f56371ce56e5f71a45cd80c16e49b43e.ppt
- Количество слайдов: 33
 Qualitative Modeling and Simulation of Genetic Regulatory Networks Hidde de Jong Projet HELIX Institut National de Recherche en Informatique et en Automatique Unité de Recherche Rhône-Alpes 655, avenue de l’Europe Montbonnot, 38334 Saint Ismier CEDEX Email: Hidde. de-Jong@inrialpes. fr
Qualitative Modeling and Simulation of Genetic Regulatory Networks Hidde de Jong Projet HELIX Institut National de Recherche en Informatique et en Automatique Unité de Recherche Rhône-Alpes 655, avenue de l’Europe Montbonnot, 38334 Saint Ismier CEDEX Email: Hidde. de-Jong@inrialpes. fr
 Overview 1. Introduction 2. Modeling and simulation of genetic regulatory networks 3. Genetic Network Analyzer (GNA) 4. Applications Initiation of sporulation in Bacillus subtilis Nutritional stress response in Escherichia coli 5. Validation of models of genetic regulatory networks 6. Conclusions 2
Overview 1. Introduction 2. Modeling and simulation of genetic regulatory networks 3. Genetic Network Analyzer (GNA) 4. Applications Initiation of sporulation in Bacillus subtilis Nutritional stress response in Escherichia coli 5. Validation of models of genetic regulatory networks 6. Conclusions 2
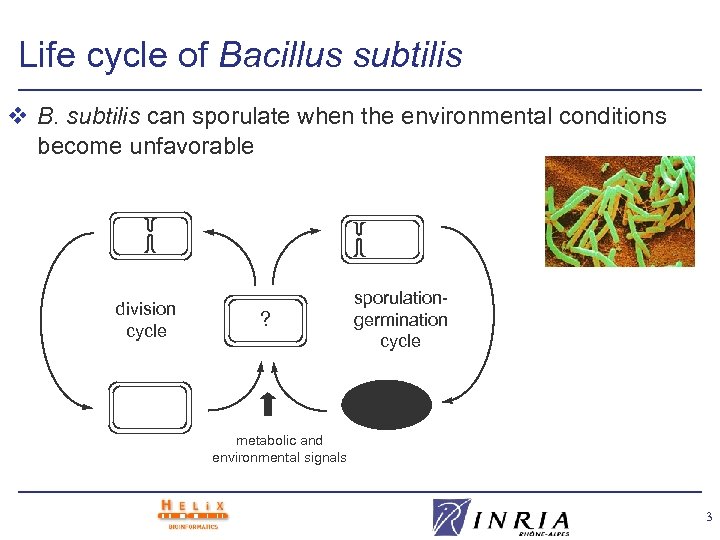 Life cycle of Bacillus subtilis v B. subtilis can sporulate when the environmental conditions become unfavorable division cycle ? sporulationgermination cycle metabolic and environmental signals 3
Life cycle of Bacillus subtilis v B. subtilis can sporulate when the environmental conditions become unfavorable division cycle ? sporulationgermination cycle metabolic and environmental signals 3
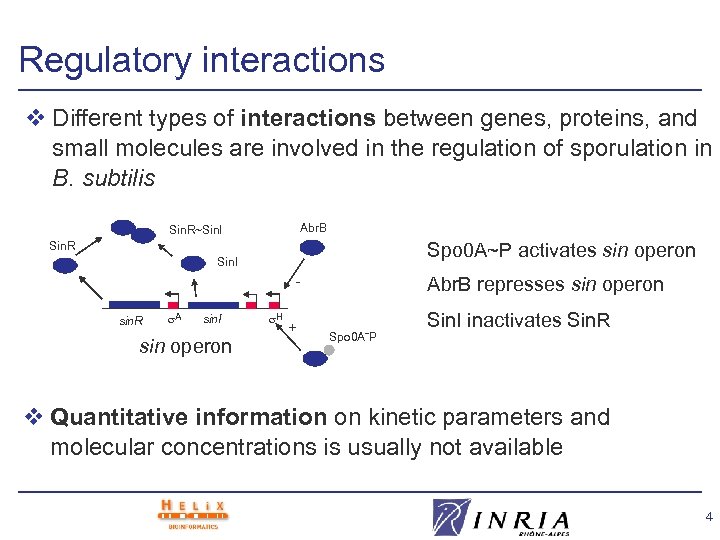 Regulatory interactions v Different types of interactions between genes, proteins, and small molecules are involved in the regulation of sporulation in B. subtilis Abr. B Sin. R~Sin. I Spo 0 A~P activates sin operon Sin. R Sin. I sin. R A sin. I sin operon H + Abr. B represses sin operon Spo 0 A˜P Sin. I inactivates Sin. R v Quantitative information on kinetic parameters and molecular concentrations is usually not available 4
Regulatory interactions v Different types of interactions between genes, proteins, and small molecules are involved in the regulation of sporulation in B. subtilis Abr. B Sin. R~Sin. I Spo 0 A~P activates sin operon Sin. R Sin. I sin. R A sin. I sin operon H + Abr. B represses sin operon Spo 0 A˜P Sin. I inactivates Sin. R v Quantitative information on kinetic parameters and molecular concentrations is usually not available 4
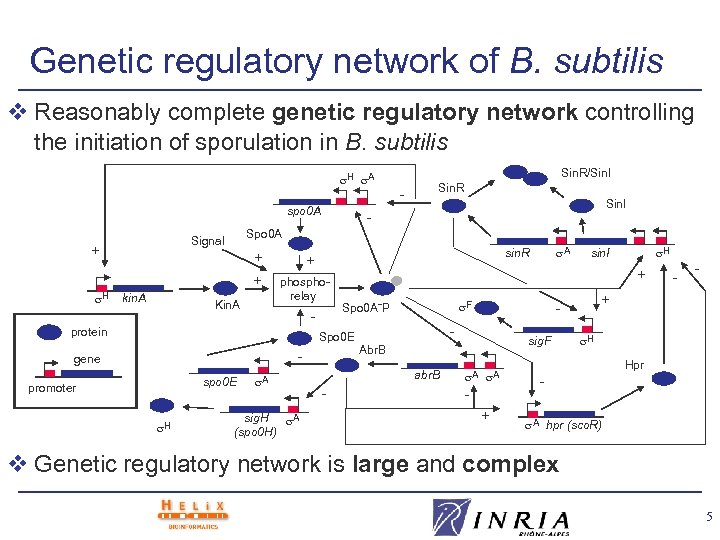 Genetic regulatory network of B. subtilis v Reasonably complete genetic regulatory network controlling the initiation of sporulation in B. subtilis H A spo 0 A H + + kin. A Kin. A - phosphorelay - gene spo 0 E A sig. H A (spo 0 H) A sin. R - sig. F Abr. B A A + - - + - F Spo 0 A˜P abr. B - H sin. I + Spo 0 E H Sin. I + protein promoter - Spo 0 A Signal + Sin. R/Sin. I Sin. R H Hpr A hpr (sco. R) v Genetic regulatory network is large and complex 5
Genetic regulatory network of B. subtilis v Reasonably complete genetic regulatory network controlling the initiation of sporulation in B. subtilis H A spo 0 A H + + kin. A Kin. A - phosphorelay - gene spo 0 E A sig. H A (spo 0 H) A sin. R - sig. F Abr. B A A + - - + - F Spo 0 A˜P abr. B - H sin. I + Spo 0 E H Sin. I + protein promoter - Spo 0 A Signal + Sin. R/Sin. I Sin. R H Hpr A hpr (sco. R) v Genetic regulatory network is large and complex 5
 Qualitative modeling and simulation v Computer support indispensable for dynamical analysis of genetic regulatory networks: modeling and simulation l precise and unambiguous description of network l systematic derivation of behavior predictions v Method for qualitative simulation of large and complex genetic regulatory networks de Jong, Gouzé et al. (2004), Bull. Math. Biol. , 66(2): 301 -340 v Method exploits related work in a variety of domains: l l l mathematical and theoretical biology qualitative reasoning about physical systems control theory and hybrid systems 6
Qualitative modeling and simulation v Computer support indispensable for dynamical analysis of genetic regulatory networks: modeling and simulation l precise and unambiguous description of network l systematic derivation of behavior predictions v Method for qualitative simulation of large and complex genetic regulatory networks de Jong, Gouzé et al. (2004), Bull. Math. Biol. , 66(2): 301 -340 v Method exploits related work in a variety of domains: l l l mathematical and theoretical biology qualitative reasoning about physical systems control theory and hybrid systems 6
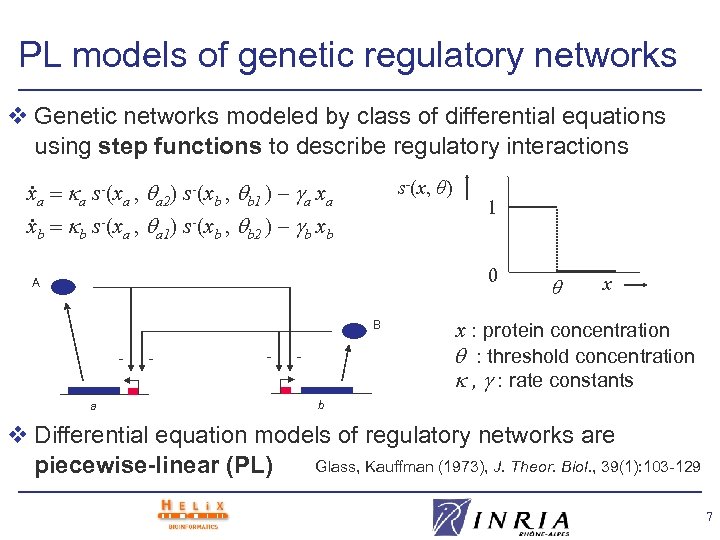 PL models of genetic regulatory networks v Genetic networks modeled by class of differential equations using step functions to describe regulatory interactions . s-(x, θ) xa a s-(xa , a 2) s-(xb , b 1 ) – a xa. xb b s-(xa , a 1) s-(xb , b 2 ) – b xb 1 0 A B a - - - x x : protein concentration : threshold concentration , : rate constants b v Differential equation models of regulatory networks are Glass, Kauffman (1973), J. Theor. Biol. , 39(1): 103 -129 piecewise-linear (PL) 7
PL models of genetic regulatory networks v Genetic networks modeled by class of differential equations using step functions to describe regulatory interactions . s-(x, θ) xa a s-(xa , a 2) s-(xb , b 1 ) – a xa. xb b s-(xa , a 1) s-(xb , b 2 ) – b xb 1 0 A B a - - - x x : protein concentration : threshold concentration , : rate constants b v Differential equation models of regulatory networks are Glass, Kauffman (1973), J. Theor. Biol. , 39(1): 103 -129 piecewise-linear (PL) 7
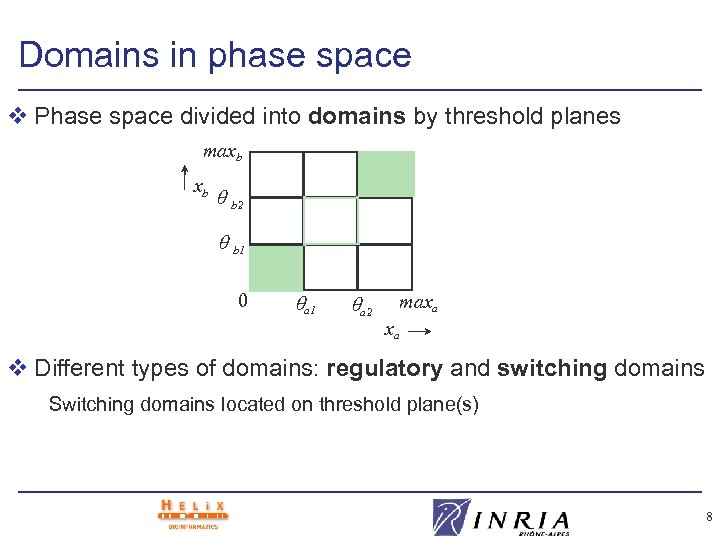 Domains in phase space v Phase space divided into domains by threshold planes maxb xb b 2 b 1 0 . . a 1 a 2 maxa xa v Different types of domains: regulatory and switching domains Switching domains located on threshold plane(s) 8
Domains in phase space v Phase space divided into domains by threshold planes maxb xb b 2 b 1 0 . . a 1 a 2 maxa xa v Different types of domains: regulatory and switching domains Switching domains located on threshold plane(s) 8
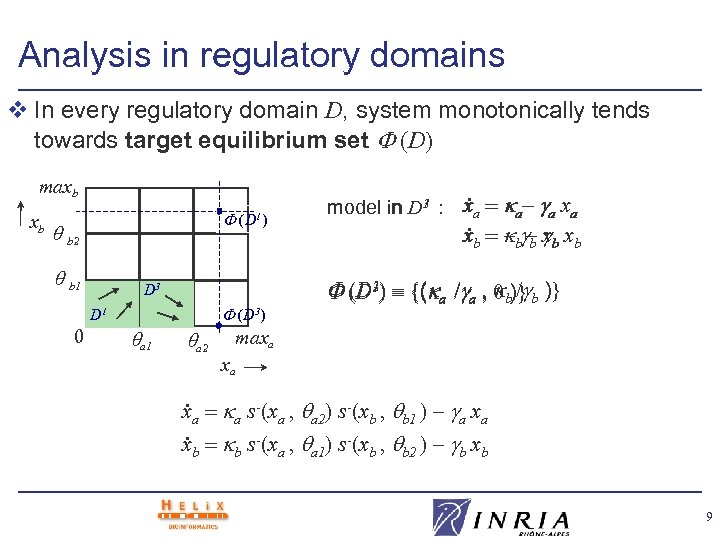 Analysis in regulatory domains v In every regulatory domain D, system monotonically tends towards target equilibrium set (D) maxb xb (D 1) b 2 b b 1 D 3 (D 3) D 1 0 . . x 1 model in D 3 : xa a– a xa a 1 a 2 – b b – xb xb 1 / (D 3) {( a / a , 0 b)} b )} maxa xa a s-(xa , a 2) s-(xb , b 1 ) – a xa. xb b s-(xa , a 1) s-(xb , b 2 ) – b xb 9
Analysis in regulatory domains v In every regulatory domain D, system monotonically tends towards target equilibrium set (D) maxb xb (D 1) b 2 b b 1 D 3 (D 3) D 1 0 . . x 1 model in D 3 : xa a– a xa a 1 a 2 – b b – xb xb 1 / (D 3) {( a / a , 0 b)} b )} maxa xa a s-(xa , a 2) s-(xb , b 1 ) – a xa. xb b s-(xa , a 1) s-(xb , b 2 ) – b xb 9
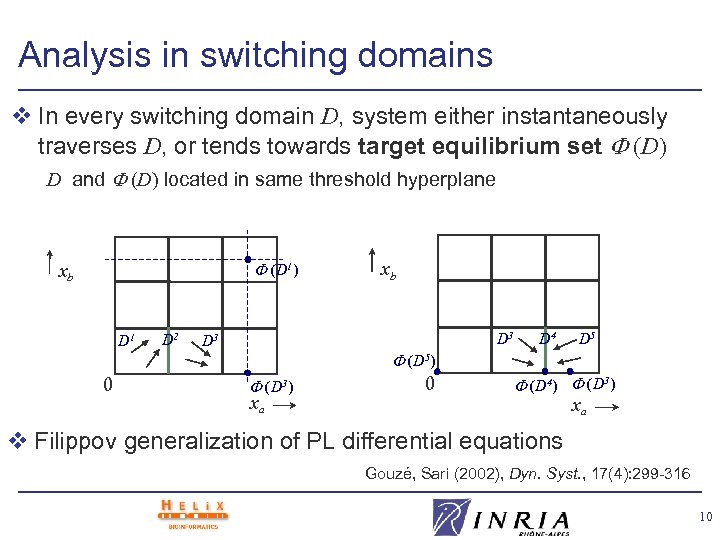 Analysis in switching domains v In every switching domain D, system either instantaneously traverses D, or tends towards target equilibrium set (D) D and (D) located in same threshold hyperplane (D 1) xb D 1 D 2 xb D 3 D 4 D 5 (D 5) 0 (D 3) xa 0 (D 4) (D 3) xa v Filippov generalization of PL differential equations Gouzé, Sari (2002), Dyn. Syst. , 17(4): 299 -316 10
Analysis in switching domains v In every switching domain D, system either instantaneously traverses D, or tends towards target equilibrium set (D) D and (D) located in same threshold hyperplane (D 1) xb D 1 D 2 xb D 3 D 4 D 5 (D 5) 0 (D 3) xa 0 (D 4) (D 3) xa v Filippov generalization of PL differential equations Gouzé, Sari (2002), Dyn. Syst. , 17(4): 299 -316 10
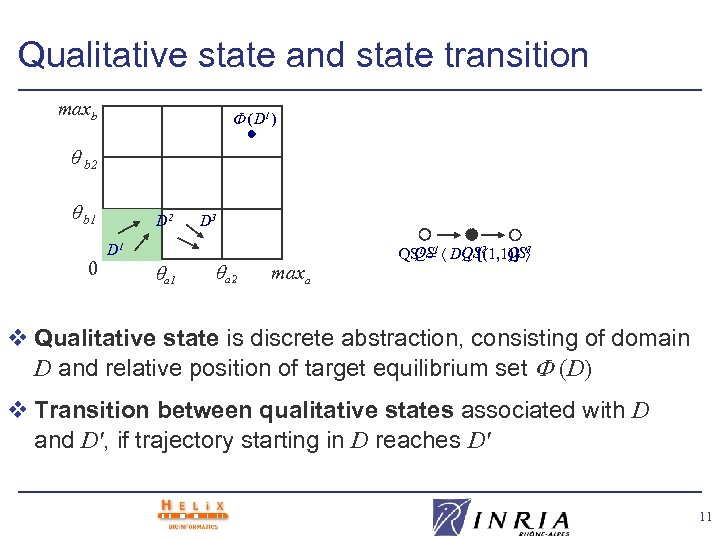 Qualitative state and state transition maxb (D 1) b 2 b 1 0 D 2 D 1 a 1 D 3 a 2 maxa 1, {(1, 1)} 3 QS QS 1 1 DQS 2 QS v Qualitative state is discrete abstraction, consisting of domain D and relative position of target equilibrium set (D) v Transition between qualitative states associated with D and D', if trajectory starting in D reaches D' 11
Qualitative state and state transition maxb (D 1) b 2 b 1 0 D 2 D 1 a 1 D 3 a 2 maxa 1, {(1, 1)} 3 QS QS 1 1 DQS 2 QS v Qualitative state is discrete abstraction, consisting of domain D and relative position of target equilibrium set (D) v Transition between qualitative states associated with D and D', if trajectory starting in D reaches D' 11
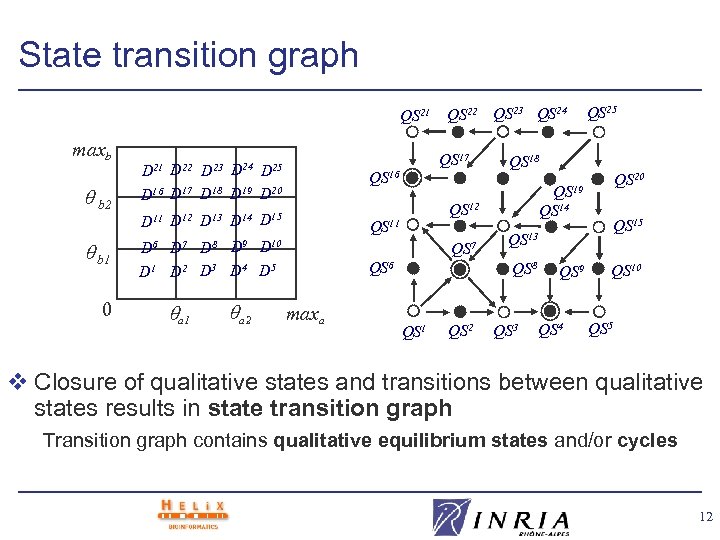 State transition graph QS 21 maxb b 2 D 21 D 22 D 23 D 24 D 25 D 16 D 17 D 18 D 19 D 20 D 15 b 1 0 QS 17 QS 16 QS 11 D 12 D 13 D 14 D 6 D 7 D 8 D 9 D 10 D 1 D 2 D 3 D 4 D 5 a 1 a 2 QS 23 QS 24 QS 6 maxa QS 18 QS 2 QS 15 QS 13 QS 8 QS 1 QS 20 QS 19 QS 14 QS 12 QS 7 QS 25 QS 3 QS 9 QS 4 QS 10 QS 5 v Closure of qualitative states and transitions between qualitative states results in state transition graph Transition graph contains qualitative equilibrium states and/or cycles 12
State transition graph QS 21 maxb b 2 D 21 D 22 D 23 D 24 D 25 D 16 D 17 D 18 D 19 D 20 D 15 b 1 0 QS 17 QS 16 QS 11 D 12 D 13 D 14 D 6 D 7 D 8 D 9 D 10 D 1 D 2 D 3 D 4 D 5 a 1 a 2 QS 23 QS 24 QS 6 maxa QS 18 QS 2 QS 15 QS 13 QS 8 QS 1 QS 20 QS 19 QS 14 QS 12 QS 7 QS 25 QS 3 QS 9 QS 4 QS 10 QS 5 v Closure of qualitative states and transitions between qualitative states results in state transition graph Transition graph contains qualitative equilibrium states and/or cycles 12
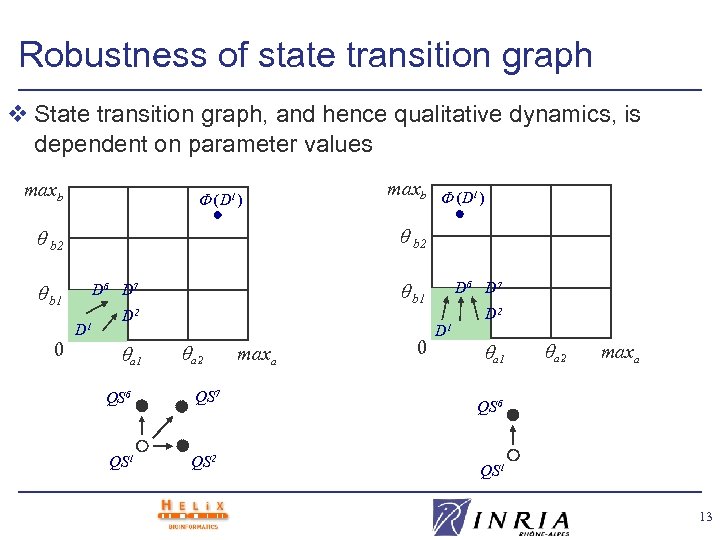 Robustness of state transition graph v State transition graph, and hence qualitative dynamics, is dependent on parameter values maxb (D 1) b 2 b 1 D 6 D 7 D 1 0 maxb (D 1) D 2 a 1 a 2 QS 6 QS 7 QS 1 QS 2 maxa 0 D 6 D 7 D 1 D 2 a 1 a 2 maxa QS 6 QS 1 13
Robustness of state transition graph v State transition graph, and hence qualitative dynamics, is dependent on parameter values maxb (D 1) b 2 b 1 D 6 D 7 D 1 0 maxb (D 1) D 2 a 1 a 2 QS 6 QS 7 QS 1 QS 2 maxa 0 D 6 D 7 D 1 D 2 a 1 a 2 maxa QS 6 QS 1 13
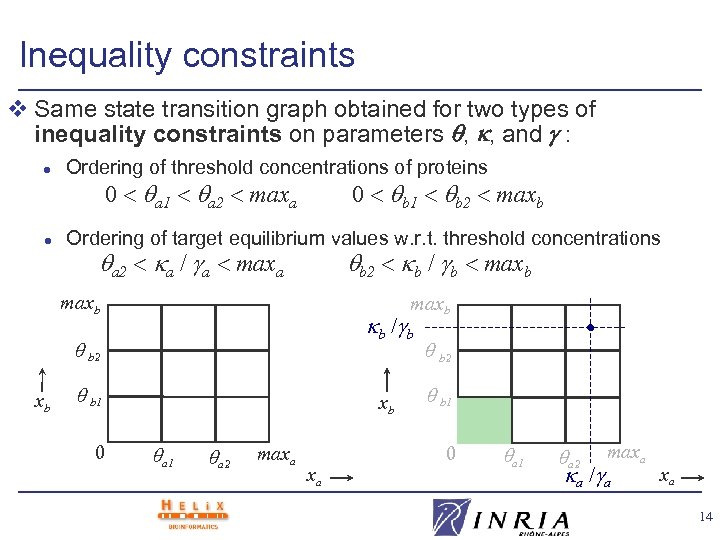 Inequality constraints v Same state transition graph obtained for two types of inequality constraints on parameters , , and : l Ordering of threshold concentrations of proteins l Ordering of target equilibrium values w. r. t. threshold concentrations 0 < a 1 < a 2 < maxa 0 < b 1 < b 2 < maxb a 2 < a / a < maxa b 2 < b / b < maxb b / b b 2 xb b 1 0 xb a 1 a 2 maxa xa b 2 b 1 0 a 1 a 2 maxa a / a xa 14
Inequality constraints v Same state transition graph obtained for two types of inequality constraints on parameters , , and : l Ordering of threshold concentrations of proteins l Ordering of target equilibrium values w. r. t. threshold concentrations 0 < a 1 < a 2 < maxa 0 < b 1 < b 2 < maxb a 2 < a / a < maxa b 2 < b / b < maxb b / b b 2 xb b 1 0 xb a 1 a 2 maxa xa b 2 b 1 0 a 1 a 2 maxa a / a xa 14
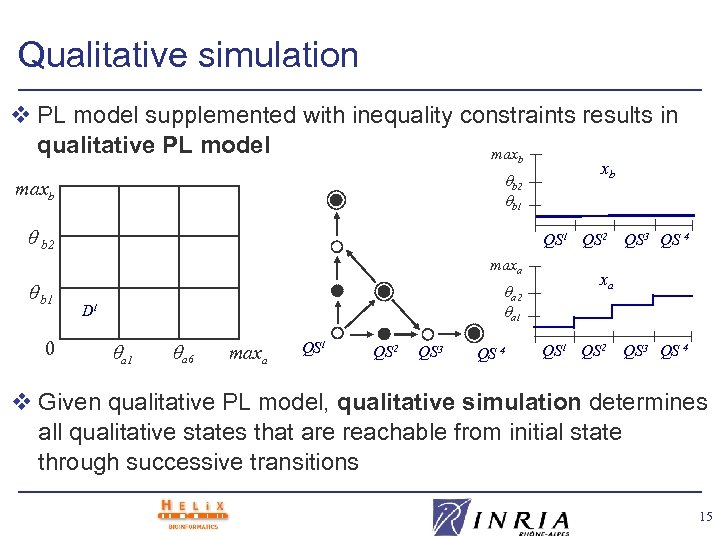 Qualitative simulation v PL model supplemented with inequality constraints results in qualitative PL model maxb b 2 b 1 maxb b 2 QS 1 QS 2 maxa b 1 0 xb a 2 a 1 D 1 a 6 maxa QS 1 QS 2 QS 3 QS 4 v Given qualitative PL model, qualitative simulation determines all qualitative states that are reachable from initial state through successive transitions 15
Qualitative simulation v PL model supplemented with inequality constraints results in qualitative PL model maxb b 2 b 1 maxb b 2 QS 1 QS 2 maxa b 1 0 xb a 2 a 1 D 1 a 6 maxa QS 1 QS 2 QS 3 QS 4 v Given qualitative PL model, qualitative simulation determines all qualitative states that are reachable from initial state through successive transitions 15
 Genetic Network Analyzer (GNA) v Qualitative simulation method implemented in Java 1. 4: Genetic Network Analyzer (GNA) Graphical interface to control simulation and analyze results de Jong et al. (2003), Bioinformatics, 19(3): 336 -344 16
Genetic Network Analyzer (GNA) v Qualitative simulation method implemented in Java 1. 4: Genetic Network Analyzer (GNA) Graphical interface to control simulation and analyze results de Jong et al. (2003), Bioinformatics, 19(3): 336 -344 16
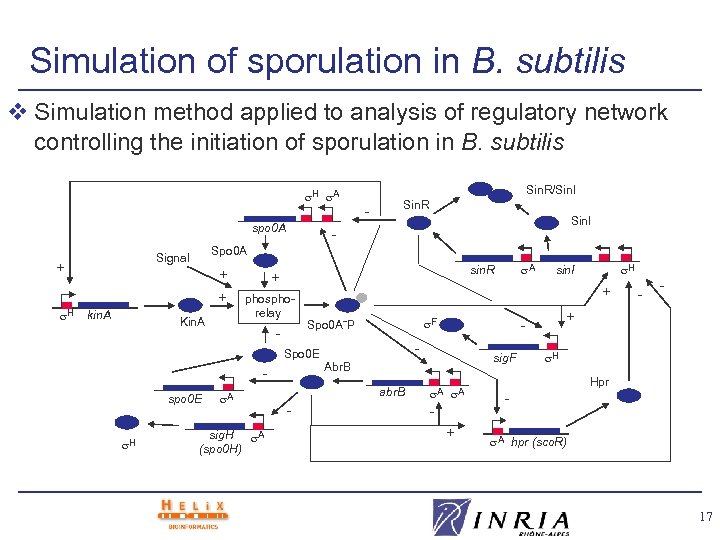 Simulation of sporulation in B. subtilis v Simulation method applied to analysis of regulatory network controlling the initiation of sporulation in B. subtilis H A spo 0 A H + + kin. A Kin. A Sin. R Sin. I - phosphorelay - spo 0 E A sig. H A (spo 0 H) A sin. R + - sig. F Abr. B A A + - - + - F Spo 0 A˜P abr. B - H sin. I + Spo 0 E H - Spo 0 A Signal + Sin. R/Sin. I H Hpr A hpr (sco. R) 17
Simulation of sporulation in B. subtilis v Simulation method applied to analysis of regulatory network controlling the initiation of sporulation in B. subtilis H A spo 0 A H + + kin. A Kin. A Sin. R Sin. I - phosphorelay - spo 0 E A sig. H A (spo 0 H) A sin. R + - sig. F Abr. B A A + - - + - F Spo 0 A˜P abr. B - H sin. I + Spo 0 E H - Spo 0 A Signal + Sin. R/Sin. I H Hpr A hpr (sco. R) 17
 Model of sporulation network v Essential part of sporulation network has been modeled by qualitative PL model: 11 differential equations, with 59 inequality constraints v Most interactions incorporated in model have been characterized on genetic and/or molecular level v With few exceptions, inequality constraints are uniquely determined by biological data If several alternative constraints are consistent with biological data, every alternative considered de Jong, Geiselmann et al. (2004), Bull. Math. Biol. , 66(2): 261 -300 18
Model of sporulation network v Essential part of sporulation network has been modeled by qualitative PL model: 11 differential equations, with 59 inequality constraints v Most interactions incorporated in model have been characterized on genetic and/or molecular level v With few exceptions, inequality constraints are uniquely determined by biological data If several alternative constraints are consistent with biological data, every alternative considered de Jong, Geiselmann et al. (2004), Bull. Math. Biol. , 66(2): 261 -300 18
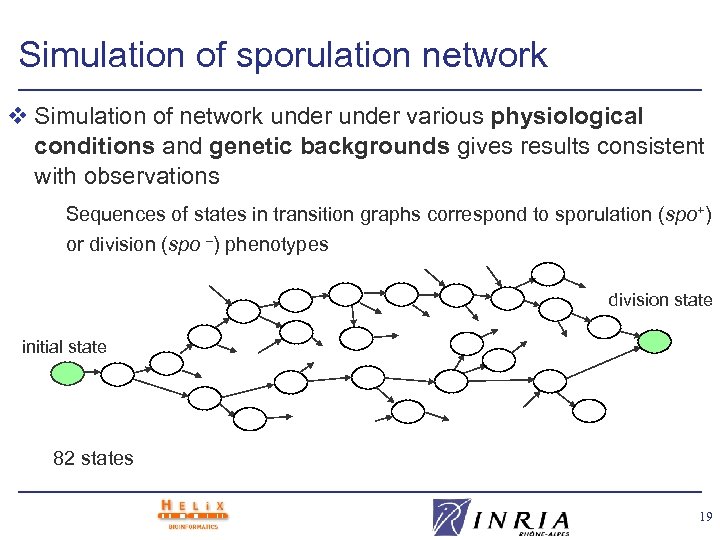 Simulation of sporulation network v Simulation of network under various physiological conditions and genetic backgrounds gives results consistent with observations Sequences of states in transition graphs correspond to sporulation (spo+) or division (spo –) phenotypes division state initial state 82 states 19
Simulation of sporulation network v Simulation of network under various physiological conditions and genetic backgrounds gives results consistent with observations Sequences of states in transition graphs correspond to sporulation (spo+) or division (spo –) phenotypes division state initial state 82 states 19
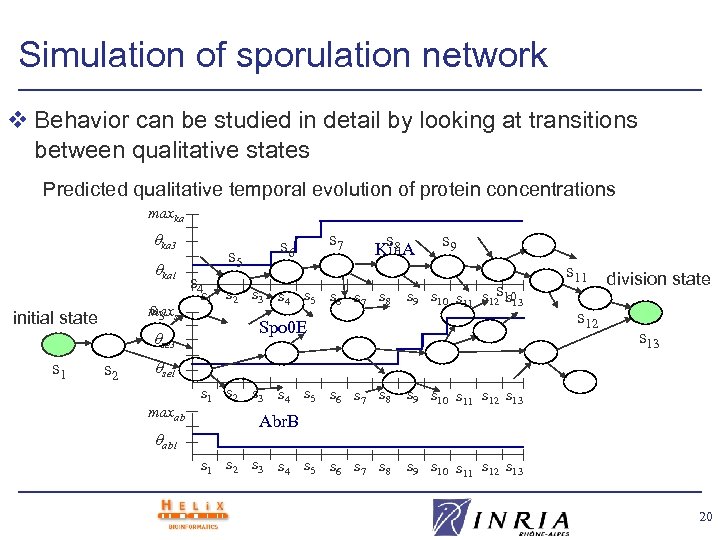 Simulation of sporulation network v Behavior can be studied in detail by looking at transitions between qualitative states Predicted qualitative temporal evolution of protein concentrations maxka ka 3 ka 1 s 3 maxse initial state s 1 se 3 s 2 s 5 s 6 s 7 s 8 Kin. A s 4 s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s s 13 s 9 s 10 s 11 s 12 10 Spo 0 E s 11 division state s 12 s 13 se 1 maxab ab 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Abr. B s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 20
Simulation of sporulation network v Behavior can be studied in detail by looking at transitions between qualitative states Predicted qualitative temporal evolution of protein concentrations maxka ka 3 ka 1 s 3 maxse initial state s 1 se 3 s 2 s 5 s 6 s 7 s 8 Kin. A s 4 s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s s 13 s 9 s 10 s 11 s 12 10 Spo 0 E s 11 division state s 12 s 13 se 1 maxab ab 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Abr. B s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 20
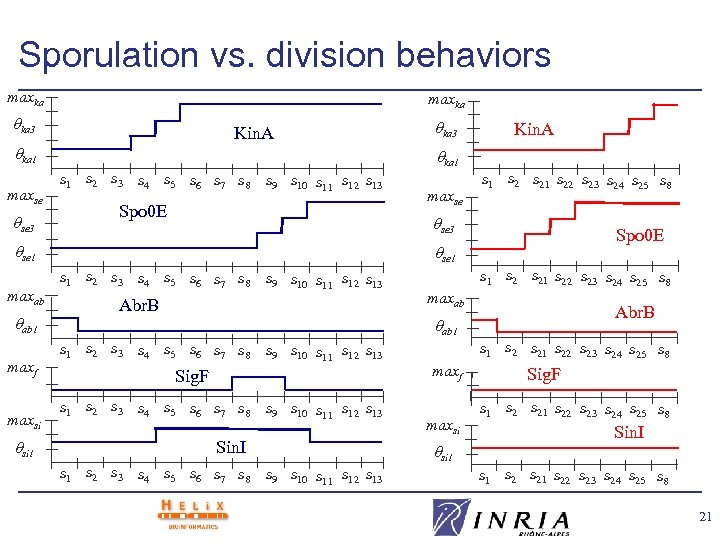 Sporulation vs. division behaviors maxka ka 3 Kin. A ka 1 maxse se 3 ab 1 maxf maxsi si 1 Kin. A ka 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Spo 0 E maxse s 1 s 21 s 22 s 23 s 24 s 25 s 8 se 3 se 1 maxab ka 3 Spo 0 E se 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Abr. B s 1 s 21 s 22 s 23 s 24 s 25 s 8 maxab Abr. B ab 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Sig. F s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Sin. I s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 1 s 21 s 22 s 23 s 24 s 25 s 8 maxf maxsi si 1 s 9 s 10 s 11 s 12 s 13 Sig. F s 1 s 21 s 22 s 23 s 24 s 25 s 8 Sin. I s 1 s 21 s 22 s 23 s 24 s 25 s 8 21
Sporulation vs. division behaviors maxka ka 3 Kin. A ka 1 maxse se 3 ab 1 maxf maxsi si 1 Kin. A ka 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Spo 0 E maxse s 1 s 21 s 22 s 23 s 24 s 25 s 8 se 3 se 1 maxab ka 3 Spo 0 E se 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Abr. B s 1 s 21 s 22 s 23 s 24 s 25 s 8 maxab Abr. B ab 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Sig. F s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 9 s 10 s 11 s 12 s 13 Sin. I s 1 s 2 s 3 s 4 s 5 s 6 s 7 s 8 s 1 s 21 s 22 s 23 s 24 s 25 s 8 maxf maxsi si 1 s 9 s 10 s 11 s 12 s 13 Sig. F s 1 s 21 s 22 s 23 s 24 s 25 s 8 Sin. I s 1 s 21 s 22 s 23 s 24 s 25 s 8 21
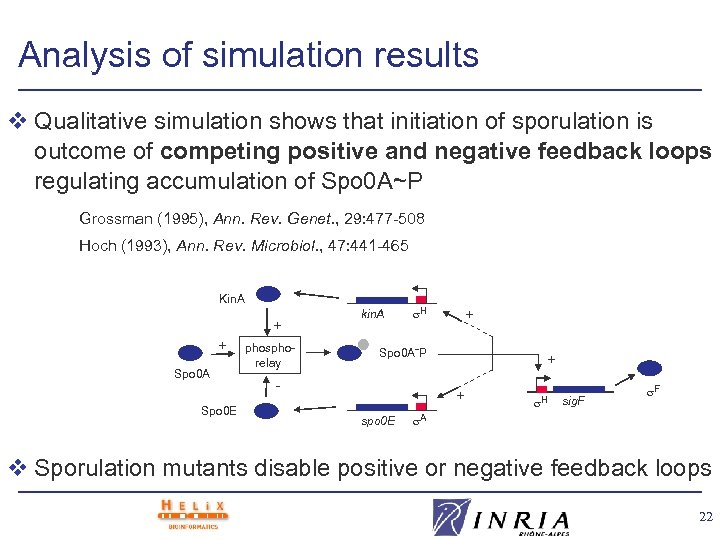 Analysis of simulation results v Qualitative simulation shows that initiation of sporulation is outcome of competing positive and negative feedback loops regulating accumulation of Spo 0 A~P Grossman (1995), Ann. Rev. Genet. , 29: 477 -508 Hoch (1993), Ann. Rev. Microbiol. , 47: 441 -465 Kin. A + + Spo 0 A Spo 0 E phosphorelay kin. A H + Spo 0 A˜P - + + spo 0 E H sig. F F A v Sporulation mutants disable positive or negative feedback loops 22
Analysis of simulation results v Qualitative simulation shows that initiation of sporulation is outcome of competing positive and negative feedback loops regulating accumulation of Spo 0 A~P Grossman (1995), Ann. Rev. Genet. , 29: 477 -508 Hoch (1993), Ann. Rev. Microbiol. , 47: 441 -465 Kin. A + + Spo 0 A Spo 0 E phosphorelay kin. A H + Spo 0 A˜P - + + spo 0 E H sig. F F A v Sporulation mutants disable positive or negative feedback loops 22
 Nutritional stress response in E. coli v Response of E. coli to nutritional stress conditions controlled by network of global regulators of transcription Fis, Crp, H-NS, Lrp, Rpo. S, … v Network only partially known and no global view of its functioning available v Computational and experimental study directed at understanding of: l How network controls gene expression to adapt cell to stress conditions l How network evolves over time to adapt to environment v Projects: inter-EPST, ARC INRIA, and ACI IMPBio ENS, Paris ; INRIA ; UJF, Grenoble ; UHA, Mulhouse 23
Nutritional stress response in E. coli v Response of E. coli to nutritional stress conditions controlled by network of global regulators of transcription Fis, Crp, H-NS, Lrp, Rpo. S, … v Network only partially known and no global view of its functioning available v Computational and experimental study directed at understanding of: l How network controls gene expression to adapt cell to stress conditions l How network evolves over time to adapt to environment v Projects: inter-EPST, ARC INRIA, and ACI IMPBio ENS, Paris ; INRIA ; UJF, Grenoble ; UHA, Mulhouse 23
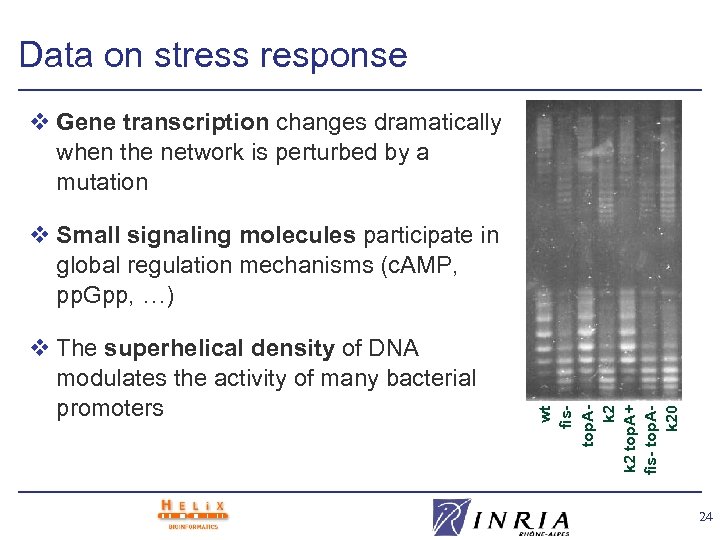 Data on stress response v Gene transcription changes dramatically when the network is perturbed by a mutation v The superhelical density of DNA modulates the activity of many bacterial promoters wt fistop. Ak 2 top. A+ fis- top. Ak 20 v Small signaling molecules participate in global regulation mechanisms (c. AMP, pp. Gpp, …) 24
Data on stress response v Gene transcription changes dramatically when the network is perturbed by a mutation v The superhelical density of DNA modulates the activity of many bacterial promoters wt fistop. Ak 2 top. A+ fis- top. Ak 20 v Small signaling molecules participate in global regulation mechanisms (c. AMP, pp. Gpp, …) 24
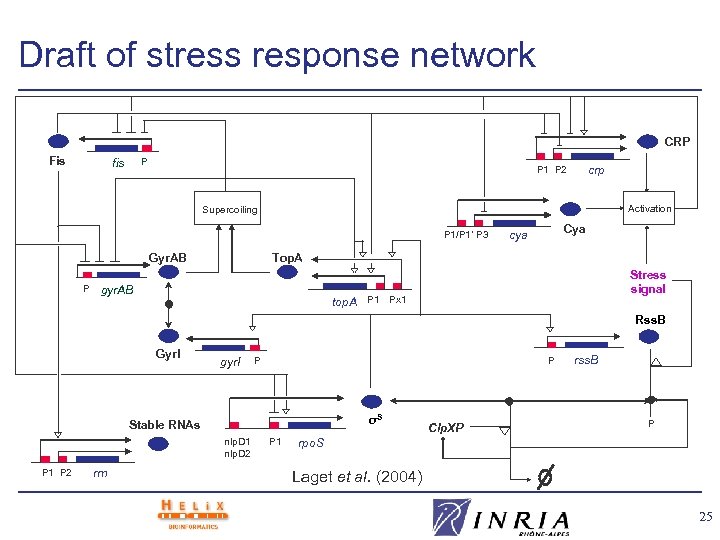 Draft of stress response network CRP Fis P fis crp P 1 P 2 Activation Supercoiling P 1/P 1’ P 3 Top. A Gyr. AB P Cya cya gyr. AB Stress signal top. A P 1 Px 1 Rss. B Gyr. I gyr. I P P σS Stable RNAs nlp. D 1 nlp. D 2 P 1 P 2 rrn P 1 Clp. XP rss. B P rpo. S Laget et al. (2004) 25
Draft of stress response network CRP Fis P fis crp P 1 P 2 Activation Supercoiling P 1/P 1’ P 3 Top. A Gyr. AB P Cya cya gyr. AB Stress signal top. A P 1 Px 1 Rss. B Gyr. I gyr. I P P σS Stable RNAs nlp. D 1 nlp. D 2 P 1 P 2 rrn P 1 Clp. XP rss. B P rpo. S Laget et al. (2004) 25
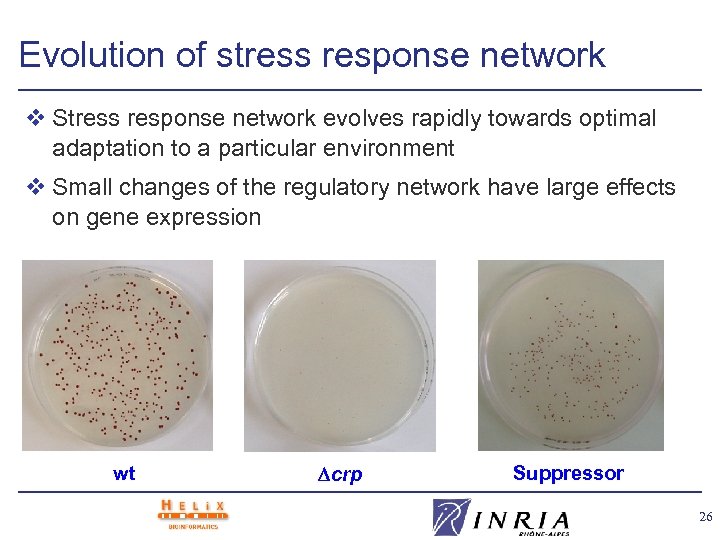 Evolution of stress response network v Stress response network evolves rapidly towards optimal adaptation to a particular environment v Small changes of the regulatory network have large effects on gene expression wt crp Suppressor 26
Evolution of stress response network v Stress response network evolves rapidly towards optimal adaptation to a particular environment v Small changes of the regulatory network have large effects on gene expression wt crp Suppressor 26
 Validation of network models v Bottleneck of qualitative simulation: visual inspection of large state transition graphs v Goal: develop a method that can test if state transition graph satisfies certain properties Is transition graph consistent with observed behavior? v Model checking is automated technique for verifying that finite state system satisfies certain properties Clarke et al. (1999), Model Checking, MIT Press v Computer tools are available to perform automated, efficient and reliable model checking (Nu. SMV) 27
Validation of network models v Bottleneck of qualitative simulation: visual inspection of large state transition graphs v Goal: develop a method that can test if state transition graph satisfies certain properties Is transition graph consistent with observed behavior? v Model checking is automated technique for verifying that finite state system satisfies certain properties Clarke et al. (1999), Model Checking, MIT Press v Computer tools are available to perform automated, efficient and reliable model checking (Nu. SMV) 27
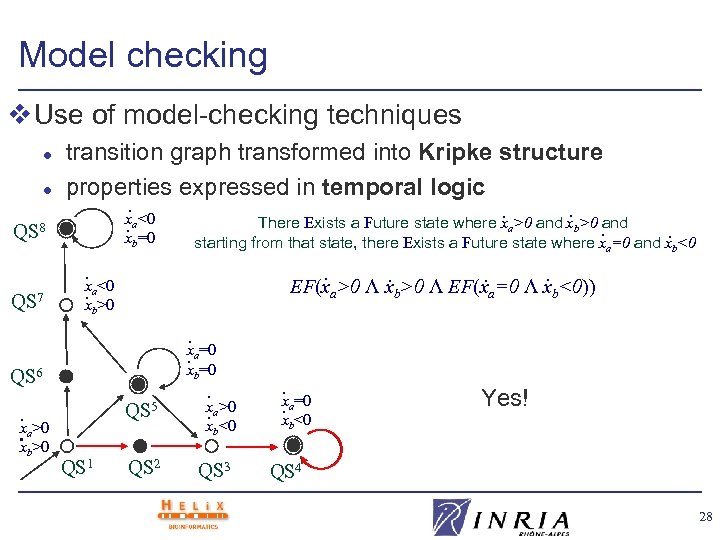 Model checking v Use of model-checking techniques l l QS 8 QS 7 transition graph transformed into Kripke structure properties expressed in temporal logic. . . x There Exists a Future state where xa>0 and xb>0 and. a<0. . xb=0 starting from that state, there Exists a Future state where x =0 and x <0 a . <0 x. a>0 x . . . b . EF(xa>0 Λ xb>0 Λ EF(xa=0 Λ xb<0)) b . x. a=0 xb=0 QS 6 . x. a>0 x >0 b QS 5 . x. a>0 x <0 b QS 1 QS 2 QS 3 . =0 x. a xb<0 Yes! QS 4 28
Model checking v Use of model-checking techniques l l QS 8 QS 7 transition graph transformed into Kripke structure properties expressed in temporal logic. . . x There Exists a Future state where xa>0 and xb>0 and. a<0. . xb=0 starting from that state, there Exists a Future state where x =0 and x <0 a . <0 x. a>0 x . . . b . EF(xa>0 Λ xb>0 Λ EF(xa=0 Λ xb<0)) b . x. a=0 xb=0 QS 6 . x. a>0 x >0 b QS 5 . x. a>0 x <0 b QS 1 QS 2 QS 3 . =0 x. a xb<0 Yes! QS 4 28
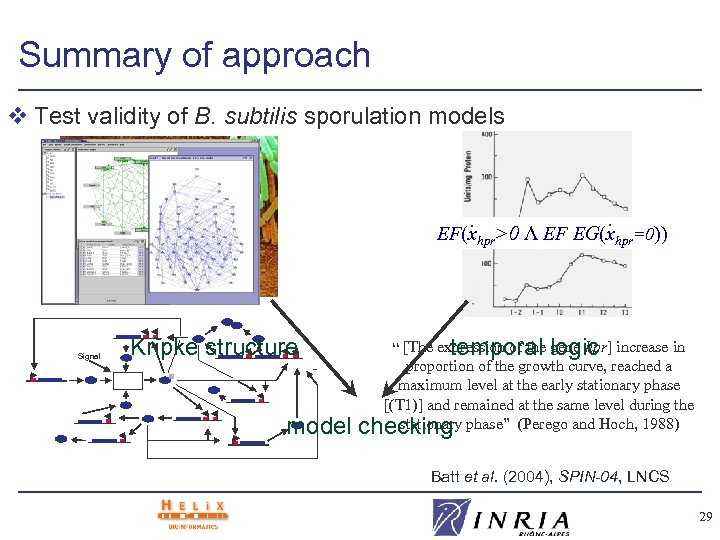 Summary of approach v Test validity of B. subtilis sporulation models . . EF(xhpr>0 Λ EF EG(xhpr=0)) Signal Kripke structure temporal logic “ [The expression of the gene hpr] increase in - model proportion of the growth curve, reached a maximum level at the early stationary phase [(T 1)] and remained at the same level during the stationary checking phase” (Perego and Hoch, 1988) Batt et al. (2004), SPIN-04, LNCS 29
Summary of approach v Test validity of B. subtilis sporulation models . . EF(xhpr>0 Λ EF EG(xhpr=0)) Signal Kripke structure temporal logic “ [The expression of the gene hpr] increase in - model proportion of the growth curve, reached a maximum level at the early stationary phase [(T 1)] and remained at the same level during the stationary checking phase” (Perego and Hoch, 1988) Batt et al. (2004), SPIN-04, LNCS 29
 Conclusions v Implemented method for qualitative simulation of large and complex genetic regulatory networks Method based on work in mathematical biology and qualitative reasoning v Method validated by analysis of regulatory network underlying initiation of sporulation in B. subtilis Simulation results consistent with observations v Method currently applied to analysis of regulatory network controlling stress adaptation in E. coli Simulation yields predictions that can be tested in the laboratory 30
Conclusions v Implemented method for qualitative simulation of large and complex genetic regulatory networks Method based on work in mathematical biology and qualitative reasoning v Method validated by analysis of regulatory network underlying initiation of sporulation in B. subtilis Simulation results consistent with observations v Method currently applied to analysis of regulatory network controlling stress adaptation in E. coli Simulation yields predictions that can be tested in the laboratory 30
 Work in progress v Validation of models of regulatory networks using gene expression data Model-checking techniques v Search of attractors in phase space and determination of their stability v Further development of computer tool GNA Connection with biological knowledge bases, … v Study of bacterial regulatory networks Sporulation in B. subtilis, phage Mu infection of E. coli, signal transduction in Synechocystis, stress adaptation in E. coli 31
Work in progress v Validation of models of regulatory networks using gene expression data Model-checking techniques v Search of attractors in phase space and determination of their stability v Further development of computer tool GNA Connection with biological knowledge bases, … v Study of bacterial regulatory networks Sporulation in B. subtilis, phage Mu infection of E. coli, signal transduction in Synechocystis, stress adaptation in E. coli 31
 Contributors Grégory Batt INRIA Rhône-Alpes Hidde de Jong INRIA Rhône-Alpes Hans Geiselmann Université Joseph Fourier, Grenoble Jean-Luc Gouzé INRIA Sophia-Antipolis Céline Hernandez INRIA Rhône-Alpes, now at SIB, Genève Eva Laget INRIA Rhône-Alpes and INSA Lyon Michel Page INRIA Rhône-Alpes and Université Pierre Mendès France, Grenoble Delphine Ropers INRIA Rhône-Alpes Tewfik Sari Université de Haute Alsace, Mulhouse Dominique Schneider Université Joseph Fourier, Grenoble 32
Contributors Grégory Batt INRIA Rhône-Alpes Hidde de Jong INRIA Rhône-Alpes Hans Geiselmann Université Joseph Fourier, Grenoble Jean-Luc Gouzé INRIA Sophia-Antipolis Céline Hernandez INRIA Rhône-Alpes, now at SIB, Genève Eva Laget INRIA Rhône-Alpes and INSA Lyon Michel Page INRIA Rhône-Alpes and Université Pierre Mendès France, Grenoble Delphine Ropers INRIA Rhône-Alpes Tewfik Sari Université de Haute Alsace, Mulhouse Dominique Schneider Université Joseph Fourier, Grenoble 32
 References de Jong, H. (2002), Modeling and simulation of genetic regulatory systems: A literature review, J. Comp. Biol. , 9(1): 69 -105. de Jong, H. , J. Geiselmann & D. Thieffry (2003), Qualitative modelling and simulation of developmental regulatory networks, On Growth, Form, and Computers, Academic Press, 109 -134. Gouzé, J. -L. & T. Sari (2002), A class of piecewise-linear differential equations arising in biological models, Dyn. Syst. , 17(4): 299 -316. de Jong, H. , J. -L. Gouzé, C. Hernandez, M. Page, T. Sari & J. Geiselmann (2004), Qualitative simulation of genetic regulatory networks using piecewise-linear models, Bull. Math. Biol. , 66(2): 301 -340. de Jong, H. , J. Geiselmann, C. Hernandez & M. Page (2003), Genetic Network Analyzer: Qualitative simulation of genetic regulatory networks, Bioinformatics, 19(3): 336 -344. de Jong, H. , J. Geiselmann, G. Batt, C. Hernandez & M. Page (2004), Qualitative simulation of the initiation of sporulation in B. subtilis, Bull. Math. Biol. , 66(2): 261 -340. GNA web site: http: //www-helix. inrialpes. fr/article 122. html 33
References de Jong, H. (2002), Modeling and simulation of genetic regulatory systems: A literature review, J. Comp. Biol. , 9(1): 69 -105. de Jong, H. , J. Geiselmann & D. Thieffry (2003), Qualitative modelling and simulation of developmental regulatory networks, On Growth, Form, and Computers, Academic Press, 109 -134. Gouzé, J. -L. & T. Sari (2002), A class of piecewise-linear differential equations arising in biological models, Dyn. Syst. , 17(4): 299 -316. de Jong, H. , J. -L. Gouzé, C. Hernandez, M. Page, T. Sari & J. Geiselmann (2004), Qualitative simulation of genetic regulatory networks using piecewise-linear models, Bull. Math. Biol. , 66(2): 301 -340. de Jong, H. , J. Geiselmann, C. Hernandez & M. Page (2003), Genetic Network Analyzer: Qualitative simulation of genetic regulatory networks, Bioinformatics, 19(3): 336 -344. de Jong, H. , J. Geiselmann, G. Batt, C. Hernandez & M. Page (2004), Qualitative simulation of the initiation of sporulation in B. subtilis, Bull. Math. Biol. , 66(2): 261 -340. GNA web site: http: //www-helix. inrialpes. fr/article 122. html 33


