Phage display analysis of the IgOme.pptx
- Количество слайдов: 49
 Phage display analysis of the Ig. Ome Jonathan M. Gershoni
Phage display analysis of the Ig. Ome Jonathan M. Gershoni

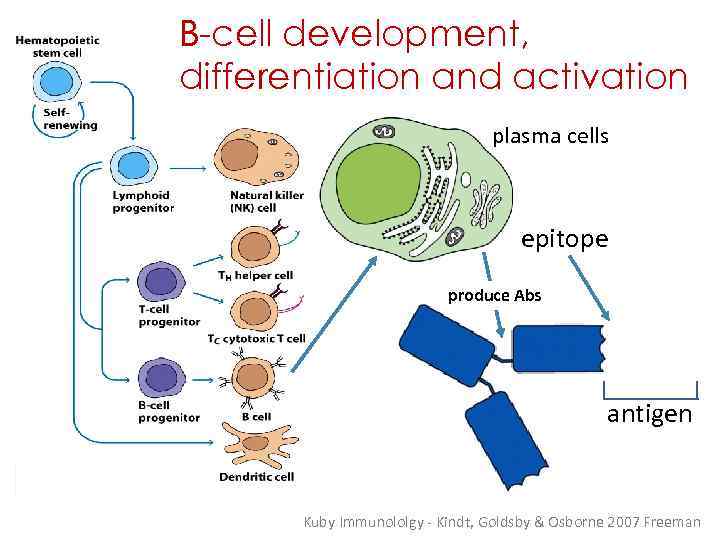 B-cell development, differentiation and activation plasma cells epitope produce Abs antigen Kuby Immunololgy - Kindt, Goldsby & Osborne 2007 Freeman
B-cell development, differentiation and activation plasma cells epitope produce Abs antigen Kuby Immunololgy - Kindt, Goldsby & Osborne 2007 Freeman
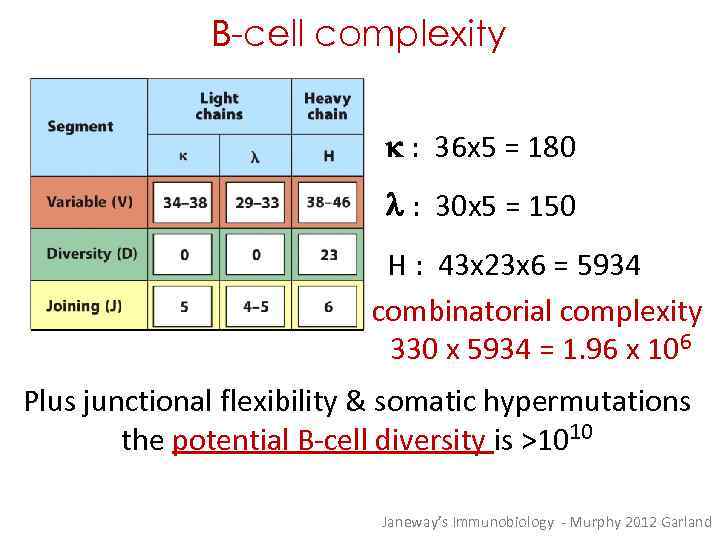 B-cell complexity k : 36 x 5 = 180 l : 30 x 5 = 150 H : 43 x 23 x 6 = 5934 combinatorial complexity 330 x 5934 = 1. 96 x 106 Plus junctional flexibility & somatic hypermutations the potential B-cell diversity is >1010 Janeway’s Immunobiology - Murphy 2012 Garland
B-cell complexity k : 36 x 5 = 180 l : 30 x 5 = 150 H : 43 x 23 x 6 = 5934 combinatorial complexity 330 x 5934 = 1. 96 x 106 Plus junctional flexibility & somatic hypermutations the potential B-cell diversity is >1010 Janeway’s Immunobiology - Murphy 2012 Garland
 Interrogation of the Ig. Ome: two approaches DIRECT -- Physically profile the entire antibody repertoire INDIRECT -- Map the landscape of the epitopes they bind
Interrogation of the Ig. Ome: two approaches DIRECT -- Physically profile the entire antibody repertoire INDIRECT -- Map the landscape of the epitopes they bind
 Mapping single epitopes Prince Charming’s riddle: How do you find the foot corresponding to the internal image of a glass slipper?
Mapping single epitopes Prince Charming’s riddle: How do you find the foot corresponding to the internal image of a glass slipper?
 Mapping single epitopes • Antigen fragmentation • Geysen pepscan • Point mutagenesis • Physical co-crystallization • Survey the universe of random peptides
Mapping single epitopes • Antigen fragmentation • Geysen pepscan • Point mutagenesis • Physical co-crystallization • Survey the universe of random peptides
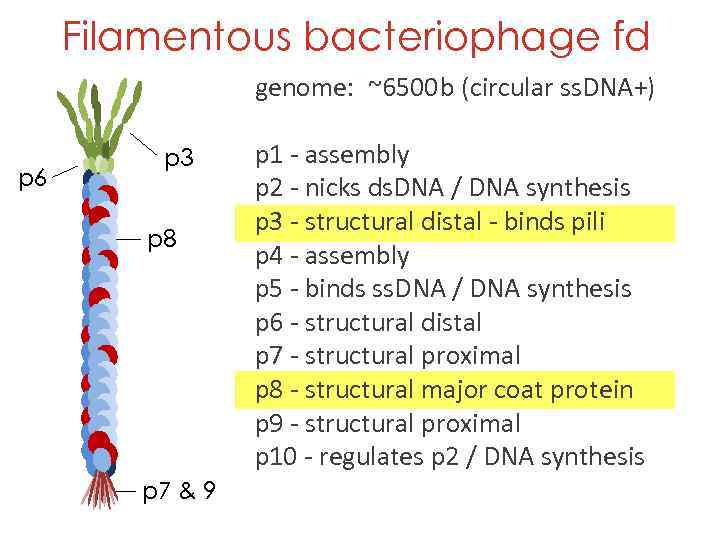 Filamentous bacteriophage fd genome: ~6500 b (circular ss. DNA+) p 6 p 3 p 8 p 7 & 9 p 1 - assembly p 2 - nicks ds. DNA / DNA synthesis p 3 - structural distal - binds pili p 4 - assembly p 5 - binds ss. DNA / DNA synthesis p 6 - structural distal p 7 - structural proximal p 8 - structural major coat protein p 9 - structural proximal p 10 - regulates p 2 / DNA synthesis
Filamentous bacteriophage fd genome: ~6500 b (circular ss. DNA+) p 6 p 3 p 8 p 7 & 9 p 1 - assembly p 2 - nicks ds. DNA / DNA synthesis p 3 - structural distal - binds pili p 4 - assembly p 5 - binds ss. DNA / DNA synthesis p 6 - structural distal p 7 - structural proximal p 8 - structural major coat protein p 9 - structural proximal p 10 - regulates p 2 / DNA synthesis
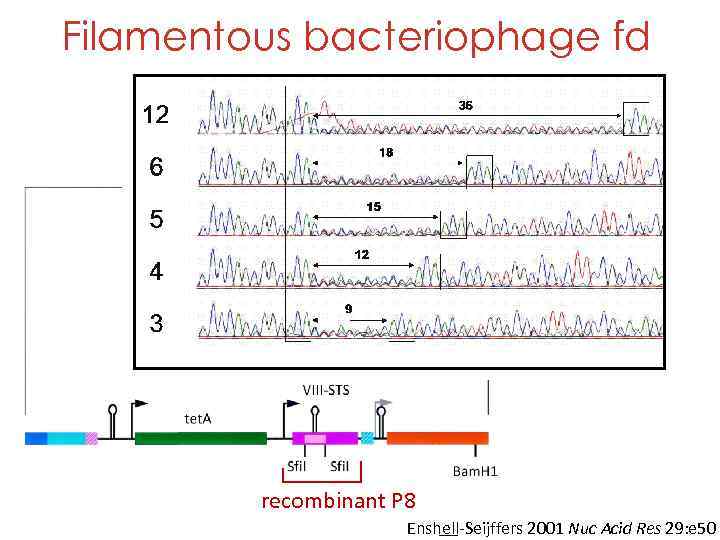 Filamentous bacteriophage fd NNK: N=GATC K=GT (UAA, UGA) Sup. E: UAG=Q recombinant P 8 Enshell-Seijffers 2001 Nuc Acid Res 29: e 50
Filamentous bacteriophage fd NNK: N=GATC K=GT (UAA, UGA) Sup. E: UAG=Q recombinant P 8 Enshell-Seijffers 2001 Nuc Acid Res 29: e 50
 Like panning for gold
Like panning for gold
 “Biopanning” 1 Apply m. Ab to plate 2 Apply phage library 3 Wash away unbound phages 4 Elute affinity selected phages
“Biopanning” 1 Apply m. Ab to plate 2 Apply phage library 3 Wash away unbound phages 4 Elute affinity selected phages
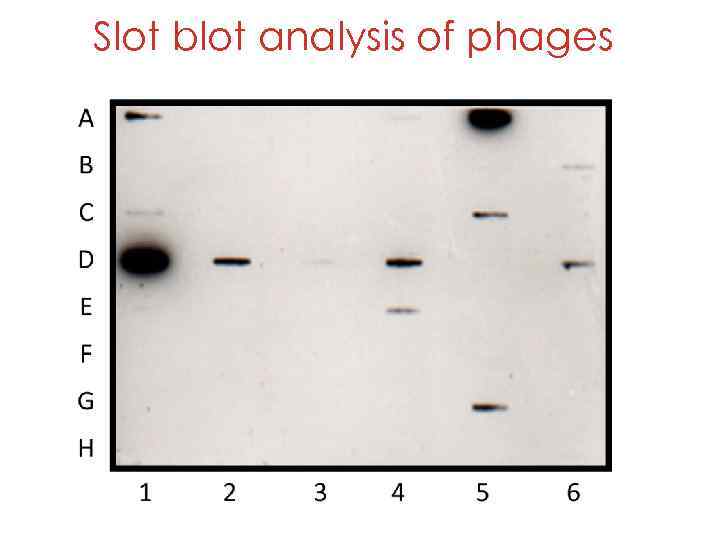 Slot blot analysis of phages
Slot blot analysis of phages
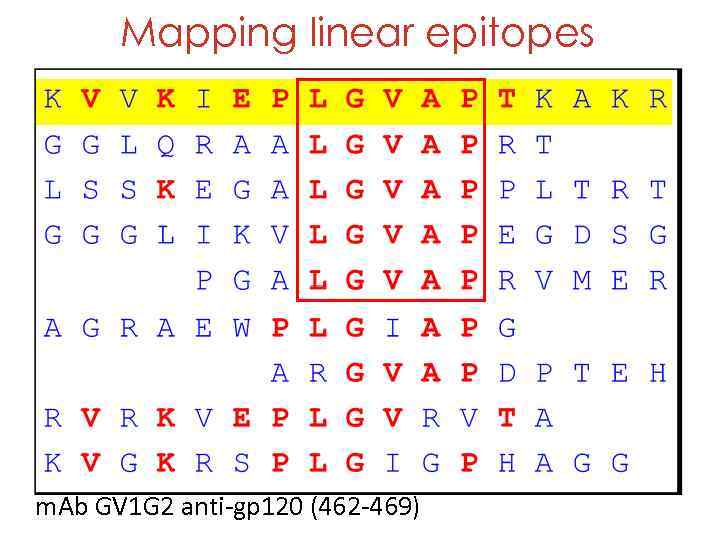 Mapping linear epitopes m. Ab GV 1 G 2 anti-gp 120 (462 -469)
Mapping linear epitopes m. Ab GV 1 G 2 anti-gp 120 (462 -469)
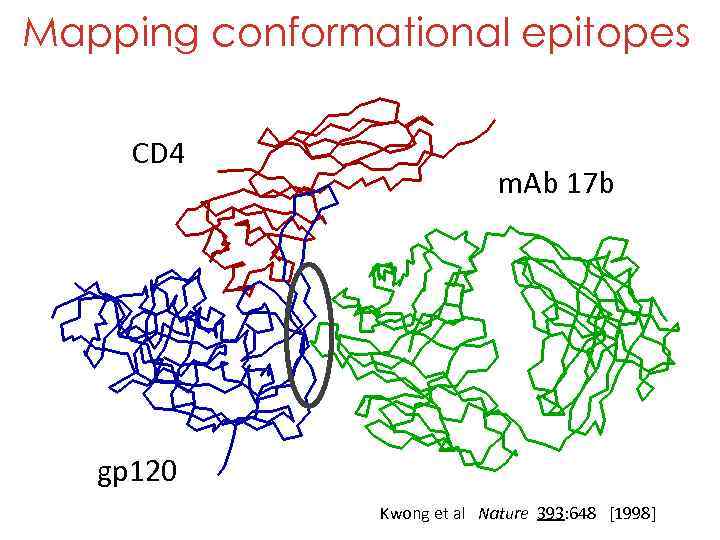 Mapping conformational epitopes CD 4 m. Ab 17 b gp 120 Kwong et al Nature 393: 648 [1998]
Mapping conformational epitopes CD 4 m. Ab 17 b gp 120 Kwong et al Nature 393: 648 [1998]
 Screen m. Abs against phage display combinatorial random peptide libraries m. Ab 17 b
Screen m. Abs against phage display combinatorial random peptide libraries m. Ab 17 b
 17 b specific phage displayed peptides Premise #1: the peptides collectively represent the epitope… somehow?
17 b specific phage displayed peptides Premise #1: the peptides collectively represent the epitope… somehow?
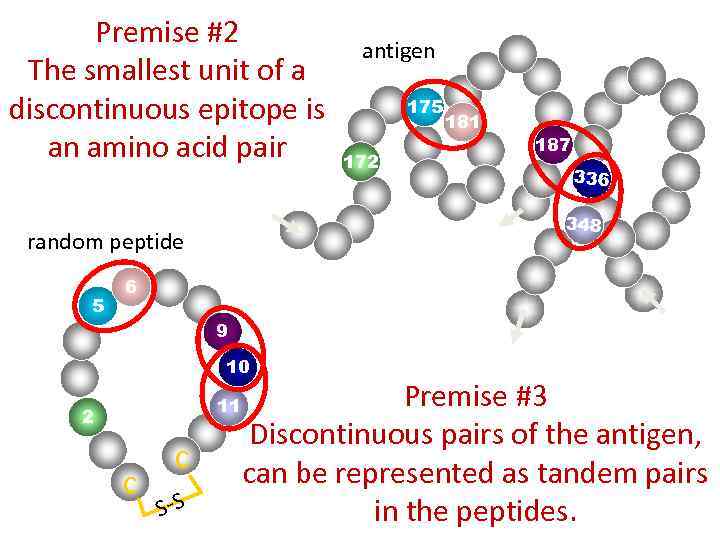 Premise #2 The smallest unit of a discontinuous epitope is an amino acid pair 175 172 181 187 336 348 random peptide 5 antigen 6 9 10 11 2 c c S-S Premise #3 Discontinuous pairs of the antigen, can be represented as tandem pairs in the peptides.
Premise #2 The smallest unit of a discontinuous epitope is an amino acid pair 175 172 181 187 336 348 random peptide 5 antigen 6 9 10 11 2 c c S-S Premise #3 Discontinuous pairs of the antigen, can be represented as tandem pairs in the peptides.
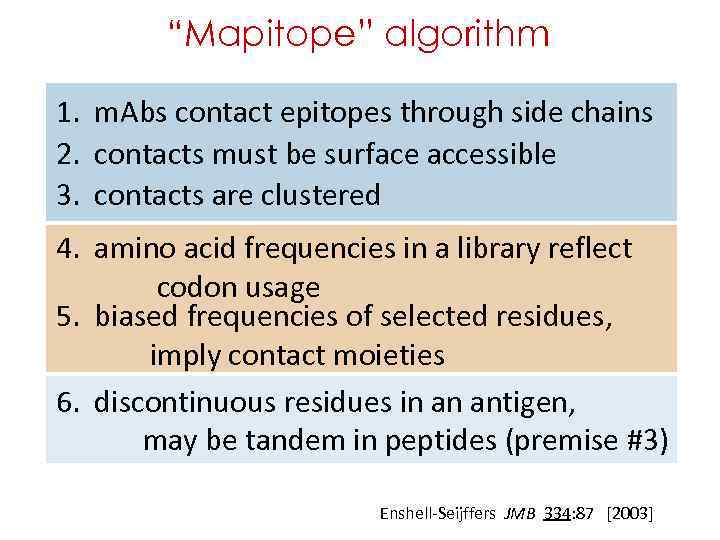 “Mapitope” algorithm 1. m. Abs contact epitopes through side chains 2. contacts must be surface accessible 3. contacts are clustered 4. amino acid frequencies in a library reflect codon usage 5. biased frequencies of selected residues, imply contact moieties 6. discontinuous residues in an antigen, may be tandem in peptides (premise #3) Enshell-Seijffers JMB 334: 87 [2003]
“Mapitope” algorithm 1. m. Abs contact epitopes through side chains 2. contacts must be surface accessible 3. contacts are clustered 4. amino acid frequencies in a library reflect codon usage 5. biased frequencies of selected residues, imply contact moieties 6. discontinuous residues in an antigen, may be tandem in peptides (premise #3) Enshell-Seijffers JMB 334: 87 [2003]
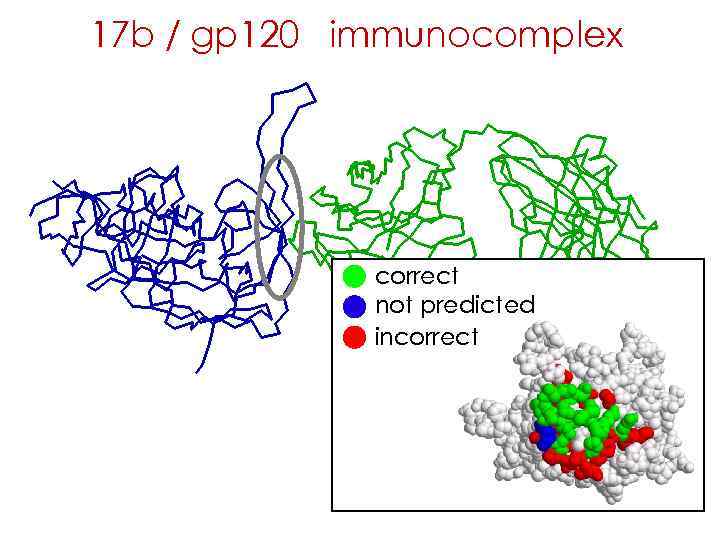 17 b / gp 120 immunocomplex correct not predicted incorrect
17 b / gp 120 immunocomplex correct not predicted incorrect
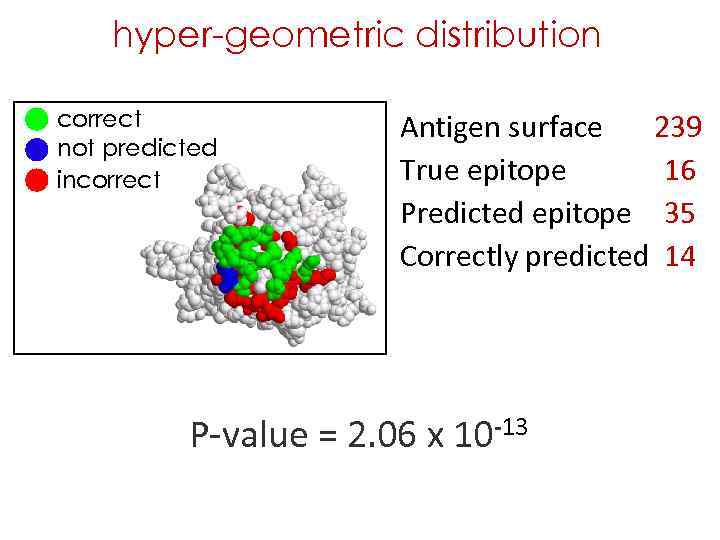 hyper-geometric distribution correct not predicted incorrect Antigen surface 239 True epitope 16 Predicted epitope 35 Correctly predicted 14 P-value = 2. 06 x 10 -13
hyper-geometric distribution correct not predicted incorrect Antigen surface 239 True epitope 16 Predicted epitope 35 Correctly predicted 14 P-value = 2. 06 x 10 -13
 Polyclonal serum is a vast collection of m. Abs
Polyclonal serum is a vast collection of m. Abs
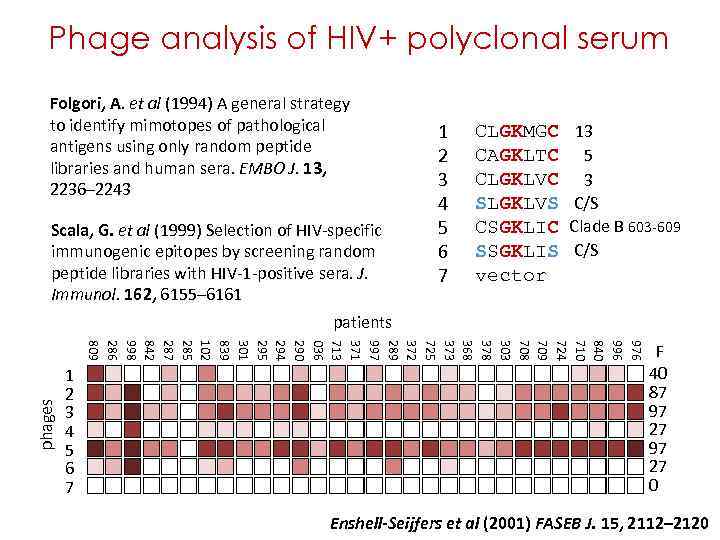 polyclonal serum Phage analysis of HIV+ polyclonal serum Folgori, A. et al (1994) A general strategy to identify mimotopes of pathological antigens using only random peptide libraries and human sera. EMBO J. 13, 2236– 2243 CLGKMGC 1 CAGKLTC 2 CLGKLVC 3 SLGKLVS 4 5 Scala, G. et al (1999) Selection of HIV-specific CSGKLIC immunogenic epitopes by screening random SSGKLIS 6 peptide libraries with HIV-1 -positive sera. J. 7 vector 13 5 3 C/S Clade B 603 -609 C/S Immunol. 162, 6155– 6161 patients phages 976 996 840 710 724 709 708 303 378 368 373 725 372 289 997 371 713 036 290 294 295 301 839 102 285 287 842 998 286 809 1 2 3 4 5 6 7 F 40 87 97 27 0 Enshell-Seijfers et al (2001) FASEB J. 15, 2112– 2120
polyclonal serum Phage analysis of HIV+ polyclonal serum Folgori, A. et al (1994) A general strategy to identify mimotopes of pathological antigens using only random peptide libraries and human sera. EMBO J. 13, 2236– 2243 CLGKMGC 1 CAGKLTC 2 CLGKLVC 3 SLGKLVS 4 5 Scala, G. et al (1999) Selection of HIV-specific CSGKLIC immunogenic epitopes by screening random SSGKLIS 6 peptide libraries with HIV-1 -positive sera. J. 7 vector 13 5 3 C/S Clade B 603 -609 C/S Immunol. 162, 6155– 6161 patients phages 976 996 840 710 724 709 708 303 378 368 373 725 372 289 997 371 713 036 290 294 295 301 839 102 285 287 842 998 286 809 1 2 3 4 5 6 7 F 40 87 97 27 0 Enshell-Seijfers et al (2001) FASEB J. 15, 2112– 2120
 Profiling the epitopes of tens of m. Abs requires affinity selection of thousands of peptides Deep Panning combines Phage Display with Next Generation Sequencing
Profiling the epitopes of tens of m. Abs requires affinity selection of thousands of peptides Deep Panning combines Phage Display with Next Generation Sequencing
 Next Generation Sequencing Second Generation – Cyclic DNA Sequencing Balasubramanian, Klenerman and Bentley University of Cambridge United Kingdom Bentley et al Nature 456: 53 -59 [2008]
Next Generation Sequencing Second Generation – Cyclic DNA Sequencing Balasubramanian, Klenerman and Bentley University of Cambridge United Kingdom Bentley et al Nature 456: 53 -59 [2008]
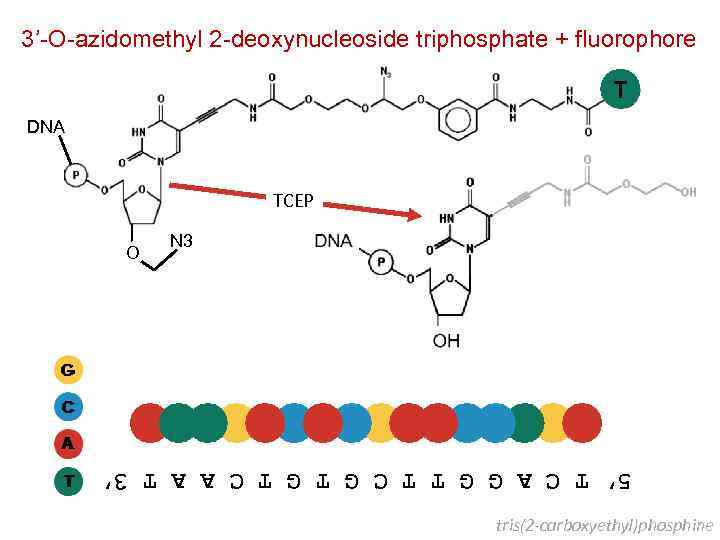 3’-O-azidomethyl 2 -deoxynucleoside triphosphate + fluorophore T DNA TCEP O N 3 5’A T T G A C G A A C C T G A 3’ 5’ T C A G G T T C G T C A A T 3’ tris(2 -carboxyethyl)phosphine
3’-O-azidomethyl 2 -deoxynucleoside triphosphate + fluorophore T DNA TCEP O N 3 5’A T T G A C G A A C C T G A 3’ 5’ T C A G G T T C G T C A A T 3’ tris(2 -carboxyethyl)phosphine
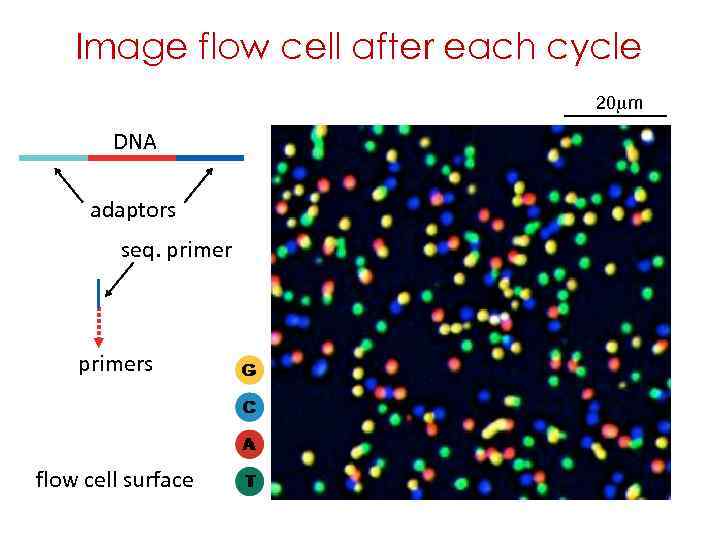 Image flow cell after each cycle 20 mm DNA adaptors seq. primers flow cell surface
Image flow cell after each cycle 20 mm DNA adaptors seq. primers flow cell surface
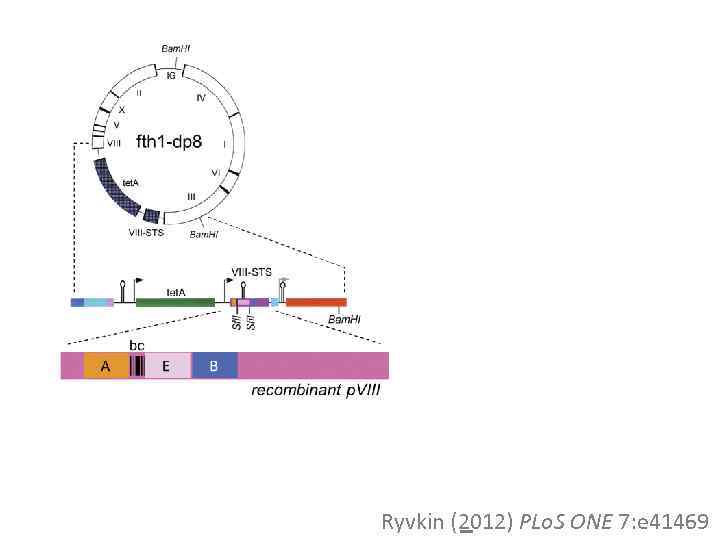 A B 44 aa Ryvkin (2012) PLo. S ONE 7: e 41469
A B 44 aa Ryvkin (2012) PLo. S ONE 7: e 41469
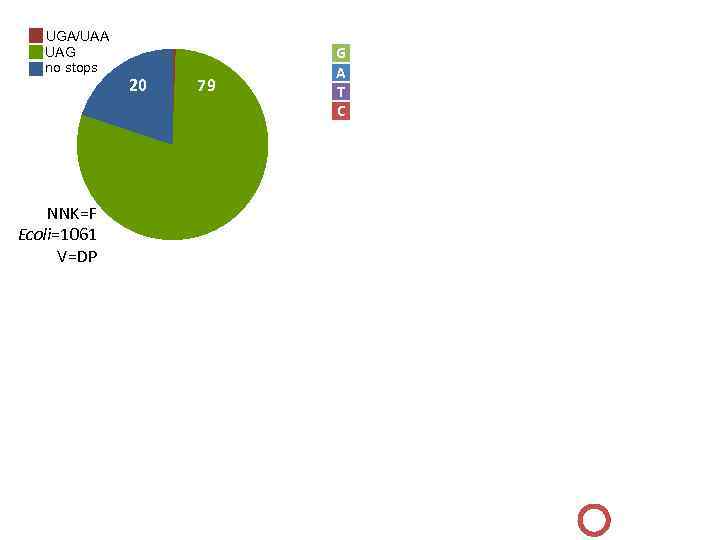 UGA/UAA UAG no stops 1 G A T C 0. 9 20 79 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=1061 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 TAG 0. 16 0. 14 0. 12 GGG 0. 1 0. 08 TGG GTG GAG 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
UGA/UAA UAG no stops 1 G A T C 0. 9 20 79 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=1061 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 TAG 0. 16 0. 14 0. 12 GGG 0. 1 0. 08 TGG GTG GAG 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
 EXPERIMENT 1 - Deep Panning with a m. Ab The random 7 mer peptide library was panned three consecutive times against GV 4 H 3 Samples were taken after Capture 1, 2 and 3 and sent to sequence
EXPERIMENT 1 - Deep Panning with a m. Ab The random 7 mer peptide library was panned three consecutive times against GV 4 H 3 Samples were taken after Capture 1, 2 and 3 and sent to sequence
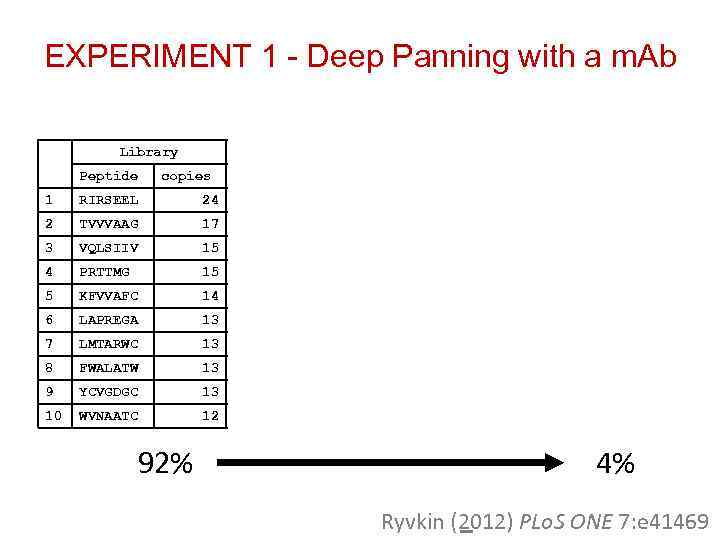 EXPERIMENT 1 - Deep Panning with a m. Ab Library Peptide Capture #1 copies Peptide copies Capture #2 Peptide Capture #3 copies Peptide Copies 1 RIRSEEL 24 ADGIVGW 3105 ADGIVGW 11489 ADGIVGW 65003 2 TVVVAAG 17 LAGAVW 3039 SAGFAME 5173 SAGFAME 6630 3 VQLSIIV 15 SAGFAME 2944 LAAGAVW 3140 VGWAVLE 2409 4 PRTTMG 15 VSLCSSR 2925 VTPHTGF 1172 VTPHTGF 2366 5 KFVVAFC 14 SVTDYVE 2855 GAHVAGG 672 LAAGAVW 2364 6 LAPREGA 13 RMGIRAL 2659 LTACTGF 660 SAGVAME 2242 7 LMTARWC 13 DDDGLDG 1931 LGWAVLD 506 ADGIVGG 1009 8 FWALATW 13 SAARVFM 1723 GTASVGF 444 EVGWAVH 905 9 YCVGDGC 13 QLYGARE 1716 SLGWAVP 355 WVGWAVQ 834 10 WVNAATC 12 NRSREMG 1707 GVGWAVP 344 LGWAVLD 768 92% 4% Ryvkin (2012) PLo. S ONE 7: e 41469
EXPERIMENT 1 - Deep Panning with a m. Ab Library Peptide Capture #1 copies Peptide copies Capture #2 Peptide Capture #3 copies Peptide Copies 1 RIRSEEL 24 ADGIVGW 3105 ADGIVGW 11489 ADGIVGW 65003 2 TVVVAAG 17 LAGAVW 3039 SAGFAME 5173 SAGFAME 6630 3 VQLSIIV 15 SAGFAME 2944 LAAGAVW 3140 VGWAVLE 2409 4 PRTTMG 15 VSLCSSR 2925 VTPHTGF 1172 VTPHTGF 2366 5 KFVVAFC 14 SVTDYVE 2855 GAHVAGG 672 LAAGAVW 2364 6 LAPREGA 13 RMGIRAL 2659 LTACTGF 660 SAGVAME 2242 7 LMTARWC 13 DDDGLDG 1931 LGWAVLD 506 ADGIVGG 1009 8 FWALATW 13 SAARVFM 1723 GTASVGF 444 EVGWAVH 905 9 YCVGDGC 13 QLYGARE 1716 SLGWAVP 355 WVGWAVQ 834 10 WVNAATC 12 NRSREMG 1707 GVGWAVP 344 LGWAVLD 768 92% 4% Ryvkin (2012) PLo. S ONE 7: e 41469
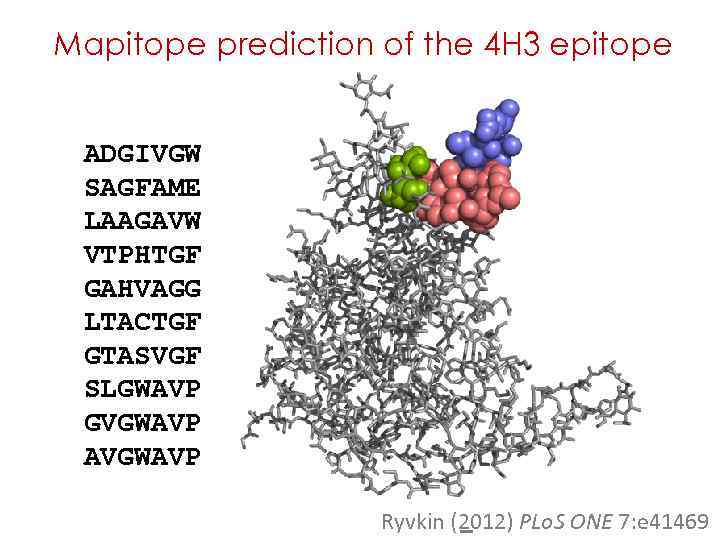 Mapitope prediction of the 4 H 3 epitope ADGIVGW SAGFAME LAAGAVW VTPHTGF GAHVAGG LTACTGF GTASVGF SLGWAVP GVGWAVP AVGWAVP Ryvkin (2012) PLo. S ONE 7: e 41469
Mapitope prediction of the 4 H 3 epitope ADGIVGW SAGFAME LAAGAVW VTPHTGF GAHVAGG LTACTGF GTASVGF SLGWAVP GVGWAVP AVGWAVP Ryvkin (2012) PLo. S ONE 7: e 41469
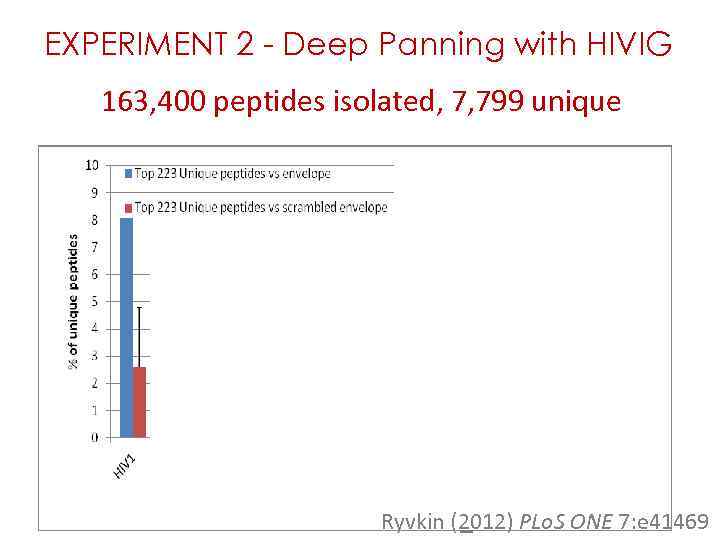 EXPERIMENT 2 - Deep Panning with HIVIG 163, 400 peptides isolated, 7, 799 unique Ryvkin (2012) PLo. S ONE 7: e 41469
EXPERIMENT 2 - Deep Panning with HIVIG 163, 400 peptides isolated, 7, 799 unique Ryvkin (2012) PLo. S ONE 7: e 41469
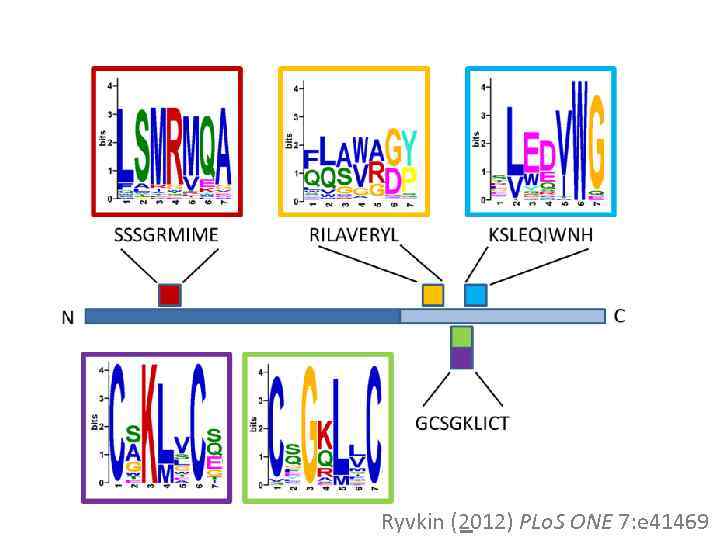 Ryvkin (2012) PLo. S ONE 7: e 41469
Ryvkin (2012) PLo. S ONE 7: e 41469
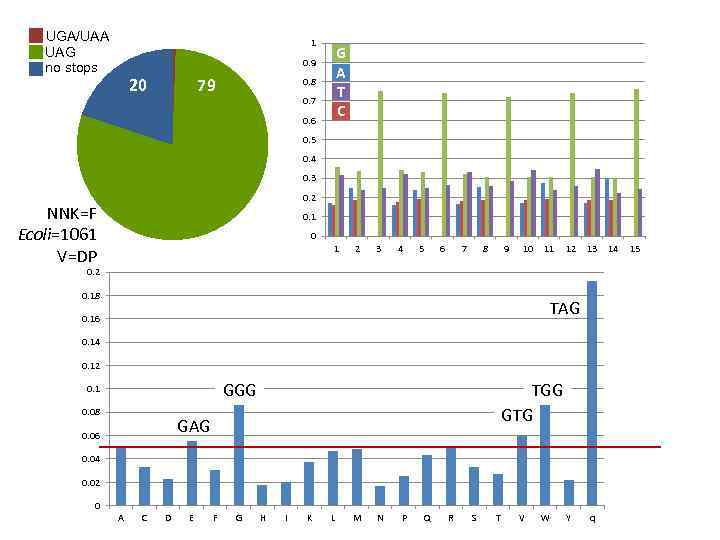 UGA/UAA UAG no stops 1 G A T C 0. 9 20 79 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=1061 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 TAG 0. 16 0. 14 0. 12 GGG 0. 1 0. 08 TGG GTG GAG 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
UGA/UAA UAG no stops 1 G A T C 0. 9 20 79 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=1061 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 TAG 0. 16 0. 14 0. 12 GGG 0. 1 0. 08 TGG GTG GAG 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
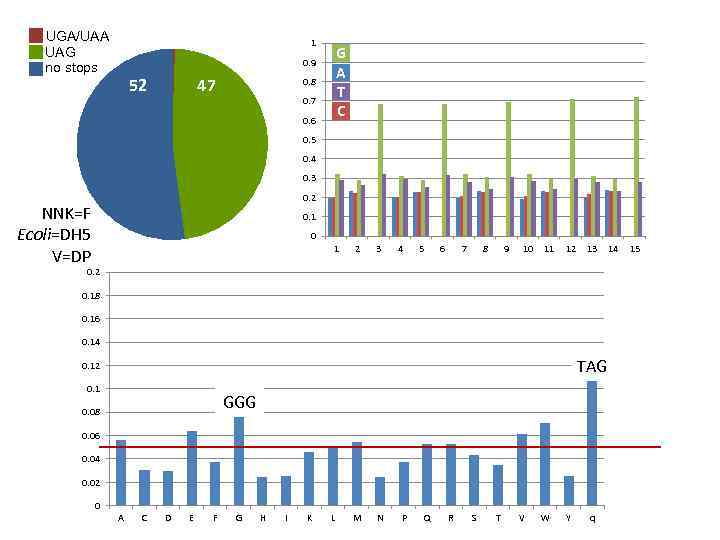 UGA/UAA UAG no stops 1 G A T C 0. 9 52 47 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=DH 5 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 0. 16 0. 14 TAG 0. 12 0. 1 GGG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
UGA/UAA UAG no stops 1 G A T C 0. 9 52 47 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=DH 5 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 0. 16 0. 14 TAG 0. 12 0. 1 GGG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
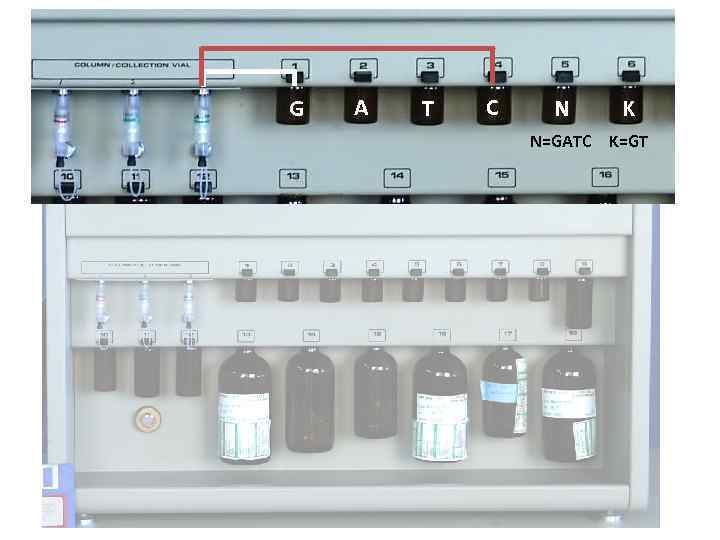 G A T C N K N=GATC K=GT
G A T C N K N=GATC K=GT
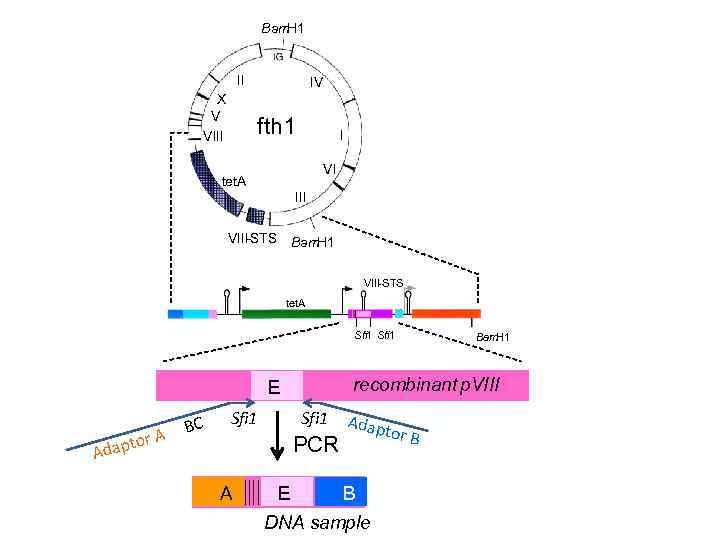 Bam H 1 II X V VIII IV fth 18233 bp tet. A I VI III VIII-STS Bam H 1 VIII-STS tet. A Sfi 1 A A aptor Ad BC B recombinant p. VIII EE Sfi 1 PCR A Bam. H 1 EE Adap tor B B DNA sample
Bam H 1 II X V VIII IV fth 18233 bp tet. A I VI III VIII-STS Bam H 1 VIII-STS tet. A Sfi 1 A A aptor Ad BC B recombinant p. VIII EE Sfi 1 PCR A Bam. H 1 EE Adap tor B B DNA sample
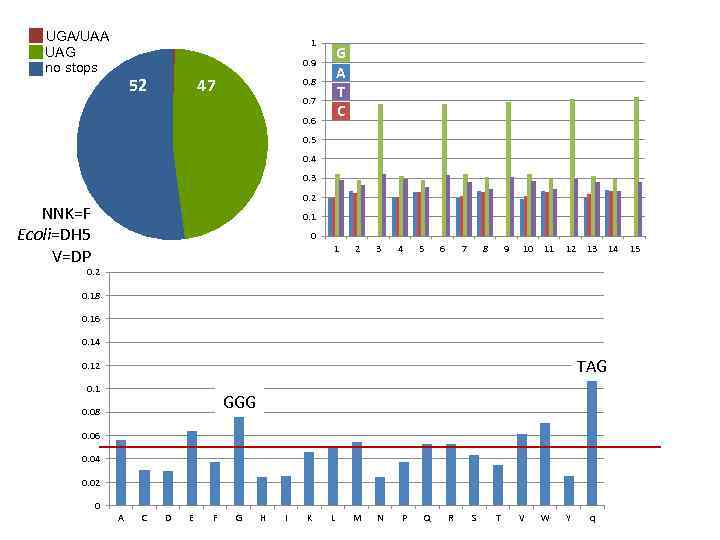 UGA/UAA UAG no stops 1 G A T C 0. 9 52 47 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=DH 5 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 0. 16 0. 14 TAG 0. 12 0. 1 GGG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
UGA/UAA UAG no stops 1 G A T C 0. 9 52 47 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=F Ecoli=DH 5 V=DP 0. 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 0. 2 0. 18 0. 16 0. 14 TAG 0. 12 0. 1 GGG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 14 15
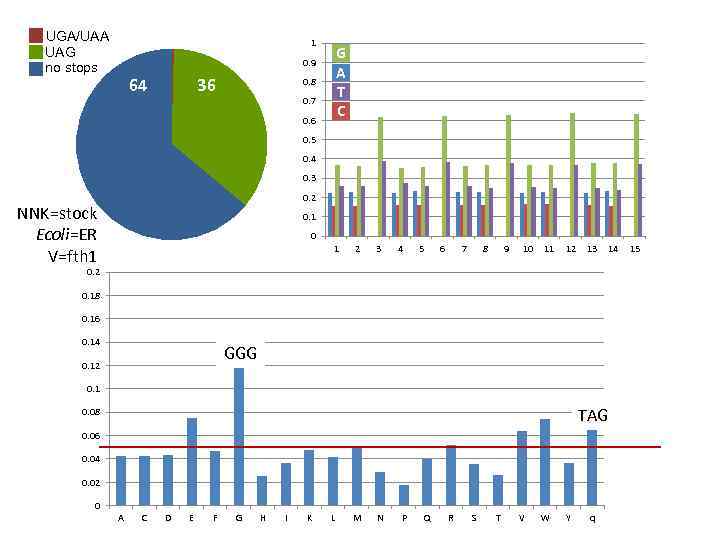 UGA/UAA UAG no stops 1 G A T C 0. 9 64 36 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=stock Ecoli=ER V=fth 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 0. 2 0. 18 0. 16 0. 14 GGG 0. 12 0. 1 TAG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 15
UGA/UAA UAG no stops 1 G A T C 0. 9 64 36 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=stock Ecoli=ER V=fth 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 0. 2 0. 18 0. 16 0. 14 GGG 0. 12 0. 1 TAG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 15
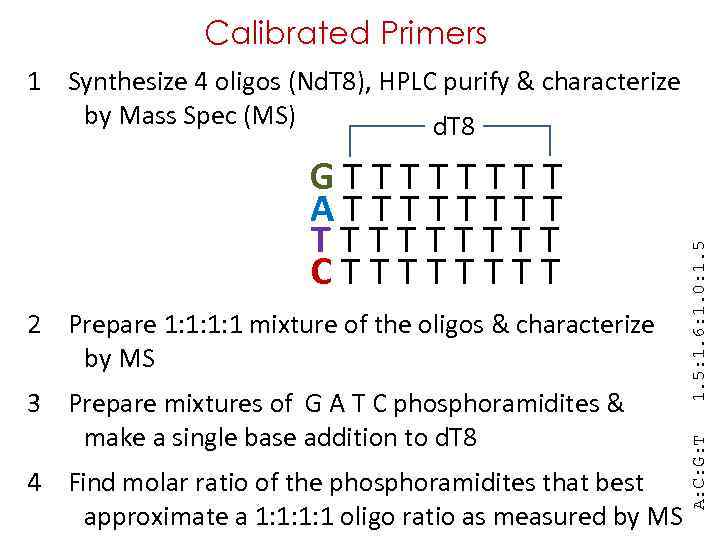 Calibrated Primers G T T T T A T T T T T C T T T T 2 Prepare 1: 1: 1: 1 mixture of the oligos & characterize by MS 3 Prepare mixtures of G A T C phosphoramidites & make a single base addition to d. T 8 4 Find molar ratio of the phosphoramidites that best approximate a 1: 1: 1: 1 oligo ratio as measured by MS A: C: G: T 1. 5: 1. 6: 1. 0: 1. 5 1 Synthesize 4 oligos (Nd. T 8), HPLC purify & characterize by Mass Spec (MS) d. T 8
Calibrated Primers G T T T T A T T T T T C T T T T 2 Prepare 1: 1: 1: 1 mixture of the oligos & characterize by MS 3 Prepare mixtures of G A T C phosphoramidites & make a single base addition to d. T 8 4 Find molar ratio of the phosphoramidites that best approximate a 1: 1: 1: 1 oligo ratio as measured by MS A: C: G: T 1. 5: 1. 6: 1. 0: 1. 5 1 Synthesize 4 oligos (Nd. T 8), HPLC purify & characterize by Mass Spec (MS) d. T 8
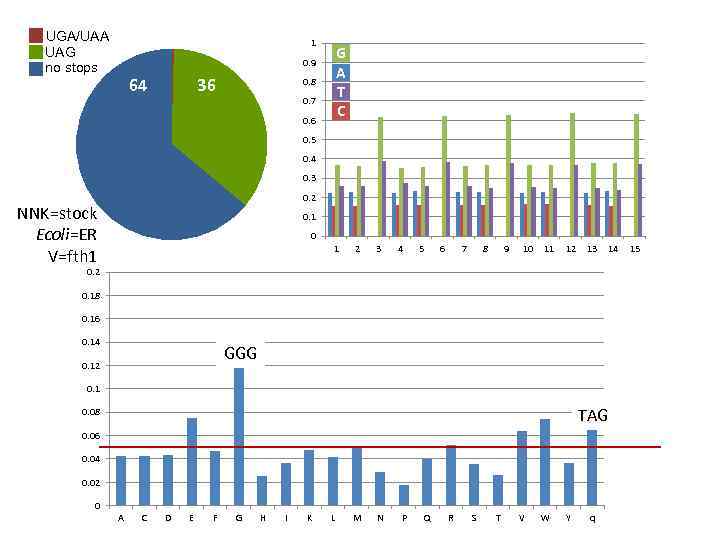 UGA/UAA UAG no stops 1 G A T C 0. 9 64 36 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=stock Ecoli=ER V=fth 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 0. 2 0. 18 0. 16 0. 14 GGG 0. 12 0. 1 TAG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 15
UGA/UAA UAG no stops 1 G A T C 0. 9 64 36 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 NNK=stock Ecoli=ER V=fth 1 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 0. 2 0. 18 0. 16 0. 14 GGG 0. 12 0. 1 TAG 0. 08 0. 06 0. 04 0. 02 0 A C D E F G H I K L M N P Q R S T V W Y q 15
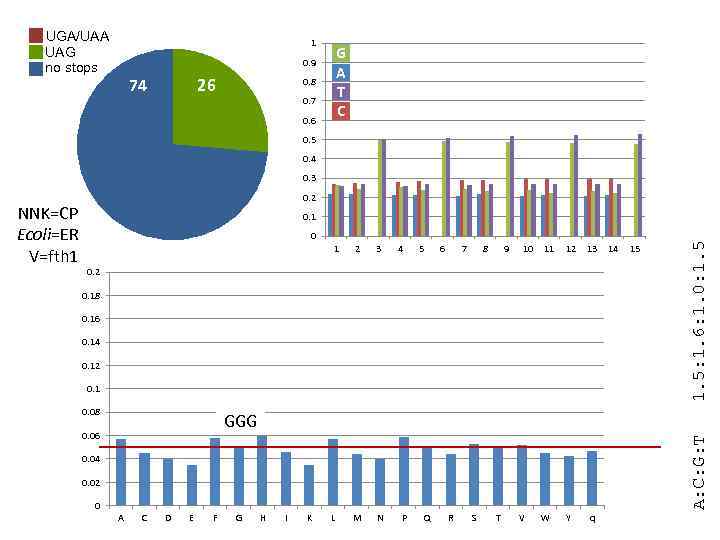 UGA/UAA UAG no stops 1 G A T C 0. 9 74 26 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 1 0 1 2 3 L M N 4 5 6 7 8 9 10 11 12 13 Y q 0. 2 0. 18 0. 16 0. 14 0. 12 0. 1 0. 08 GGG 0. 06 0. 04 0. 02 0 A C D E F G H I K P Q R S T V W 14 15 A: C: G: T 1. 5: 1. 6: 1. 0: 1. 5 NNK=CP Ecoli=ER V=fth 1 0. 2
UGA/UAA UAG no stops 1 G A T C 0. 9 74 26 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 1 0 1 2 3 L M N 4 5 6 7 8 9 10 11 12 13 Y q 0. 2 0. 18 0. 16 0. 14 0. 12 0. 1 0. 08 GGG 0. 06 0. 04 0. 02 0 A C D E F G H I K P Q R S T V W 14 15 A: C: G: T 1. 5: 1. 6: 1. 0: 1. 5 NNK=CP Ecoli=ER V=fth 1 0. 2
 H 5 N 1 avian influenza
H 5 N 1 avian influenza
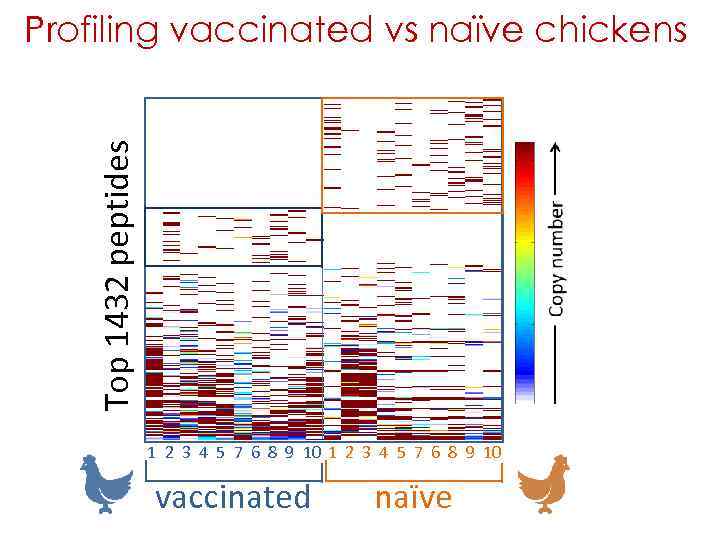
 Profiling vaccinated vs naïve chickens 10 chickens vaccinated with H 5 N 1 were compared with 10 naïve chickens 20 sera were analyzed by 3 rounds of Deep Panning vaccinated 32, 997, 026 naïve 18, 641, 551 The top 100 most frequent peptides were taken for each bird - i. e. , 2000 top peptides Redundant peptides were scored once i. e. , 1432 unique peptides
Profiling vaccinated vs naïve chickens 10 chickens vaccinated with H 5 N 1 were compared with 10 naïve chickens 20 sera were analyzed by 3 rounds of Deep Panning vaccinated 32, 997, 026 naïve 18, 641, 551 The top 100 most frequent peptides were taken for each bird - i. e. , 2000 top peptides Redundant peptides were scored once i. e. , 1432 unique peptides
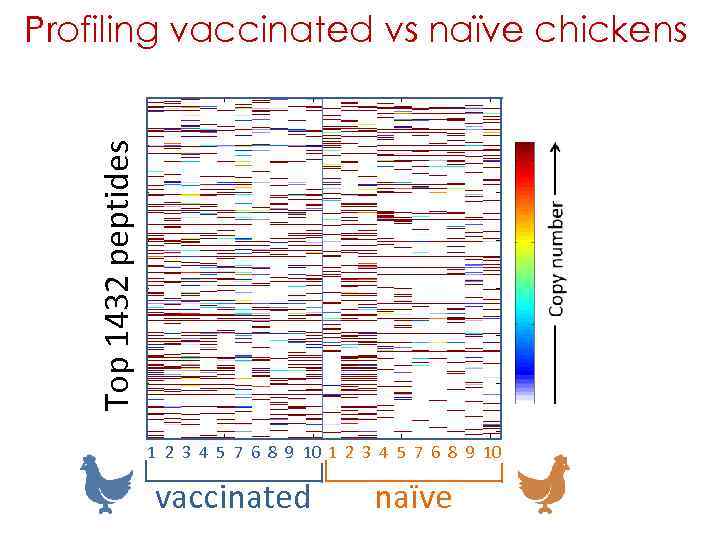 Top 1432 peptides Profiling vaccinated vs naïve chickens 1 2 3 4 5 7 6 8 9 10 vaccinated naïve
Top 1432 peptides Profiling vaccinated vs naïve chickens 1 2 3 4 5 7 6 8 9 10 vaccinated naïve
 Top 1432 peptides Profiling vaccinated vs naïve chickens 1 2 3 4 5 7 6 8 9 10 vaccinated naïve
Top 1432 peptides Profiling vaccinated vs naïve chickens 1 2 3 4 5 7 6 8 9 10 vaccinated naïve
 Conclusions Deep Panning produces meaningful Ig. Ome profiles Profiling the Ig. Ome allows - Diagnosis of infectious disease Multiplex screening of blood transfusions Discovery of the correlates of immunity Monitoring vaccination efficacy
Conclusions Deep Panning produces meaningful Ig. Ome profiles Profiling the Ig. Ome allows - Diagnosis of infectious disease Multiplex screening of blood transfusions Discovery of the correlates of immunity Monitoring vaccination efficacy
 TAU: Dr. Anna Roitburd – Berman Dr. Yael Weiss Ottolenghi Dr. Gilad Kaplan Smadar Neeman Arie Ryvkin Noy Motiee Prof. Tal Pupko Haim Ashkenazy Prof. George E. Lewis -- IHV – U Maryland Prof. Wayne Marasco -- Harvard University Dr. Avishay Lublin -- Kimron Dr. Shimon Perk -- Kimron Dr. David Swayne -- USDA Спасибо o
TAU: Dr. Anna Roitburd – Berman Dr. Yael Weiss Ottolenghi Dr. Gilad Kaplan Smadar Neeman Arie Ryvkin Noy Motiee Prof. Tal Pupko Haim Ashkenazy Prof. George E. Lewis -- IHV – U Maryland Prof. Wayne Marasco -- Harvard University Dr. Avishay Lublin -- Kimron Dr. Shimon Perk -- Kimron Dr. David Swayne -- USDA Спасибо o



