70baed108b2bfdd6d1121dcb91d8052f.ppt
- Количество слайдов: 40

Qualitative Simulation of the Carbon Starvation Response in Escherichia coli Delphine Ropers 1 Hidde de Jong 1 Geiselmann 1, 2 1 INRIA Johannes Rhône-Alpes 2 Laboratoire Adaptation Pathogénie des Microorganismes Faculté de Médecine et Pharmacie Université Joseph Fourier CNRS UMR 5163 Email: Delphine. Ropers@inrialpes. fr

Overview 1. Carbon starvation response of Escherichia coli 2. Qualitative modeling, simulation, and analysis of carbon starvation network 3. Experimental validation of carbon starvation model 2
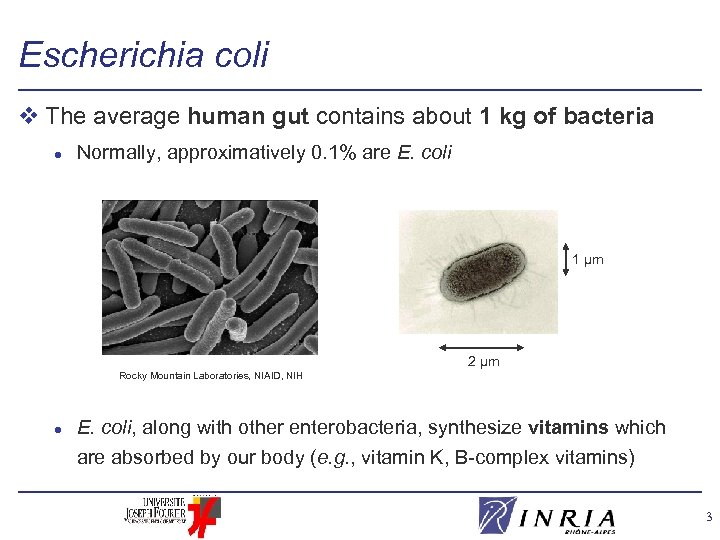
Escherichia coli v The average human gut contains about 1 kg of bacteria l Normally, approximatively 0. 1% are E. coli 1 µm 2 µm Rocky Mountain Laboratories, NIAID, NIH l E. coli, along with other enterobacteria, synthesize vitamins which are absorbed by our body (e. g. , vitamin K, B-complex vitamins) 3

Escherichia coli stress responses v E. coli is able to adapt and respond to a variety of stresses in its environment Model organism for understanding adaptation of pathogenic bacteria to their host Storz and Hengge-Aronis (2000), Bacterial Stress Responses, ASM Press Heat shock Nutritional stress Osmotic stress Cold shock … 4
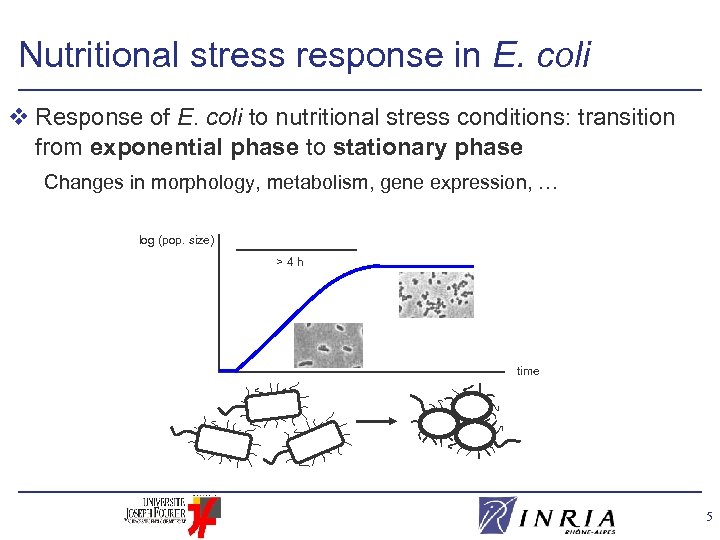
Nutritional stress response in E. coli v Response of E. coli to nutritional stress conditions: transition from exponential phase to stationary phase Changes in morphology, metabolism, gene expression, … log (pop. size) >4 h time 5
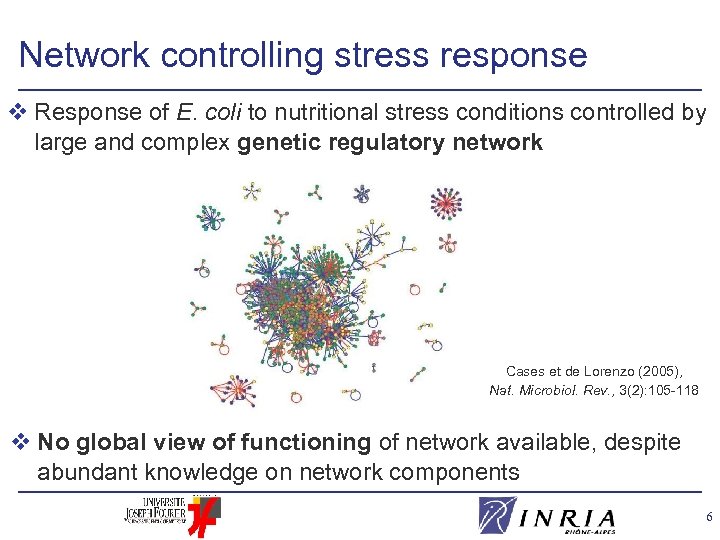

Network controlling stress response v Response of E. coli to nutritional stress conditions controlled by large and complex genetic regulatory network Cases et de Lorenzo (2005), Nat. Microbiol. Rev. , 3(2): 105 -118 v No global view of functioning of network available, despite abundant knowledge on network components 6

Analysis of carbon starvation response v Modeling and experimental studies directed at understanding how network controls carbon starvation response v Which network components and which interactions to take into account? l Impossible to model the whole network E. coli genome: ~4500 genes (~150 transcription factor genes) Start with the simplest possible representation of the carbon starvation response in E. coli l 7
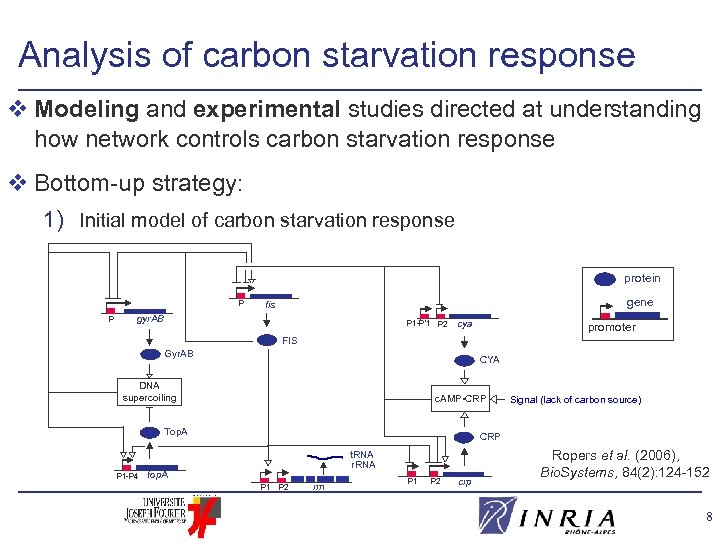
Analysis of carbon starvation response v Modeling and experimental studies directed at understanding how network controls carbon starvation response v Bottom-up strategy: 1) Initial model of carbon starvation response protein P P gene fis gyr. AB P 1 -P’ 1 P 2 cya promoter FIS Gyr. AB CYA DNA supercoiling c. AMP • CRP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp Ropers et al. (2006), Bio. Systems, 84(2): 124 -152 8

Analysis of carbon starvation response v Modeling and experimental studies directed at understanding how network controls carbon starvation response v Bottom-up strategy: 1) Initial model of the carbon starvation response 1. Search and curate data available in the literature and databases 2) Experimental verification of model predictions 3) Extension of model to take into account wrong predictions Additional global regulators: IHF, HNS, pp. Gpp, FNR, LRP, Arc. A, … 9
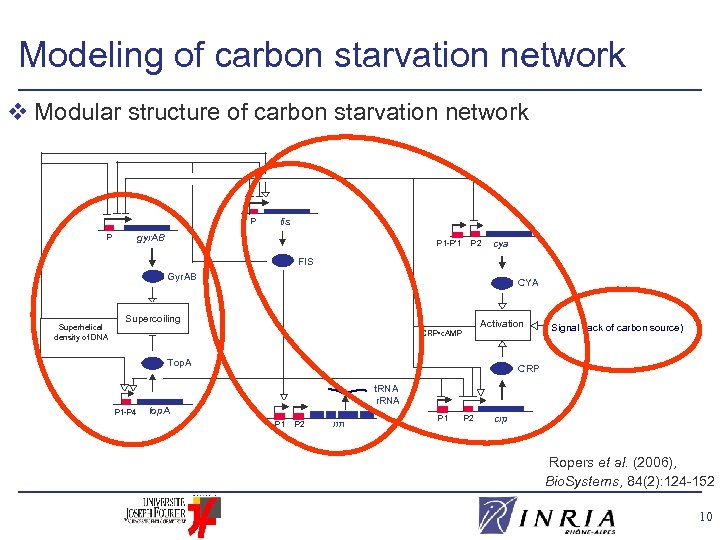
Modeling of carbon starvation network v Modular structure of carbon starvation network P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB Superhelical density of DNA CYA Supercoiling Activation CRP • c. AMP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp Ropers et al. (2006), Bio. Systems, 84(2): 124 -152 10
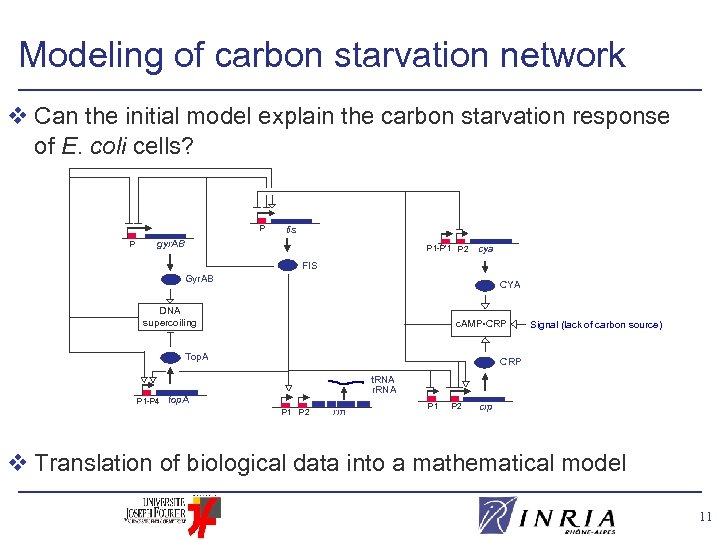
Modeling of carbon starvation network v Can the initial model explain the carbon starvation response of E. coli cells? P P fis gyr. AB P 1 -P’ 1 P 2 cya FIS Gyr. AB CYA DNA supercoiling c. AMP • CRP Top. A P 1 -P 4 Signal (lack of carbon source) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp v Translation of biological data into a mathematical model 11
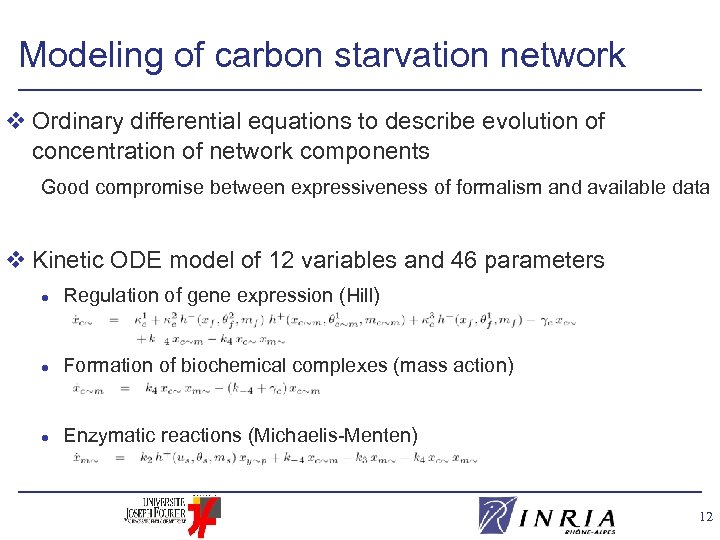

Modeling of carbon starvation network v Ordinary differential equations to describe evolution of concentration of network components Good compromise between expressiveness of formalism and available data v Kinetic ODE model of 12 variables and 46 parameters l Regulation of gene expression (Hill) l Formation of biochemical complexes (mass action) l Enzymatic reactions (Michaelis-Menten) 12
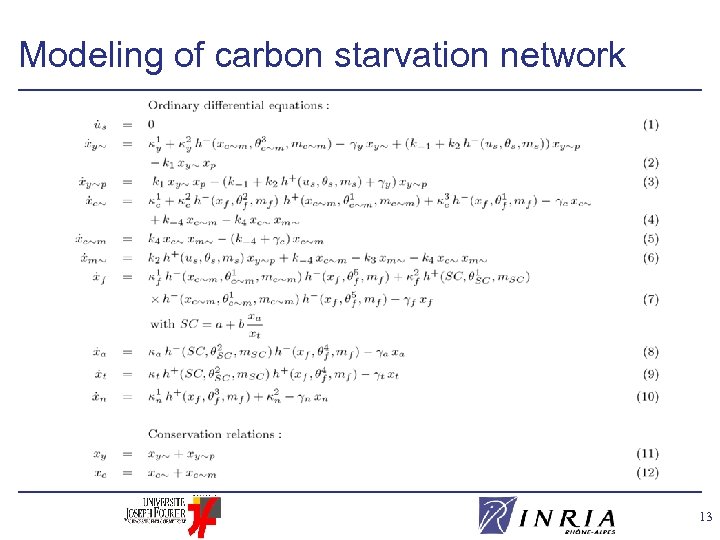

Modeling of carbon starvation network 13

Modeling of carbon starvation network v Current constraints on kinetic modeling of E. coli network: • Knowledge on molecular mechanisms incomplete • Quantitative information on kinetic parameters and molecular concentrations mostly absent v Possible strategies to overcome the constraints • Parameter sensitivity analysis • Model simplifications v Intuition: essential properties of system dynamics robust against moderate changes in kinetic parameters and rate laws 14
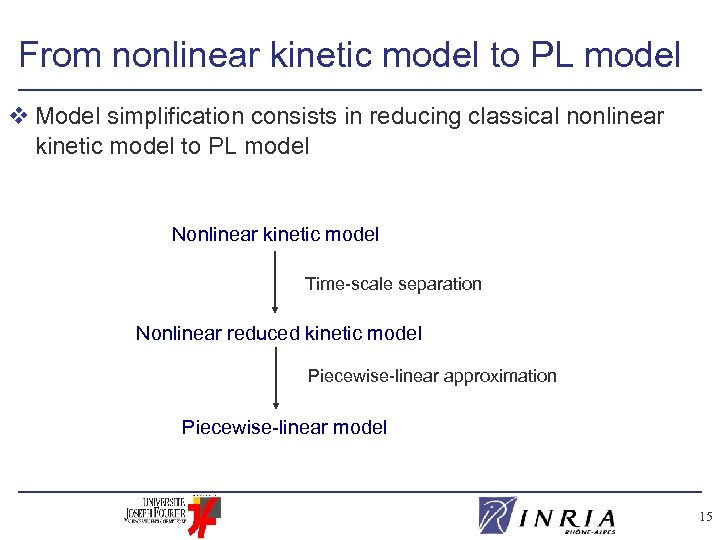

From nonlinear kinetic model to PL model v Model simplification consists in reducing classical nonlinear kinetic model to PL model Nonlinear kinetic model Time-scale separation Nonlinear reduced kinetic model Piecewise-linear approximation Piecewise-linear model 15
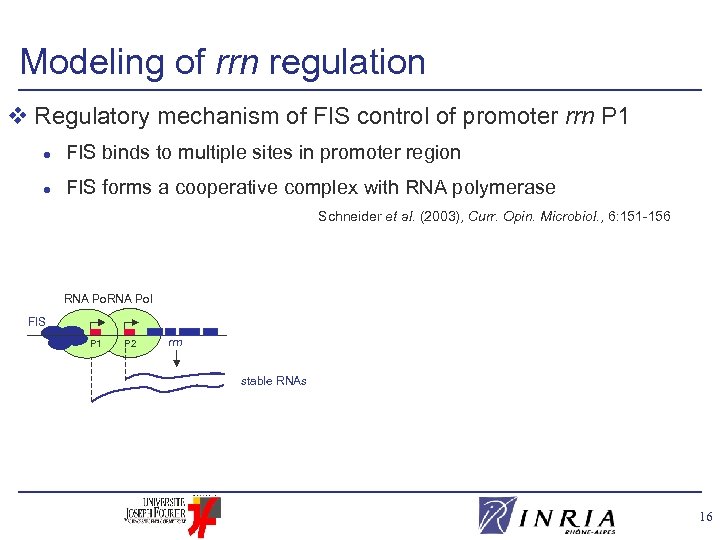
Modeling of rrn regulation v Regulatory mechanism of FIS control of promoter rrn P 1 l FIS binds to multiple sites in promoter region l FIS forms a cooperative complex with RNA polymerase Schneider et al. (2003), Curr. Opin. Microbiol. , 6: 151 -156 RNA Pol FIS P 1 P 2 rrn stable RNAs 16
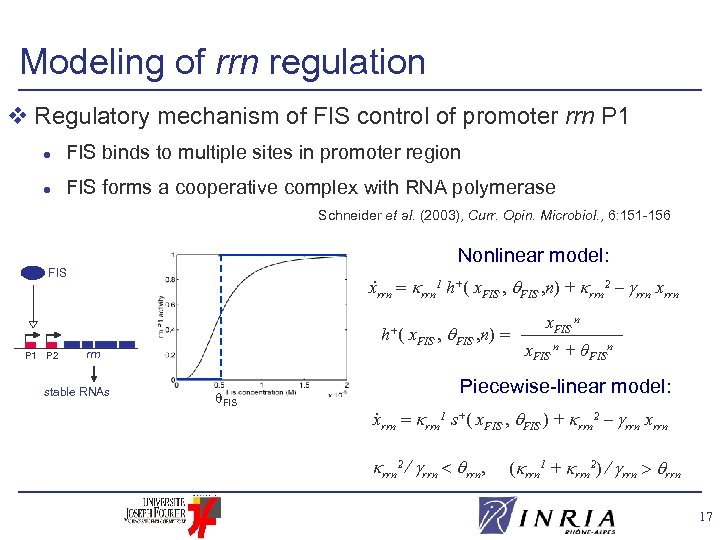

Modeling of rrn regulation v Regulatory mechanism of FIS control of promoter rrn P 1 l FIS binds to multiple sites in promoter region l FIS forms a cooperative complex with RNA polymerase Schneider et al. (2003), Curr. Opin. Microbiol. , 6: 151 -156 Nonlinear model: . FIS xrrn 1 h+( x. FIS , n) + rrn 2 – rrn xrrn h+( P 1 P 2 x. FIS , n) rrn stable RNAs FIS x. FIS n + θFISn Piecewise-linear model: . xrrn 1 s+( x. FIS , FIS ) + rrn 2 – rrn xrrn 2 rrn, ( rrn 1 + rrn 2) rrn 17
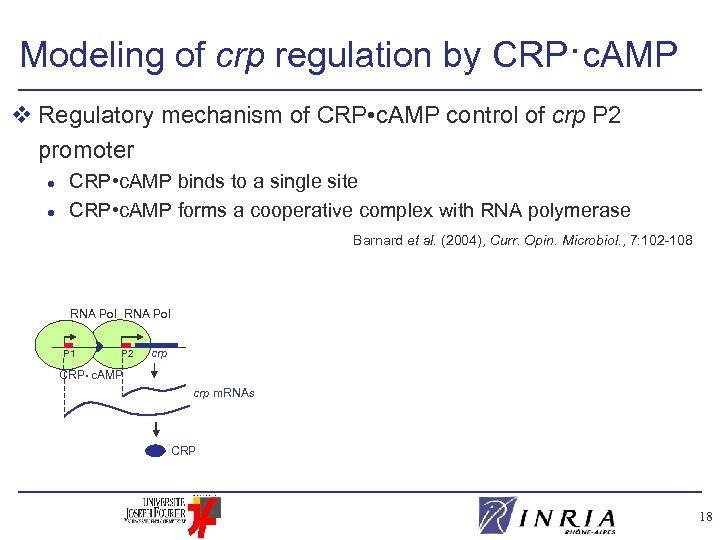

Modeling of crp regulation by CRP·c. AMP v Regulatory mechanism of CRP • c. AMP control of crp P 2 promoter l l CRP • c. AMP binds to a single site CRP • c. AMP forms a cooperative complex with RNA polymerase Barnard et al. (2004), Curr. Opin. Microbiol. , 7: 102 -108 RNA Pol P 1 P 2 crp CRP • c. AMP crp m. RNAs CRP 18
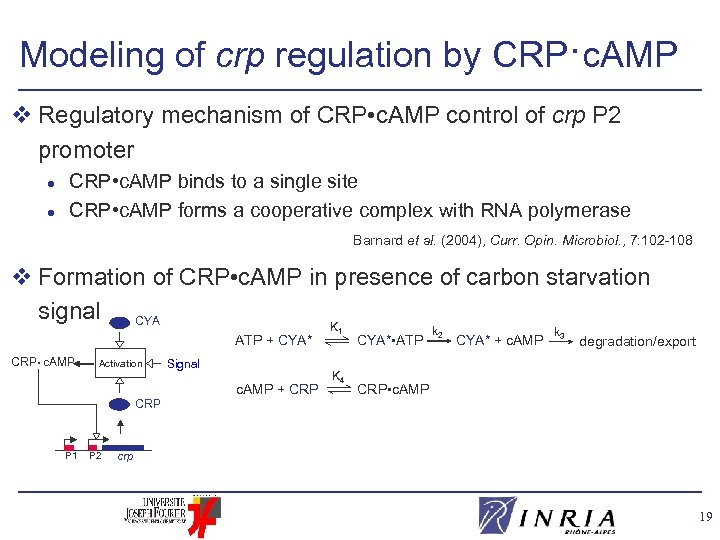

Modeling of crp regulation by CRP·c. AMP v Regulatory mechanism of CRP • c. AMP control of crp P 2 promoter l l CRP • c. AMP binds to a single site CRP • c. AMP forms a cooperative complex with RNA polymerase Barnard et al. (2004), Curr. Opin. Microbiol. , 7: 102 -108 v Formation of CRP • c. AMP in presence of carbon starvation signal CYA K ATP + CYA* CRP • c. AMP Activation Signal c. AMP + CRP P 1 P 2 1 K 4 CYA* • ATP k 2 CYA* + c. AMP k 3 degradation/export CRP • c. AMP crp 19
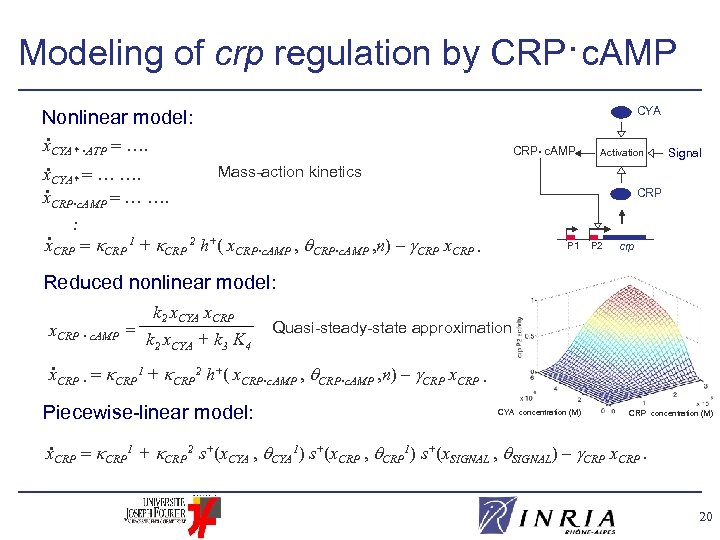
Modeling of crp regulation by CRP·c. AMP CYA Nonlinear model: . x. CYA* ·ATP …. . Mass-action kinetics x. CYA* … …. . x. CRP·c. AMP … …. . x. CRP 1 + CRP 2 h+( x. CRP·c. AMP , n) – CRP x. CRP · CRP • c. AMP Activation Signal CRP P 1 P 2 crp Reduced nonlinear model: x. CRP · c. AMP = k 2 x. CYA x. CRP k 2 x. CYA + k 3 K 4 Quasi-steady-state approximation . x. CRP · CRP 1 + CRP 2 h+( x. CRP·c. AMP , n) – CRP x. CRP · Piecewise-linear model: CYA concentration (M) CRP concentration (M) . x. CRP 1 + CRP 2 s+(x. CYA , CYA 1) s+(x. CRP , CRP 1) s+(x. SIGNAL , SIGNAL) – CRP x. CRP. 20
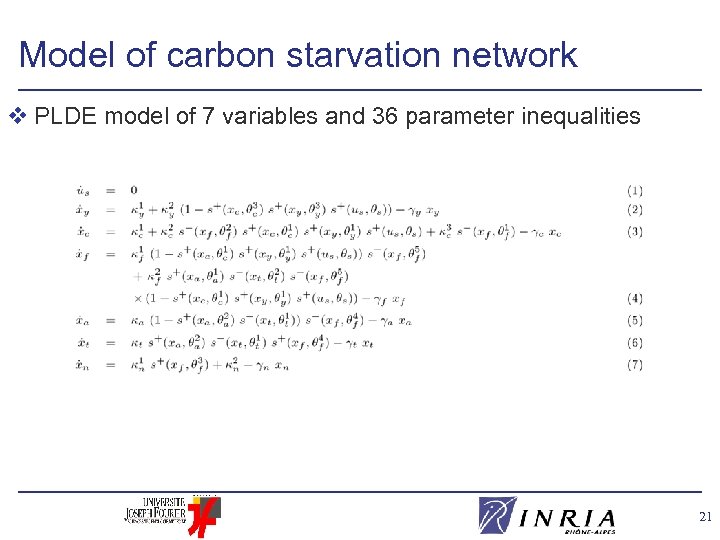

Model of carbon starvation network v PLDE model of 7 variables and 36 parameter inequalities 21
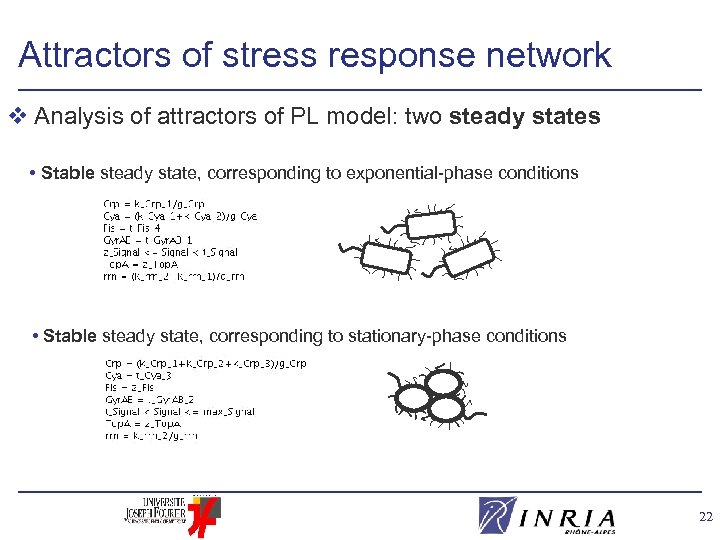

Attractors of stress response network v Analysis of attractors of PL model: two steady states • Stable steady state, corresponding to exponential-phase conditions • Stable steady state, corresponding to stationary-phase conditions 22
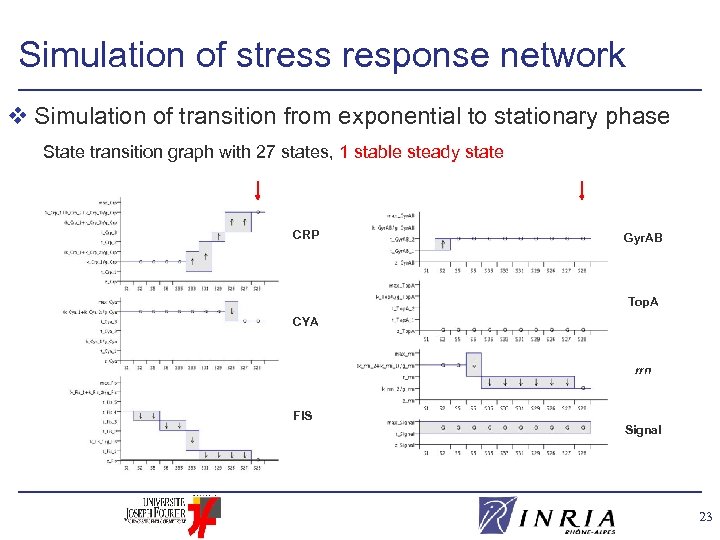

Simulation of stress response network v Simulation of transition from exponential to stationary phase State transition graph with 27 states, 1 stable steady state CRP Gyr. AB Top. A CYA rrn FIS Signal 23
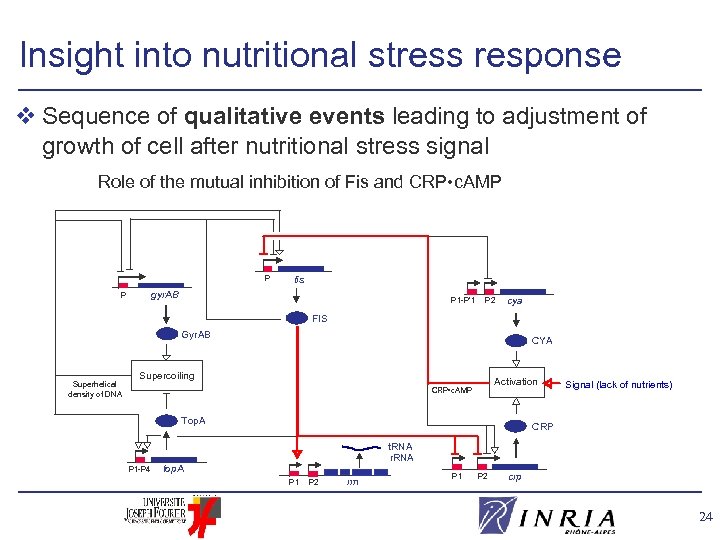
Insight into nutritional stress response v Sequence of qualitative events leading to adjustment of growth of cell after nutritional stress signal Role of the mutual inhibition of Fis and CRP • c. AMP P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB Superhelical density of DNA CYA Supercoiling Activation CRP • c. AMP Top. A P 1 -P 4 Signal (lack of nutrients) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp 24
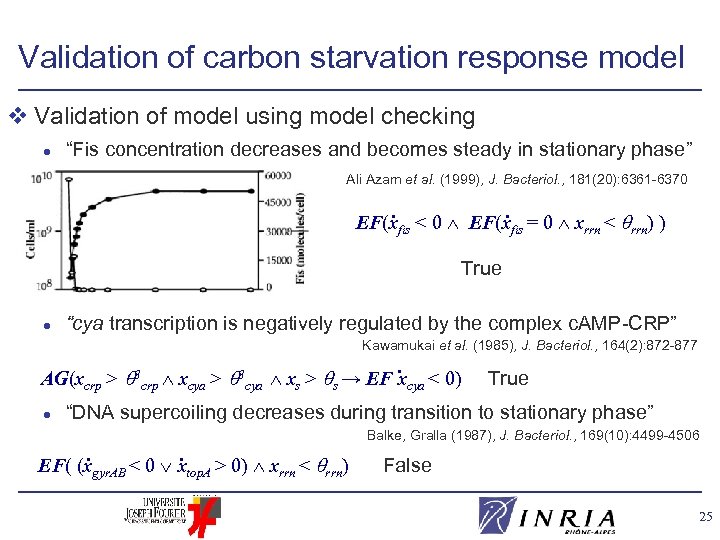
Validation of carbon starvation response model v Validation of model using model checking l “Fis concentration decreases and becomes steady in stationary phase” Ali Azam et al. (1999), J. Bacteriol. , 181(20): 6361 -6370 . . EF(xfis < 0 EF(xfis = 0 xrrn < rrn) ) True l “cya transcription is negatively regulated by the complex c. AMP-CRP” Kawamukai et al. (1985), J. Bacteriol. , 164(2): 872 -877 . AG(xcrp > 3 crp xcya > 3 cya xs > s → EF xcya < 0) l True “DNA supercoiling decreases during transition to stationary phase” . . EF( (xgyr. AB < 0 xtop. A > 0) xrrn < rrn) Balke, Gralla (1987), J. Bacteriol. , 169(10): 4499 -4506 False 25
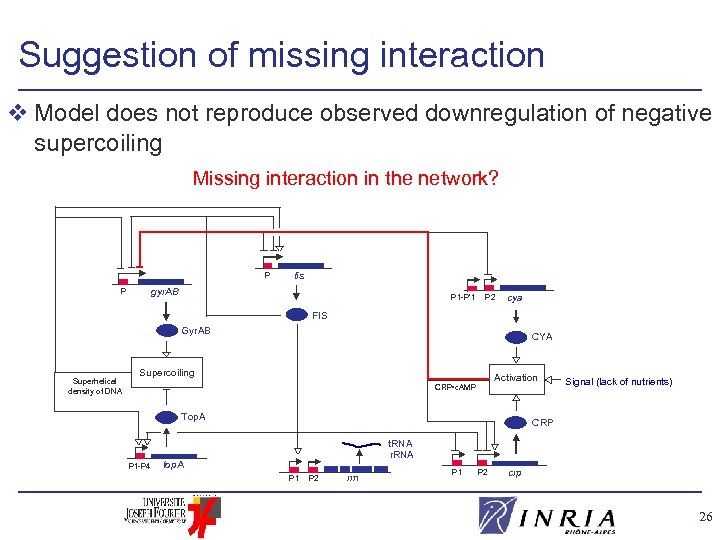
Suggestion of missing interaction v Model does not reproduce observed downregulation of negative supercoiling Missing interaction in the network? P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB Superhelical density of DNA CYA Supercoiling Activation CRP • c. AMP Top. A P 1 -P 4 Signal (lack of nutrients) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp 26
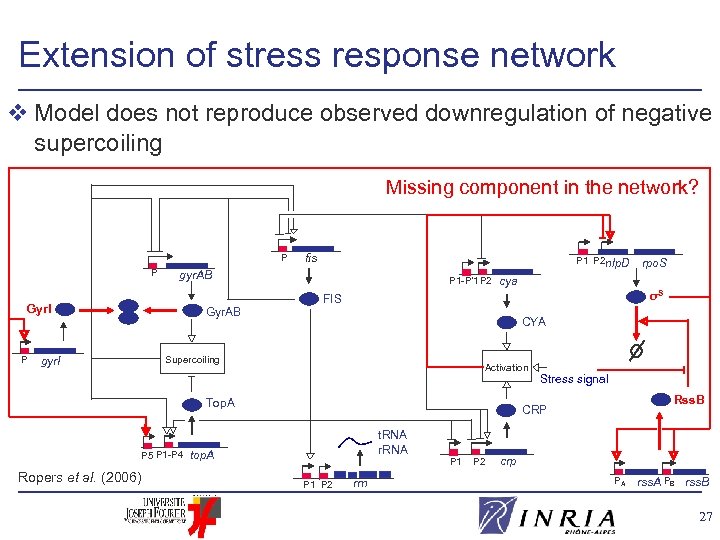
Extension of stress response network v Model does not reproduce observed downregulation of negative supercoiling Missing component in the network? P P P 1 P 2 nlp. D gyr. AB P 1 -P’ 1 P 2 σS Gyr. AB CYA Supercoiling gyr. I Activation Top. A P 5 P 1 -P 4 Ropers et al. (2006) rpo. S cya FIS Gyr. I P fis Stress signal Rss. B CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp PA rss. A PB rss. B 27
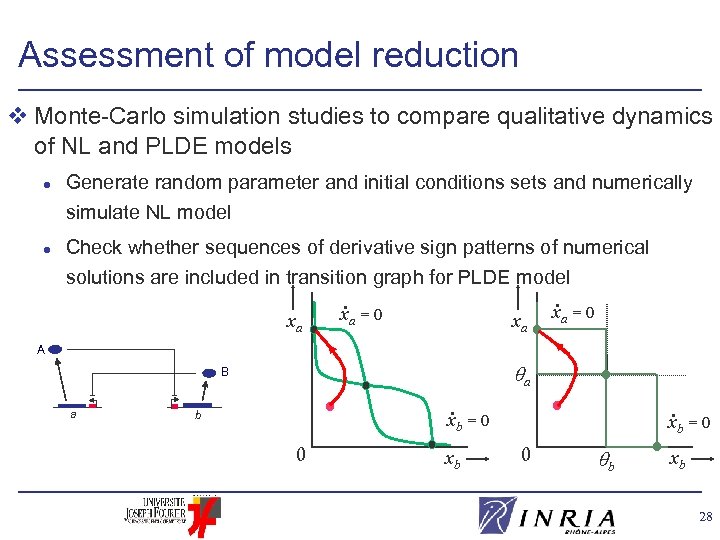
Assessment of model reduction v Monte-Carlo simulation studies to compare qualitative dynamics of NL and PLDE models l l Generate random parameter and initial conditions sets and numerically simulate NL model Check whether sequences of derivative sign patterns of numerical solutions are included in transition graph for PLDE model xa = 0 A a B a . b . xb = 0 0 xb xb = 0 0 b xb 28
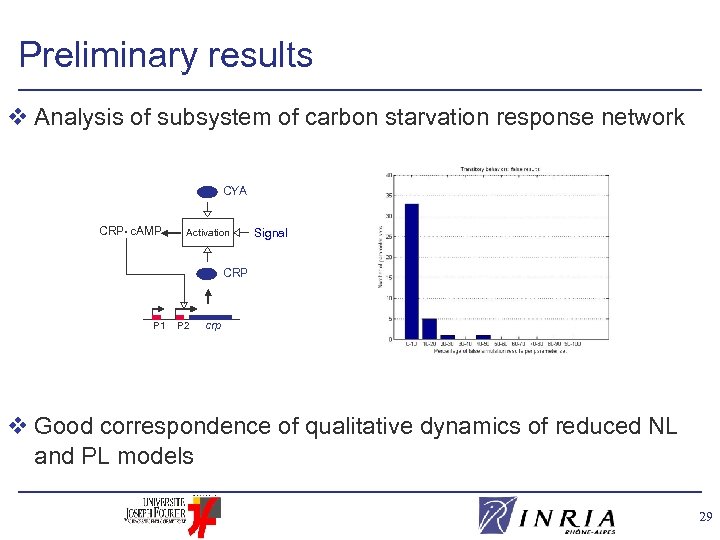
Preliminary results v Analysis of subsystem of carbon starvation response network CYA CRP • c. AMP Activation Signal CRP P 1 P 2 crp v Good correspondence of qualitative dynamics of reduced NL and PL models 29
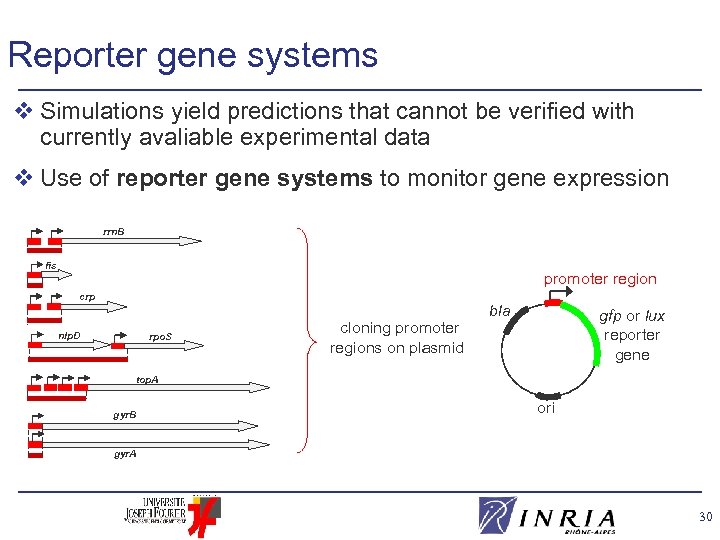
Reporter gene systems v Simulations yield predictions that cannot be verified with currently avaliable experimental data v Use of reporter gene systems to monitor gene expression rrn. B fis promoter region crp nlp. D rpo. S cloning promoter regions on plasmid bla gfp or lux reporter gene top. A gyr. B ori gyr. A 30
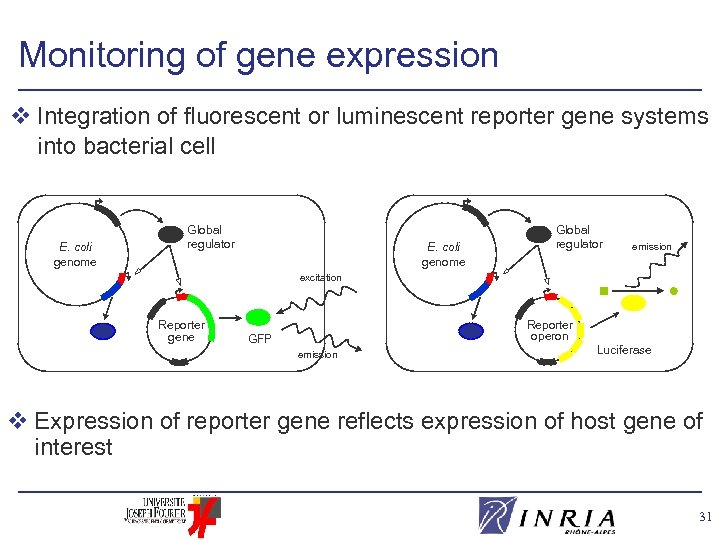
Monitoring of gene expression v Integration of fluorescent or luminescent reporter gene systems into bacterial cell E. coli genome Global regulator emission excitation Reporter gene Reporter operon GFP emission Luciferase v Expression of reporter gene reflects expression of host gene of interest 31
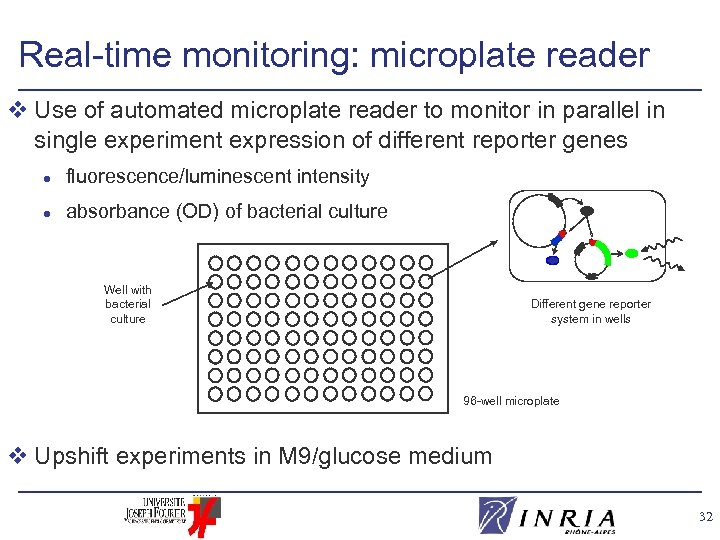

Real-time monitoring: microplate reader v Use of automated microplate reader to monitor in parallel in single experiment expression of different reporter genes l fluorescence/luminescent intensity l absorbance (OD) of bacterial culture Well with bacterial culture Different gene reporter system in wells 96 -well microplate v Upshift experiments in M 9/glucose medium 32
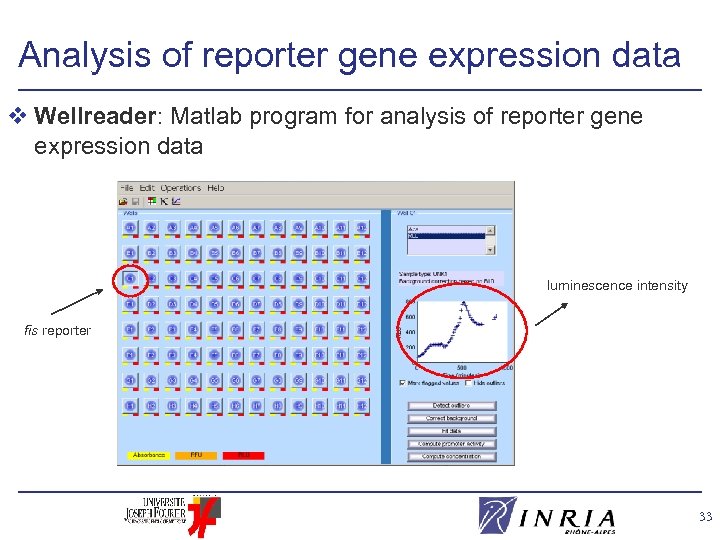
Analysis of reporter gene expression data v Wellreader: Matlab program for analysis of reporter gene expression data luminescence intensity fis reporter 33

Data analysis issues v Outlier detection v Data smoothing and interpolation by means of cubic smoothing splines v Computation of reporter concentration, promoter activity, host protein concentration 34
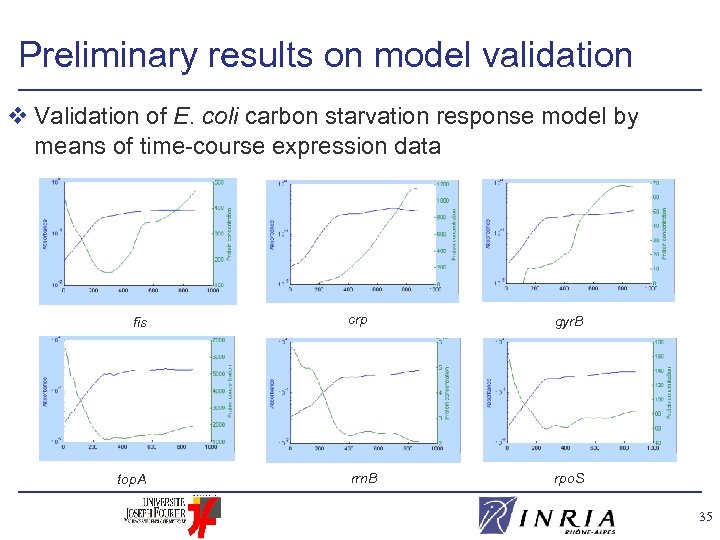
Preliminary results on model validation v Validation of E. coli carbon starvation response model by means of time-course expression data fis top. A crp gyr. B rrn. B rpo. S 35

Conclusions v Understanding of functioning and development of living organisms requires analysis of genetic regulatory networks From structure to behavior of networks v Need for mathematical methods and computer tools welladapted to available experimental data Coarse-grained models and qualitative analysis of dynamics v Biological relevance attained through integration of modeling and experiments Models guide experiments, and experiments stimulate models 36

Further work v Monitoring reporter-gene expression in single cell Collaboration with Irina Mihalcescu (Université Joseph Fourier, Grenoble) v Extensions of carbon starvation model l l Inclusion of additional global regulators involved in carbon starvation response Composite models of E. coli stress response on genetic and metabolic level Collaboration with Daniel Kahn (INRIA Rhône-Alpes, Lyon) 37

Contributors and sponsors Grégory Batt, Boston University, USA Hidde de Jong, INRIA Rhône-Alpes, France Hans Geiselmann, Université Joseph Fourier, Grenoble, France Jean-Luc Gouzé, INRIA Sophia-Antipolis, France Radu Mateescu, INRIA Rhône-Alpes, France Michel Page, INRIA Rhône-Alpes/Université Pierre Mendès France, Grenoble, France Corinne Pinel, Université Joseph Fourier, Grenoble, France Delphine Ropers, INRIA Rhône-Alpes, France Tewfik Sari, Université de Haute Alsace, Mulhouse, France Dominique Schneider, Université Joseph Fourier, Grenoble, France Ministère de la Recherche, IMPBIO program European Commission, FP 6, NEST program INRIA, ARC program Agence Nationale de la Recherche, Bio. Sys program 38

39
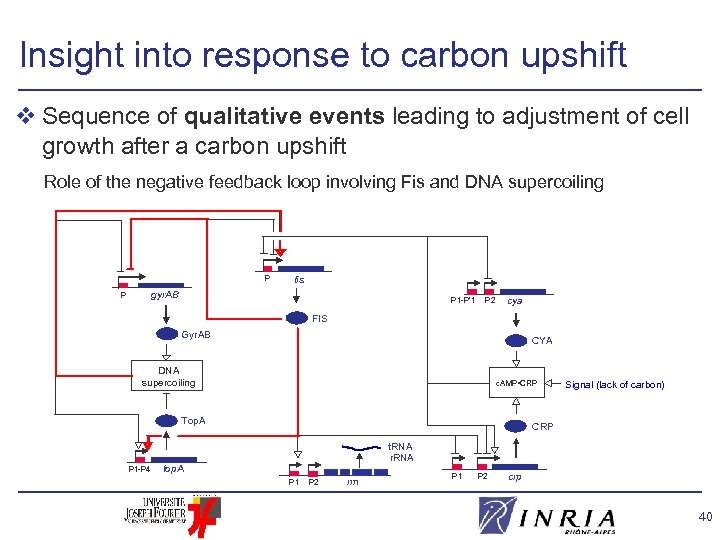
Insight into response to carbon upshift v Sequence of qualitative events leading to adjustment of cell growth after a carbon upshift Role of the negative feedback loop involving Fis and DNA supercoiling P fis gyr. AB P P 1 -P’ 1 P 2 cya FIS Gyr. AB CYA DNA supercoiling c. AMP • CRP Top. A P 1 -P 4 Signal (lack of carbon) CRP t. RNA r. RNA top. A P 1 P 2 rrn P 1 P 2 crp 40
70baed108b2bfdd6d1121dcb91d8052f.ppt