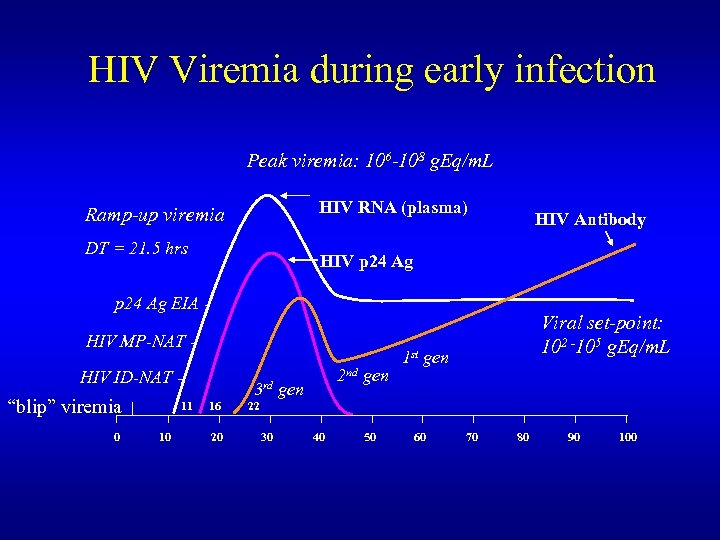 HIV Viremia during early infection Peak viremia: 106 -108 g. Eq/m. L HIV RNA (plasma) Ramp-up viremia DT = 21. 5 hrs HIV Antibody HIV p 24 Ag EIA HIV MP-NAT HIV ID-NAT - “blip” viremia 0 11 10 2 nd gen 3 rd gen 16 20 Viral set-point: 102 -105 g. Eq/m. L 1 st gen 22 30 40 50 60 70 80 90 100
HIV Viremia during early infection Peak viremia: 106 -108 g. Eq/m. L HIV RNA (plasma) Ramp-up viremia DT = 21. 5 hrs HIV Antibody HIV p 24 Ag EIA HIV MP-NAT HIV ID-NAT - “blip” viremia 0 11 10 2 nd gen 3 rd gen 16 20 Viral set-point: 102 -105 g. Eq/m. L 1 st gen 22 30 40 50 60 70 80 90 100
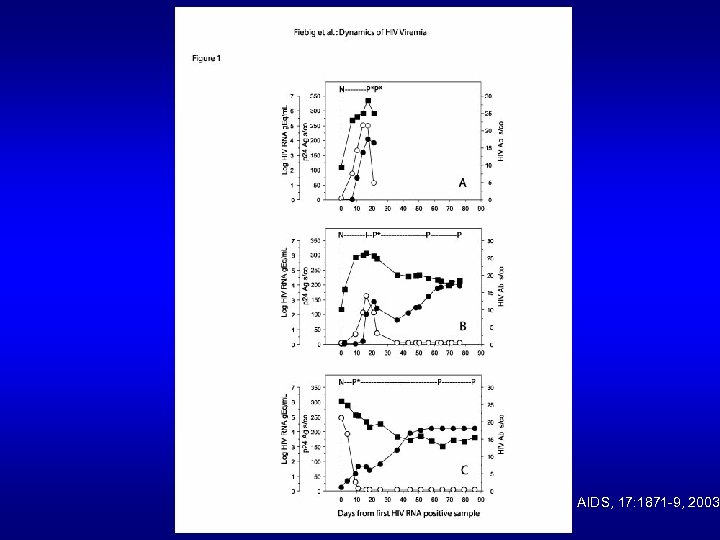 AIDS, 17: 1871 -9, 2003.
AIDS, 17: 1871 -9, 2003.
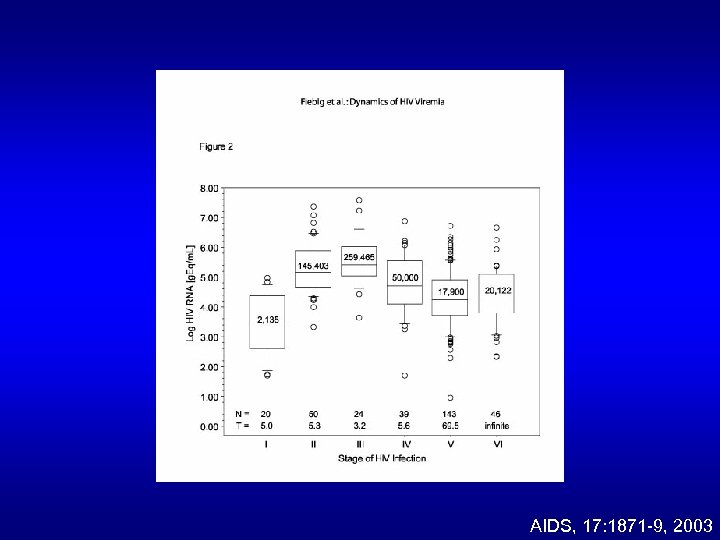 AIDS, 17: 1871 -9, 2003
AIDS, 17: 1871 -9, 2003
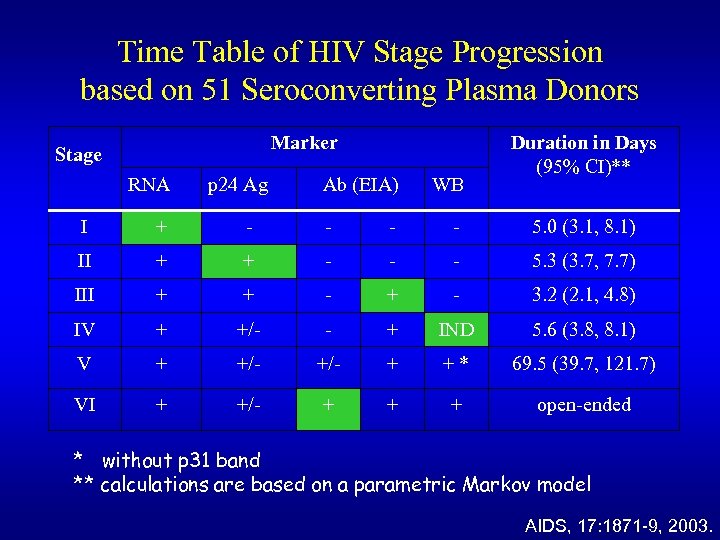 Time Table of HIV Stage Progression based on 51 Seroconverting Plasma Donors Marker Stage RNA p 24 Ag Ab (EIA) WB Duration in Days (95% CI)** I + - - 5. 0 (3. 1, 8. 1) II + + - - - 5. 3 (3. 7, 7. 7) III + + - 3. 2 (2. 1, 4. 8) IV + +/- - + IND 5. 6 (3. 8, 8. 1) V + +/- + +* 69. 5 (39. 7, 121. 7) VI + +/- + + + open-ended * without p 31 band ** calculations are based on a parametric Markov model AIDS, 17: 1871 -9, 2003.
Time Table of HIV Stage Progression based on 51 Seroconverting Plasma Donors Marker Stage RNA p 24 Ag Ab (EIA) WB Duration in Days (95% CI)** I + - - 5. 0 (3. 1, 8. 1) II + + - - - 5. 3 (3. 7, 7. 7) III + + - 3. 2 (2. 1, 4. 8) IV + +/- - + IND 5. 6 (3. 8, 8. 1) V + +/- + +* 69. 5 (39. 7, 121. 7) VI + +/- + + + open-ended * without p 31 band ** calculations are based on a parametric Markov model AIDS, 17: 1871 -9, 2003.
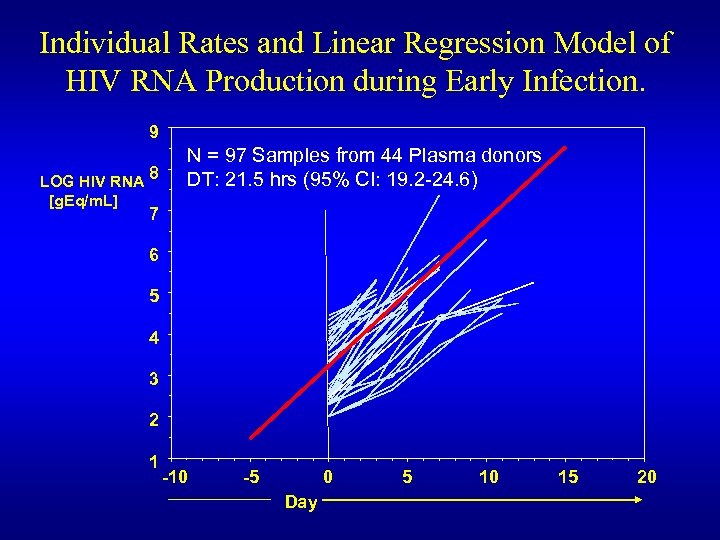 Individual Rates and Linear Regression Model of HIV RNA Production during Early Infection. 9 LOG HIV RNA 8 [g. Eq/m. L] N = 97 Samples from 44 Plasma donors DT: 21. 5 hrs (95% CI: 19. 2 -24. 6) 7 6 5 4 3 2 1 -10 -5 0 Day 5 10 15 20
Individual Rates and Linear Regression Model of HIV RNA Production during Early Infection. 9 LOG HIV RNA 8 [g. Eq/m. L] N = 97 Samples from 44 Plasma donors DT: 21. 5 hrs (95% CI: 19. 2 -24. 6) 7 6 5 4 3 2 1 -10 -5 0 Day 5 10 15 20
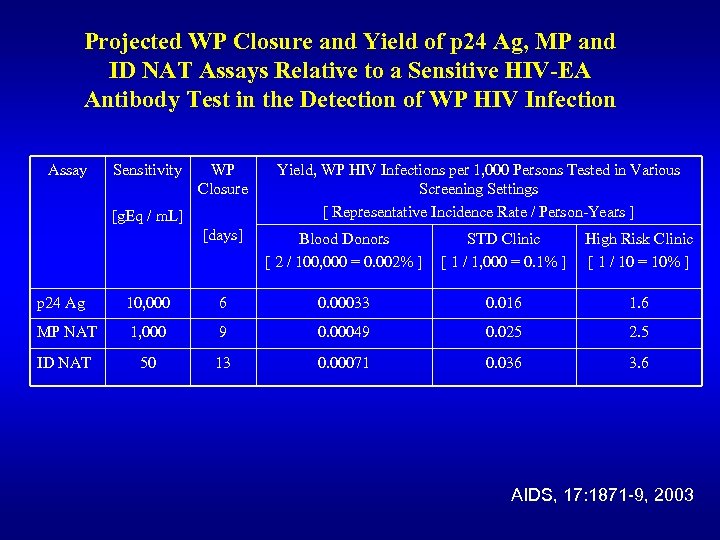 Projected WP Closure and Yield of p 24 Ag, MP and ID NAT Assays Relative to a Sensitive HIV-EA Antibody Test in the Detection of WP HIV Infection Assay Sensitivity WP Closure [g. Eq / m. L] Yield, WP HIV Infections per 1, 000 Persons Tested in Various Screening Settings [ Representative Incidence Rate / Person-Years ] [days] Blood Donors [ 2 / 100, 000 = 0. 002% ] STD Clinic [ 1 / 1, 000 = 0. 1% ] High Risk Clinic [ 1 / 10 = 10% ] p 24 Ag 10, 000 6 0. 00033 0. 016 1. 6 MP NAT 1, 000 9 0. 00049 0. 025 2. 5 ID NAT 50 13 0. 00071 0. 036 3. 6 AIDS, 17: 1871 -9, 2003
Projected WP Closure and Yield of p 24 Ag, MP and ID NAT Assays Relative to a Sensitive HIV-EA Antibody Test in the Detection of WP HIV Infection Assay Sensitivity WP Closure [g. Eq / m. L] Yield, WP HIV Infections per 1, 000 Persons Tested in Various Screening Settings [ Representative Incidence Rate / Person-Years ] [days] Blood Donors [ 2 / 100, 000 = 0. 002% ] STD Clinic [ 1 / 1, 000 = 0. 1% ] High Risk Clinic [ 1 / 10 = 10% ] p 24 Ag 10, 000 6 0. 00033 0. 016 1. 6 MP NAT 1, 000 9 0. 00049 0. 025 2. 5 ID NAT 50 13 0. 00071 0. 036 3. 6 AIDS, 17: 1871 -9, 2003
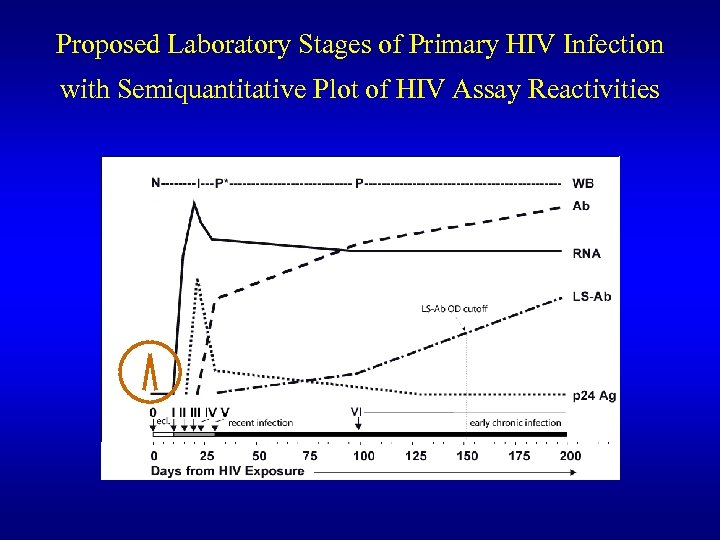 Proposed Laboratory Stages of Primary HIV Infection with Semiquantitative Plot of HIV Assay Reactivities
Proposed Laboratory Stages of Primary HIV Infection with Semiquantitative Plot of HIV Assay Reactivities
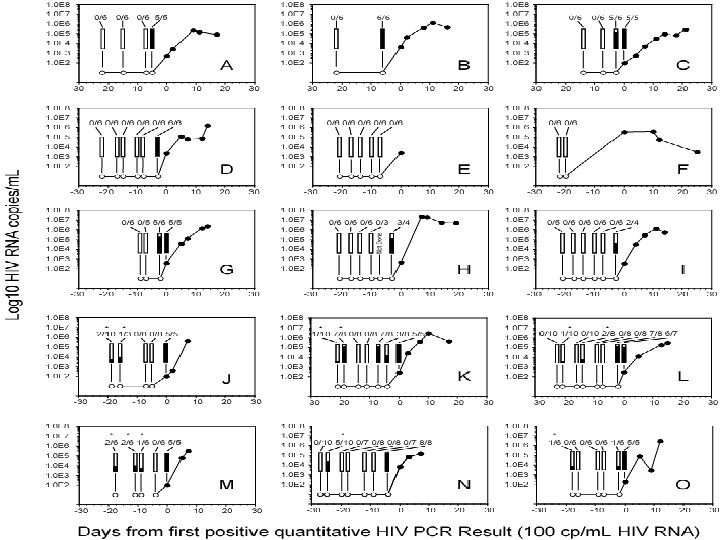
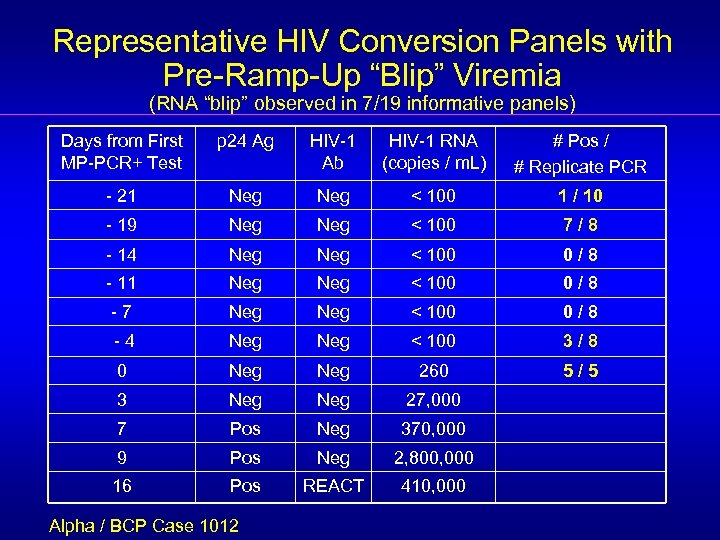 Representative HIV Conversion Panels with Pre-Ramp-Up “Blip” Viremia (RNA “blip” observed in 7/19 informative panels) Days from First MP-PCR+ Test p 24 Ag HIV-1 Ab HIV-1 RNA (copies / m. L) # Pos / # Replicate PCR - 21 Neg < 100 1 / 10 - 19 Neg < 100 7/8 - 14 Neg < 100 0/8 - 11 Neg < 100 0/8 -7 Neg < 100 0/8 -4 Neg < 100 3/8 0 Neg 260 5/5 3 Neg 27, 000 7 Pos Neg 370, 000 9 Pos Neg 2, 800, 000 16 Pos REACT 410, 000 Alpha / BCP Case 1012
Representative HIV Conversion Panels with Pre-Ramp-Up “Blip” Viremia (RNA “blip” observed in 7/19 informative panels) Days from First MP-PCR+ Test p 24 Ag HIV-1 Ab HIV-1 RNA (copies / m. L) # Pos / # Replicate PCR - 21 Neg < 100 1 / 10 - 19 Neg < 100 7/8 - 14 Neg < 100 0/8 - 11 Neg < 100 0/8 -7 Neg < 100 0/8 -4 Neg < 100 3/8 0 Neg 260 5/5 3 Neg 27, 000 7 Pos Neg 370, 000 9 Pos Neg 2, 800, 000 16 Pos REACT 410, 000 Alpha / BCP Case 1012
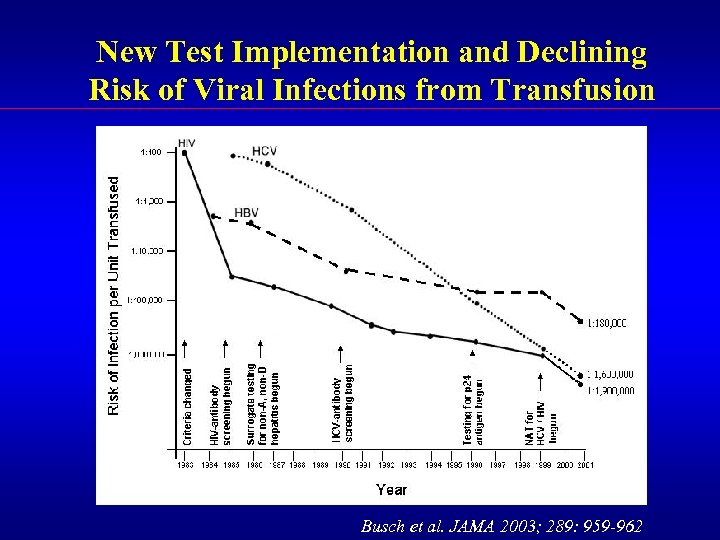 New Test Implementation and Declining Risk of Viral Infections from Transfusion Busch et al. JAMA 2003; 289: 959 -962
New Test Implementation and Declining Risk of Viral Infections from Transfusion Busch et al. JAMA 2003; 289: 959 -962
 Advances in NAT systems for blood donor screening • Fully automated platforms – Bar coded tubes to validated electronic results • Generic extraction of RNA/DNA • Internal controls to verify amplification • Multiplex or parallel detection of mulitple viruses (Taqman, beacon or DKA strategies) • Primers selected to amplify divergent subtypes • Rapid response to emerging pathogens
Advances in NAT systems for blood donor screening • Fully automated platforms – Bar coded tubes to validated electronic results • Generic extraction of RNA/DNA • Internal controls to verify amplification • Multiplex or parallel detection of mulitple viruses (Taqman, beacon or DKA strategies) • Primers selected to amplify divergent subtypes • Rapid response to emerging pathogens
 TIGRIS Fully Automated NAT System
TIGRIS Fully Automated NAT System
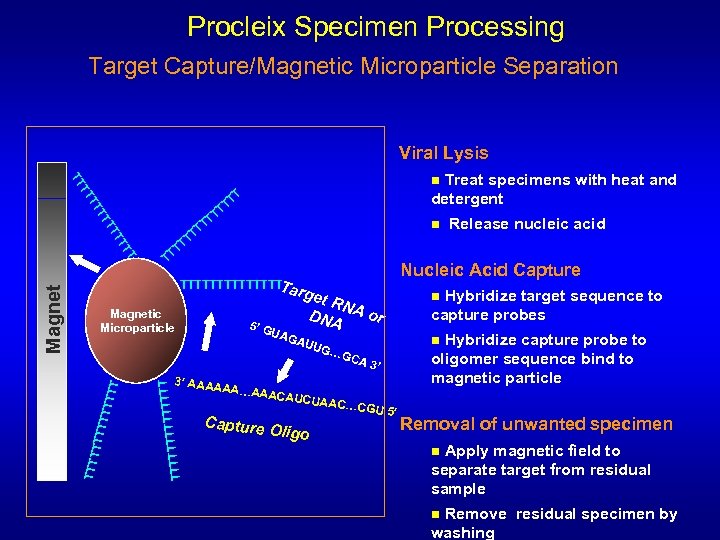 Procleix Specimen Processing Target Capture/Magnetic Microparticle Separation Viral Lysis Treat specimens with heat and detergent TT TT n TT TT TT T TT Magnet TTTTTTT a T TTTTTT T 5’ G UAG 3’ AAAA T TTTTTT TT Magnetic Microparticle Release nucleic acid Nucleic Acid Capture rget RNA or DNA AUU AA…AA Hybridize capture probe to oligomer sequence bind to magnetic particle n G… ACAUC Hybridize target sequence to capture probes n GCA 3’ Capture O UAAC… ligo CGU 5’ Removal of unwanted specimen Apply magnetic field to separate target from residual sample n Remove residual specimen by washing n
Procleix Specimen Processing Target Capture/Magnetic Microparticle Separation Viral Lysis Treat specimens with heat and detergent TT TT n TT TT TT T TT Magnet TTTTTTT a T TTTTTT T 5’ G UAG 3’ AAAA T TTTTTT TT Magnetic Microparticle Release nucleic acid Nucleic Acid Capture rget RNA or DNA AUU AA…AA Hybridize capture probe to oligomer sequence bind to magnetic particle n G… ACAUC Hybridize target sequence to capture probes n GCA 3’ Capture O UAAC… ligo CGU 5’ Removal of unwanted specimen Apply magnetic field to separate target from residual sample n Remove residual specimen by washing n
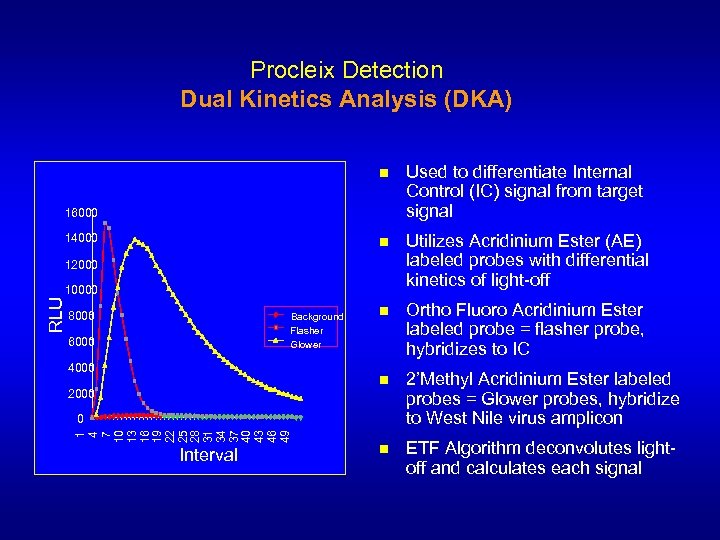 Procleix Detection Dual Kinetics Analysis (DKA) n Used to differentiate Internal Control (IC) signal from target signal n Utilizes Acridinium Ester (AE) labeled probes with differential kinetics of light-off n Ortho Fluoro Acridinium Ester labeled probe = flasher probe, hybridizes to IC n 2’Methyl Acridinium Ester labeled probes = Glower probes, hybridize to West Nile virus amplicon n ETF Algorithm deconvolutes lightoff and calculates each signal 16000 14000 12000 8000 Background Flasher Glower 6000 4000 2000 0 1 4 7 10 13 16 19 22 25 28 31 34 37 40 43 46 49 RLU 10000 Interval
Procleix Detection Dual Kinetics Analysis (DKA) n Used to differentiate Internal Control (IC) signal from target signal n Utilizes Acridinium Ester (AE) labeled probes with differential kinetics of light-off n Ortho Fluoro Acridinium Ester labeled probe = flasher probe, hybridizes to IC n 2’Methyl Acridinium Ester labeled probes = Glower probes, hybridize to West Nile virus amplicon n ETF Algorithm deconvolutes lightoff and calculates each signal 16000 14000 12000 8000 Background Flasher Glower 6000 4000 2000 0 1 4 7 10 13 16 19 22 25 28 31 34 37 40 43 46 49 RLU 10000 Interval
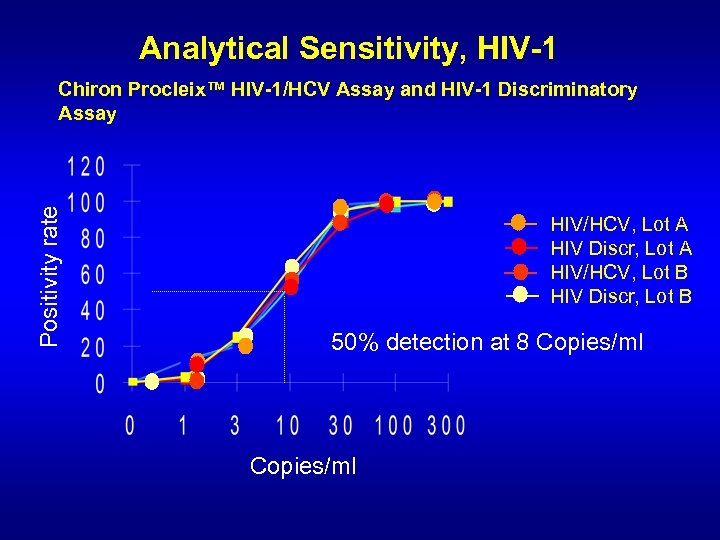 Analytical Sensitivity, HIV-1 Positivity rate Chiron Procleix™ HIV-1/HCV Assay and HIV-1 Discriminatory Assay HIV/HCV, Lot A HIV Discr, Lot A HIV/HCV, Lot B HIV Discr, Lot B 50% detection at 8 Copies/ml
Analytical Sensitivity, HIV-1 Positivity rate Chiron Procleix™ HIV-1/HCV Assay and HIV-1 Discriminatory Assay HIV/HCV, Lot A HIV Discr, Lot A HIV/HCV, Lot B HIV Discr, Lot B 50% detection at 8 Copies/ml
 Roche Fully Automated Taq. Screen™ System Hamilton Pipettor COBAS Taq. Man COBAS Ampli. Prep
Roche Fully Automated Taq. Screen™ System Hamilton Pipettor COBAS Taq. Man COBAS Ampli. Prep
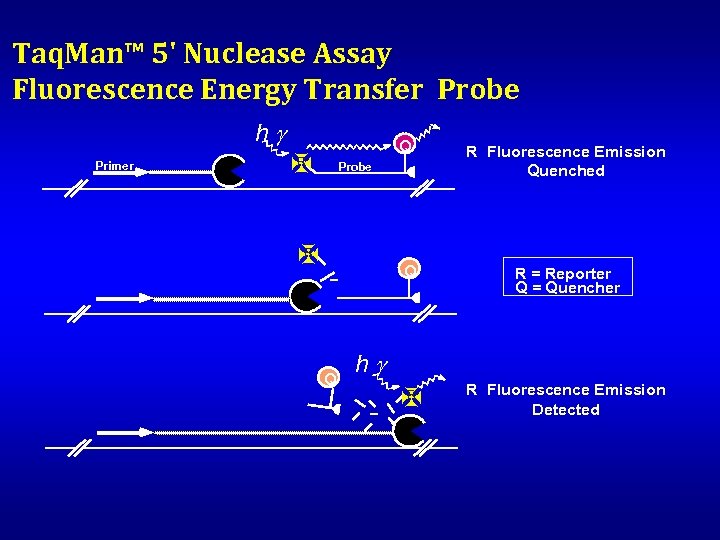 Taq. Man™ 5' Nuclease Assay Fluorescence Energy Transfer Probe hg Primer Q X R Probe X R Fluorescence Emission Quenched R Q Q R = Reporter Q = Quencher X R Fluorescence Emission Detected hg R
Taq. Man™ 5' Nuclease Assay Fluorescence Energy Transfer Probe hg Primer Q X R Probe X R Fluorescence Emission Quenched R Q Q R = Reporter Q = Quencher X R Fluorescence Emission Detected hg R
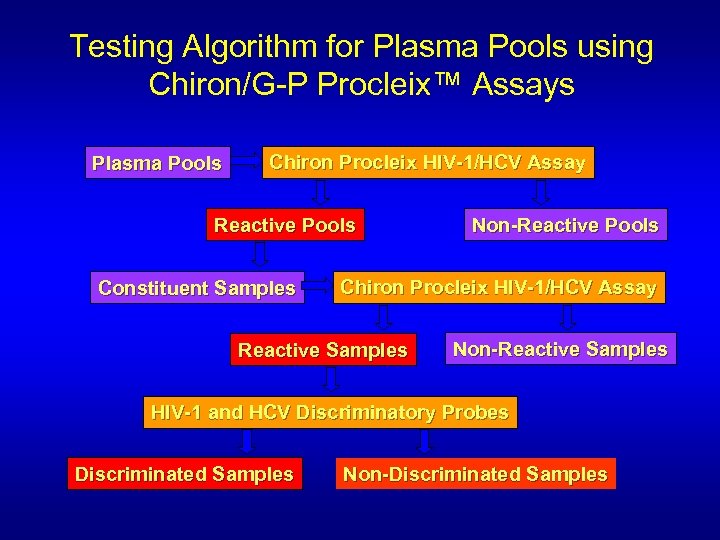 Testing Algorithm for Plasma Pools using Chiron/G-P Procleix™ Assays Plasma Pools Chiron Procleix HIV-1/HCV Assay Reactive Pools Constituent Samples Non-Reactive Pools Chiron Procleix HIV-1/HCV Assay Reactive Samples Non-Reactive Samples HIV-1 and HCV Discriminatory Probes Discriminated Samples Non-Discriminated Samples
Testing Algorithm for Plasma Pools using Chiron/G-P Procleix™ Assays Plasma Pools Chiron Procleix HIV-1/HCV Assay Reactive Pools Constituent Samples Non-Reactive Pools Chiron Procleix HIV-1/HCV Assay Reactive Samples Non-Reactive Samples HIV-1 and HCV Discriminatory Probes Discriminated Samples Non-Discriminated Samples
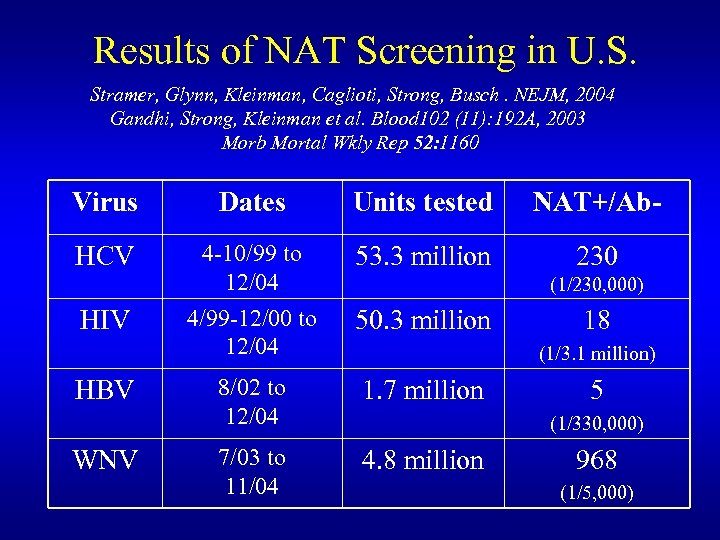 Results of NAT Screening in U. S. Stramer, Glynn, Kleinman, Caglioti, Strong, Busch. NEJM, 2004 Gandhi, Strong, Kleinman et al. Blood 102 (11): 192 A, 2003 Morb Mortal Wkly Rep 52: 1160 Virus Dates Units tested NAT+/Ab- HCV 4 -10/99 to 12/04 4/99 -12/00 to 12/04 53. 3 million 230 8/02 to 12/04 1. 7 million 7/03 to 11/04 4. 8 million HIV HBV WNV (1/230, 000) 50. 3 million 18 (1/3. 1 million) 5 (1/330, 000) 968 (1/5, 000)
Results of NAT Screening in U. S. Stramer, Glynn, Kleinman, Caglioti, Strong, Busch. NEJM, 2004 Gandhi, Strong, Kleinman et al. Blood 102 (11): 192 A, 2003 Morb Mortal Wkly Rep 52: 1160 Virus Dates Units tested NAT+/Ab- HCV 4 -10/99 to 12/04 4/99 -12/00 to 12/04 53. 3 million 230 8/02 to 12/04 1. 7 million 7/03 to 11/04 4. 8 million HIV HBV WNV (1/230, 000) 50. 3 million 18 (1/3. 1 million) 5 (1/330, 000) 968 (1/5, 000)
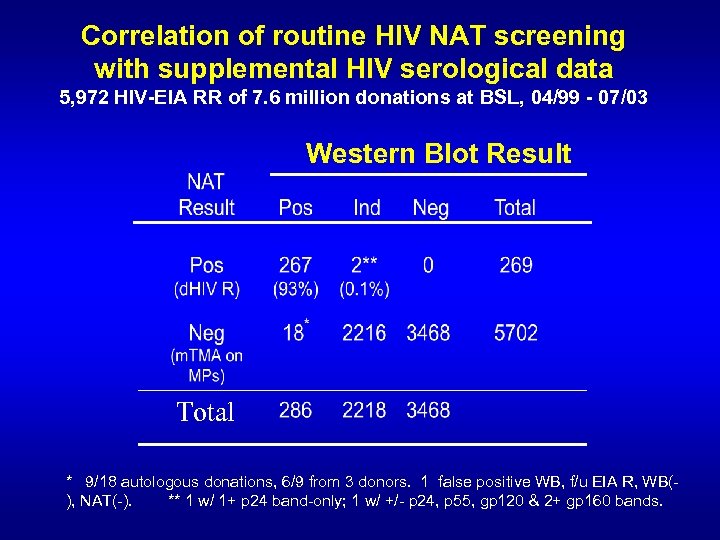 Correlation of routine HIV NAT screening with supplemental HIV serological data 5, 972 HIV-EIA RR of 7. 6 million donations at BSL, 04/99 - 07/03 Western Blot Result Total * 9/18 autologous donations, 6/9 from 3 donors. 1 false positive WB, f/u EIA R, WB(), NAT(-). ** 1 w/ 1+ p 24 band-only; 1 w/ +/- p 24, p 55, gp 120 & 2+ gp 160 bands.
Correlation of routine HIV NAT screening with supplemental HIV serological data 5, 972 HIV-EIA RR of 7. 6 million donations at BSL, 04/99 - 07/03 Western Blot Result Total * 9/18 autologous donations, 6/9 from 3 donors. 1 false positive WB, f/u EIA R, WB(), NAT(-). ** 1 w/ 1+ p 24 band-only; 1 w/ +/- p 24, p 55, gp 120 & 2+ gp 160 bands.
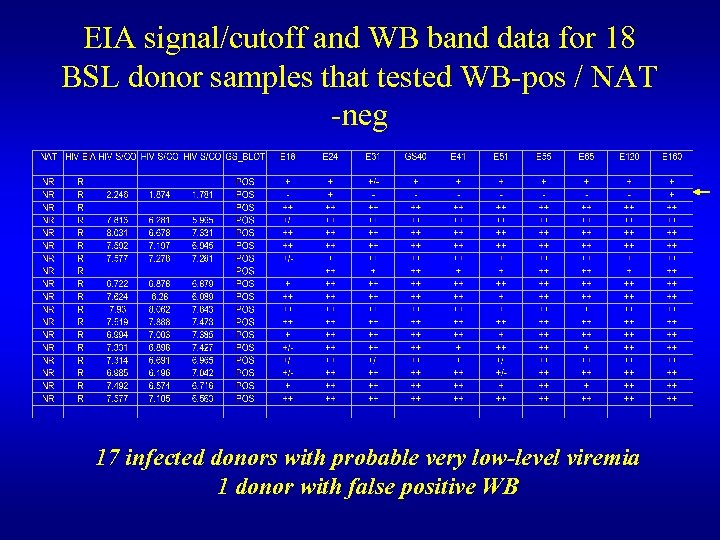 EIA signal/cutoff and WB band data for 18 BSL donor samples that tested WB-pos / NAT -neg 17 infected donors with probable very low-level viremia 1 donor with false positive WB
EIA signal/cutoff and WB band data for 18 BSL donor samples that tested WB-pos / NAT -neg 17 infected donors with probable very low-level viremia 1 donor with false positive WB
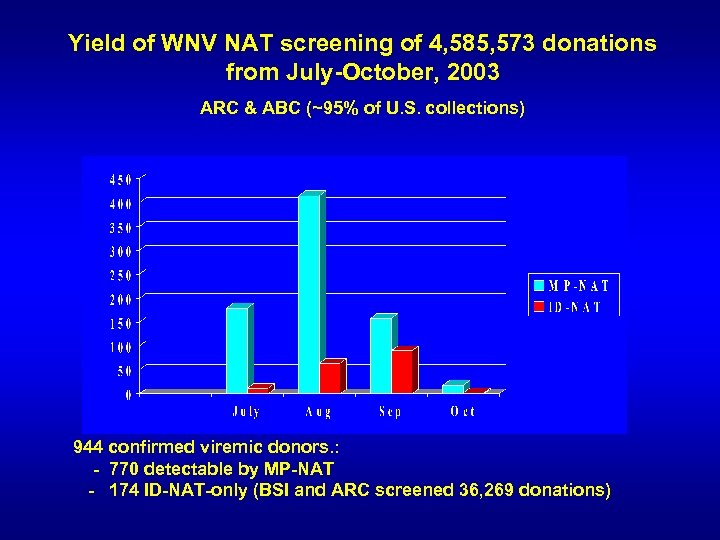 Yield of WNV NAT screening of 4, 585, 573 donations from July-October, 2003 ARC & ABC (~95% of U. S. collections) 944 confirmed viremic donors. : - 770 detectable by MP-NAT - 174 ID-NAT-only (BSI and ARC screened 36, 269 donations)
Yield of WNV NAT screening of 4, 585, 573 donations from July-October, 2003 ARC & ABC (~95% of U. S. collections) 944 confirmed viremic donors. : - 770 detectable by MP-NAT - 174 ID-NAT-only (BSI and ARC screened 36, 269 donations)
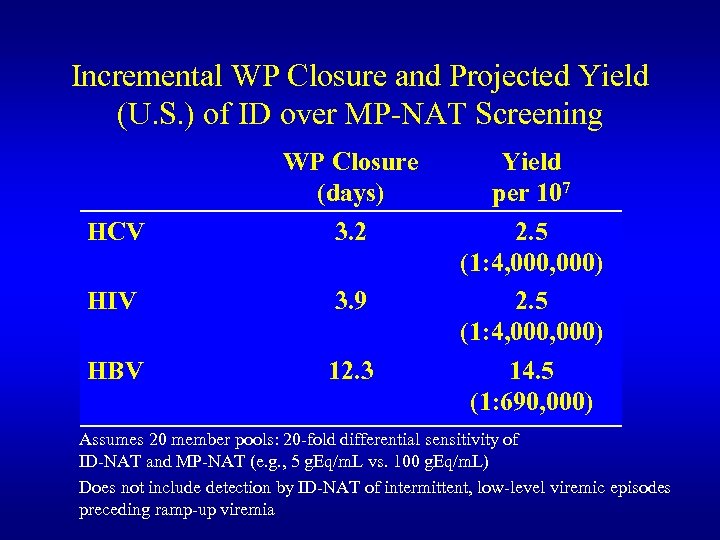 Incremental WP Closure and Projected Yield (U. S. ) of ID over MP-NAT Screening HCV WP Closure (days) 3. 2 HIV 3. 9 HBV 12. 3 Yield per 107 2. 5 (1: 4, 000, 000) 14. 5 (1: 690, 000) Assumes 20 member pools: 20 -fold differential sensitivity of ID-NAT and MP-NAT (e. g. , 5 g. Eq/m. L vs. 100 g. Eq/m. L) Does not include detection by ID-NAT of intermittent, low-level viremic episodes preceding ramp-up viremia
Incremental WP Closure and Projected Yield (U. S. ) of ID over MP-NAT Screening HCV WP Closure (days) 3. 2 HIV 3. 9 HBV 12. 3 Yield per 107 2. 5 (1: 4, 000, 000) 14. 5 (1: 690, 000) Assumes 20 member pools: 20 -fold differential sensitivity of ID-NAT and MP-NAT (e. g. , 5 g. Eq/m. L vs. 100 g. Eq/m. L) Does not include detection by ID-NAT of intermittent, low-level viremic episodes preceding ramp-up viremia
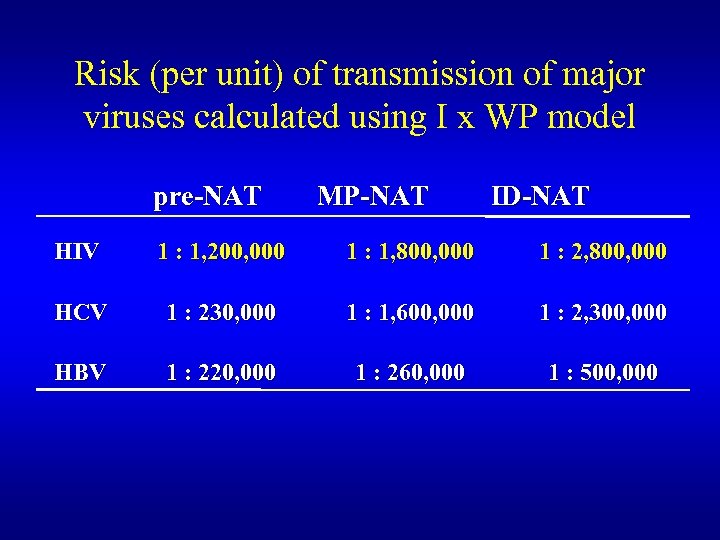 Risk (per unit) of transmission of major viruses calculated using I x WP model pre-NAT MP-NAT ID-NAT HIV 1 : 1, 200, 000 1 : 1, 800, 000 1 : 2, 800, 000 HCV 1 : 230, 000 1 : 1, 600, 000 1 : 2, 300, 000 HBV 1 : 220, 000 1 : 260, 000 1 : 500, 000
Risk (per unit) of transmission of major viruses calculated using I x WP model pre-NAT MP-NAT ID-NAT HIV 1 : 1, 200, 000 1 : 1, 800, 000 1 : 2, 800, 000 HCV 1 : 230, 000 1 : 1, 600, 000 1 : 2, 300, 000 HBV 1 : 220, 000 1 : 260, 000 1 : 500, 000
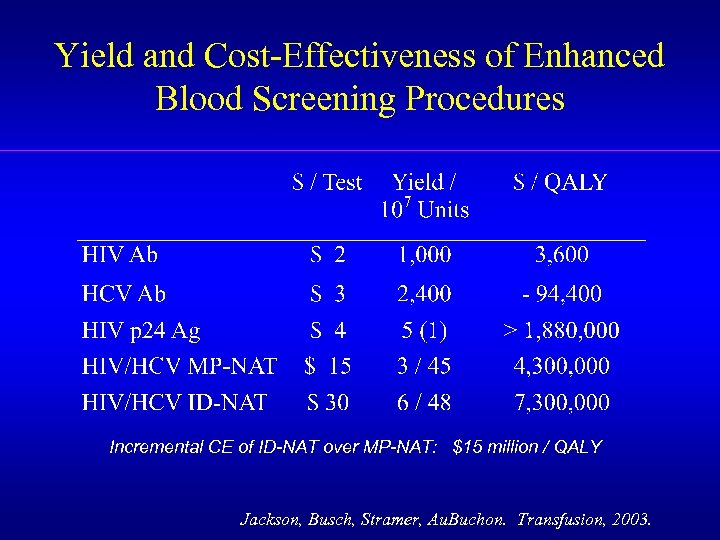 Yield and Cost-Effectiveness of Enhanced Blood Screening Procedures Incremental CE of ID-NAT over MP-NAT: $15 million / QALY Jackson, Busch, Stramer, Au. Buchon. Transfusion, 2003.
Yield and Cost-Effectiveness of Enhanced Blood Screening Procedures Incremental CE of ID-NAT over MP-NAT: $15 million / QALY Jackson, Busch, Stramer, Au. Buchon. Transfusion, 2003.
 Application of donor NAT screening assays in HIV research, diagnostic and public health settings • Pathogenesis and treatment studies (AIEDRP) – – Mechanisms of viral clearance/control Immune and viral evolution during primary infection Response of early vs delayed therapy Antiviral vs immune enhancement strategies • Vaccine trials – Establish incidence in enrolled populations – Discriminate vaccine response from breakthrough Tx – Precisely define infection dates for assessment of efficacy • PEP trial – Real-time NAT sensitivity and specificity
Application of donor NAT screening assays in HIV research, diagnostic and public health settings • Pathogenesis and treatment studies (AIEDRP) – – Mechanisms of viral clearance/control Immune and viral evolution during primary infection Response of early vs delayed therapy Antiviral vs immune enhancement strategies • Vaccine trials – Establish incidence in enrolled populations – Discriminate vaccine response from breakthrough Tx – Precisely define infection dates for assessment of efficacy • PEP trial – Real-time NAT sensitivity and specificity
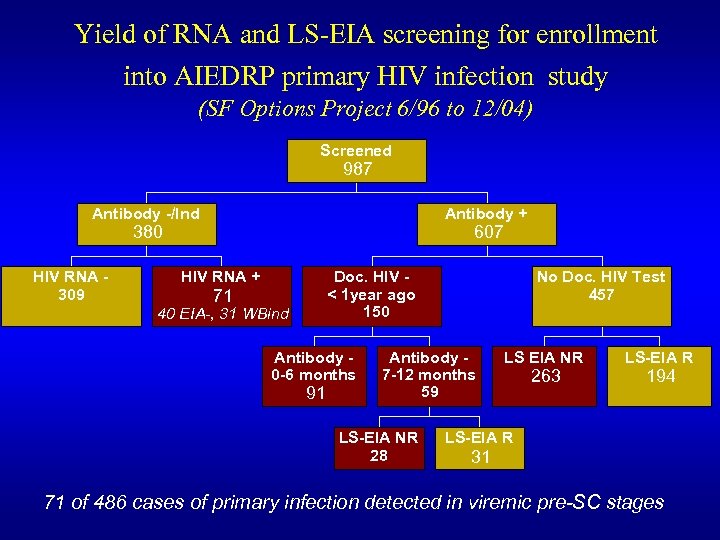 Yield of RNA and LS-EIA screening for enrollment into AIEDRP primary HIV infection study (SF Options Project 6/96 to 12/04) Screened 987 Antibody -/Ind Antibody + 380 HIV RNA 309 607 HIV RNA + Doc. HIV < 1 year ago 150 71 40 EIA-, 31 WBind Antibody 0 -6 months 91 No Doc. HIV Test 457 Antibody 7 -12 months 59 LS-EIA NR 28 LS EIA NR 263 LS-EIA R 194 LS-EIA R 31 71 of 486 cases of primary infection detected in viremic pre-SC stages
Yield of RNA and LS-EIA screening for enrollment into AIEDRP primary HIV infection study (SF Options Project 6/96 to 12/04) Screened 987 Antibody -/Ind Antibody + 380 HIV RNA 309 607 HIV RNA + Doc. HIV < 1 year ago 150 71 40 EIA-, 31 WBind Antibody 0 -6 months 91 No Doc. HIV Test 457 Antibody 7 -12 months 59 LS-EIA NR 28 LS EIA NR 263 LS-EIA R 194 LS-EIA R 31 71 of 486 cases of primary infection detected in viremic pre-SC stages
 Use of NAT Testing to Determine Time of Infection in Two Phase III Trials of an HIV Vaccine (VAX 004/005) • 30 of 711 infections (1. 8%) detected by SC occurred in volunteers who were already viremic at baseline - would have been misclassified as vaccine failures – Based on 15 -day (+/-5) NAT+/Ab- WP, baseline annualized HIV incidence estimated at 5. 4% (1. 2%-13. 7%, combined 95%CI). • Additional 34% of infections had their date of infection changed based on NAT - important in the time-to-event analyses of efficacy • No vaccine effect on rate of detection of RNA+/Absamples – no evidence for accelerated or retarded seroconversion
Use of NAT Testing to Determine Time of Infection in Two Phase III Trials of an HIV Vaccine (VAX 004/005) • 30 of 711 infections (1. 8%) detected by SC occurred in volunteers who were already viremic at baseline - would have been misclassified as vaccine failures – Based on 15 -day (+/-5) NAT+/Ab- WP, baseline annualized HIV incidence estimated at 5. 4% (1. 2%-13. 7%, combined 95%CI). • Additional 34% of infections had their date of infection changed based on NAT - important in the time-to-event analyses of efficacy • No vaccine effect on rate of detection of RNA+/Absamples – no evidence for accelerated or retarded seroconversion


