1440906541f1b2c9e276a4b4e06fe9f8.ppt
- Количество слайдов: 23
 Rational Drug Design Soma Mandal , Mee'nal Moudgil , Sanat K. Mandal
Rational Drug Design Soma Mandal , Mee'nal Moudgil , Sanat K. Mandal
 Introduction • Drug: Compounds used for the prevention and treatment of diseases like cancer, etc. • Ideal drug: 1) target: bio-molecule , involved in signaling or metabolic pathways, that are specific to disease process by either protein-protein or protein-nucleic acid interactions. 2)antagonist action-inhibiting functions of the disease causing proteins. 3) Inhibiting interactions of the proteins. 4) Activates other proteins, that are deregulated in such disease like cancer.
Introduction • Drug: Compounds used for the prevention and treatment of diseases like cancer, etc. • Ideal drug: 1) target: bio-molecule , involved in signaling or metabolic pathways, that are specific to disease process by either protein-protein or protein-nucleic acid interactions. 2)antagonist action-inhibiting functions of the disease causing proteins. 3) Inhibiting interactions of the proteins. 4) Activates other proteins, that are deregulated in such disease like cancer.
 Drug Discovery Protein targets for drugs can be known by 3 D X-ray crystallography, NMR, docking tools, Computer aided drug designing. PDB: 71550 proteins structure available. X-ray crystallography: 63124 NMR: 7985 Cryoelectron microscopy: 266 Hybrid: 42 Other: 133 Still, insignificant so many lead drugs are unknown.
Drug Discovery Protein targets for drugs can be known by 3 D X-ray crystallography, NMR, docking tools, Computer aided drug designing. PDB: 71550 proteins structure available. X-ray crystallography: 63124 NMR: 7985 Cryoelectron microscopy: 266 Hybrid: 42 Other: 133 Still, insignificant so many lead drugs are unknown.
 Drug designing is: 1)challenging 2)Expensive 3)Time consuming So, Multidisciplinary approach: Computational tools, methodologies for structure guided approach + Global gene expression data analysis by softwares. Hence, 1) Efficiency increased 2) Cost effectiveness 3) Time saved 4) Strategies to overcome toxic side effects
Drug designing is: 1)challenging 2)Expensive 3)Time consuming So, Multidisciplinary approach: Computational tools, methodologies for structure guided approach + Global gene expression data analysis by softwares. Hence, 1) Efficiency increased 2) Cost effectiveness 3) Time saved 4) Strategies to overcome toxic side effects
 Drug Design 2 ways: A) Development of ligands with desired properties for targets having known structure and functions. B) Development of ligands with predefined properties for targets whose structural information may be or may not be known. This, unknown target information can be found by global gene expression data.
Drug Design 2 ways: A) Development of ligands with desired properties for targets having known structure and functions. B) Development of ligands with predefined properties for targets whose structural information may be or may not be known. This, unknown target information can be found by global gene expression data.
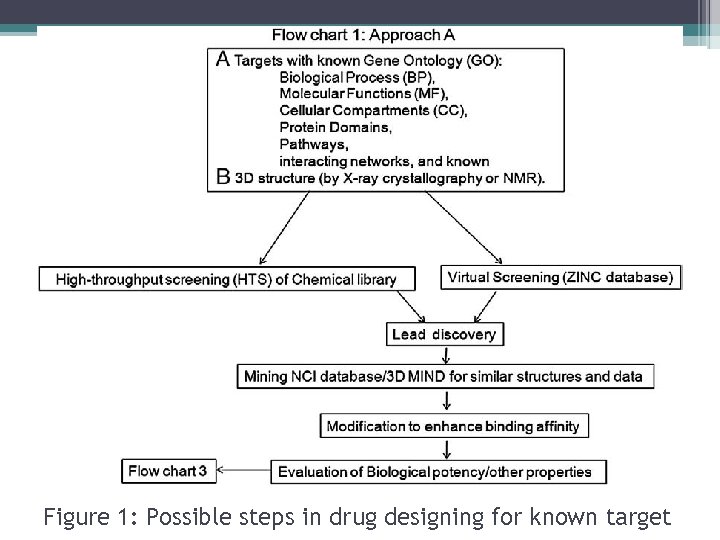 Figure 1: Possible steps in drug designing for known target
Figure 1: Possible steps in drug designing for known target
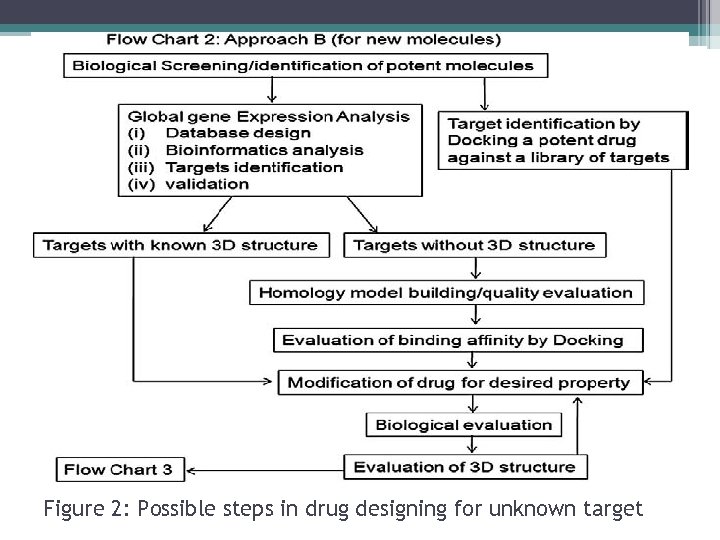 Figure 2: Possible steps in drug designing for unknown target
Figure 2: Possible steps in drug designing for unknown target
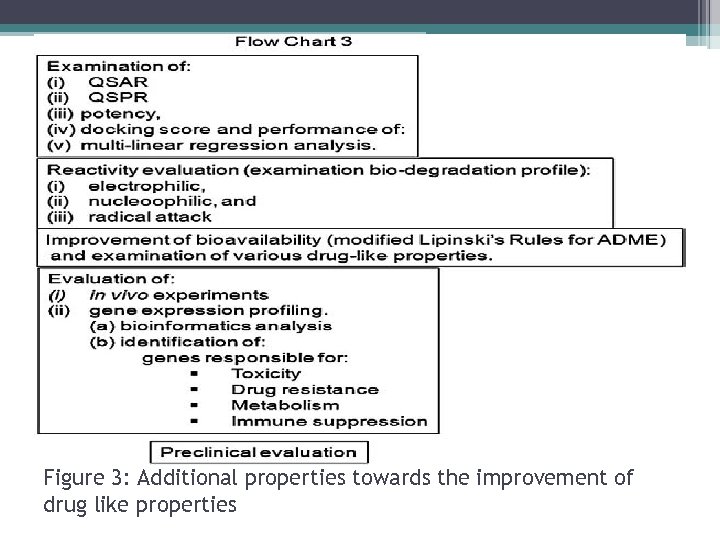 Figure 3: Additional properties towards the improvement of drug like properties
Figure 3: Additional properties towards the improvement of drug like properties
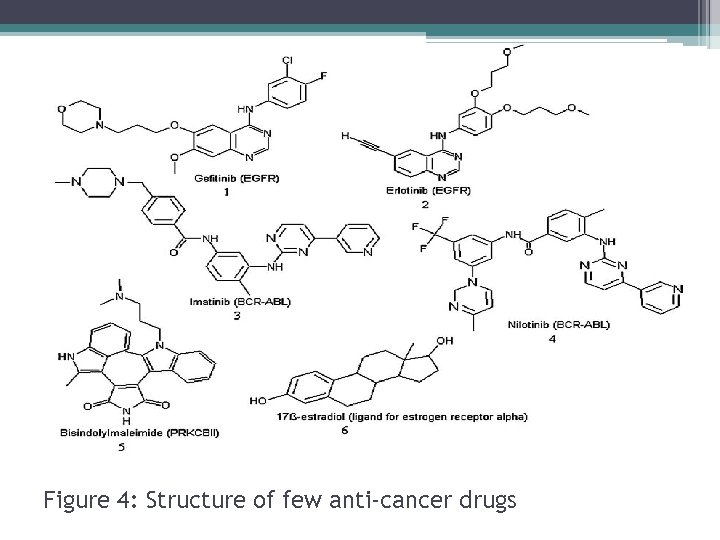 Figure 4: Structure of few anti-cancer drugs
Figure 4: Structure of few anti-cancer drugs
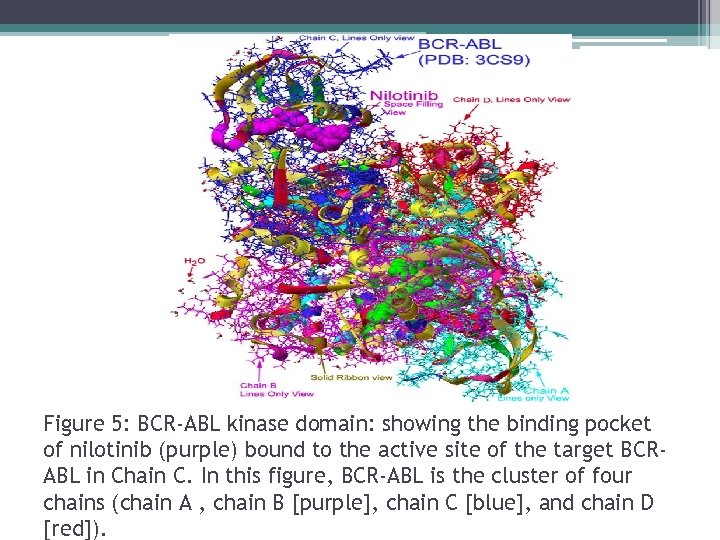 Figure 5: BCR-ABL kinase domain: showing the binding pocket of nilotinib (purple) bound to the active site of the target BCRABL in Chain C. In this figure, BCR-ABL is the cluster of four chains (chain A , chain B [purple], chain C [blue], and chain D [red]).
Figure 5: BCR-ABL kinase domain: showing the binding pocket of nilotinib (purple) bound to the active site of the target BCRABL in Chain C. In this figure, BCR-ABL is the cluster of four chains (chain A , chain B [purple], chain C [blue], and chain D [red]).
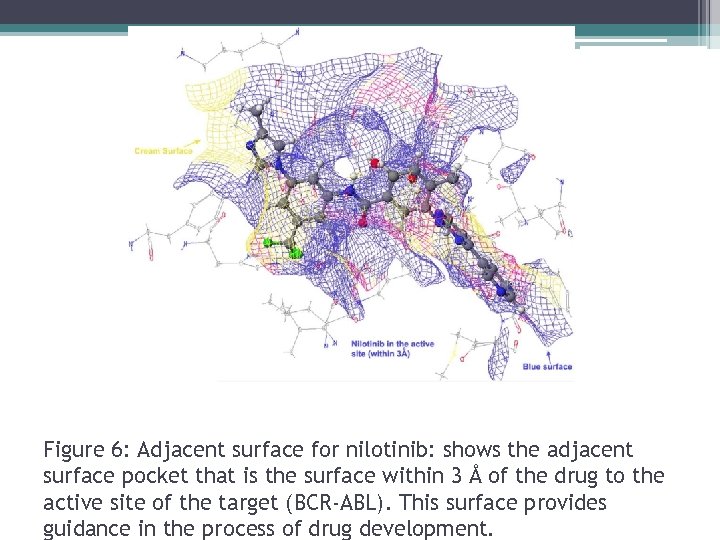 Figure 6: Adjacent surface for nilotinib: shows the adjacent surface pocket that is the surface within 3 Å of the drug to the active site of the target (BCR-ABL). This surface provides guidance in the process of drug development.
Figure 6: Adjacent surface for nilotinib: shows the adjacent surface pocket that is the surface within 3 Å of the drug to the active site of the target (BCR-ABL). This surface provides guidance in the process of drug development.
 - This adjacent surface provides information necessary for modification of bound drug. - Red= where protein needs H-acceptors - Cream= Hydrophobic surface - Pocket surface= active site Ligands that bind well, should have H-bond acceptor groups touching red surface. Reactivity: Nucleophilic, electrophilic or radical frontier density measures susceptibility of substrate by attack. -It reveals reactive sites based on electron configuration.
- This adjacent surface provides information necessary for modification of bound drug. - Red= where protein needs H-acceptors - Cream= Hydrophobic surface - Pocket surface= active site Ligands that bind well, should have H-bond acceptor groups touching red surface. Reactivity: Nucleophilic, electrophilic or radical frontier density measures susceptibility of substrate by attack. -It reveals reactive sites based on electron configuration.
 Known targets for Cancer therapy: • EGFR shows aberrant expression in cancers like lung cancer, breast cancer. • It interacts with other 151 proteins. • It is involved in 19 signaling pathways. • EGF is a natural ligand binds to EGFR and mediates signaling events. • Another drug with the EGFR as a target is gefitinib. • It attaches to protein kinase domain in EGFR, which is an important target involved in many malignancy promoting processes.
Known targets for Cancer therapy: • EGFR shows aberrant expression in cancers like lung cancer, breast cancer. • It interacts with other 151 proteins. • It is involved in 19 signaling pathways. • EGF is a natural ligand binds to EGFR and mediates signaling events. • Another drug with the EGFR as a target is gefitinib. • It attaches to protein kinase domain in EGFR, which is an important target involved in many malignancy promoting processes.
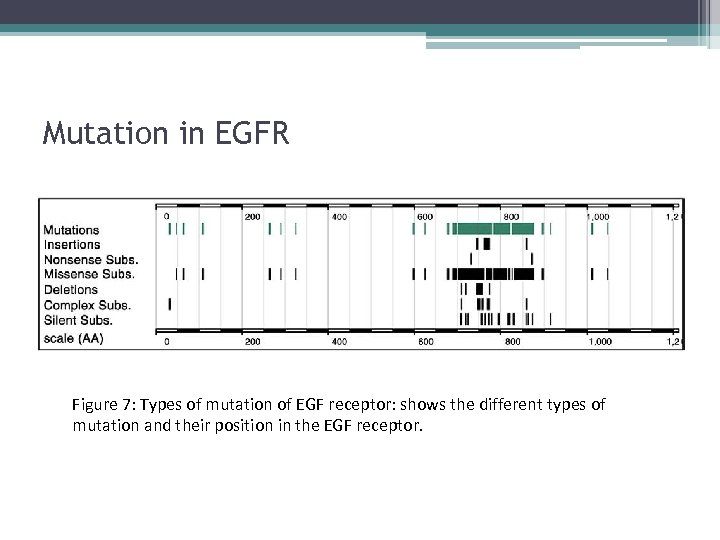 Mutation in EGFR Figure 7: Types of mutation of EGF receptor: shows the different types of mutation and their position in the EGF receptor.
Mutation in EGFR Figure 7: Types of mutation of EGF receptor: shows the different types of mutation and their position in the EGF receptor.
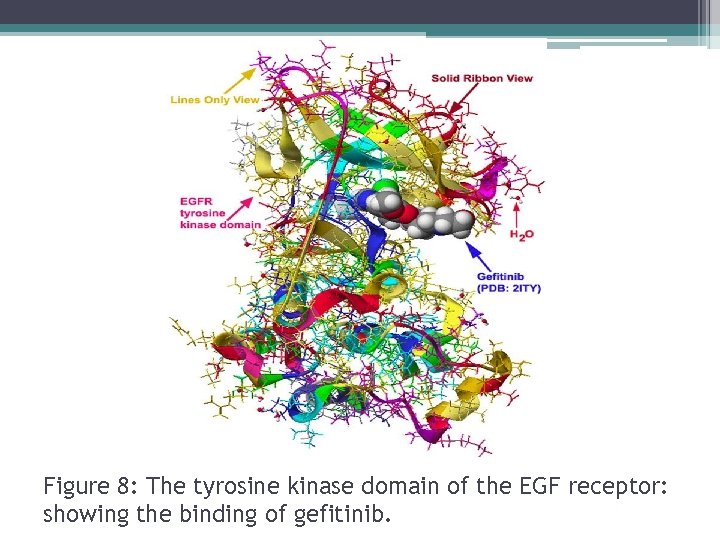 Figure 8: The tyrosine kinase domain of the EGF receptor: showing the binding of gefitinib.
Figure 8: The tyrosine kinase domain of the EGF receptor: showing the binding of gefitinib.
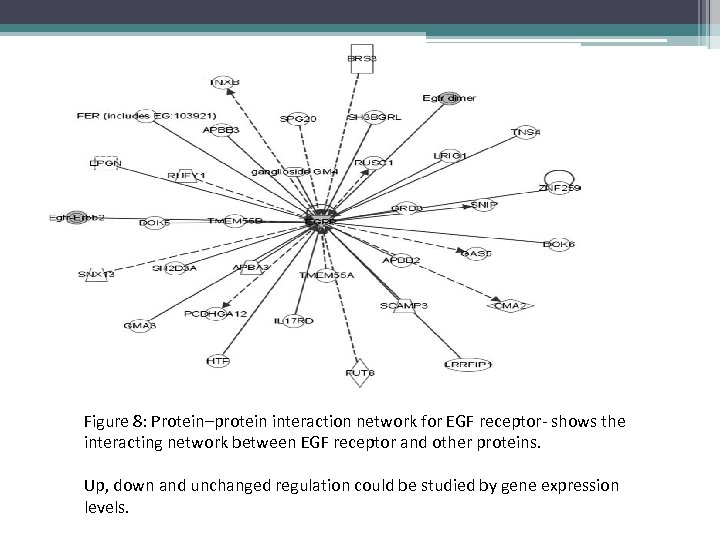 Figure 8: Protein–protein interaction network for EGF receptor- shows the interacting network between EGF receptor and other proteins. Up, down and unchanged regulation could be studied by gene expression levels.
Figure 8: Protein–protein interaction network for EGF receptor- shows the interacting network between EGF receptor and other proteins. Up, down and unchanged regulation could be studied by gene expression levels.
 Drug Designing Methods: • Databases: PDB: Experimentally determined structures. NCI: National Cancer Institute. Pubchem: Bulk data available for QSAR for lead discovery. 3 D MIND: Information about cytotoxic potency for 60 human cancer cell lines. OSIRIS: To draw chemical structures and predict drug like properties like absorption in body etc. Links at the bottom of the paper. Autodock and Dock 6 for docking
Drug Designing Methods: • Databases: PDB: Experimentally determined structures. NCI: National Cancer Institute. Pubchem: Bulk data available for QSAR for lead discovery. 3 D MIND: Information about cytotoxic potency for 60 human cancer cell lines. OSIRIS: To draw chemical structures and predict drug like properties like absorption in body etc. Links at the bottom of the paper. Autodock and Dock 6 for docking
 Currents drugs: 1) Safety concerns 2) Adverse toxic side effects 3) Cardiac Toxicity 4) Devlopment of drug resistance. So, the maximum docking score doesn’t necessarily mean it’s a potent drug. Hence, current drugs need to overcome this hurdle of toxicity.
Currents drugs: 1) Safety concerns 2) Adverse toxic side effects 3) Cardiac Toxicity 4) Devlopment of drug resistance. So, the maximum docking score doesn’t necessarily mean it’s a potent drug. Hence, current drugs need to overcome this hurdle of toxicity.
 Combination Therapy • Ancient Asian herbal medicine with combination of many herbs produces no side effects • Same approach is applied to modern medicine • We can use combination of different drugs to combat a specific disease like cancer.
Combination Therapy • Ancient Asian herbal medicine with combination of many herbs produces no side effects • Same approach is applied to modern medicine • We can use combination of different drugs to combat a specific disease like cancer.
 Multidisciplinary Approach A) Global gene expression profiling - It reveals insights into pathogenosis of diseases including cancer. Methods such as SAGE or Microarray analysis(MA) are used. To access SAGE or MA, bioinformatics tools are used. B) Global Gene Expression Analysis • Databases like GEO (for microarray data) or SAGE data depository, Stanford MA Database etc. are available. • We can ‘group the genes’ based on the data from above mentioned tools for gene annotation. • DAVID tool- gene annotation, gene ontology, protein pathways and protein domains finding. • GSEA (Gene Set Enrichment Analysis)- defines whether set of genes shows difference between normal and abnormal states of the genes or not.
Multidisciplinary Approach A) Global gene expression profiling - It reveals insights into pathogenosis of diseases including cancer. Methods such as SAGE or Microarray analysis(MA) are used. To access SAGE or MA, bioinformatics tools are used. B) Global Gene Expression Analysis • Databases like GEO (for microarray data) or SAGE data depository, Stanford MA Database etc. are available. • We can ‘group the genes’ based on the data from above mentioned tools for gene annotation. • DAVID tool- gene annotation, gene ontology, protein pathways and protein domains finding. • GSEA (Gene Set Enrichment Analysis)- defines whether set of genes shows difference between normal and abnormal states of the genes or not.
 • All these tools are free tools but many commercial tools are available to carry out specific experiments. • Gene up/down/unchanged expression is required for target finding for lead molecules. • Modification of the drug itself or applying different strategies is necessary for combination therapy.
• All these tools are free tools but many commercial tools are available to carry out specific experiments. • Gene up/down/unchanged expression is required for target finding for lead molecules. • Modification of the drug itself or applying different strategies is necessary for combination therapy.
 Conclusion • Cancer is common in all the ages and it shows metastasis, angiogenesis and cell death, etc. • So, drugs to cure cancer are necessary. • Except toxicity, drugs are quite tolerable so they are combined with other drugs. • Comparative analysis of gene expression levels between drug treated and non-treated condition should be studied. • If getting superior quality drug than the available ones, clinical trial phases can be performed on them.
Conclusion • Cancer is common in all the ages and it shows metastasis, angiogenesis and cell death, etc. • So, drugs to cure cancer are necessary. • Except toxicity, drugs are quite tolerable so they are combined with other drugs. • Comparative analysis of gene expression levels between drug treated and non-treated condition should be studied. • If getting superior quality drug than the available ones, clinical trial phases can be performed on them.
 • QUESTIONS? ? ?
• QUESTIONS? ? ?


