5f93e2bcda59ae60d40c70c04778581a.ppt
- Количество слайдов: 21

PDBe. Chem The Ligand Database EBI is an Outstation of the European Molecular Biology Laboratory.

Why PDBe. Chem? • • Link between protein and chemistry Reference dictionary for the chemical definition of 3 letter coded single residues in the PDB (ww. PDB CCD) Holds the single residue definitions for the PDB and represents it for all other databases at the EBI (PDBe. Chem database) How else will you find ligand structures !!!!! (PDBe. Chem search system)
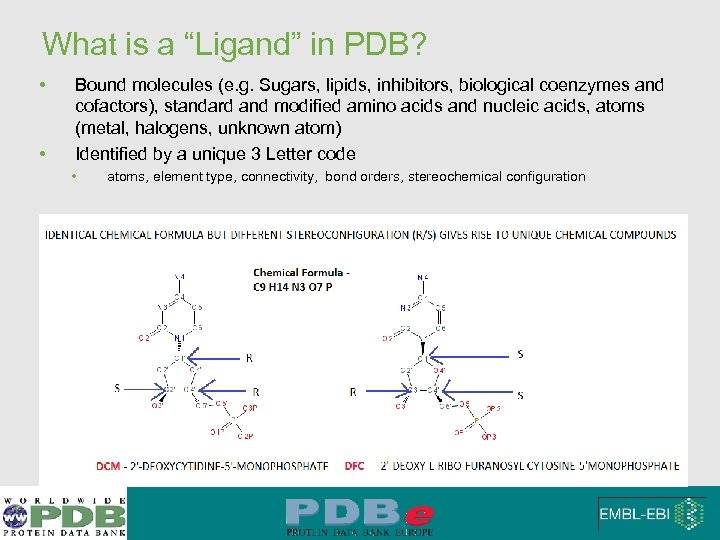

What is a “Ligand” in PDB? • • Bound molecules (e. g. Sugars, lipids, inhibitors, biological coenzymes and cofactors), standard and modified amino acids and nucleic acids, atoms (metal, halogens, unknown atom) Identified by a unique 3 Letter code • atoms, element type, connectivity, bond orders, stereochemical configuration
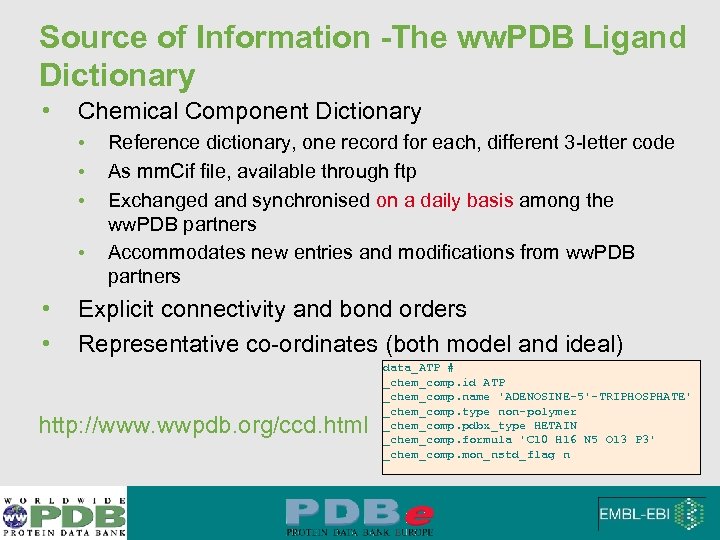

Source of Information -The ww. PDB Ligand Dictionary • Chemical Component Dictionary • • • Reference dictionary, one record for each, different 3 -letter code As mm. Cif file, available through ftp Exchanged and synchronised on a daily basis among the ww. PDB partners Accommodates new entries and modifications from ww. PDB partners Explicit connectivity and bond orders Representative co-ordinates (both model and ideal) http: //www. wwpdb. org/ccd. html data_ATP # _chem_comp. id ATP _chem_comp. name 'ADENOSINE-5'-TRIPHOSPHATE' _chem_comp. type non-polymer _chem_comp. pdbx_type HETAIN _chem_comp. formula 'C 10 H 16 N 5 O 13 P 3' _chem_comp. mon_nstd_flag n
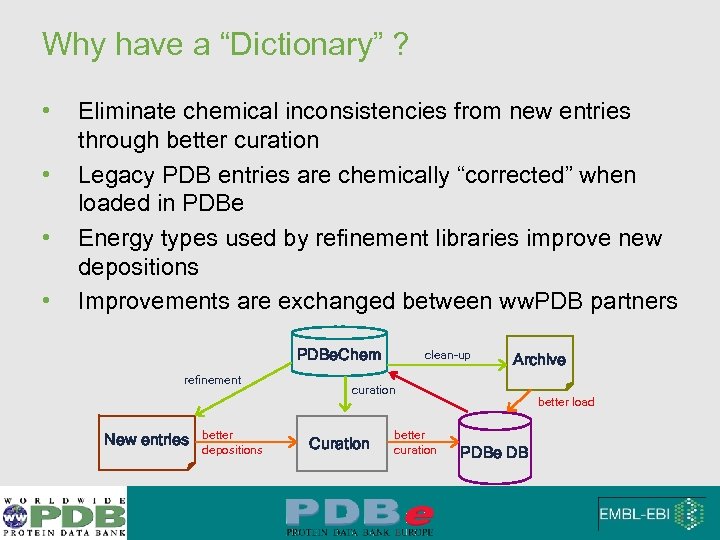

Why have a “Dictionary” ? • • Eliminate chemical inconsistencies from new entries through better curation Legacy PDB entries are chemically “corrected” when loaded in PDBe Energy types used by refinement libraries improve new depositions Improvements are exchanged between ww. PDB partners PDBe. Chem refinement New entries better depositions clean-up Archive curation Curation better curation better load PDBe DB

The PDBe. Chem database • Collection of all chemical species and small molecules in the PDB • • A ligand is a distinct stereo-isomer • • Complete, up to date Atoms and element types Bonds and bond orders Stereo configuration of atoms and bonds in cases of stereoisomers (R/S – E/Z) Names and co-ordinates not fundamental • But there is a consistent set of identifiers • Atoms, bond order, and stereochemistry • Derived properties
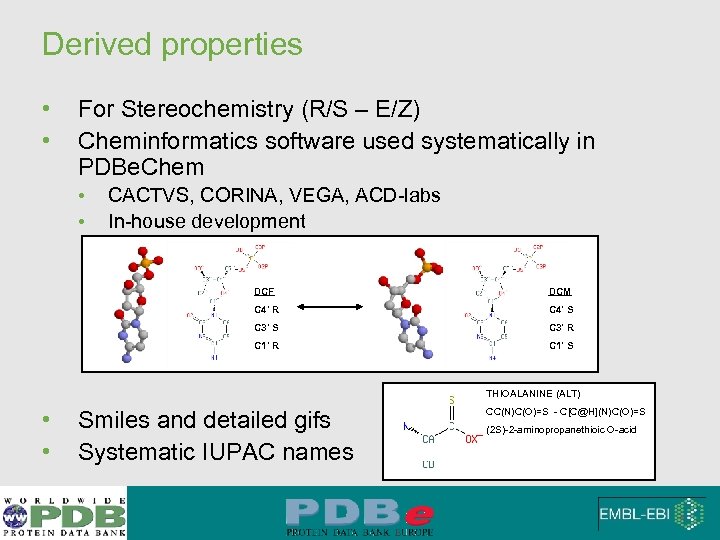
Derived properties • • For Stereochemistry (R/S – E/Z) Cheminformatics software used systematically in PDBe. Chem • • CACTVS, CORINA, VEGA, ACD-labs In-house development DCF DCM C 4' R C 4' S C 3' R C 1' S THIOALANINE (ALT) • • Smiles and detailed gifs Systematic IUPAC names CC(N)C(O)=S - C[C@H](N)C(O)=S (2 S)-2 -aminopropanethioic O-acid

Search options • • By ligand code By ligand name or synonym By formula or formula range By non-stereo smile By exact stereo or nonstereo structure By fingerprint similarity By fragment expression PDBe. Chem Some uses: • Search for drug or ligands. • Understand chemistry of ligand. • Download ‘ideal’ coordinates for own analysis (docking etc) and study.
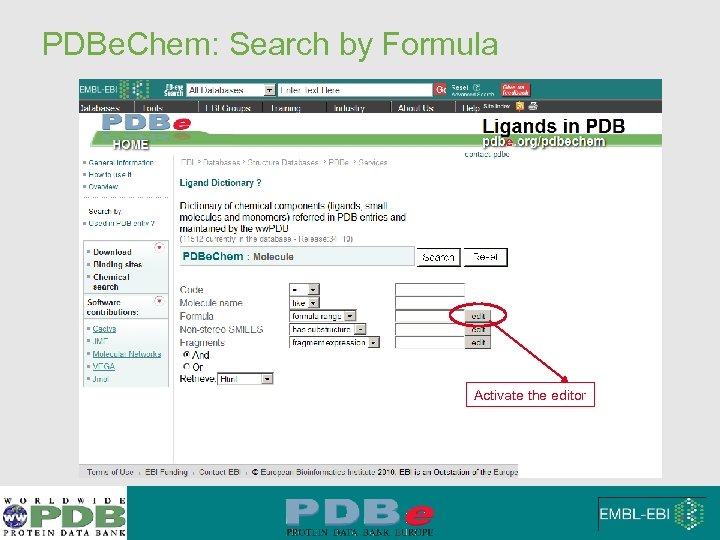
PDBe. Chem: Search by Formula Activate the editor
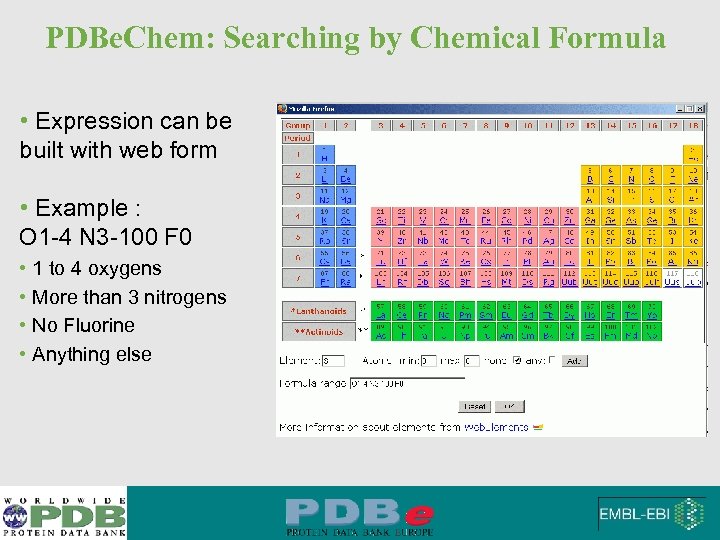

PDBe. Chem: Searching by Chemical Formula • Expression can be built with web form • Example : O 1 -4 N 3 -100 F 0 • 1 to 4 oxygens • More than 3 nitrogens • No Fluorine • Anything else
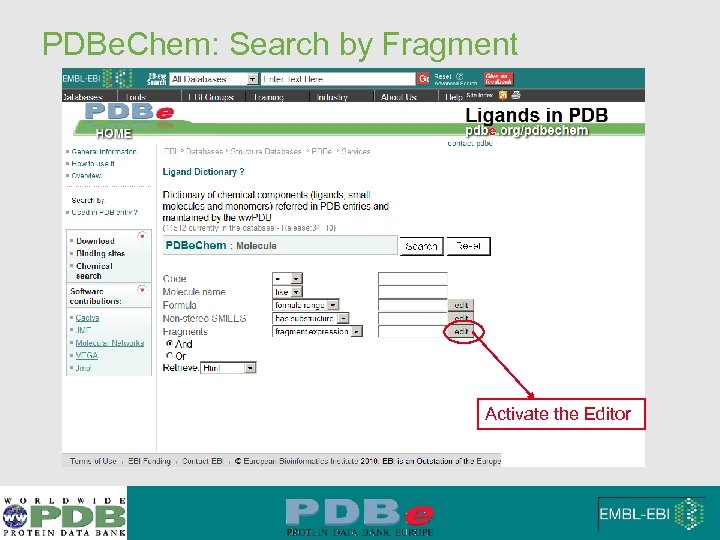
PDBe. Chem: Search by Fragment Activate the Editor
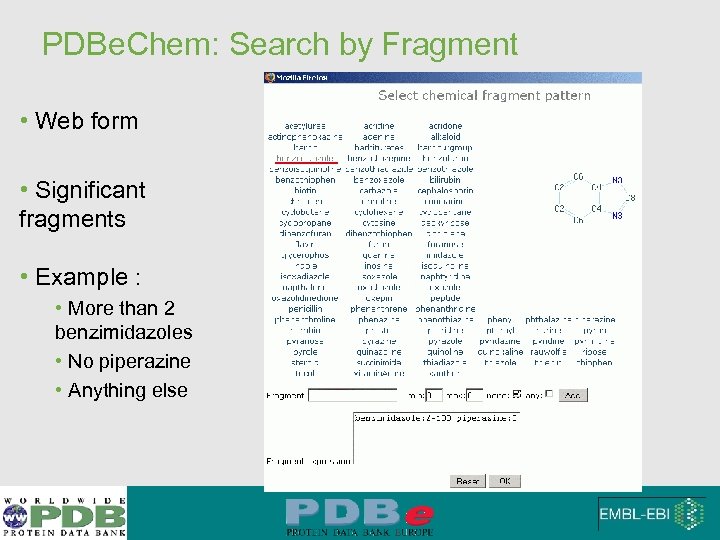

PDBe. Chem: Search by Fragment • Web form • Significant fragments • Example : • More than 2 benzimidazoles • No piperazine • Anything else
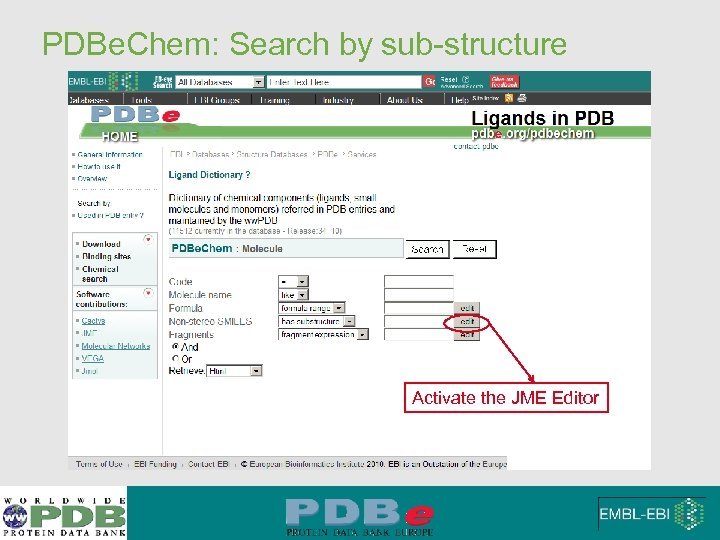
PDBe. Chem: Search by sub-structure Activate the JME Editor
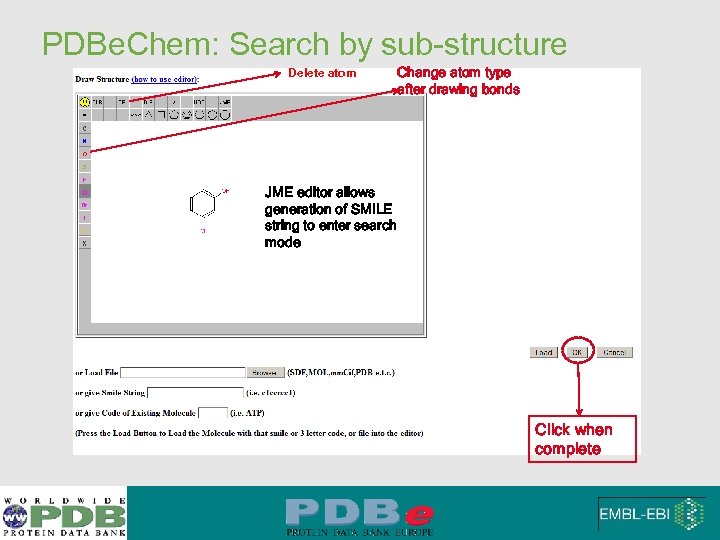

PDBe. Chem: Search by sub-structure Delete atom Change atom type after drawing bonds JME editor allows generation of SMILE string to enter search mode Click when complete
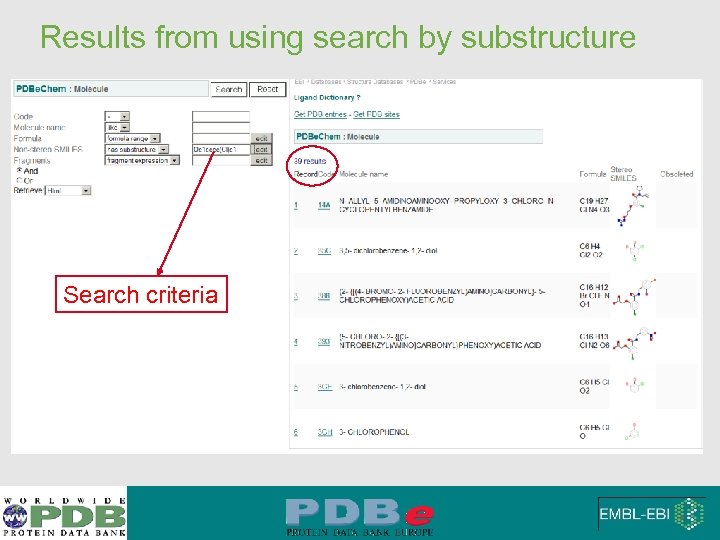
Results from using search by substructure Search criteria
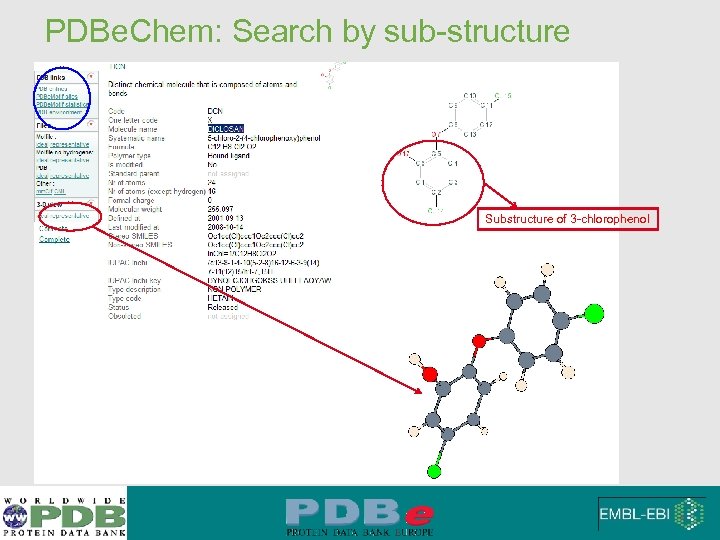
PDBe. Chem: Search by sub-structure Substructure of 3 -chlorophenol
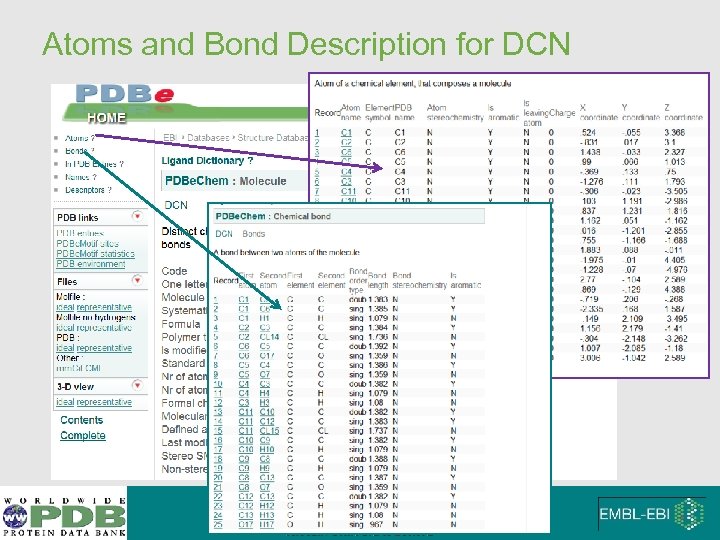

Atoms and Bond Description for DCN
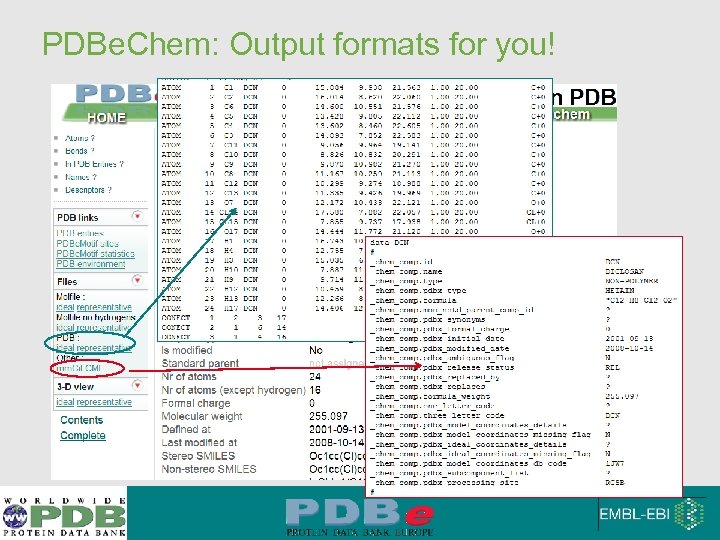

PDBe. Chem: Output formats for you!
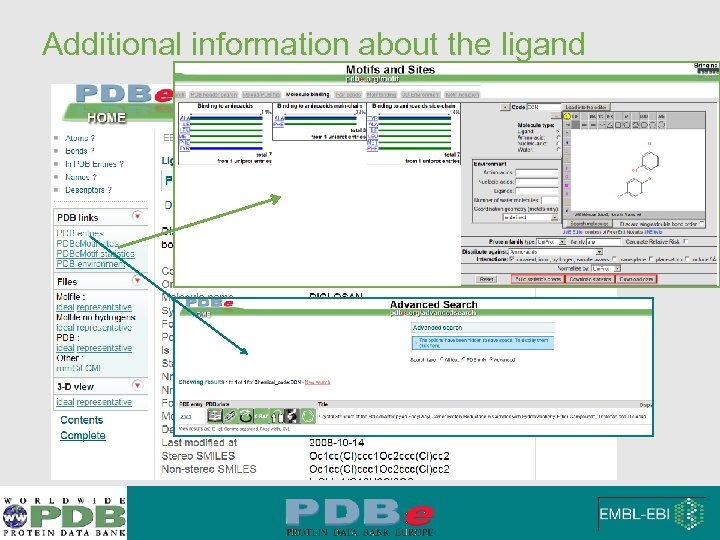
Additional information about the ligand
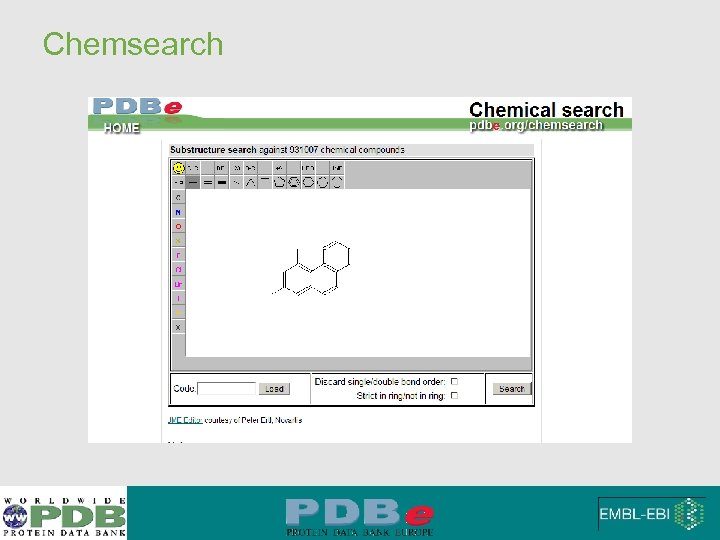
Chemsearch
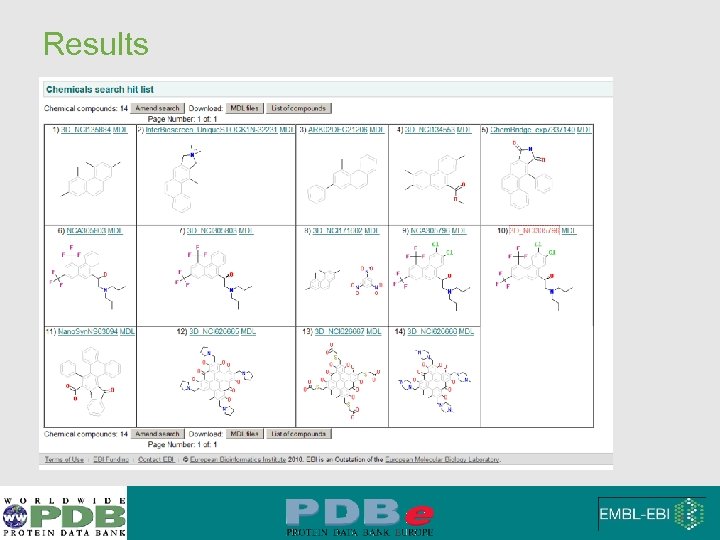
Results
5f93e2bcda59ae60d40c70c04778581a.ppt