bc1a0be419e970dc3d97ab19c79a997a.ppt
- Количество слайдов: 21

Pathogenic and attenuated rabies viruses induces differential host protein expression in the central nervous system: Implication of neuronal dysfunction Zhen F. Fu Department of Pathology University of Georgia
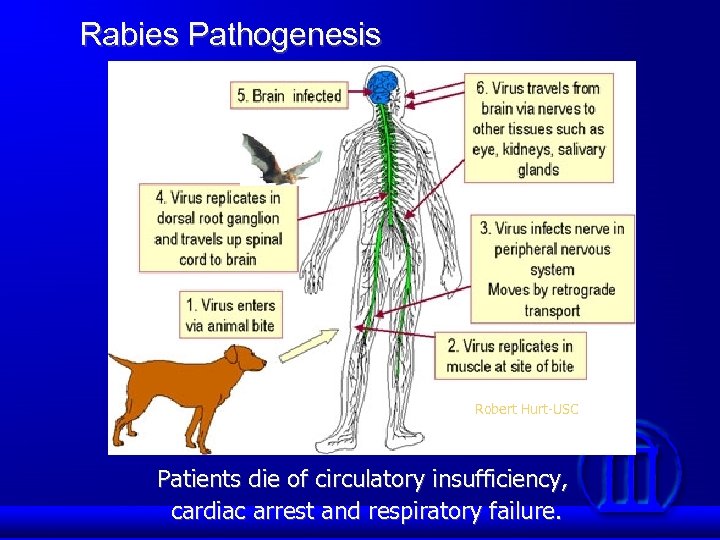
Rabies Pathogenesis Robert Hurt-USC Patients die of circulatory insufficiency, cardiac arrest and respiratory failure.

Despite extensive research in the past 100 years, we still know very little about the pathogenic mechanism by which rabies virus infection of neurons causes rabies. There are scarce neuropathology with mild inflammation and little neuronal loss, which cannot explain the lethality of the disease. It has been hypothesized that rabies results from neuronal dysfunction rather than structural damage. However, it is not known how RV infection leads to neuronal dysfunction. To better understand rabies pathogenesis, we initiate a project to determine how the host responds to rabies virus infections using one street and one fixed virus. This is accomplished by using proteomics technologies.

Two viruses were used in this study: SHBRV: Wt virus, normally circulating in silver-haired bats and responsible for most of the human rabies in the US. CVS-B 2 C: Lab-adapted attenuated virus derived from CVS-24 by passaging in BHK cells.
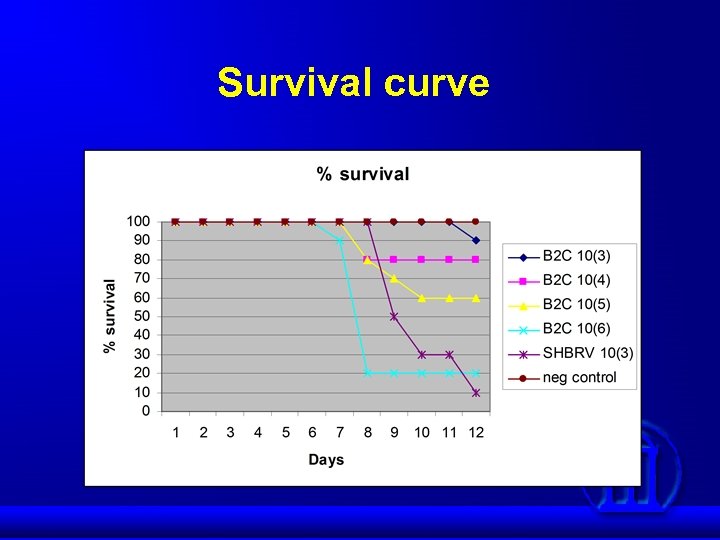
Survival curve
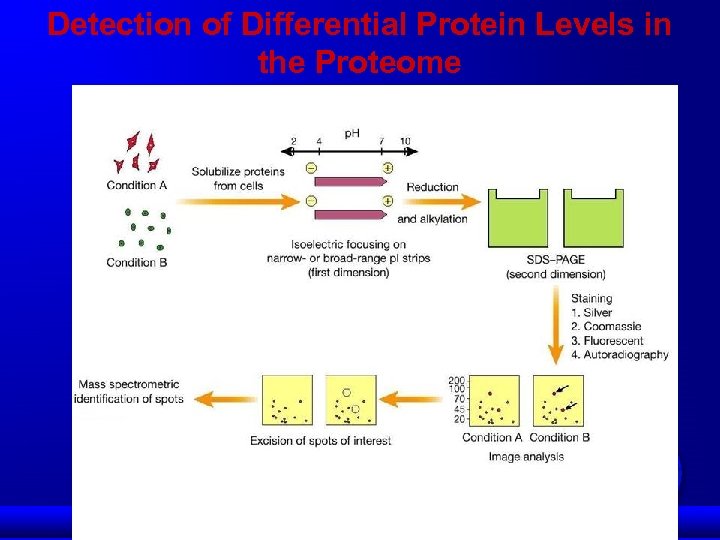
Detection of Differential Protein Levels in the Proteome
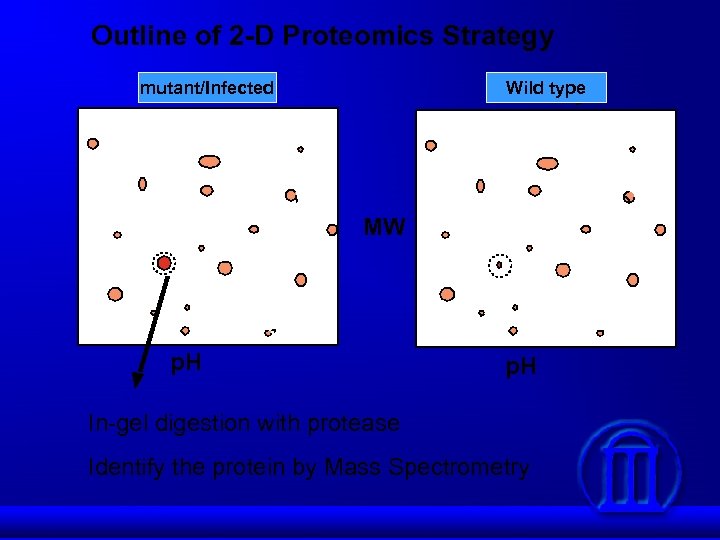
Outline of 2 -D Proteomics Strategy mutant/Infected Wild type MW p. H In-gel digestion with protease Identify the protein by Mass Spectrometry
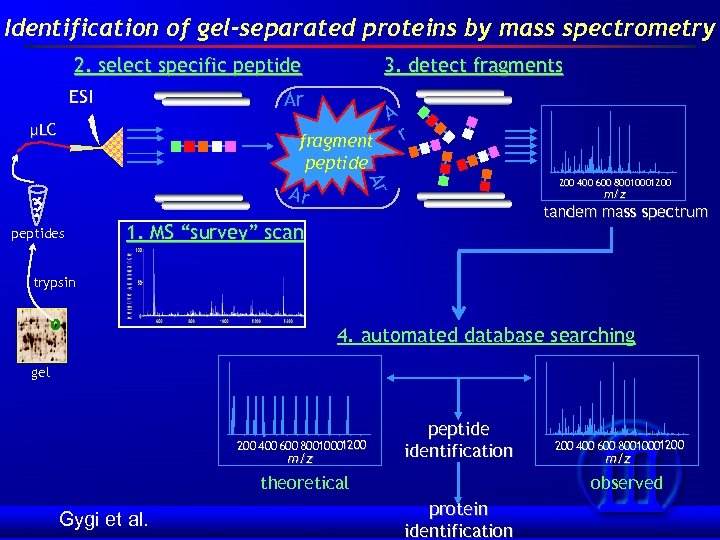
Identification of gel-separated proteins by mass spectrometry 2. select specific peptide ESI 3. detect fragments Ar µLC A fragment peptide 200 400 600 80010001200 Ar Ar peptides r m/z tandem mass spectrum 1. MS “survey” scan trypsin 4. automated database searching gel 1200 400 600 80010001200 m/z peptide identification theoretical Gygi et al. 1200 400 600 80010001200 m/z observed protein identification

The situation to avoid… Software is essential !
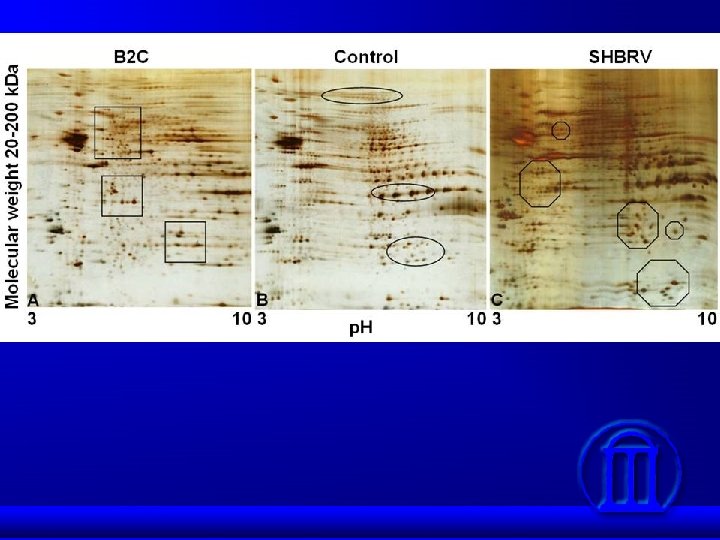
Proteins differentially expressed in response to SHBRV infection Category Protein PI Molecular weight (k. Da) Ion Homeostasis Na+/K+ ATPase 5. 3 110 ↑ +3. 0 H+ ATPase subunit a isoform 1 6. 0 96 ↑ +2. 6 TRIM 9 6. 6 90 ↓ -3. 0 -SNAP 5. 3 33 ↓ -3. 0 Pallidin (Syntaxin 13) 5. 8 20 ↓ -2. 2 Synexin 5. 9 50 ↓ -2. 0 Syntaxin 18 5. 2 35 ↓ -1. 8 Synaptic Physiology Up/downregulation Fold change
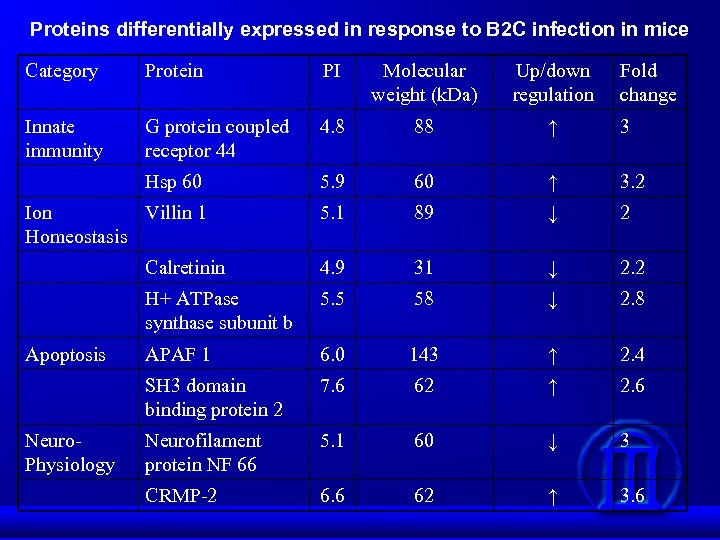
Proteins differentially expressed in response to B 2 C infection in mice Category Protein PI Molecular weight (k. Da) Up/down regulation Innate immunity G protein coupled receptor 44 4. 8 88 ↑ 3 Hsp 60 5. 9 60 ↑ 3. 2 Ion Villin 1 Homeostasis 5. 1 89 ↓ 2 Calretinin 4. 9 31 ↓ 2. 2 H+ ATPase synthase subunit b 5. 5 58 ↓ 2. 8 APAF 1 6. 0 143 ↑ 2. 4 SH 3 domain binding protein 2 7. 6 62 ↑ 2. 6 Neurofilament protein NF 66 5. 1 60 ↓ 3 CRMP-2 6. 6 62 ↑ 3. 6 Apoptosis Neuro. Physiology Fold change
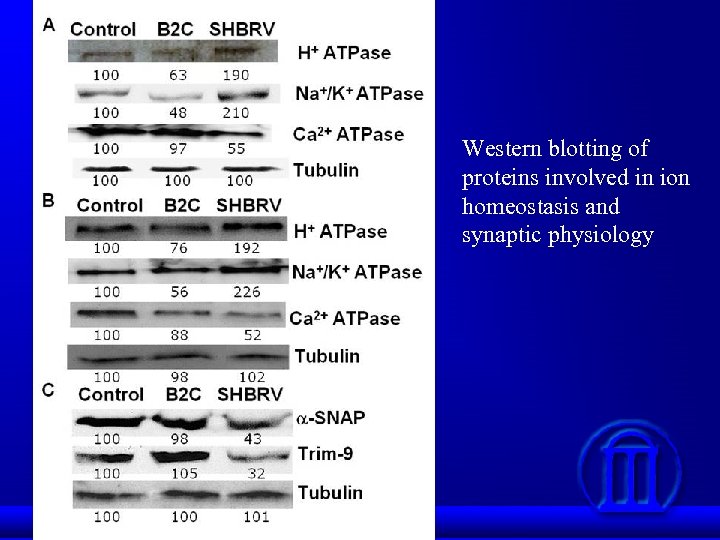
Western blotting of proteins involved in ion homeostasis and synaptic physiology
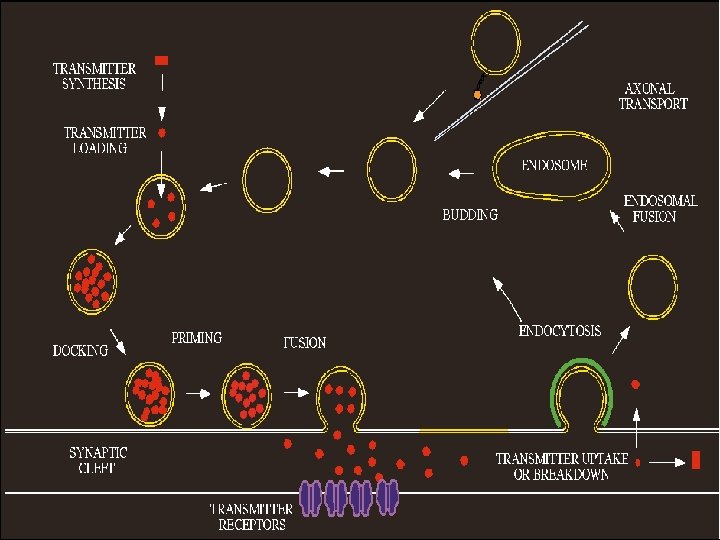
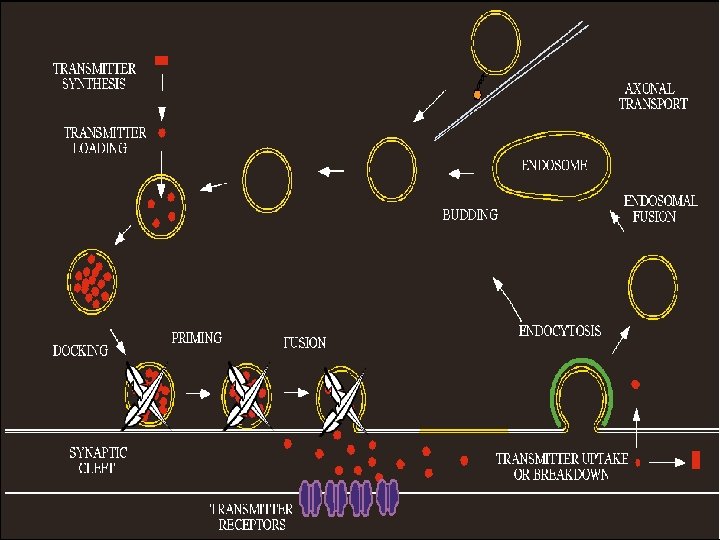
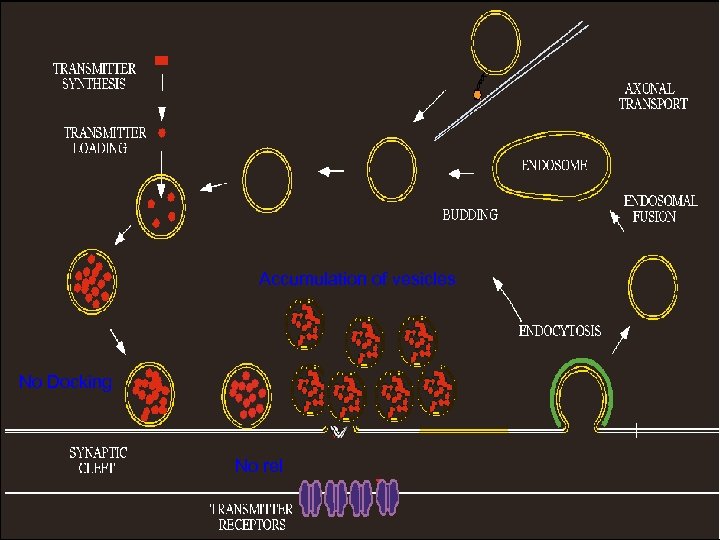
Accumulation of vesicles No Docking No release of Neurotransmitters
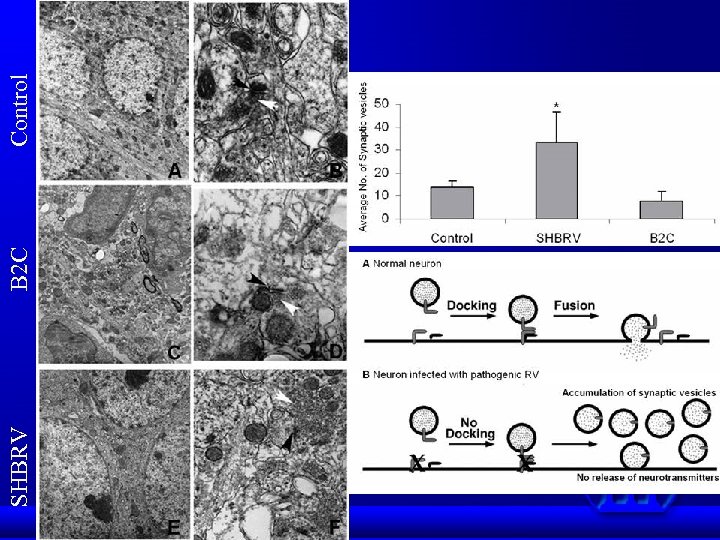
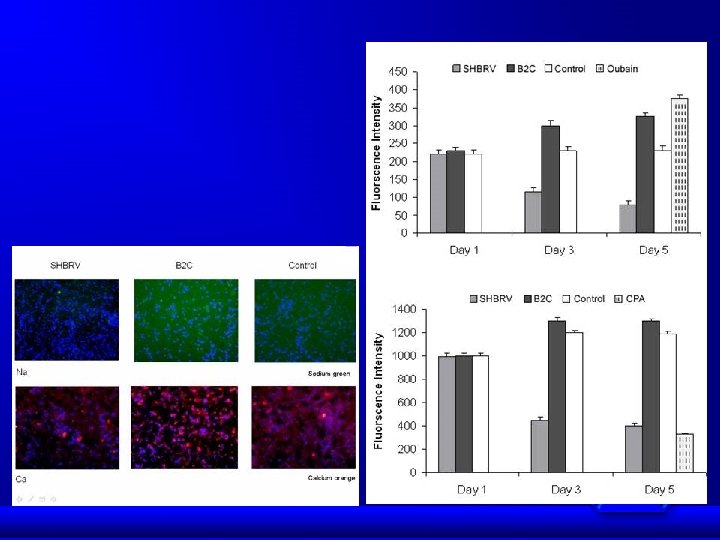
SHBRV B 2 C Control

Conclusions Proteomics data indicate that wt RV infection resulted in up-regulation of proteins involved in ion homeostasis and down-regulation of synaptic proteins. The altered protein expression as detected by 2 D-gel analysis is confirmed by Western blotting in animals infected either ic or im as well as in primary neuron. Up-regulation of Na/K-ATPase leads to decrease in Na+ concentrations in infected cells. Likewise, down-regulation of Ca-ATPase resulted in decrease of Ca++ concentration in infected cells. Changes in Na/Ca concentration affects membrane potential and thus leading to alteration of neuronal transmission. Synaptic proteins such as syntaxin, -SANP, and TRIM 9 play important roles in synaptic-vesicle fusion and docking of synaptic vesicles. Down-regulation of these proteins prevented the docking and fusion of synaptic vesicles with presynaptic membrane, thus resulting in accumulation of synaptic vesicles in the presynapses. Thus our data may provide structural and metabolic basis by which RV infection causes neuronal dysfunction.

Acknowledgements Vikas Dhingra Xia-qing Li Luciana Sarmento UGA Proteomics Facility Tracy Andachtc

Thank you!!
bc1a0be419e970dc3d97ab19c79a997a.ppt