90cd2e41425f5b21885302bea5774201.ppt
- Количество слайдов: 23

Principles of Bioinorganic Chemistry - 2004 The grade for this course will be determined by a term exam (35%), a written research paper with oral presentation (55%), and problem sets (10%). The oral presentations will be held in research conference style at an all-day symposium at MIT on Saturday, October 30 th. Please reserve the date for there are no excused absences. Papers are due October 28 th. Problem sets are due one week after their assigned date. Recitations are held at 5 PM on Mondays. WEB SITE: web. mit. edu/5. 062/www/
![Control and Use of Metal Ion Concentrations PRINCIPLES: • Homeostasis: maintain [M+ ] in Control and Use of Metal Ion Concentrations PRINCIPLES: • Homeostasis: maintain [M+ ] in](https://present5.com/presentation/90cd2e41425f5b21885302bea5774201/image-2.jpg)
Control and Use of Metal Ion Concentrations PRINCIPLES: • Homeostasis: maintain [M+ ] in proper range • Detoxification: remove excess and/or unnatural metal ions • Extracellular carriers • Passive transport • Ion channels/pumps • Metalloregulation • Binding and release of metal ions to receptors controlled by p. H and redox changes • Ion concentration gradients - used to transmit energy and information

Note hinge motion that accompanies iron/carbonate binding
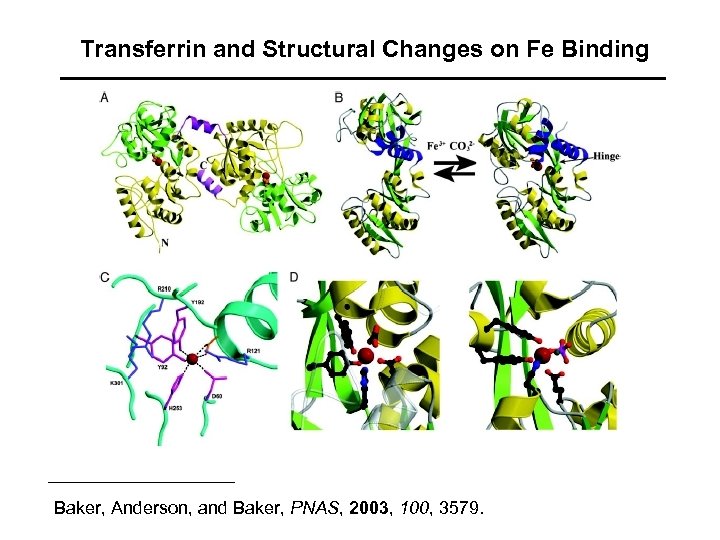
Transferrin and Structural Changes on Fe Binding Baker, Anderson, and Baker, PNAS, 2003, 100, 3579.
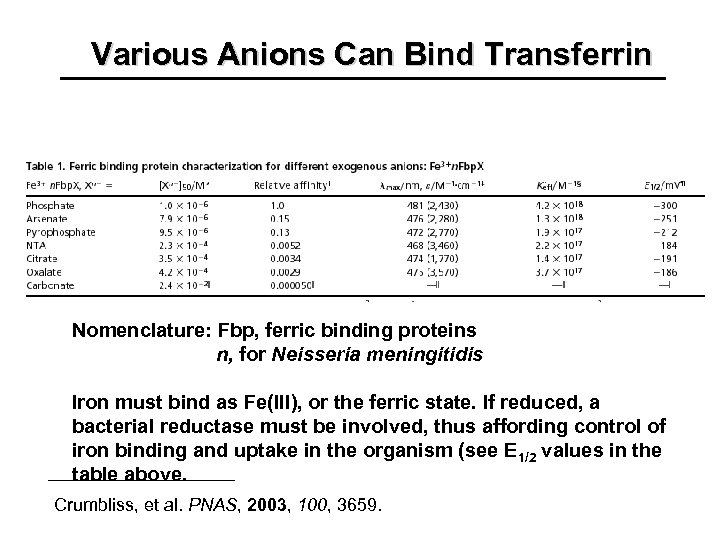



Various Anions Can Bind Transferrin Nomenclature: Fbp, ferric binding proteins n, for Neisseria meningitidis Iron must bind as Fe(III), or the ferric state. If reduced, a bacterial reductase must be involved, thus affording control of iron binding and uptake in the organism (see E 1/2 values in the table above. Crumbliss, et al. PNAS, 2003, 100, 3659.
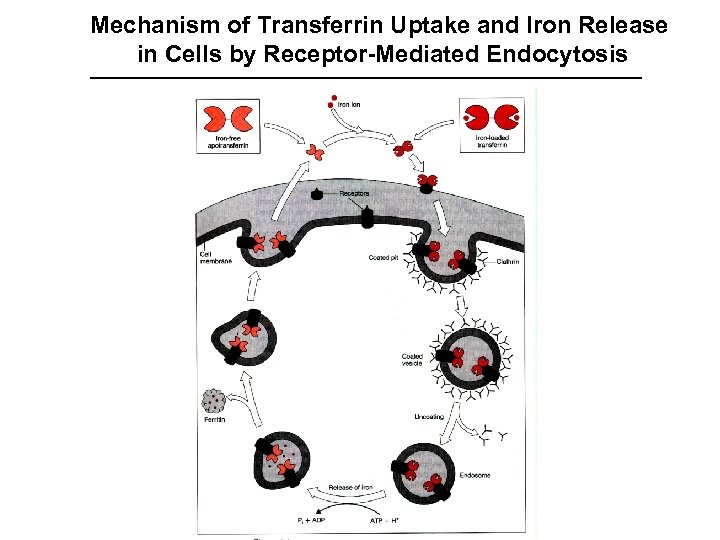
Mechanism of Transferrin Uptake and Iron Release in Cells by Receptor-Mediated Endocytosis

Metal Regulation of Gene Expression PRINCIPLES: • Metal-mediated protein structure changes affect transcription • Metal-mediated protein structure changes affect translation • Apo vs holo metalloproteins bind DNA/RNA differently • Metalloregulatory protein is the sensor - inorganic chemistry • Metal-induced protein structure changes also activate enzyme • Metal-induced protein structure changes are metal-specific ILLUSTRATIONS: • Iron regulatory proteins (IRPs); control Ft and Tf translation • Regulation of a toxic metal, mercury • Zinc finger proteins control transcription • Ca 2+, a second messenger and sentinel at the synapse

Regulation of Iron Levels in Cells The Players: • Ferritin, the iron storage protein: 24 -subunits, ~175 aa each; has cubic symmetry; apo. Ft can house 1000 iron atoms in its central core; a ferroxidase center loads the iron into the protein • Transferrin, the uptake protein, discussed previously Metalloregulation: • In bacteria, occurs at the transcriptional level • In mammals, the synthesis of apoferritin and of the transferrin receptor are regulated at the level of translation, not transcription Central dogma of molecular biology: DNA transcription m. RNA translation Protein
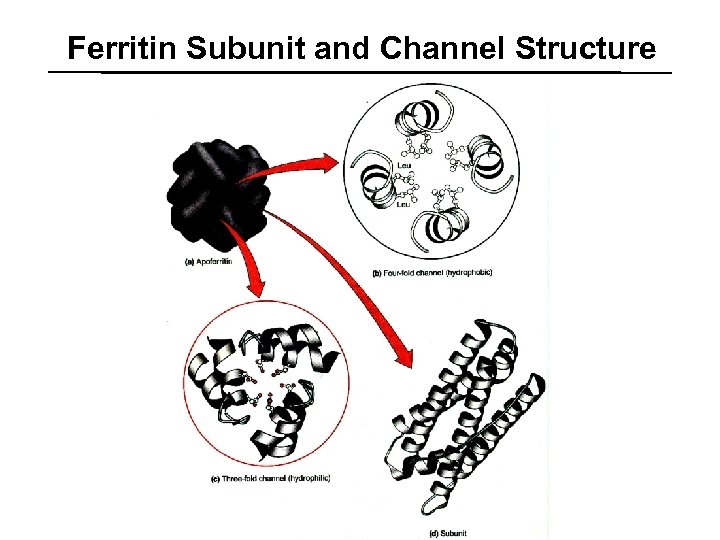
Ferritin Subunit and Channel Structure
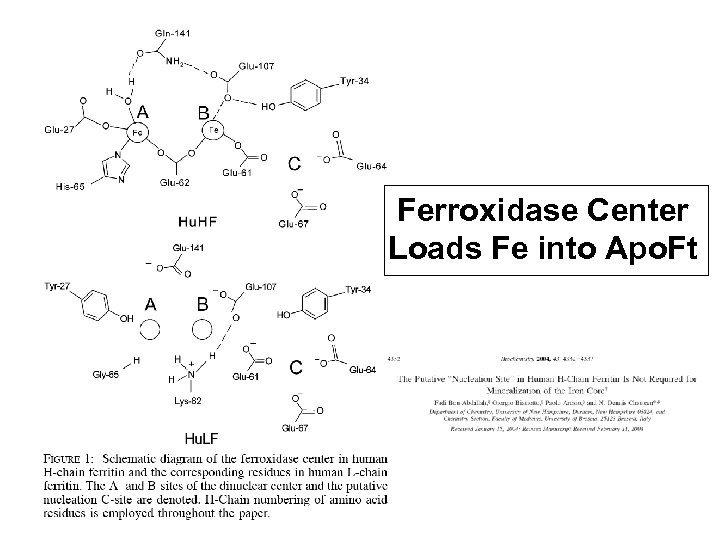

Ferroxidase Center Loads Fe into Apo. Ft
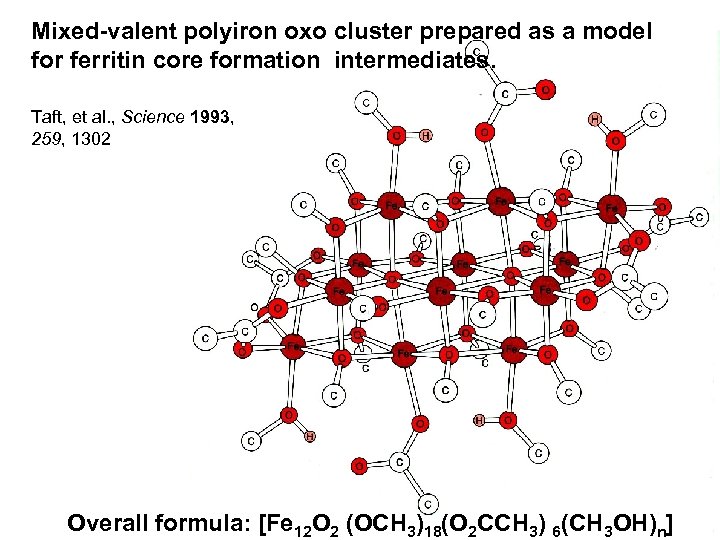
Mixed-valent polyiron oxo cluster prepared as a model for ferritin core formation intermediates. Taft, et al. , Science 1993, 259, 1302 Overall formula: [Fe 12 O 2 (OCH 3)18(O 2 CCH 3) 6(CH 3 OH)n]
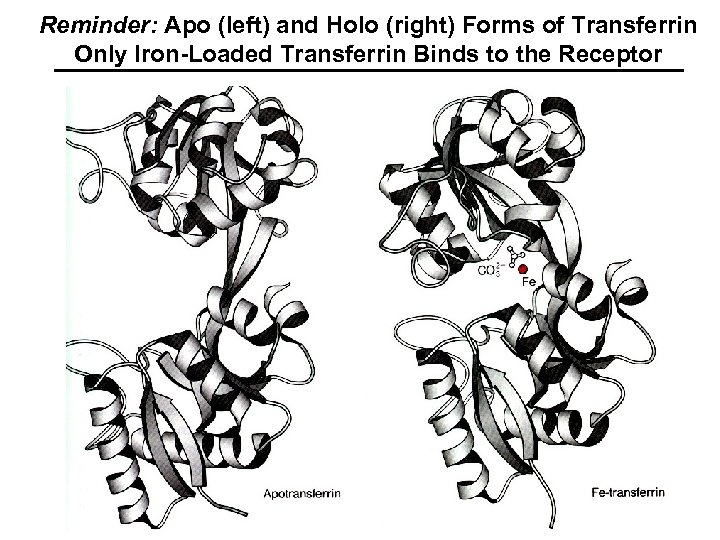
Reminder: Apo (left) and Holo (right) Forms of Transferrin Only Iron-Loaded Transferrin Binds to the Receptor

Metalloregulation of Iron Uptake and Storage Bacteria: A single protein, Fur (for iron uptake regulator), controls the transcription of genes involved in siderophore biosynthesis. Fur is a dimer with subunits of Mr 17 k. Da. At high iron levels, the Fur protein has bound metal and interacts specifically with DNA repressing transcription. Mammals: Expression of ferritin and the transferrin receptor is regulated at the translational level.

Components of the Metalloregulatory System Stemloop structure in the m. RNA IRP Ironresponsive protein (IRP)
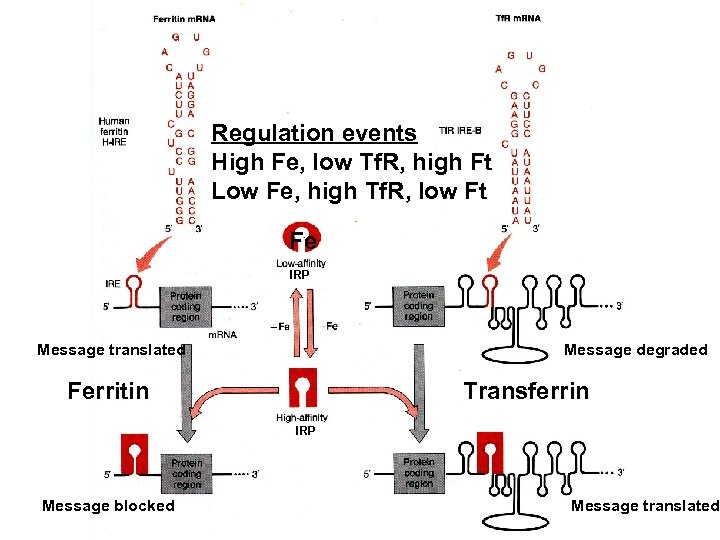
Regulation events High Fe, low Tf. R, high Ft Low Fe, high Tf. R, low Ft Fe IRP Message translated Message degraded Ferritin Transferrin IRP Message blocked Message translated

IRP 1 is the Cytosolic Aconitase Contains an Fe 4 S 4 Cluster assembled in protein, which then dissociates from m. RNA Apoprotein stays associated with m. RNA

Regulation of a Toxic Metal, Mercury The problem: Mercury in the environment of industrial plants is converted by bacterial to harmful organomercury compounds. Fish and other plant and animal life assimilate the mercury which ultimately enters the human food chain. Bacteria defend themselves against the mercury by using the proteins listed below. The players: Organomercurial lyase Mercuric ion reductase Mer. R, the intracellular mercuric ion sensor The implications: Transcription of the genes encoding the proteins is controlled by Mer. R in response to mercury levels

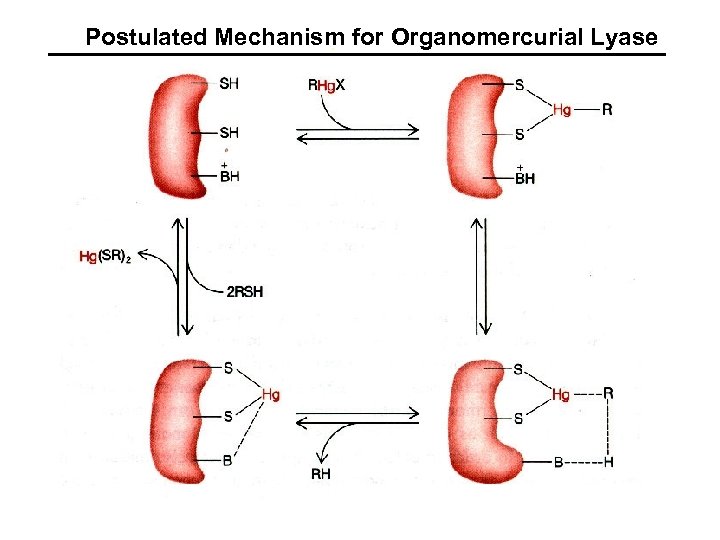
Postulated Mechanism for Organomercurial Lyase

Mer. R and Mercuric Ion Reductase Properties Reductase: no structural or detailed mechanistic information Mer. R EXAFS spectroscopy and chemical modification experiments indicate that Hg-Mer. R has a 3 -coordinate, Hg(S-Cys)3 environment with an average Hg–S distance of 2. 43 Å. This unusual tridentate heavy metal receptor site is consistent with thermodynamic stability of [Hg(SR) 3]- complexes and may account both for the high affinity of the Hg(II) binding and for the selectivity for Hg(II) over other soft metal ions that prefer tetrahedral metal-thiolate coordination.
![Effect of [Hg 2+] on Transcription Activity Effect of [Hg 2+] on Transcription Activity](https://present5.com/presentation/90cd2e41425f5b21885302bea5774201/image-23.jpg)
Effect of [Hg 2+] on Transcription Activity
90cd2e41425f5b21885302bea5774201.ppt