f044f0d9b7469d5946ed03992391ec66.ppt
- Количество слайдов: 56
 Reaction Networks with Fluctuations: From Inter-stellar Chemistry to Intra-cellular Biology Ofer Biham The Hebrew University Azi Lipshtat Baruch Barzel Adina Lederhendler Adiel Loinger Hagai Perets Hilik Frank Amir Zait Nadav Katz Gian Vidali Eric Herbst Raz Kupferman Nathalie Balaban Joachim Krug Evelyne Roueff Jacques Le Bourlot Franck Le Petit Israel Science Foundation The Adler Foundation for Space Research The Center for Complexity Science US-Israel Binational Science Foundation
Reaction Networks with Fluctuations: From Inter-stellar Chemistry to Intra-cellular Biology Ofer Biham The Hebrew University Azi Lipshtat Baruch Barzel Adina Lederhendler Adiel Loinger Hagai Perets Hilik Frank Amir Zait Nadav Katz Gian Vidali Eric Herbst Raz Kupferman Nathalie Balaban Joachim Krug Evelyne Roueff Jacques Le Bourlot Franck Le Petit Israel Science Foundation The Adler Foundation for Space Research The Center for Complexity Science US-Israel Binational Science Foundation
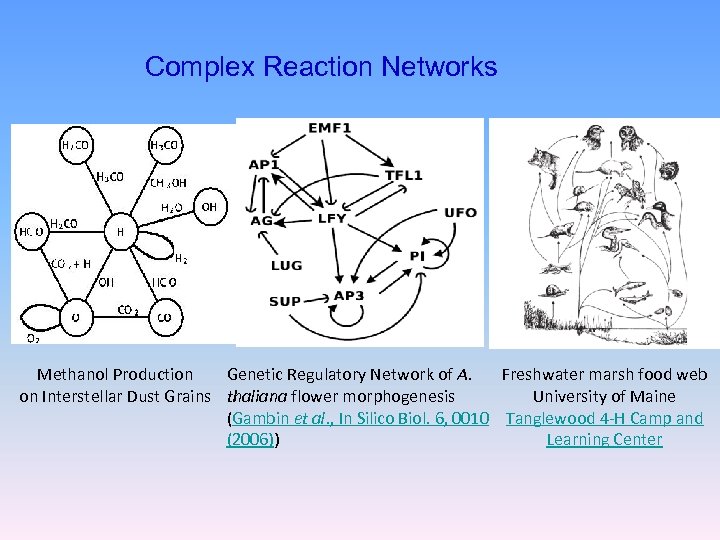 Complex Reaction Networks Methanol Production Genetic Regulatory Network of A. Freshwater marsh food web on Interstellar Dust Grains thaliana flower morphogenesis University of Maine (Gambin et al. , In Silico Biol. 6, 0010 Tanglewood 4 -H Camp and (2006)) Learning Center
Complex Reaction Networks Methanol Production Genetic Regulatory Network of A. Freshwater marsh food web on Interstellar Dust Grains thaliana flower morphogenesis University of Maine (Gambin et al. , In Silico Biol. 6, 0010 Tanglewood 4 -H Camp and (2006)) Learning Center
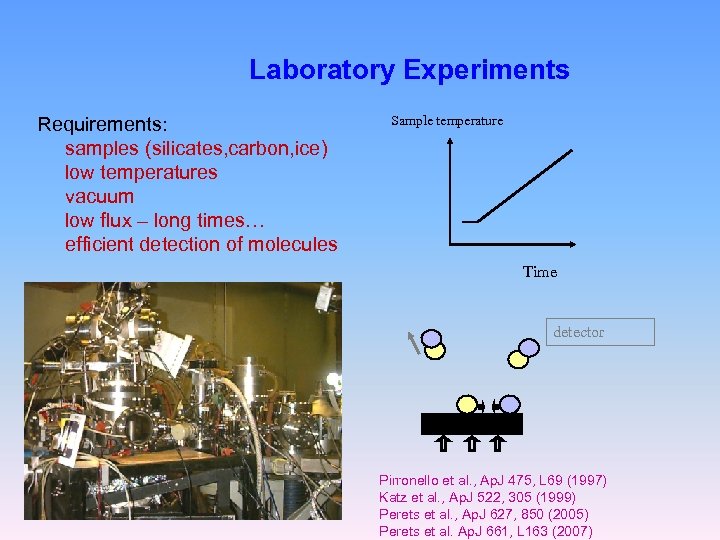 Laboratory Experiments Requirements: samples (silicates, carbon, ice) low temperatures vacuum low flux – long times… efficient detection of molecules Sample temperature Time detector Pirronello et al. , Ap. J 475, L 69 (1997) Katz et al. , Ap. J 522, 305 (1999) Perets et al. , Ap. J 627, 850 (2005) Perets et al. Ap. J 661, L 163 (2007)
Laboratory Experiments Requirements: samples (silicates, carbon, ice) low temperatures vacuum low flux – long times… efficient detection of molecules Sample temperature Time detector Pirronello et al. , Ap. J 475, L 69 (1997) Katz et al. , Ap. J 522, 305 (1999) Perets et al. , Ap. J 627, 850 (2005) Perets et al. Ap. J 661, L 163 (2007)
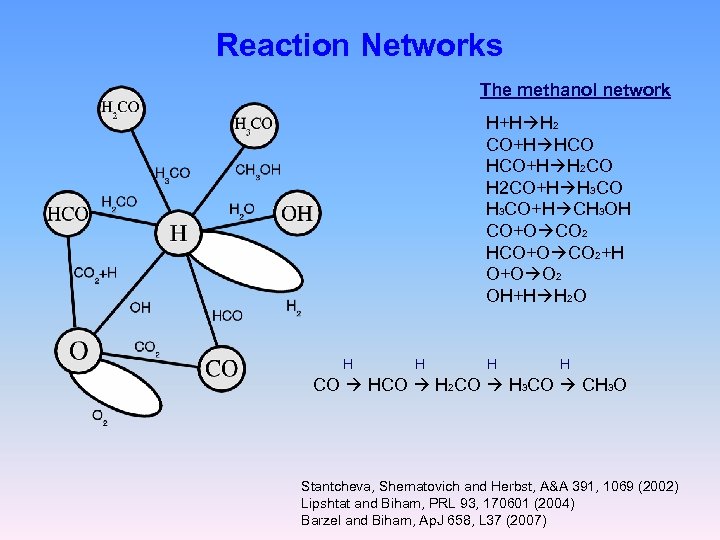 Reaction Networks The methanol network H+H H 2 CO+H HCO+H H 2 CO+H H 3 CO+H CH 3 OH CO+O CO 2 HCO+O CO 2+H O+O O 2 OH+H H 2 O H H CO H 2 CO H 3 CO CH 3 O Stantcheva, Shematovich and Herbst, A&A 391, 1069 (2002) Lipshtat and Biham, PRL 93, 170601 (2004) Barzel and Biham, Ap. J 658, L 37 (2007)
Reaction Networks The methanol network H+H H 2 CO+H HCO+H H 2 CO+H H 3 CO+H CH 3 OH CO+O CO 2 HCO+O CO 2+H O+O O 2 OH+H H 2 O H H CO H 2 CO H 3 CO CH 3 O Stantcheva, Shematovich and Herbst, A&A 391, 1069 (2002) Lipshtat and Biham, PRL 93, 170601 (2004) Barzel and Biham, Ap. J 658, L 37 (2007)
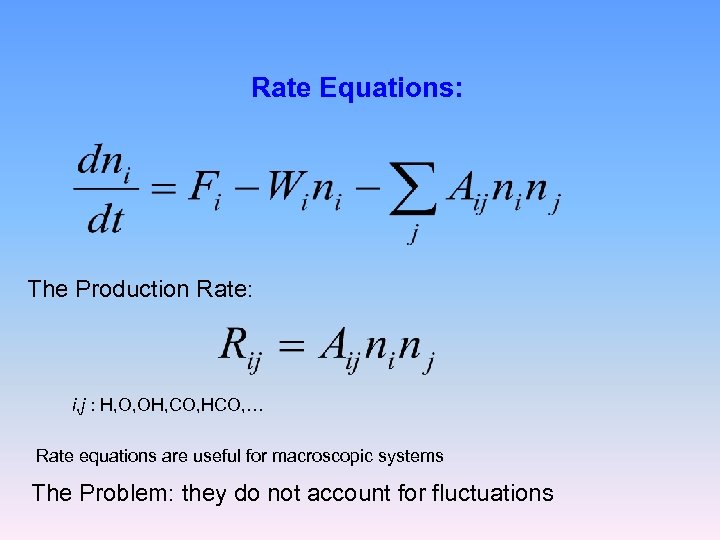 Rate Equations: The Production Rate: i, j : H, O, OH, CO, HCO, … Rate equations are useful for macroscopic systems The Problem: they do not account for fluctuations
Rate Equations: The Production Rate: i, j : H, O, OH, CO, HCO, … Rate equations are useful for macroscopic systems The Problem: they do not account for fluctuations
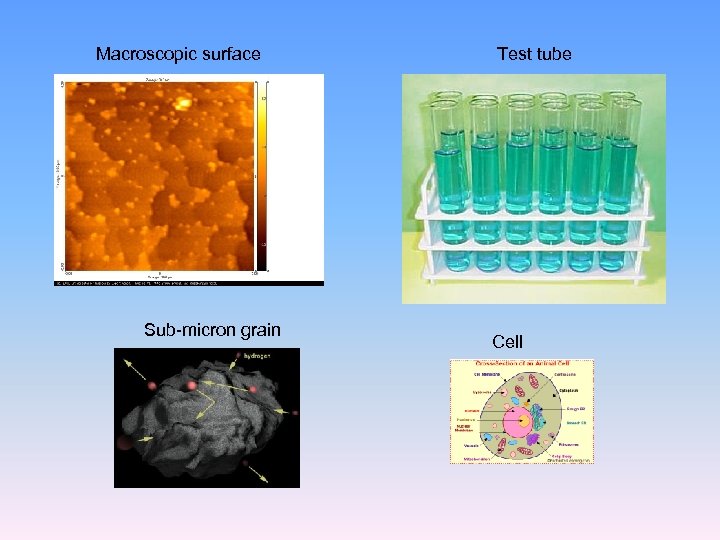 Macroscopic surface Sub-micron grain Test tube Cell
Macroscopic surface Sub-micron grain Test tube Cell
 Rate Equation Master Equation For small grains under low flux the typical population size of H atoms on a grain may go down to n~1 Thus: the rate equation (mean field approximation) fails One needs to take into account: The discreteness of the H atoms The fluctuations in the populations of H atoms on grains → Master Equation for the probability distribution P(n), n=0, 1, 2, 3, …
Rate Equation Master Equation For small grains under low flux the typical population size of H atoms on a grain may go down to n~1 Thus: the rate equation (mean field approximation) fails One needs to take into account: The discreteness of the H atoms The fluctuations in the populations of H atoms on grains → Master Equation for the probability distribution P(n), n=0, 1, 2, 3, …
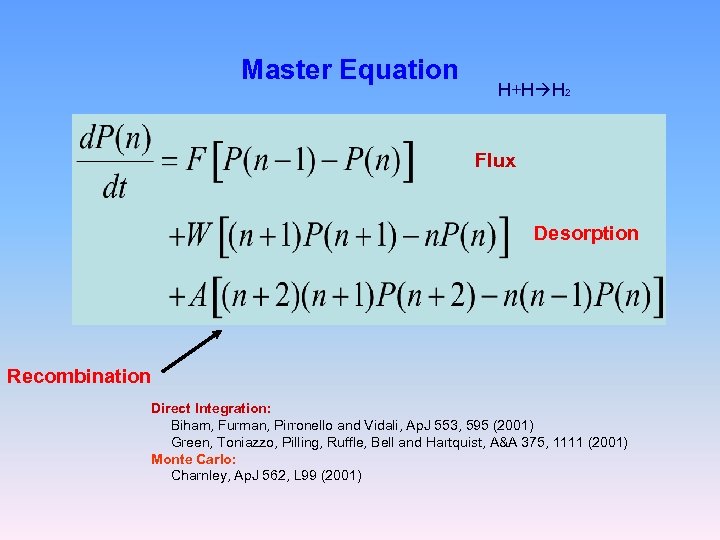 Master Equation H+H H 2 Flux Desorption Recombination Direct Integration: Biham, Furman, Pirronello and Vidali, Ap. J 553, 595 (2001) Green, Toniazzo, Pilling, Ruffle, Bell and Hartquist, A&A 375, 1111 (2001) Monte Carlo: Charnley, Ap. J 562, L 99 (2001)
Master Equation H+H H 2 Flux Desorption Recombination Direct Integration: Biham, Furman, Pirronello and Vidali, Ap. J 553, 595 (2001) Green, Toniazzo, Pilling, Ruffle, Bell and Hartquist, A&A 375, 1111 (2001) Monte Carlo: Charnley, Ap. J 562, L 99 (2001)
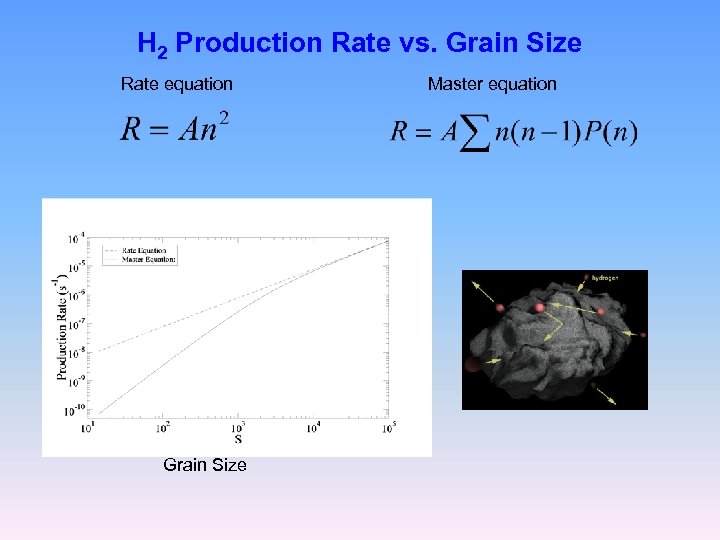 H 2 Production Rate vs. Grain Size Rate equation Grain Size Master equation
H 2 Production Rate vs. Grain Size Rate equation Grain Size Master equation
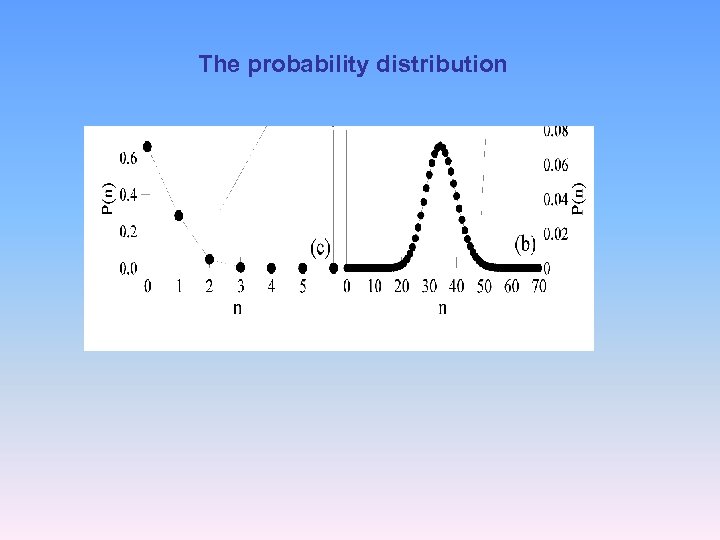 The probability distribution
The probability distribution
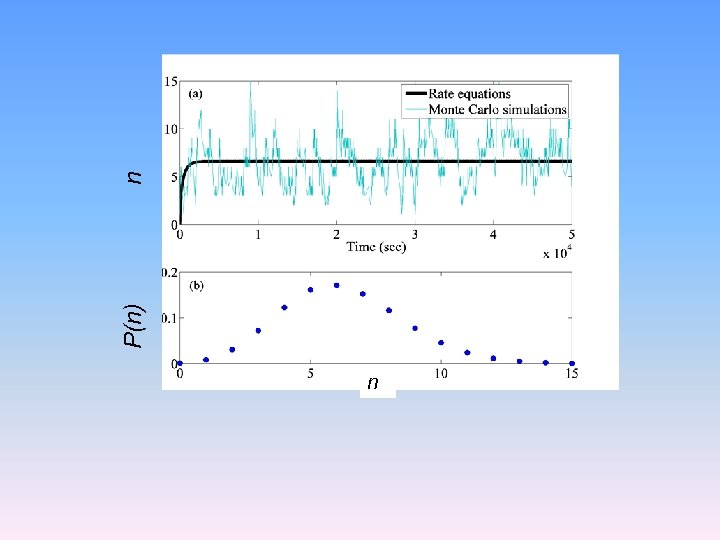 n P(n) n
n P(n) n
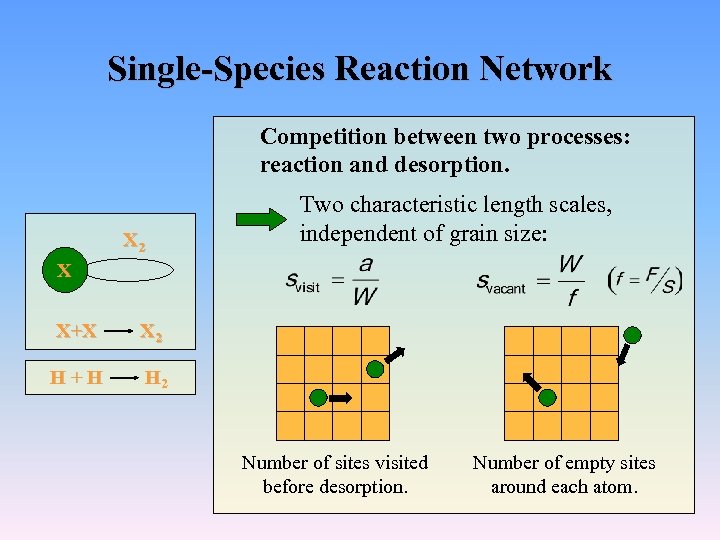 Single-Species Reaction Network Competition between two processes: reaction and desorption. X 2 Two characteristic length scales, independent of grain size: X X+X X 2 H+H H 2 Number of sites visited before desorption. Number of empty sites around each atom.
Single-Species Reaction Network Competition between two processes: reaction and desorption. X 2 Two characteristic length scales, independent of grain size: X X+X X 2 H+H H 2 Number of sites visited before desorption. Number of empty sites around each atom.
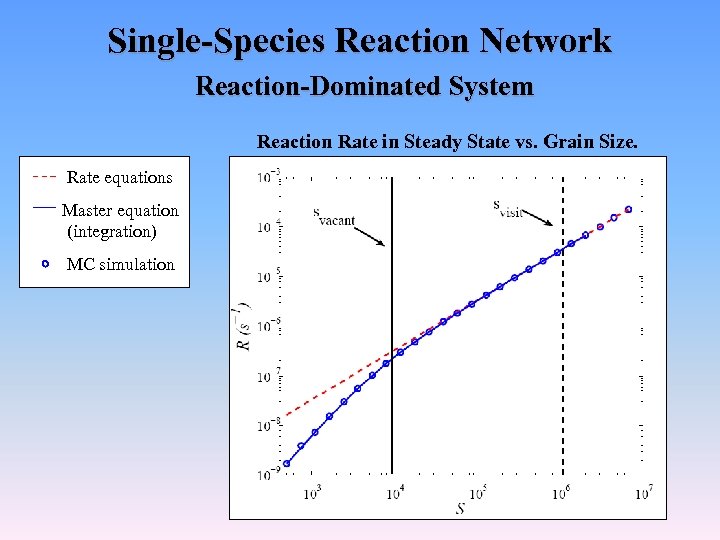 Single-Species Reaction Network Reaction-Dominated System Reaction Rate in Steady State vs. Grain Size. Rate equations Master equation (integration) MC simulation
Single-Species Reaction Network Reaction-Dominated System Reaction Rate in Steady State vs. Grain Size. Rate equations Master equation (integration) MC simulation
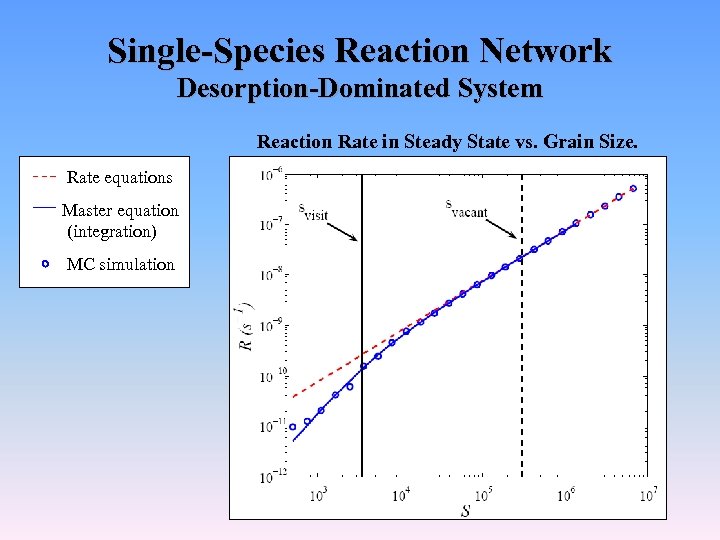 Single-Species Reaction Network Desorption-Dominated System Reaction Rate in Steady State vs. Grain Size. Rate equations Master equation (integration) MC simulation
Single-Species Reaction Network Desorption-Dominated System Reaction Rate in Steady State vs. Grain Size. Rate equations Master equation (integration) MC simulation
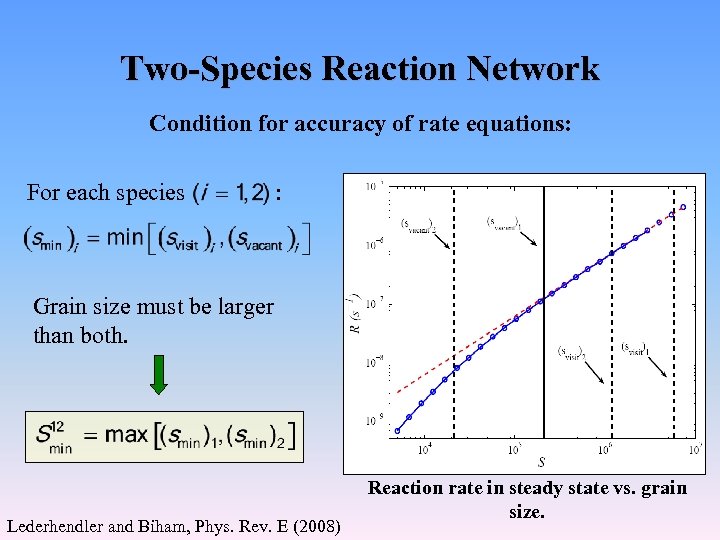 Two-Species Reaction Network Condition for accuracy of rate equations: For each species : Grain size must be larger than both. Lederhendler and Biham, Phys. Rev. E (2008) Reaction rate in steady state vs. grain size.
Two-Species Reaction Network Condition for accuracy of rate equations: For each species : Grain size must be larger than both. Lederhendler and Biham, Phys. Rev. E (2008) Reaction rate in steady state vs. grain size.
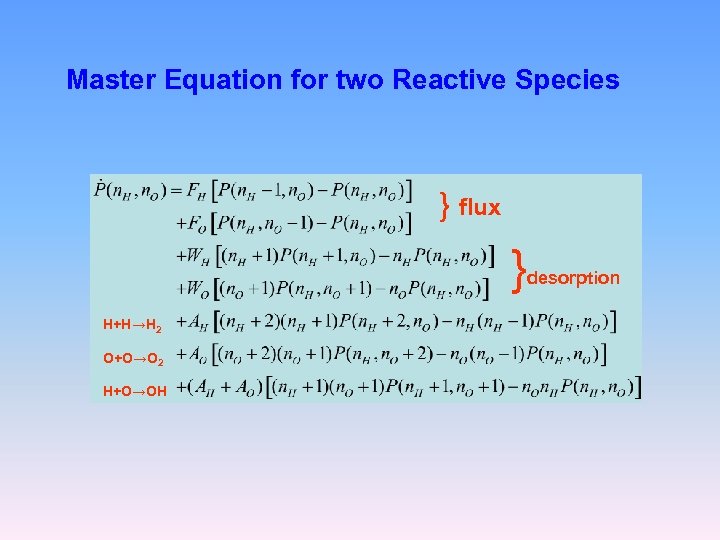 Master Equation for two Reactive Species } flux } H+H→H 2 O+O→O 2 H+O→OH desorption
Master Equation for two Reactive Species } flux } H+H→H 2 O+O→O 2 H+O→OH desorption
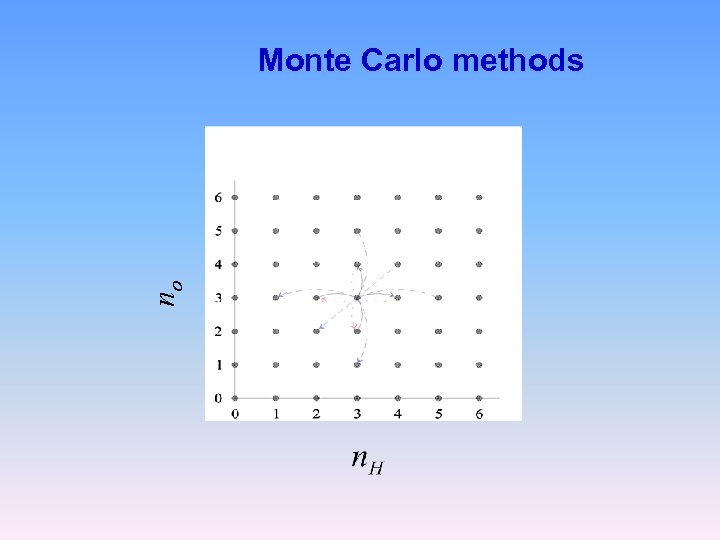 no Monte Carlo methods
no Monte Carlo methods
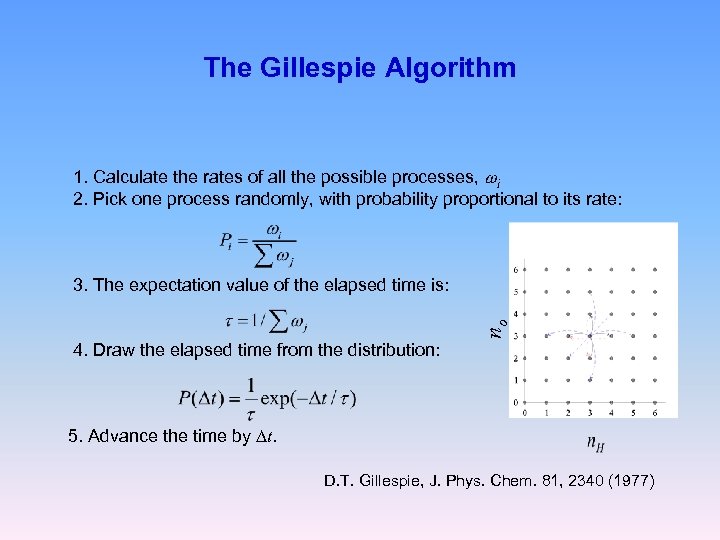 The Gillespie Algorithm 1. Calculate the rates of all the possible processes, wi 2. Pick one process randomly, with probability proportional to its rate: no 3. The expectation value of the elapsed time is: 4. Draw the elapsed time from the distribution: 5. Advance the time by Dt. D. T. Gillespie, J. Phys. Chem. 81, 2340 (1977)
The Gillespie Algorithm 1. Calculate the rates of all the possible processes, wi 2. Pick one process randomly, with probability proportional to its rate: no 3. The expectation value of the elapsed time is: 4. Draw the elapsed time from the distribution: 5. Advance the time by Dt. D. T. Gillespie, J. Phys. Chem. 81, 2340 (1977)
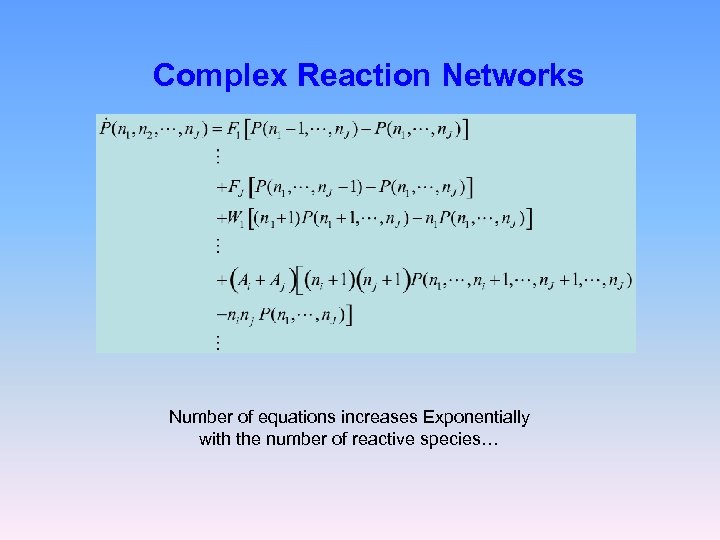 Complex Reaction Networks Number of equations increases Exponentially with the number of reactive species…
Complex Reaction Networks Number of equations increases Exponentially with the number of reactive species…
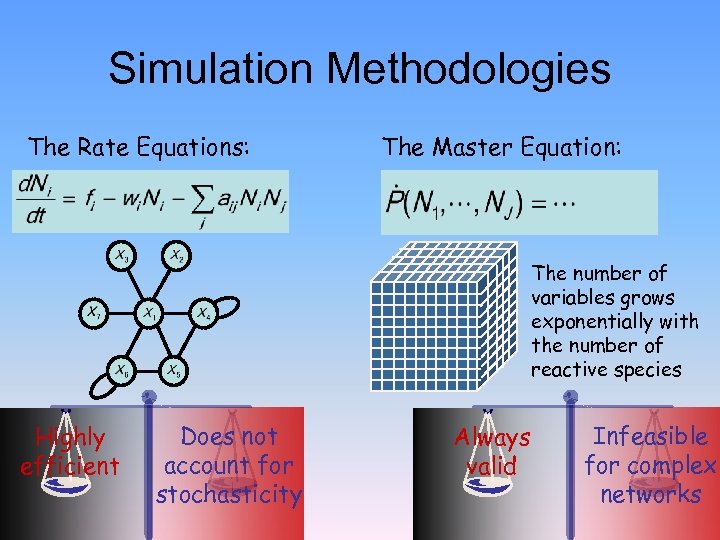 Simulation Methodologies The Rate Equations: The Master Equation: The number of variables grows exponentially with the number of reactive species Highly efficient Does not account for stochasticity Always valid Infeasible for complex networks
Simulation Methodologies The Rate Equations: The Master Equation: The number of variables grows exponentially with the number of reactive species Highly efficient Does not account for stochasticity Always valid Infeasible for complex networks
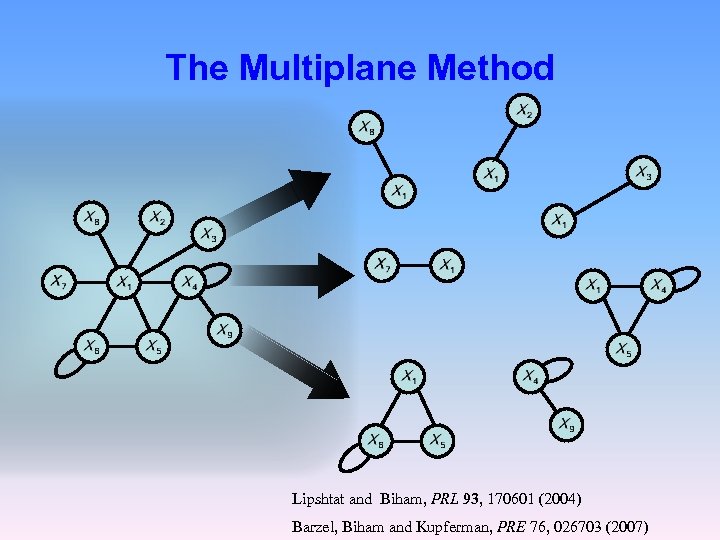 The Multiplane Method Lipshtat and Biham, PRL 93, 170601 (2004) Barzel, Biham and Kupferman, PRE 76, 026703 (2007)
The Multiplane Method Lipshtat and Biham, PRL 93, 170601 (2004) Barzel, Biham and Kupferman, PRE 76, 026703 (2007)
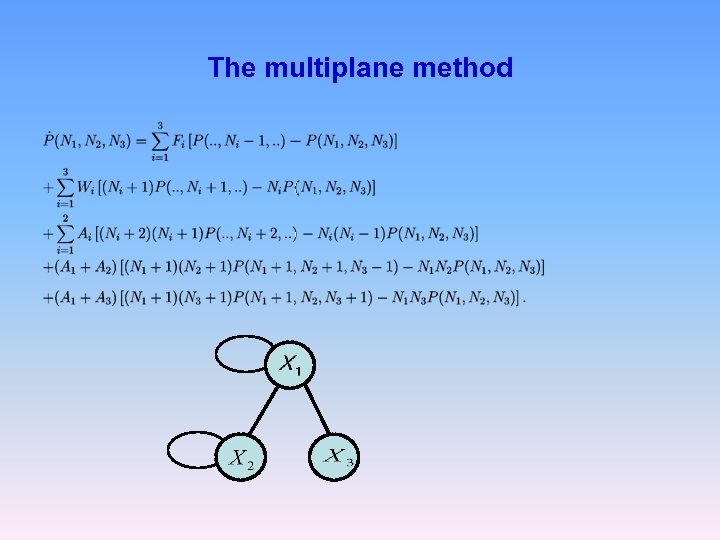 The multiplane method
The multiplane method
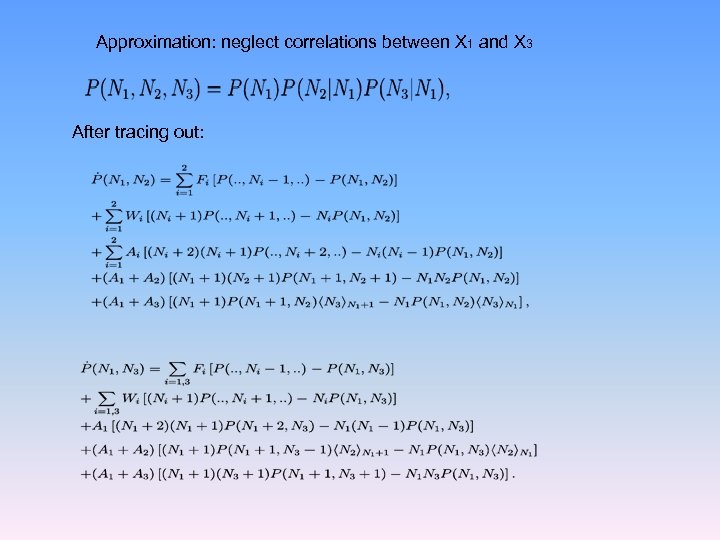 Approximation: neglect correlations between X 1 and X 3 After tracing out:
Approximation: neglect correlations between X 1 and X 3 After tracing out:
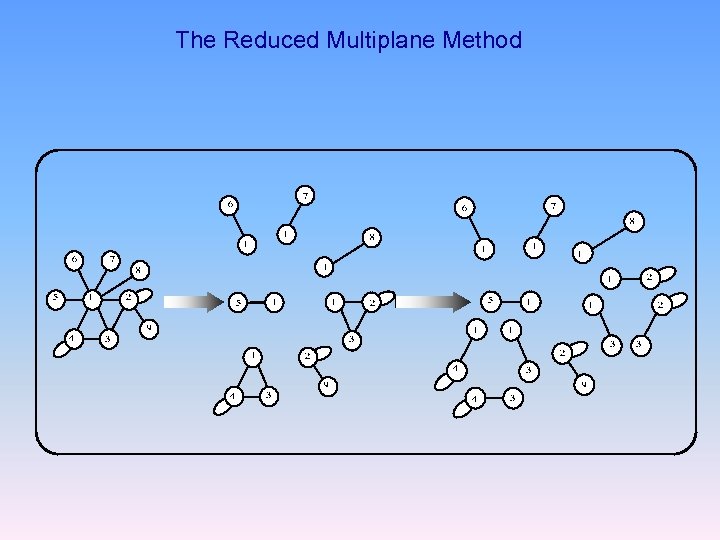 The Reduced Multiplane Method
The Reduced Multiplane Method
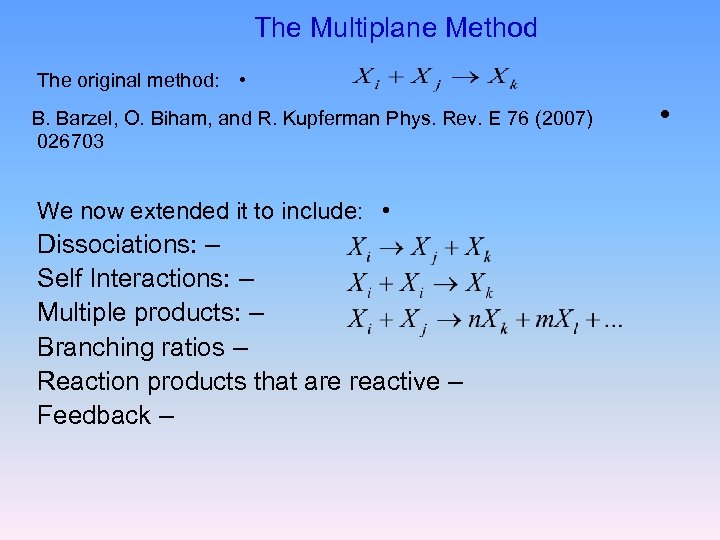 The Multiplane Method The original method: • B. Barzel, O. Biham, and R. Kupferman Phys. Rev. E 76 (2007) 026703 We now extended it to include: • Dissociations: – Self Interactions: – Multiple products: – Branching ratios – Reaction products that are reactive – Feedback – •
The Multiplane Method The original method: • B. Barzel, O. Biham, and R. Kupferman Phys. Rev. E 76 (2007) 026703 We now extended it to include: • Dissociations: – Self Interactions: – Multiple products: – Branching ratios – Reaction products that are reactive – Feedback – •
![System Size [S] Relative Errors Population Size (#/s) Population Size System Size [S] Relative Errors Population Size (#/s) Population Size](https://present5.com/presentation/f044f0d9b7469d5946ed03992391ec66/image-26.jpg) System Size [S] Relative Errors Population Size (#/s) Population Size
System Size [S] Relative Errors Population Size (#/s) Population Size
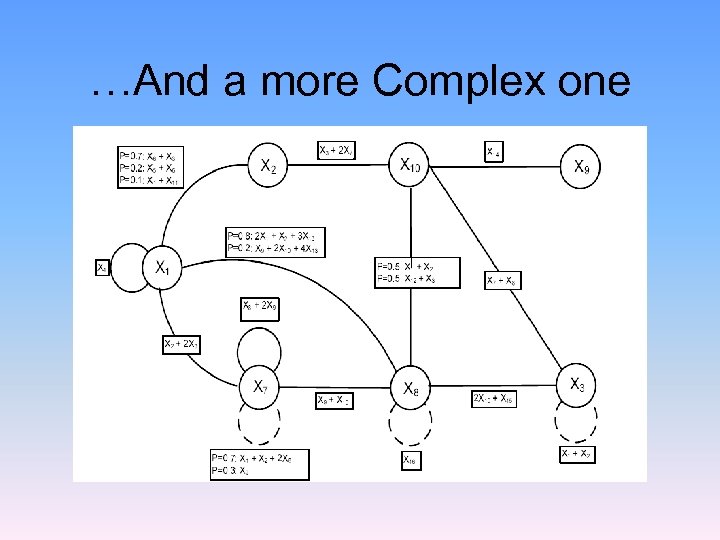 …And a more Complex one
…And a more Complex one
![System Size [S] Production Rate (#s) Population Size System Size [S] Production Rate (#s) Population Size](https://present5.com/presentation/f044f0d9b7469d5946ed03992391ec66/image-28.jpg) System Size [S] Production Rate (#s) Population Size
System Size [S] Production Rate (#s) Population Size
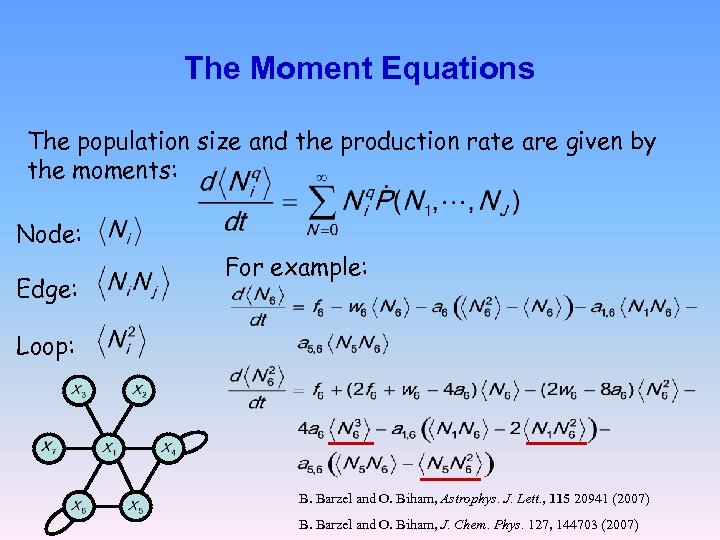 The Moment Equations The population size and the production rate are given by the moments: Node: Edge: For example: Loop: B. Barzel and O. Biham, Astrophys. J. Lett. , 115 20941 (2007) B. Barzel and O. Biham, J. Chem. Phys. 127, 144703 (2007)
The Moment Equations The population size and the production rate are given by the moments: Node: Edge: For example: Loop: B. Barzel and O. Biham, Astrophys. J. Lett. , 115 20941 (2007) B. Barzel and O. Biham, J. Chem. Phys. 127, 144703 (2007)
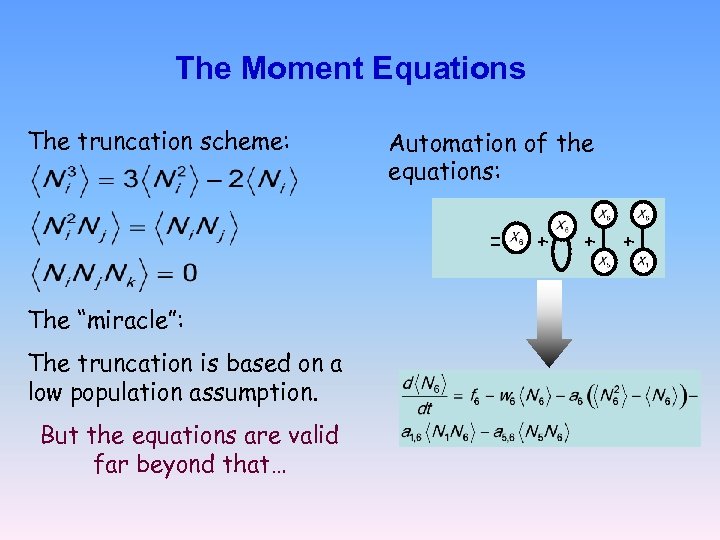 The Moment Equations The truncation scheme: Automation of the equations: = The “miracle”: The truncation is based on a low population assumption. But the equations are valid far beyond that… + + +
The Moment Equations The truncation scheme: Automation of the equations: = The “miracle”: The truncation is based on a low population assumption. But the equations are valid far beyond that… + + +
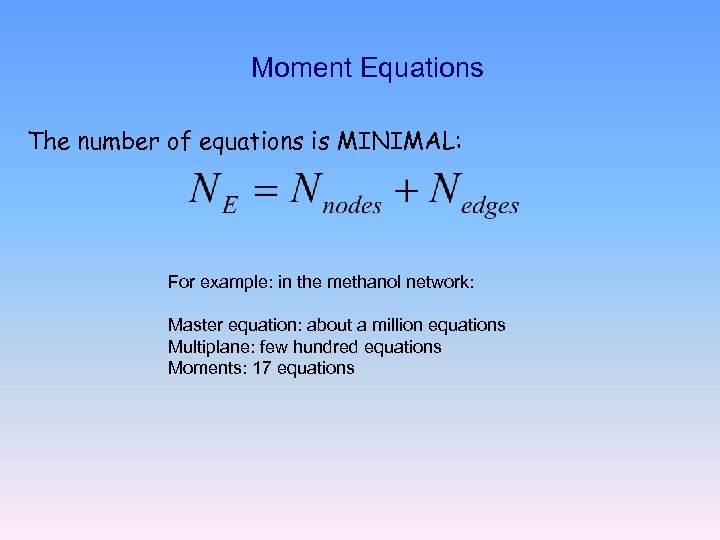 Moment Equations The number of equations is MINIMAL: For example: in the methanol network: Master equation: about a million equations Multiplane: few hundred equations Moments: 17 equations
Moment Equations The number of equations is MINIMAL: For example: in the methanol network: Master equation: about a million equations Multiplane: few hundred equations Moments: 17 equations
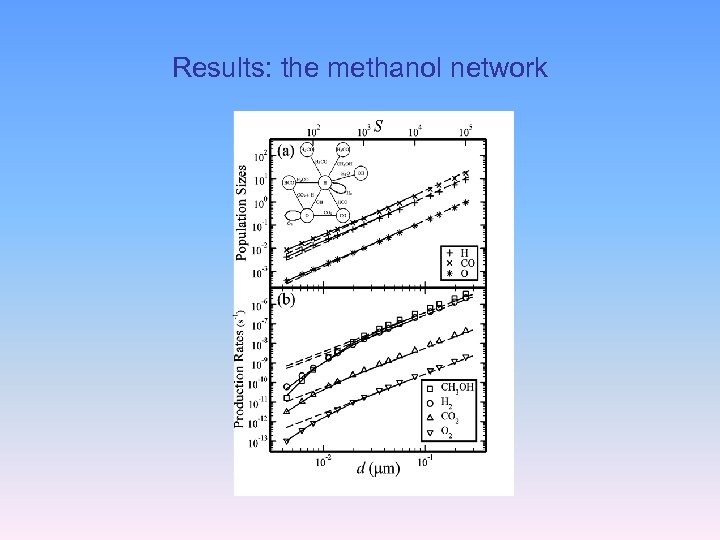 Results: the methanol network
Results: the methanol network
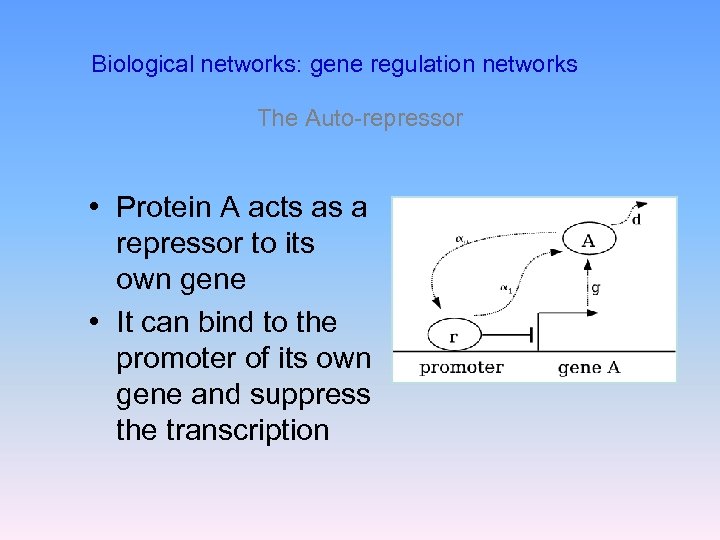 Biological networks: gene regulation networks The Auto-repressor • Protein A acts as a repressor to its own gene • It can bind to the promoter of its own gene and suppress the transcription
Biological networks: gene regulation networks The Auto-repressor • Protein A acts as a repressor to its own gene • It can bind to the promoter of its own gene and suppress the transcription
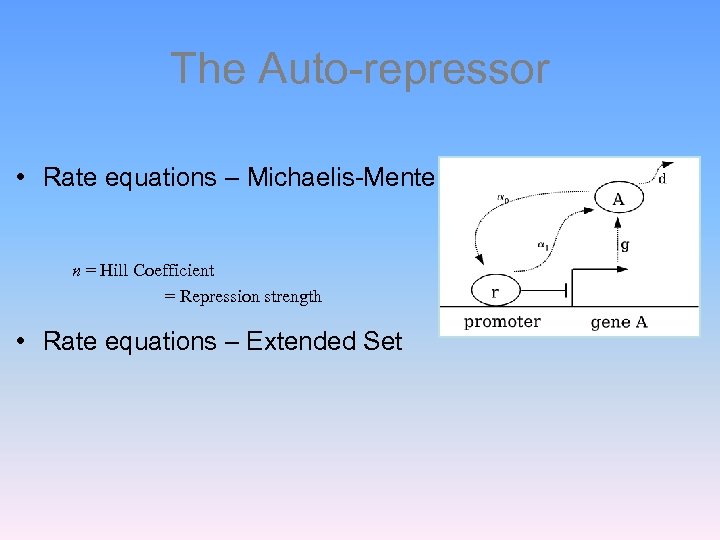 The Auto-repressor • Rate equations – Michaelis-Menten form n = Hill Coefficient = Repression strength • Rate equations – Extended Set
The Auto-repressor • Rate equations – Michaelis-Menten form n = Hill Coefficient = Repression strength • Rate equations – Extended Set
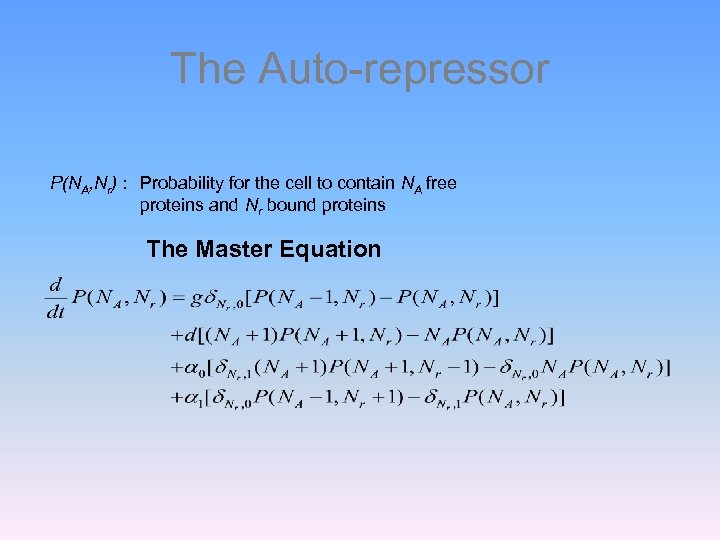 The Auto-repressor P(NA, Nr) : Probability for the cell to contain NA free proteins and Nr bound proteins The Master Equation
The Auto-repressor P(NA, Nr) : Probability for the cell to contain NA free proteins and Nr bound proteins The Master Equation
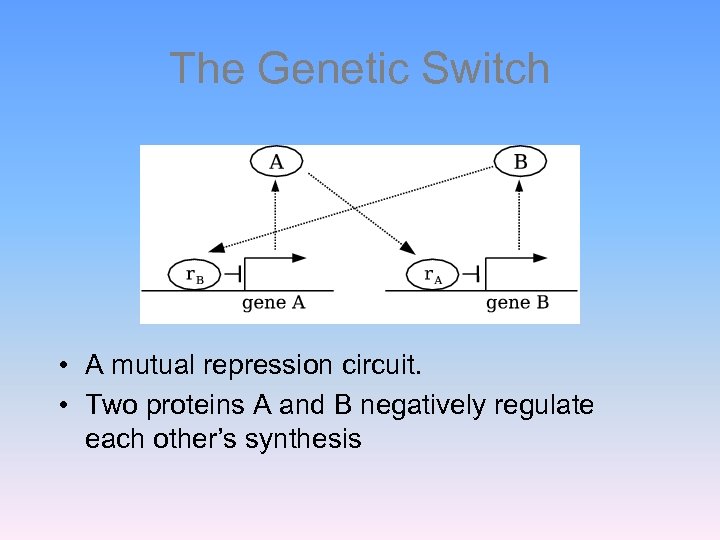 The Genetic Switch • A mutual repression circuit. • Two proteins A and B negatively regulate each other’s synthesis
The Genetic Switch • A mutual repression circuit. • Two proteins A and B negatively regulate each other’s synthesis
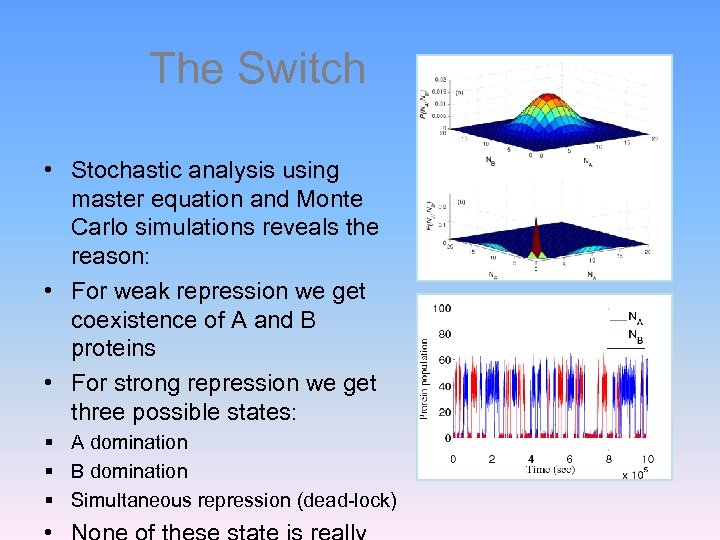 The Switch • Stochastic analysis using master equation and Monte Carlo simulations reveals the reason: • For weak repression we get coexistence of A and B proteins • For strong repression we get three possible states: § A domination § B domination § Simultaneous repression (dead-lock)
The Switch • Stochastic analysis using master equation and Monte Carlo simulations reveals the reason: • For weak repression we get coexistence of A and B proteins • For strong repression we get three possible states: § A domination § B domination § Simultaneous repression (dead-lock)
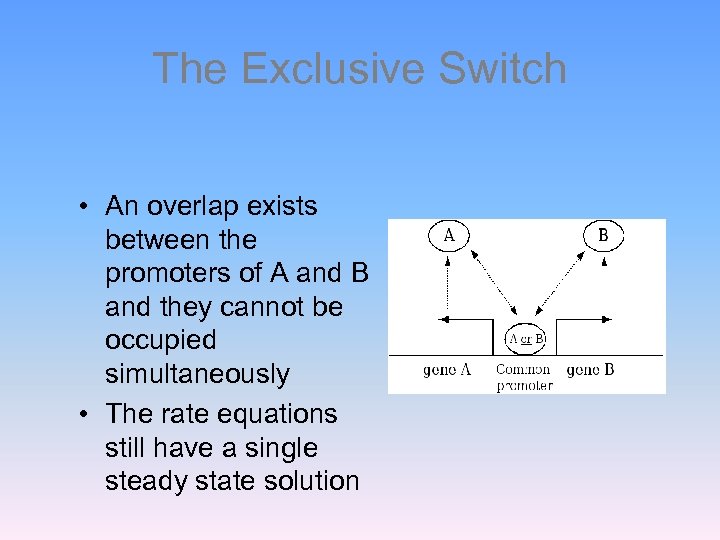 The Exclusive Switch • An overlap exists between the promoters of A and B and they cannot be occupied simultaneously • The rate equations still have a single steady state solution
The Exclusive Switch • An overlap exists between the promoters of A and B and they cannot be occupied simultaneously • The rate equations still have a single steady state solution
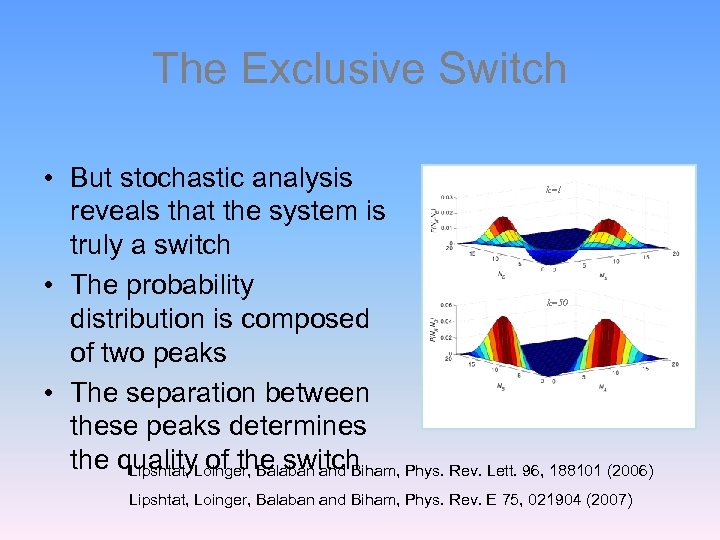 The Exclusive Switch • But stochastic analysis reveals that the system is truly a switch • The probability distribution is composed of two peaks • The separation between these peaks determines the quality. Loinger, Balaban and Biham, Phys. Rev. Lett. 96, 188101 (2006) Lipshtat, of the switch k=1 k=50 Lipshtat, Loinger, Balaban and Biham, Phys. Rev. E 75, 021904 (2007)
The Exclusive Switch • But stochastic analysis reveals that the system is truly a switch • The probability distribution is composed of two peaks • The separation between these peaks determines the quality. Loinger, Balaban and Biham, Phys. Rev. Lett. 96, 188101 (2006) Lipshtat, of the switch k=1 k=50 Lipshtat, Loinger, Balaban and Biham, Phys. Rev. E 75, 021904 (2007)
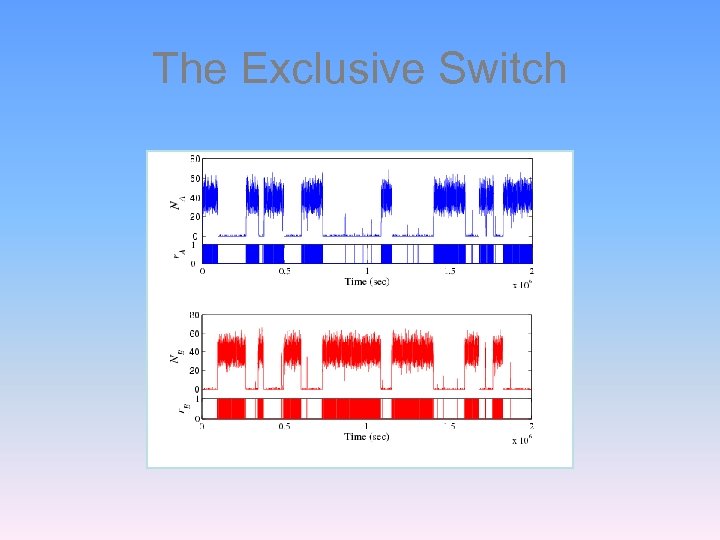 The Exclusive Switch
The Exclusive Switch
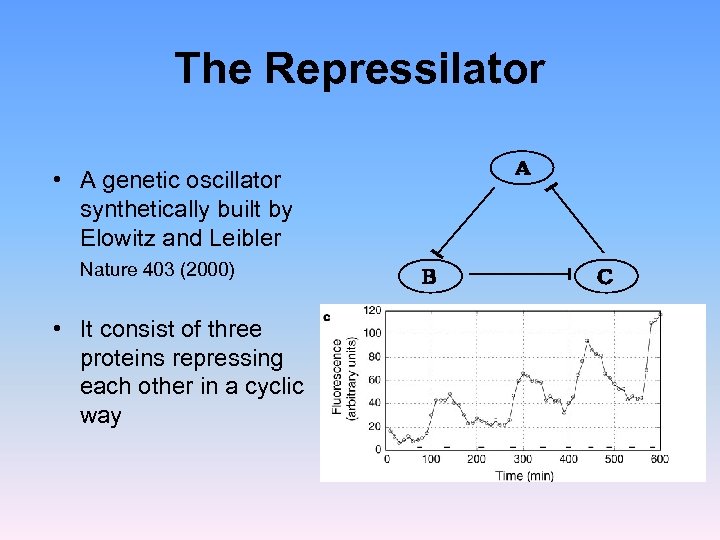 The Repressilator • A genetic oscillator synthetically built by Elowitz and Leibler Nature 403 (2000) • It consist of three proteins repressing each other in a cyclic way
The Repressilator • A genetic oscillator synthetically built by Elowitz and Leibler Nature 403 (2000) • It consist of three proteins repressing each other in a cyclic way
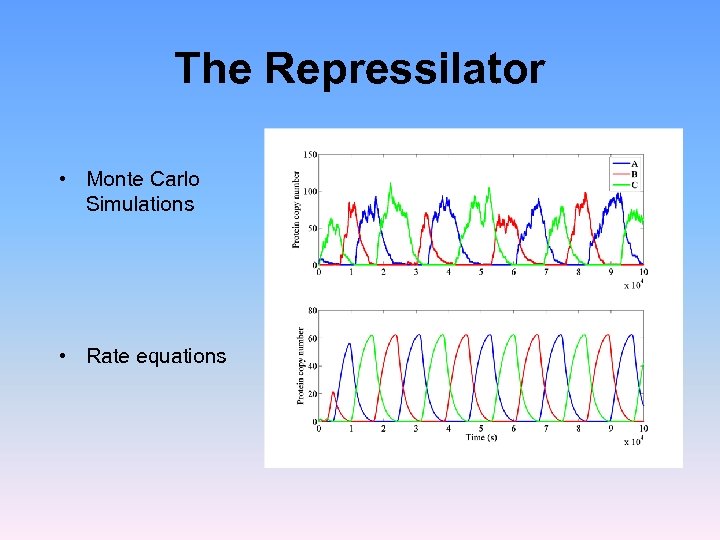 The Repressilator • Monte Carlo Simulations • Rate equations
The Repressilator • Monte Carlo Simulations • Rate equations
 Plasmids • The repressilator and synthetic toggle switch were encoded on plasmids in E. coli. • Plasmids are circular self replicating DNA molecules which include only few genes. • The number of plasmids in a cell can be controlled. How does the number of plasmids affect the dynamical behavior?
Plasmids • The repressilator and synthetic toggle switch were encoded on plasmids in E. coli. • Plasmids are circular self replicating DNA molecules which include only few genes. • The number of plasmids in a cell can be controlled. How does the number of plasmids affect the dynamical behavior?
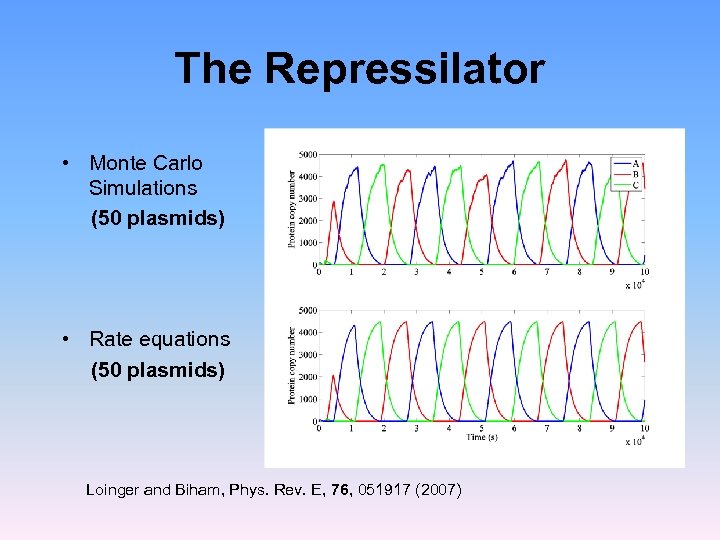 The Repressilator • Monte Carlo Simulations (50 plasmids) • Rate equations (50 plasmids) Loinger and Biham, Phys. Rev. E, 76, 051917 (2007)
The Repressilator • Monte Carlo Simulations (50 plasmids) • Rate equations (50 plasmids) Loinger and Biham, Phys. Rev. E, 76, 051917 (2007)
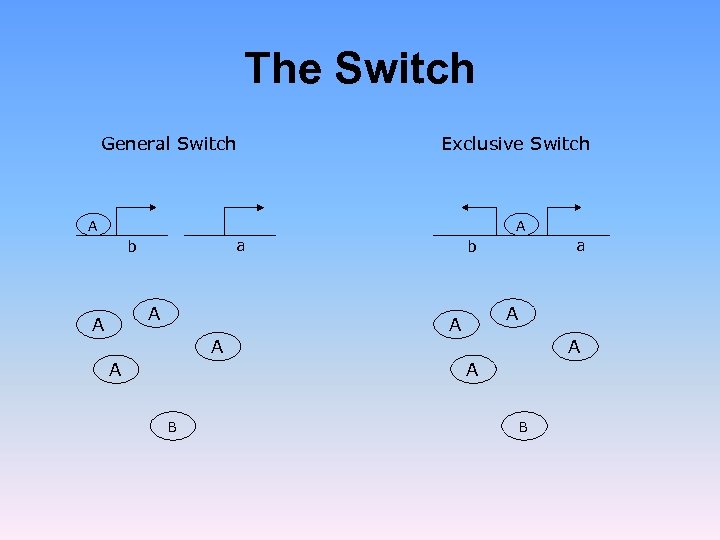 The Switch General Switch Exclusive Switch A A a b A A A B B
The Switch General Switch Exclusive Switch A A a b A A A B B
![The Switch Warren and ten Wolde [PRL 92, 128101 (2004)] showed that for a The Switch Warren and ten Wolde [PRL 92, 128101 (2004)] showed that for a](https://present5.com/presentation/f044f0d9b7469d5946ed03992391ec66/image-47.jpg) The Switch Warren and ten Wolde [PRL 92, 128101 (2004)] showed that for a single plasmid with h = 2, the exclusive switch is more stable than the general switch. This does not hold for a high plasmid copy number Loinger and Biham, Phys. Rev. Lett. , 103, 068104 (2009)
The Switch Warren and ten Wolde [PRL 92, 128101 (2004)] showed that for a single plasmid with h = 2, the exclusive switch is more stable than the general switch. This does not hold for a high plasmid copy number Loinger and Biham, Phys. Rev. Lett. , 103, 068104 (2009)
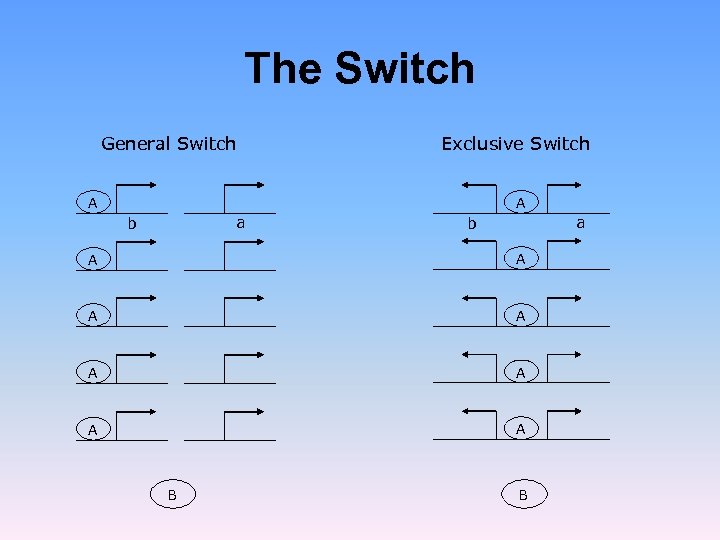 The Switch General Switch Exclusive Switch A A a b A A A A B B
The Switch General Switch Exclusive Switch A A a b A A A A B B
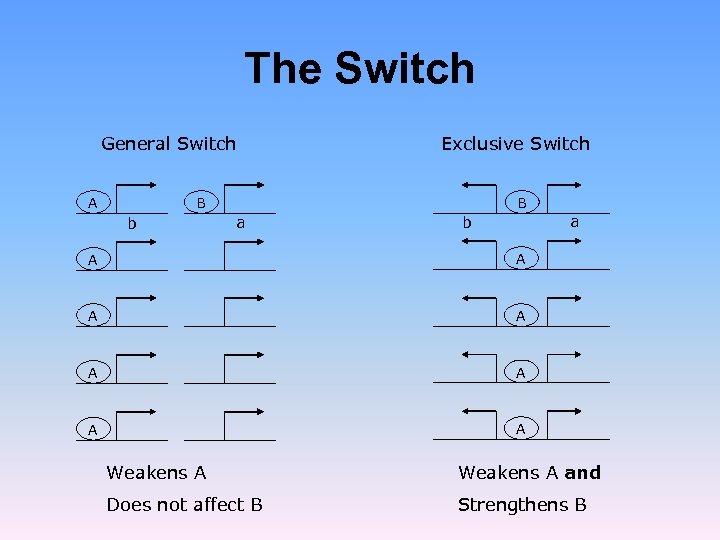 The Switch General Switch A Exclusive Switch B b B a b A A A A a A Weakens A and Does not affect B Strengthens B
The Switch General Switch A Exclusive Switch B b B a b A A A A a A Weakens A and Does not affect B Strengthens B
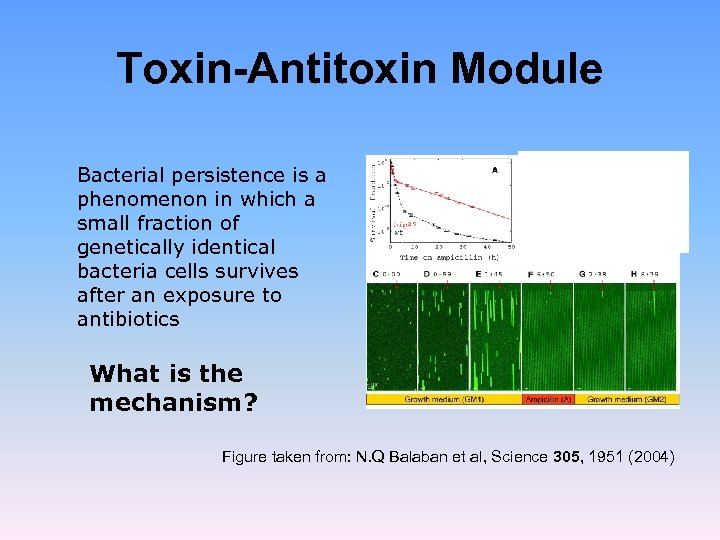 Toxin-Antitoxin Module Bacterial persistence is a phenomenon in which a small fraction of genetically identical bacteria cells survives after an exposure to antibiotics What is the mechanism? Figure taken from: N. Q Balaban et al, Science 305, 1951 (2004)
Toxin-Antitoxin Module Bacterial persistence is a phenomenon in which a small fraction of genetically identical bacteria cells survives after an exposure to antibiotics What is the mechanism? Figure taken from: N. Q Balaban et al, Science 305, 1951 (2004)
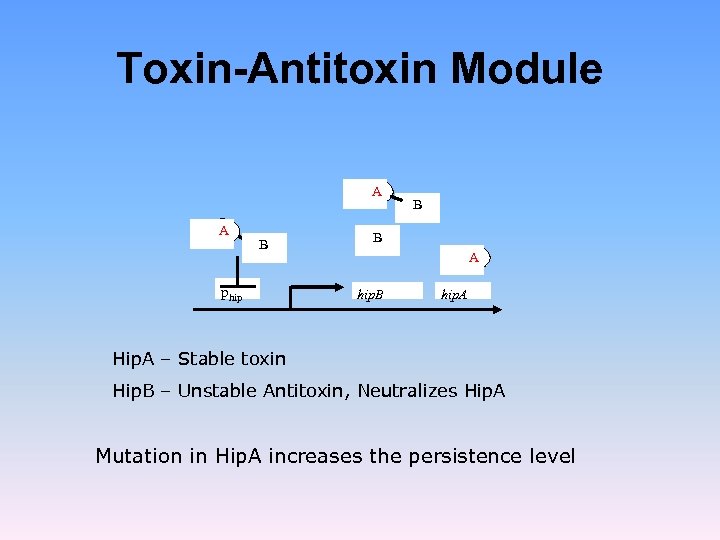 Toxin-Antitoxin Module A A B phip B B A hip. B hip. A Hip. A – Stable toxin Hip. B – Unstable Antitoxin, Neutralizes Hip. A Mutation in Hip. A increases the persistence level
Toxin-Antitoxin Module A A B phip B B A hip. B hip. A Hip. A – Stable toxin Hip. B – Unstable Antitoxin, Neutralizes Hip. A Mutation in Hip. A increases the persistence level
 Toxin-Antitoxin Module
Toxin-Antitoxin Module
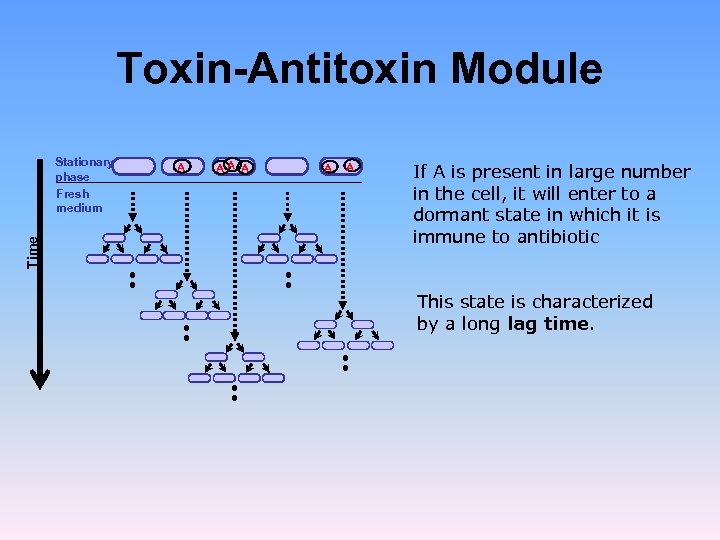 Toxin-Antitoxin Module Time Stationary phase Fresh medium A AA A If A is present in large number in the cell, it will enter to a dormant state in which it is immune to antibiotic This state is characterized by a long lag time.
Toxin-Antitoxin Module Time Stationary phase Fresh medium A AA A If A is present in large number in the cell, it will enter to a dormant state in which it is immune to antibiotic This state is characterized by a long lag time.
![Lag time [a. u. ] Lag time [min] Toxin-Antitoxin Module Hip. A expression level Lag time [a. u. ] Lag time [min] Toxin-Antitoxin Module Hip. A expression level](https://present5.com/presentation/f044f0d9b7469d5946ed03992391ec66/image-54.jpg) Lag time [a. u. ] Lag time [min] Toxin-Antitoxin Module Hip. A expression level (m. Cherry fluorescence) [a. u. ] Hip. A expression level (Atot) [a. u. ] Threshold behavior Dormant/persister state if free A is large enough
Lag time [a. u. ] Lag time [min] Toxin-Antitoxin Module Hip. A expression level (m. Cherry fluorescence) [a. u. ] Hip. A expression level (Atot) [a. u. ] Threshold behavior Dormant/persister state if free A is large enough
![Toxin-Antitoxin Module Fraction of persisters = Prob(A>A ) 0 [Threshold] Dependence on A and Toxin-Antitoxin Module Fraction of persisters = Prob(A>A ) 0 [Threshold] Dependence on A and](https://present5.com/presentation/f044f0d9b7469d5946ed03992391ec66/image-55.jpg) Toxin-Antitoxin Module Fraction of persisters = Prob(A>A ) 0 [Threshold] Dependence on A and B production Experiment P(Free A > A 0) Fraction of persisters Theory Hip. A expression level (Atot) [a. u. ] Hip. A expression level (m. Cherry fluorescence) [a. u. ] Rotem, Loinger, Ronin, Levin-Reisman, Gabay, Biham and Balaban, preprint
Toxin-Antitoxin Module Fraction of persisters = Prob(A>A ) 0 [Threshold] Dependence on A and B production Experiment P(Free A > A 0) Fraction of persisters Theory Hip. A expression level (Atot) [a. u. ] Hip. A expression level (m. Cherry fluorescence) [a. u. ] Rotem, Loinger, Ronin, Levin-Reisman, Gabay, Biham and Balaban, preprint
 Summary • Direct integration of the master equation and Monte Carlo simulations are not feasible for the simulation of complex reaction networks. • The moment equations provide an efficient method for the evaluation of reaction rates. The multiplane method also provides a good approximation of the distribution. • The number of equations is dramatically reduced compared to the master equation. The moment equations include one equation for each species and one equation for each reaction. • The moment equations are easy to construct using a diagrammatic method.
Summary • Direct integration of the master equation and Monte Carlo simulations are not feasible for the simulation of complex reaction networks. • The moment equations provide an efficient method for the evaluation of reaction rates. The multiplane method also provides a good approximation of the distribution. • The number of equations is dramatically reduced compared to the master equation. The moment equations include one equation for each species and one equation for each reaction. • The moment equations are easy to construct using a diagrammatic method.


