8cd3a98418b9e445bcba054453faf305.ppt
- Количество слайдов: 26
 P 300 Marks Active Enhancers Ruijuan Li Chao He Rui Fu
P 300 Marks Active Enhancers Ruijuan Li Chao He Rui Fu
 Main contents • Background and conclusion • Experimental approaches • Data processing • Summary and discussion
Main contents • Background and conclusion • Experimental approaches • Data processing • Summary and discussion
 Background • Problems ØEnhancers • Previous researches ØExisting work • Authors introduction ØResearch interests
Background • Problems ØEnhancers • Previous researches ØExisting work • Authors introduction ØResearch interests
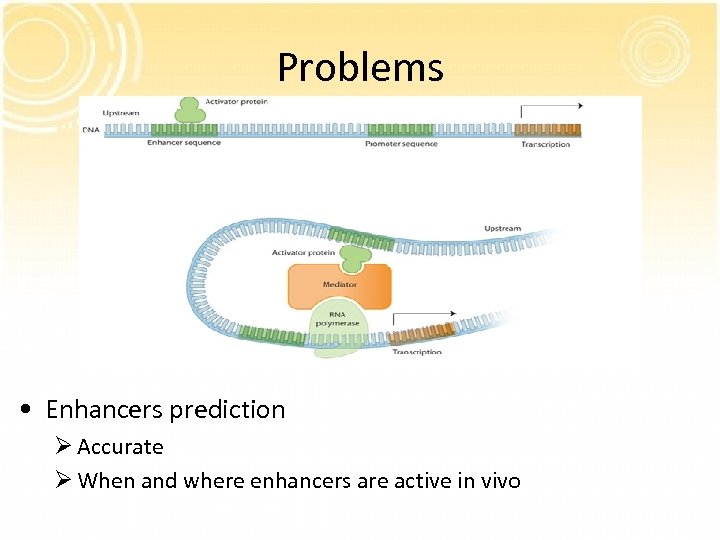 Problems • Enhancers prediction Ø Accurate Ø When and where enhancers are active in vivo
Problems • Enhancers prediction Ø Accurate Ø When and where enhancers are active in vivo
 Previous researches • Comparative genome methods ØEvolutionary sequence constraint failed to reveal when and where enhancers are active in vivo ØSome regulatory elements are not sufficiently conserved to be detectable. • Conservation-independent approach ØCh. IP-seq with an antibody specific for an enhancerbinding protein • P 300 has been showed to be associated with enhancers
Previous researches • Comparative genome methods ØEvolutionary sequence constraint failed to reveal when and where enhancers are active in vivo ØSome regulatory elements are not sufficiently conserved to be detectable. • Conservation-independent approach ØCh. IP-seq with an antibody specific for an enhancerbinding protein • P 300 has been showed to be associated with enhancers
 Authors introduction • Axel Visel – Staff scientist in the Genomics Division, Lawrence Berkeley National Laboratory – Comparative genomics, sequencing-based chromatin studies (Ch. IP-seq), and transgenic reporter assays – Systematic identification and functional characterization of distant-acting enhancers • Matthew J. Blow – Comparative genomics, RNA editing
Authors introduction • Axel Visel – Staff scientist in the Genomics Division, Lawrence Berkeley National Laboratory – Comparative genomics, sequencing-based chromatin studies (Ch. IP-seq), and transgenic reporter assays – Systematic identification and functional characterization of distant-acting enhancers • Matthew J. Blow – Comparative genomics, RNA editing
 • Prabhakar S, Visel A, . . . , Pennacchio LA, Rubin EM, Noonan JP (13 authors). Human-specific gain of function in a developmental enhancer. Science 2008 • Visel A, . . . , Rubin EM, Pennacchio LA (10 authors). Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nature Genet 2008 • Rahimov F, Marazita ML, Visel A, . . . , Murray JC (23 authors). Disruption of an AP-2 alpha binding site in an IRF 6 enhancer is associated with cleft lip. Nature Genet 2008 • De Val S, . . . , Visel A, . . . , Black BL (15 authors). Combinatorial regulation of endothelial gene expression by ets and forkhead transcription factors. Cell 2008 • Lein ES, Hawrylycz MJ, . . . , Visel A, . . . , Jones AR (108 authors). Genomewide atlas of gene expression in the adult mouse brain. Nature 2007 • Pennacchio LA, . . . , Visel A, Rubin EM (19 authors). In vivo enhancer analysis of human conserved non-coding sequences. Nature 2006
• Prabhakar S, Visel A, . . . , Pennacchio LA, Rubin EM, Noonan JP (13 authors). Human-specific gain of function in a developmental enhancer. Science 2008 • Visel A, . . . , Rubin EM, Pennacchio LA (10 authors). Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nature Genet 2008 • Rahimov F, Marazita ML, Visel A, . . . , Murray JC (23 authors). Disruption of an AP-2 alpha binding site in an IRF 6 enhancer is associated with cleft lip. Nature Genet 2008 • De Val S, . . . , Visel A, . . . , Black BL (15 authors). Combinatorial regulation of endothelial gene expression by ets and forkhead transcription factors. Cell 2008 • Lein ES, Hawrylycz MJ, . . . , Visel A, . . . , Jones AR (108 authors). Genomewide atlas of gene expression in the adult mouse brain. Nature 2007 • Pennacchio LA, . . . , Visel A, Rubin EM (19 authors). In vivo enhancer analysis of human conserved non-coding sequences. Nature 2006
 Corresponding Author • Len A. Pennacchio • Molecular biologist, Senior staff scientist in the Genomics Division, Lawrence Berkeley National Laboratory • Head of the Genetic Analysis Program and the Genomic Technologies Program, Joint Genome Institute • 2007 White House Presidential Early Career Award for Scientists and Engineers (PECASE); • Contributed to the human genome project with an analysis of human chromosome 16. • Gene regulation, the genetic basis of differences in body shape between different individuals, conserved sequences in the genome, and connections between junk and heart disease
Corresponding Author • Len A. Pennacchio • Molecular biologist, Senior staff scientist in the Genomics Division, Lawrence Berkeley National Laboratory • Head of the Genetic Analysis Program and the Genomic Technologies Program, Joint Genome Institute • 2007 White House Presidential Early Career Award for Scientists and Engineers (PECASE); • Contributed to the human genome project with an analysis of human chromosome 16. • Gene regulation, the genetic basis of differences in body shape between different individuals, conserved sequences in the genome, and connections between junk and heart disease
 Conclusion • P 300 binding sites accurately identifies enhancers and their associated activities in vivo. • The data will be useful to study the role of tissue-specific enhancers in human biology and disease on a genome-wide scale.
Conclusion • P 300 binding sites accurately identifies enhancers and their associated activities in vivo. • The data will be useful to study the role of tissue-specific enhancers in human biology and disease on a genome-wide scale.
 Experimental approaches Ch. IP-seq: map p 300 binding sequences in vivo Transgenic mouse enhancer assay: test the activity of predicted p 300 peaks
Experimental approaches Ch. IP-seq: map p 300 binding sequences in vivo Transgenic mouse enhancer assay: test the activity of predicted p 300 peaks
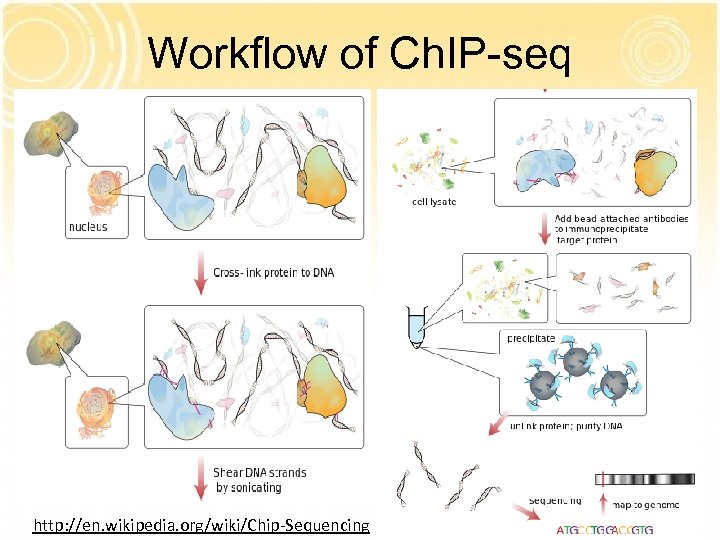 Workflow of Ch. IP-seq http: //en. wikipedia. org/wiki/Chip-Sequencing
Workflow of Ch. IP-seq http: //en. wikipedia. org/wiki/Chip-Sequencing
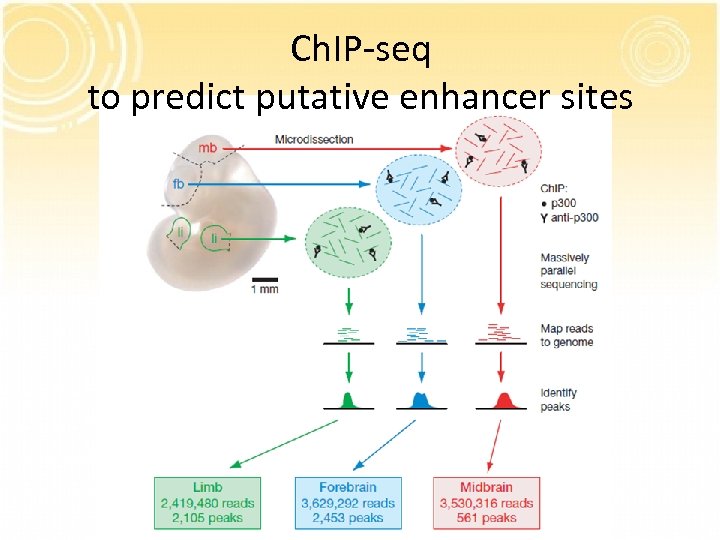 Ch. IP-seq to predict putative enhancer sites
Ch. IP-seq to predict putative enhancer sites
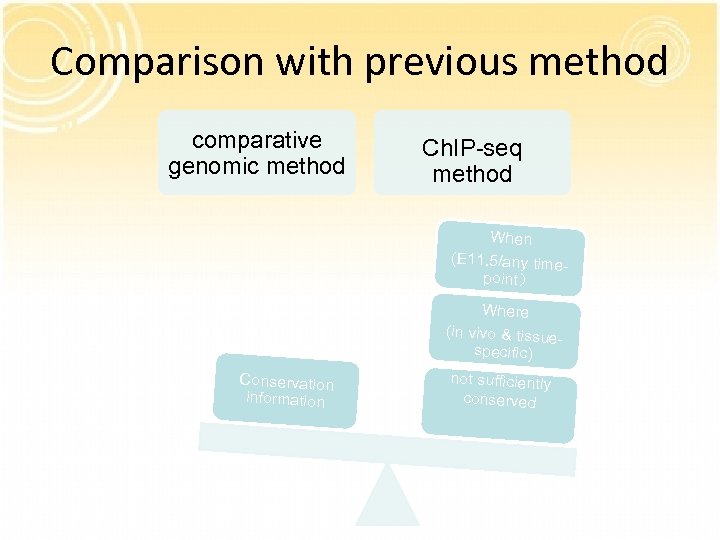 Comparison with previous method comparative genomic method Ch. IP-seq method When (E 11. 5/any time- point) Conservation information Where (in vivo & tissue. Regulatory specific) elements not sufficiently conserved
Comparison with previous method comparative genomic method Ch. IP-seq method When (E 11. 5/any time- point) Conservation information Where (in vivo & tissue. Regulatory specific) elements not sufficiently conserved
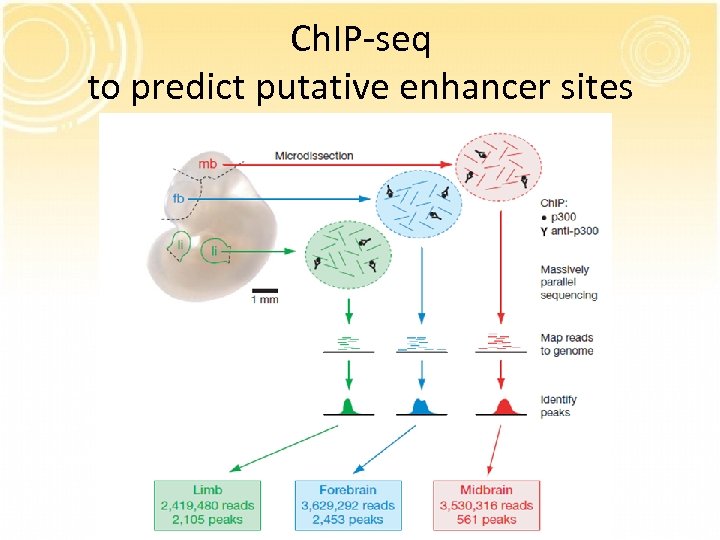 Ch. IP-seq to predict putative enhancer sites
Ch. IP-seq to predict putative enhancer sites
 Workflow of transgenic mouse enhancer assay Selection of 86 candidate regions for testing Cloning human genomic sequences into Hsp 68 -Lac. Z reporter vector Pronuclear injection and staining at E 11. 5
Workflow of transgenic mouse enhancer assay Selection of 86 candidate regions for testing Cloning human genomic sequences into Hsp 68 -Lac. Z reporter vector Pronuclear injection and staining at E 11. 5
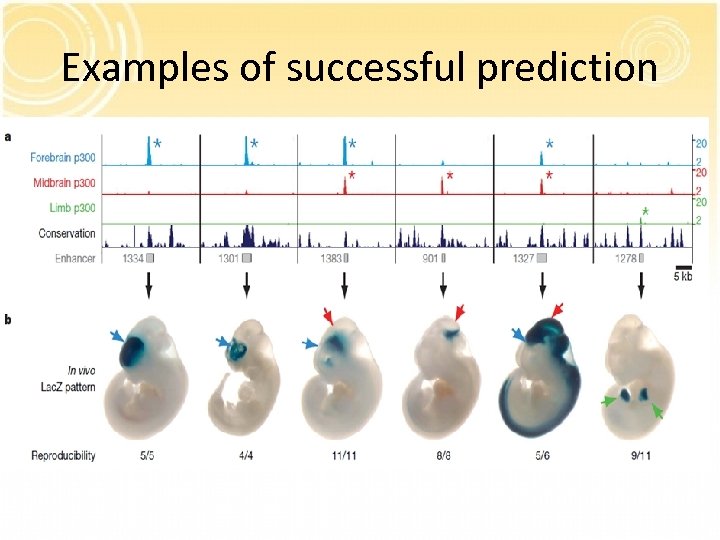 Examples of successful prediction
Examples of successful prediction
 Transgenic mouse enhancer assay • Advantage accurate • Disadvantage effect of endogenous regulatory elements
Transgenic mouse enhancer assay • Advantage accurate • Disadvantage effect of endogenous regulatory elements
 Data processing • Peak Calling to identify p 300 binding sites • Validate p 300 binding sites are active enhancers
Data processing • Peak Calling to identify p 300 binding sites • Validate p 300 binding sites are active enhancers
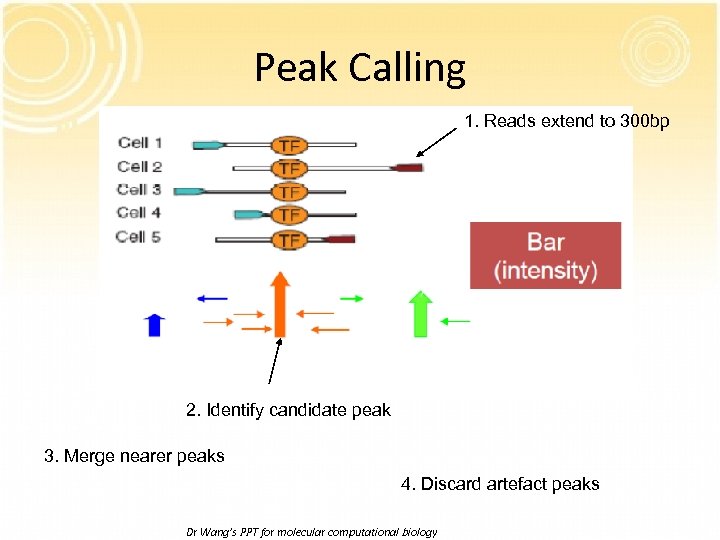 Peak Calling 1. Reads extend to 300 bp 2. Identify candidate peak 3. Merge nearer peaks 4. Discard artefact peaks Dr Wang’s PPT for molecular computational biology
Peak Calling 1. Reads extend to 300 bp 2. Identify candidate peak 3. Merge nearer peaks 4. Discard artefact peaks Dr Wang’s PPT for molecular computational biology
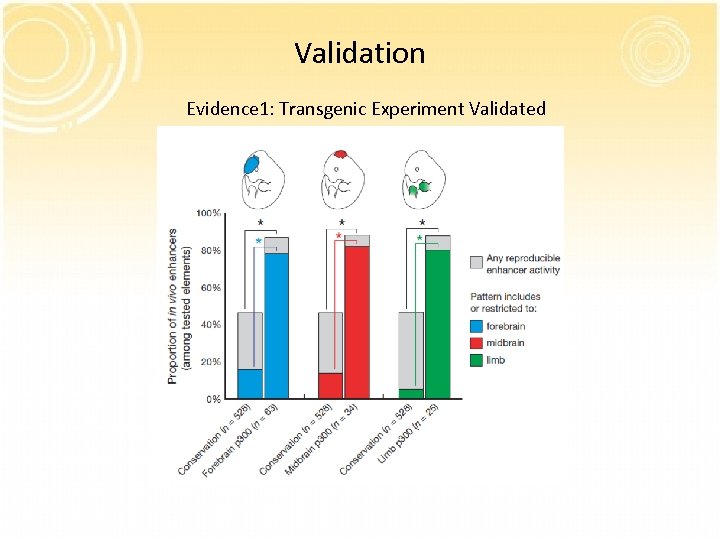 Validation Evidence 1: Transgenic Experiment Validated
Validation Evidence 1: Transgenic Experiment Validated
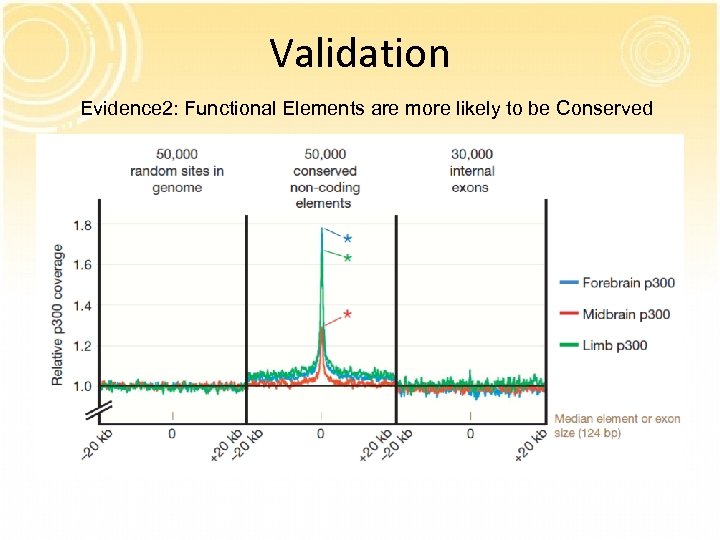 Validation Evidence 2: Functional Elements are more likely to be Conserved
Validation Evidence 2: Functional Elements are more likely to be Conserved
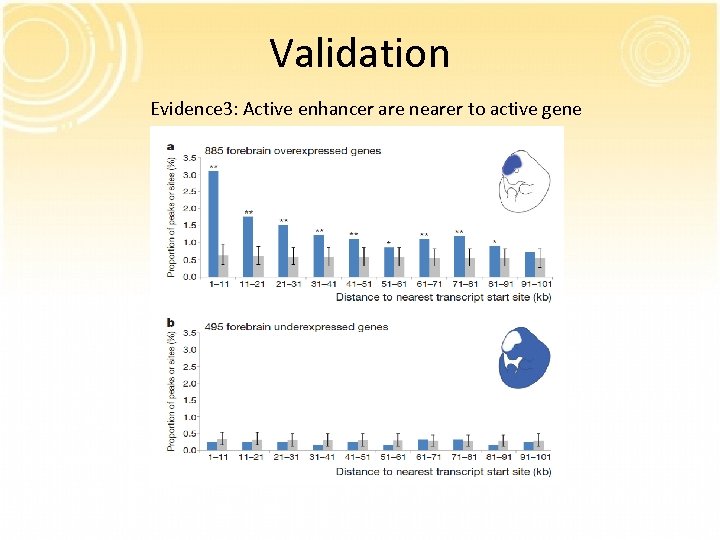 Validation Evidence 3: Active enhancer are nearer to active gene
Validation Evidence 3: Active enhancer are nearer to active gene
 Summary • p 300 binding sites are probably to be celltype specific enhancer. • Most p 300 -bound regions are conserved • p 300 binding sites are significant nearer to expressed genes than random sites.
Summary • p 300 binding sites are probably to be celltype specific enhancer. • Most p 300 -bound regions are conserved • p 300 binding sites are significant nearer to expressed genes than random sites.
 Improvement • Peak Calling Arbitrary to extend the reads to 300 bp No control to test peak quality • Active enhancer prediction Sensitivity is low Still some errors
Improvement • Peak Calling Arbitrary to extend the reads to 300 bp No control to test peak quality • Active enhancer prediction Sensitivity is low Still some errors
 Nowadays • More marks: H 3 K 4 me 1, H 3 K 27 ac • Enhancers are transcribing bidirectional sm. RNA • Locate enhancers target promoters by new experiments
Nowadays • More marks: H 3 K 4 me 1, H 3 K 27 ac • Enhancers are transcribing bidirectional sm. RNA • Locate enhancers target promoters by new experiments
 Thank you
Thank you


