86aa9df2c38eab1a2872f6a309769ef0.ppt
- Количество слайдов: 54

Part III Systems Biology DAH 3. 1 Mass Spectrometry Kathryn Lilley Cambridge Centre for Proteomics Department of Biochemistry University of Cambridge k. s. lilley@bioc. cam. ac. uk www. bio. cam. ac. uk/proteomics/

C a m b r i d g e C e n t r e f o r P r o t e o m i c s Definition of the Proteome Why bother studying it? ? The analysis of the entire PROTEin complement expressed by a gen. OME. Wasinger et al Electrophoresis 16 (1995) Could be: Cellular extract Secreted fluid Tissue Whole organism

A Proteomicist’s Tools • Mass spectrometry • Protein and peptide separation methods • Databases and software • Validation tools – Western blotting – GFP tagging and confocal microscopy

Instruments for mass analysis Mass Spectrometers measure m/z of gaseous ion Mass spectrometers comprise: A source which is responsible for ionising the sample, e. g. electrospray, laser desorption An analyser which separates and carries the ions to the detector e. g. Quadrupole, Ion-trap (mass range 2000 -4000 m/z) TOF (time of flight) (mass range 0 -200, 000+ m/z) A detector e. g. electron multiplier

Outline • Proteomics workflows • Protein identification • Post translational modification

Protein analysis on pure proteins/complexes Mass of protein Modification status Low through put methods Usually carried out on pure proteins or complexes can be difficult to deconvolute with many isoforms Already know what the protein is Higher order structures ability to spray whole complexes and look at components and stoichiometries

Protein analysis on complex mixtures For more complex samples you cannot purify each one and then analyse it. Methods need to be applied where proteins can be analysed simultaneously Proteins can be separated then analysed or converted to peptides which are then analysed The peptides act as surrogates for the protein
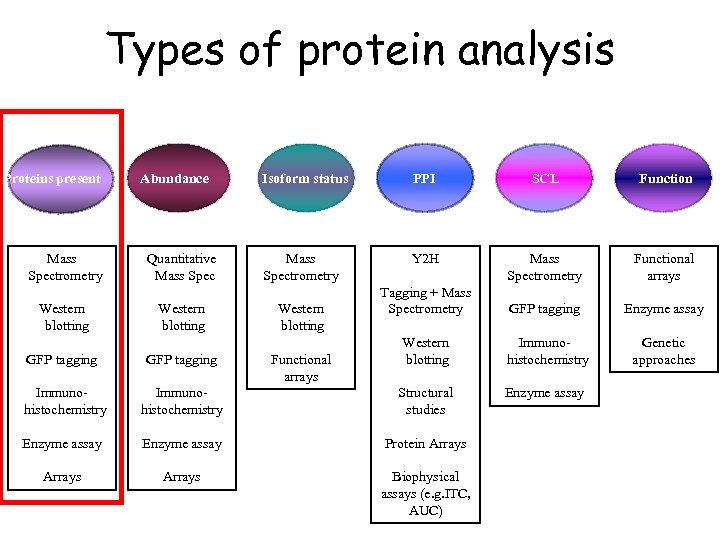
Types of protein analysis Proteins present Mass Spectrometry Western blotting Abundance Quantitative Mass Spec Western blotting Isoform status PPI SCL Function Mass Spectrometry Y 2 H Mass Spectrometry Functional arrays Tagging + Mass Spectrometry GFP tagging Enzyme assay Western blotting Immunohistochemistry Genetic approaches Enzyme assay Western blotting GFP tagging Functional arrays Immunohistochemistry Structural studies Enzyme assay Protein Arrays Biophysical assays (e. g. ITC, AUC)
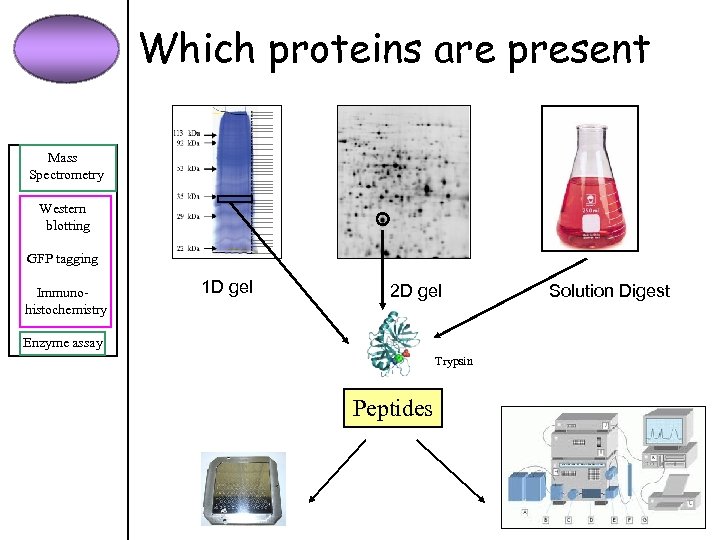
Which proteins are present Mass Spectrometry Western blotting GFP tagging Immunohistochemistry 1 D gel 2 D gel Enzyme assay Trypsin Peptides Solution Digest
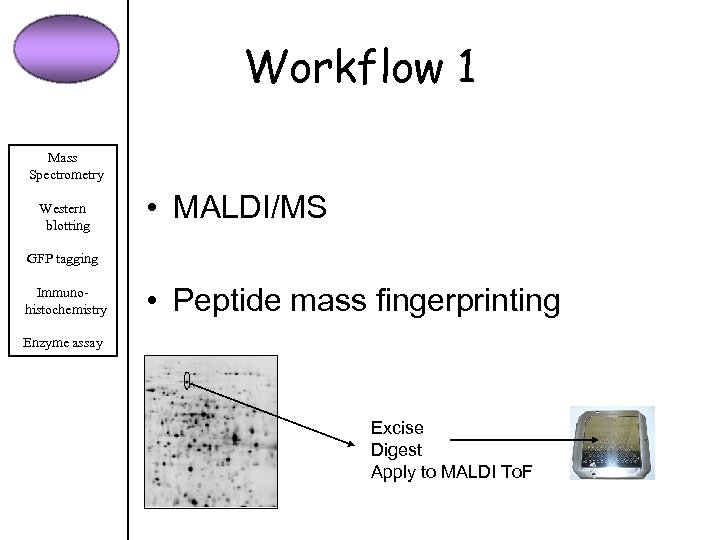
Workflow 1 Mass Spectrometry Western blotting • MALDI/MS GFP tagging Immunohistochemistry • Peptide mass fingerprinting Enzyme assay Excise Digest Apply to MALDI To. F
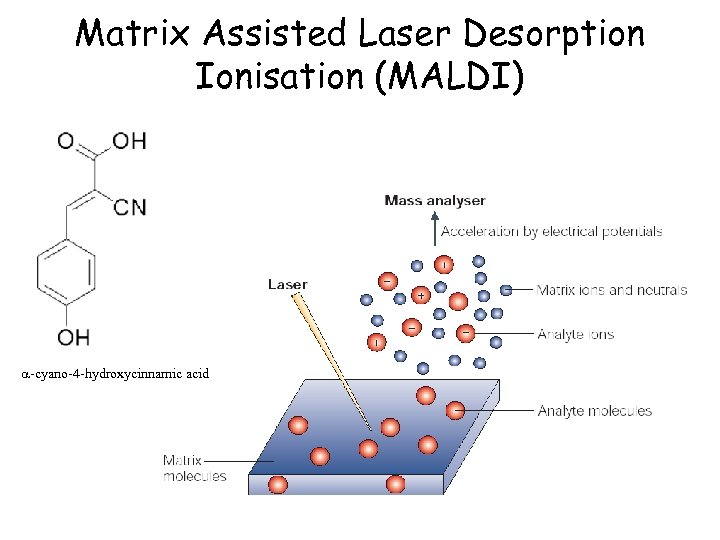
Matrix Assisted Laser Desorption Ionisation (MALDI) a-cyano-4 -hydroxycinnamic acid
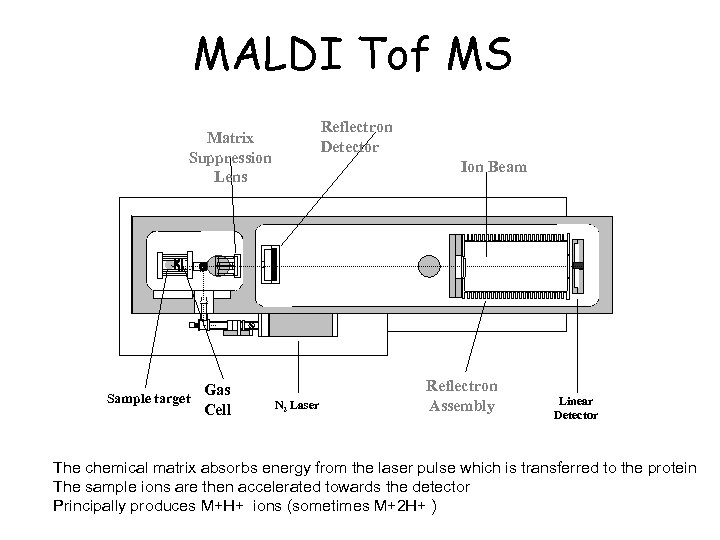
MALDI Tof MS Reflectron Detector Matrix Suppression Lens Sample target Gas Cell Ion Beam N 2 Laser Reflectron Assembly Linear Detector The chemical matrix absorbs energy from the laser pulse which is transferred to the protein The sample ions are then accelerated towards the detector Principally produces M+H+ ions (sometimes M+2 H+ )
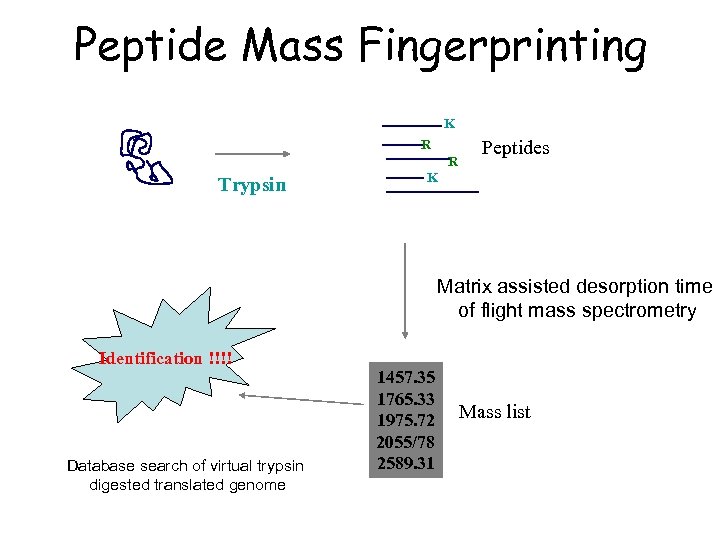
Peptide Mass Fingerprinting K R R Trypsin Peptides K Matrix assisted desorption time of flight mass spectrometry Identification !!!! Database search of virtual trypsin digested translated genome 1457. 35 1765. 33 1975. 72 2055/78 2589. 31 Mass list
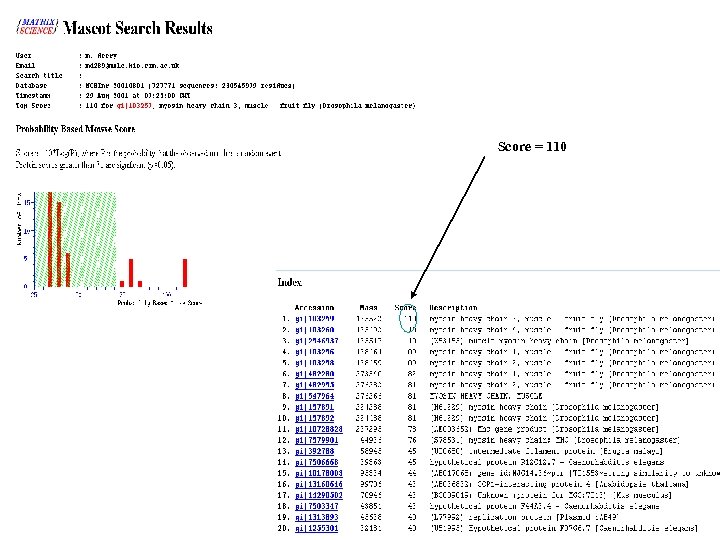
Score = 110

Strengths Quick Cheap Limitations Only works well for purified proteins Require well annotated genome
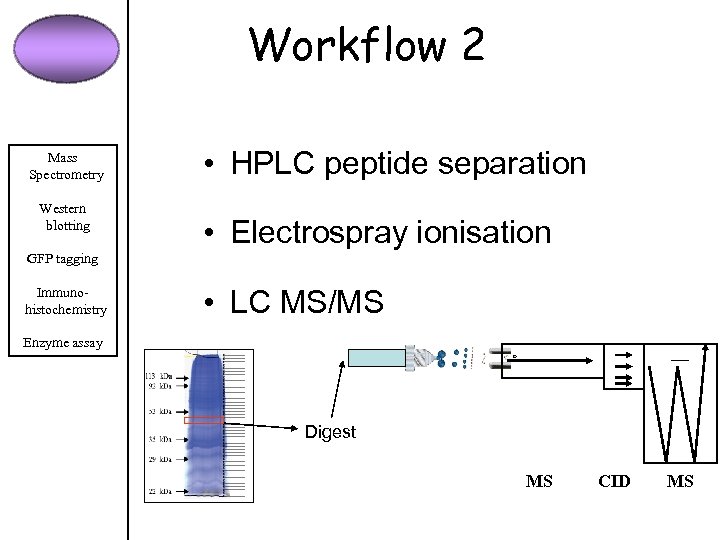
Workflow 2 Mass Spectrometry Western blotting • HPLC peptide separation • Electrospray ionisation GFP tagging Immunohistochemistry • LC MS/MS Enzyme assay Digest MS CID MS
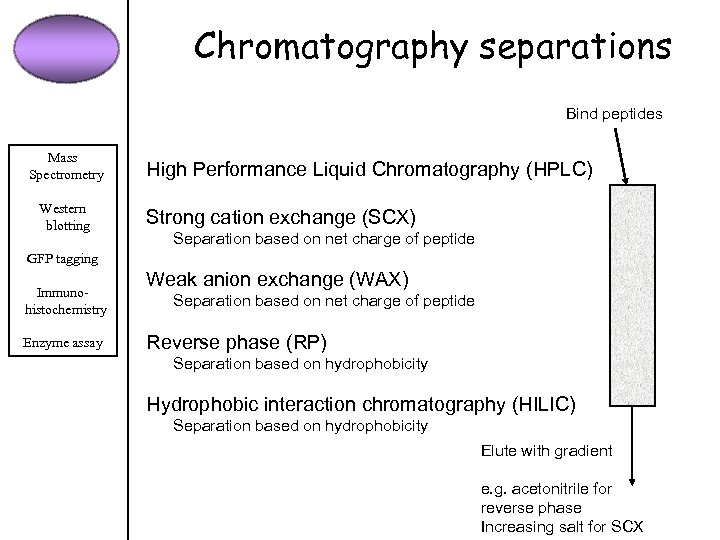
Chromatography separations Bind peptides Mass Spectrometry Western blotting High Performance Liquid Chromatography (HPLC) Strong cation exchange (SCX) Separation based on net charge of peptide GFP tagging Immunohistochemistry Enzyme assay Weak anion exchange (WAX) Separation based on net charge of peptide Reverse phase (RP) Separation based on hydrophobicity Hydrophobic interaction chromatography (HILIC) Separation based on hydrophobicity Elute with gradient e. g. acetonitrile for reverse phase Increasing salt for SCX
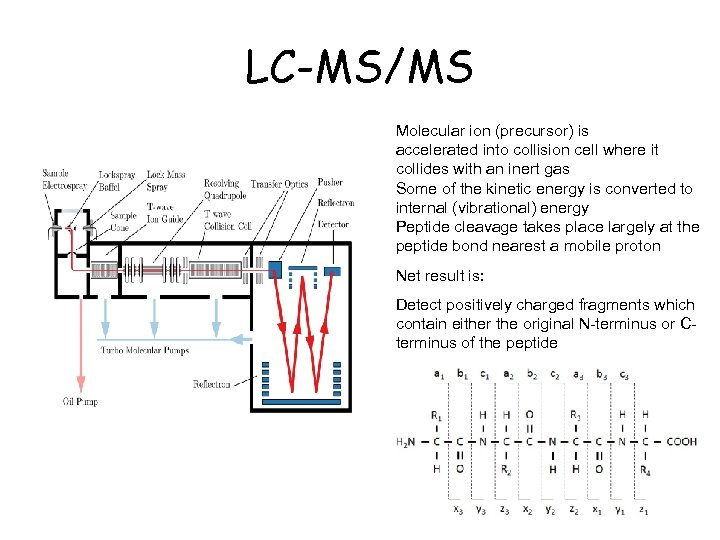
LC-MS/MS Molecular ion (precursor) is accelerated into collision cell where it collides with an inert gas Some of the kinetic energy is converted to internal (vibrational) energy Peptide cleavage takes place largely at the peptide bond nearest a mobile proton Net result is: Detect positively charged fragments which contain either the original N-terminus or Cterminus of the peptide
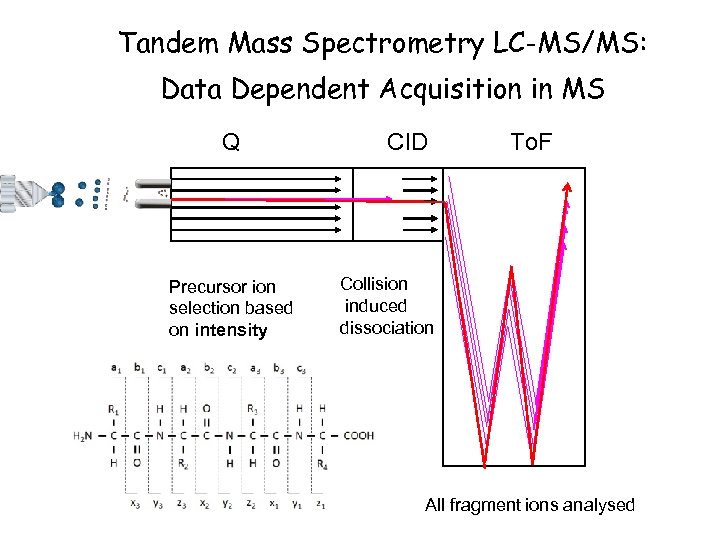
Tandem Mass Spectrometry LC-MS/MS: Data Dependent Acquisition in MS Q Precursor ion selection based on intensity CID To. F Collision induced dissociation Precursors scanned out of first quad. All fragment ions analysed

Typical output • List of peptide masses – Precursor mass (parent ion mass) • Fragment ion masses – y-ions – b-ions

Protein identification • Search engines MASCOT - http: //www. matrixscience. com SEQUEST - fields. scripps. edu/sequest/ X ! Tandem -www. thegpm. org/tandem/index. html Phenyx- www. genebio. com/products/phenyx/
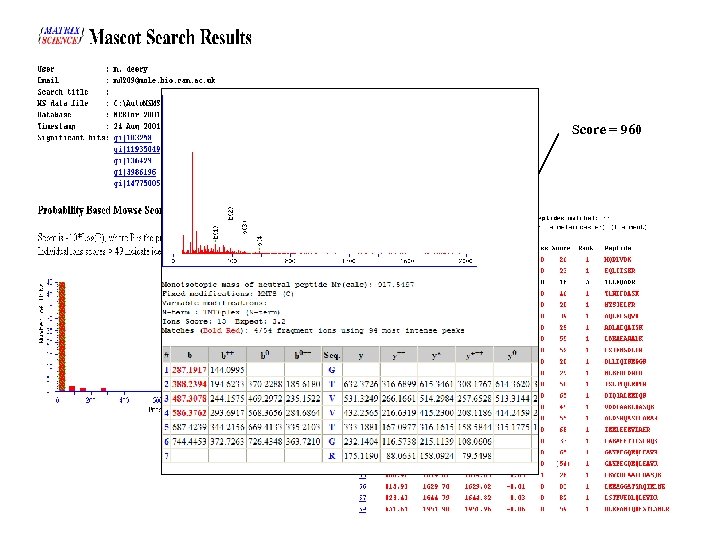
Score = 960
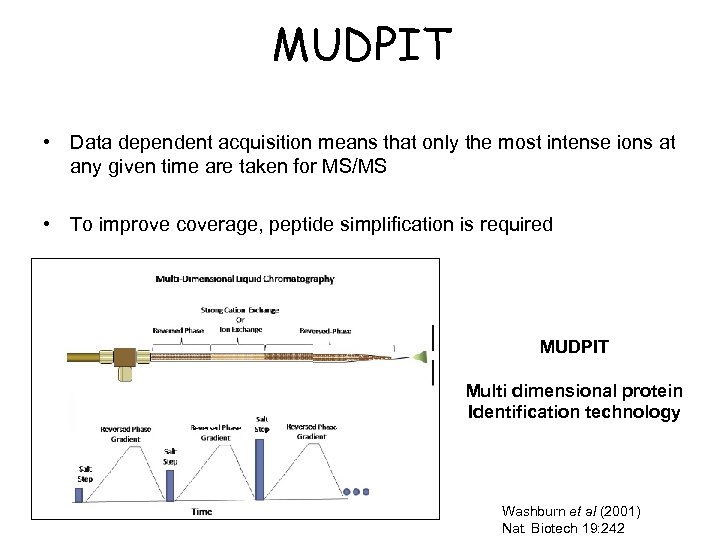
MUDPIT • Data dependent acquisition means that only the most intense ions at any given time are taken for MS/MS • To improve coverage, peptide simplification is required MUDPIT Multi dimensional protein Identification technology Washburn et al (2001) Nat. Biotech 19: 242

Strengths and weaknesses • Can be used with very complex mixtures of proteins • If the genome is not sequenced then sequence returned may show similarity or identity to related organisms • De novo sequencing • More time consuming • Equipment more expensive
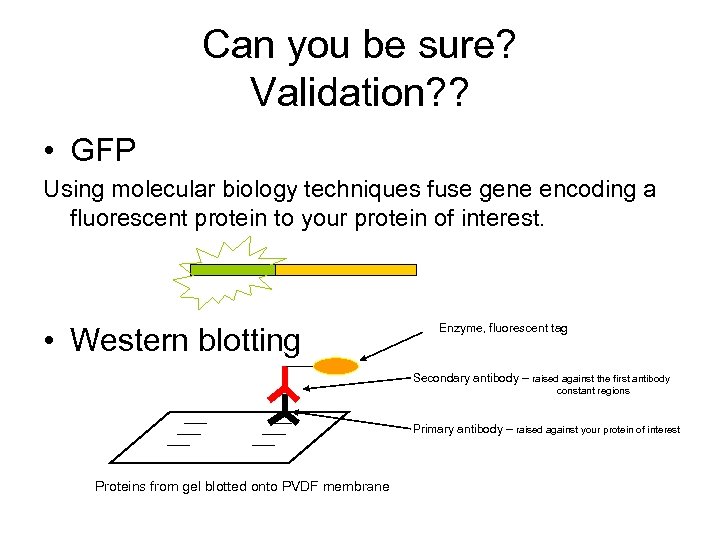

Can you be sure? Validation? ? • GFP Using molecular biology techniques fuse gene encoding a fluorescent protein to your protein of interest. • Western blotting Enzyme, fluorescent tag Secondary antibody – raised against the first antibody constant regions Primary antibody – raised against your protein of interest Proteins from gel blotted onto PVDF membrane
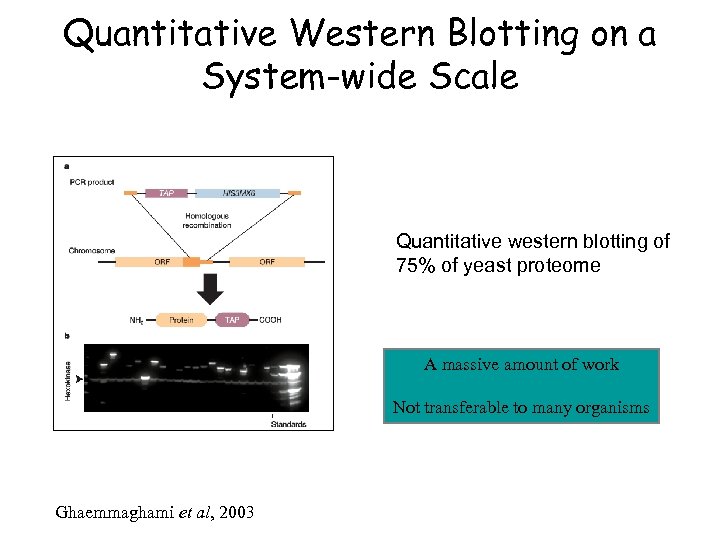
Quantitative Western Blotting on a System-wide Scale Quantitative western blotting of 75% of yeast proteome A massive amount of work Not transferable to many organisms Ghaemmaghami et al, 2003
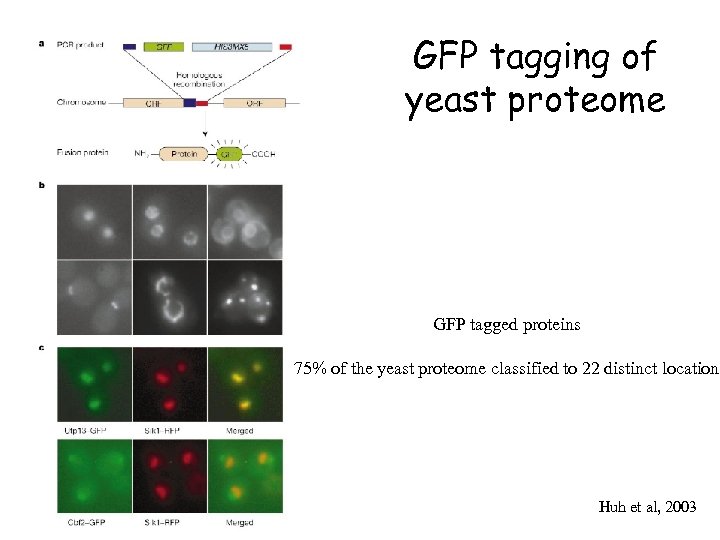
GFP tagging of yeast proteome GFP tagged proteins 75% of the yeast proteome classified to 22 distinct location Huh et al, 2003
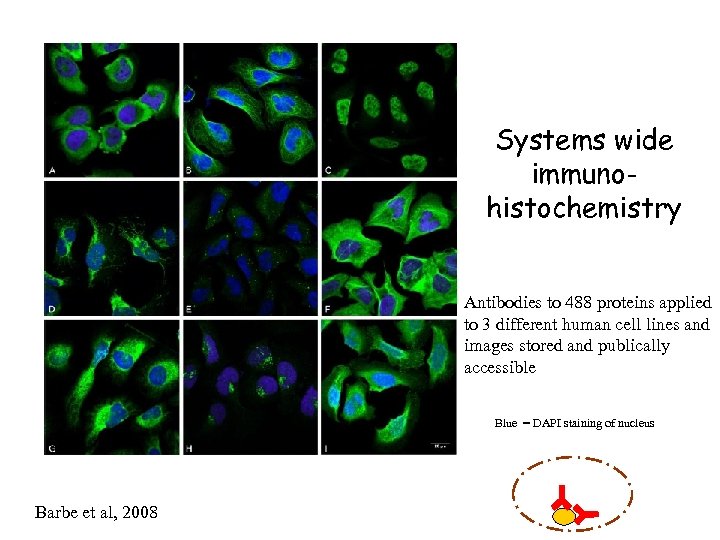
Systems wide immunohistochemistry Antibodies to 488 proteins applied to 3 different human cell lines and images stored and publically accessible Blue = DAPI staining of nucleus Barbe et al, 2008
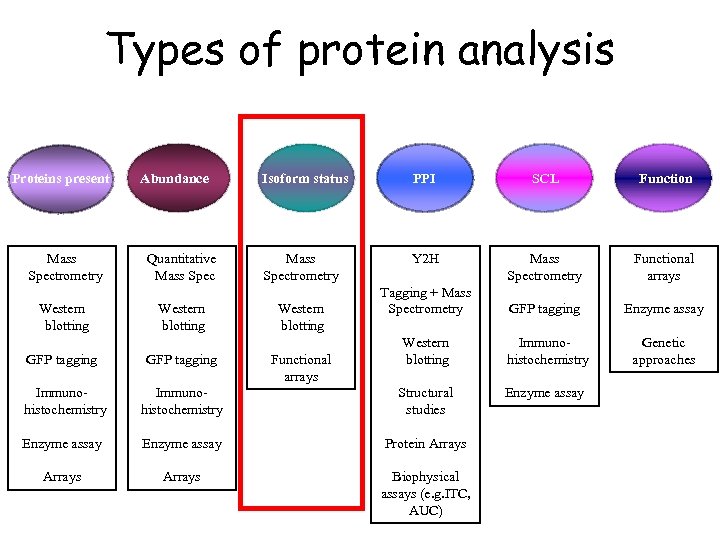
Types of protein analysis Proteins present Mass Spectrometry Western blotting Abundance Quantitative Mass Spec Western blotting Isoform status PPI SCL Function Mass Spectrometry Y 2 H Mass Spectrometry Functional arrays Tagging + Mass Spectrometry GFP tagging Enzyme assay Western blotting Immunohistochemistry Genetic approaches Enzyme assay Western blotting GFP tagging Functional arrays Immunohistochemistry Structural studies Enzyme assay Protein Arrays Biophysical assays (e. g. ITC, AUC)
Isoform status Mass Spectrometry Protein Isoform Analysis Proteins may be: Western blotting Functional arrays – Covalently modified – Truncated – Dimerised
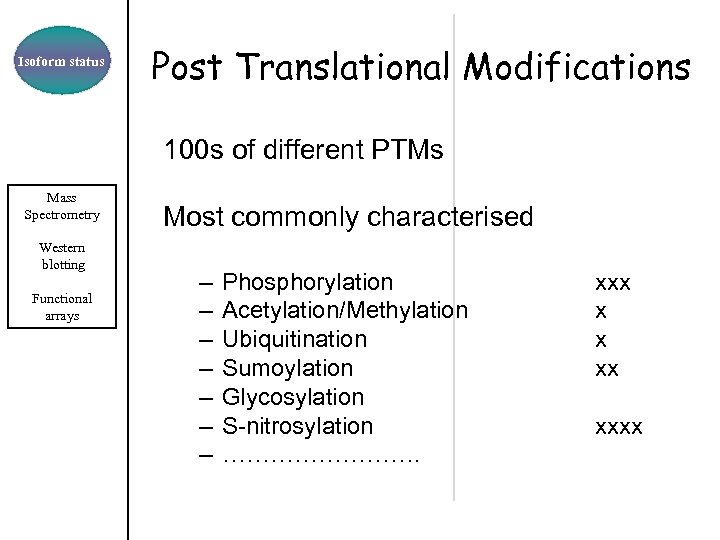

Isoform status Post Translational Modifications 100 s of different PTMs Mass Spectrometry Western blotting Functional arrays Most commonly characterised – – – – Phosphorylation Acetylation/Methylation Ubiquitination Sumoylation Glycosylation S-nitrosylation …………. xxx x x xx xxxx
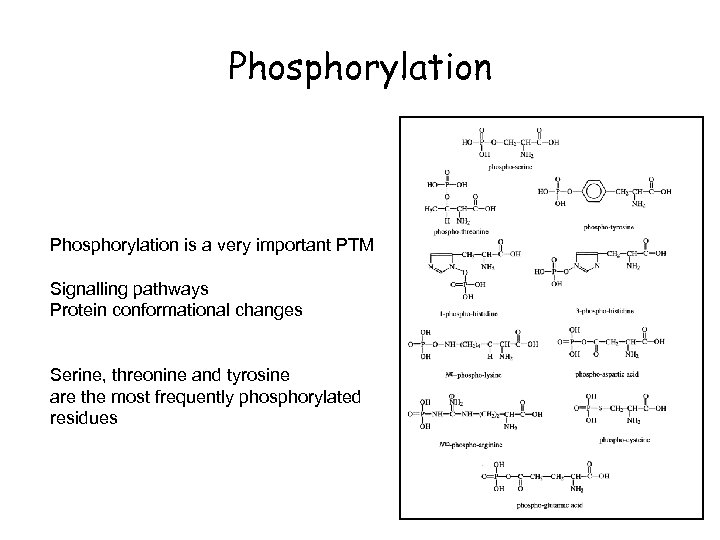
Phosphorylation is a very important PTM Signalling pathways Protein conformational changes Serine, threonine and tyrosine are the most frequently phosphorylated residues
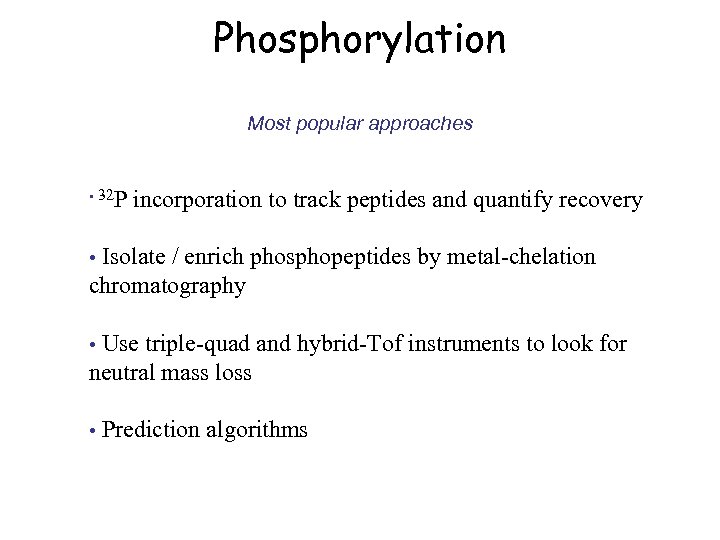

Phosphorylation Most popular approaches • 32 P incorporation to track peptides and quantify recovery Isolate / enrich phosphopeptides by metal-chelation chromatography • Use triple-quad and hybrid-Tof instruments to look for neutral mass loss • • Prediction algorithms

Problems with Phosphoproteomics • Phospho groups are highly dynamic • Phospho tyrosine is very rare • Phosphopeptides ionise poorly, they tend to be very acidic • The phosphate group tends to fall off p. Ser and p. Thr during MS/MS

Phospho-protein and -peptide enrichment • Phospho-tyrosine – good antibodies • Phospho-serine and phospho-threonine – Metal chelate chromatography – Ion exchange chromatography
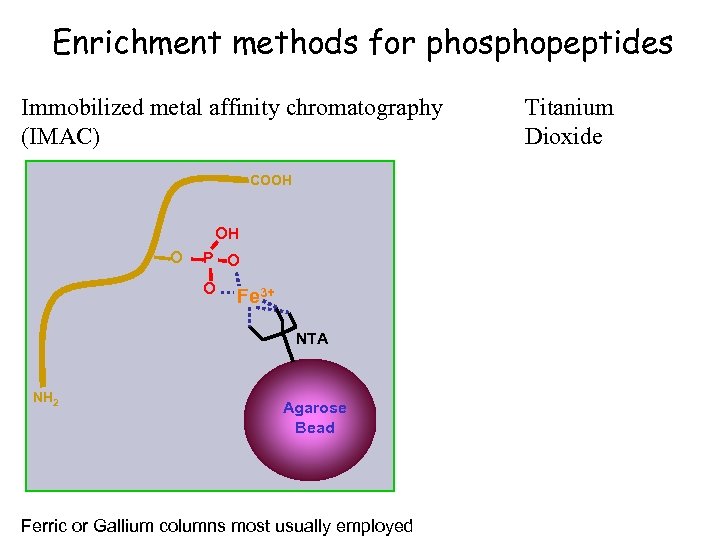
Enrichment methods for phosphopeptides Immobilized metal affinity chromatography (IMAC) COOH OH O P O O Fe 3+ NTA NH 2 Agarose Bead Ferric or Gallium columns most usually employed Titanium Dioxide
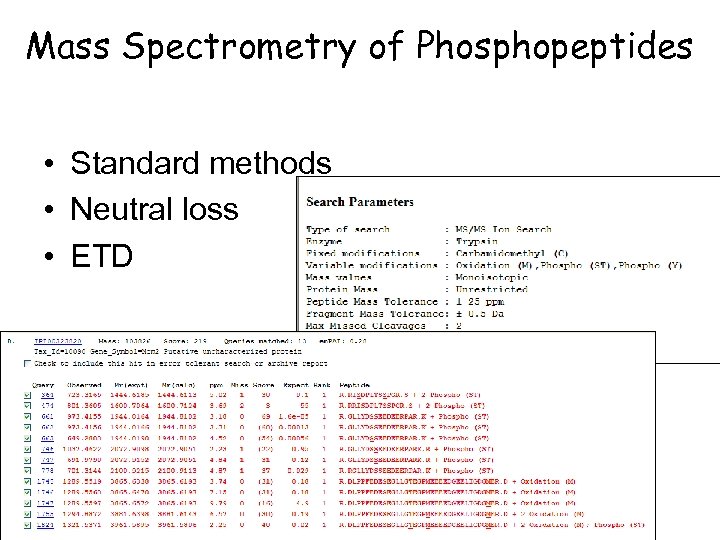

Mass Spectrometry of Phosphopeptides • Standard methods • Neutral loss • ETD
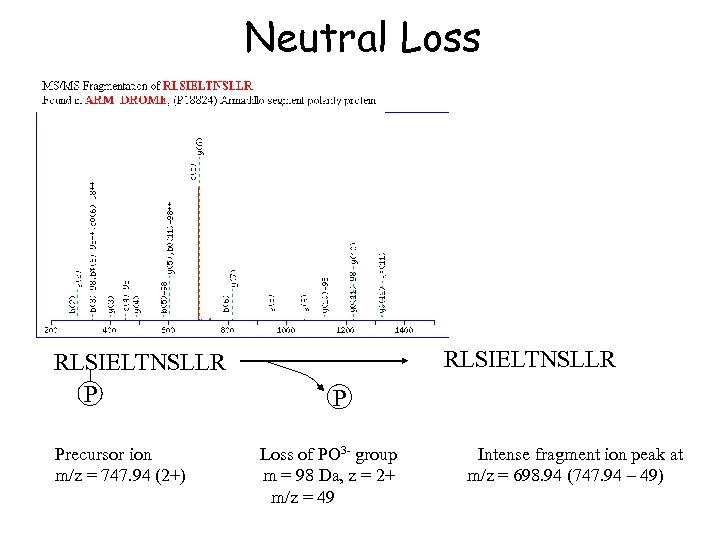
Neutral Loss RLSIELTNSLLR P Precursor ion m/z = 747. 94 (2+) RLSIELTNSLLR P Loss of PO 3 - group m = 98 Da, z = 2+ m/z = 49 Intense fragment ion peak at m/z = 698. 94 (747. 94 – 49)

Electron Transfer Dissociation (ETD). § Gentler fragmentation than CID § Preserves post-translational modifications, such as phosphorylation § Produces c and z ions § Better sequence coverage than CID
![Electron Transfer Dissociation (ETD) [M + 3 H]3+ + A[M + 3 H]2+ • Electron Transfer Dissociation (ETD) [M + 3 H]3+ + A[M + 3 H]2+ •](https://present5.com/presentation/86aa9df2c38eab1a2872f6a309769ef0/image-40.jpg)
Electron Transfer Dissociation (ETD) [M + 3 H]3+ + A[M + 3 H]2+ • + A [C+2 H]1+ + [Z+H]1+ • C Z R Fluoranthene Radical Anion (Good Electron Donor/ETD Reagent) >1 e. V Electron “Thermal” e- R + -
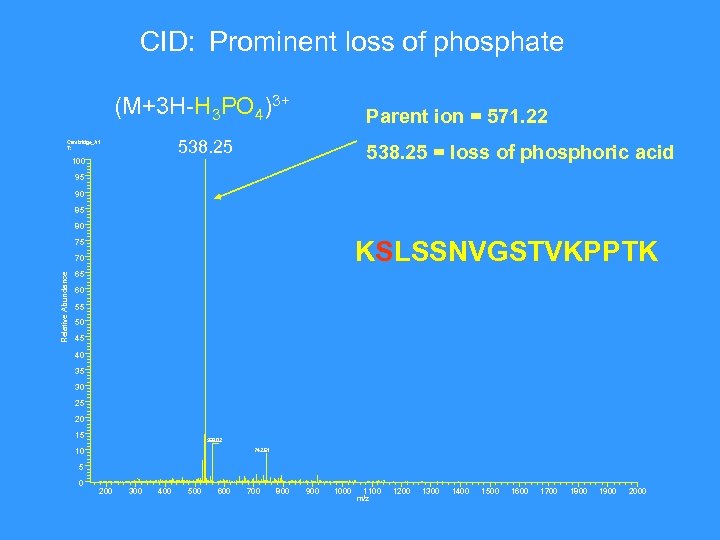
CID: Prominent loss of phosphate (M+3 H-H 3 PO 4)3+ Parent ion = 571. 22 538. 25 Cambridge_A 1 T: 100 538. 25 = loss of phosphoric acid 95 90 85 80 KSLSSNVGSTVKPPTK 75 Relative Abundance 70 65 60 55 50 45 40 35 30 25 20 15 559. 02 10 742. 61 5 0 200 300 400 500 600 700 800 900 1000 1100 m/z 1200 1300 1400 1500 1600 1700 1800 1900 2000
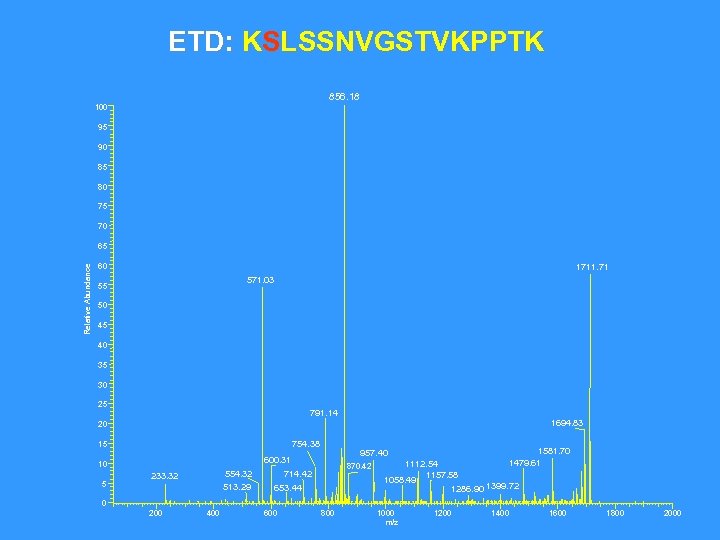
ETD: KSLSSNVGSTVKPPTK 856. 18 100 95 90 85 80 75 70 Relative Abundance 65 60 1711. 71 571. 03 55 50 45 40 35 30 25 791. 14 1694. 83 20 754. 38 15 5 554. 32 513. 29 233. 32 1581. 70 1479. 61 1112. 54 1157. 58 1058. 49 1286. 90 1399. 72 957. 40 600. 31 10 870. 42 714. 42 653. 44 0 200 400 600 800 1000 m/z 1200 1400 1600 1800 2000

Methylation/Acetylation More straightforward, but will be issues with digestion rates if trypsin is used Can enrich, antibody affinity capture to acetyl-lysine Pang et al (2010) Identification of arginine- and lysinemethylation in the proteome of Saccharomyces cerevisiae and its functional implications BMC Genomics 2010, 11: 92 Choudhary, et al. (2009) Lysine acetylation targets protein complexes and co-regulates major cellular functions, Science 325, 834 -840.
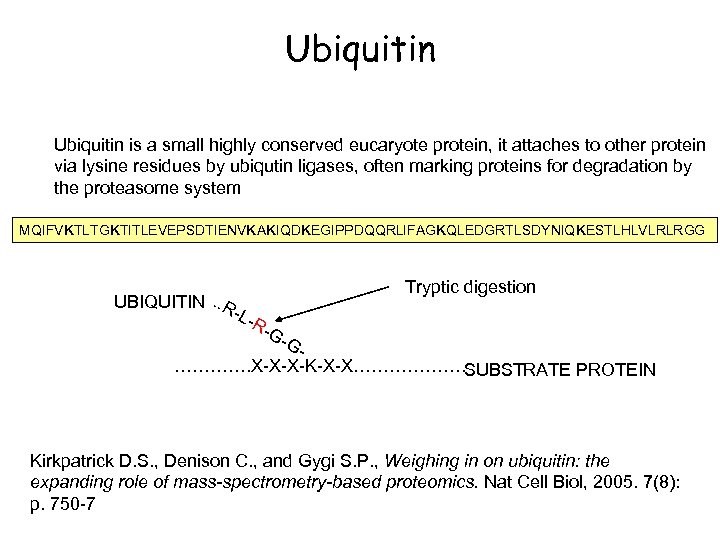
Ubiquitin is a small highly conserved eucaryote protein, it attaches to other protein via lysine residues by ubiqutin ligases, often marking proteins for degradation by the proteasome system MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGG Tryptic digestion UBIQUITIN. . R -L- R- G- G…………. X-X-X-K-X-X………………. SUBSTRATE PROTEIN Kirkpatrick D. S. , Denison C. , and Gygi S. P. , Weighing in on ubiquitin: the expanding role of mass-spectrometry-based proteomics. Nat Cell Biol, 2005. 7(8): p. 750 -7
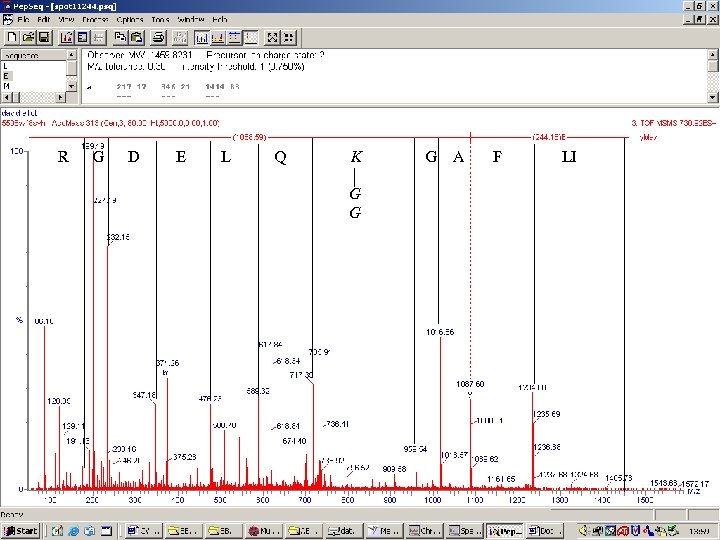
R G D E L Q K G G G A F LI

SUMO Small Ubiquitin-like Modifier 3 (4) versions Many functions including stability, nuclear-cytosolic transport, and transcriptional regulation Sadly there is no well placed tryptic site or site for any other common protease near the point at which it attaches to its modification target MSMS spectra are thus a mess, as two sets of –b and –y ions will be produced per with SUMO modification Automated identification of SUMOylation sites using mass spectrometry and SUMm. On pattern recognition software. Pedrioli PG, et al. Nat Methods. 2006 3(7): 533 -9.
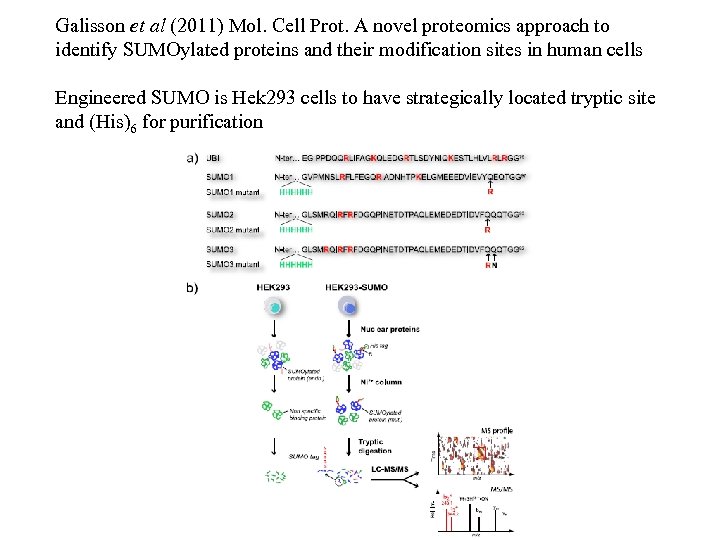
Galisson et al (2011) Mol. Cell Prot. A novel proteomics approach to identify SUMOylated proteins and their modification sites in human cells Engineered SUMO is Hek 293 cells to have strategically located tryptic site and (His)6 for purification
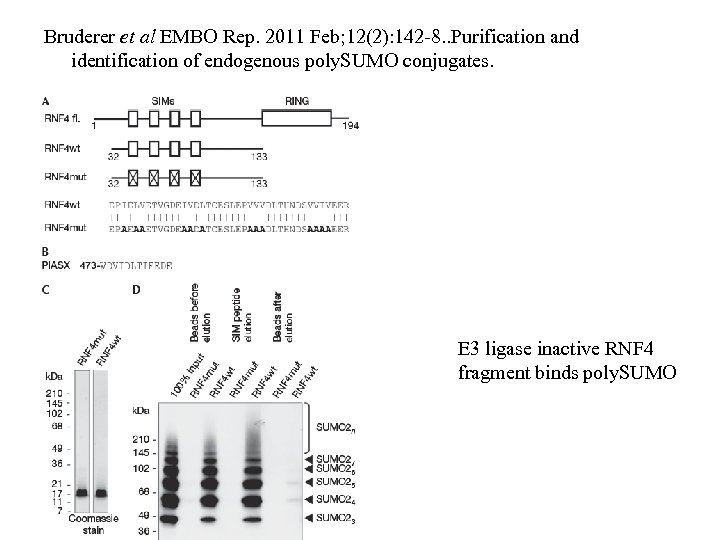
Bruderer et al EMBO Rep. 2011 Feb; 12(2): 142 -8. . Purification and identification of endogenous poly. SUMO conjugates. E 3 ligase inactive RNF 4 fragment binds poly. SUMO

Glycosylation Common, up to 50% of human proteins are glycosylated Need to fond site of attached, using N-linked (Asn) or O-linked (Ser) Also need to determine structure of glycosyl group Very complex, highly combinatorial The most challenging PTM for high throughput proteomics Enrichment possible with lectin affinity chromatography
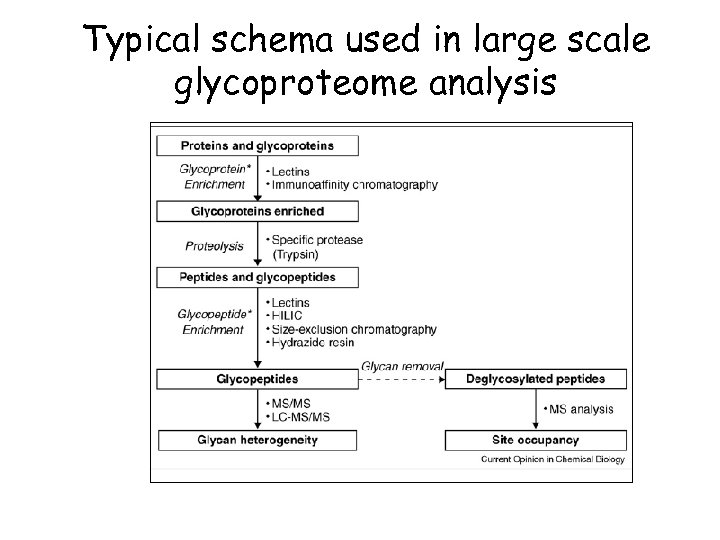
Typical schema used in large scale glycoproteome analysis
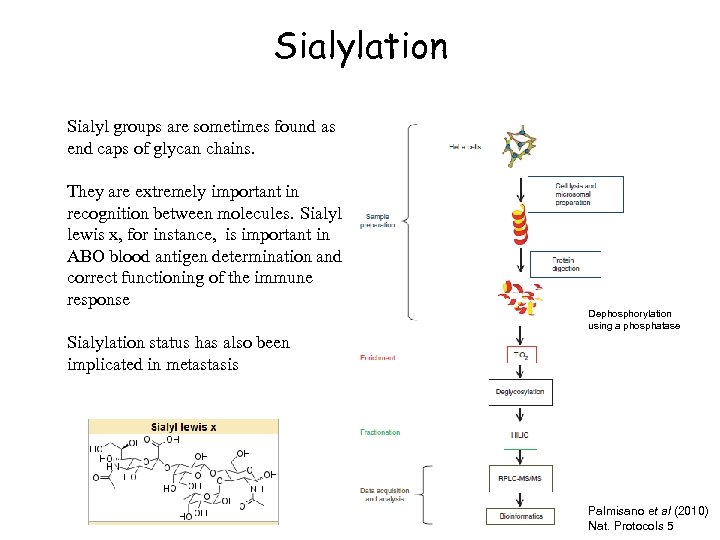
Sialylation Sialyl groups are sometimes found as end caps of glycan chains. They are extremely important in recognition between molecules. Sialyl lewis x, for instance, is important in ABO blood antigen determination and correct functioning of the immune response Dephosphorylation using a phosphatase Sialylation status has also been implicated in metastasis Palmisano et al (2010) Nat. Protocols 5
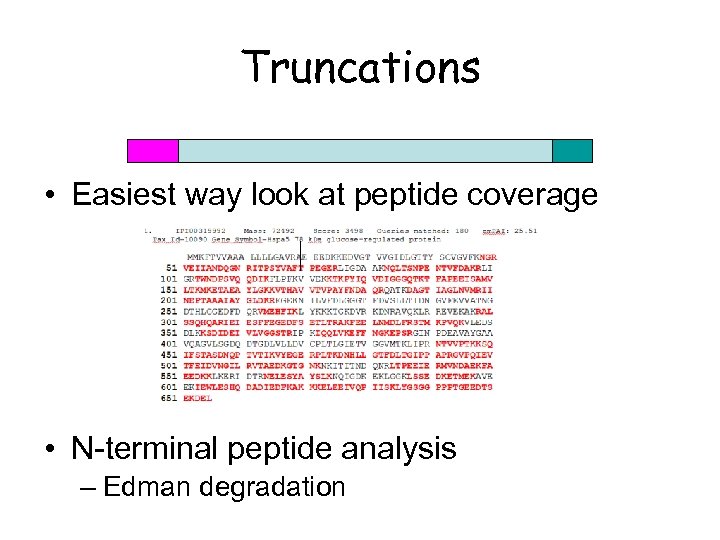
Truncations • Easiest way look at peptide coverage • N-terminal peptide analysis – Edman degradation
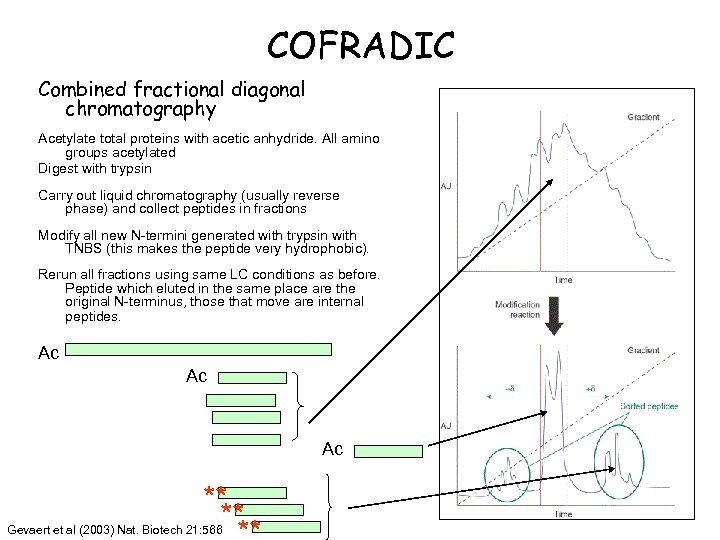
COFRADIC Combined fractional diagonal chromatography Acetylate total proteins with acetic anhydride. All amino groups acetylated Digest with trypsin Carry out liquid chromatography (usually reverse phase) and collect peptides in fractions Modify all new N-termini generated with trypsin with TNBS (this makes the peptide very hydrophobic). Rerun all fractions using same LC conditions as before. Peptide which eluted in the same place are the original N-terminus, those that move are internal peptides. Ac Ac Ac ** ** Gevaert et al (2003) Nat. Biotech 21: 566

This week • Tuesday: – 1 pm Seminar by Matthias Mann – 2. 15 pm Q and A session with Prof. Mann and lecture for me • Wednesday: – 9 am – Practical Class – 4 pm Lecture from me
86aa9df2c38eab1a2872f6a309769ef0.ppt