e4e38ddcc0c771a46369bccc42cd0e05.ppt
- Количество слайдов: 32

Qualitative Simulation of the Carbon Starvation Response in Escherichia coli Delphine Ropers INRIA Rhône-Alpes 655 avenue de l’Europe Montbonnot, 38334 Saint Ismier CEDEX, France Email: Delphine. Ropers@inrialpes. fr Web: http: //www-helix. inrialpes. fr/article 593. html

Overview 1. Introduction: nutritional stress response in E. coli 2. Qualitative modeling and simulation of genetic regulatory networks 3. Modeling of carbon starvation response in E. coli 4. Experimental validation of model predictions 5. Work in progress 2

Stress response in Escherichia coli v Bacteria able to adapt to a variety of changing environmental conditions Heat shock Nutritional stress Osmotic stress Cold shock … v Stress response in E. coli has been much studied Model for understanding adaptation of pathogenic bacteria to their host 3
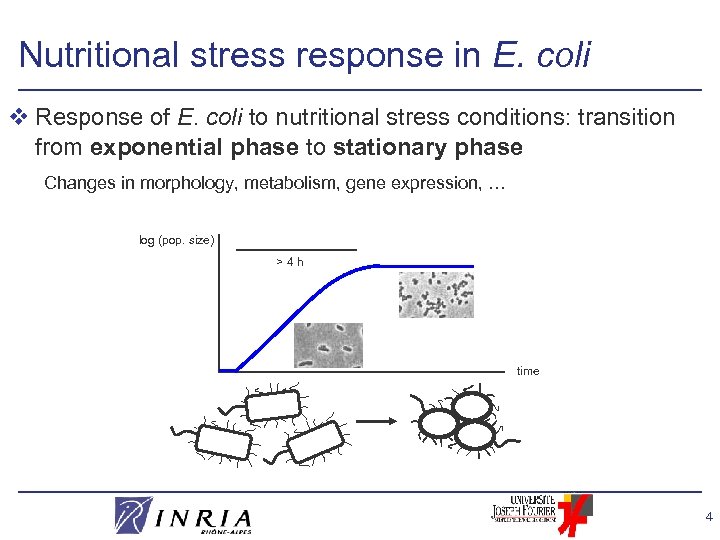
Nutritional stress response in E. coli v Response of E. coli to nutritional stress conditions: transition from exponential phase to stationary phase Changes in morphology, metabolism, gene expression, … log (pop. size) >4 h time 4
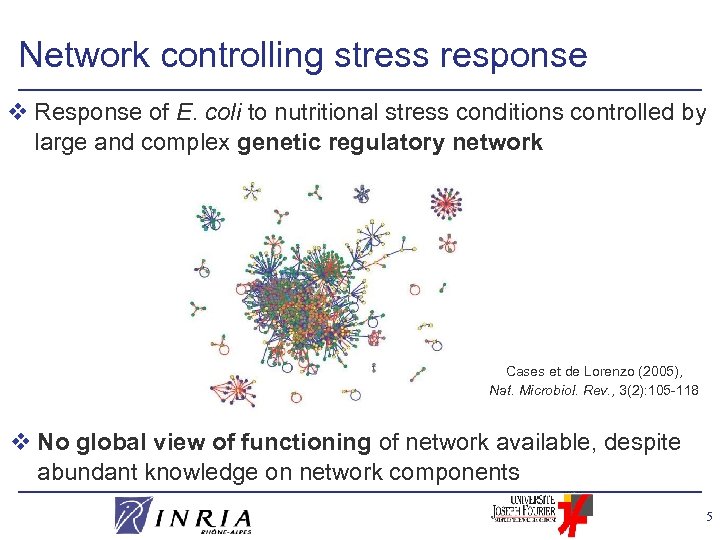

Network controlling stress response v Response of E. coli to nutritional stress conditions controlled by large and complex genetic regulatory network Cases et de Lorenzo (2005), Nat. Microbiol. Rev. , 3(2): 105 -118 v No global view of functioning of network available, despite abundant knowledge on network components 5
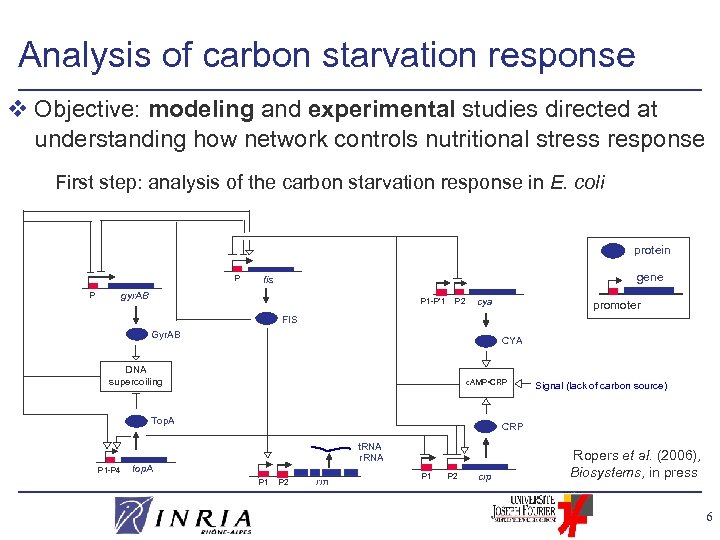
Analysis of carbon starvation response v Objective: modeling and experimental studies directed at understanding how network controls nutritional stress response First step: analysis of the carbon starvation response in E. coli protein P gene fis gyr. AB P P 1 -P’ 1 P 2 cya promoter FIS Gyr. AB CYA DNA supercoiling c. AMP • CRP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp Ropers et al. (2006), Biosystems, in press 6

Qualitative modeling and simulation v Current constraints on modeling and simulation: l l Knowledge on molecular mechanisms rare Quantitative information on kinetic parameters and molecular concentrations absent v Method for qualitative simulation of large and complex genetic regulatory networks using coarse-grained models de Jong, Gouzé et al. (2004), Bull. Math. Biol. , 66(2): 301 -340 Batt G. et al. (2005), Hybrid Systems: Computation and Control, LNCS 3414, 134 -150. v Method used to simulate initiation of sporulation in Bacillus subtilis and quorum sensing of Pseudomonas aeruginosa de Jong et al. (2004), Bull. Math. Biol. , 66(2): 261 -300 Viretta and Fussenegger (2004), Biotechnol. Prog. , 20(3): 670 -8 7
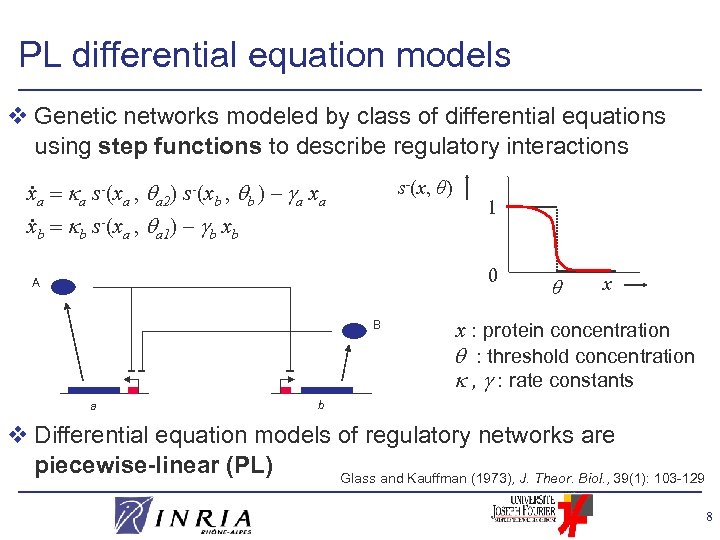

PL differential equation models v Genetic networks modeled by class of differential equations using step functions to describe regulatory interactions . s-(x, θ) xa a s-(xa , a 2) s-(xb , b ) – a xa. xb b s-(xa , a 1) – b xb 1 0 A B a x x : protein concentration : threshold concentration , : rate constants b v Differential equation models of regulatory networks are piecewise-linear (PL) Glass and Kauffman (1973), J. Theor. Biol. , 39(1): 103 -129 8
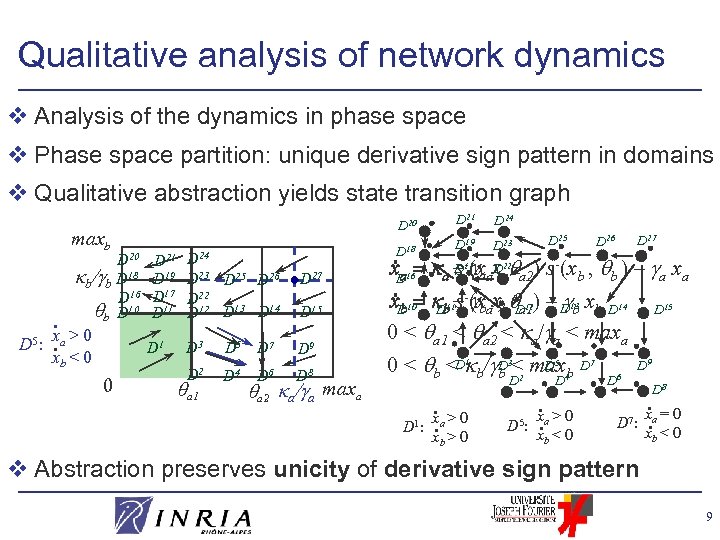

Qualitative analysis of network dynamics v Analysis of the dynamics in phase space v Phase space partition: unique derivative sign pattern in domains v Qualitative abstraction yields state transition graph maxb . D 20 . D 5 : x a > 0. xb < 0 0 D 21 D 19 D 17 D 11 D 24 D 23 D 25 D 26 D 22 D 13 D 14 D 15 D 3 D 5 D 7 D 9 D 2 D 4 D 8 a 1 D 6 D 27 a 2 a/ a maxa D 24 D 25 D 27 D 26 D 19 D 23 D- a D 22 s 17 , a – (xaxa a 2) s (xb , b ) – a xa b s b , a 1 b D 11– (xaxb D 12) – D 13 xb D 14 D 15 . x. D 16 a. x. D 10 b 0 < a 1 < a 2 < a/ a < maxa 1 D 5 0 < b <D b/ D 3< max 4 b D 7 6 b 2 D 18 D 1 D 20 b/ b D 18 D 16 b D 10 D 21 D D . x >0 D 1: . a xb > 0 . x >0 D 5: . a xb < 0 D D 9 D 7: D 8 . xa = 0. xb < 0 v Abstraction preserves unicity of derivative sign pattern 9
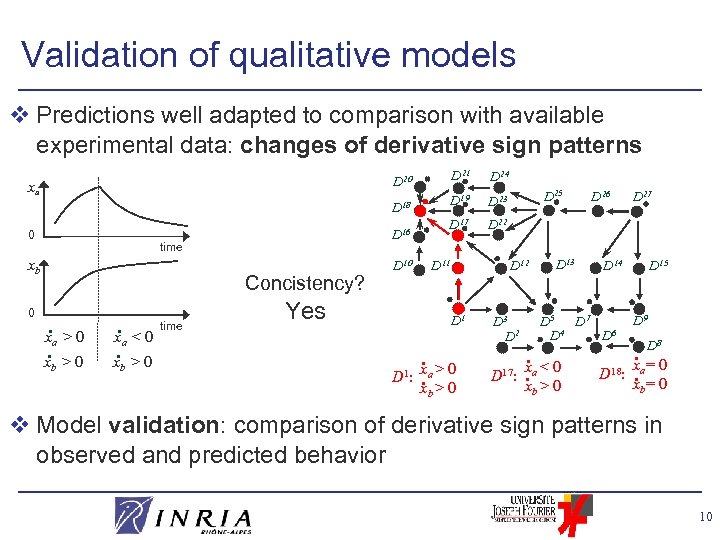

Validation of qualitative models v Predictions well adapted to comparison with available experimental data: changes of derivative sign patterns D 21 D 19 D 23 D 17 D 18 0 D 16 time xb 0 D 24 D 22 D 20 xa Concistency? . x >0 a b . x <0. x >0 a b time D 11 D 10 Yes D 12 D 1 . D 25 D 1: xa > 0. xb > 0 D 3 D 2 . D 13 D 26 D 15 D 14 D 5 D 7 D 4 D 6 x <0 D 17: . a xb > 0 D 27 D 18: D 9 . D 8 xa= 0. xb= 0 v Model validation: comparison of derivative sign patterns in observed and predicted behavior 10
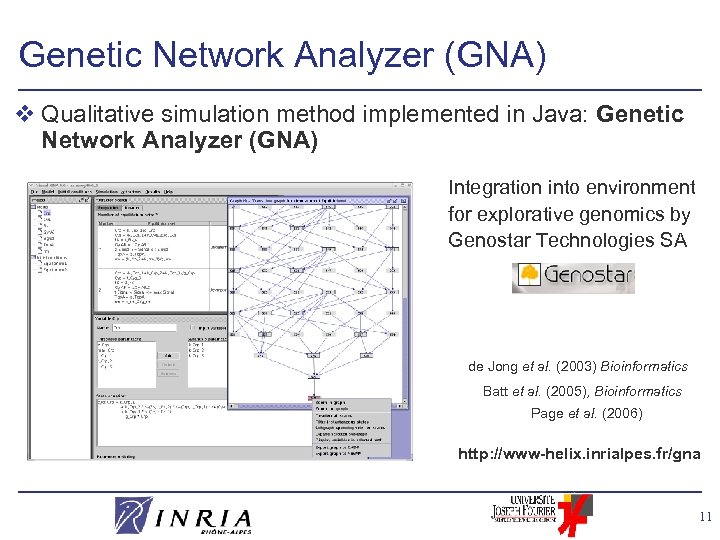

Genetic Network Analyzer (GNA) v Qualitative simulation method implemented in Java: Genetic Network Analyzer (GNA) Integration into environment for explorative genomics by Genostar Technologies SA de Jong et al. (2003) Bioinformatics Batt et al. (2005), Bioinformatics Page et al. (2006) http: //www-helix. inrialpes. fr/gna 11
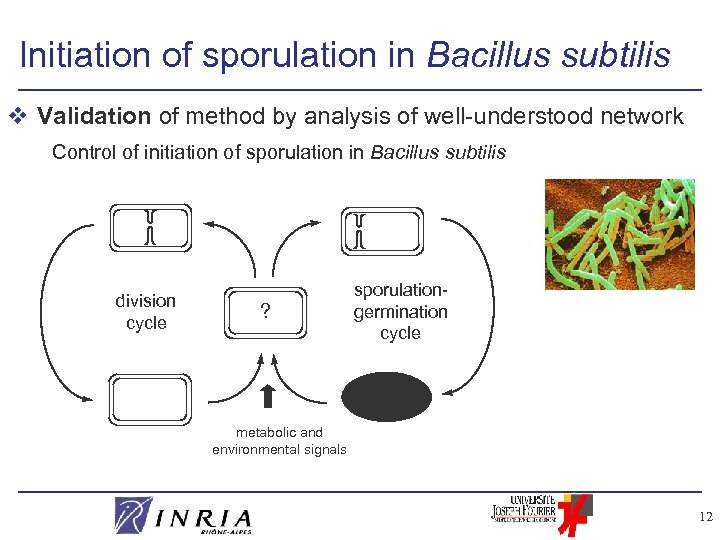

Initiation of sporulation in Bacillus subtilis v Validation of method by analysis of well-understood network Control of initiation of sporulation in Bacillus subtilis division cycle ? sporulationgermination cycle metabolic and environmental signals 12
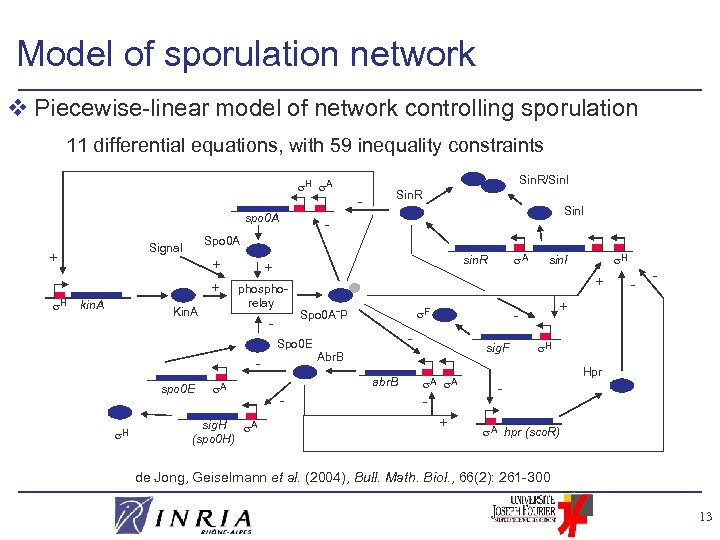
Model of sporulation network v Piecewise-linear model of network controlling sporulation 11 differential equations, with 59 inequality constraints H A spo 0 A H + + kin. A Kin. A Sin. I - phosphorelay - spo 0 E A sig. H A (spo 0 H) A sin. R + - sig. F Abr. B A A + - - + - F Spo 0 A˜P abr. B - H sin. I + Spo 0 E H - Spo 0 A Signal + Sin. R/Sin. I Sin. R H Hpr A hpr (sco. R) de Jong, Geiselmann et al. (2004), Bull. Math. Biol. , 66(2): 261 -300 13
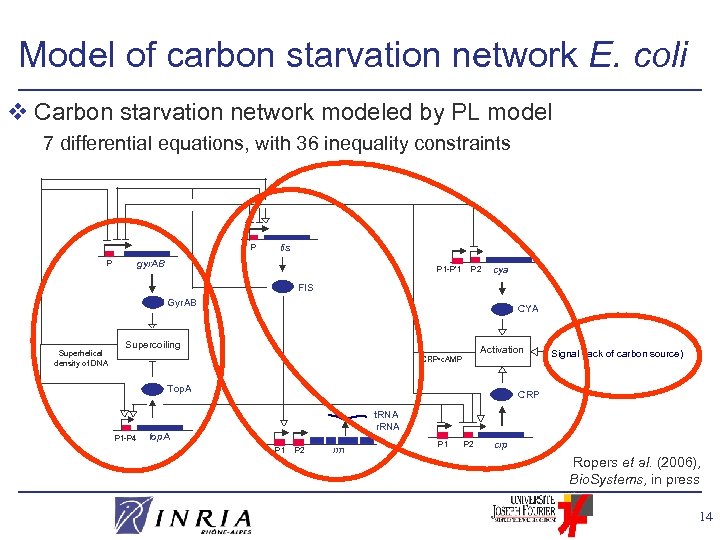
Model of carbon starvation network E. coli v Carbon starvation network modeled by PL model 7 differential equations, with 36 inequality constraints P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB Superhelical density of DNA CYA Supercoiling Activation CRP • c. AMP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp Ropers et al. (2006), Bio. Systems, in press 14
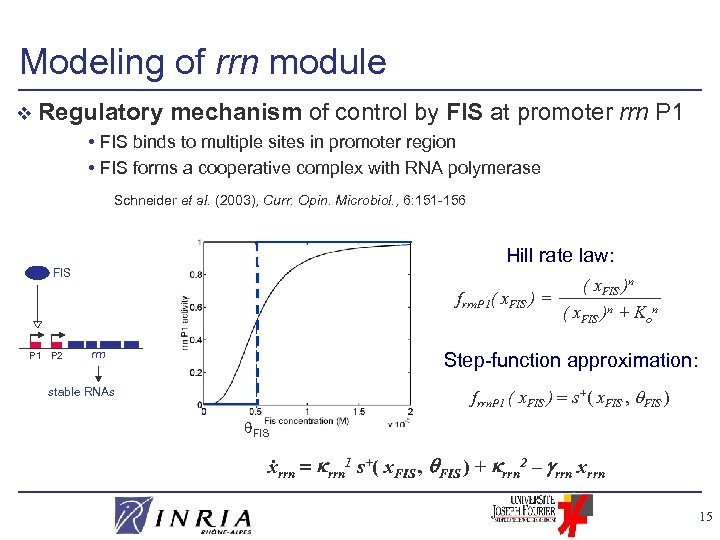
Modeling of rrn module v Regulatory mechanism of control by FIS at promoter • FIS binds to multiple sites in promoter region • FIS forms a cooperative complex with RNA polymerase rrn P 1 Schneider et al. (2003), Curr. Opin. Microbiol. , 6: 151 -156 Hill rate law: FIS frrn. P 1( x. FIS ) = P 1 P 2 rrn ( x. FIS )n + Kon Step-function approximation: frrn. P 1 ( x. FIS ) s+( x. FIS , FIS ) stable RNAs FIS . xrrn 1 s+( x. FIS , FIS ) + rrn 2 – rrn xrrn 15
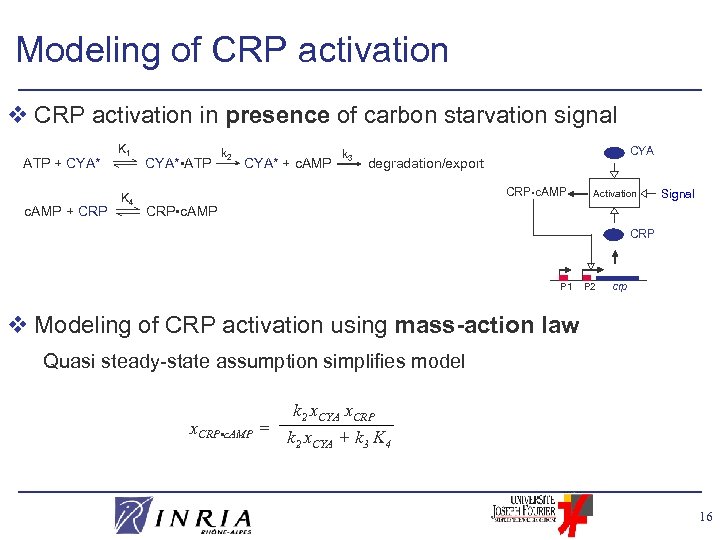

Modeling of CRP activation v CRP activation in presence of carbon starvation signal ATP + CYA* c. AMP + CRP K 1 K 4 CYA* • ATP k 2 CYA* + c. AMP k 3 CYA degradation/export CRP • c. AMP Activation Signal CRP • c. AMP CRP P 1 P 2 crp v Modeling of CRP activation using mass-action law Quasi steady-state assumption simplifies model x. CRP • c. AMP = k 2 x. CYA x. CRP k 2 x. CYA + k 3 K 4 16
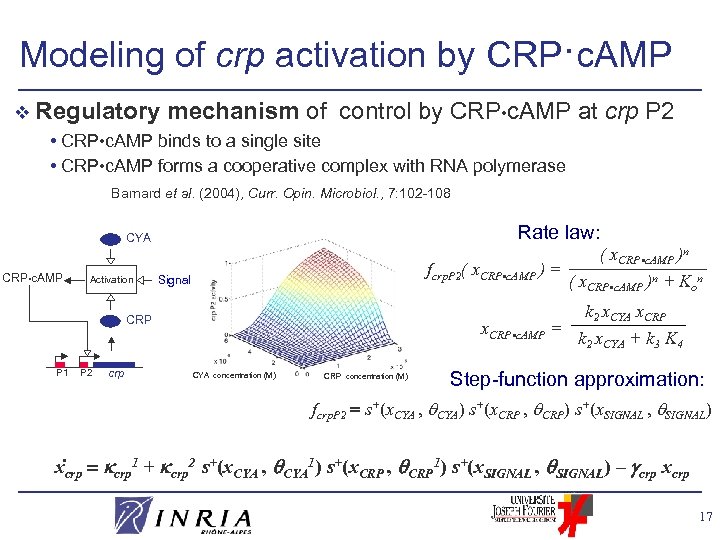
Modeling of crp activation by CRP·c. AMP v Regulatory mechanism of control by CRP • c. AMP • CRP • c. AMP binds to a single site • CRP • c. AMP forms a cooperative complex with RNA polymerase at crp P 2 Barnard et al. (2004), Curr. Opin. Microbiol. , 7: 102 -108 Rate law: CYA CRP • c. AMP Activation ( x. CRP • c. AMP )n fcrp. P 2( x. CRP • c. AMP ) = ( x. CRP • c. AMP )n + Kon Signal CRP P 1 P 2 crp x. CRP • c. AMP = CYA concentration (M) k 2 x. CYA x. CRP k 2 x. CYA + k 3 K 4 Step-function approximation: fcrp. P 2 s+(x. CYA , CYA) s+(x. CRP , CRP) s+(x. SIGNAL , SIGNAL) CRP concentration (M) . xcrp 1 + crp 2 s+(x. CYA , CYA 1) s+(x. CRP , CRP 1) s+(x. SIGNAL , SIGNAL) – crp xcrp 17
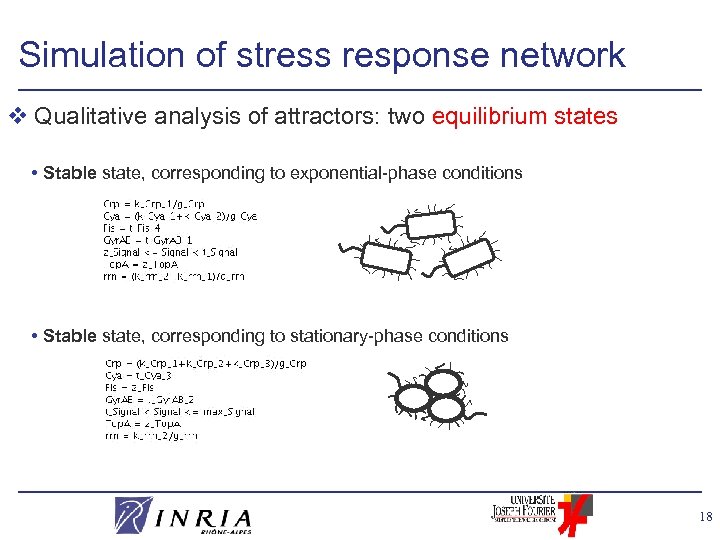

Simulation of stress response network v Qualitative analysis of attractors: two equilibrium states • Stable state, corresponding to exponential-phase conditions • Stable state, corresponding to stationary-phase conditions 18
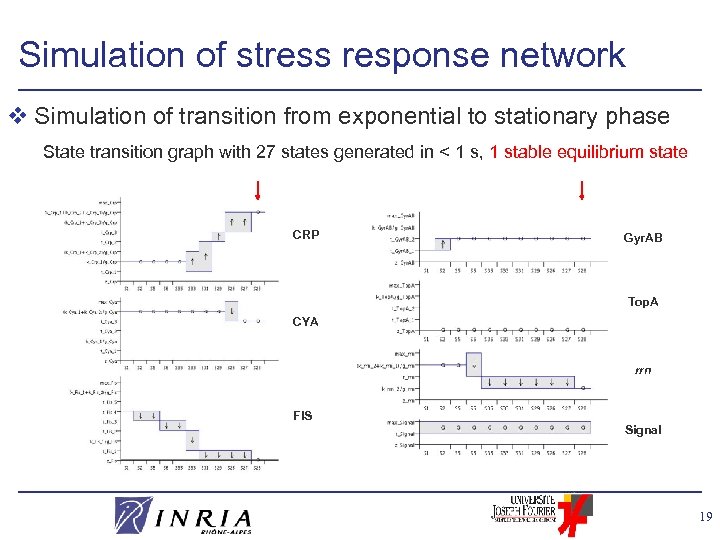

Simulation of stress response network v Simulation of transition from exponential to stationary phase State transition graph with 27 states generated in < 1 s, 1 stable equilibrium state CRP Gyr. AB Top. A CYA rrn FIS Signal 19
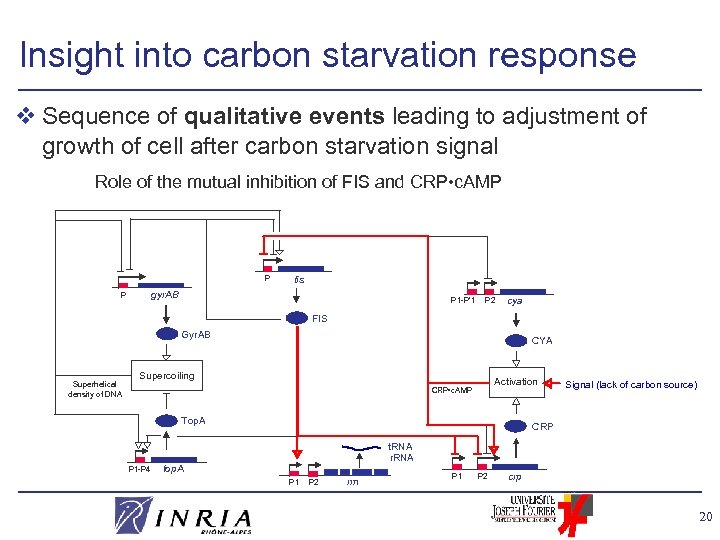
Insight into carbon starvation response v Sequence of qualitative events leading to adjustment of growth of cell after carbon starvation signal Role of the mutual inhibition of FIS and CRP • c. AMP P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB Superhelical density of DNA CYA Supercoiling Activation CRP • c. AMP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp 20
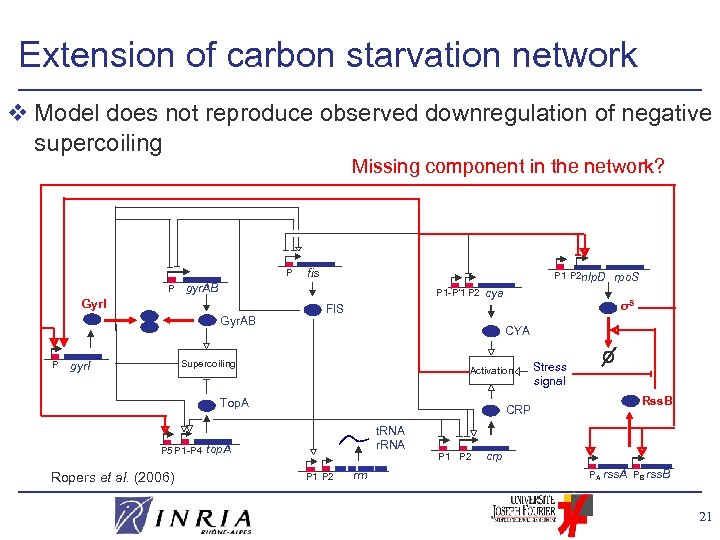
Extension of carbon starvation network v Model does not reproduce observed downregulation of negative supercoiling Missing component in the network? P P P 1 P 2 nlp. D gyr. AB P 1 -P’ 1 P 2 Gyr. I Gyr. AB P fis CYA Activation Top. A P 5 P 1 -P 4 Ropers et al. (2006) σS FIS Supercoiling gyr. I cya rpo. S Stress signal Rss. B CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp PA rss. A PB rss. B 21
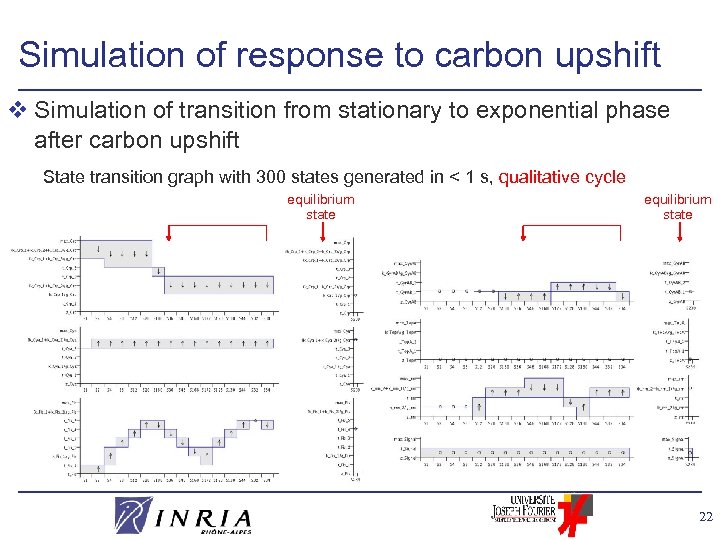
Simulation of response to carbon upshift v Simulation of transition from stationary to exponential phase after carbon upshift State transition graph with 300 states generated in < 1 s, qualitative cycle equilibrium state CRP Gyr. AB CYA Top. A rrn FIS Signal 22
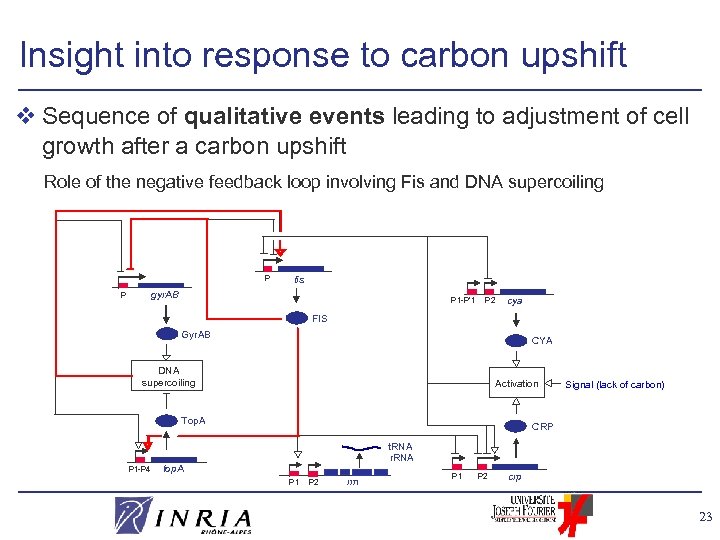
Insight into response to carbon upshift v Sequence of qualitative events leading to adjustment of cell growth after a carbon upshift Role of the negative feedback loop involving Fis and DNA supercoiling P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB CYA DNA supercoiling Activation Top. A P 1 -P 4 Signal (lack of carbon) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp 23
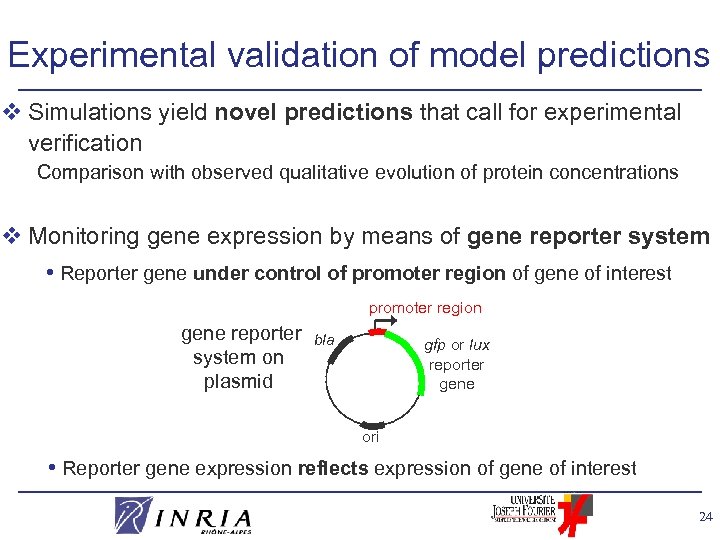

Experimental validation of model predictions v Simulations yield novel predictions that call for experimental verification Comparison with observed qualitative evolution of protein concentrations v Monitoring gene expression by means of gene reporter system • Reporter gene under control of promoter region of gene of interest promoter region gene reporter system on plasmid bla gfp or lux reporter gene ori • Reporter gene expression reflects expression of gene of interest 24
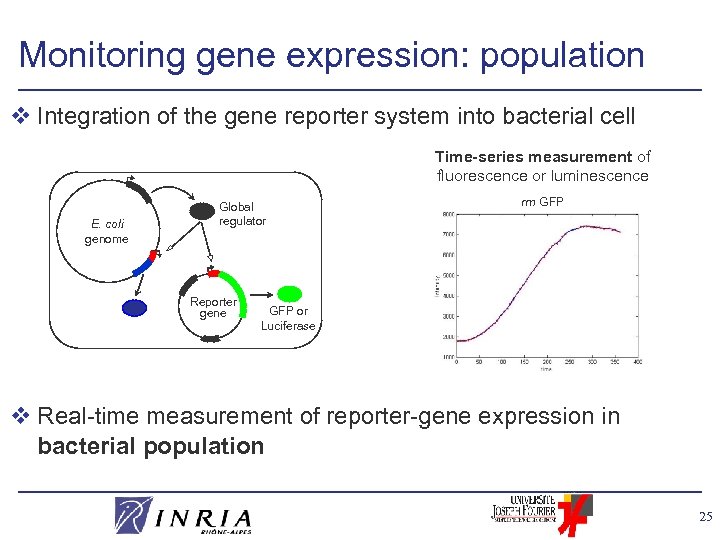
Monitoring gene expression: population v Integration of the gene reporter system into bacterial cell Time-series measurement of fluorescence or luminescence E. coli genome Global regulator Reporter gene rrn GFP or Luciferase v Real-time measurement of reporter-gene expression in bacterial population 25
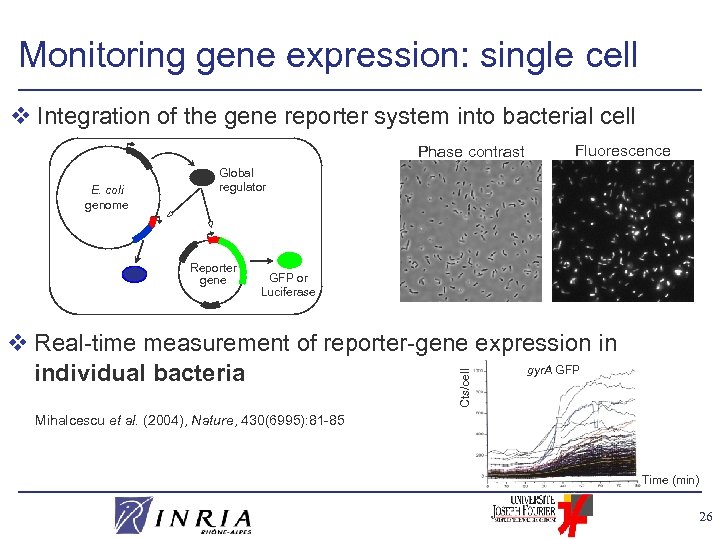
Monitoring gene expression: single cell v Integration of the gene reporter system into bacterial cell Phase contrast E. coli genome Fluorescence Global regulator Reporter gene GFP or Luciferase Cts/cell v Real-time measurement of reporter-gene expression in gyr. A GFP individual bacteria Mihalcescu et al. (2004), Nature, 430(6995): 81 -85 Time (min) 26
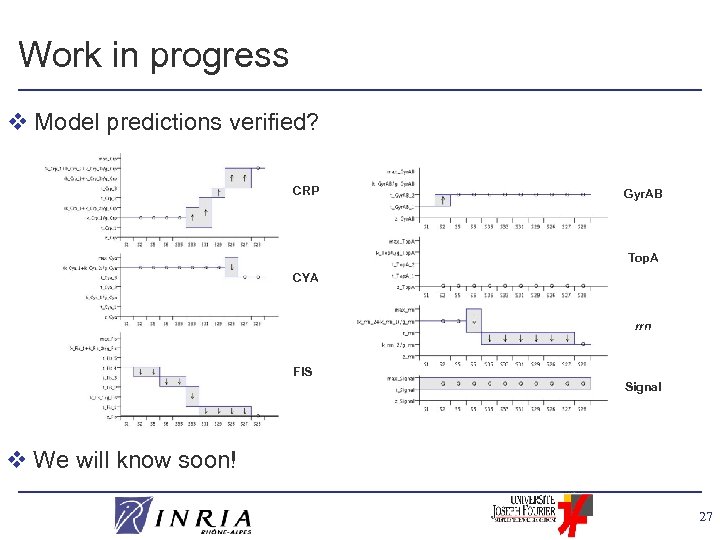

Work in progress v Model predictions verified? CRP Gyr. AB Top. A CYA rrn FIS Signal v We will know soon! 27

Conclusions v Understanding of functioning and development of living organisms requires analysis of genetic regulatory networks From structure to behavior of networks v Need for mathematical methods and computer tools welladapted to available experimental data Coarse-grained models and qualitative analysis of dynamics v Biological relevance attained through integration of modeling and experiments Models guide experiments, and experiments stimulate models 28

Contributors Grégory Batt, INRIA Rhône-Alpes, France Danielle Bonaccio, Université Joseph Fourier, Grenoble, France Hidde de Jong, INRIA Rhône-Alpes, France Hans Geiselmann, Université Joseph Fourier, Grenoble, France Jean-Luc Gouzé, INRIA Sophia-Antipolis, France Irina Mihalcescu, Université Joseph Fourier, Grenoble, France Michel Page, INRIA Rhône-Alpes/Université Pierre Mendès France, Grenoble, France Corinne Pinel, Université Joseph Fourier, Grenoble, France Delphine Ropers, INRIA Rhône-Alpes, France Tewfik Sari Université de Haute Alsace, Mulhouse, France Dominique Schneider Université Joseph Fourier, Grenoble, France 29

30
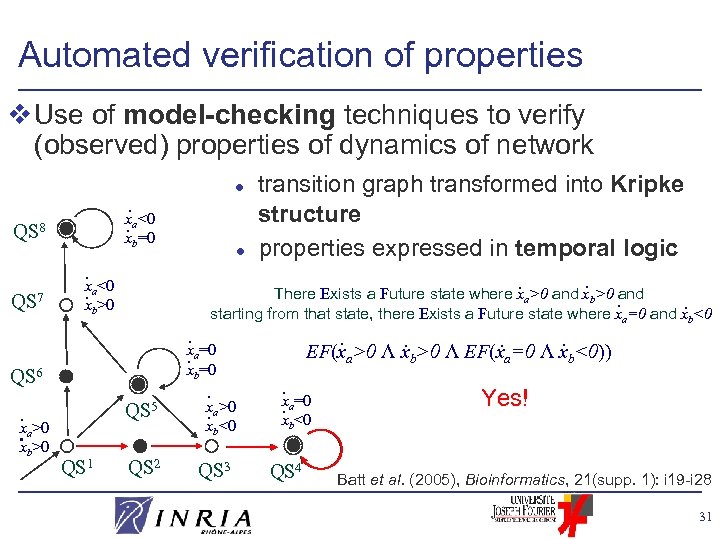

Automated verification of properties v Use of model-checking techniques to verify (observed) properties of dynamics of network. x. a<0 x =0 QS 8 QS 7 b . <0 x. a>0 x b . x. a=0 xb=0 QS 6 . x. a>0 x >0 b transition graph transformed into Kripke structure l properties expressed in temporal logic. . There Exists a Future state where xa>0 and xb>0 and. . starting from that state, there Exists a Future state where x =0 and x <0 l QS 5 . x. a>0 x <0 b QS 1 QS 2 QS 3 . . a b EF(xa>0 Λ xb>0 Λ EF(xa=0 Λ xb<0)) . =0 x. a xb<0 QS 4 Yes! Batt et al. (2005), Bioinformatics, 21(supp. 1): i 19 -i 28 31
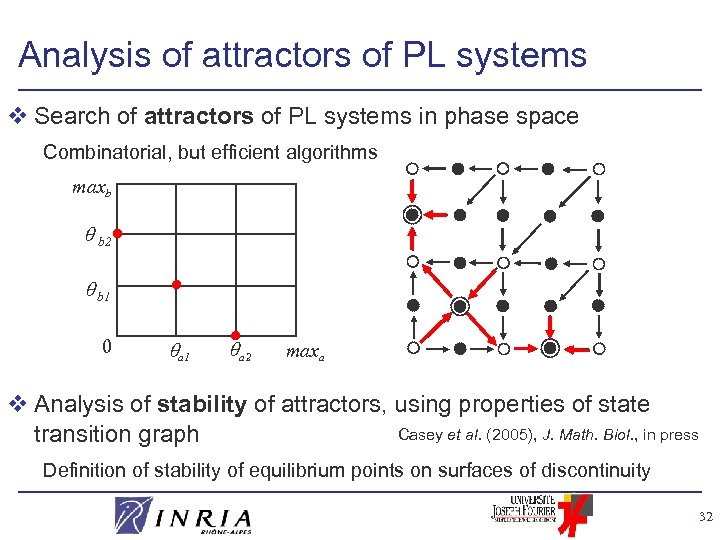

Analysis of attractors of PL systems v Search of attractors of PL systems in phase space Combinatorial, but efficient algorithms maxb b 2 b 1 0 a 1 a 2 maxa v Analysis of stability of attractors, using properties of state Casey et al. (2005), J. Math. Biol. , in press transition graph Definition of stability of equilibrium points on surfaces of discontinuity 32
e4e38ddcc0c771a46369bccc42cd0e05.ppt