7cbc8a4d39803b681f683fbc34283531.ppt
- Количество слайдов: 45
 Research In the Post-Genomics Era Martina Mc. Gloughlin, Biotechnology Program and Life Sciences Informatics Program UC Davis
Research In the Post-Genomics Era Martina Mc. Gloughlin, Biotechnology Program and Life Sciences Informatics Program UC Davis
 UC Davis Biotechnology Program UC Systemwide Life Sciences Informatics Program “Biology in the 21 st century will increasingly become an information science” Leroy Hood, Jan 11, 1999 “Any cell has in it a billion years of experimentation by its ancestors” Max Delbruck, 1949 2
UC Davis Biotechnology Program UC Systemwide Life Sciences Informatics Program “Biology in the 21 st century will increasingly become an information science” Leroy Hood, Jan 11, 1999 “Any cell has in it a billion years of experimentation by its ancestors” Max Delbruck, 1949 2
 Genomics w The massive interest and commitment of resources in both the public and private sectors flows from the generally-held perception that genomics will be the single most fruitful approach to the acquisition of new information in basic and applied biology in the next several decades. w If genomics were only to be a tool for the basic biologist, the benefits of this approach would be staggering, yielding new insights into fundamental processes such as cell division, differentiation, transformation, the development and reproduction of organisms and the diversity of populations. w The rewards in applied biology, however, have clearly attracted the private sector and public interest. These include the promise of facile new approaches for drug discovery, new understanding of metabolic processes and new approaches to determining qualitative and quantitative traits in plants and animals for breeding and genetic engineering. 3
Genomics w The massive interest and commitment of resources in both the public and private sectors flows from the generally-held perception that genomics will be the single most fruitful approach to the acquisition of new information in basic and applied biology in the next several decades. w If genomics were only to be a tool for the basic biologist, the benefits of this approach would be staggering, yielding new insights into fundamental processes such as cell division, differentiation, transformation, the development and reproduction of organisms and the diversity of populations. w The rewards in applied biology, however, have clearly attracted the private sector and public interest. These include the promise of facile new approaches for drug discovery, new understanding of metabolic processes and new approaches to determining qualitative and quantitative traits in plants and animals for breeding and genetic engineering. 3
 One Human Sequence Typed in 10 -pitch font, one human sequence would stretch for more than 5, 000 miles. Digitally formatted, it could be stored on one CD-ROM. Biologically encoded, it fits easily within a single cell. 4
One Human Sequence Typed in 10 -pitch font, one human sequence would stretch for more than 5, 000 miles. Digitally formatted, it could be stored on one CD-ROM. Biologically encoded, it fits easily within a single cell. 4
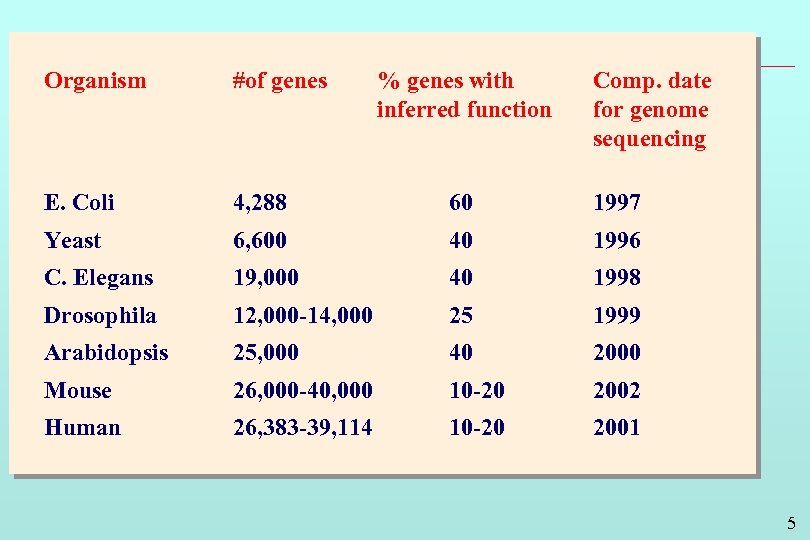 Organism #of genes % genes with inferred function Comp. date for genome sequencing E. Coli 4, 288 60 1997 Yeast 6, 600 40 1996 C. Elegans 19, 000 40 1998 Drosophila 12, 000 -14, 000 25 1999 Arabidopsis 25, 000 40 2000 Mouse 26, 000 -40, 000 10 -20 2002 Human 26, 383 -39, 114 10 -20 2001 5
Organism #of genes % genes with inferred function Comp. date for genome sequencing E. Coli 4, 288 60 1997 Yeast 6, 600 40 1996 C. Elegans 19, 000 40 1998 Drosophila 12, 000 -14, 000 25 1999 Arabidopsis 25, 000 40 2000 Mouse 26, 000 -40, 000 10 -20 2002 Human 26, 383 -39, 114 10 -20 2001 5
 Paradigm Shift in Biology The new paradigm, now emerging, is that all the ‘genes’ will be known (in the sense of being resident in databases available electronically), and that the starting point of a biological investigation will be theoretical. An individual scientist will begin with a theoretical conjecture, only then turning to experiment to follow or test that hypothesis. Walter Gilbert. 1991. Towards a paradigm shift in biology. Nature, 349: 99. 6
Paradigm Shift in Biology The new paradigm, now emerging, is that all the ‘genes’ will be known (in the sense of being resident in databases available electronically), and that the starting point of a biological investigation will be theoretical. An individual scientist will begin with a theoretical conjecture, only then turning to experiment to follow or test that hypothesis. Walter Gilbert. 1991. Towards a paradigm shift in biology. Nature, 349: 99. 6
![Paradigm Shift in Biology To use [the] flood of knowledge, which will pour across Paradigm Shift in Biology To use [the] flood of knowledge, which will pour across](https://present5.com/presentation/7cbc8a4d39803b681f683fbc34283531/image-7.jpg) Paradigm Shift in Biology To use [the] flood of knowledge, which will pour across the computer networks of the world, biologists not only must become computer literate, but also change their approach to the problem of understanding life. Walter Gilbert. 1991. Towards a paradigm shift in biology. Nature, 349: 99. 7
Paradigm Shift in Biology To use [the] flood of knowledge, which will pour across the computer networks of the world, biologists not only must become computer literate, but also change their approach to the problem of understanding life. Walter Gilbert. 1991. Towards a paradigm shift in biology. Nature, 349: 99. 7
 What’s Really Next The post-genome era in biological research will take for granted ready access to huge amounts of genomic data. The challenge will be understanding those data and using the understanding to solve real-world problems. . . 8
What’s Really Next The post-genome era in biological research will take for granted ready access to huge amounts of genomic data. The challenge will be understanding those data and using the understanding to solve real-world problems. . . 8
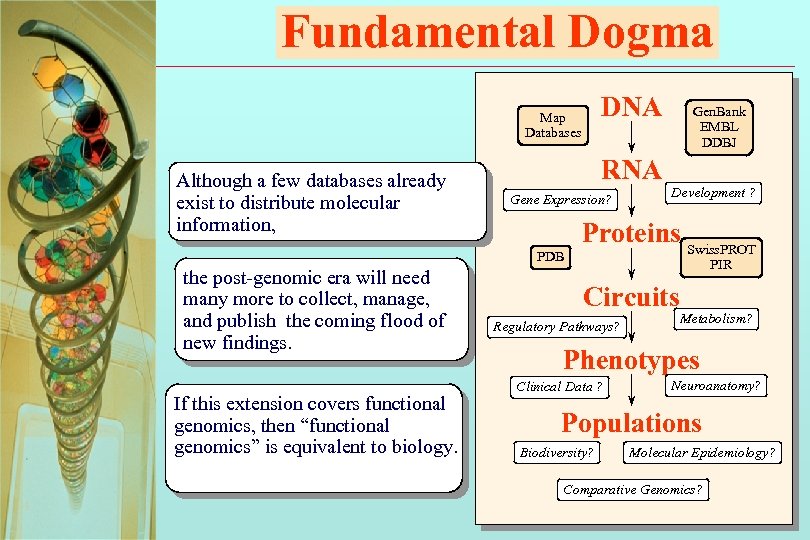 Fundamental Dogma DNA Map Databases Although a few databases already exist to distribute molecular information, RNA Gene Expression? PDB the post-genomic era will need many more to collect, manage, and publish the coming flood of new findings. If this extension covers functional genomics, then “functional genomics” is equivalent to biology. Gen. Bank EMBL DDBJ Development ? Proteins Swiss. PROT PIR Circuits Regulatory Pathways? Metabolism? Phenotypes Clinical Data ? Neuroanatomy? Populations Biodiversity? Molecular Epidemiology? Comparative Genomics? 9
Fundamental Dogma DNA Map Databases Although a few databases already exist to distribute molecular information, RNA Gene Expression? PDB the post-genomic era will need many more to collect, manage, and publish the coming flood of new findings. If this extension covers functional genomics, then “functional genomics” is equivalent to biology. Gen. Bank EMBL DDBJ Development ? Proteins Swiss. PROT PIR Circuits Regulatory Pathways? Metabolism? Phenotypes Clinical Data ? Neuroanatomy? Populations Biodiversity? Molecular Epidemiology? Comparative Genomics? 9
 There are Problems with the HGP. . . • The actual sequence data makes up only 16% of the content; the other 84% is annotated. • The is leads to a number of issues: – How to structure databases for mining – How to establish control vocabularies to establish integrity of searches – What new algorithms are needed to facilitate processing and correlating the petabytes (1015 bytes) of information – How can protein function be extracted for the purposes of diagnostics, drug discovery and therapeutics 10
There are Problems with the HGP. . . • The actual sequence data makes up only 16% of the content; the other 84% is annotated. • The is leads to a number of issues: – How to structure databases for mining – How to establish control vocabularies to establish integrity of searches – What new algorithms are needed to facilitate processing and correlating the petabytes (1015 bytes) of information – How can protein function be extracted for the purposes of diagnostics, drug discovery and therapeutics 10
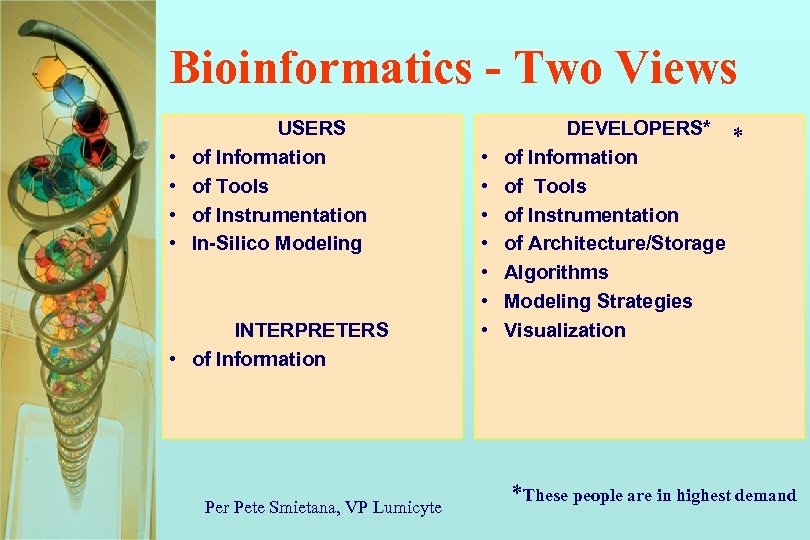 Bioinformatics - Two Views • • USERS of Information of Tools of Instrumentation In-Silico Modeling INTERPRETERS • of Information Per Pete Smietana, VP Lumicyte • • DEVELOPERS* of Information of Tools of Instrumentation of Architecture/Storage Algorithms Modeling Strategies Visualization * *These people are in highest demand
Bioinformatics - Two Views • • USERS of Information of Tools of Instrumentation In-Silico Modeling INTERPRETERS • of Information Per Pete Smietana, VP Lumicyte • • DEVELOPERS* of Information of Tools of Instrumentation of Architecture/Storage Algorithms Modeling Strategies Visualization * *These people are in highest demand
 Typical Bioinformatics Multi-Disciplinary Training • Scientists – Biology, Molecular Genetics, Clinical Biochemistry, Protein Structure Chemistry • Mathematicians – Statistics, Algorithms, Image processing • Computer Scientists – Database, User Interface/Visualizations, Networking (Internets/Intranets), Instrument Control 12
Typical Bioinformatics Multi-Disciplinary Training • Scientists – Biology, Molecular Genetics, Clinical Biochemistry, Protein Structure Chemistry • Mathematicians – Statistics, Algorithms, Image processing • Computer Scientists – Database, User Interface/Visualizations, Networking (Internets/Intranets), Instrument Control 12
 Typical Bioinformatics Multi-Disciplinary Functions • Scientists – Experimental Design & Interpretation – Laboratory Protocols & Standards/Controls • Mathematicians – Analysis & Correlation of Data – Validation methodologies • Computer Scientists – Information Storage / Control Vocabulary – Data Mining 13
Typical Bioinformatics Multi-Disciplinary Functions • Scientists – Experimental Design & Interpretation – Laboratory Protocols & Standards/Controls • Mathematicians – Analysis & Correlation of Data – Validation methodologies • Computer Scientists – Information Storage / Control Vocabulary – Data Mining 13
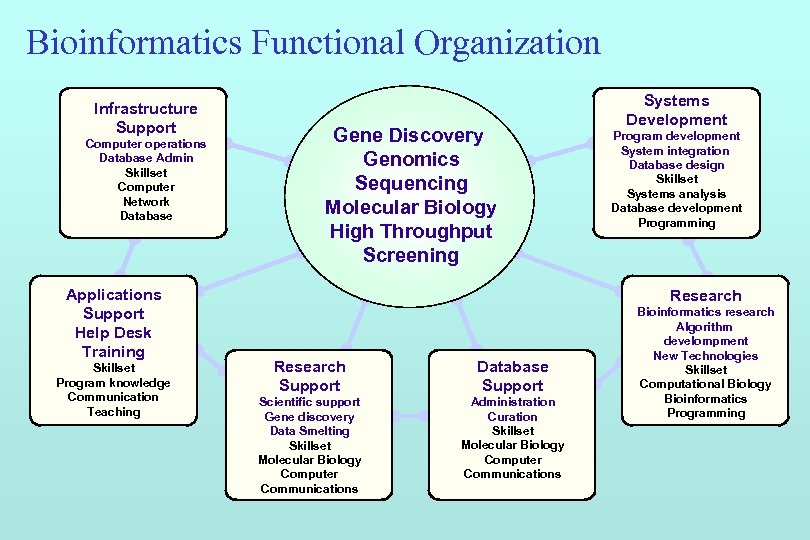 Bioinformatics Functional Organization Infrastructure Support Computer operations Database Admin Skillset Computer Network Database Applications Support Help Desk Training Skillset Program knowledge Communication Teaching Gene Discovery Genomics Sequencing Molecular Biology High Throughput Screening Systems Development Program development System integration Database design Skillset Systems analysis Database development Programming Research Support Database Support Scientific support Gene discovery Data Smelting Skillset Molecular Biology Computer Communications Administration Curation Skillset Molecular Biology Computer Communications Bioinformatics research Algorithm develompment New Technologies Skillset Computational Biology Bioinformatics Programming
Bioinformatics Functional Organization Infrastructure Support Computer operations Database Admin Skillset Computer Network Database Applications Support Help Desk Training Skillset Program knowledge Communication Teaching Gene Discovery Genomics Sequencing Molecular Biology High Throughput Screening Systems Development Program development System integration Database design Skillset Systems analysis Database development Programming Research Support Database Support Scientific support Gene discovery Data Smelting Skillset Molecular Biology Computer Communications Administration Curation Skillset Molecular Biology Computer Communications Bioinformatics research Algorithm develompment New Technologies Skillset Computational Biology Bioinformatics Programming
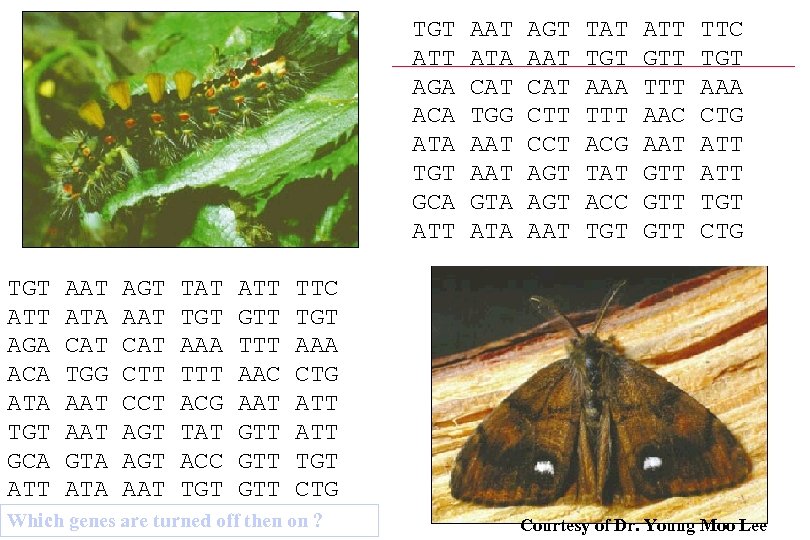 TGT ATT AGA ACA ATA TGT GCA ATT AAT ATA CAT TGG AAT AAT GTA ATA AGT AAT CAT CTT CCT AGT AGT AAT TAT TGT AAA TTT ACG TAT ACC TGT ATT GTT TTT AAC AAT GTT GTT GTT TTC TGT AAA CTG ATT ATT TGT CTG Which genes are turned off then on ? Courtesy of Dr. Young Moo Lee 15
TGT ATT AGA ACA ATA TGT GCA ATT AAT ATA CAT TGG AAT AAT GTA ATA AGT AAT CAT CTT CCT AGT AGT AAT TAT TGT AAA TTT ACG TAT ACC TGT ATT GTT TTT AAC AAT GTT GTT GTT TTC TGT AAA CTG ATT ATT TGT CTG Which genes are turned off then on ? Courtesy of Dr. Young Moo Lee 15
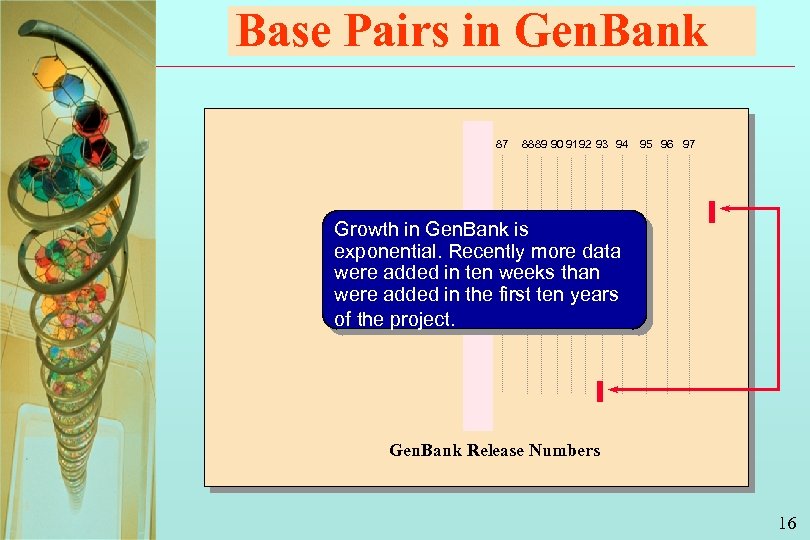 Base Pairs in Gen. Bank 87 88 89 90 91 92 93 94 95 96 97 Growth in Gen. Bank is exponential. Recently more data were added in ten weeks than were added in the first ten years of the project. Gen. Bank Release Numbers 16
Base Pairs in Gen. Bank 87 88 89 90 91 92 93 94 95 96 97 Growth in Gen. Bank is exponential. Recently more data were added in ten weeks than were added in the first ten years of the project. Gen. Bank Release Numbers 16
 Rhetorical Question Which is likely to be more complex: • identifying, documenting, and tracking the whereabouts of all parcels in transit in the US at one time • identifying, documenting, and analyzing the structure and function of all individual genes in all economically significant organisms; then analyzing all significant gene-gene and geneenvironment interactions in those organisms and their environments 17
Rhetorical Question Which is likely to be more complex: • identifying, documenting, and tracking the whereabouts of all parcels in transit in the US at one time • identifying, documenting, and analyzing the structure and function of all individual genes in all economically significant organisms; then analyzing all significant gene-gene and geneenvironment interactions in those organisms and their environments 17
 Business Factoids United Parcel Service: • uses two redundant 3 Terabyte (yes, 3000 GB) databases to track all packages in transit. • has 4, 000 full-time employees dedicated to IT • • spends one billion dollars per year on IT has an income of 1. 1 billion dollars, against revenues of 22. 4 billion dollars 18
Business Factoids United Parcel Service: • uses two redundant 3 Terabyte (yes, 3000 GB) databases to track all packages in transit. • has 4, 000 full-time employees dedicated to IT • • spends one billion dollars per year on IT has an income of 1. 1 billion dollars, against revenues of 22. 4 billion dollars 18
 Examples of Biotech/IT Fusion Technologies ¨ Genomics, proteomics and bioinformatics ¨ Combinatorial –chemistry ¨ Peptide libraries- tea bags, beads ¨ Combinatorial -biology ¨ Directed evolution ¨ DNA Shuffling, Molecular Breeding ¨ High throughput analysis ¨ Nucleic Acid based Sequencing Microarrays • Photolithography • Mirrors • Spotted Chips • Semi-conductor ¨ Protein based 2 -D, electrospray/nanospray MS: MALDI-TOF, LC/MS/MS, SELDI ¨ Imaging/optical biology ¨ Biosensors, Bioelectronics and Bionetworks (Nanotechnology) 19
Examples of Biotech/IT Fusion Technologies ¨ Genomics, proteomics and bioinformatics ¨ Combinatorial –chemistry ¨ Peptide libraries- tea bags, beads ¨ Combinatorial -biology ¨ Directed evolution ¨ DNA Shuffling, Molecular Breeding ¨ High throughput analysis ¨ Nucleic Acid based Sequencing Microarrays • Photolithography • Mirrors • Spotted Chips • Semi-conductor ¨ Protein based 2 -D, electrospray/nanospray MS: MALDI-TOF, LC/MS/MS, SELDI ¨ Imaging/optical biology ¨ Biosensors, Bioelectronics and Bionetworks (Nanotechnology) 19
 Genomics, Proteomics and Bioinformatics ¨ Genomics is operationally defined as investigations into the structure and function of very large numbers of genes undertaken in a simultaneous fashion. ¨ Structural genomics includes the genetic mapping, physical mapping and sequencing of entire genomes. ¨ Comparative genomics means information gained in one organism can have application in other even distantly related organisms. This enables the application of information gained from facile model systems to agricultural and medical problems. The nature and significance of differences between genomes also provides a powerful tool for determining the relationship between genotype and phenotype through comparative genomics and morphological and physiological studies. ¨ Functional genomics Phenotype is logically the subject of functional genomics. Genome sequencing for most organisms of interest will be complete within the near future, ushering in the so called "post-genome era. " Walter Gilbert directly speculated on the nature of biology in the "post-genome era": "The new paradigm, now emerging, is that all genes will be known (in the sense of being resident in databases available electronically), and that the starting point of a biological investigation will be theoretical. “ 20
Genomics, Proteomics and Bioinformatics ¨ Genomics is operationally defined as investigations into the structure and function of very large numbers of genes undertaken in a simultaneous fashion. ¨ Structural genomics includes the genetic mapping, physical mapping and sequencing of entire genomes. ¨ Comparative genomics means information gained in one organism can have application in other even distantly related organisms. This enables the application of information gained from facile model systems to agricultural and medical problems. The nature and significance of differences between genomes also provides a powerful tool for determining the relationship between genotype and phenotype through comparative genomics and morphological and physiological studies. ¨ Functional genomics Phenotype is logically the subject of functional genomics. Genome sequencing for most organisms of interest will be complete within the near future, ushering in the so called "post-genome era. " Walter Gilbert directly speculated on the nature of biology in the "post-genome era": "The new paradigm, now emerging, is that all genes will be known (in the sense of being resident in databases available electronically), and that the starting point of a biological investigation will be theoretical. “ 20
 Genomics, Proteomics and Bioinformatics ¨ Proteomics At the molecular level, phenotype includes all temporal and spatial aspects of gene expression as well as related aspects of the expression, structure, function and spatial localization of proteins. The Proteome is the set of all expressed proteins for a given organism. ¨ The next hierarchical level of phenotype considers how the proteome within and among cells cooperates to produce the biochemistry and physiology of individual cells and organisms. “Physiomics" is a descriptor for this approach. “Phenomics" The final hierarchical levels of phenotype include anatomy and function for cells and whole organisms. ¨ Bioinformatics: Computational or algorithmic approaches to the production of information from large amounts of biological data, include prediction of protein structure, dynamic modeling of complex physiological systems or the statistical treatment of quantitative traits in populations in order to determine the genetic basis for these traits. ¨ Unquestionably, bioinformatics will be an essential component of all research activities utilizing structural and functional genomics approaches 21
Genomics, Proteomics and Bioinformatics ¨ Proteomics At the molecular level, phenotype includes all temporal and spatial aspects of gene expression as well as related aspects of the expression, structure, function and spatial localization of proteins. The Proteome is the set of all expressed proteins for a given organism. ¨ The next hierarchical level of phenotype considers how the proteome within and among cells cooperates to produce the biochemistry and physiology of individual cells and organisms. “Physiomics" is a descriptor for this approach. “Phenomics" The final hierarchical levels of phenotype include anatomy and function for cells and whole organisms. ¨ Bioinformatics: Computational or algorithmic approaches to the production of information from large amounts of biological data, include prediction of protein structure, dynamic modeling of complex physiological systems or the statistical treatment of quantitative traits in populations in order to determine the genetic basis for these traits. ¨ Unquestionably, bioinformatics will be an essential component of all research activities utilizing structural and functional genomics approaches 21
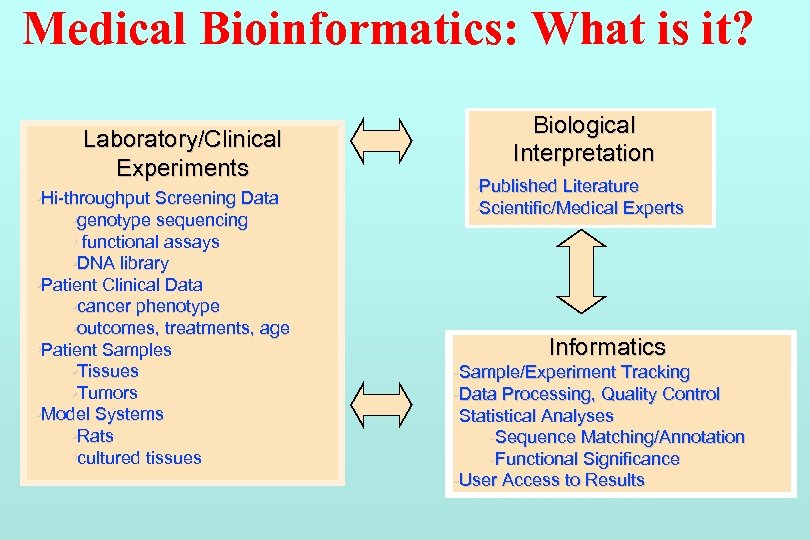 Medical Bioinformatics: What is it? Laboratory/Clinical Experiments • Hi-throughput Screening Data • genotype sequencing • functional assays • DNA library • Patient Clinical Data • cancer phenotype • outcomes, treatments, age • Patient Samples • Tissues • Tumors • Model Systems • Rats • cultured tissues Biological Interpretation • Published Literature • Scientific/Medical Experts Informatics • Sample/Experiment Tracking • Data Processing, Quality Control • Statistical Analyses • Sequence Matching/Annotation • Functional Significance • User Access to Results
Medical Bioinformatics: What is it? Laboratory/Clinical Experiments • Hi-throughput Screening Data • genotype sequencing • functional assays • DNA library • Patient Clinical Data • cancer phenotype • outcomes, treatments, age • Patient Samples • Tissues • Tumors • Model Systems • Rats • cultured tissues Biological Interpretation • Published Literature • Scientific/Medical Experts Informatics • Sample/Experiment Tracking • Data Processing, Quality Control • Statistical Analyses • Sequence Matching/Annotation • Functional Significance • User Access to Results
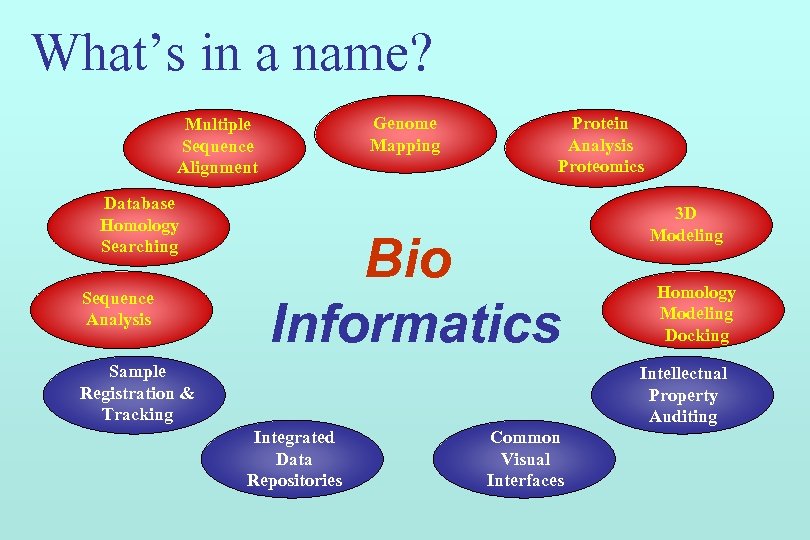 What’s in a name? Genome Mapping Multiple Sequence Alignment Database Homology Searching Sequence Analysis Protein Analysis Proteomics Bio Informatics Sample Registration & Tracking 3 D Modeling Homology Modeling Docking Intellectual Property Auditing Integrated Data Repositories Common Visual Interfaces
What’s in a name? Genome Mapping Multiple Sequence Alignment Database Homology Searching Sequence Analysis Protein Analysis Proteomics Bio Informatics Sample Registration & Tracking 3 D Modeling Homology Modeling Docking Intellectual Property Auditing Integrated Data Repositories Common Visual Interfaces
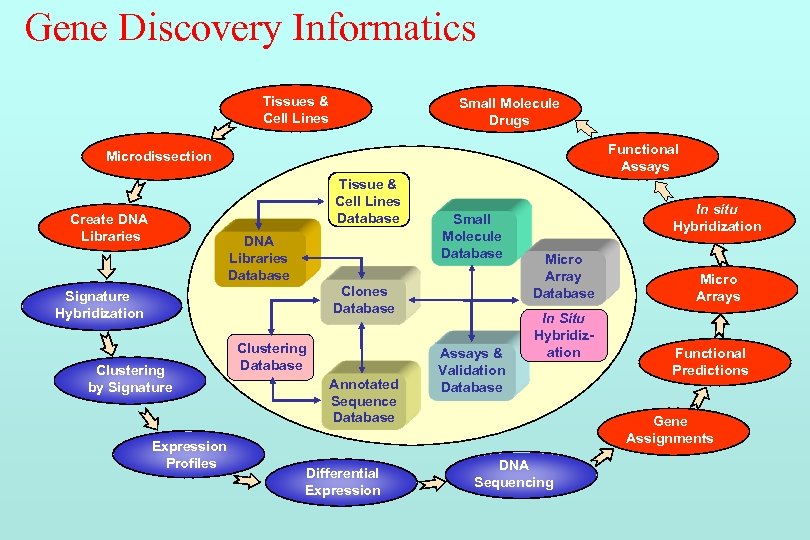 Gene Discovery Informatics Tissues & Cell Lines Small Molecule Drugs Functional Assays Microdissection Tissue & Cell Lines Database Create DNA Libraries Database Small Molecule Database Clones Database Signature Hybridization Clustering by Signature Expression Profiles Clustering Database Annotated Sequence Database Differential Expression Assays & Validation Database In situ Hybridization Micro Array Database In Situ Hybridization Micro Arrays Functional Predictions Gene Assignments DNA Sequencing
Gene Discovery Informatics Tissues & Cell Lines Small Molecule Drugs Functional Assays Microdissection Tissue & Cell Lines Database Create DNA Libraries Database Small Molecule Database Clones Database Signature Hybridization Clustering by Signature Expression Profiles Clustering Database Annotated Sequence Database Differential Expression Assays & Validation Database In situ Hybridization Micro Array Database In Situ Hybridization Micro Arrays Functional Predictions Gene Assignments DNA Sequencing
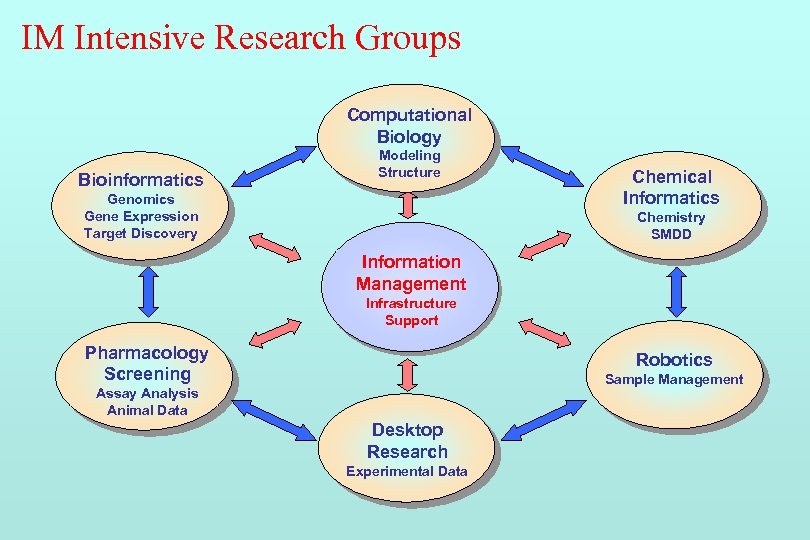 IM Intensive Research Groups Computational Biology Bioinformatics Modeling Structure Genomics Gene Expression Target Discovery Chemical Informatics Chemistry SMDD Information Management Infrastructure Support Pharmacology Screening Robotics Sample Management Assay Analysis Animal Data Desktop Research Experimental Data
IM Intensive Research Groups Computational Biology Bioinformatics Modeling Structure Genomics Gene Expression Target Discovery Chemical Informatics Chemistry SMDD Information Management Infrastructure Support Pharmacology Screening Robotics Sample Management Assay Analysis Animal Data Desktop Research Experimental Data
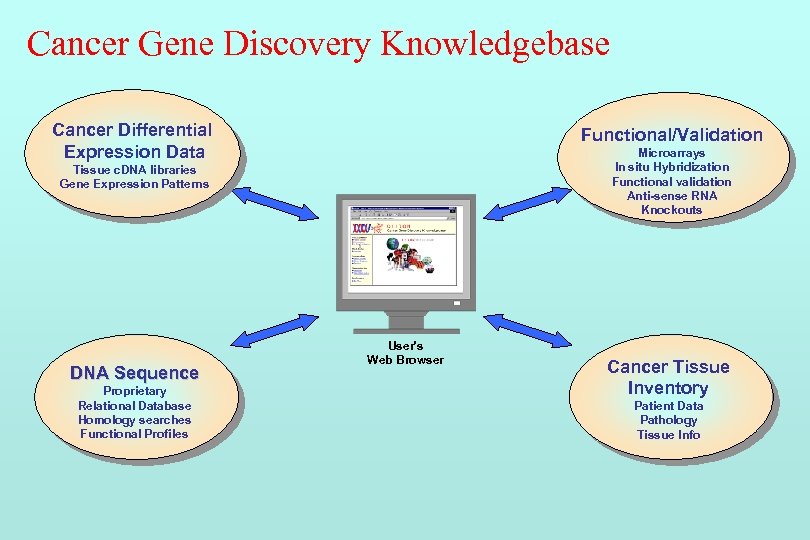 Cancer Gene Discovery Knowledgebase Cancer Differential Expression Data Functional/Validation Microarrays In situ Hybridization Functional validation Anti-sense RNA Knockouts Tissue c. DNA libraries Gene Expression Patterns DNA Sequence Proprietary Relational Database Homology searches Functional Profiles User's Web Browser Cancer Tissue Inventory Patient Data Pathology Tissue Info
Cancer Gene Discovery Knowledgebase Cancer Differential Expression Data Functional/Validation Microarrays In situ Hybridization Functional validation Anti-sense RNA Knockouts Tissue c. DNA libraries Gene Expression Patterns DNA Sequence Proprietary Relational Database Homology searches Functional Profiles User's Web Browser Cancer Tissue Inventory Patient Data Pathology Tissue Info
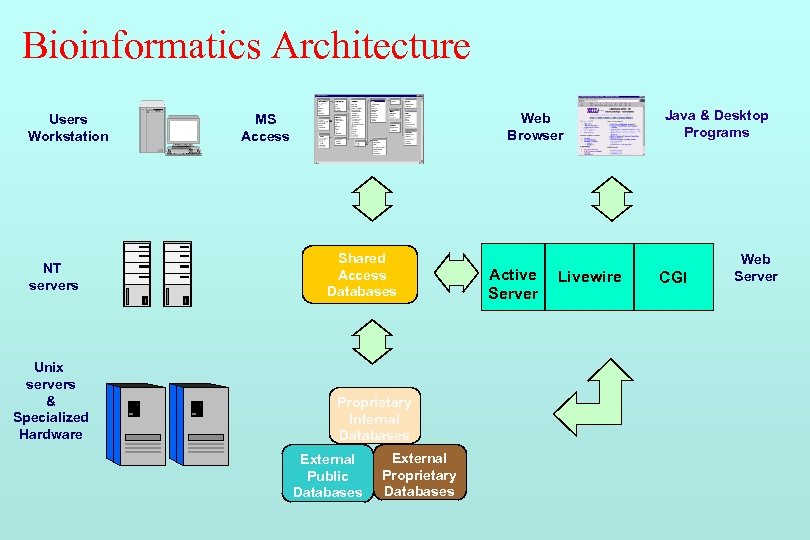 Bioinformatics Architecture Users Workstation NT servers Unix servers & Specialized Hardware Web Browser MS Access Shared Access Databases Proprietary Internal Databases External Public Databases External Proprietary Databases Active Server Livewire Java & Desktop Programs CGI Web Server
Bioinformatics Architecture Users Workstation NT servers Unix servers & Specialized Hardware Web Browser MS Access Shared Access Databases Proprietary Internal Databases External Public Databases External Proprietary Databases Active Server Livewire Java & Desktop Programs CGI Web Server
 Challenges in High Throughput Biotechnology R&D Ø Volume of Data is Growing Rapidly Ø Technology is Evolving Rapidly § Instrumentation, Informatics Ø Biological Definitions are Constantly Updated § New Interactions and Functions Discovered Daily Ø New Genes § Sometimes Homologous to Known Genes ->re-evaluate old data Ø Full Value is Realized by Integrating Multiple High-Throughput Platforms § Sequencing, Functional Screens, Small Molecule Activity Ø Up-Front Design of Data and Quality Control Databases is Crucial to Success § High Data Quality is Essential § Financially Impossible to Repeat Experiments § Requires Informatics Specialist who Understands Laboratory Techniques
Challenges in High Throughput Biotechnology R&D Ø Volume of Data is Growing Rapidly Ø Technology is Evolving Rapidly § Instrumentation, Informatics Ø Biological Definitions are Constantly Updated § New Interactions and Functions Discovered Daily Ø New Genes § Sometimes Homologous to Known Genes ->re-evaluate old data Ø Full Value is Realized by Integrating Multiple High-Throughput Platforms § Sequencing, Functional Screens, Small Molecule Activity Ø Up-Front Design of Data and Quality Control Databases is Crucial to Success § High Data Quality is Essential § Financially Impossible to Repeat Experiments § Requires Informatics Specialist who Understands Laboratory Techniques
 Recent Informatics Job Description Title: Information Specialist DUTIES: The role of this position is to provide scientific bioinformatics support for gene discovery research utilizing DNA microarray technology at Chiron. The successful candidate will participate in research projects within a bioinformatics team that will provide analysis and data management resources necessary to optimize research activities. REQUIREMENTS: A BS/MS in molecular biology, biochemistry and/or a field related to bioinformatics. A minimum of 1 year of biotechnology research experience working with projects and scientists in the field and 1 year experience utilizing data analysis tools in biotechnology research, specifically in the field of microarrays. Proficiency with bioinformatics programs and algorithms including both unix and desktop systems. Proficiency in data analysis programs such as Excel and statistical analysis packages used in the analysis of microarray data. Proficiency in SQL and working with relational database systems. Strong communication and teaching skills for working with colleagues and project researchers. Scientist II, Research Microarrays DUTIES; Develop and apply data analysis methods for interpreting high-throughput microarray experiments. Disseminate research results in presentations and writing. Should be flexible and work well in a team environment. REQUIREMENTS: Ph. D. in Physical Sciences, Computing Sciences or Statistics. Previous experience analyzing large data sets, modeling laboratory experiments, developing quantitative assessments of data reliability, and associated computer programming tasks.
Recent Informatics Job Description Title: Information Specialist DUTIES: The role of this position is to provide scientific bioinformatics support for gene discovery research utilizing DNA microarray technology at Chiron. The successful candidate will participate in research projects within a bioinformatics team that will provide analysis and data management resources necessary to optimize research activities. REQUIREMENTS: A BS/MS in molecular biology, biochemistry and/or a field related to bioinformatics. A minimum of 1 year of biotechnology research experience working with projects and scientists in the field and 1 year experience utilizing data analysis tools in biotechnology research, specifically in the field of microarrays. Proficiency with bioinformatics programs and algorithms including both unix and desktop systems. Proficiency in data analysis programs such as Excel and statistical analysis packages used in the analysis of microarray data. Proficiency in SQL and working with relational database systems. Strong communication and teaching skills for working with colleagues and project researchers. Scientist II, Research Microarrays DUTIES; Develop and apply data analysis methods for interpreting high-throughput microarray experiments. Disseminate research results in presentations and writing. Should be flexible and work well in a team environment. REQUIREMENTS: Ph. D. in Physical Sciences, Computing Sciences or Statistics. Previous experience analyzing large data sets, modeling laboratory experiments, developing quantitative assessments of data reliability, and associated computer programming tasks.
 Funding UC-Industry research collaborations in • bioinformatics • food and agricultural informatics • environmental informatics Slide 30 • medical informatics • computational aspects of imaging & modeling LSI Research Proposals January 26, 2001 May 22, 2001 October 2, 2001 Learn more about it… http: //lsi. ucdavis. edu Opportunity Awards Year round 30
Funding UC-Industry research collaborations in • bioinformatics • food and agricultural informatics • environmental informatics Slide 30 • medical informatics • computational aspects of imaging & modeling LSI Research Proposals January 26, 2001 May 22, 2001 October 2, 2001 Learn more about it… http: //lsi. ucdavis. edu Opportunity Awards Year round 30
 “The two technologies that will shape the next century are biotechnology and information technology” Bill Gates “The two technologies that will have the greatest impact on each other in the new millennium are biotechnology and information technology” Martina Mc. Gloughlin 31
“The two technologies that will shape the next century are biotechnology and information technology” Bill Gates “The two technologies that will have the greatest impact on each other in the new millennium are biotechnology and information technology” Martina Mc. Gloughlin 31
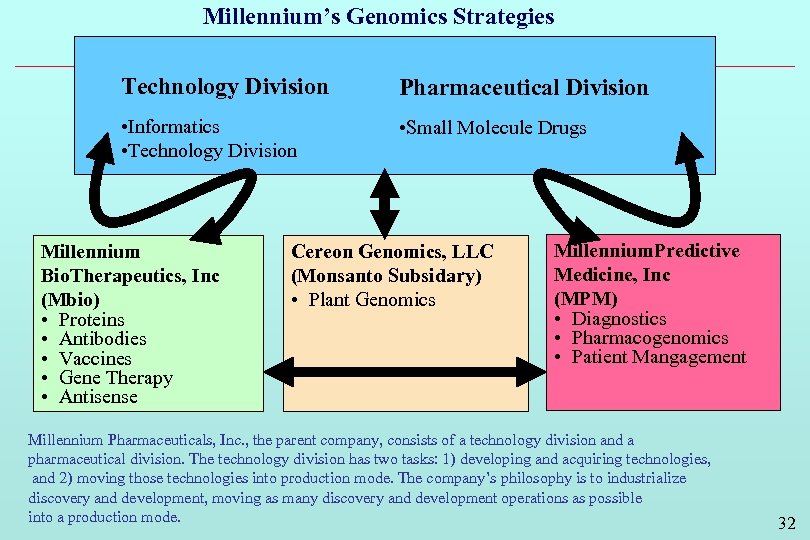 Millennium’s Genomics Strategies Technology Division Pharmaceutical Division • Informatics • Technology Division • Small Molecule Drugs Millennium Bio. Therapeutics, Inc (Mbio) • Proteins • Antibodies • Vaccines • Gene Therapy • Antisense Cereon Genomics, LLC (Monsanto Subsidary) • Plant Genomics Millennium. Predictive Medicine, Inc (MPM) • Diagnostics • Pharmacogenomics • Patient Mangagement Millennium Pharmaceuticals, Inc. , the parent company, consists of a technology division and a pharmaceutical division. The technology division has two tasks: 1) developing and acquiring technologies, and 2) moving those technologies into production mode. The company’s philosophy is to industrialize discovery and development, moving as many discovery and development operations as possible into a production mode. 32
Millennium’s Genomics Strategies Technology Division Pharmaceutical Division • Informatics • Technology Division • Small Molecule Drugs Millennium Bio. Therapeutics, Inc (Mbio) • Proteins • Antibodies • Vaccines • Gene Therapy • Antisense Cereon Genomics, LLC (Monsanto Subsidary) • Plant Genomics Millennium. Predictive Medicine, Inc (MPM) • Diagnostics • Pharmacogenomics • Patient Mangagement Millennium Pharmaceuticals, Inc. , the parent company, consists of a technology division and a pharmaceutical division. The technology division has two tasks: 1) developing and acquiring technologies, and 2) moving those technologies into production mode. The company’s philosophy is to industrialize discovery and development, moving as many discovery and development operations as possible into a production mode. 32
 33
33
 Expression Technologies There are currently four commonly used approaches to high throughput, comprehensive analysis of relative transcript expression levels. The enumeration of expressed sequence tags (ESTs), Serial Analysis of Gene Expression (SAGE), Differential Display Approaches, Array-based hybridization ¨ ¨ The enumeration of expressed sequence tags (ESTs) from representative c. DNA libraries. A method of approximating the relative representation of the gene transcript within the starting cell population. Gene. Trace Systems, HHMI, IMAGE Consortium, Incyte, The Institute for Genomic Research Serial Analysis of Gene Expression. The enumeration of serially concatenated 9 -11 base tags from specially prepared c. DNA libraries. The frequency of particular transcripts within the starting cell population is reflected by the number of times the associated sequence tag is encountered within the sequence pop Genzyme Molecular Oncology, Johns Hopkins University 34
Expression Technologies There are currently four commonly used approaches to high throughput, comprehensive analysis of relative transcript expression levels. The enumeration of expressed sequence tags (ESTs), Serial Analysis of Gene Expression (SAGE), Differential Display Approaches, Array-based hybridization ¨ ¨ The enumeration of expressed sequence tags (ESTs) from representative c. DNA libraries. A method of approximating the relative representation of the gene transcript within the starting cell population. Gene. Trace Systems, HHMI, IMAGE Consortium, Incyte, The Institute for Genomic Research Serial Analysis of Gene Expression. The enumeration of serially concatenated 9 -11 base tags from specially prepared c. DNA libraries. The frequency of particular transcripts within the starting cell population is reflected by the number of times the associated sequence tag is encountered within the sequence pop Genzyme Molecular Oncology, Johns Hopkins University 34
 Expression Technologies ¨ Differential Display Approaches Fragments defined by specific sequence delimiters can be used as unique identifiers of genes, when coupled with information about fragment length or fragment location within the expressed gene. The relative representation of an expressed gene within a cell can then be estimated based on the relative representation of the fragment associated with that gene within the pool of all possible fragments. A number of different approaches have been developed to exploit this hypothesis for comprehensive expression analysis. ¨ Curagen Corporation - Quantitative Expression Analysis (QEA) Digital Gene Technologies, Inc. - Total Gene expression Analysis (TOGA) Display Systems Biotech - Restriction Fragment Differential Display-PCR (RFDD-PCR) Genaissance Gene. Logic - Restriction Enzyme Analysis of Differentiallyexpressed Sequences (READS) ¨ ¨ 35
Expression Technologies ¨ Differential Display Approaches Fragments defined by specific sequence delimiters can be used as unique identifiers of genes, when coupled with information about fragment length or fragment location within the expressed gene. The relative representation of an expressed gene within a cell can then be estimated based on the relative representation of the fragment associated with that gene within the pool of all possible fragments. A number of different approaches have been developed to exploit this hypothesis for comprehensive expression analysis. ¨ Curagen Corporation - Quantitative Expression Analysis (QEA) Digital Gene Technologies, Inc. - Total Gene expression Analysis (TOGA) Display Systems Biotech - Restriction Fragment Differential Display-PCR (RFDD-PCR) Genaissance Gene. Logic - Restriction Enzyme Analysis of Differentiallyexpressed Sequences (READS) ¨ ¨ 35
 Expression Technologies Array-based hybridization Based on the exquisite specificity of nucleotide interactions oligonucleotides or c. DNA can be used to selectively identify or capture DNA or RNA of specific sequence composition. The primary approaches include array- based technologies that can identify specific expressed gene products on high density formats, including filters, microscope slides, or microchips, and solutionbased technologies relying on spectroscopic analyses, such as mass spectrometry. Affymetrix, Axon Instruments, Inc, Bio. Discovery Inc. Bio. Robotics, Cartesian Technologies, Clontech General Scanning Inc. , Gene. Machines, Genetic Micro. Systems Inc. , Gene. Trace Systems, Genometrix, Genomic Solution, Hyseq, Inc. Hyseq/ Applied Biosystems Division of Perkin Elmer Incyte, Intelligent Automation Systems/Intelligent Bio. Instruments, Molecular Dynamics, NHGRI Laboratory of Cancer Genetics, NEN Life Science Products Protogene, Radius Bio. Sciences, Research Genetics, Inc. Stanford University, Dr. Pat Brown, Synteni, Tele. Chem International, Rosetta Inpharmatics (Lee. Hood) 36
Expression Technologies Array-based hybridization Based on the exquisite specificity of nucleotide interactions oligonucleotides or c. DNA can be used to selectively identify or capture DNA or RNA of specific sequence composition. The primary approaches include array- based technologies that can identify specific expressed gene products on high density formats, including filters, microscope slides, or microchips, and solutionbased technologies relying on spectroscopic analyses, such as mass spectrometry. Affymetrix, Axon Instruments, Inc, Bio. Discovery Inc. Bio. Robotics, Cartesian Technologies, Clontech General Scanning Inc. , Gene. Machines, Genetic Micro. Systems Inc. , Gene. Trace Systems, Genometrix, Genomic Solution, Hyseq, Inc. Hyseq/ Applied Biosystems Division of Perkin Elmer Incyte, Intelligent Automation Systems/Intelligent Bio. Instruments, Molecular Dynamics, NHGRI Laboratory of Cancer Genetics, NEN Life Science Products Protogene, Radius Bio. Sciences, Research Genetics, Inc. Stanford University, Dr. Pat Brown, Synteni, Tele. Chem International, Rosetta Inpharmatics (Lee. Hood) 36
 Expression Technologies- Proteomics Most processes manifest themselves at the level of protein activity, but until recently, high throughput analysis of proteins was not possible. Several technologies now makes it feasible to perform mass screening of proteins ¨ 2 - D Gel Electrophoresis- Life. Protein Expression Database provides a bioinformatics platform for investigating 2 D gel images sequence data/annotation. (Incyte, Oxford Bioscience) Immobiline, Image. Master software Amersham/ Pharmacia/Molecular Dynamics, Bio. Rad ¨ LC-MS/MS, MALDI-TOF mass spectrometer offers fast and reliable protein identification for high throughput proteomic studies. MD, Perkin Elmer ¨ Per. Septive Biosystems, PE Biosystems venture, integrates robotics, mass spec, data searching technologies into 1 system for HT ID proteins, peptides ¨ SELDI (Surface-Enhanced Laser Desorption/ Ionization) Protein. Chip technology rapid separation, detection and analysis of proteins at the femtomole level directly from biological samples- Ciphergen ¨ Variants on yeast two-hybrid system, which is widely used for analyzing protein–protein interactions in vivo ¨ Phage Display 37
Expression Technologies- Proteomics Most processes manifest themselves at the level of protein activity, but until recently, high throughput analysis of proteins was not possible. Several technologies now makes it feasible to perform mass screening of proteins ¨ 2 - D Gel Electrophoresis- Life. Protein Expression Database provides a bioinformatics platform for investigating 2 D gel images sequence data/annotation. (Incyte, Oxford Bioscience) Immobiline, Image. Master software Amersham/ Pharmacia/Molecular Dynamics, Bio. Rad ¨ LC-MS/MS, MALDI-TOF mass spectrometer offers fast and reliable protein identification for high throughput proteomic studies. MD, Perkin Elmer ¨ Per. Septive Biosystems, PE Biosystems venture, integrates robotics, mass spec, data searching technologies into 1 system for HT ID proteins, peptides ¨ SELDI (Surface-Enhanced Laser Desorption/ Ionization) Protein. Chip technology rapid separation, detection and analysis of proteins at the femtomole level directly from biological samples- Ciphergen ¨ Variants on yeast two-hybrid system, which is widely used for analyzing protein–protein interactions in vivo ¨ Phage Display 37
 Genomics Center w Some of the 25 new genomics faculty will belong to the UC Davis Genome Center, the first new product of the Genomics Initiative. ($20 m set aside for faculty) w Designed to establish the campus as an international leader in functional and comparative genomics, the center will include scientists specializing in gene studies from a multitude of disciplines, including human and animal medicine, engineering, agriculture, mathematics and the biological and physical sciences. w The Genome Center will also include a revitalized pharmacology and toxicology department in the School of Medicine and a group of bioinformatics faculty members who will provide the computational biology and informatics research needed to analyze the enormous amounts of data generated by the genomics research. 38
Genomics Center w Some of the 25 new genomics faculty will belong to the UC Davis Genome Center, the first new product of the Genomics Initiative. ($20 m set aside for faculty) w Designed to establish the campus as an international leader in functional and comparative genomics, the center will include scientists specializing in gene studies from a multitude of disciplines, including human and animal medicine, engineering, agriculture, mathematics and the biological and physical sciences. w The Genome Center will also include a revitalized pharmacology and toxicology department in the School of Medicine and a group of bioinformatics faculty members who will provide the computational biology and informatics research needed to analyze the enormous amounts of data generated by the genomics research. 38
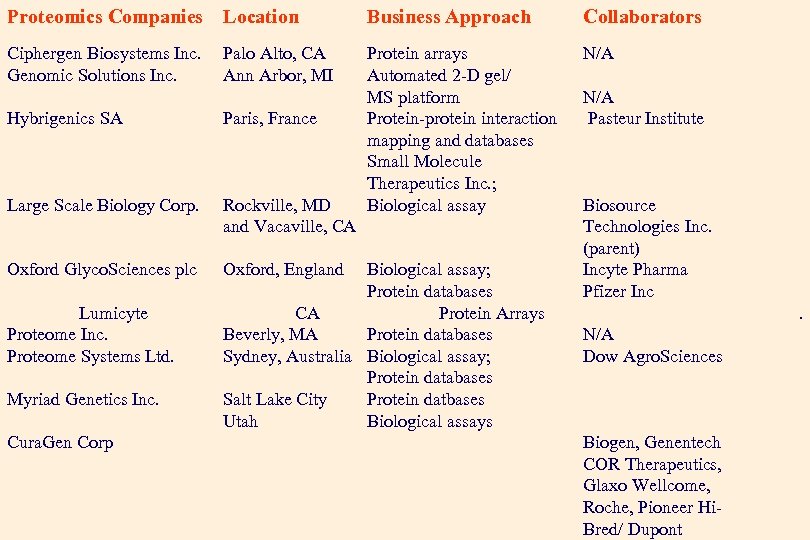 Proteomics Companies Location Business Approach Collaborators Ciphergen Biosystems Inc. Genomic Solutions Inc. Protein arrays Automated 2 -D gel/ MS platform Protein-protein interaction mapping and databases Small Molecule Therapeutics Inc. ; Biological assay N/A Palo Alto, CA Ann Arbor, MI Hybrigenics SA Paris, France Large Scale Biology Corp. Rockville, MD and Vacaville, CA Oxford Glyco. Sciences plc Oxford, England Lumicyte Proteome Inc. Proteome Systems Ltd. Myriad Genetics Inc. Cura. Gen Corp Biological assay; Protein databases CA Protein Arrays Beverly, MA Protein databases Sydney, Australia Biological assay; Protein databases Salt Lake City Protein datbases Utah Biological assays N/A Pasteur Institute Biosource Technologies Inc. (parent) Incyte Pharma Pfizer Inc. N/A Dow Agro. Sciences Biogen, Genentech COR Therapeutics, Glaxo Wellcome, Roche, Pioneer Hi. Bred/ Dupont 39
Proteomics Companies Location Business Approach Collaborators Ciphergen Biosystems Inc. Genomic Solutions Inc. Protein arrays Automated 2 -D gel/ MS platform Protein-protein interaction mapping and databases Small Molecule Therapeutics Inc. ; Biological assay N/A Palo Alto, CA Ann Arbor, MI Hybrigenics SA Paris, France Large Scale Biology Corp. Rockville, MD and Vacaville, CA Oxford Glyco. Sciences plc Oxford, England Lumicyte Proteome Inc. Proteome Systems Ltd. Myriad Genetics Inc. Cura. Gen Corp Biological assay; Protein databases CA Protein Arrays Beverly, MA Protein databases Sydney, Australia Biological assay; Protein databases Salt Lake City Protein datbases Utah Biological assays N/A Pasteur Institute Biosource Technologies Inc. (parent) Incyte Pharma Pfizer Inc. N/A Dow Agro. Sciences Biogen, Genentech COR Therapeutics, Glaxo Wellcome, Roche, Pioneer Hi. Bred/ Dupont 39
 Companies dependant on Informatics Affymtrix Agilent Alpha Gene Alpha Innotech Amersham Pharmacia Biotech Axon Instruments Bio Discovery Bio Roboti c s Biospace Mesure s Cartesian Technologies Cellomics Ciphergen Clinical Micro Sensors CLONTECH Cura. Gen Display Systems Biotech Double Twist Gene. Data Gene. Focus Gene. Machines Genetic Micro Systems Genometrix Genomic Solutions GSI Lumonics Imaging Research Iris Bio. Technologies Incyte Pharmaceutical s LION Ag Lumicyte Micronics Nanogen NEN Life Science Pro d u c t s PHASE 1 Molecular Toxicology Phoretix International Proteome Protogene Laboratories R&D Systems Radius Biosciences Research Genetics Scanalyticsl Sigma - Genosys Silicon Genetics Tele. Chem International Universal Imaging V&P Scientific Virtek Vysis 40
Companies dependant on Informatics Affymtrix Agilent Alpha Gene Alpha Innotech Amersham Pharmacia Biotech Axon Instruments Bio Discovery Bio Roboti c s Biospace Mesure s Cartesian Technologies Cellomics Ciphergen Clinical Micro Sensors CLONTECH Cura. Gen Display Systems Biotech Double Twist Gene. Data Gene. Focus Gene. Machines Genetic Micro Systems Genometrix Genomic Solutions GSI Lumonics Imaging Research Iris Bio. Technologies Incyte Pharmaceutical s LION Ag Lumicyte Micronics Nanogen NEN Life Science Pro d u c t s PHASE 1 Molecular Toxicology Phoretix International Proteome Protogene Laboratories R&D Systems Radius Biosciences Research Genetics Scanalyticsl Sigma - Genosys Silicon Genetics Tele. Chem International Universal Imaging V&P Scientific Virtek Vysis 40
 w w w w IP Challenges June 5, 1995 --Human Genome Sciences applies for a patent on a gene that produces a "receptor" protein that is later called CCR 5. HGS has no idea that CCR 5 is an HIV receptor. December 1995 --U. S. researcher Robert Gallo, the co-discoverer of HIV, and colleagues find three chemicals that inhibit the AIDS virus. Don’t how the chemicals work. February 1996 --Edward Berger at the NIH discovers that Gallo's inhibitors work in late-stage AIDS by blocking a receptor on the surface of T-cells. June 1996 --In a period of just 10 days, five groups of scientists publish papers saying CCR 5 is the receptor for virtually all strains of HIV. January 2000 --Schering-Plough researchers tell a San Francisco AIDS conference they have discovered new inhibitors. Merck researchers are known to have made similar discoveries. Feb. 15, 2000 --The U. S. Patent and Trademark Office grants HGS a patent on the gene that makes CCR 5 and on techniques for producing CCR 5 artificially. The decision sends HGS stock flying and dismays researchers. HGS: identified in whole or in part 95% of the 100, 000 or so human genes. 100 human gene patents 7, 500 pending. 41
w w w w IP Challenges June 5, 1995 --Human Genome Sciences applies for a patent on a gene that produces a "receptor" protein that is later called CCR 5. HGS has no idea that CCR 5 is an HIV receptor. December 1995 --U. S. researcher Robert Gallo, the co-discoverer of HIV, and colleagues find three chemicals that inhibit the AIDS virus. Don’t how the chemicals work. February 1996 --Edward Berger at the NIH discovers that Gallo's inhibitors work in late-stage AIDS by blocking a receptor on the surface of T-cells. June 1996 --In a period of just 10 days, five groups of scientists publish papers saying CCR 5 is the receptor for virtually all strains of HIV. January 2000 --Schering-Plough researchers tell a San Francisco AIDS conference they have discovered new inhibitors. Merck researchers are known to have made similar discoveries. Feb. 15, 2000 --The U. S. Patent and Trademark Office grants HGS a patent on the gene that makes CCR 5 and on techniques for producing CCR 5 artificially. The decision sends HGS stock flying and dismays researchers. HGS: identified in whole or in part 95% of the 100, 000 or so human genes. 100 human gene patents 7, 500 pending. 41
 The Shape of the Wave – 1999 » JGI releases 150 Mbases draft » Celera releases the sequence of Drosophila (140 Mb) » Public “draft” effort reaches halfway point (1, 500 Mb) » 20 more Microbial genomes completed (80 Mb but 60, 000 genes) » First release of Celera “shotgun” (9, 000 Mb) – 2000 » Public “draft” completed (1, 500 Mb) » Mouse “draft” begins (500 Mb - comparisons with human) » Two more Celera shotgun releases ( 18, 000 Mb) » 40 more Microbial genomes sequenced (160 Mb -120, 000 genes) 42
The Shape of the Wave – 1999 » JGI releases 150 Mbases draft » Celera releases the sequence of Drosophila (140 Mb) » Public “draft” effort reaches halfway point (1, 500 Mb) » 20 more Microbial genomes completed (80 Mb but 60, 000 genes) » First release of Celera “shotgun” (9, 000 Mb) – 2000 » Public “draft” completed (1, 500 Mb) » Mouse “draft” begins (500 Mb - comparisons with human) » Two more Celera shotgun releases ( 18, 000 Mb) » 40 more Microbial genomes sequenced (160 Mb -120, 000 genes) 42
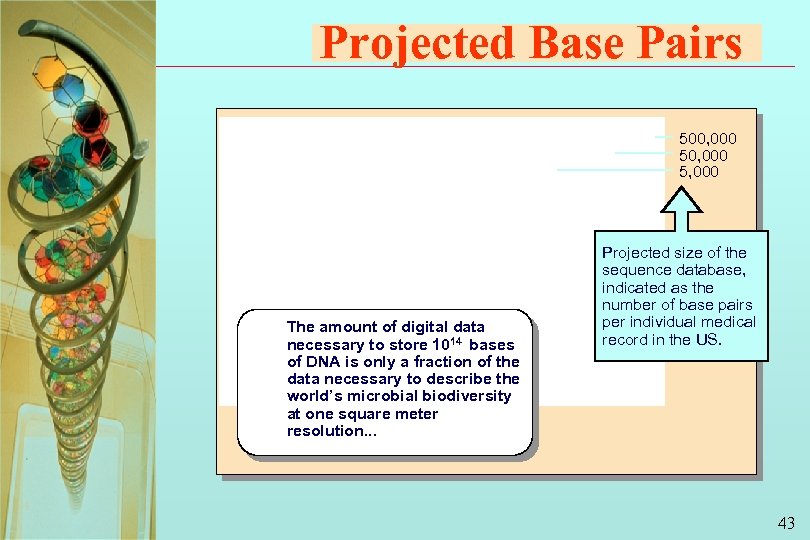 Projected Base Pairs 500, 000 5, 000 The amount of digital data necessary to store 1014 bases of DNA is only a fraction of the data necessary to describe the world’s microbial biodiversity at one square meter Year resolution. . . Projected size of the sequence database, indicated as the number of base pairs per individual medical record in the US. 43
Projected Base Pairs 500, 000 5, 000 The amount of digital data necessary to store 1014 bases of DNA is only a fraction of the data necessary to describe the world’s microbial biodiversity at one square meter Year resolution. . . Projected size of the sequence database, indicated as the number of base pairs per individual medical record in the US. 43
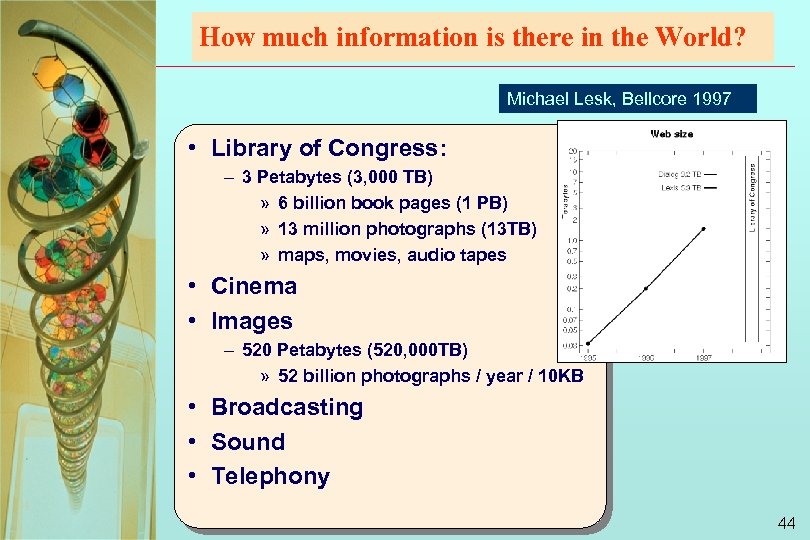 How much information is there in the World? Michael Lesk, Bellcore 1997 • Library of Congress: – 3 Petabytes (3, 000 TB) » 6 billion book pages (1 PB) » 13 million photographs (13 TB) » maps, movies, audio tapes • Cinema • Images – 520 Petabytes (520, 000 TB) » 52 billion photographs / year / 10 KB • Broadcasting • Sound • Telephony 44
How much information is there in the World? Michael Lesk, Bellcore 1997 • Library of Congress: – 3 Petabytes (3, 000 TB) » 6 billion book pages (1 PB) » 13 million photographs (13 TB) » maps, movies, audio tapes • Cinema • Images – 520 Petabytes (520, 000 TB) » 52 billion photographs / year / 10 KB • Broadcasting • Sound • Telephony 44
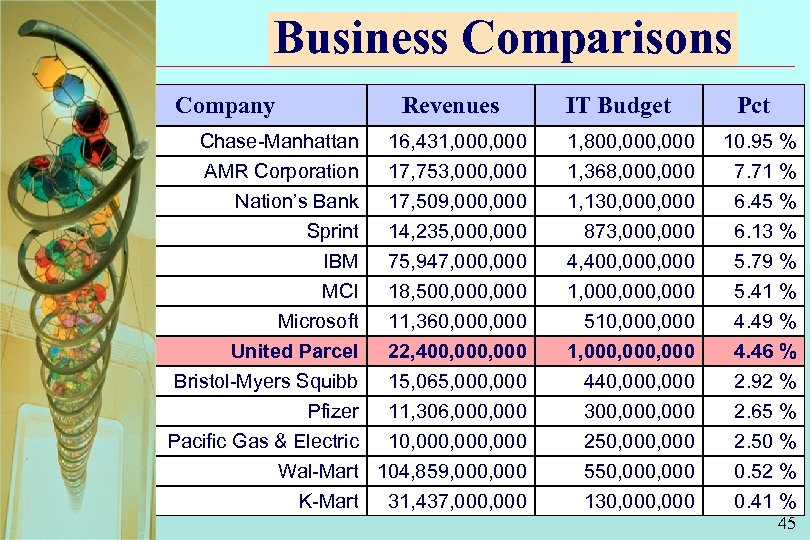 Business Comparisons Company Revenues IT Budget Pct Chase-Manhattan AMR Corporation Nation’s Bank Sprint 16, 431, 000 17, 753, 000 17, 509, 000 14, 235, 000 1, 800, 000 1, 368, 000 1, 130, 000 873, 000 10. 95 % 7. 71 % 6. 45 % 6. 13 % IBM MCI Microsoft United Parcel Bristol-Myers Squibb Pfizer Pacific Gas & Electric 75, 947, 000 18, 500, 000 11, 360, 000 22, 400, 000 15, 065, 000 11, 306, 000 10, 000, 000 4, 400, 000 1, 000, 000 510, 000 1, 000, 000 440, 000 300, 000 250, 000 5. 79 % 5. 41 % 4. 49 % 4. 46 % 2. 92 % 2. 65 % 2. 50 % Wal-Mart 104, 859, 000 K-Mart 31, 437, 000 550, 000 130, 000 0. 52 % 0. 41 % 45
Business Comparisons Company Revenues IT Budget Pct Chase-Manhattan AMR Corporation Nation’s Bank Sprint 16, 431, 000 17, 753, 000 17, 509, 000 14, 235, 000 1, 800, 000 1, 368, 000 1, 130, 000 873, 000 10. 95 % 7. 71 % 6. 45 % 6. 13 % IBM MCI Microsoft United Parcel Bristol-Myers Squibb Pfizer Pacific Gas & Electric 75, 947, 000 18, 500, 000 11, 360, 000 22, 400, 000 15, 065, 000 11, 306, 000 10, 000, 000 4, 400, 000 1, 000, 000 510, 000 1, 000, 000 440, 000 300, 000 250, 000 5. 79 % 5. 41 % 4. 49 % 4. 46 % 2. 92 % 2. 65 % 2. 50 % Wal-Mart 104, 859, 000 K-Mart 31, 437, 000 550, 000 130, 000 0. 52 % 0. 41 % 45


