65ef8846124d0c3c4c234fc173c7cbb7.ppt
- Количество слайдов: 56
 Order of progress reports March 11 1. Yamrus/Wirth 2. Terry/Kolansky 3. Hoffert/Sluhocki March 18 1. Pero/Bartlow 2. Janosov/Jescavage/Keiser 3. Ciaston/Suchocki/Yuhas March 25 1. Marx 2. Bird
Order of progress reports March 11 1. Yamrus/Wirth 2. Terry/Kolansky 3. Hoffert/Sluhocki March 18 1. Pero/Bartlow 2. Janosov/Jescavage/Keiser 3. Ciaston/Suchocki/Yuhas March 25 1. Marx 2. Bird
 How to make a poster Dina Mandoli, U Washington http: //www. aspb. org/education/poster. cfm
How to make a poster Dina Mandoli, U Washington http: //www. aspb. org/education/poster. cfm
 Characters of good posters 1. Readable: ideas flow easily
Characters of good posters 1. Readable: ideas flow easily
 Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet
Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet
 Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet 3. Well-organized: elements are arranged logically
Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet 3. Well-organized: elements are arranged logically
 Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet 3. Well-organized: elements are arranged logically 4. Succinct: you have 11 seconds to get your audience’s atention
Characters of good posters 1. Readable: ideas flow easily 2. Legible: can easily read from 10 feet 3. Well-organized: elements are arranged logically 4. Succinct: you have 11 seconds to get your audience’s atention
 Getting started 1. Choose the main message
Getting started 1. Choose the main message
 Getting started 1. Choose the main message 2. Lay out panels crudely to fit space available
Getting started 1. Choose the main message 2. Lay out panels crudely to fit space available
 Getting started 1. Choose the main message 2. Lay out panels crudely to fit space available 3. KISS ( keep it simple, stupid!) • Remove all extraneous material
Getting started 1. Choose the main message 2. Lay out panels crudely to fit space available 3. KISS ( keep it simple, stupid!) • Remove all extraneous material
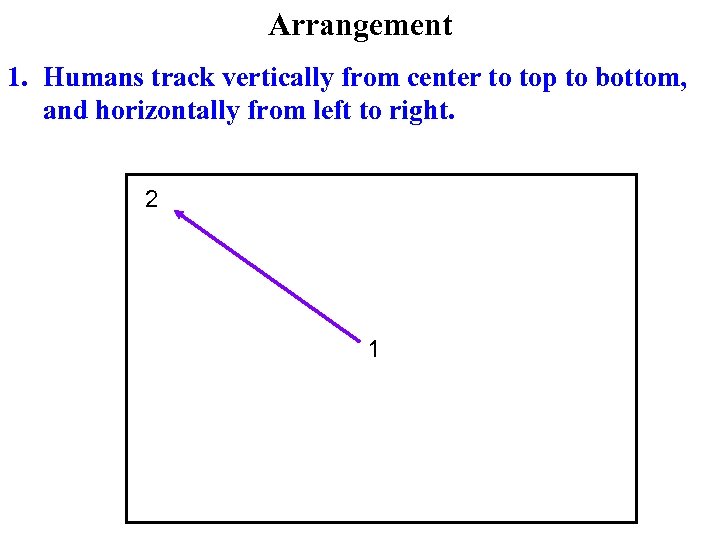 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 1
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 1
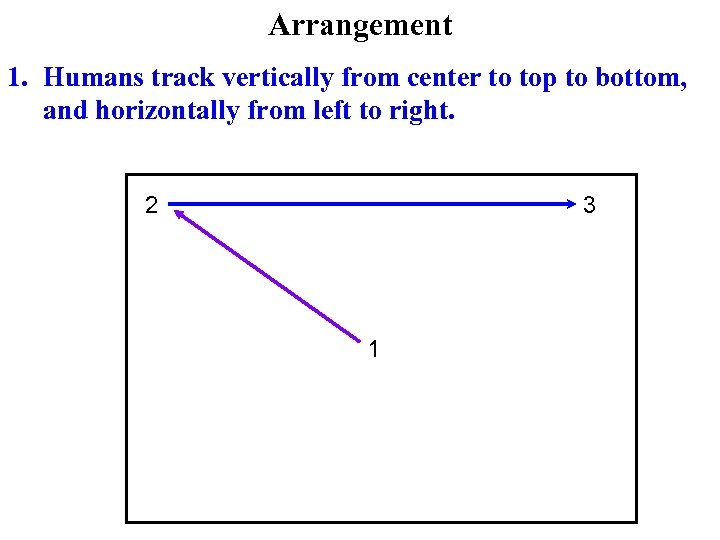 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 3 1
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 3 1
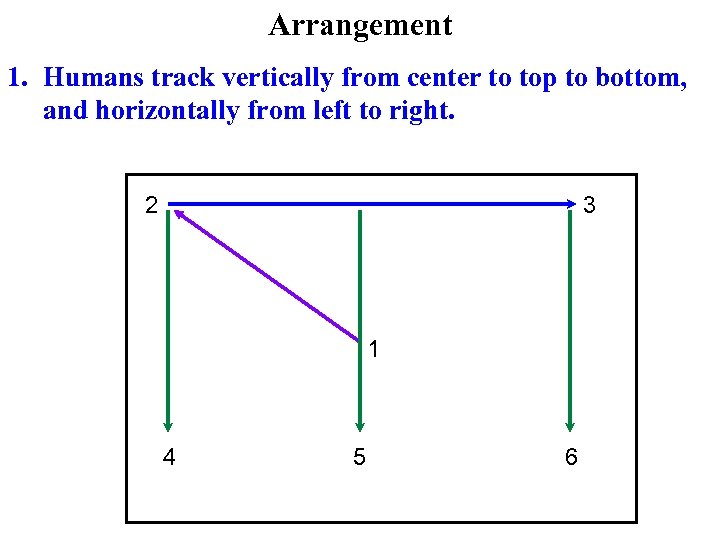 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 3 1 4 5 6
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. 2 3 1 4 5 6
 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title Naked photo at Sydney Opera House highlights openness 1
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title Naked photo at Sydney Opera House highlights openness 1
 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 people posed naked for a Spencer Tunick photo to show that Australia is a free & equal society 1
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 people posed naked for a Spencer Tunick photo to show that Australia is a free & equal society 1
 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia is a free & equal society 1
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia is a free & equal society 1
 Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia 2) Many is a free & equal 1 society Aussies support gays
Arrangement 1. Humans track vertically from center to top to bottom, and horizontally from left to right. • Put your best pictures in center, just below title • Put your key text elements in upper corners Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia 2) Many is a free & equal 1 society Aussies support gays
 • • • Arrangement Put your best pictures in center, just below title Put your key text elements in upper corners Put your remaining material down the page in decreasing order of importance Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia 2) Many is a free & equal 1 society Aussies Introduction: support gays Discussion:
• • • Arrangement Put your best pictures in center, just below title Put your key text elements in upper corners Put your remaining material down the page in decreasing order of importance Naked photo at Sydney Opera House highlights openness Abstract: >5, 000 Conclusions people posed naked 1) Australia for a Spencer is a free & Tunick photo to equal society show that Australia 2) Many is a free & equal 1 society Aussies Introduction: support gays Discussion:
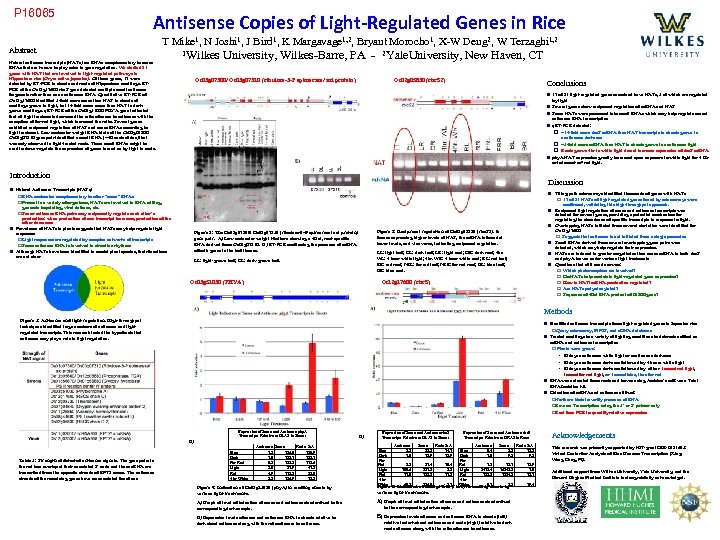 P 16065 Antisense Copies of Light-Regulated Genes in Rice Abstract T Mike 1, N Joshi 1, J Bird 1, K Margavage 1, 2, Bryant Morocho 1, X-W Deng 2, W Terzaghi 1, 2 Natural antisense transcripts (NATs) are RNAs complementary to sense RNAs that are known to play roles in gene regulation. We studied 21 genes with NAT that are involved in light-regulated pathways in Nipponbare rice (Oryza sativa japonica). Of these genes, 17 were detected by RT-PCR in shoots and roots of Nipponbare seedlings. RTPCR of the Os 12 g 17600 rbc. S gene detected multiple small antisense fragments rather than one continuous RNA. Quantitative RT-PCR of Os 12 g 17600 identified 5 -fold more sense than NAT in shoots of seedlings grown in light, but 14 -fold more sense than NAT in darkgrown seedlings. q. RT-PCR of the Os 03 g 51030 PHYA gene indicated that all light treatments decreased the ratio of sense to antisense with the exception of far-red light, which increased the ratio. Several genes exhibited reciprocal regulation of NAT and sense RNAs according to light treatment. Low molecular weight RNA blots of the Os 03 g 07300/ Os 03 g 07310 gene pair identified a small RNA (~40 nucleotides) that was only observed in light-treated roots. These small RNAs might be used to down-regulate the expression of genes turned on by light in roots. 1 Wilkes University, Wilkes-Barre, PA - 2 Yale. University, New Haven, CT Os 03 g 07300/ Os 03 g 07310 (ribulose-3 -P epimerase/ axi protein) Os 02 g 05830 (rbc. S 2) Conclusions n 17 of 21 light-regulated genes examined have NATs, 5 of which are regulated by light n Several genes show reciprocal regulation of m. RNA and NAT n Some NATs were processed into small RNAs which may help regulate sense/ antisense RNA transcription n q. RT-PCR detected: q ~14 -fold more rbc. S m. RNA than NAT transcripts in shoots grown in continuous darkness q ~5 -fold more m. RNA than NAT in shoots grown in continuous light q Roots grown 4 hr in white light do not increase expression of rbc. S m. RNA n phy. A NAT expression greatly increased upon exposure to white light for 4 Hr or to 1 mmol. m 2 red light. Introduction Discussion n Natural Antisense Transcripts (NATs) q RNA molecules complementary to other “sense” RNAs q Present in a variety of organisms, NATs are involved in RNA editing, genomic imprinting, viral defense, etc. q Sense-antisense RNA pairs may reciprocally regulate each other’s production: when production of one transcript increases, production of the other decreases n Prevalence of NATs in plants suggests that NATs may help regulate light responses q Light responses are regulated by complex networks of transcripts q Some antisense RNA is involved in circadian rhythms n Although NATs have been identified in model plant species, their functions are not clear n Tiling-path microarrays identified thousands of genes with NATs q 17 of 21 NATs of light-regulated genes found by microarrays were confirmed, validating this high-throughput approach. n Reciprocal light regulation of sense and antisense transcripts was detected for several genes, providing a potential mechanism for regulating the abundance of specific transcripts in response to light. n Overlapping NATs initiated from several start sites were identified for Os 12 g 17600 q Suggests that antisense is not initiated from a single promoter. n Small RNAs derived from several overlapping gene pairs were detected, which may help regulate their expression. n NATs are induced in greater magnitudes than sense m. RNA in both rbc. S and phy. A leaves under various light treatments n Questions that still need answers: q Which photoreceptors are involved? q Do NATs help modulate light-regulated gene expression? q How is NAT/ m. RNA production regulated? q Are NATs polyadenylated? q Sequence of 40 nt RNA product of 07300 gene? Figure 3: Reciprocal regulation of Os 02 g 05830 (rbc. S 2). In tissues expressing higher levels of NAT, the m. RNA is found at lower levels, and vice versa, indicating reciprocal regulation. Figure 2: The Os 03 g 07300/ Os 03 g 07310 (ribulose-3 -P epimerase/ axi protein) gene pair. A) Low molecular weight Northern showing a 40 nt, root-specific RNA derived from Os 03 g 07310. B) RT-PCR confirming the presence of m. RNA of both genes in the leaf tissues. LL: light leaf; DL: dark leaf; LR: light root; DR: dark root; 4 hr. WL: 4 hour white light; 4 hr. WR: 4 hour white root; RL: red leaf; RR: red root; FRL: far red leaf; FRR: far red root; BL: blue leaf; BR: blue root. LL: light-grown leaf; DL: dark-grown leaf. Os 03 g 51030 (PHYA) Os 12 g 17600 (rbc. S) Methods Figure 1: Antisense and light regulation. High-throughput techniques identified large numbers of antisense and lightregulated transcripts. This research tested the hypothesis that antisense may play a role in light regulation. n Identified antisense transcripts from light-regulated genes in Japonica rice q Query microarray, MPSS, and c. DNA databases n Treated seedlings to a variety of lighting conditions to determine effect on m. RNA and antisense transcription q Plants were grown: § 10 days continuous white light or continuous darkness § 10 days continuous darkness followed by 4 hours white light § 10 days continuous darkness followed by either 1 mmol red light, 1 mmol far red light, or 1 mmol blue, then far red n RNA was extracted from roots and leaves using Ambion’s mi. Rvana Total RNA Isolation kit. n Detection of m. RNA and antisense utilized: q. Northern blots to verify presence of RNA q. Reverse Transcription using the 5’ or 3’ primer only q. Real time PCR to quantify relative expression Expression of Sense and Antisense phy. A Transcripts Relative to DLAS in Shoots B) Table 1: Strength of detected antisense signals. The gene pairs in the red box overlap at their annotated 3’ ends and the sno. RNA are transcribed from the opposite strands of RPT 2 exons. The antisense strands of the remaining genes have no annotated functions. Blue Dark Far Red Light Red 4 hr White Antisense Sense Ratio S: A 1. 2 156. 0 130. 9 1. 0 152. 1 0. 8 132. 2 172. 6 2. 0 81. 9 41. 8 4. 9 112. 3 23. 0 2. 5 186. 9 75. 3 B) Expression of Sense and Antisense rbc. S Transcripts Relative to DLAS in Shoots Blue Dark Far Red Light Red 4 hr White Antisense 5. 8 1. 0 Sense Ratio S: A 85. 3 14. 7 13. 9 5. 5 100. 6 11. 0 57. 4 527. 3 122. 5 10. 4 5. 2 11. 2 40. 2 284. 0 7. 1 Expression of Sense and Antisense rbc. S Transcripts Relative to DRAS in Roots Blue Dark Far Red Light Red 4 hr White Antisense 0. 4 1. 0 Sense Ratio S: A 5. 5 12. 5 9. 8 1. 2 2418. 4 153. 8 15. 7 16842. 5 1854. 2 12. 9 7. 0 12. 1 0. 1 2. 5 19. 4 Figure 4: Induction of Os 03 g 51030 (phy. A) in seedling shoots by various light treatments. Figure 5: Induction of Os 12 g 17600 (rbc. S) in seedling shoots by various light treatments. A) Graph of level of induction of sense and antisense standardized to the corresponding dark sample. A) Graph of level of induction of sense and antisense standardized B) Expression levels of sense and antisense RNA in shoots relative to dark shoot antisense along with the ratio of sense to antisense. B) Expression levels of sense and antisense RNA in shoots (left) to the corresponding dark sample. relative to dark shoot antisense and roots (right) relative to dark root antisense along with the ratio of sense to antisense. Acknowledgements This research was primarily supported by NSF grant DBI-0421675: Virtual Center for Analysis of Rice Genome Transcription (Xing. Wang Deng, PI). Additional support from Wilkes University, Yale University, and the Howard Hughes Medical Institute is also gratefully acknowledged.
P 16065 Antisense Copies of Light-Regulated Genes in Rice Abstract T Mike 1, N Joshi 1, J Bird 1, K Margavage 1, 2, Bryant Morocho 1, X-W Deng 2, W Terzaghi 1, 2 Natural antisense transcripts (NATs) are RNAs complementary to sense RNAs that are known to play roles in gene regulation. We studied 21 genes with NAT that are involved in light-regulated pathways in Nipponbare rice (Oryza sativa japonica). Of these genes, 17 were detected by RT-PCR in shoots and roots of Nipponbare seedlings. RTPCR of the Os 12 g 17600 rbc. S gene detected multiple small antisense fragments rather than one continuous RNA. Quantitative RT-PCR of Os 12 g 17600 identified 5 -fold more sense than NAT in shoots of seedlings grown in light, but 14 -fold more sense than NAT in darkgrown seedlings. q. RT-PCR of the Os 03 g 51030 PHYA gene indicated that all light treatments decreased the ratio of sense to antisense with the exception of far-red light, which increased the ratio. Several genes exhibited reciprocal regulation of NAT and sense RNAs according to light treatment. Low molecular weight RNA blots of the Os 03 g 07300/ Os 03 g 07310 gene pair identified a small RNA (~40 nucleotides) that was only observed in light-treated roots. These small RNAs might be used to down-regulate the expression of genes turned on by light in roots. 1 Wilkes University, Wilkes-Barre, PA - 2 Yale. University, New Haven, CT Os 03 g 07300/ Os 03 g 07310 (ribulose-3 -P epimerase/ axi protein) Os 02 g 05830 (rbc. S 2) Conclusions n 17 of 21 light-regulated genes examined have NATs, 5 of which are regulated by light n Several genes show reciprocal regulation of m. RNA and NAT n Some NATs were processed into small RNAs which may help regulate sense/ antisense RNA transcription n q. RT-PCR detected: q ~14 -fold more rbc. S m. RNA than NAT transcripts in shoots grown in continuous darkness q ~5 -fold more m. RNA than NAT in shoots grown in continuous light q Roots grown 4 hr in white light do not increase expression of rbc. S m. RNA n phy. A NAT expression greatly increased upon exposure to white light for 4 Hr or to 1 mmol. m 2 red light. Introduction Discussion n Natural Antisense Transcripts (NATs) q RNA molecules complementary to other “sense” RNAs q Present in a variety of organisms, NATs are involved in RNA editing, genomic imprinting, viral defense, etc. q Sense-antisense RNA pairs may reciprocally regulate each other’s production: when production of one transcript increases, production of the other decreases n Prevalence of NATs in plants suggests that NATs may help regulate light responses q Light responses are regulated by complex networks of transcripts q Some antisense RNA is involved in circadian rhythms n Although NATs have been identified in model plant species, their functions are not clear n Tiling-path microarrays identified thousands of genes with NATs q 17 of 21 NATs of light-regulated genes found by microarrays were confirmed, validating this high-throughput approach. n Reciprocal light regulation of sense and antisense transcripts was detected for several genes, providing a potential mechanism for regulating the abundance of specific transcripts in response to light. n Overlapping NATs initiated from several start sites were identified for Os 12 g 17600 q Suggests that antisense is not initiated from a single promoter. n Small RNAs derived from several overlapping gene pairs were detected, which may help regulate their expression. n NATs are induced in greater magnitudes than sense m. RNA in both rbc. S and phy. A leaves under various light treatments n Questions that still need answers: q Which photoreceptors are involved? q Do NATs help modulate light-regulated gene expression? q How is NAT/ m. RNA production regulated? q Are NATs polyadenylated? q Sequence of 40 nt RNA product of 07300 gene? Figure 3: Reciprocal regulation of Os 02 g 05830 (rbc. S 2). In tissues expressing higher levels of NAT, the m. RNA is found at lower levels, and vice versa, indicating reciprocal regulation. Figure 2: The Os 03 g 07300/ Os 03 g 07310 (ribulose-3 -P epimerase/ axi protein) gene pair. A) Low molecular weight Northern showing a 40 nt, root-specific RNA derived from Os 03 g 07310. B) RT-PCR confirming the presence of m. RNA of both genes in the leaf tissues. LL: light leaf; DL: dark leaf; LR: light root; DR: dark root; 4 hr. WL: 4 hour white light; 4 hr. WR: 4 hour white root; RL: red leaf; RR: red root; FRL: far red leaf; FRR: far red root; BL: blue leaf; BR: blue root. LL: light-grown leaf; DL: dark-grown leaf. Os 03 g 51030 (PHYA) Os 12 g 17600 (rbc. S) Methods Figure 1: Antisense and light regulation. High-throughput techniques identified large numbers of antisense and lightregulated transcripts. This research tested the hypothesis that antisense may play a role in light regulation. n Identified antisense transcripts from light-regulated genes in Japonica rice q Query microarray, MPSS, and c. DNA databases n Treated seedlings to a variety of lighting conditions to determine effect on m. RNA and antisense transcription q Plants were grown: § 10 days continuous white light or continuous darkness § 10 days continuous darkness followed by 4 hours white light § 10 days continuous darkness followed by either 1 mmol red light, 1 mmol far red light, or 1 mmol blue, then far red n RNA was extracted from roots and leaves using Ambion’s mi. Rvana Total RNA Isolation kit. n Detection of m. RNA and antisense utilized: q. Northern blots to verify presence of RNA q. Reverse Transcription using the 5’ or 3’ primer only q. Real time PCR to quantify relative expression Expression of Sense and Antisense phy. A Transcripts Relative to DLAS in Shoots B) Table 1: Strength of detected antisense signals. The gene pairs in the red box overlap at their annotated 3’ ends and the sno. RNA are transcribed from the opposite strands of RPT 2 exons. The antisense strands of the remaining genes have no annotated functions. Blue Dark Far Red Light Red 4 hr White Antisense Sense Ratio S: A 1. 2 156. 0 130. 9 1. 0 152. 1 0. 8 132. 2 172. 6 2. 0 81. 9 41. 8 4. 9 112. 3 23. 0 2. 5 186. 9 75. 3 B) Expression of Sense and Antisense rbc. S Transcripts Relative to DLAS in Shoots Blue Dark Far Red Light Red 4 hr White Antisense 5. 8 1. 0 Sense Ratio S: A 85. 3 14. 7 13. 9 5. 5 100. 6 11. 0 57. 4 527. 3 122. 5 10. 4 5. 2 11. 2 40. 2 284. 0 7. 1 Expression of Sense and Antisense rbc. S Transcripts Relative to DRAS in Roots Blue Dark Far Red Light Red 4 hr White Antisense 0. 4 1. 0 Sense Ratio S: A 5. 5 12. 5 9. 8 1. 2 2418. 4 153. 8 15. 7 16842. 5 1854. 2 12. 9 7. 0 12. 1 0. 1 2. 5 19. 4 Figure 4: Induction of Os 03 g 51030 (phy. A) in seedling shoots by various light treatments. Figure 5: Induction of Os 12 g 17600 (rbc. S) in seedling shoots by various light treatments. A) Graph of level of induction of sense and antisense standardized to the corresponding dark sample. A) Graph of level of induction of sense and antisense standardized B) Expression levels of sense and antisense RNA in shoots relative to dark shoot antisense along with the ratio of sense to antisense. B) Expression levels of sense and antisense RNA in shoots (left) to the corresponding dark sample. relative to dark shoot antisense and roots (right) relative to dark root antisense along with the ratio of sense to antisense. Acknowledgements This research was primarily supported by NSF grant DBI-0421675: Virtual Center for Analysis of Rice Genome Transcription (Xing. Wang Deng, PI). Additional support from Wilkes University, Yale University, and the Howard Hughes Medical Institute is also gratefully acknowledged.
 Appearance (10 pts total) Type, figures large enough to see from 10 feet? Figures and tables clear, legible, and nicely presented? Good ratio of figures to text? Artistic impression Effectiveness of presentation (40 pts total) Effective title? Readability? Organization? Is abstract concise and prominently displayed? Does abstract have all elements & summarize the poster? Do conclusions concisely summarize the major results? Are conclusions supported by data given on the poster? Is introduction concise, yet adequate? Do figure and table titles concisely summarize key results? Are captions adequate & distinct from figure and table titles? Are figures and tables clear and easily understood? Are the various elements effectively arranged? Is discussion concise, yet explain each result? English (spelling, grammar) Overall: did it effectively deliver the key points? 1 pts ______ 2 pts ______ 3 pts ______ 4 pts_______ 1 pt ____________ 3 pts_______ 2 pts_____________ 4 pts_______ 2 pts_______ 3 pts_______ 1 pts_____________ 2 pts_____________ 3 pts_______ 5 pts_______
Appearance (10 pts total) Type, figures large enough to see from 10 feet? Figures and tables clear, legible, and nicely presented? Good ratio of figures to text? Artistic impression Effectiveness of presentation (40 pts total) Effective title? Readability? Organization? Is abstract concise and prominently displayed? Does abstract have all elements & summarize the poster? Do conclusions concisely summarize the major results? Are conclusions supported by data given on the poster? Is introduction concise, yet adequate? Do figure and table titles concisely summarize key results? Are captions adequate & distinct from figure and table titles? Are figures and tables clear and easily understood? Are the various elements effectively arranged? Is discussion concise, yet explain each result? English (spelling, grammar) Overall: did it effectively deliver the key points? 1 pts ______ 2 pts ______ 3 pts ______ 4 pts_______ 1 pt ____________ 3 pts_______ 2 pts_____________ 4 pts_______ 2 pts_______ 3 pts_______ 1 pts_____________ 2 pts_____________ 3 pts_______ 5 pts_______
 / 10 Team / 1 Organization? / 1 Clarity? / 1 Introduction? / 1 Were the results clearly presented? / 1 Were the conclusions clearly presented? / 1 Discussion? / 1 Understanding? / 1 Future Plans? / 1 Time? / 1 Teamwork? / 10 Individual / 4 Contribution to overall effort / 1 Role in presentation / 1 Level of understanding / 1 Role in question/answer / 1 Quality of answers / 6 Mechanics / 1 Confidence / 1 Diction & volume / 1 Interaction with audience / 1 Pace / 1 poise, mannerisms / 1 use of images
/ 10 Team / 1 Organization? / 1 Clarity? / 1 Introduction? / 1 Were the results clearly presented? / 1 Were the conclusions clearly presented? / 1 Discussion? / 1 Understanding? / 1 Future Plans? / 1 Time? / 1 Teamwork? / 10 Individual / 4 Contribution to overall effort / 1 Role in presentation / 1 Level of understanding / 1 Role in question/answer / 1 Quality of answers / 6 Mechanics / 1 Confidence / 1 Diction & volume / 1 Interaction with audience / 1 Pace / 1 poise, mannerisms / 1 use of images
 Formal Report 1. Title Page: should include the title, name of each author, and the institution at which the work was performed. • Text should be centered. DWA 3, an Arabidopsis DWD protein, acts as a negative regulator in ABA signal transduction Jae-Hoon Lee, a William Terzaghi, a, b and Xing Wang Deng a, * a Department of Molecular, Cellular, and Developmental Biology, Yale University, New Haven, Connecticut 06520 -8104, U. S. A. b Department of Biology, Wilkes University, Wilkes-Barre, PA 18766, U. S. A. * Corresponding author, Fax: +1 203 432 3204 E-mail address: xingwang. deng@yale. edu Keywords: CUL 4 -based E 3 ligase; DWD; ABA; negative regulator; signal transduction pathway; Arabidopsis
Formal Report 1. Title Page: should include the title, name of each author, and the institution at which the work was performed. • Text should be centered. DWA 3, an Arabidopsis DWD protein, acts as a negative regulator in ABA signal transduction Jae-Hoon Lee, a William Terzaghi, a, b and Xing Wang Deng a, * a Department of Molecular, Cellular, and Developmental Biology, Yale University, New Haven, Connecticut 06520 -8104, U. S. A. b Department of Biology, Wilkes University, Wilkes-Barre, PA 18766, U. S. A. * Corresponding author, Fax: +1 203 432 3204 E-mail address: xingwang. deng@yale. edu Keywords: CUL 4 -based E 3 ligase; DWD; ABA; negative regulator; signal transduction pathway; Arabidopsis
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
Formal Report 2. Abstract: summarizes the paper in 250 words or less. DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods DWD proteins have been reported as substrate receptors for cullin– RING ubiquitin ligase 4 (CRL 4). Upon screening T-DNA mutants of DWD genes for abscisic acid (ABA) responses we obtained several candidates which exhibited ABA-hypersensitivity and one was named DWA 3 (DWD hypersensitive to ABA 3). DWA 3 associated with the CRL 4 components DDB 1 and CUL 4, indicating that DWA 3 may function as a substrate receptor for CRL 4. ABA-inducible transcription factors (ABI 5 and At. MYC 2) and their downstream genes were hyperinduced by ABA in dwa 3. Taken together, we suggest DWA 3 is a negative regulator of ABA responses and may be involved in protein degradation mediated by CRL 4.
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data • 1 -2 sentences of Discussion
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data • 1 -2 sentences of Discussion
 Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data • 1 -2 sentences of Discussion • rarely cite references. If they do, full citation must be included.
Formal Report 2. Abstract: summarizes the paper in 250 words or less. • mini-papers that are often published separately • should have 1 or 2 sentences of Introduction • 1 - 2 sentences of Materials and Methods • 3 -4 sentences of Results, including quantitative data • 1 -2 sentences of Discussion • rarely cite references. If they do, full citation must be included.
 Formal Report 3. Introduction: explains why the experiment was done.
Formal Report 3. Introduction: explains why the experiment was done.
 Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are
Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are
 Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are • specifically states the hypothesis that was tested.
Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are • specifically states the hypothesis that was tested.
 Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are • specifically states the hypothesis that was tested. • I write it last, to guide readers to my results
Formal Report 3. Introduction: explains why the experiment was done. • provides enough detail about what was previously known to explain what the outstanding questions are • specifically states the hypothesis that was tested. • I write it last, to guide readers to my results
 Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment.
Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment.
 Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc
Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc
 Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Instead, cite the reference from which the recipe was obtained.
Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Instead, cite the reference from which the recipe was obtained.
 Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Do cite the manufacturers of esoteric reagents or equipment
Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Do cite the manufacturers of esoteric reagents or equipment
 Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Do cite the manufacturers of esoteric reagents or equipment • Both Materials and Methods and Results should be written in the past tense. Routine calculations are not described here, unless they are done in an unusual way.
Formal Report 4. Materials and Methods: provide sufficient detail that another biologist could repeat your experiment. • list the organisms used (including the latin binomial) the names of the reagents, procedures followed, etc • do not give the recipe for each reagent. • Do cite the manufacturers of esoteric reagents or equipment • Both Materials and Methods and Results should be written in the past tense. Routine calculations are not described here, unless they are done in an unusual way.
 Formal Report 5. Results: devote one paragraph to each figure or table
Formal Report 5. Results: devote one paragraph to each figure or table
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features.
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features.
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited”
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited”
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited” • each figure or table should have its own title and caption
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited” • each figure or table should have its own title and caption
 Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited” • each figure or table should have its own title and caption • caption gives sufficient information that a reader can figure out what it is about without reading the text. i. e. summarizes M&M & identifies panels, symbols, etc
Formal Report 5. Results: devote one paragraph to each figure or table • start with a sentence explaining the purpose of the expt • second sentence should summarize the methods • third sentence states where the results are presented • remaining sentences point out the key features. • Results can be presented as figures or tables, which should be presented on separate pages attached after the “literature cited” • each figure or table should have its own title and caption • caption gives sufficient information that a reader can figure out what it is about without reading the text. i. e. summarizes M&M & identifies panels, symbols, etc
 Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end
Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end
 Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results
Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results
 Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results • Remaining sentences explain them, and may propose ways to test these explanations
Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results • Remaining sentences explain them, and may propose ways to test these explanations
 Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results • Remaining sentences explain them, and may propose ways to test these explanations • Last paragraph discusses global implications: now that we know this, how does it change our world?
Formal Report 6. Discussion: one paragraph per figure or table + a global discussion at end • First sentence summarizes results • Remaining sentences explain them, and may propose ways to test these explanations • Last paragraph discusses global implications: now that we know this, how does it change our world?
 Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper.
Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper.
 Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals.
Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals.
 Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals. • For your report use the format (Smith, 2008) in the text
Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals. • For your report use the format (Smith, 2008) in the text
 Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals. • For your report use the format (Smith, 2008) in the text • use this format in the “Literature Cited: ” Smith, E. J. (2008) BRF 3 encodes a novel ubiquitin ligase. Molecular Plant 3: 345 -361
Formal Report 7. Literature cited: all citations listed in the text should be listed at the end of the paper. • Formats for citing and listing references vary among journals. • For your report use the format (Smith, 2008) in the text • use this format in the “Literature Cited: ” Smith, E. J. (2008) BRF 3 encodes a novel ubiquitin ligase. Molecular Plant 3: 345 -361
 Bio 392: Manuscript Draft Grading Checklist 1) Abstract (6 pts) Were all elements (I, M&M, R & D) present? Were all elements adequately explained? Was all information relevant? Was all information clearly and succinctly explained? 2) Introduction (6 pts) Was the hypothesis (or purpose) clearly stated? Was adequate background information provided? Was all information relevant? Was all information clearly explained? 3) Materials and Methods (4 pts) Were all procedures clearly and accurately explained? Was all information relevant?
Bio 392: Manuscript Draft Grading Checklist 1) Abstract (6 pts) Were all elements (I, M&M, R & D) present? Were all elements adequately explained? Was all information relevant? Was all information clearly and succinctly explained? 2) Introduction (6 pts) Was the hypothesis (or purpose) clearly stated? Was adequate background information provided? Was all information relevant? Was all information clearly explained? 3) Materials and Methods (4 pts) Were all procedures clearly and accurately explained? Was all information relevant?
 4) Results (10 pts) Is there a separate paragraph for each expt? Does the first sentence of each para state the purpose? Does the second sentence summarize the methods? Does the third sentence state where the results are? Do the remaining sentences explain the figure/table and point out key results? Do figures and tables do the job? Titles and captions? 5) Discussion (8 pts) Were key results summarized? Were key results discussed? Were further experiments proposed? Were broader implications discussed?
4) Results (10 pts) Is there a separate paragraph for each expt? Does the first sentence of each para state the purpose? Does the second sentence summarize the methods? Does the third sentence state where the results are? Do the remaining sentences explain the figure/table and point out key results? Do figures and tables do the job? Titles and captions? 5) Discussion (8 pts) Were key results summarized? Were key results discussed? Were further experiments proposed? Were broader implications discussed?
 6) Literature cited (2 pts) Were references used correctly in the text? Were all citations made in text listed in correct format? Were any references not cited in the text? 7) Writing (4 pts) Organization Clarity Conciseness Grammar and spelling
6) Literature cited (2 pts) Were references used correctly in the text? Were all citations made in text listed in correct format? Were any references not cited in the text? 7) Writing (4 pts) Organization Clarity Conciseness Grammar and spelling


