7d0e4ad1fb945b072542ab59ae755471.ppt
- Количество слайдов: 42

Pathways of technology adoption and adaptation Saheer Gharbia Applied and Functional Genomics Centre for Infection Health Protection Agency

Methods and approaches to elucidate the spread of infectious diseases are as old as human civilisation and stretch in their logic from magical, philosophical to scientific.

Pheno-, Sero-, phage and biotypes extremely valuable, able to identify cellular components associated with virulence. Limited discriminatory power, resolving pathogens into only a few types. Challenged by Diversity, mutations and acquisition of new traits

Technology in diagnostics is a composite of: • Refining what is established • Exploring what is new (concept or tool) • Adapting the new to the function

Examples of Streamlining: VIDAS®: automated system for Rapid Pathogen Monitoring VITEK® 2 Compact: automated system for microbial identification API® / ID 32/ APIWEB®: microbial identification with internet-base tool TEMPO®: enumeration of quality indicators in food air IDEAL®: aerobiocontamination control Count-Tact™ range: monitoring of surface and air biocontamination Bac. T/ALERT® 3 D microbial detection system Multiplex Fluorescent Immunoassay Ra. PET Rapid Particle Enhanced Technology

Salvador Luria and Max Delbruck (1943) Provided a statistical demonstration that inheritance in bacteria follows Darwinian principles. Mutations occur randomly in bacterial populations. They occur in small numbers in some populations and in large numbers in other cultures. awarded the Nobel Prize in Medicine or Physiology in 1969.

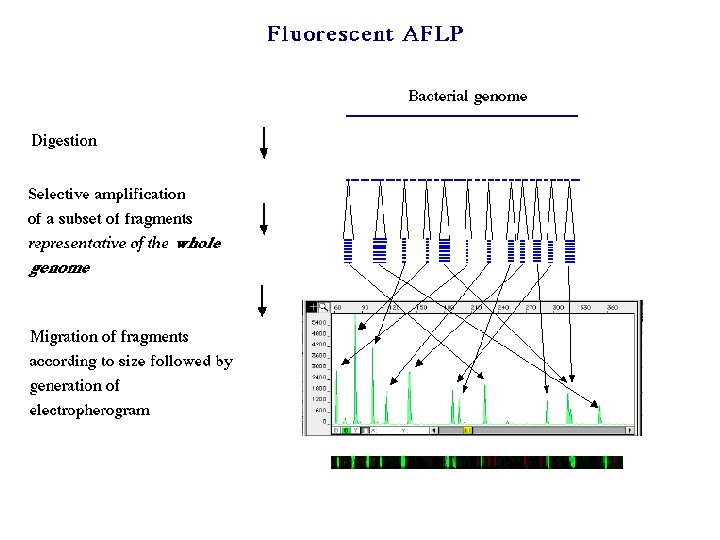
Classical Molecular Genetics for the analysis of infectious agents was a natural follow up especially with the publication of the Watson-Crick DNA model. Gross DNA structure: Restriction Diget, southern hybridisation, RFLP, AFLP, plasmid profiling, PFGE, f. AFLP and ribotyping

PCR lead to the next phase by accelerating the process, adding higher resolution, selective amplification and quantitative measurements

The power of PCR-based methods is the ease with which they can be applied to many bacterial pathogens and their multilocus discrimination. Such methods have proven valuable for genetic dissection of pathogens for which previous methods have failed.
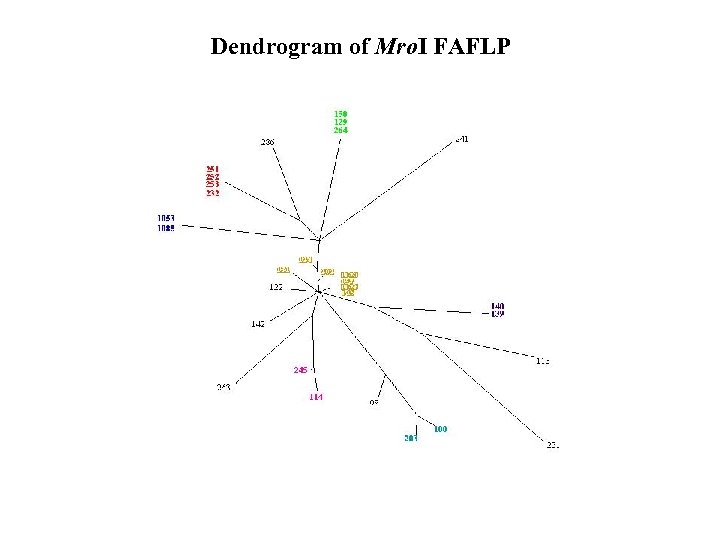
Dendrogram of Mro. I FAFLP


However, a limitation of many PCRbased approaches is the biallelic (binary) nature of their data resulting from the presence or absence of a marker fragment.
Draft Microbial Genomes The first pathogens to be sequenced under the current program are members of the Bacillus, Brucella, Clostridium, Francisella, Shigella, and Yersinia groups. In many of these groups, several strains or related species will be sequenced, for example, two strains of Bacillus anthracis (anthrax) and one of the similar species Bacillus thuringiensis.

Sequence-derived DNA typing is more rapid and has an even greater capacity for genetic dissection of bacterial pathogens. It is limited only by the genome size and the technology. Because most microbial genomes consist of millions of nucleotides, technology is invariably limiting.

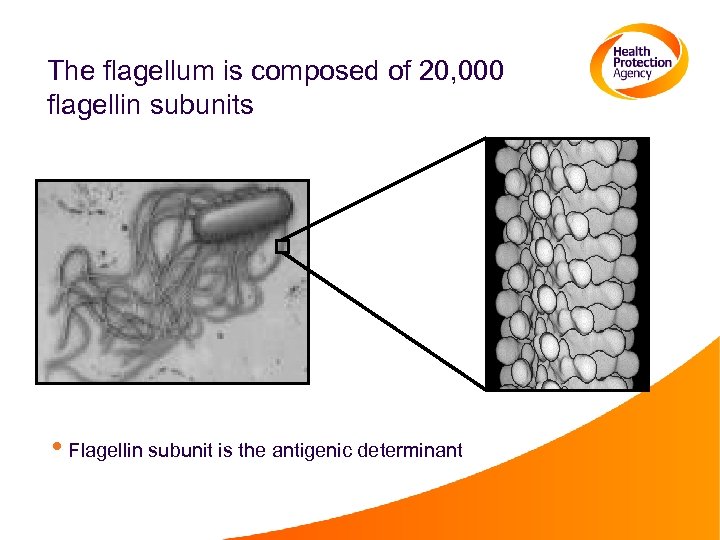
The flagellum is composed of 20, 000 flagellin subunits • Flagellin subunit is the antigenic determinant

H antigen typing of Salmonella Kauffmann-White scheme • Expression of antigens is determined by agglutination with specific antisera • In accordance with this scheme, routine clinical laboratories classify Salmonella by their particular combination of flagellar (H) and somatic (O) antigens. • O antigens (60 have been distinguished) • H 1 antigens (63 have been distinguished) • H 2 antigens, not always present (37 have been distinguished) • H-antigens are designated by letters of the alphabet (a, to z, z 1, z 2 etc. ) and by Arabic numerals.
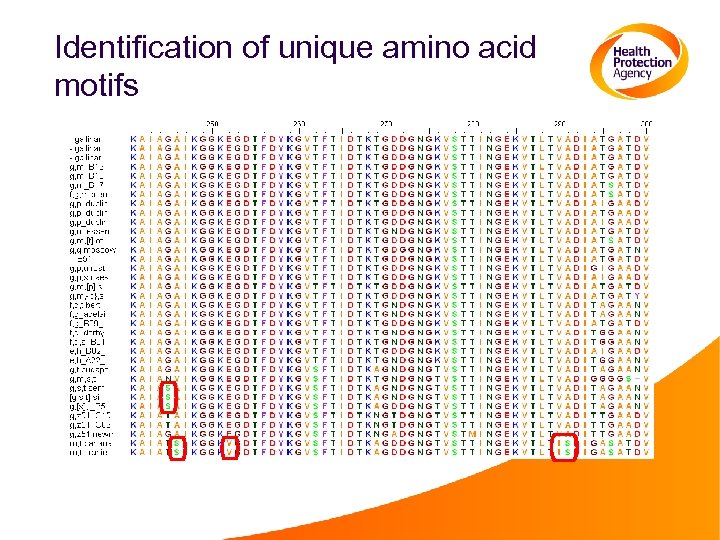

Identification of unique amino acid motifs

Sequence distances of fli. C of different serotypes g-complex sequences “non-g” sequences

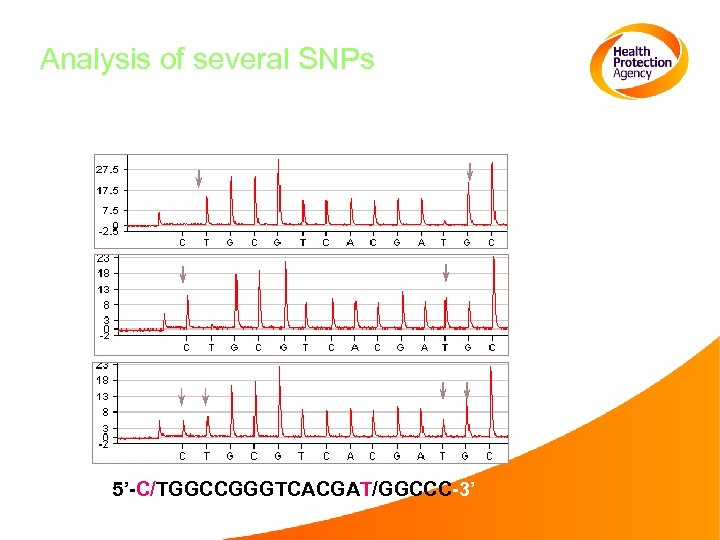
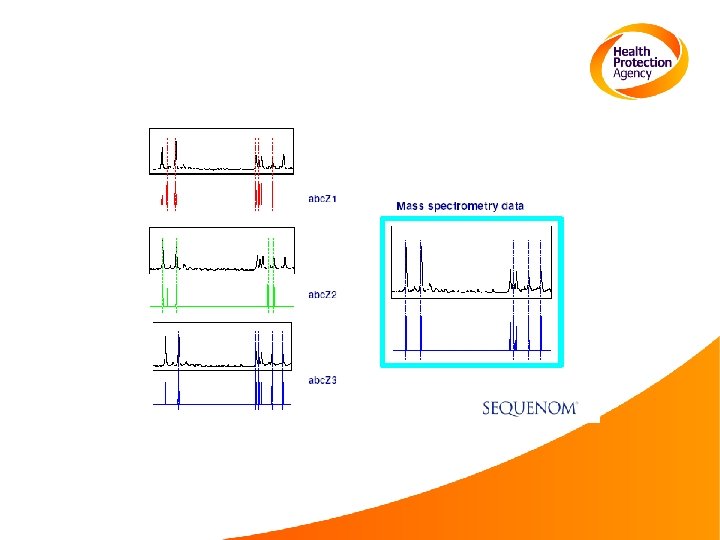
Analysis of several SNPs 5’-C/TGGCCGGGTCACGAT/GGCCC-3’
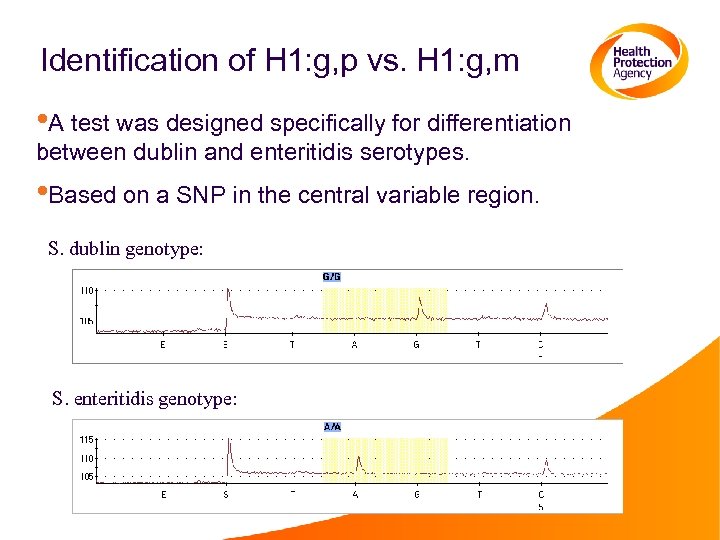

Identification of H 1: g, p vs. H 1: g, m • A test was designed specifically for differentiation between dublin and enteritidis serotypes. • Based on a SNP in the central variable region. S. dublin genotype: S. enteritidis genotype:


• One of the most recent developments in molecular analysis involves the analysis of VNTR sequences. • Short nucleotide sequences that are repeated multiple times often vary in copy number, creating length polymorphisms that can be detected by PCR using flanking primers. • Satellites: Spanning megabases • Minisatellites: Repeat units 6 -100 bp (spans 100’s bps • Microsatellites: Repeat units 1 -5 bp (spans 10’s bps)
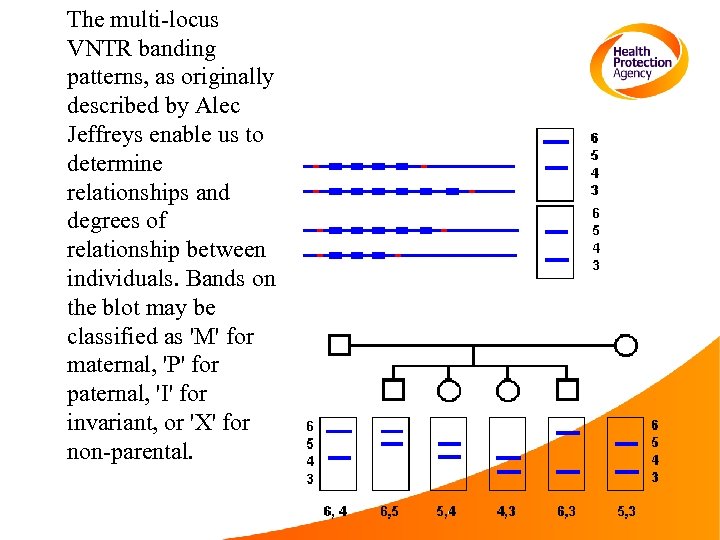
The multi-locus VNTR banding patterns, as originally described by Alec Jeffreys enable us to determine relationships and degrees of relationship between individuals. Bands on the blot may be classified as 'M' for maternal, 'P' for paternal, 'I' for invariant, or 'X' for non-parental.
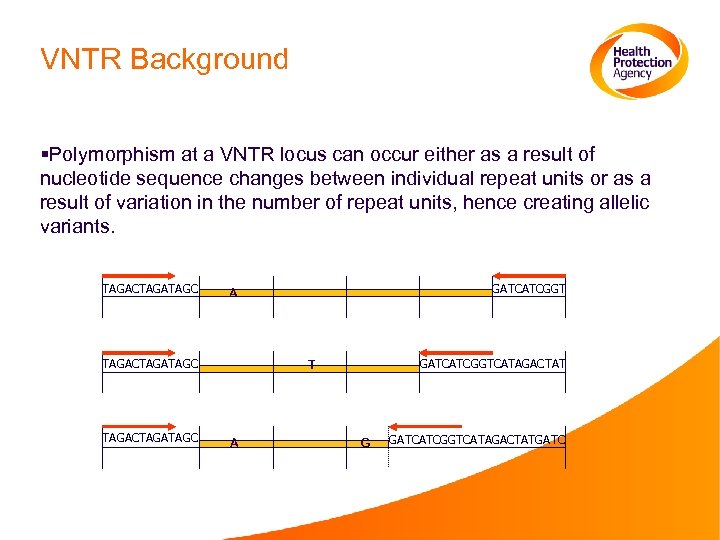
VNTR Background §Polymorphism at a VNTR locus can occur either as a result of nucleotide sequence changes between individual repeat units or as a result of variation in the number of repeat units, hence creating allelic variants. TAGACTAGATAGC GATCATCGGT A GATCATCGGTCATAGACTAT T A G GATCATCGGTCATAGACTATGATC
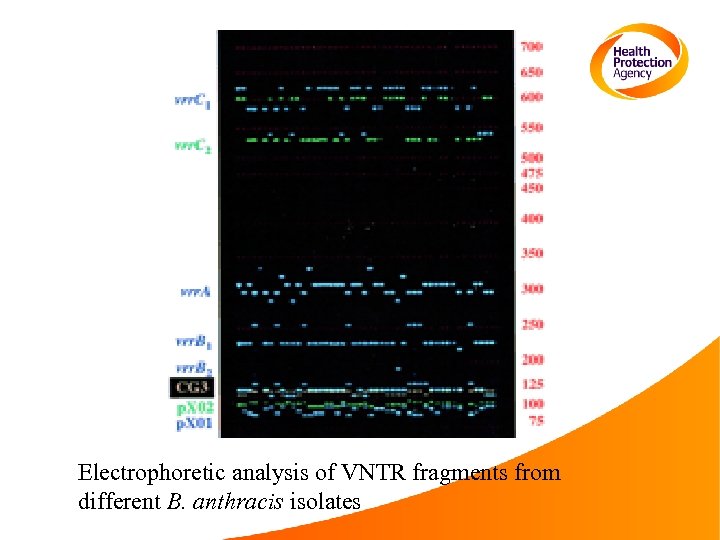
Electrophoretic analysis of VNTR fragments from different B. anthracis isolates
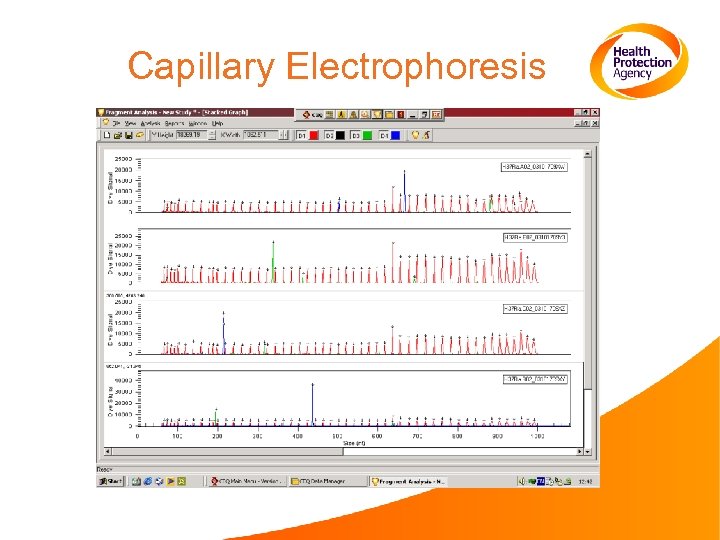
Capillary Electrophoresis
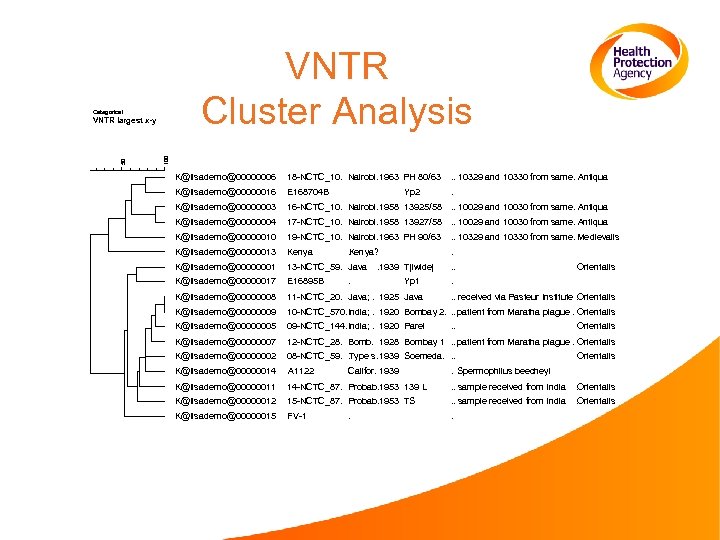
VNTR Cluster Analysis Categorical 100 50 VNTR largest x-y K@lisademo@00000006 18 -NCTC_10. Nairobi. 1963 PH 80/63 . . 10329 and 10330 from same. Antiqua . K@lisademo@00000016 E 168704 B K@lisademo@00000003 16 -NCTC_10. Nairobi. 1958 13925/58 . . 10029 and 10030 from same. Antiqua . K@lisademo@00000004 17 -NCTC_10. Nairobi. 1958 13927/58. K@lisademo@00000010 19 -NCTC_10. Nairobi. 1963 PH 90/63 . . 10329 and 10330 from same. Medievalis. K@lisademo@00000013 Kenya K@lisademo@00000001 13 -NCTC_59. . Java . 1939 Tjiwidej . . K@lisademo@00000017 E 16895 B . K@lisademo@00000008 11 -NCTC_20. . Java; . 1925 Java K@lisademo@00000009 10 -NCTC_570. India; . 1920 Bombay 2. . . patient from Maratha plague. Orientalis. K@lisademo@00000005 09 -NCTC_144. India; . 1920 Parel . K@lisademo@00000007 12 -NCTC_28. Bomb. 1928 Bombay 1 . . patient from Maratha plague. Orientalis. K@lisademo@00000002 08 -NCTC_59. . Type s. 1939 Soemeda. . . K@lisademo@00000014 A 1122 K@lisademo@00000011 14 -NCTC_87. . Probab. 1953 139 L . . sample received from India . Orientalis K@lisademo@00000012 15 -NCTC_87. . Probab. 1953 TS . . sample received from India . Orientalis K@lisademo@00000015 FV-1 . Yp 2 . Kenya? . . . 10029 and 10030 from same. Antiqua. Yp 1 . Califor. 1939 . . Orientalis . . received via Pasteur Institute. Orientalis. . Orientalis . Spermophilus beecheyi

Microarrays

Array Technology High density oligonucleotide arrays High density ‘spotted’ arrays Low density ‘line-probe’ arrays Low density addressable arrays

Pathogenesis The CDC, NAID: SARS, smallpox Disease Susceptibility “susceptibility genes” in HIV Vaccine Development to examine transcriptional activity of all genes of pathogenic microorganisms under in vivo conditions Pathogen Identification Identify pathogens using ribosomal DNA sequences Drug Response identify the presence of drug resistance genes catalog individual genetic variations in drug resistance

Imagine a world where microscopic medical implants patrol our arteries, diagnosing ailments and fighting disease; where military battle-suits deflect explosions; where computer chips are no bigger than specks of dust; and where clouds of miniature space probes transmit data from the atmospheres of Mars or Titan. Many incredible claims have been made about the future's nanotechnological applications, but what exactly does nano mean, and why has controversy plagued this emerging technology?

Nanotechnology is science and engineering at the scale of atoms and molecules. It is the manipulation and use of materials and devices so tiny that nothing can be built any smaller.
Green Emission Quantum Dot Fluorene Oligomer for Electronic Applications Water Soluble & Functional NIR Dyes

Molecular diagnostics is becoming a driving force in drug development. Applications have spread from identifying infections to include screening for cancer, hepatitis and genetic disorders Personalised Medicine The scientific community is progressing quite rapidly in developing molecular diagnostics, and industry is developing assay prototypes and conducting larger validation studies to advance this research to full clinical utility.

The risk associated with the newer technologies is that the accuracy and precision of the data generated will be compromised. Highly specific assay formats are required to detect DNA sequence variations To achieve such specificity, Careful assay design, Validation, Quality Control Standards are required
7d0e4ad1fb945b072542ab59ae755471.ppt