ae257273e42b8bd8e459f55586866b16.ppt
- Количество слайдов: 49

Principles of Bioinorganic Chemistry - 2003 The grade for this course will be determined by a term exam (35%), a written research paper with oral presentation (45%), problem sets (12%) and classroom participation (8%). The oral presentations will be held in research conference style at MIT's Endicott House estate in Dedham, MA, on Saturday, October 18. Please reserve the date for there are no excused absences. Papers will be due approximately one week earlier. WEB SITE: web. mit. edu/5. 062/www/

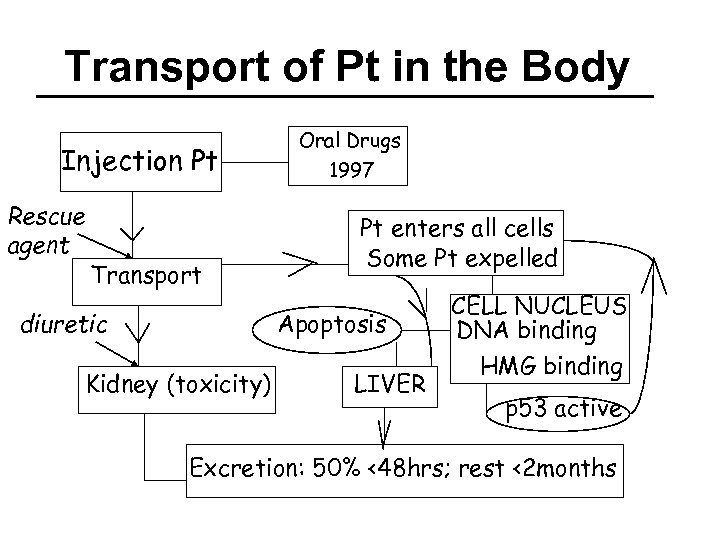
Transport of Pt in the Body Injection Pt Rescue agent Transport Oral Drugs 1997 Pt enters all cells Some Pt expelled Apoptosis diuretic Kidney (toxicity) LIVER CELL NUCLEUS DNA binding HMG binding p 53 active Excretion: 50% <48 hrs; rest <2 months

Obstacles for Cisplatin On Route to DNA • Reagents in blood plasm: proteins, protective agents • Receptors at cell wall • Reagents in cellular membrane • Reagents inside the cell, such as glutathione, S-donor peptides • Reagents in the nuclear membrane

Transport from Outside to Inside Cell • • Cell receptors? Active or passive cell-wall transport? Relationships with resistance? ? Carrier molecule? YES: PS (phos/ser)! • Inside cell: glutathione-like ligands take over; can some Pt species escape to the nucleus? YES: transfer proved


Pt Pt Pt

Structures of the 1, 2 -d(Gp. G) Intrastrand Cisplatin Adduct d(p. G) adduct duplex DNA adduct Sherman et al. (1985) Science 230, 412 Takahara et al. (1995) Nature 377, 649.

Structure of a {Pt(R, R-DACH)}2+ Intrastrand Cross. Link in a Duplex Dodecamer Showing the G*G* Step A very similar structure occurs for the 3’ orientational isomer of a {Pt(NH 3)(NH 2 Cy)}2+ G*G* cross-link on the same duplex dodecamer.
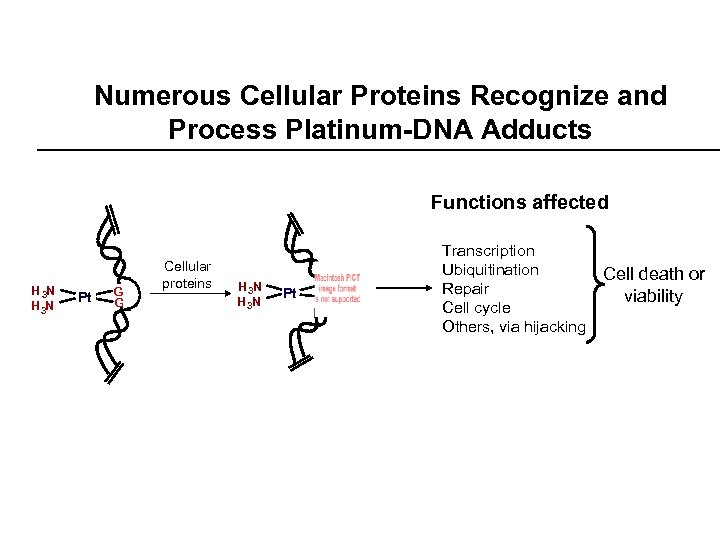
Numerous Cellular Proteins Recognize and Process Platinum-DNA Adducts Functions affected H 3 N Pt GG G Cellular proteins H 3 N Pt G G G Transcription Ubiquitination Repair Cell cycle Others, via hijacking Cell death or viability
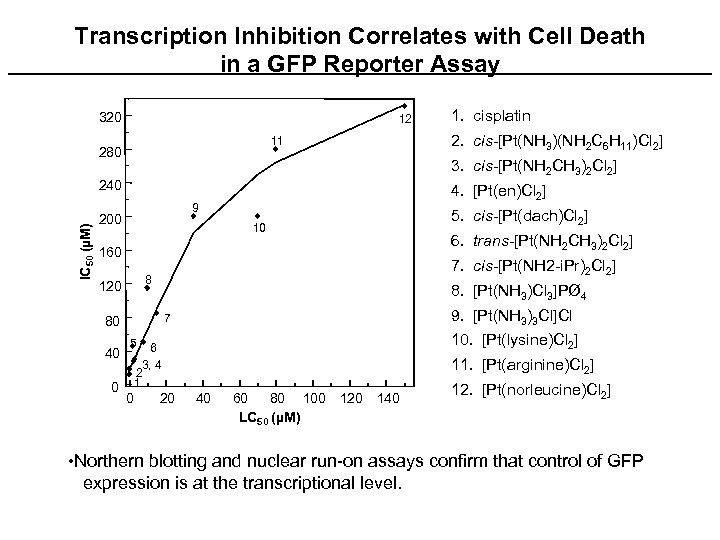

Transcription Inhibition Correlates with Cell Death in a GFP Reporter Assay 320 12 11 280 IC 50 (µM) 4. [Pt(en)Cl 2] 9 200 10 160 6. trans-[Pt(NH 2 CH 3)2 Cl 2] 8. [Pt(NH 3)Cl 3]PØ 4 9. [Pt(NH 3)3 Cl]Cl 7 80 10. [Pt(lysine)Cl 2] 5 6 3, 4 2 1 0 5. cis-[Pt(dach)Cl 2] 7. cis-[Pt(NH 2 -i. Pr)2 Cl 2] 8 120 0 2. cis-[Pt(NH 3)(NH 2 C 6 H 11)Cl 2] 3. cis-[Pt(NH 2 CH 3)2 Cl 2] 240 40 1. cisplatin 20 11. [Pt(arginine)Cl 2] 40 60 80 100 120 140 LC 50 (µM) 12. [Pt(norleucine)Cl 2] • Northern blotting and nuclear run-on assays confirm that control of GFP expression is at the transcriptional level.
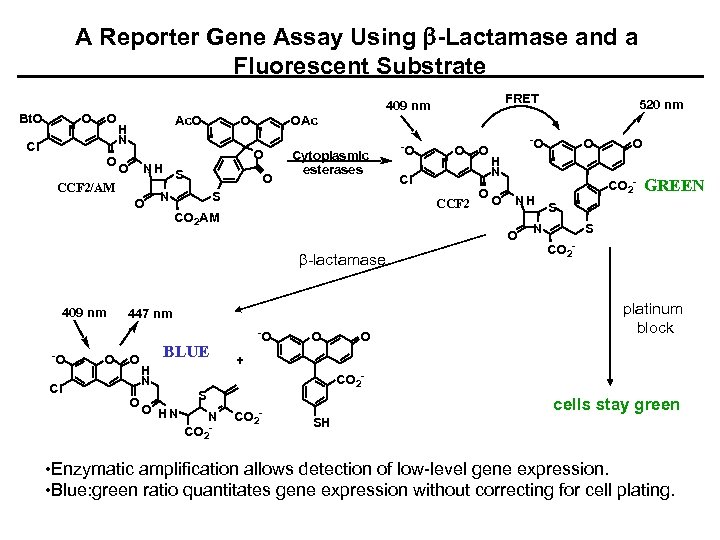
A Reporter Gene Assay Using b-Lactamase and a Fluorescent Substrate Bt. O O O Cl O FRET 409 nm Ac. O H N O CCF 2/AM O OAc O O NH S O N Cytoplasmic esterases S -O O Cl CCF 2 CO 2 AM O O -O b-lactamase -O Cl O 447 nm O O BLUE H N O HN -O O H N O NH O 409 nm 520 nm O CO 2 - GREEN S CO 2 - platinum block + CO 2 - S N CO 2 - cells stay green SH • Enzymatic amplification allows detection of low-level gene expression. • Blue: green ratio quantitates gene expression without correcting for cell plating.

Cisplatin Inhibits b-Lactamase Gene Expression Control 40 µM cisplatin 37°C, 24 h 1 µM CCF 2/AM
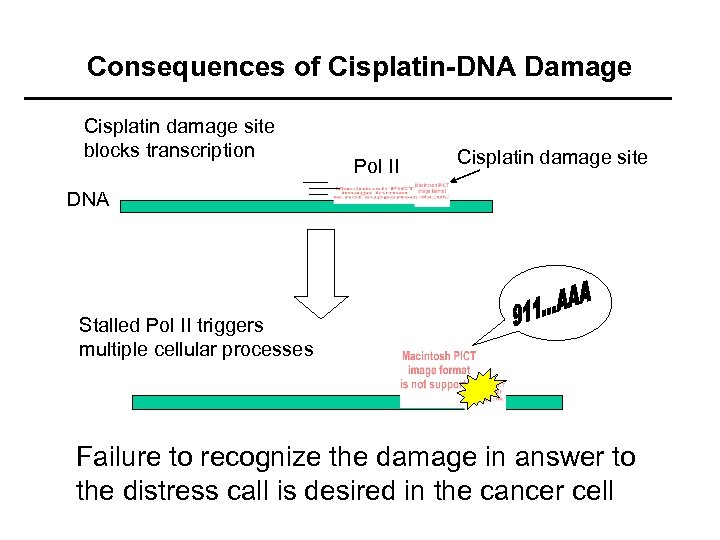
Consequences of Cisplatin-DNA Damage Cisplatin damage site blocks transcription Pol II Cisplatin damage site DNA Stalled Pol II triggers multiple cellular processes Failure to recognize the damage in answer to the distress call is desired in the cancer cell
Consequences of Cisplatin-DNA Damage Ubiquitinated Pol II is replaced. Ub Cellular repair machinery is recruited Repair team Recognition and repair of the damage in answer to the distress call is desired in healthy cells.
Consequence of Cisplatin Damage Restart transcription OR Cell death Dead End Selective cell death of cancer cells is the goal!



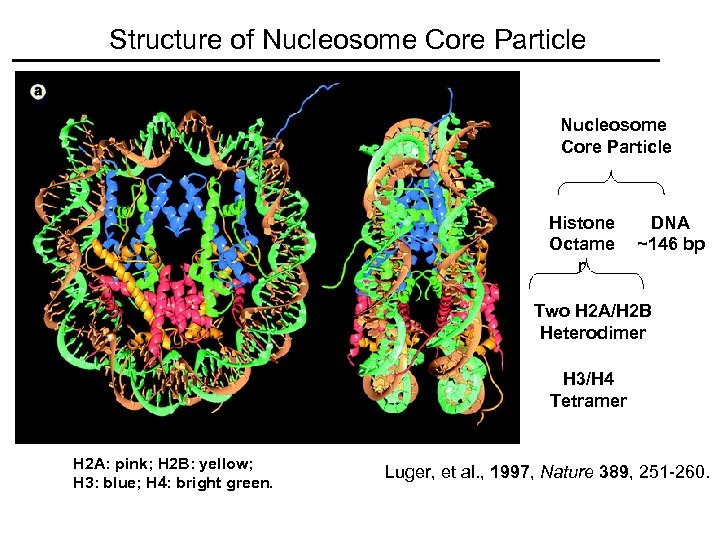
Structure of Nucleosome Core Particle Histone Octame r DNA ~146 bp Two H 2 A/H 2 B Heterodimer H 3/H 4 Tetramer H 2 A: pink; H 2 B: yellow; H 3: blue; H 4: bright green. Luger, et al. , 1997, Nature 389, 251 -260.
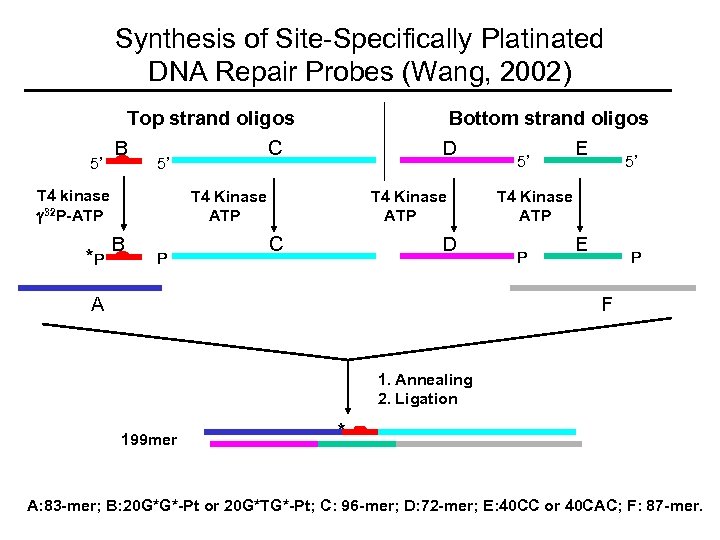
Synthesis of Site-Specifically Platinated DNA Repair Probes (Wang, 2002) Top strand oligos 5’ B *P C 5’ T 4 kinase 32 P-ATP Bottom strand oligos D T 4 Kinase ATP B P T 4 Kinase ATP C D A 5’ E 5’ T 4 Kinase ATP P E P F 1. Annealing 2. Ligation 199 mer * A: 83 -mer; B: 20 G*G*-Pt or 20 G*TG*-Pt; C: 96 -mer; D: 72 -mer; E: 40 CC or 40 CAC; F: 87 -mer.
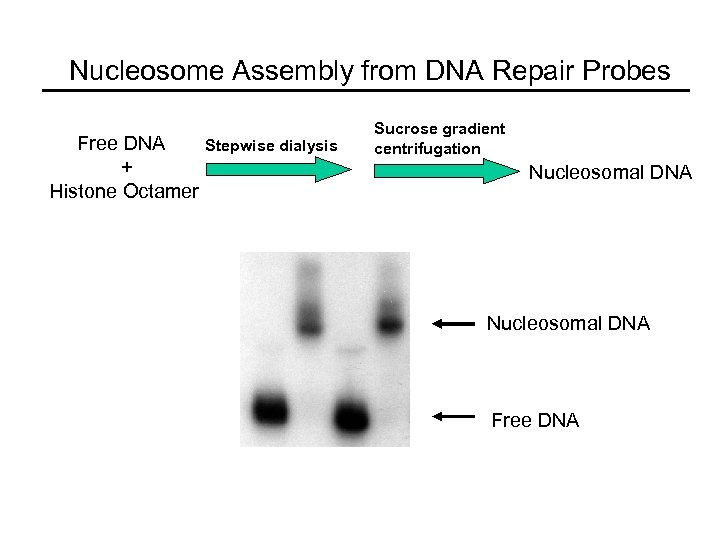
Nucleosome Assembly from DNA Repair Probes Free DNA + Histone Octamer Stepwise dialysis Sucrose gradient centrifugation Nucleosomal DNA Free DNA
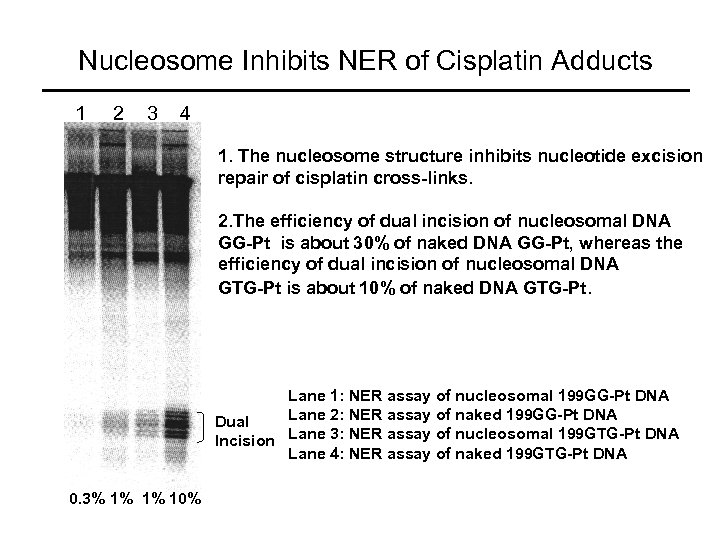
Nucleosome Inhibits NER of Cisplatin Adducts 1 2 3 4 1. The nucleosome structure inhibits nucleotide excision repair of cisplatin cross-links. 2. The efficiency of dual incision of nucleosomal DNA GG-Pt is about 30% of naked DNA GG-Pt, whereas the efficiency of dual incision of nucleosomal DNA GTG-Pt is about 10% of naked DNA GTG-Pt. Lane 1: NER assay of nucleosomal 199 GG-Pt DNA Lane 2: NER assay of naked 199 GG-Pt DNA Dual Incision Lane 3: NER assay of nucleosomal 199 GTG-Pt DNA Lane 4: NER assay of naked 199 GTG-Pt DNA 0. 3% 1% 1% 10%
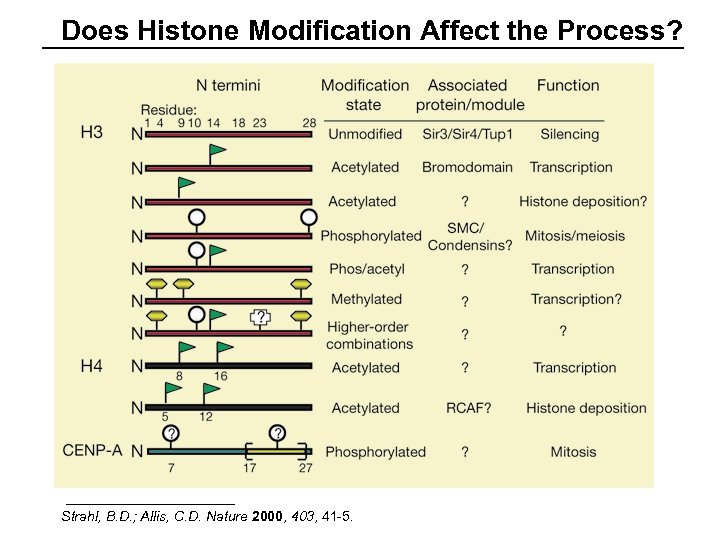

Does Histone Modification Affect the Process? Strahl, B. D. ; Allis, C. D. Nature 2000, 403, 41 -5.
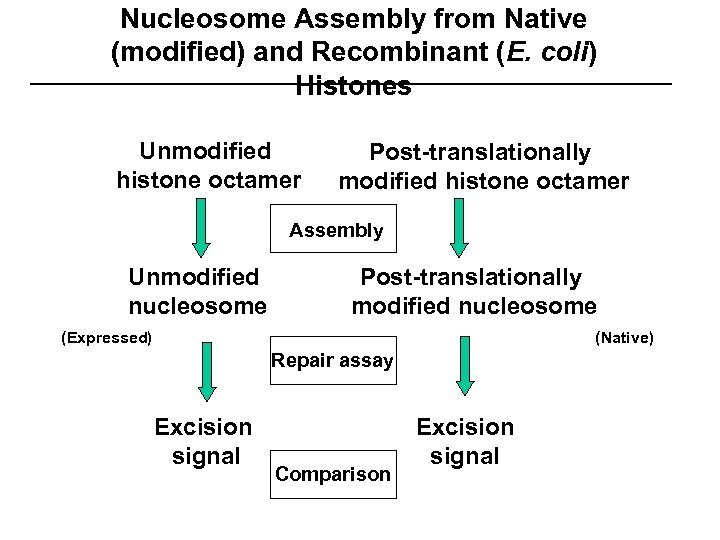
Nucleosome Assembly from Native (modified) and Recombinant (E. coli) Histones Unmodified histone octamer Post-translationally modified histone octamer Assembly Unmodified nucleosome Post-translationally modified nucleosome (Expressed) (Native) Repair assay Excision signal Comparison Excision signal
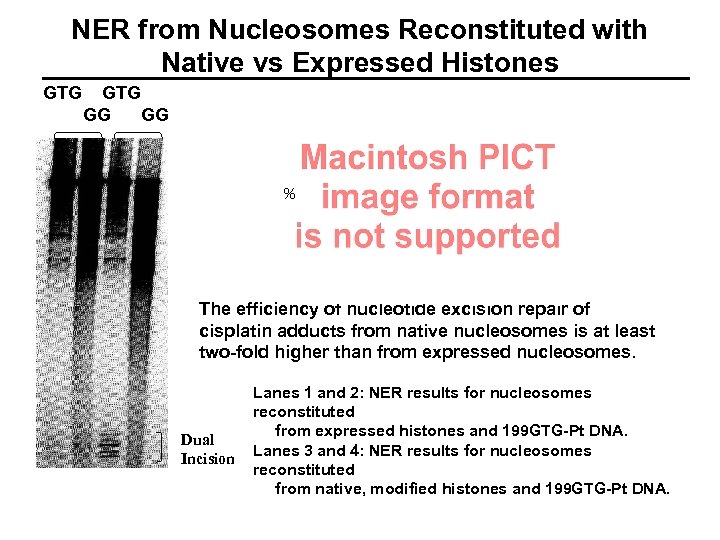
NER from Nucleosomes Reconstituted with Native vs Expressed Histones GTG GG GG % The efficiency of nucleotide excision repair of cisplatin adducts from native nucleosomes is at least two-fold higher than from expressed nucleosomes. Dual Incision Lanes 1 and 2: NER results for nucleosomes reconstituted from expressed histones and 199 GTG-Pt DNA. Lanes 3 and 4: NER results for nucleosomes reconstituted from native, modified histones and 199 GTG-Pt DNA.
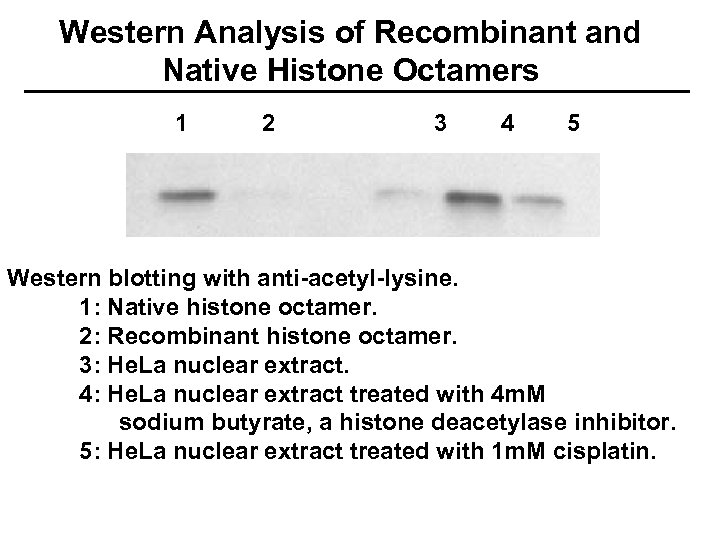

Western Analysis of Recombinant and Native Histone Octamers 1 2 3 4 5 Western blotting with anti-acetyl-lysine. 1: Native histone octamer. 2: Recombinant histone octamer. 3: He. La nuclear extract. 4: He. La nuclear extract treated with 4 m. M sodium butyrate, a histone deacetylase inhibitor. 5: He. La nuclear extract treated with 1 m. M cisplatin.
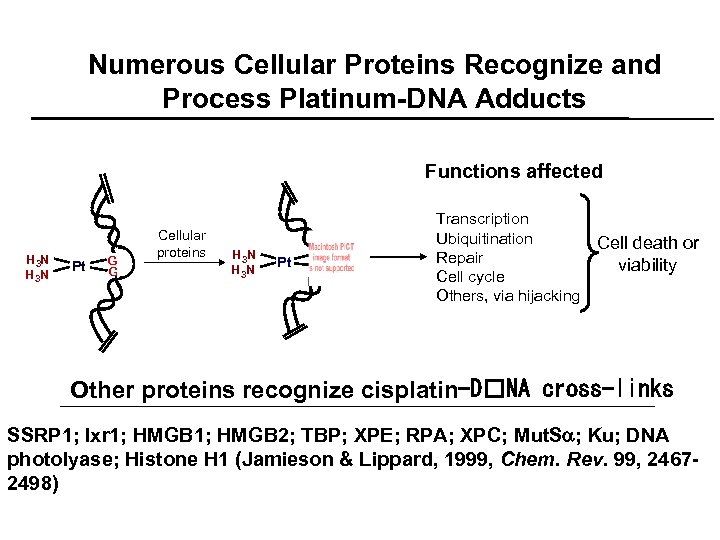
Numerous Cellular Proteins Recognize and Process Platinum-DNA Adducts Functions affected H 3 N Pt GG G Cellular proteins H 3 N Pt G G G Transcription Ubiquitination Repair Cell cycle Others, via hijacking Cell death or viability Other proteins recognize cisplatin-D NA cross-links SSRP 1; Ixr 1; HMGB 2; TBP; XPE; RPA; XPC; Mut. Sa; Ku; DNA photolyase; Histone H 1 (Jamieson & Lippard, 1999, Chem. Rev. 99, 24672498)
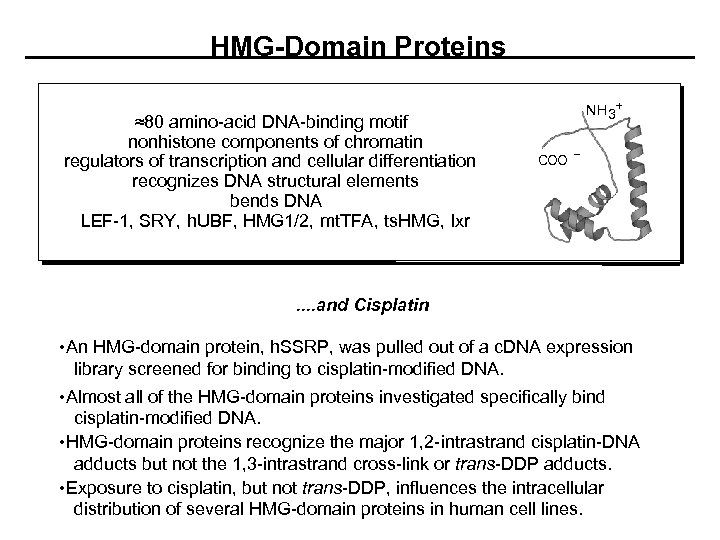
HMG-Domain Proteins ≈80 amino-acid DNA-binding motif nonhistone components of chromatin regulators of transcription and cellular differentiation recognizes DNA structural elements bends DNA LEF-1, SRY, h. UBF, HMG 1/2, mt. TFA, ts. HMG, Ixr NH 3+ COO . . and Cisplatin • An HMG-domain protein, h. SSRP, was pulled out of a c. DNA expression library screened for binding to cisplatin-modified DNA. • Almost all of the HMG-domain proteins investigated specifically bind cisplatin-modified DNA. • HMG-domain proteins recognize the major 1, 2 -intrastrand cisplatin-DNA adducts but not the 1, 3 -intrastrand cross-link or trans-DDP adducts. • Exposure to cisplatin, but not trans-DDP, influences the intracellular distribution of several HMG-domain proteins in human cell lines.

Structure of a Complex of HMGB 1 Domain A with Cisplatin-Modified Duplex DNA HMG-box proteins bind specifically to cisplatin 1, 2 intrastrand cross-links. These major adducts are shielded from nucleotide excision repair in vitro and in vivo. Individual A and B domains of HMGB 1 are responsible for the recognition of cisplatinmodified DNA.
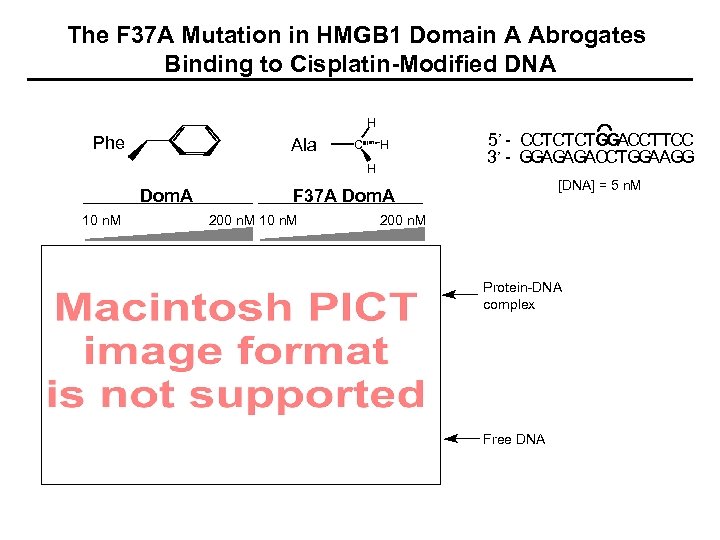
The F 37 A Mutation in HMGB 1 Domain A Abrogates Binding to Cisplatin-Modified DNA H Phe Ala C H H Dom. A 10 n. M 5’ - CCTCTCTGGACCTTCC 3’ - GGAGAGACCTGGAAGG [DNA] = 5 n. M F 37 A Dom. A 200 n. M 10 n. M 200 n. M Protein-DNA complex Free DNA
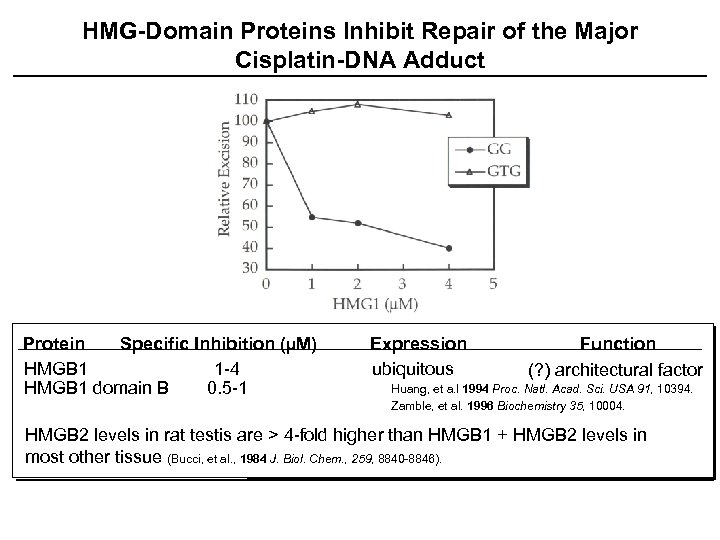
HMG-Domain Proteins Inhibit Repair of the Major Cisplatin-DNA Adduct Protein Specific Inhibition (µM) HMGB 1 1 -4 HMGB 1 domain B 0. 5 -1 Expression ubiquitous Function (? ) architectural factor Huang, et a. l 1994 Proc. Natl. Acad. Sci. USA 91, 10394. Zamble, et al. 1996 Biochemistry 35, 10004. HMGB 2 levels in rat testis are > 4 -fold higher than HMGB 1 + HMGB 2 levels in most other tissue (Bucci, et al. , 1984 J. Biol. Chem. , 259, 8840 -8846).

Repair Shielding by HMG-Domain Protein Overexpression of an HMG-domain protein may sensitize cells to cisplatin.
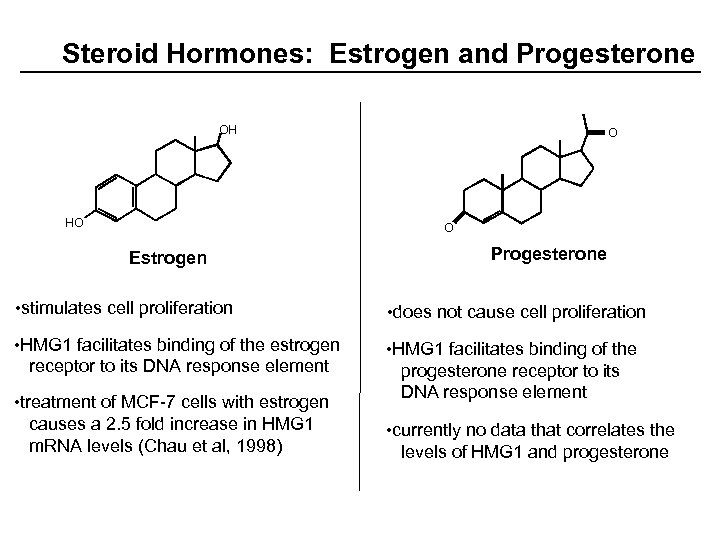
Steroid Hormones: Estrogen and Progesterone OH HO O O Estrogen Progesterone • stimulates cell proliferation • does not cause cell proliferation • HMG 1 facilitates binding of the estrogen receptor to its DNA response element • HMG 1 facilitates binding of the progesterone receptor to its DNA response element • treatment of MCF-7 cells with estrogen causes a 2. 5 fold increase in HMG 1 m. RNA levels (Chau et al, 1998) • currently no data that correlates the levels of HMG 1 and progesterone
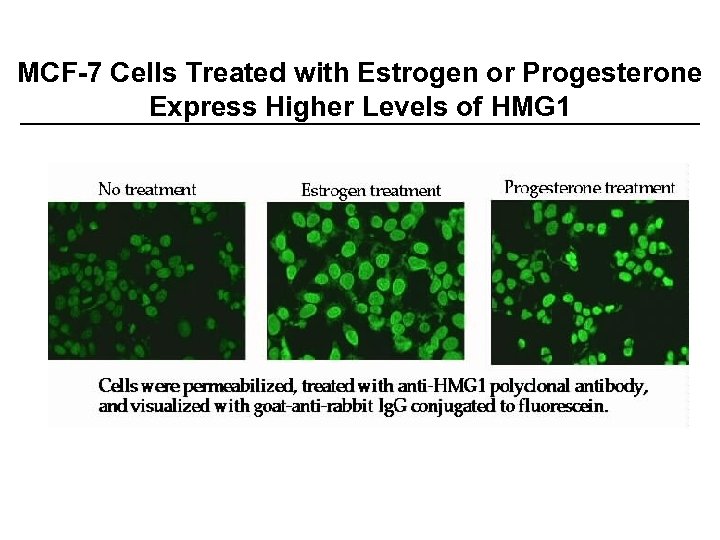
MCF-7 Cells Treated with Estrogen or Progesterone Express Higher Levels of HMG 1
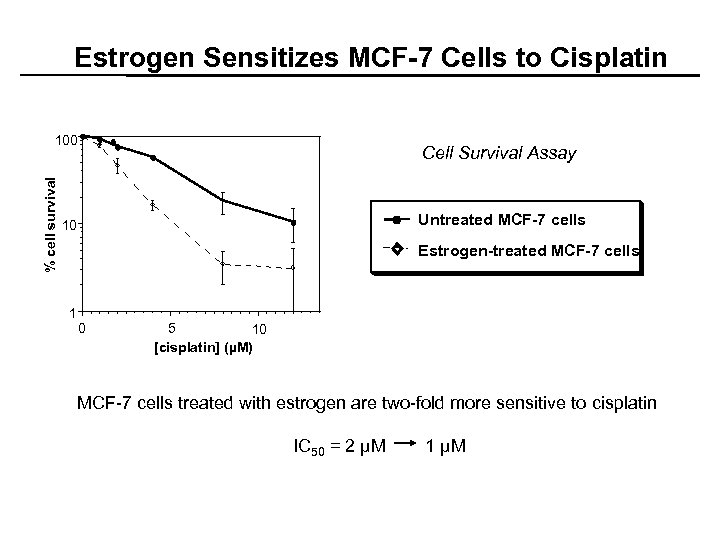
Estrogen Sensitizes MCF-7 Cells to Cisplatin % cell survival 100 Cell Survival Assay Untreated MCF-7 cells 10 Estrogen-treated MCF-7 cells 1 0 5 10 [cisplatin] (µM) MCF-7 cells treated with estrogen are two-fold more sensitive to cisplatin IC 50 = 2 µM 1 µM
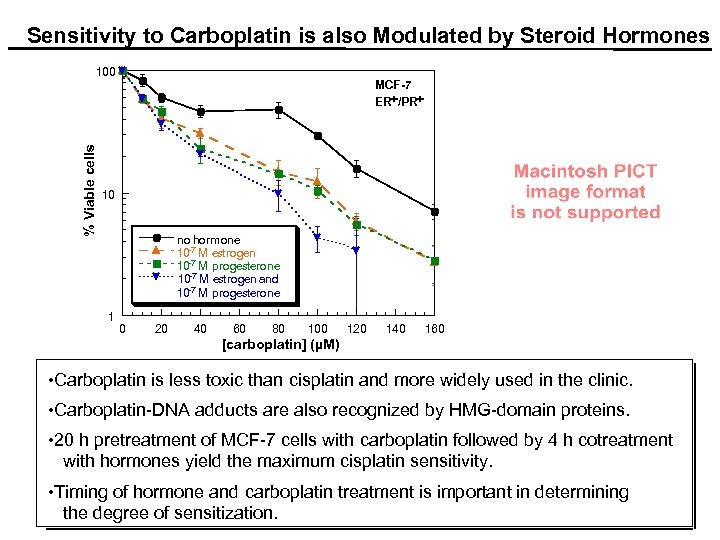
. Sensitivity to Carboplatin is also Modulated by Steroid Hormones % Viable cells 100 MCF-7 ER+/PR+ 10 no hormone 10 -7 M estrogen 10 -7 M progesterone 10 -7 M estrogen and 10 -7 M progesterone 1 0 20 40 60 80 100 120 140 160 [carboplatin] (µM) • Carboplatin is less toxic than cisplatin and more widely used in the clinic. • Carboplatin-DNA adducts are also recognized by HMG-domain proteins. • 20 h pretreatment of MCF-7 cells with carboplatin followed by 4 h cotreatment with hormones yield the maximum cisplatin sensitivity. • Timing of hormone and carboplatin treatment is important in determining the degree of sensitization.
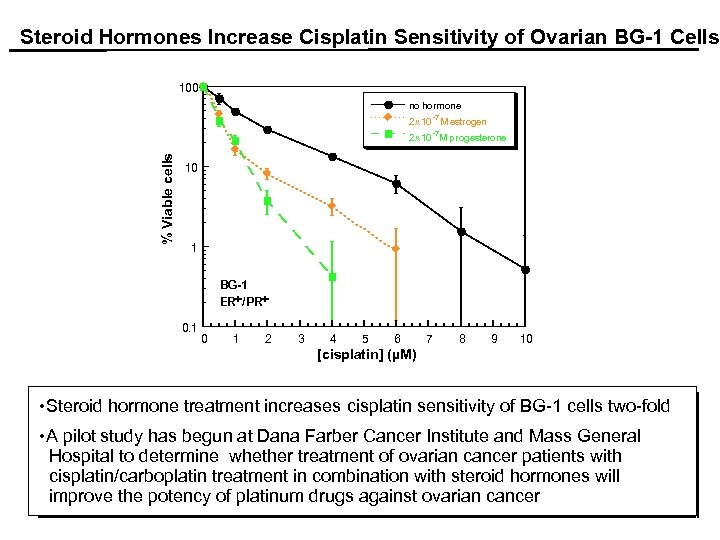
Steroid Hormones Increase Cisplatin Sensitivity of Ovarian BG-1 Cells. 100 no hormone -7 2 x 10 M estrogen -7 % Viable cells 2 x 10 M progesterone 10 1 BG-1 ER+/PR+ 0. 1 0 1 2 3 4 5 6 7 8 9 10 [cisplatin] (µM) • Steroid hormone treatment increases cisplatin sensitivity of BG-1 cells two-fold • A pilot study has begun at Dana Farber Cancer Institute and Mass General Hospital to determine whether treatment of ovarian cancer patients with cisplatin/carboplatin treatment in combination with steroid hormones will improve the potency of platinum drugs against ovarian cancer

Why Use Pt(IV)? • Pt(IV) complexes are kinetically inert – Facilitates synthetic manipulations – Allows for oral administration • Different pharmacological and pharmaco-kinetic properties – Spectrum of activity – Reduced side effects – Drug resistance – Reduction in vivo to reactive Pt(II)
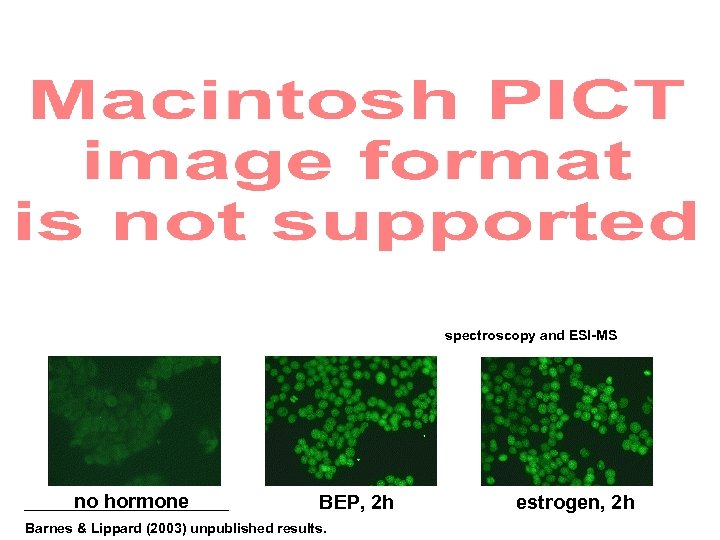
Full characterization by NMR spectroscopy and ESI-MS no hormone BEP, 2 h Barnes & Lippard (2003) unpublished results. estrogen, 2 h

Cytotoxicity Studies: BEP 1 IC 50: 3. 7 M (MCF-7), 3. 8 M (HCC-1937) Thus HMGB 1 overexpression does not sensitize the ER(+) cells. Barnes & Lippard (2003) unpublished results.

BEP 1 Cytotoxicity: Why are ER(+) cells not sensitized compared to the ER(-) cells? • Kinetics of HMGB 1 upregulation are not optimized for repairshielding of cisplatin adducts • Concentration of estrogen delivered to the cell is not suitable for desired HMGB 1 upregulation – Estrogen-induced cell proliferation • Estrogen-compounds derivatized at the 17 -position are not recognized by the estrogen-receptor with strong affinity

Strategy to Address Kinetics Issue: Vary the Length of the Linker to Estrogen Moiety Barnes & Lippard (2003) unpublished results.

Cytotoxicity Studies: BEP 2, BEP 3, BEP 4, BEP 5 Optimal kinetics

Summary of Major Findings Structures of cisplatin-DNA 1, 2 -intrastrand cross-link, and in complex with HMG-domain A, reveal hydrophobic notch and Phe intercalation. Adduct blocks transcription and leads to ubiquitination of RNA Pol II large subunit. Nucleotide excision repair removes the major 1, 2 -intrastrand cross-links; repair is less efficient from nucleosomes. Posttranslational histone modification stimulates NER. Cisplatin treatment of cells stimulates histone acetylation. HMG-domain proteins shield cisplatin intrastrand d(Gp. G) cross-links from nucleotide excision repair. Steroid hormones stimulate HMGB 1 expression and sensitize cells to cisplatin and carboplatin. Phase I clinical trial has commenced at DFCI and MGH. Novel linked Pt(IV) estradiol complex strategy for new drug candidates.

Electron Transfer (ET) in Living Systems PRINCIPLES: • M-binding sites tailored to minimize structural changes upon ET • One-electron transfer processes preferred • Coupling of H+ with electron transfer controls redox potential • ET can occur over long distances; ~ 11 -13 Å is most common • Parameters: distance, driving force, reorganizational energy TOPICS: • Three major bioinorganic ET units: Fen. Sn clusters; Cu; hemes • Long-distance electron transfer: dependence on distance, driving force, reorganization energy • Electron supply in the methane monooxygenase system
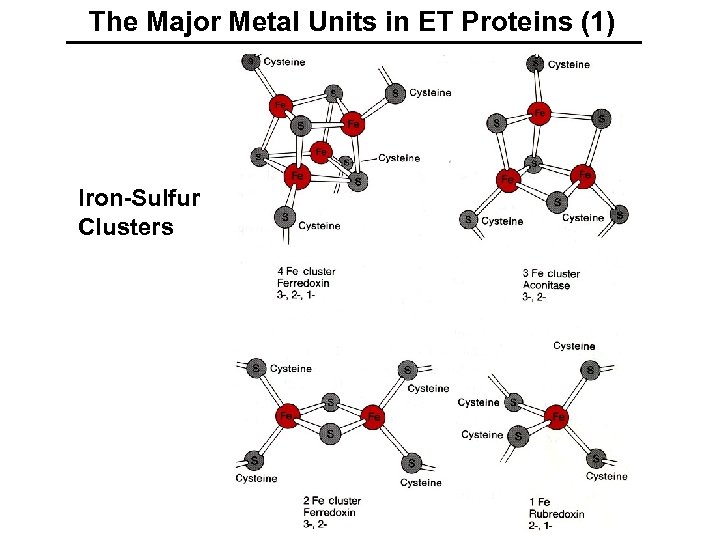
The Major Metal Units in ET Proteins (1) Iron-Sulfur Clusters
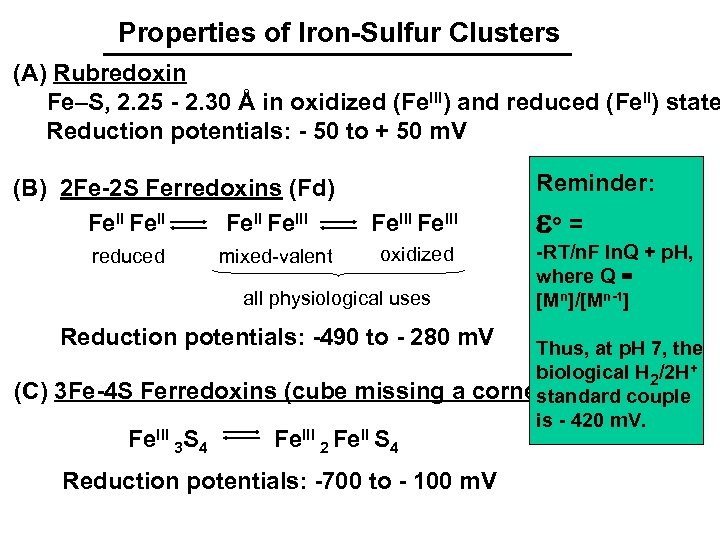

Properties of Iron-Sulfur Clusters (A) Rubredoxin Fe–S, 2. 25 - 2. 30 Å in oxidized (Fe. III) and reduced (Fe. II) state Reduction potentials: - 50 to + 50 m. V (B) 2 Fe-2 S Ferredoxins (Fd) Fe. III reduced mixed-valent Reminder: eo = Fe. III -RT/n. F ln. Q + p. H, where Q = [Mn]/[Mn-1] oxidized all physiological uses Reduction potentials: -490 to - 280 m. V (C) 3 Fe-4 S Ferredoxins (cube missing a Fe. III 3 S 4 Fe. III 2 Fe. II S 4 Thus, at p. H 7, the biological H 2/2 H+ corner) standard couple is - 420 m. V. Reduction potentials: -700 to - 100 m. V
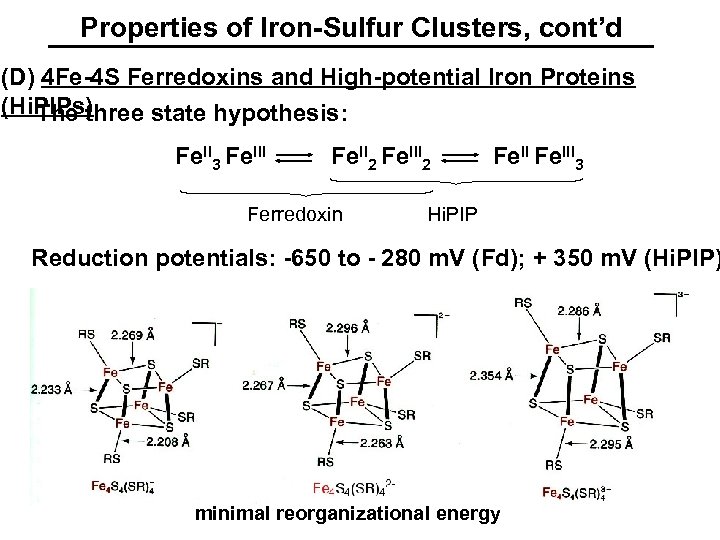
Properties of Iron-Sulfur Clusters, cont’d (D) 4 Fe-4 S Ferredoxins and High-potential Iron Proteins (Hi. PIPs) The three state hypothesis: Fe. II 3 Fe. III Fe. II 2 Fe. III 2 Ferredoxin Fe. III 3 Hi. PIP Reduction potentials: -650 to - 280 m. V (Fd); + 350 m. V (Hi. PIP) minimal reorganizational energy
ae257273e42b8bd8e459f55586866b16.ppt