8093b4135ddb904554c047c65c9d8933.ppt
- Количество слайдов: 82
 Recognition of Protein Features Limsoon Wong Institute for Infocomm Research BI 6103 guest lecture on ? ? March 2004 Copyright 2003 limsoon wong
Recognition of Protein Features Limsoon Wong Institute for Infocomm Research BI 6103 guest lecture on ? ? March 2004 Copyright 2003 limsoon wong
 Lecture Plan • Membrane proteins • Subcellular localization Copyright 2003 limsoon wong
Lecture Plan • Membrane proteins • Subcellular localization Copyright 2003 limsoon wong
 Recognition of Transmembrane Helices Copyright 2003 limsoon wong
Recognition of Transmembrane Helices Copyright 2003 limsoon wong
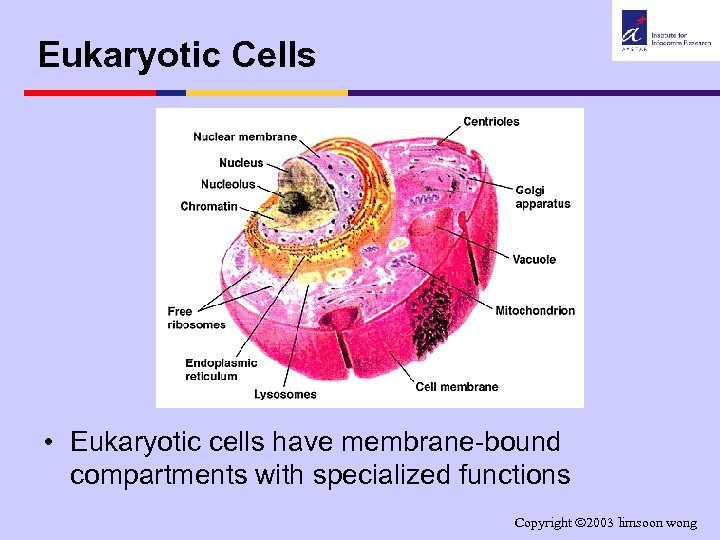 Eukaryotic Cells • Eukaryotic cells have membrane-bound compartments with specialized functions Copyright 2003 limsoon wong
Eukaryotic Cells • Eukaryotic cells have membrane-bound compartments with specialized functions Copyright 2003 limsoon wong
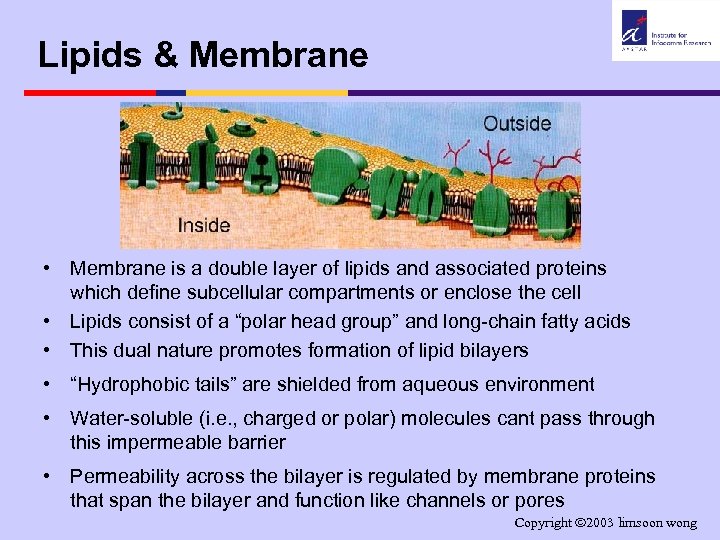 Lipids & Membrane • Membrane is a double layer of lipids and associated proteins which define subcellular compartments or enclose the cell • Lipids consist of a “polar head group” and long-chain fatty acids • This dual nature promotes formation of lipid bilayers • “Hydrophobic tails” are shielded from aqueous environment • Water-soluble (i. e. , charged or polar) molecules cant pass through this impermeable barrier • Permeability across the bilayer is regulated by membrane proteins that span the bilayer and function like channels or pores Copyright 2003 limsoon wong
Lipids & Membrane • Membrane is a double layer of lipids and associated proteins which define subcellular compartments or enclose the cell • Lipids consist of a “polar head group” and long-chain fatty acids • This dual nature promotes formation of lipid bilayers • “Hydrophobic tails” are shielded from aqueous environment • Water-soluble (i. e. , charged or polar) molecules cant pass through this impermeable barrier • Permeability across the bilayer is regulated by membrane proteins that span the bilayer and function like channels or pores Copyright 2003 limsoon wong
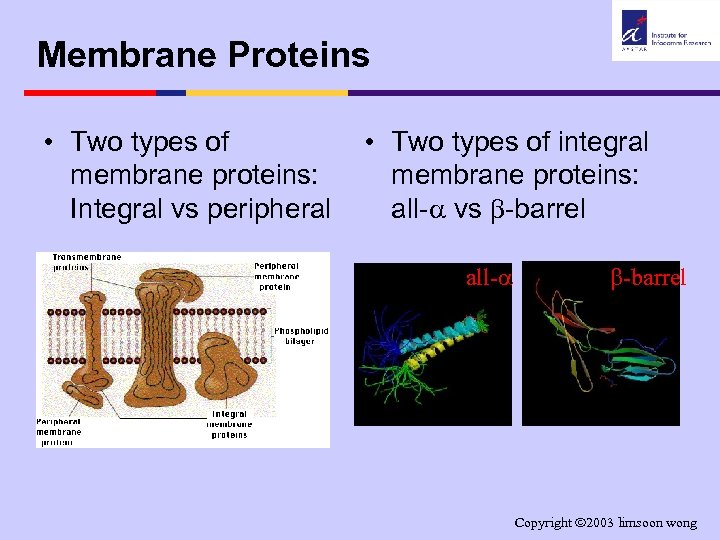 Membrane Proteins • Two types of membrane proteins: Integral vs peripheral • Two types of integral membrane proteins: all- vs -barrel all- -barrel Copyright 2003 limsoon wong
Membrane Proteins • Two types of membrane proteins: Integral vs peripheral • Two types of integral membrane proteins: all- vs -barrel all- -barrel Copyright 2003 limsoon wong
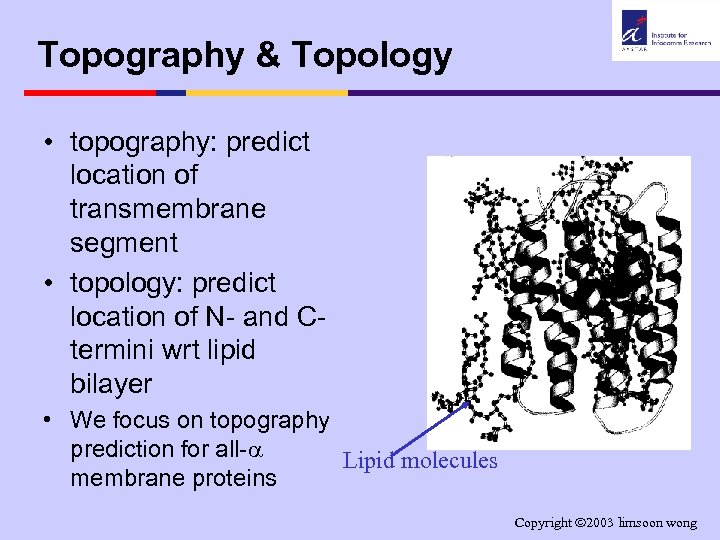 Topography & Topology • topography: predict location of transmembrane segment • topology: predict location of N- and Ctermini wrt lipid bilayer • We focus on topography prediction for all- Lipid molecules membrane proteins Copyright 2003 limsoon wong
Topography & Topology • topography: predict location of transmembrane segment • topology: predict location of N- and Ctermini wrt lipid bilayer • We focus on topography prediction for all- Lipid molecules membrane proteins Copyright 2003 limsoon wong
 Datasets • Jayasinghe et al. Protein Sci, 10: 455 -458, 2001 – 59 high resolution membrane proteins – www. biocomp. unibo. it/gigi/ENSEMBLE • Moller et al. Bioinformatics, 16: 1159 --1160, 2000 – 151 low resolution membrane proteins • Jones et al. , Biochem. , 33(10): 3038 --3049, 1994 – 38 multi-spanning and 45 single-spanning membrane proteins – topologies experimentally determined • Sonnhammer et al. , ISMB, 6: 175 -182, 1998 – 108 multi-spanning and 52 single-spanning membrane proteins – most of experimentally determined topologies, but less reliably determined than Jones et al. Copyright 2003 limsoon wong
Datasets • Jayasinghe et al. Protein Sci, 10: 455 -458, 2001 – 59 high resolution membrane proteins – www. biocomp. unibo. it/gigi/ENSEMBLE • Moller et al. Bioinformatics, 16: 1159 --1160, 2000 – 151 low resolution membrane proteins • Jones et al. , Biochem. , 33(10): 3038 --3049, 1994 – 38 multi-spanning and 45 single-spanning membrane proteins – topologies experimentally determined • Sonnhammer et al. , ISMB, 6: 175 -182, 1998 – 108 multi-spanning and 52 single-spanning membrane proteins – most of experimentally determined topologies, but less reliably determined than Jones et al. Copyright 2003 limsoon wong
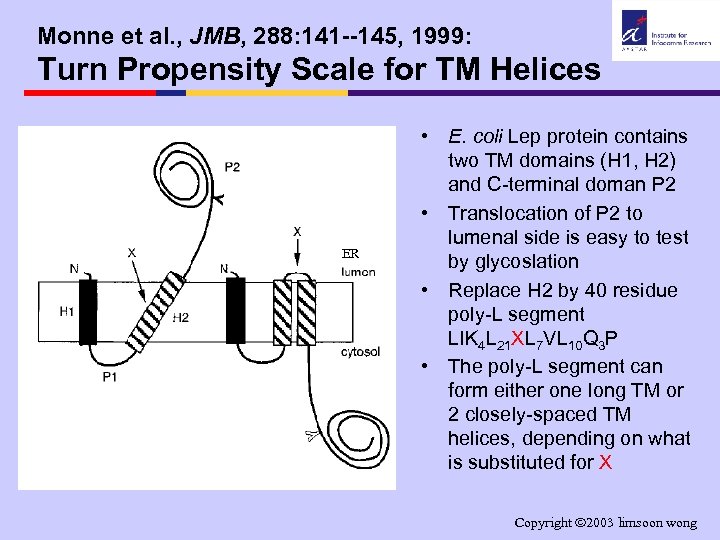 Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices ER • E. coli Lep protein contains two TM domains (H 1, H 2) and C-terminal doman P 2 • Translocation of P 2 to lumenal side is easy to test by glycoslation • Replace H 2 by 40 residue poly-L segment LIK 4 L 21 XL 7 VL 10 Q 3 P • The poly-L segment can form either one long TM or 2 closely-spaced TM helices, depending on what is substituted for X Copyright 2003 limsoon wong
Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices ER • E. coli Lep protein contains two TM domains (H 1, H 2) and C-terminal doman P 2 • Translocation of P 2 to lumenal side is easy to test by glycoslation • Replace H 2 by 40 residue poly-L segment LIK 4 L 21 XL 7 VL 10 Q 3 P • The poly-L segment can form either one long TM or 2 closely-spaced TM helices, depending on what is substituted for X Copyright 2003 limsoon wong
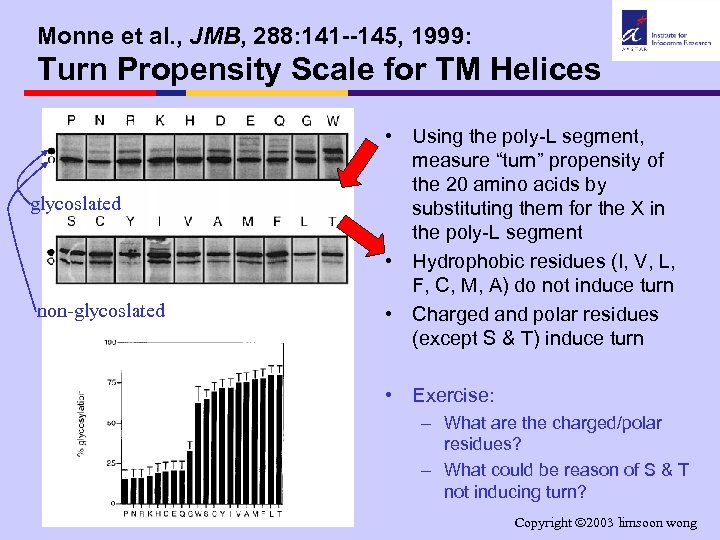 Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices glycoslated non-glycoslated • Using the poly-L segment, measure “turn” propensity of the 20 amino acids by substituting them for the X in the poly-L segment • Hydrophobic residues (I, V, L, F, C, M, A) do not induce turn • Charged and polar residues (except S & T) induce turn • Exercise: – What are the charged/polar residues? – What could be reason of S & T not inducing turn? Copyright 2003 limsoon wong
Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices glycoslated non-glycoslated • Using the poly-L segment, measure “turn” propensity of the 20 amino acids by substituting them for the X in the poly-L segment • Hydrophobic residues (I, V, L, F, C, M, A) do not induce turn • Charged and polar residues (except S & T) induce turn • Exercise: – What are the charged/polar residues? – What could be reason of S & T not inducing turn? Copyright 2003 limsoon wong
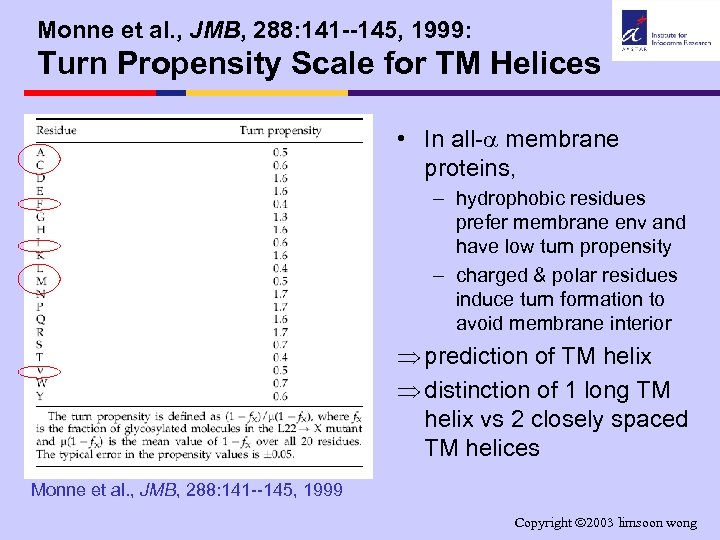 Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices • In all- membrane proteins, – hydrophobic residues prefer membrane env and have low turn propensity – charged & polar residues induce turn formation to avoid membrane interior Þ prediction of TM helix Þ distinction of 1 long TM helix vs 2 closely spaced TM helices Monne et al. , JMB, 288: 141 --145, 1999 Copyright 2003 limsoon wong
Monne et al. , JMB, 288: 141 --145, 1999: Turn Propensity Scale for TM Helices • In all- membrane proteins, – hydrophobic residues prefer membrane env and have low turn propensity – charged & polar residues induce turn formation to avoid membrane interior Þ prediction of TM helix Þ distinction of 1 long TM helix vs 2 closely spaced TM helices Monne et al. , JMB, 288: 141 --145, 1999 Copyright 2003 limsoon wong
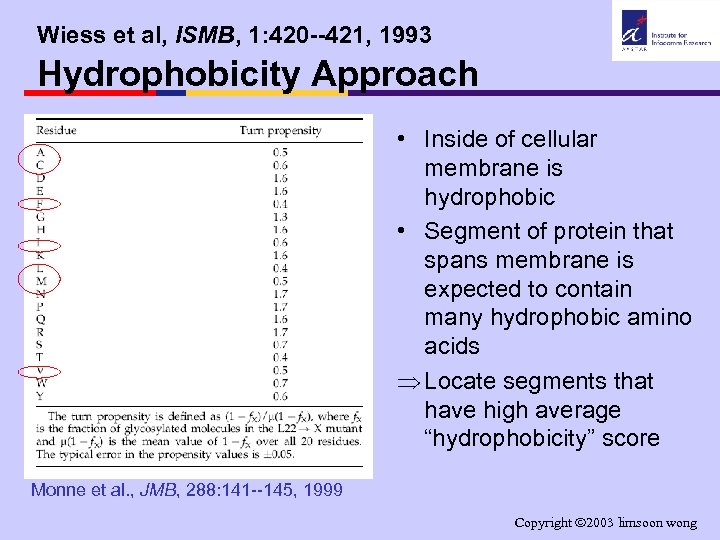 Wiess et al, ISMB, 1: 420 --421, 1993 Hydrophobicity Approach • Inside of cellular membrane is hydrophobic • Segment of protein that spans membrane is expected to contain many hydrophobic amino acids Þ Locate segments that have high average “hydrophobicity” score Monne et al. , JMB, 288: 141 --145, 1999 Copyright 2003 limsoon wong
Wiess et al, ISMB, 1: 420 --421, 1993 Hydrophobicity Approach • Inside of cellular membrane is hydrophobic • Segment of protein that spans membrane is expected to contain many hydrophobic amino acids Þ Locate segments that have high average “hydrophobicity” score Monne et al. , JMB, 288: 141 --145, 1999 Copyright 2003 limsoon wong
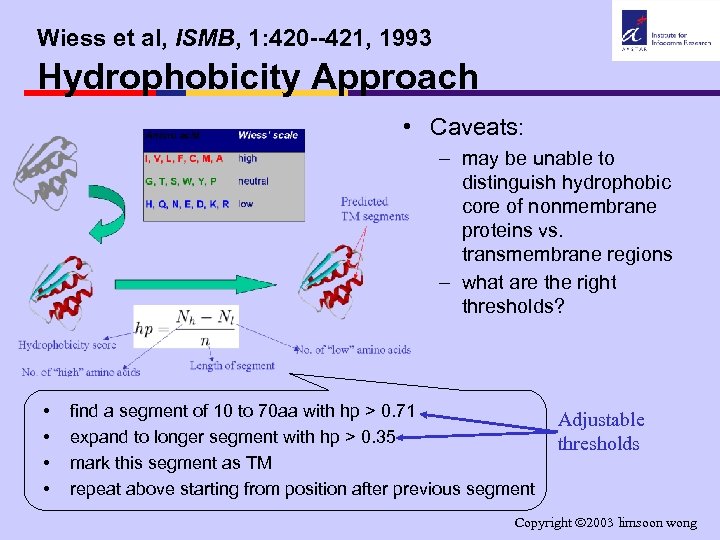 Wiess et al, ISMB, 1: 420 --421, 1993 Hydrophobicity Approach • Caveats: – may be unable to distinguish hydrophobic core of nonmembrane proteins vs. transmembrane regions – what are the right thresholds? • • find a segment of 10 to 70 aa with hp > 0. 71 expand to longer segment with hp > 0. 35 mark this segment as TM repeat above starting from position after previous segment Adjustable thresholds Copyright 2003 limsoon wong
Wiess et al, ISMB, 1: 420 --421, 1993 Hydrophobicity Approach • Caveats: – may be unable to distinguish hydrophobic core of nonmembrane proteins vs. transmembrane regions – what are the right thresholds? • • find a segment of 10 to 70 aa with hp > 0. 71 expand to longer segment with hp > 0. 35 mark this segment as TM repeat above starting from position after previous segment Adjustable thresholds Copyright 2003 limsoon wong
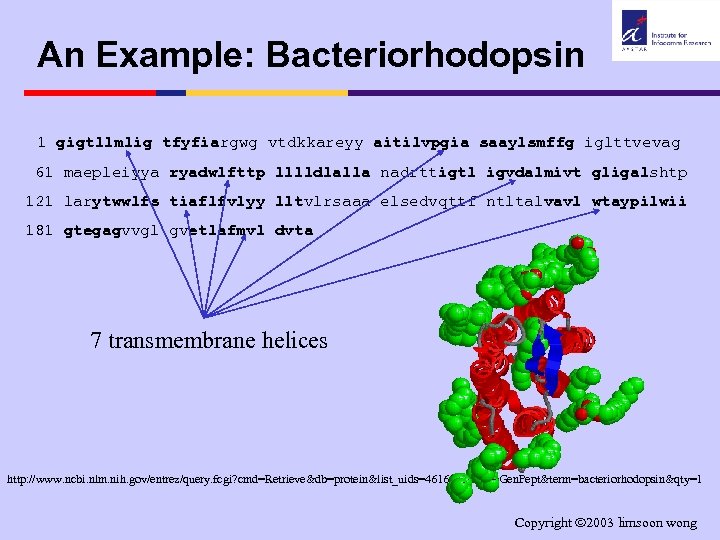 An Example: Bacteriorhodopsin 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta 7 transmembrane helices http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi? cmd=Retrieve&db=protein&list_uids=461610&dopt=Gen. Pept&term=bacteriorhodopsin&qty=1 Copyright 2003 limsoon wong
An Example: Bacteriorhodopsin 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta 7 transmembrane helices http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi? cmd=Retrieve&db=protein&list_uids=461610&dopt=Gen. Pept&term=bacteriorhodopsin&qty=1 Copyright 2003 limsoon wong
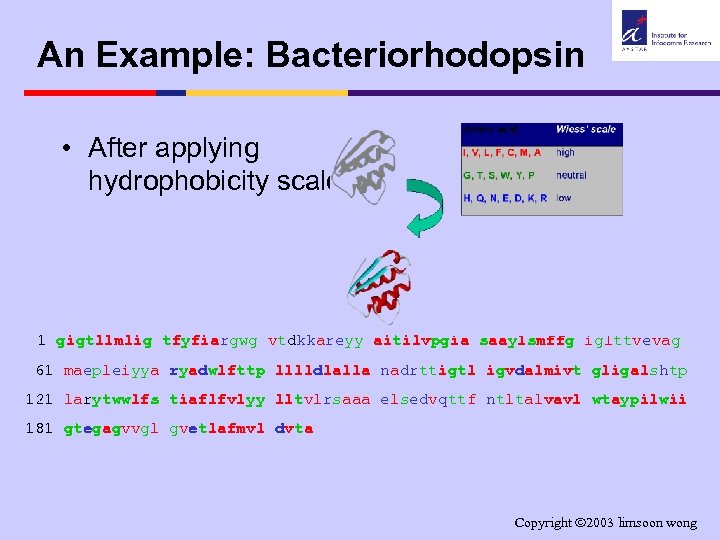 An Example: Bacteriorhodopsin • After applying hydrophobicity scale. . . 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta Copyright 2003 limsoon wong
An Example: Bacteriorhodopsin • After applying hydrophobicity scale. . . 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta Copyright 2003 limsoon wong
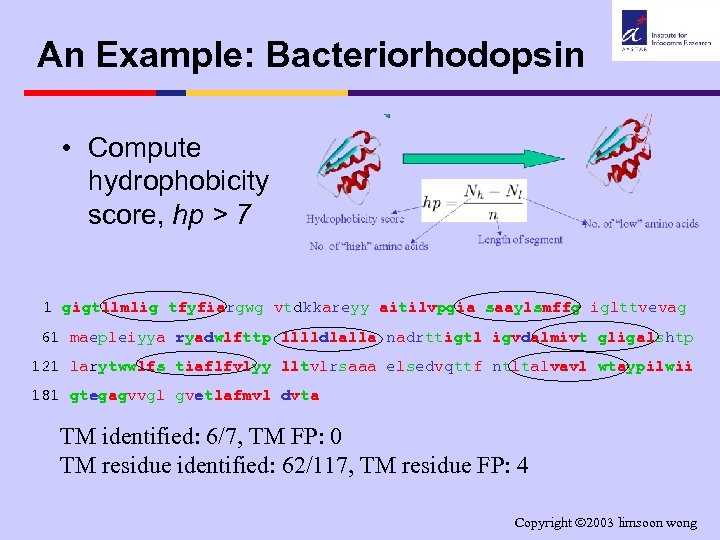 An Example: Bacteriorhodopsin • Compute hydrophobicity score, hp > 7 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta TM identified: 6/7, TM FP: 0 TM residue identified: 62/117, TM residue FP: 4 Copyright 2003 limsoon wong
An Example: Bacteriorhodopsin • Compute hydrophobicity score, hp > 7 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta TM identified: 6/7, TM FP: 0 TM residue identified: 62/117, TM residue FP: 4 Copyright 2003 limsoon wong
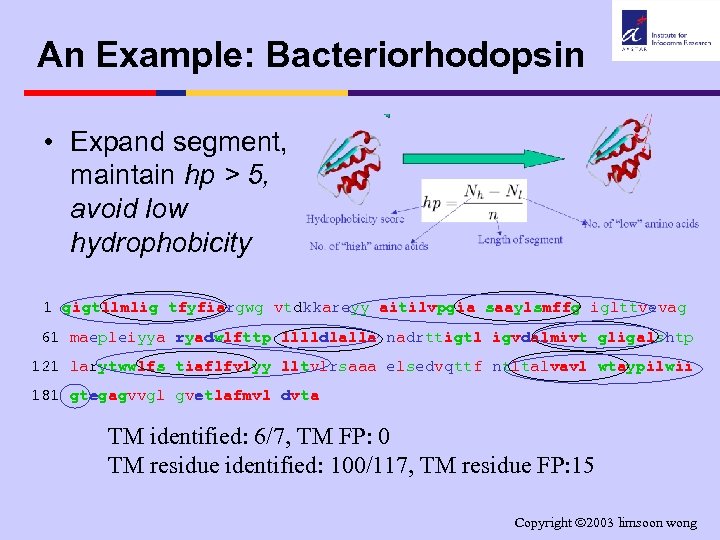 An Example: Bacteriorhodopsin • Expand segment, maintain hp > 5, avoid low hydrophobicity 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta TM identified: 6/7, TM FP: 0 TM residue identified: 100/117, TM residue FP: 15 Copyright 2003 limsoon wong
An Example: Bacteriorhodopsin • Expand segment, maintain hp > 5, avoid low hydrophobicity 1 gigtllmlig tfyfiargwg vtdkkareyy aitilvpgia saaylsmffg iglttvevag 61 maepleiyya ryadwlfttp lllldlalla nadrttigtl igvdalmivt gligalshtp 121 larytwwlfs tiaflfvlyy lltvlrsaaa elsedvqttf ntltalvavl wtaypilwii 181 gtegagvvgl gvetlafmvl dvta TM identified: 6/7, TM FP: 0 TM residue identified: 100/117, TM residue FP: 15 Copyright 2003 limsoon wong
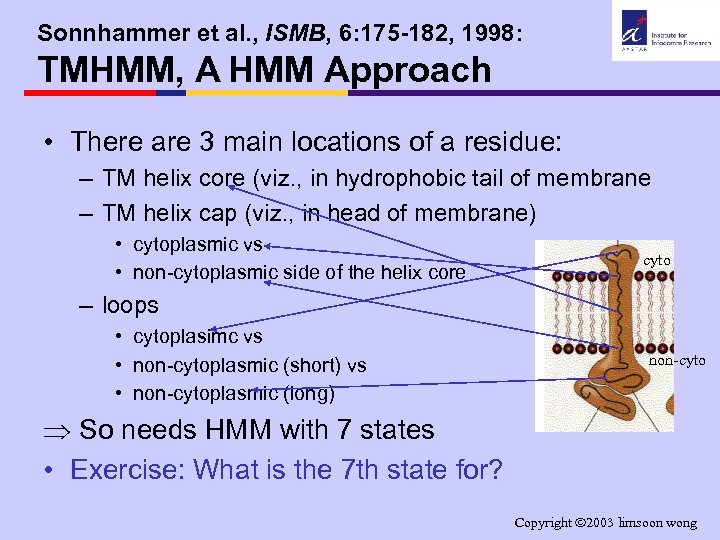 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, A HMM Approach • There are 3 main locations of a residue: – TM helix core (viz. , in hydrophobic tail of membrane – TM helix cap (viz. , in head of membrane) • cytoplasmic vs • non-cytoplasmic side of the helix core cyto – loops • cytoplasimc vs • non-cytoplasmic (short) vs • non-cytoplasmic (long) non-cyto Þ So needs HMM with 7 states • Exercise: What is the 7 th state for? Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, A HMM Approach • There are 3 main locations of a residue: – TM helix core (viz. , in hydrophobic tail of membrane – TM helix cap (viz. , in head of membrane) • cytoplasmic vs • non-cytoplasmic side of the helix core cyto – loops • cytoplasimc vs • non-cytoplasmic (short) vs • non-cytoplasmic (long) non-cyto Þ So needs HMM with 7 states • Exercise: What is the 7 th state for? Copyright 2003 limsoon wong
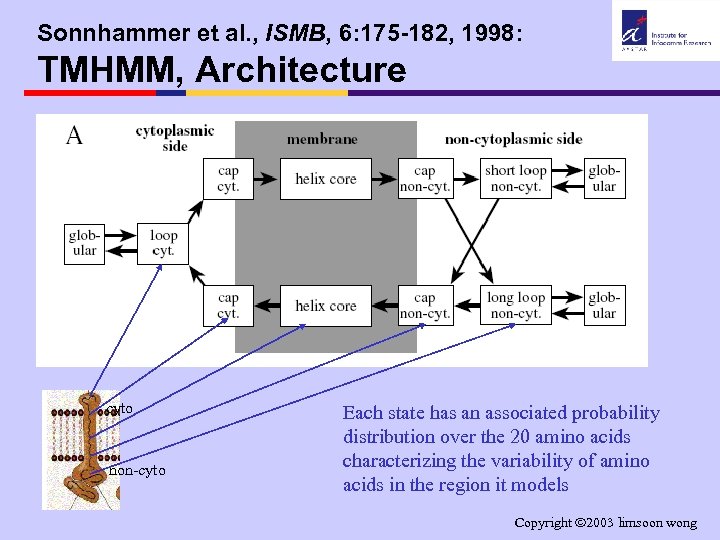 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture cyto non-cyto Each state has an associated probability distribution over the 20 amino acids characterizing the variability of amino acids in the region it models Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture cyto non-cyto Each state has an associated probability distribution over the 20 amino acids characterizing the variability of amino acids in the region it models Copyright 2003 limsoon wong
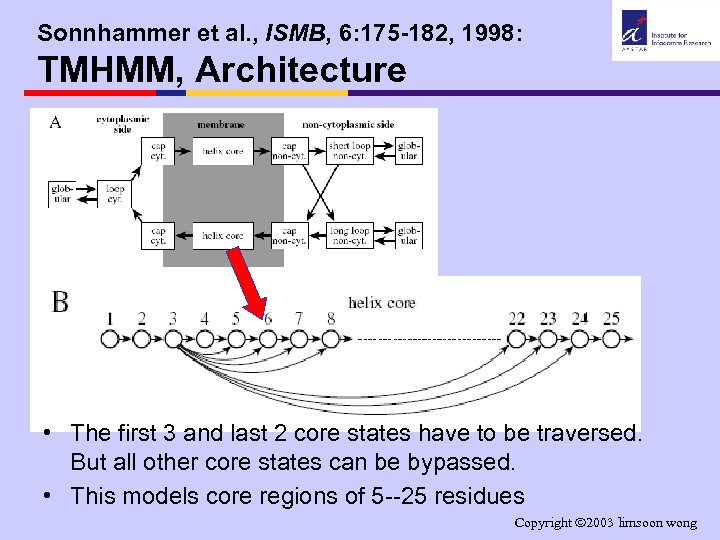 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture • The first 3 and last 2 core states have to be traversed. But all other core states can be bypassed. • This models core regions of 5 --25 residues Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture • The first 3 and last 2 core states have to be traversed. But all other core states can be bypassed. • This models core regions of 5 --25 residues Copyright 2003 limsoon wong
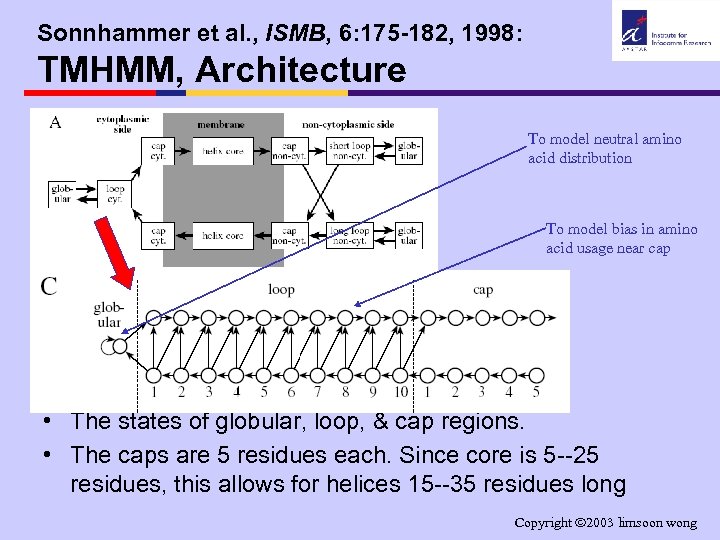 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture To model neutral amino acid distribution To model bias in amino acid usage near cap • The states of globular, loop, & cap regions. • The caps are 5 residues each. Since core is 5 --25 residues, this allows for helices 15 --35 residues long Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Architecture To model neutral amino acid distribution To model bias in amino acid usage near cap • The states of globular, loop, & cap regions. • The caps are 5 residues each. Since core is 5 --25 residues, this allows for helices 15 --35 residues long Copyright 2003 limsoon wong
 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Training the HMM • Stage 1: Baum-Welch is used for maximum likelihood estimation from “diluted” labeled training data. As precise end of TM is only approximately known, we “dilute” by unlabeling 3 residues on each side of a helix boundary to accommodate this • Stage 2: Baum-Welch is used for maximum likelihood estimation from “relabeled” training data. The original training data are diluted as by unlabeling 5 residues on each side of a helix boundary. Model from Stage 1 is used to produce “relabeled training data” by relabeling this part under constraints of remaining labels • Stage 3: Model from Stage 2 is further tuned by a method for “discriminative” training, to maximize probability of correct prediction (Krogh, ISMB, 5: 179 --186, 1997) Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Training the HMM • Stage 1: Baum-Welch is used for maximum likelihood estimation from “diluted” labeled training data. As precise end of TM is only approximately known, we “dilute” by unlabeling 3 residues on each side of a helix boundary to accommodate this • Stage 2: Baum-Welch is used for maximum likelihood estimation from “relabeled” training data. The original training data are diluted as by unlabeling 5 residues on each side of a helix boundary. Model from Stage 1 is used to produce “relabeled training data” by relabeling this part under constraints of remaining labels • Stage 3: Model from Stage 2 is further tuned by a method for “discriminative” training, to maximize probability of correct prediction (Krogh, ISMB, 5: 179 --186, 1997) Copyright 2003 limsoon wong
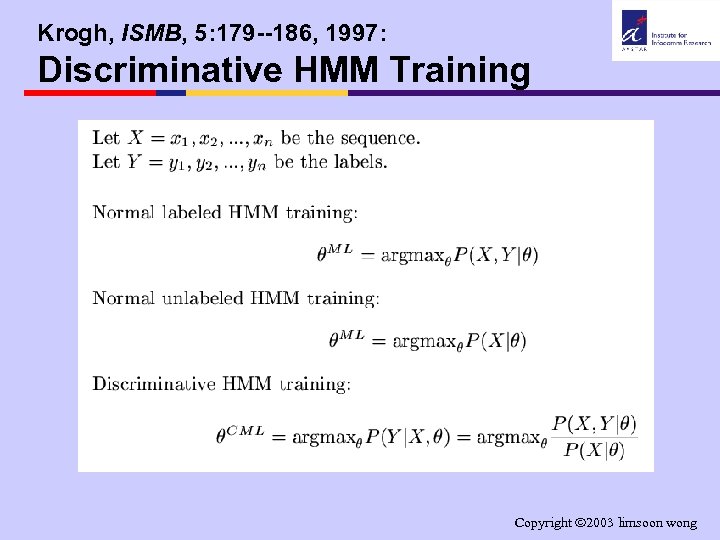 Krogh, ISMB, 5: 179 --186, 1997: Discriminative HMM Training Copyright 2003 limsoon wong
Krogh, ISMB, 5: 179 --186, 1997: Discriminative HMM Training Copyright 2003 limsoon wong
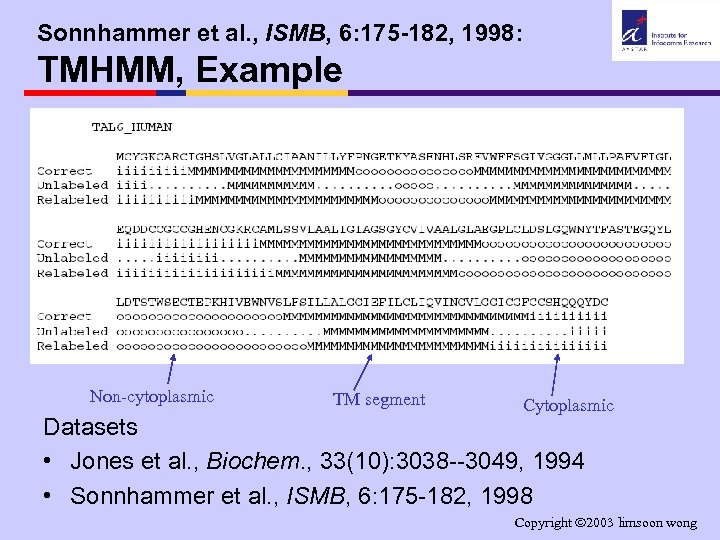 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Example Non-cytoplasmic TM segment Cytoplasmic Datasets • Jones et al. , Biochem. , 33(10): 3038 --3049, 1994 • Sonnhammer et al. , ISMB, 6: 175 -182, 1998 Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Example Non-cytoplasmic TM segment Cytoplasmic Datasets • Jones et al. , Biochem. , 33(10): 3038 --3049, 1994 • Sonnhammer et al. , ISMB, 6: 175 -182, 1998 Copyright 2003 limsoon wong
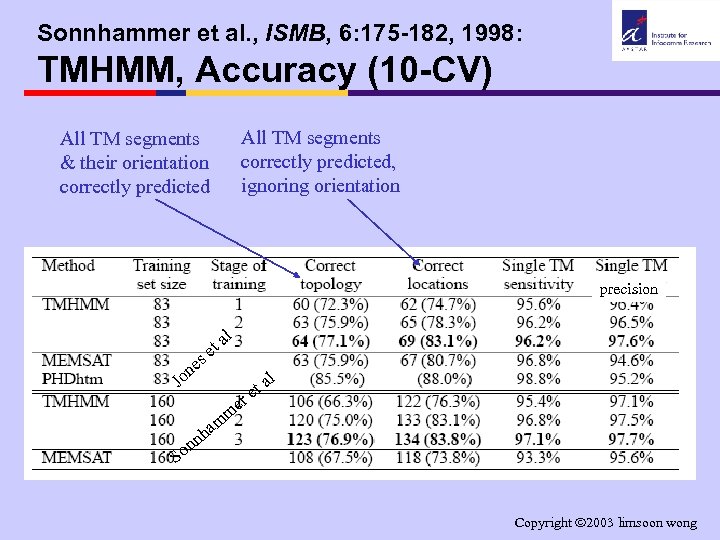 Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Accuracy (10 -CV) All TM segments correctly predicted, ignoring orientation All TM segments & their orientation correctly predicted precision l a et s ne l Jo r n n So me m ha a et Copyright 2003 limsoon wong
Sonnhammer et al. , ISMB, 6: 175 -182, 1998: TMHMM, Accuracy (10 -CV) All TM segments correctly predicted, ignoring orientation All TM segments & their orientation correctly predicted precision l a et s ne l Jo r n n So me m ha a et Copyright 2003 limsoon wong
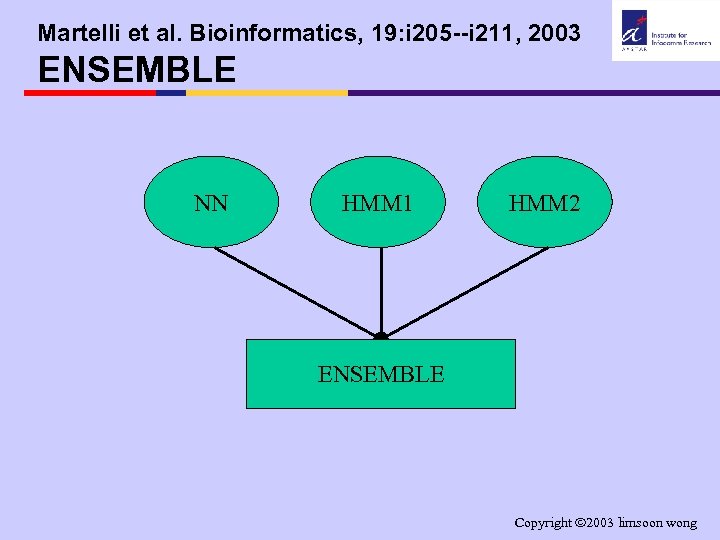 Martelli et al. Bioinformatics, 19: i 205 --i 211, 2003 ENSEMBLE NN HMM 1 HMM 2 ENSEMBLE Copyright 2003 limsoon wong
Martelli et al. Bioinformatics, 19: i 205 --i 211, 2003 ENSEMBLE NN HMM 1 HMM 2 ENSEMBLE Copyright 2003 limsoon wong
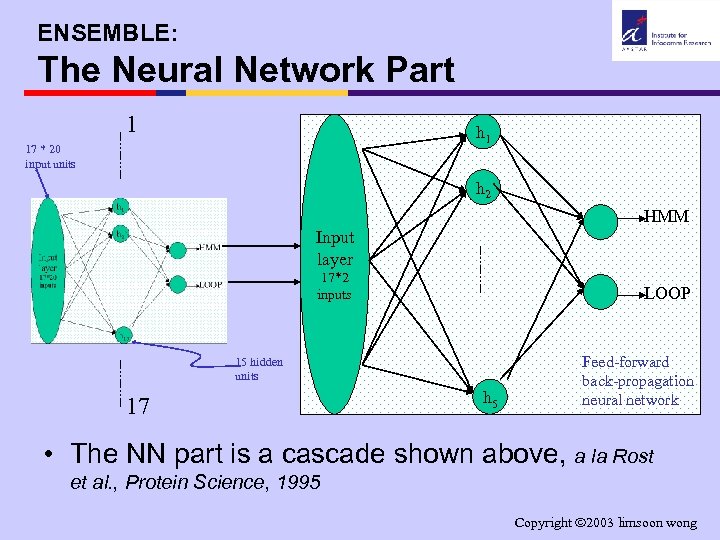 ENSEMBLE: The Neural Network Part 1 h 1 17 * 20 input units h 2 HMM Input layer 17*2 inputs LOOP 15 hidden units 17 h 5 Feed-forward back-propagation neural network • The NN part is a cascade shown above, a la Rost et al. , Protein Science, 1995 Copyright 2003 limsoon wong
ENSEMBLE: The Neural Network Part 1 h 1 17 * 20 input units h 2 HMM Input layer 17*2 inputs LOOP 15 hidden units 17 h 5 Feed-forward back-propagation neural network • The NN part is a cascade shown above, a la Rost et al. , Protein Science, 1995 Copyright 2003 limsoon wong
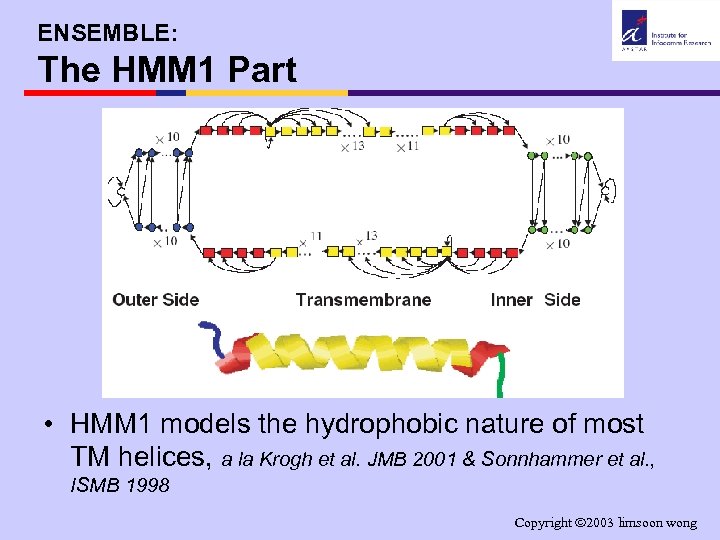 ENSEMBLE: The HMM 1 Part • HMM 1 models the hydrophobic nature of most TM helices, a la Krogh et al. JMB 2001 & Sonnhammer et al. , ISMB 1998 Copyright 2003 limsoon wong
ENSEMBLE: The HMM 1 Part • HMM 1 models the hydrophobic nature of most TM helices, a la Krogh et al. JMB 2001 & Sonnhammer et al. , ISMB 1998 Copyright 2003 limsoon wong
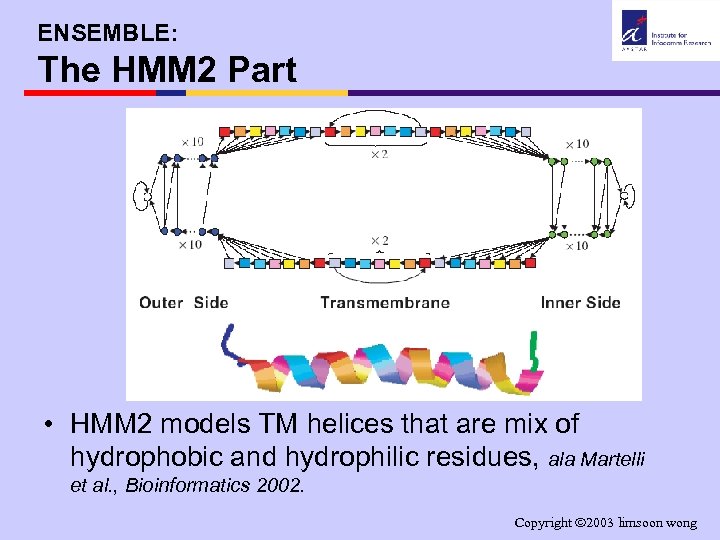 ENSEMBLE: The HMM 2 Part • HMM 2 models TM helices that are mix of hydrophobic and hydrophilic residues, ala Martelli et al. , Bioinformatics 2002. Copyright 2003 limsoon wong
ENSEMBLE: The HMM 2 Part • HMM 2 models TM helices that are mix of hydrophobic and hydrophilic residues, ala Martelli et al. , Bioinformatics 2002. Copyright 2003 limsoon wong
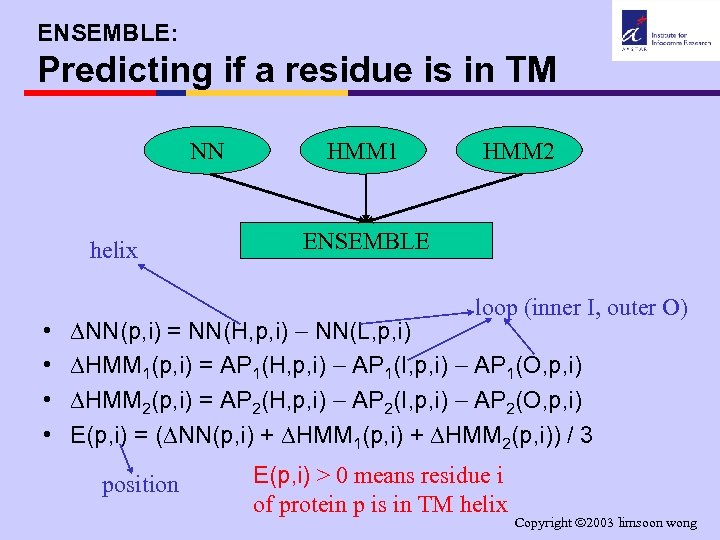 ENSEMBLE: Predicting if a residue is in TM NN helix • • HMM 1 HMM 2 ENSEMBLE loop (inner I, outer O) NN(p, i) = NN(H, p, i) NN(L, p, i) HMM 1(p, i) = AP 1(H, p, i) AP 1(I, p, i) AP 1(O, p, i) HMM 2(p, i) = AP 2(H, p, i) AP 2(I, p, i) AP 2(O, p, i) E(p, i) = ( NN(p, i) + HMM 1(p, i) + HMM 2(p, i)) / 3 position E(p, i) > 0 means residue i of protein p is in TM helix Copyright 2003 limsoon wong
ENSEMBLE: Predicting if a residue is in TM NN helix • • HMM 1 HMM 2 ENSEMBLE loop (inner I, outer O) NN(p, i) = NN(H, p, i) NN(L, p, i) HMM 1(p, i) = AP 1(H, p, i) AP 1(I, p, i) AP 1(O, p, i) HMM 2(p, i) = AP 2(H, p, i) AP 2(I, p, i) AP 2(O, p, i) E(p, i) = ( NN(p, i) + HMM 1(p, i) + HMM 2(p, i)) / 3 position E(p, i) > 0 means residue i of protein p is in TM helix Copyright 2003 limsoon wong
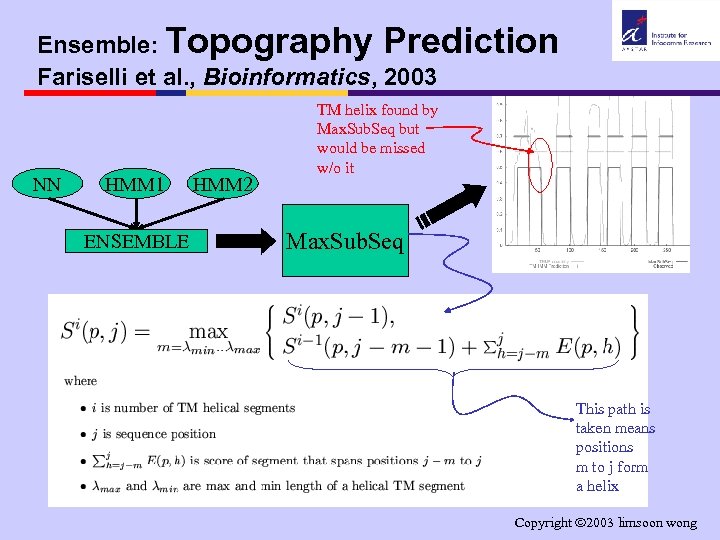 Ensemble: Topography Prediction Fariselli et al. , Bioinformatics, 2003 NN HMM 1 ENSEMBLE HMM 2 TM helix found by Max. Sub. Seq but would be missed w/o it Max. Sub. Seq This path is taken means positions m to j form a helix Copyright 2003 limsoon wong
Ensemble: Topography Prediction Fariselli et al. , Bioinformatics, 2003 NN HMM 1 ENSEMBLE HMM 2 TM helix found by Max. Sub. Seq but would be missed w/o it Max. Sub. Seq This path is taken means positions m to j form a helix Copyright 2003 limsoon wong
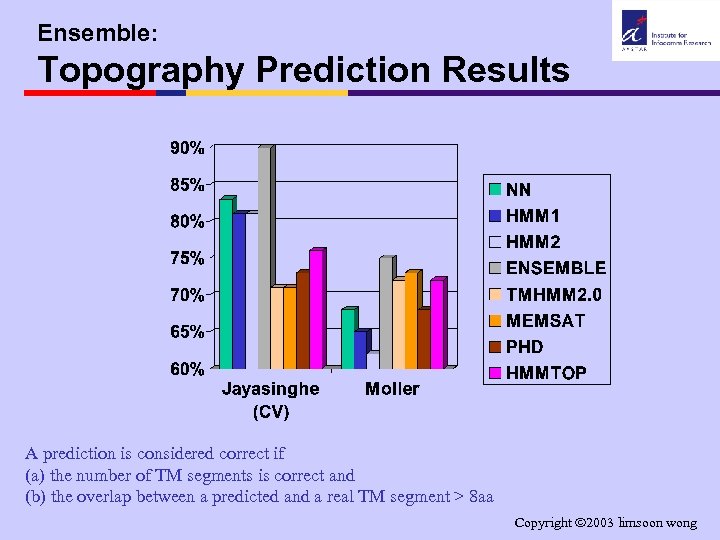 Ensemble: Topography Prediction Results A prediction is considered correct if (a) the number of TM segments is correct and (b) the overlap between a predicted and a real TM segment > 8 aa Copyright 2003 limsoon wong
Ensemble: Topography Prediction Results A prediction is considered correct if (a) the number of TM segments is correct and (b) the overlap between a predicted and a real TM segment > 8 aa Copyright 2003 limsoon wong
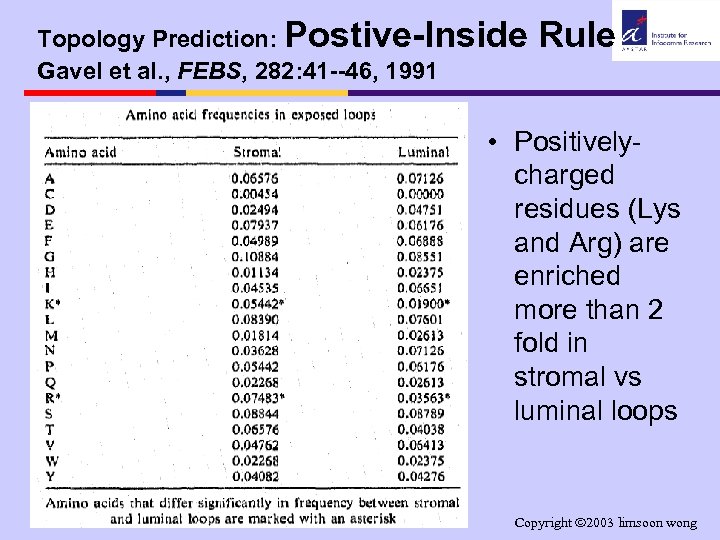 Topology Prediction: Postive-Inside Gavel et al. , FEBS, 282: 41 --46, 1991 Rule • Positivelycharged residues (Lys and Arg) are enriched more than 2 fold in stromal vs luminal loops Copyright 2003 limsoon wong
Topology Prediction: Postive-Inside Gavel et al. , FEBS, 282: 41 --46, 1991 Rule • Positivelycharged residues (Lys and Arg) are enriched more than 2 fold in stromal vs luminal loops Copyright 2003 limsoon wong
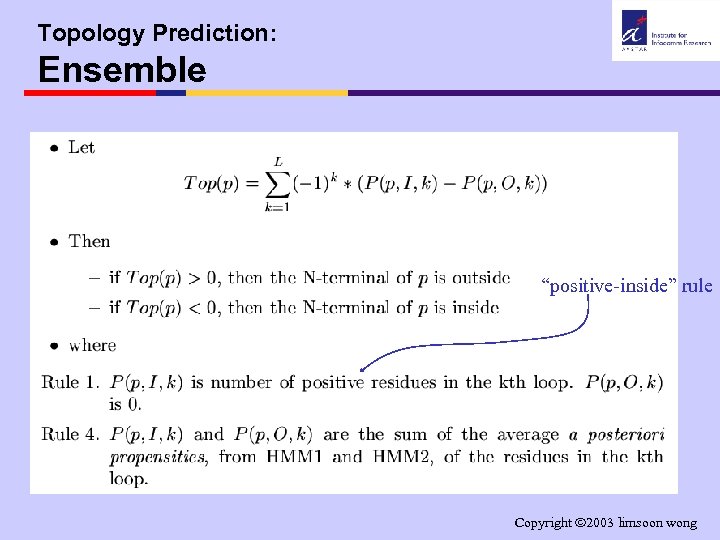 Topology Prediction: Ensemble “positive-inside” rule Copyright 2003 limsoon wong
Topology Prediction: Ensemble “positive-inside” rule Copyright 2003 limsoon wong
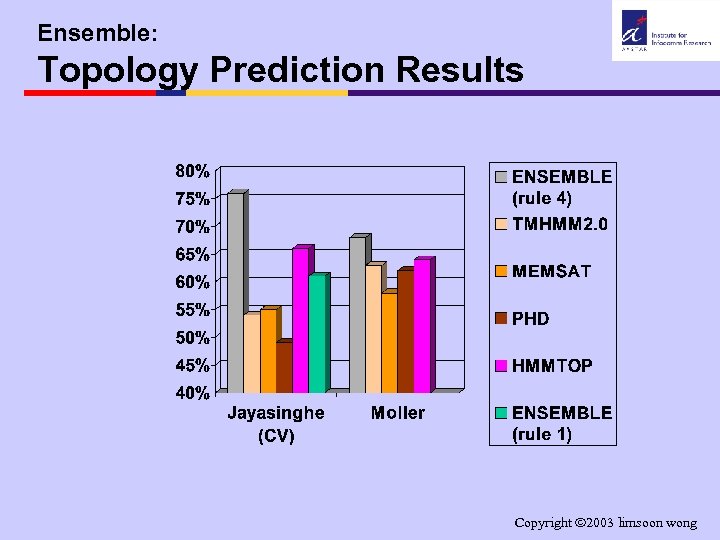 Ensemble: Topology Prediction Results Copyright 2003 limsoon wong
Ensemble: Topology Prediction Results Copyright 2003 limsoon wong
 Short Break Copyright 2003 limsoon wong
Short Break Copyright 2003 limsoon wong
 Subcellular Localization Copyright 2003 limsoon wong
Subcellular Localization Copyright 2003 limsoon wong
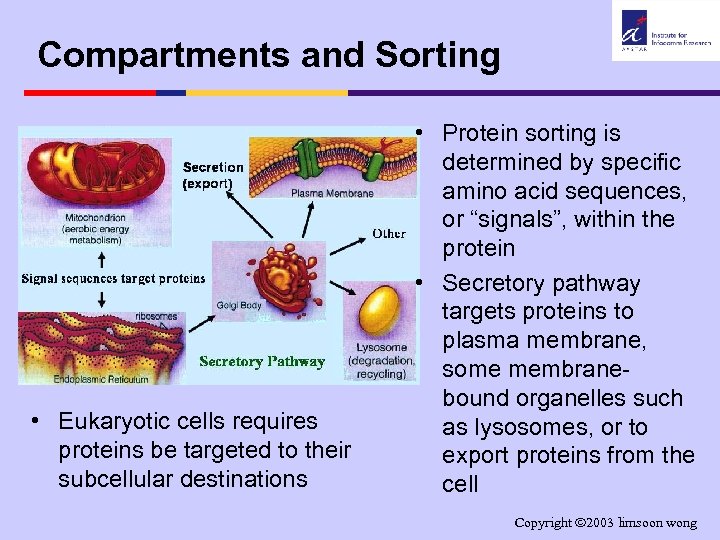 Compartments and Sorting • Eukaryotic cells requires proteins be targeted to their subcellular destinations • Protein sorting is determined by specific amino acid sequences, or “signals”, within the protein • Secretory pathway targets proteins to plasma membrane, some membranebound organelles such as lysosomes, or to export proteins from the cell Copyright 2003 limsoon wong
Compartments and Sorting • Eukaryotic cells requires proteins be targeted to their subcellular destinations • Protein sorting is determined by specific amino acid sequences, or “signals”, within the protein • Secretory pathway targets proteins to plasma membrane, some membranebound organelles such as lysosomes, or to export proteins from the cell Copyright 2003 limsoon wong
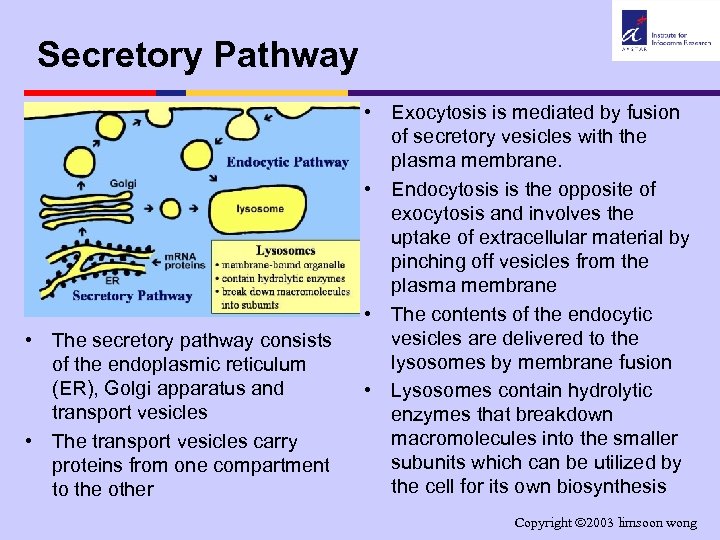 Secretory Pathway • The secretory pathway consists of the endoplasmic reticulum (ER), Golgi apparatus and transport vesicles • The transport vesicles carry proteins from one compartment to the other • Exocytosis is mediated by fusion of secretory vesicles with the plasma membrane. • Endocytosis is the opposite of exocytosis and involves the uptake of extracellular material by pinching off vesicles from the plasma membrane • The contents of the endocytic vesicles are delivered to the lysosomes by membrane fusion • Lysosomes contain hydrolytic enzymes that breakdown macromolecules into the smaller subunits which can be utilized by the cell for its own biosynthesis Copyright 2003 limsoon wong
Secretory Pathway • The secretory pathway consists of the endoplasmic reticulum (ER), Golgi apparatus and transport vesicles • The transport vesicles carry proteins from one compartment to the other • Exocytosis is mediated by fusion of secretory vesicles with the plasma membrane. • Endocytosis is the opposite of exocytosis and involves the uptake of extracellular material by pinching off vesicles from the plasma membrane • The contents of the endocytic vesicles are delivered to the lysosomes by membrane fusion • Lysosomes contain hydrolytic enzymes that breakdown macromolecules into the smaller subunits which can be utilized by the cell for its own biosynthesis Copyright 2003 limsoon wong
 Datasets • Reinhartdt & Hubbard, NAR, 26: 2230 --2236, 1998 – 2427 eukaryotic proteins for 4 locations (cytoplasmic, extracellular, nuclear, & mitochondrial) – 997 prokaryotic proteins for 3 locations (cytoplasmic, extracellular, & periplasmic) • Park & Kanehisa, Bioinformatics, 19: 1656 --1663, 2003 – 7589 eukaryotic proteins from 709 organisms for 12 locations (chloroplast, cytoplasmic, cytoskeleton, ER, extracellular, golgi, lysosomal, mitochondrial, nuclear, peroxisomal, plasma membrane, vacuolar) • Chou & Cai, JBC. , 277: 45765 --45769, 2002 – 2191 proteins for 12 locations • Emanuelsson et al. , JMB, 300: 1005 --1016, 2000 • Gardy et al. , NAR, 31: 3613 --3617, 2003 Copyright 2003 limsoon wong
Datasets • Reinhartdt & Hubbard, NAR, 26: 2230 --2236, 1998 – 2427 eukaryotic proteins for 4 locations (cytoplasmic, extracellular, nuclear, & mitochondrial) – 997 prokaryotic proteins for 3 locations (cytoplasmic, extracellular, & periplasmic) • Park & Kanehisa, Bioinformatics, 19: 1656 --1663, 2003 – 7589 eukaryotic proteins from 709 organisms for 12 locations (chloroplast, cytoplasmic, cytoskeleton, ER, extracellular, golgi, lysosomal, mitochondrial, nuclear, peroxisomal, plasma membrane, vacuolar) • Chou & Cai, JBC. , 277: 45765 --45769, 2002 – 2191 proteins for 12 locations • Emanuelsson et al. , JMB, 300: 1005 --1016, 2000 • Gardy et al. , NAR, 31: 3613 --3617, 2003 Copyright 2003 limsoon wong
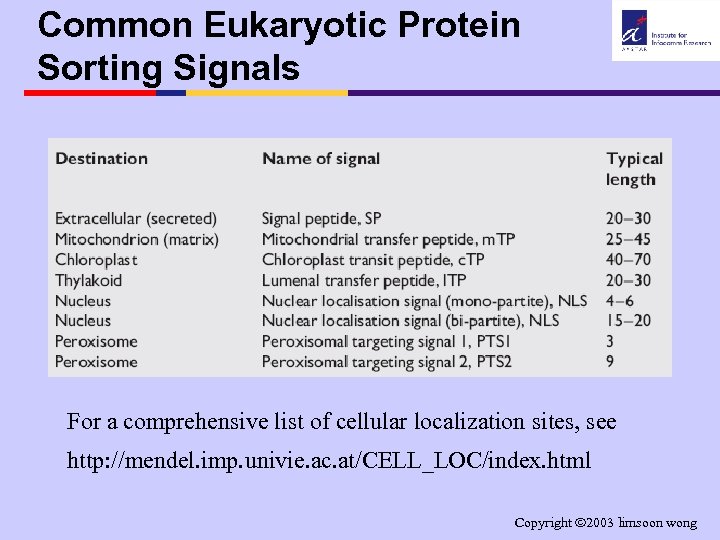 Common Eukaryotic Protein Sorting Signals For a comprehensive list of cellular localization sites, see http: //mendel. imp. univie. ac. at/CELL_LOC/index. html Copyright 2003 limsoon wong
Common Eukaryotic Protein Sorting Signals For a comprehensive list of cellular localization sites, see http: //mendel. imp. univie. ac. at/CELL_LOC/index. html Copyright 2003 limsoon wong
 ~25 aa Schematic View of Sorting Signals cleavage site Copyright 2003 limsoon wong
~25 aa Schematic View of Sorting Signals cleavage site Copyright 2003 limsoon wong
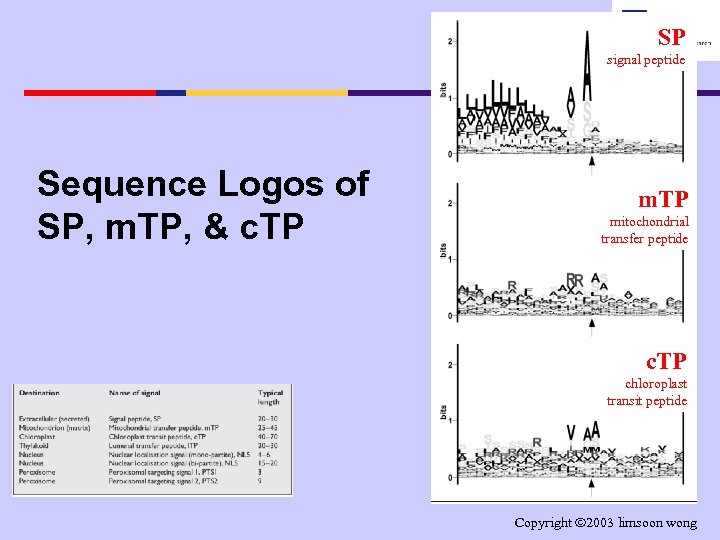 SP signal peptide Sequence Logos of SP, m. TP, & c. TP mitochondrial transfer peptide c. TP chloroplast transit peptide Copyright 2003 limsoon wong
SP signal peptide Sequence Logos of SP, m. TP, & c. TP mitochondrial transfer peptide c. TP chloroplast transit peptide Copyright 2003 limsoon wong
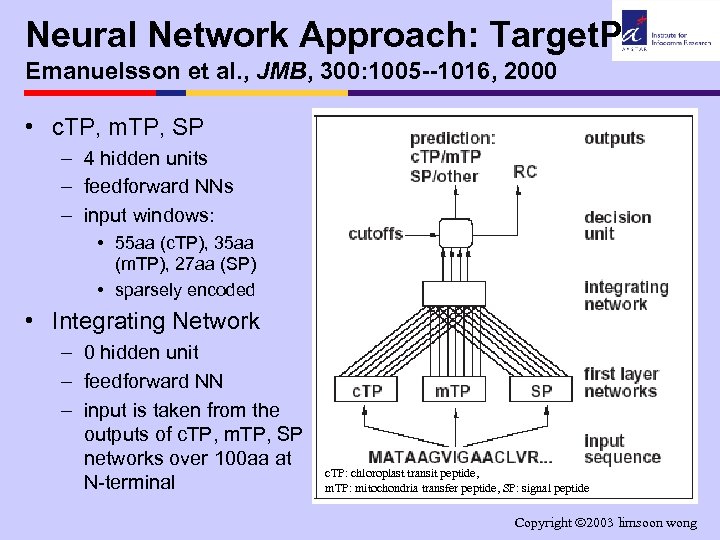 Neural Network Approach: Target. P Emanuelsson et al. , JMB, 300: 1005 --1016, 2000 • c. TP, m. TP, SP – 4 hidden units – feedforward NNs – input windows: • 55 aa (c. TP), 35 aa (m. TP), 27 aa (SP) • sparsely encoded • Integrating Network – 0 hidden unit – feedforward NN – input is taken from the outputs of c. TP, m. TP, SP networks over 100 aa at N-terminal c. TP: chloroplast transit peptide, m. TP: mitochondria transfer peptide, SP: signal peptide Copyright 2003 limsoon wong
Neural Network Approach: Target. P Emanuelsson et al. , JMB, 300: 1005 --1016, 2000 • c. TP, m. TP, SP – 4 hidden units – feedforward NNs – input windows: • 55 aa (c. TP), 35 aa (m. TP), 27 aa (SP) • sparsely encoded • Integrating Network – 0 hidden unit – feedforward NN – input is taken from the outputs of c. TP, m. TP, SP networks over 100 aa at N-terminal c. TP: chloroplast transit peptide, m. TP: mitochondria transfer peptide, SP: signal peptide Copyright 2003 limsoon wong
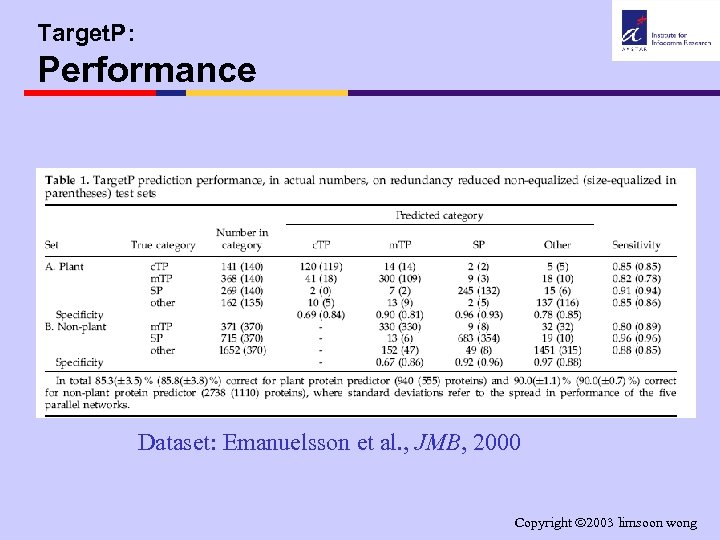 Target. P: Performance Dataset: Emanuelsson et al. , JMB, 2000 Copyright 2003 limsoon wong
Target. P: Performance Dataset: Emanuelsson et al. , JMB, 2000 Copyright 2003 limsoon wong
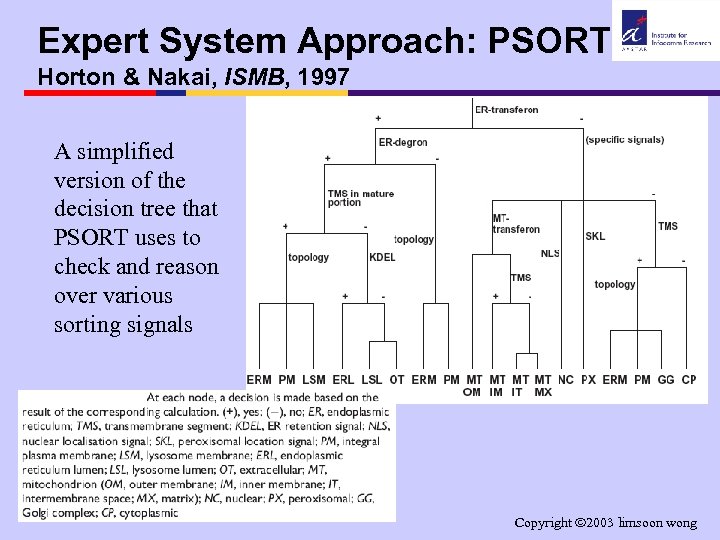 Expert System Approach: PSORT Horton & Nakai, ISMB, 1997 A simplified version of the decision tree that PSORT uses to check and reason over various sorting signals Copyright 2003 limsoon wong
Expert System Approach: PSORT Horton & Nakai, ISMB, 1997 A simplified version of the decision tree that PSORT uses to check and reason over various sorting signals Copyright 2003 limsoon wong
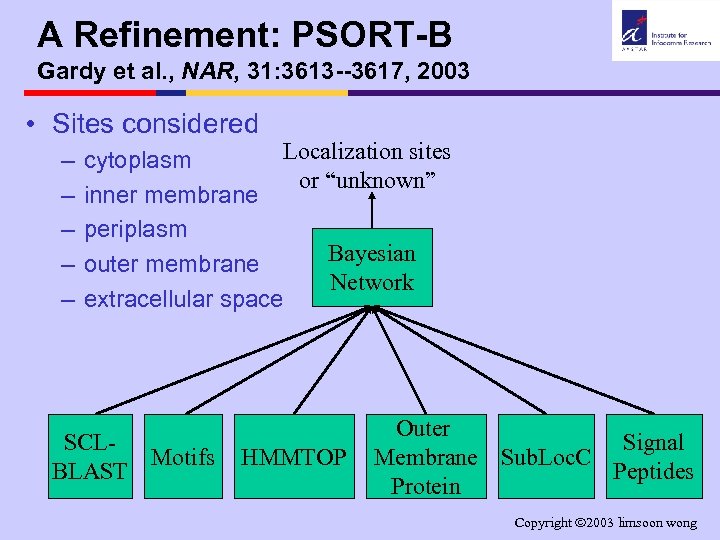 A Refinement: PSORT-B Gardy et al. , NAR, 31: 3613 --3617, 2003 • Sites considered – – – Localization sites cytoplasm or “unknown” inner membrane periplasm Bayesian outer membrane Network extracellular space SCLMotifs BLAST HMMTOP Outer Signal Membrane Sub. Loc. C Peptides Protein Copyright 2003 limsoon wong
A Refinement: PSORT-B Gardy et al. , NAR, 31: 3613 --3617, 2003 • Sites considered – – – Localization sites cytoplasm or “unknown” inner membrane periplasm Bayesian outer membrane Network extracellular space SCLMotifs BLAST HMMTOP Outer Signal Membrane Sub. Loc. C Peptides Protein Copyright 2003 limsoon wong
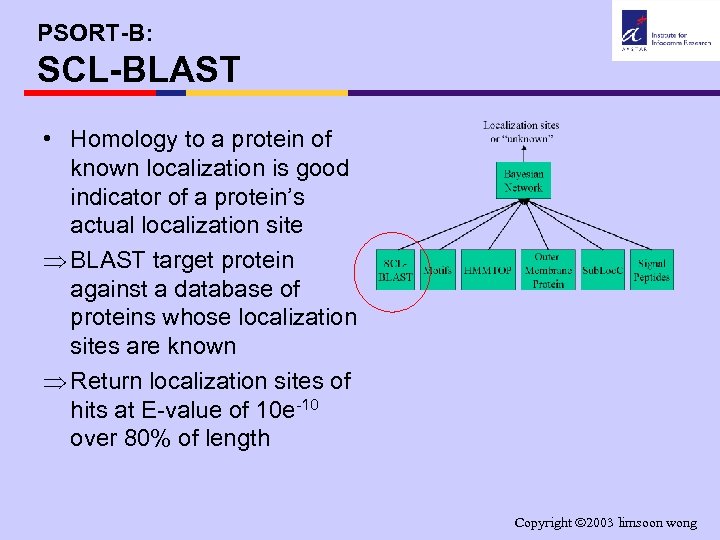 PSORT-B: SCL-BLAST • Homology to a protein of known localization is good indicator of a protein’s actual localization site Þ BLAST target protein against a database of proteins whose localization sites are known Þ Return localization sites of hits at E-value of 10 e-10 over 80% of length Copyright 2003 limsoon wong
PSORT-B: SCL-BLAST • Homology to a protein of known localization is good indicator of a protein’s actual localization site Þ BLAST target protein against a database of proteins whose localization sites are known Þ Return localization sites of hits at E-value of 10 e-10 over 80% of length Copyright 2003 limsoon wong
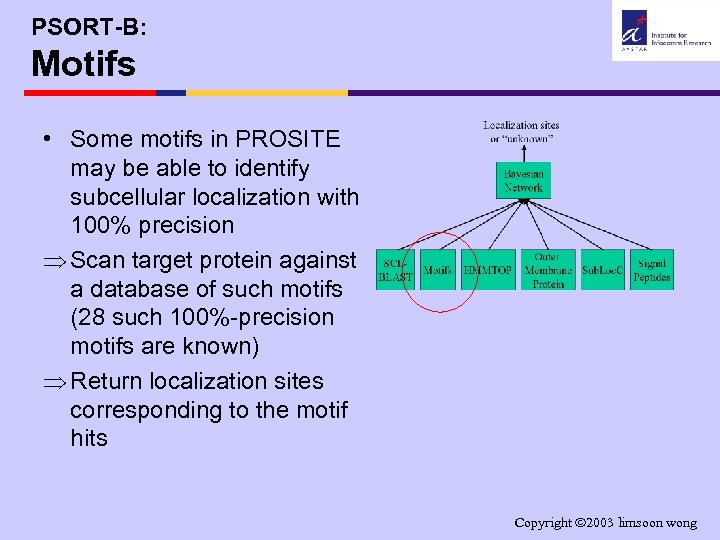 PSORT-B: Motifs • Some motifs in PROSITE may be able to identify subcellular localization with 100% precision Þ Scan target protein against a database of such motifs (28 such 100%-precision motifs are known) Þ Return localization sites corresponding to the motif hits Copyright 2003 limsoon wong
PSORT-B: Motifs • Some motifs in PROSITE may be able to identify subcellular localization with 100% precision Þ Scan target protein against a database of such motifs (28 such 100%-precision motifs are known) Þ Return localization sites corresponding to the motif hits Copyright 2003 limsoon wong
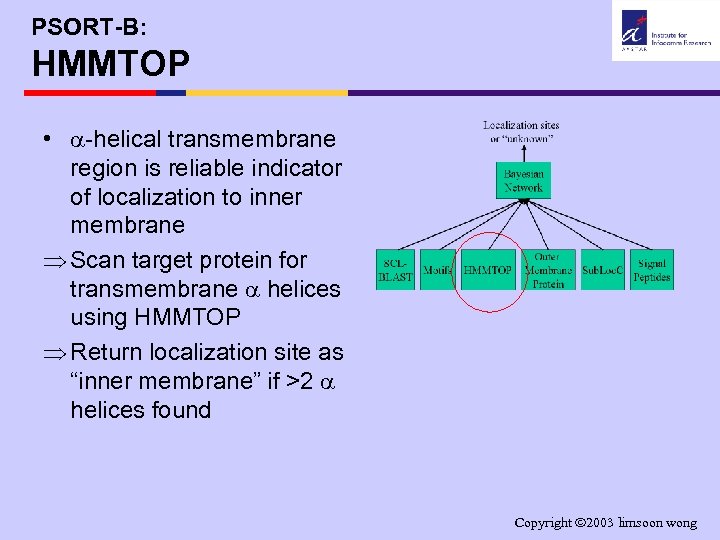 PSORT-B: HMMTOP • -helical transmembrane region is reliable indicator of localization to inner membrane Þ Scan target protein for transmembrane helices using HMMTOP Þ Return localization site as “inner membrane” if >2 helices found Copyright 2003 limsoon wong
PSORT-B: HMMTOP • -helical transmembrane region is reliable indicator of localization to inner membrane Þ Scan target protein for transmembrane helices using HMMTOP Þ Return localization site as “inner membrane” if >2 helices found Copyright 2003 limsoon wong
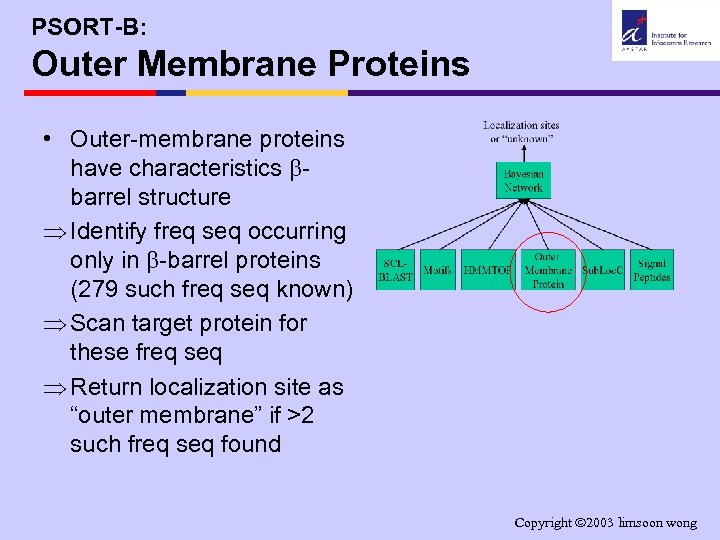 PSORT-B: Outer Membrane Proteins • Outer-membrane proteins have characteristics barrel structure Þ Identify freq seq occurring only in -barrel proteins (279 such freq seq known) Þ Scan target protein for these freq seq Þ Return localization site as “outer membrane” if >2 such freq seq found Copyright 2003 limsoon wong
PSORT-B: Outer Membrane Proteins • Outer-membrane proteins have characteristics barrel structure Þ Identify freq seq occurring only in -barrel proteins (279 such freq seq known) Þ Scan target protein for these freq seq Þ Return localization site as “outer membrane” if >2 such freq seq found Copyright 2003 limsoon wong
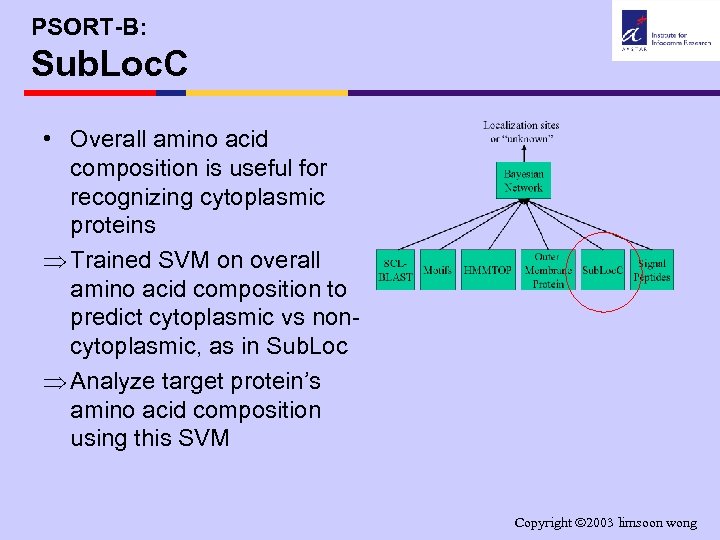 PSORT-B: Sub. Loc. C • Overall amino acid composition is useful for recognizing cytoplasmic proteins Þ Trained SVM on overall amino acid composition to predict cytoplasmic vs noncytoplasmic, as in Sub. Loc Þ Analyze target protein’s amino acid composition using this SVM Copyright 2003 limsoon wong
PSORT-B: Sub. Loc. C • Overall amino acid composition is useful for recognizing cytoplasmic proteins Þ Trained SVM on overall amino acid composition to predict cytoplasmic vs noncytoplasmic, as in Sub. Loc Þ Analyze target protein’s amino acid composition using this SVM Copyright 2003 limsoon wong
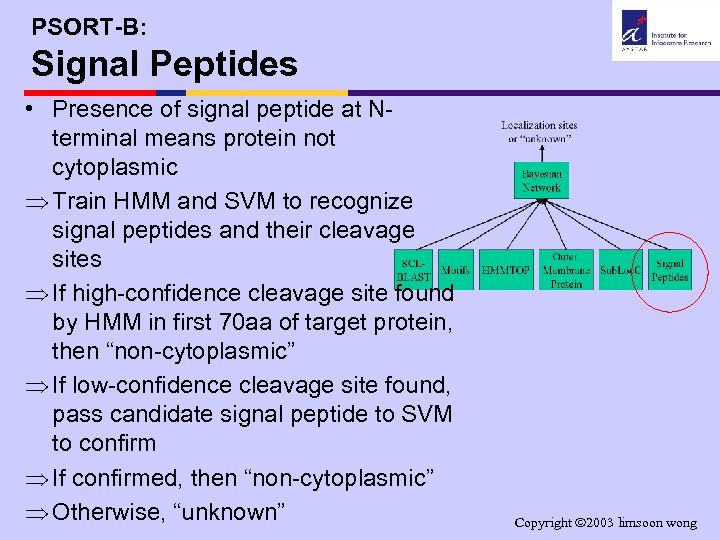 PSORT-B: Signal Peptides • Presence of signal peptide at Nterminal means protein not cytoplasmic Þ Train HMM and SVM to recognize signal peptides and their cleavage sites Þ If high-confidence cleavage site found by HMM in first 70 aa of target protein, then “non-cytoplasmic” Þ If low-confidence cleavage site found, pass candidate signal peptide to SVM to confirm Þ If confirmed, then “non-cytoplasmic” Þ Otherwise, “unknown” Copyright 2003 limsoon wong
PSORT-B: Signal Peptides • Presence of signal peptide at Nterminal means protein not cytoplasmic Þ Train HMM and SVM to recognize signal peptides and their cleavage sites Þ If high-confidence cleavage site found by HMM in first 70 aa of target protein, then “non-cytoplasmic” Þ If low-confidence cleavage site found, pass candidate signal peptide to SVM to confirm Þ If confirmed, then “non-cytoplasmic” Þ Otherwise, “unknown” Copyright 2003 limsoon wong
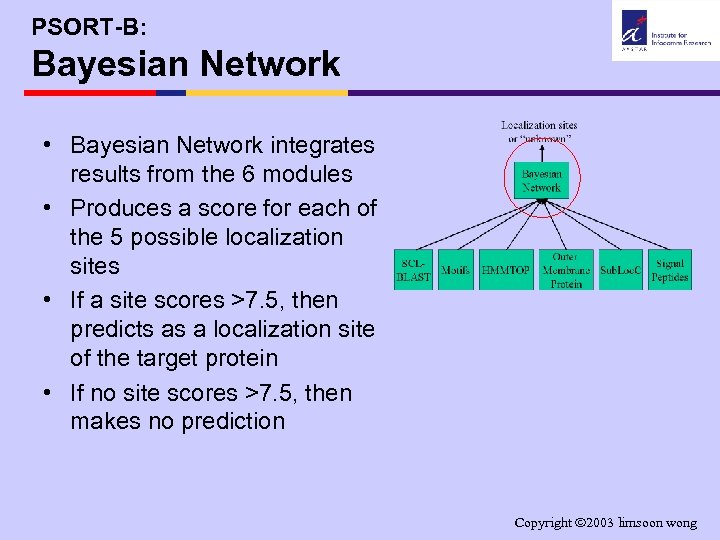 PSORT-B: Bayesian Network • Bayesian Network integrates results from the 6 modules • Produces a score for each of the 5 possible localization sites • If a site scores >7. 5, then predicts as a localization site of the target protein • If no site scores >7. 5, then makes no prediction Copyright 2003 limsoon wong
PSORT-B: Bayesian Network • Bayesian Network integrates results from the 6 modules • Produces a score for each of the 5 possible localization sites • If a site scores >7. 5, then predicts as a localization site of the target protein • If no site scores >7. 5, then makes no prediction Copyright 2003 limsoon wong
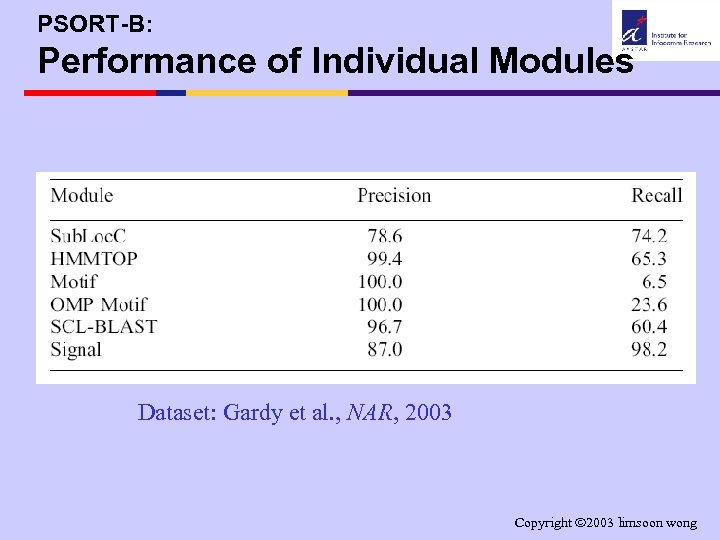 PSORT-B: Performance of Individual Modules Dataset: Gardy et al. , NAR, 2003 Copyright 2003 limsoon wong
PSORT-B: Performance of Individual Modules Dataset: Gardy et al. , NAR, 2003 Copyright 2003 limsoon wong
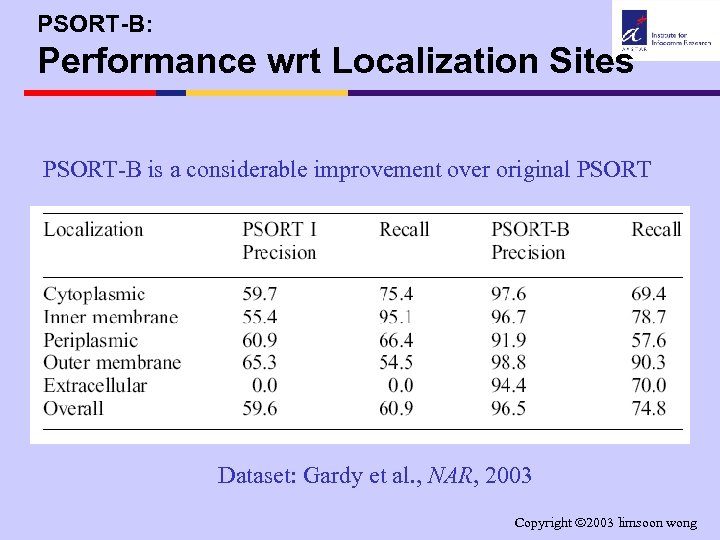 PSORT-B: Performance wrt Localization Sites PSORT-B is a considerable improvement over original PSORT Dataset: Gardy et al. , NAR, 2003 Copyright 2003 limsoon wong
PSORT-B: Performance wrt Localization Sites PSORT-B is a considerable improvement over original PSORT Dataset: Gardy et al. , NAR, 2003 Copyright 2003 limsoon wong
 PSORT vs PSORT-B: Some Remarks • PSORT considers various signal/features in a top-down way driven by its reasoning tree • PSORT-B generates all signal/features in a bottom-up way, then integrate them for decision making using Bayesian Network • Machine learning “beats” human expert? Probably the number of features/rules needed is too much/complicated Copyright 2003 limsoon wong
PSORT vs PSORT-B: Some Remarks • PSORT considers various signal/features in a top-down way driven by its reasoning tree • PSORT-B generates all signal/features in a bottom-up way, then integrate them for decision making using Bayesian Network • Machine learning “beats” human expert? Probably the number of features/rules needed is too much/complicated Copyright 2003 limsoon wong
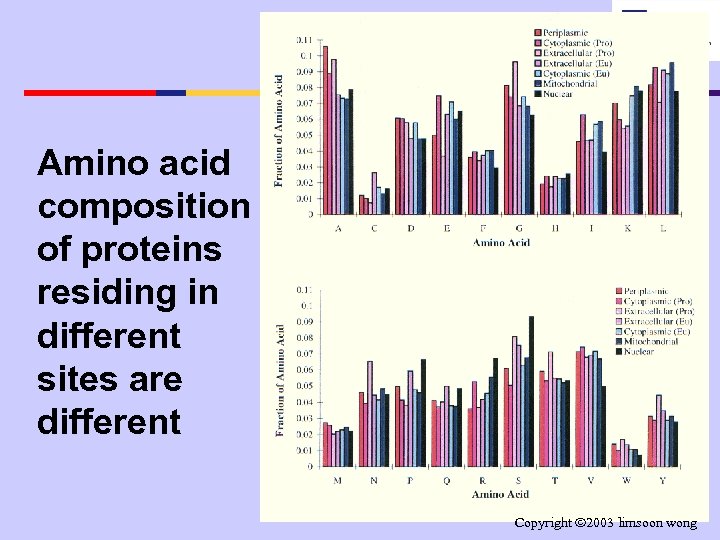 Amino acid composition of proteins residing in different sites are different Copyright 2003 limsoon wong
Amino acid composition of proteins residing in different sites are different Copyright 2003 limsoon wong
 Amino Acid Composition Differences • each cellular location • If the above is true, has own characteristic the amino acid physio-chemical composition environment differences wrt cellular location sites • proteins in each should be more location have adapted pronounced on thru evolution to that protein surfaces than environment protein interior • thus reflected in the • Exercise: Why? protein structure and amino acid composition Copyright 2003 limsoon wong
Amino Acid Composition Differences • each cellular location • If the above is true, has own characteristic the amino acid physio-chemical composition environment differences wrt cellular location sites • proteins in each should be more location have adapted pronounced on thru evolution to that protein surfaces than environment protein interior • thus reflected in the • Exercise: Why? protein structure and amino acid composition Copyright 2003 limsoon wong
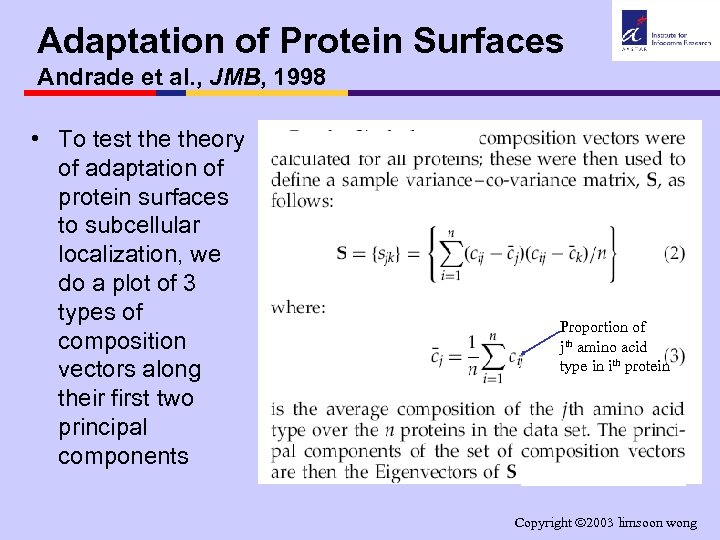 Adaptation of Protein Surfaces Andrade et al. , JMB, 1998 • To test theory of adaptation of protein surfaces to subcellular localization, we do a plot of 3 types of composition vectors along their first two principal components Proportion of jth amino acid type in ith protein Copyright 2003 limsoon wong
Adaptation of Protein Surfaces Andrade et al. , JMB, 1998 • To test theory of adaptation of protein surfaces to subcellular localization, we do a plot of 3 types of composition vectors along their first two principal components Proportion of jth amino acid type in ith protein Copyright 2003 limsoon wong
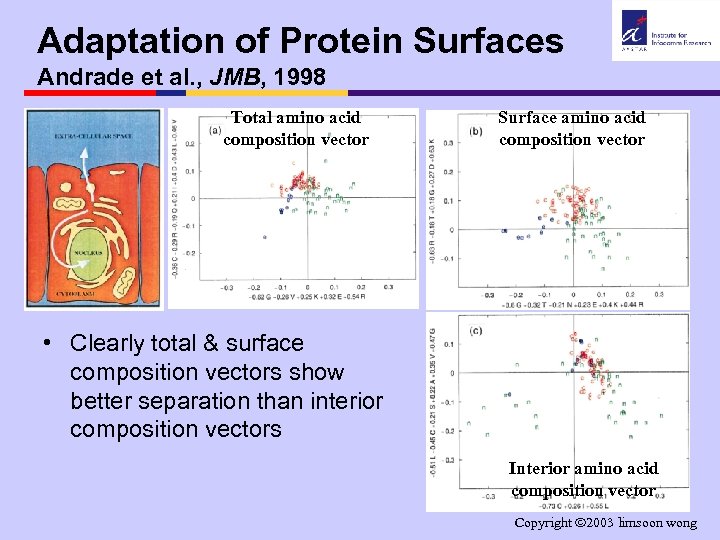 Adaptation of Protein Surfaces Andrade et al. , JMB, 1998 Total amino acid composition vector Surface amino acid composition vector • Clearly total & surface composition vectors show better separation than interior composition vectors Interior amino acid composition vector Copyright 2003 limsoon wong
Adaptation of Protein Surfaces Andrade et al. , JMB, 1998 Total amino acid composition vector Surface amino acid composition vector • Clearly total & surface composition vectors show better separation than interior composition vectors Interior amino acid composition vector Copyright 2003 limsoon wong
 Amino Acid Composition • This means can use amino acid composition vectors, especially those from protein surfaces, to predict subcellular localization! • Let’s see how this turn out…. Copyright 2003 limsoon wong
Amino Acid Composition • This means can use amino acid composition vectors, especially those from protein surfaces, to predict subcellular localization! • Let’s see how this turn out…. Copyright 2003 limsoon wong
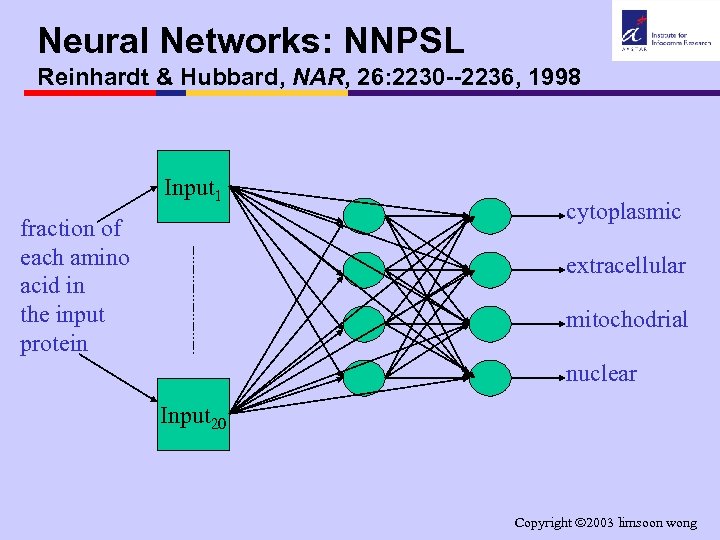 Neural Networks: NNPSL Reinhardt & Hubbard, NAR, 26: 2230 --2236, 1998 Input 1 fraction of each amino acid in the input protein cytoplasmic extracellular mitochodrial nuclear Input 20 Copyright 2003 limsoon wong
Neural Networks: NNPSL Reinhardt & Hubbard, NAR, 26: 2230 --2236, 1998 Input 1 fraction of each amino acid in the input protein cytoplasmic extracellular mitochodrial nuclear Input 20 Copyright 2003 limsoon wong
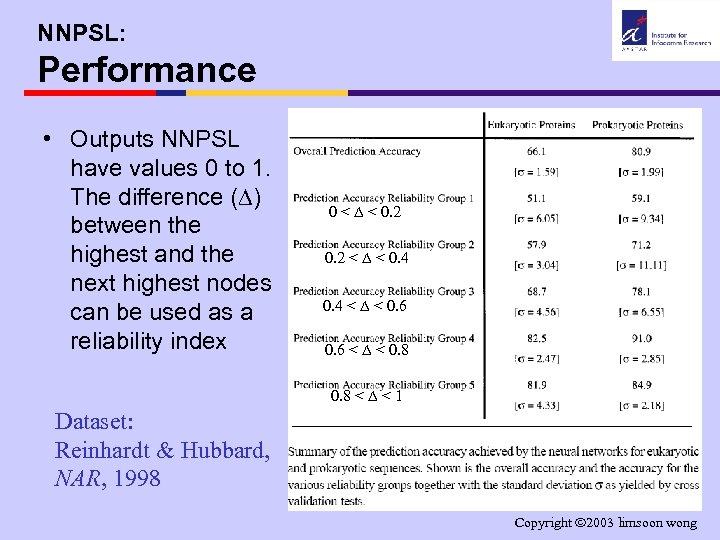 NNPSL: Performance • Outputs NNPSL have values 0 to 1. The difference ( ) between the highest and the next highest nodes can be used as a reliability index 0 < < 0. 2 < < 0. 4 < < 0. 6 < < 0. 8 < < 1 Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
NNPSL: Performance • Outputs NNPSL have values 0 to 1. The difference ( ) between the highest and the next highest nodes can be used as a reliability index 0 < < 0. 2 < < 0. 4 < < 0. 6 < < 0. 8 < < 1 Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
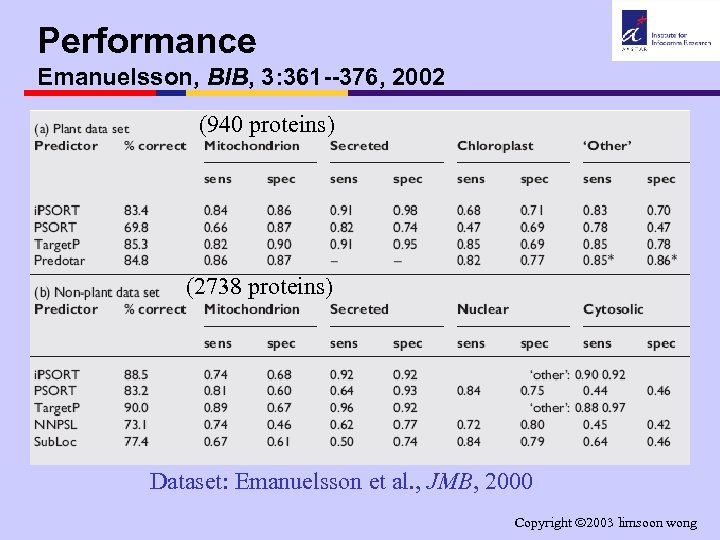 Performance Emanuelsson, BIB, 3: 361 --376, 2002 (940 proteins) (2738 proteins) Dataset: Emanuelsson et al. , JMB, 2000 Copyright 2003 limsoon wong
Performance Emanuelsson, BIB, 3: 361 --376, 2002 (940 proteins) (2738 proteins) Dataset: Emanuelsson et al. , JMB, 2000 Copyright 2003 limsoon wong
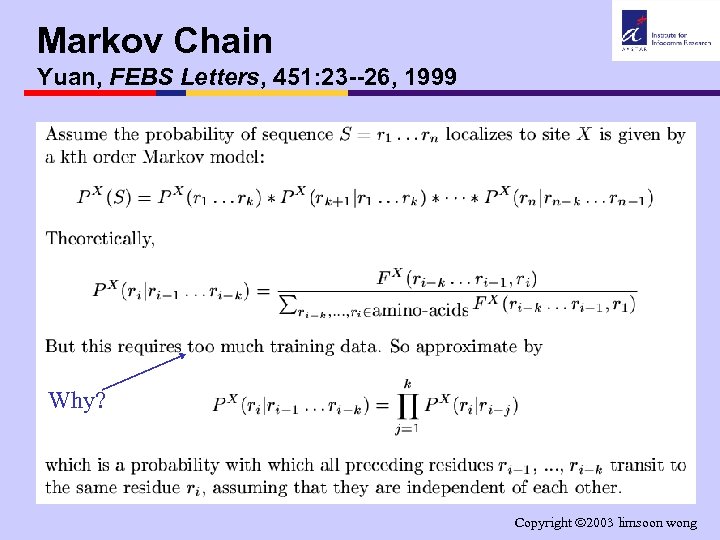 Markov Chain Yuan, FEBS Letters, 451: 23 --26, 1999 Why? Copyright 2003 limsoon wong
Markov Chain Yuan, FEBS Letters, 451: 23 --26, 1999 Why? Copyright 2003 limsoon wong
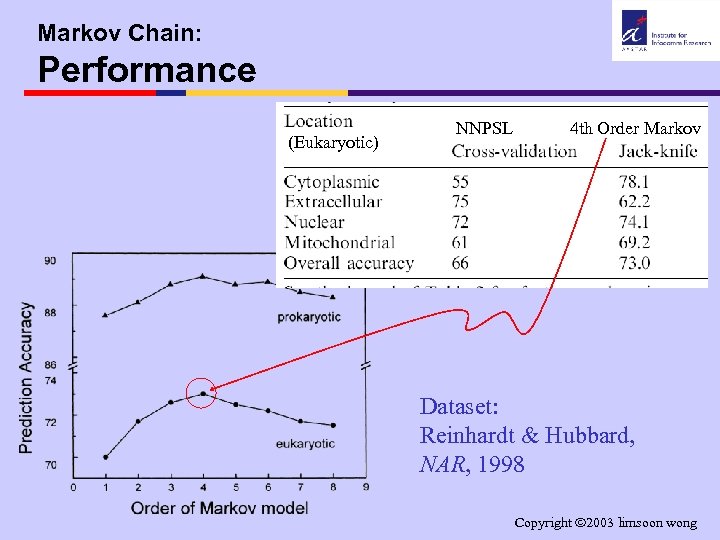 Markov Chain: Performance (Eukaryotic) NNPSL 4 th Order Markov Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
Markov Chain: Performance (Eukaryotic) NNPSL 4 th Order Markov Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
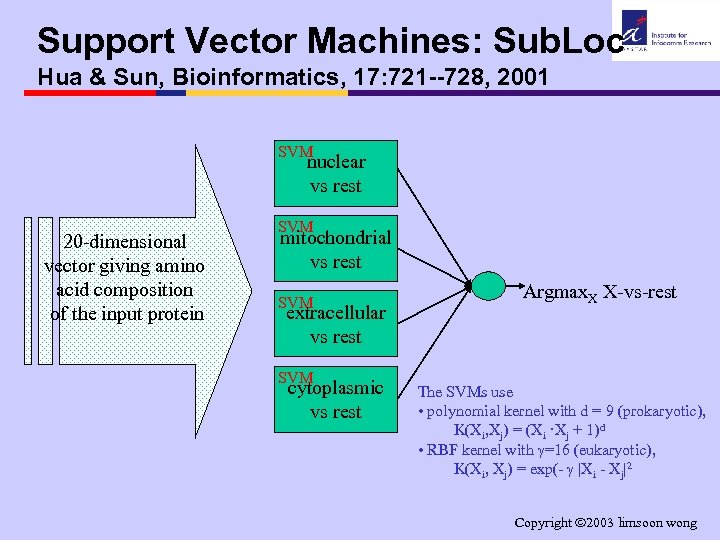 Support Vector Machines: Sub. Loc Hua & Sun, Bioinformatics, 17: 721 --728, 2001 SVM nuclear vs rest 20 -dimensional vector giving amino acid composition of the input protein SVM mitochondrial vs rest SVM extracellular vs rest SVM cytoplasmic vs rest Argmax. X X-vs-rest The SVMs use • polynomial kernel with d = 9 (prokaryotic), K(Xi, Xj) = (Xi ·Xj + 1)d • RBF kernel with =16 (eukaryotic), K(Xi, Xj) = exp(- |Xi - Xj|2 Copyright 2003 limsoon wong
Support Vector Machines: Sub. Loc Hua & Sun, Bioinformatics, 17: 721 --728, 2001 SVM nuclear vs rest 20 -dimensional vector giving amino acid composition of the input protein SVM mitochondrial vs rest SVM extracellular vs rest SVM cytoplasmic vs rest Argmax. X X-vs-rest The SVMs use • polynomial kernel with d = 9 (prokaryotic), K(Xi, Xj) = (Xi ·Xj + 1)d • RBF kernel with =16 (eukaryotic), K(Xi, Xj) = exp(- |Xi - Xj|2 Copyright 2003 limsoon wong
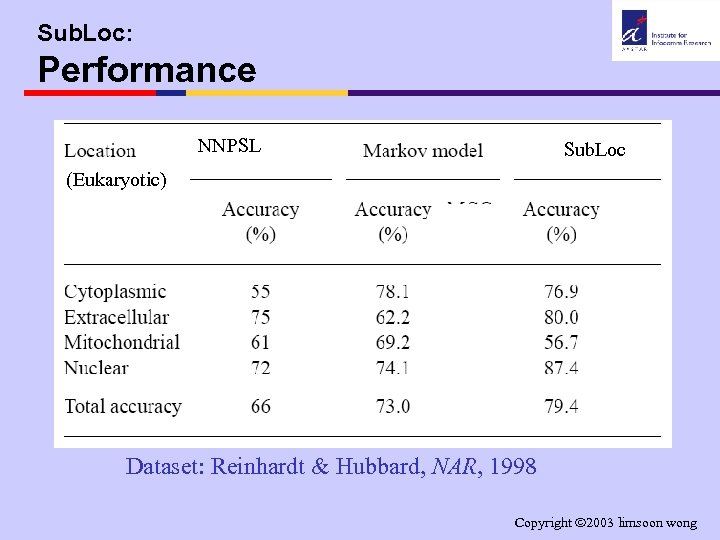 Sub. Loc: Performance NNPSL Sub. Loc (Eukaryotic) Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
Sub. Loc: Performance NNPSL Sub. Loc (Eukaryotic) Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
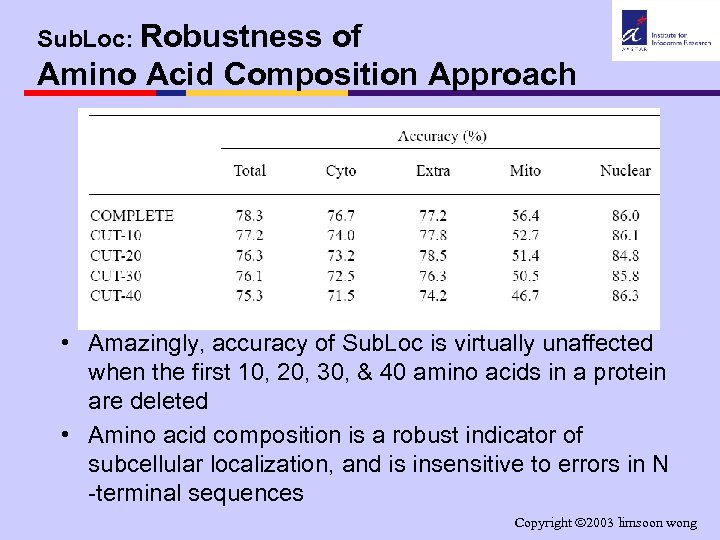 Sub. Loc: Robustness of Amino Acid Composition Approach • Amazingly, accuracy of Sub. Loc is virtually unaffected when the first 10, 20, 30, & 40 amino acids in a protein are deleted • Amino acid composition is a robust indicator of subcellular localization, and is insensitive to errors in N -terminal sequences Copyright 2003 limsoon wong
Sub. Loc: Robustness of Amino Acid Composition Approach • Amazingly, accuracy of Sub. Loc is virtually unaffected when the first 10, 20, 30, & 40 amino acids in a protein are deleted • Amino acid composition is a robust indicator of subcellular localization, and is insensitive to errors in N -terminal sequences Copyright 2003 limsoon wong
 Amino Acid Composition: Taking it Further • How about pairs of consecutive amino acids? (a. k. a 2 -grams) How about 3 grams, …, k-grams? • How about pseudo amino acid composition? • How about presence of entire functional domains? (I. e. think of the presence/absence of a functional domain as a summary of amino acid sequence info. . . ) Copyright 2003 limsoon wong
Amino Acid Composition: Taking it Further • How about pairs of consecutive amino acids? (a. k. a 2 -grams) How about 3 grams, …, k-grams? • How about pseudo amino acid composition? • How about presence of entire functional domains? (I. e. think of the presence/absence of a functional domain as a summary of amino acid sequence info. . . ) Copyright 2003 limsoon wong
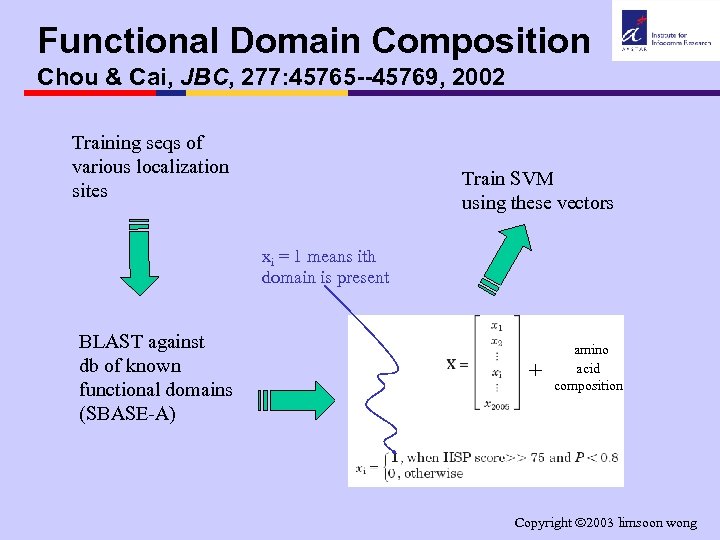 Functional Domain Composition Chou & Cai, JBC, 277: 45765 --45769, 2002 Training seqs of various localization sites Train SVM using these vectors xi = 1 means ith domain is present BLAST against db of known functional domains (SBASE-A) + amino acid composition Copyright 2003 limsoon wong
Functional Domain Composition Chou & Cai, JBC, 277: 45765 --45769, 2002 Training seqs of various localization sites Train SVM using these vectors xi = 1 means ith domain is present BLAST against db of known functional domains (SBASE-A) + amino acid composition Copyright 2003 limsoon wong
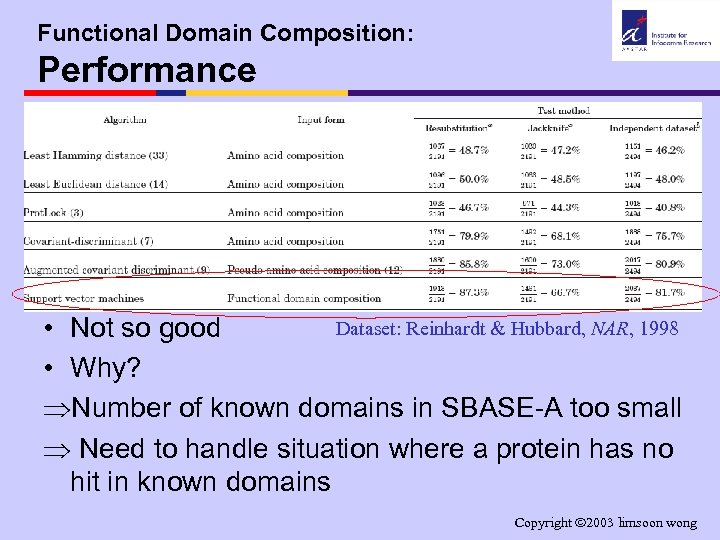 Functional Domain Composition: Performance Dataset: Reinhardt & Hubbard, NAR, 1998 • Not so good • Why? ÞNumber of known domains in SBASE-A too small Þ Need to handle situation where a protein has no hit in known domains Copyright 2003 limsoon wong
Functional Domain Composition: Performance Dataset: Reinhardt & Hubbard, NAR, 1998 • Not so good • Why? ÞNumber of known domains in SBASE-A too small Þ Need to handle situation where a protein has no hit in known domains Copyright 2003 limsoon wong
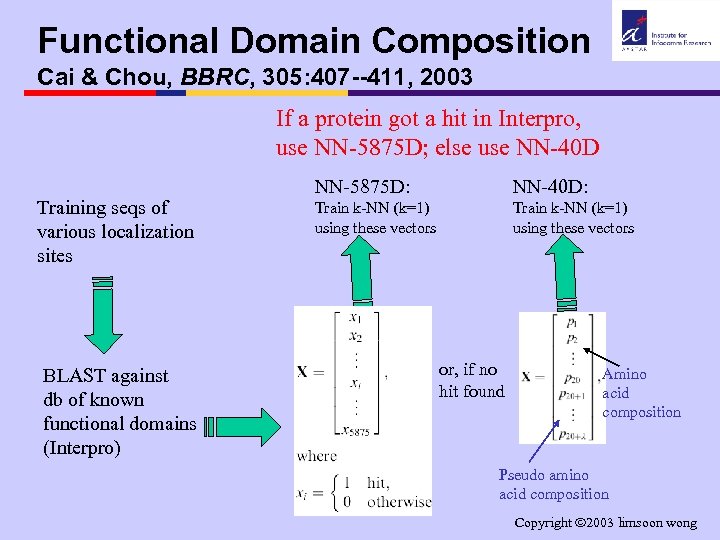 Functional Domain Composition Cai & Chou, BBRC, 305: 407 --411, 2003 If a protein got a hit in Interpro, use NN-5875 D; else use NN-40 D Training seqs of various localization sites BLAST against db of known functional domains (Interpro) NN-5875 D: NN-40 D: Train k-NN (k=1) using these vectors or, if no hit found Amino acid composition Pseudo amino acid composition Copyright 2003 limsoon wong
Functional Domain Composition Cai & Chou, BBRC, 305: 407 --411, 2003 If a protein got a hit in Interpro, use NN-5875 D; else use NN-40 D Training seqs of various localization sites BLAST against db of known functional domains (Interpro) NN-5875 D: NN-40 D: Train k-NN (k=1) using these vectors or, if no hit found Amino acid composition Pseudo amino acid composition Copyright 2003 limsoon wong
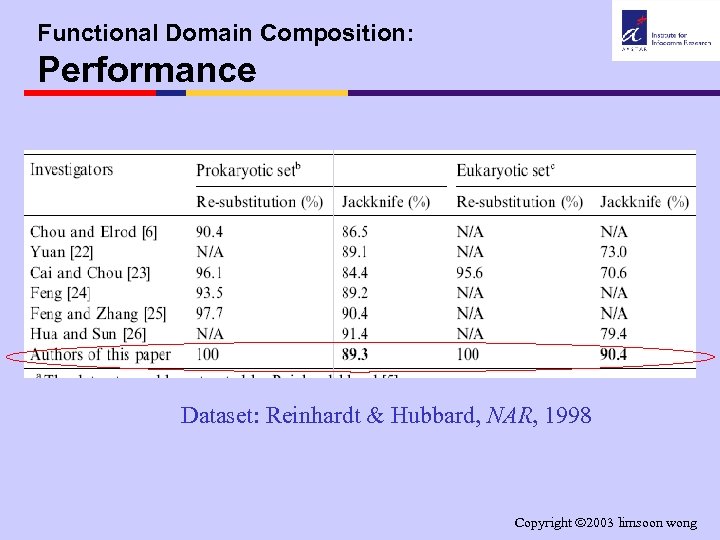 Functional Domain Composition: Performance Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
Functional Domain Composition: Performance Dataset: Reinhardt & Hubbard, NAR, 1998 Copyright 2003 limsoon wong
 Notes Copyright 2003 limsoon wong
Notes Copyright 2003 limsoon wong
 References (Transmembrane) • Wiess et al. “Transmembrane segment prediction from protein sequence data”, ISMB, 420 --421, 1993 • Gavel et al. “The positive-inside rule applies to thylakoid membrane proteins”, FEBS 282: 41 --46, 1991 • Monne et al. “A turn propensity scale for transmembrane helices”, JMB, 288: 141 --145, 1999 • Sonnhammer et al. “A hidden Markov model for predicting transmembrane helices in protein sequences”, ISMB, 6: 175 --182, 1998 • Martelli et al. “An ENSEMBLE machine learning approach for the prediction of all-alpha membrane proteins”, Bioinformatics, 19(suppl): i 205 --i 211, 2003 Copyright 2003 limsoon wong
References (Transmembrane) • Wiess et al. “Transmembrane segment prediction from protein sequence data”, ISMB, 420 --421, 1993 • Gavel et al. “The positive-inside rule applies to thylakoid membrane proteins”, FEBS 282: 41 --46, 1991 • Monne et al. “A turn propensity scale for transmembrane helices”, JMB, 288: 141 --145, 1999 • Sonnhammer et al. “A hidden Markov model for predicting transmembrane helices in protein sequences”, ISMB, 6: 175 --182, 1998 • Martelli et al. “An ENSEMBLE machine learning approach for the prediction of all-alpha membrane proteins”, Bioinformatics, 19(suppl): i 205 --i 211, 2003 Copyright 2003 limsoon wong
 References (Transmembrane) • Von Heijne. “Membrane protein structure prediction”, JMB, 225: 487 --494, 1992 • Jacoboni et al. “Prediction of the transmembrane regions of beta-barrel membrane proteins with a neural network-based predictor”, Protein Sci. , 10: 779 --787, 2001 • Martelli et al. “a sequence-profile-based HMM for predicting and discriminating beta barrel membrane proteins”, Bioinformatics, 18: S 46 --S 53, 2002 • Moller et al. “Evaluation of methods for the prediction of membrane spanning regions”, Bioinformatics, 17: 646 --653, 2001 • Fariselli et al. “Max. Sub. Seq: an algorithm for segmentlength optimization. The case study of the transmembrane spanning segments”, Bioinformatics, 19: 500 --505, 2003 Copyright 2003 limsoon wong
References (Transmembrane) • Von Heijne. “Membrane protein structure prediction”, JMB, 225: 487 --494, 1992 • Jacoboni et al. “Prediction of the transmembrane regions of beta-barrel membrane proteins with a neural network-based predictor”, Protein Sci. , 10: 779 --787, 2001 • Martelli et al. “a sequence-profile-based HMM for predicting and discriminating beta barrel membrane proteins”, Bioinformatics, 18: S 46 --S 53, 2002 • Moller et al. “Evaluation of methods for the prediction of membrane spanning regions”, Bioinformatics, 17: 646 --653, 2001 • Fariselli et al. “Max. Sub. Seq: an algorithm for segmentlength optimization. The case study of the transmembrane spanning segments”, Bioinformatics, 19: 500 --505, 2003 Copyright 2003 limsoon wong
 References (Transmembrane) • Rost et al. “Transmembrane helices predicted at 95% accuracy”, Protein Sci. , 4: 521 --533, 1995 • Krogh et al. “Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes”, JMB, 305: 567 --580, 2001 • Andersson et al. “Different positively charged amino acids have similar effectson the topology of a polytopic transmembrane protein in E. coli”, JBC, 267: 1491 --1495, 1992 Copyright 2003 limsoon wong
References (Transmembrane) • Rost et al. “Transmembrane helices predicted at 95% accuracy”, Protein Sci. , 4: 521 --533, 1995 • Krogh et al. “Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes”, JMB, 305: 567 --580, 2001 • Andersson et al. “Different positively charged amino acids have similar effectson the topology of a polytopic transmembrane protein in E. coli”, JBC, 267: 1491 --1495, 1992 Copyright 2003 limsoon wong
 References (Subcellular Localization) • Horton & Nakai, “Better prediction of protein cellular localization sites with the k-nearest neighbours classifier”, ISMB, 5: 147 --152, 1997 • Gardy et al. , “PSORT-B: Improving protein subcellular localization for Gram-negative bacteria”, NAR, 31: 3613 --3617, 2003 • Emanuelsson, “Predicting protein subcellular localization from amino acid sequence information”, BIB, 3: 361 --376, 2002 • Andrade et al. , “Adaptation of protein surfaces to subcellular location”, JMB, 276: 517 --525, 1998 • Yuan, “Prediction of protein subcellular locations using Markov chain models”, FEBS Letters, 451: 23 --26, 1999 Copyright 2003 limsoon wong
References (Subcellular Localization) • Horton & Nakai, “Better prediction of protein cellular localization sites with the k-nearest neighbours classifier”, ISMB, 5: 147 --152, 1997 • Gardy et al. , “PSORT-B: Improving protein subcellular localization for Gram-negative bacteria”, NAR, 31: 3613 --3617, 2003 • Emanuelsson, “Predicting protein subcellular localization from amino acid sequence information”, BIB, 3: 361 --376, 2002 • Andrade et al. , “Adaptation of protein surfaces to subcellular location”, JMB, 276: 517 --525, 1998 • Yuan, “Prediction of protein subcellular locations using Markov chain models”, FEBS Letters, 451: 23 --26, 1999 Copyright 2003 limsoon wong
 References (Subcellular Localization) • Emanuelsson et al. , “Chloro. P, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites”, Protein Sci. , 8: 978 --984, 1999 • Emanuelsson et al. , "Predicting subcellular localization of proteins based on their N-terminal amino acid sequence", JMB, 300: 1005 -1016, 2000 • Hua & Sun, “Support vector machine approach for protein subcellular localization prediction”, Bioinformatics, 17: 721 --728, 2001 • Reinhardt & Hubbard, “Using neural networks for prediction of the subcellular location of proteins”, NAR, 26: 2230 --2236, 1998 Copyright 2003 limsoon wong
References (Subcellular Localization) • Emanuelsson et al. , “Chloro. P, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites”, Protein Sci. , 8: 978 --984, 1999 • Emanuelsson et al. , "Predicting subcellular localization of proteins based on their N-terminal amino acid sequence", JMB, 300: 1005 -1016, 2000 • Hua & Sun, “Support vector machine approach for protein subcellular localization prediction”, Bioinformatics, 17: 721 --728, 2001 • Reinhardt & Hubbard, “Using neural networks for prediction of the subcellular location of proteins”, NAR, 26: 2230 --2236, 1998 Copyright 2003 limsoon wong
 References (Subcellular Localization) • Cai & Chou, “Nearest neighbour algorithm for predicting protein subcellular location by combining functional domain composition and pseudo-amino acid composition”, BBRC, 305: 407 --411, 2003 • Chou & Cai, “Using functional domain composition and support vector machines for prediction of protein subcellular location”, JBC, 277: 45765 --45769, 2002 • Park & Kanehisa, “Prediction of protein subcellular locations by support vector machines using compositions of amino acids and amino acid pairs”, Bioinformatics, 19: 1656 --1663, 2003 Copyright 2003 limsoon wong
References (Subcellular Localization) • Cai & Chou, “Nearest neighbour algorithm for predicting protein subcellular location by combining functional domain composition and pseudo-amino acid composition”, BBRC, 305: 407 --411, 2003 • Chou & Cai, “Using functional domain composition and support vector machines for prediction of protein subcellular location”, JBC, 277: 45765 --45769, 2002 • Park & Kanehisa, “Prediction of protein subcellular locations by support vector machines using compositions of amino acids and amino acid pairs”, Bioinformatics, 19: 1656 --1663, 2003 Copyright 2003 limsoon wong


