66bbe799c9a9bfb291d6b0117c7369d0.ppt
- Количество слайдов: 18

Ontology Engineering approaches based on semi-automated curation of the primary literature Gully APC Burns, Tommy Ingulfsen, Donghui Feng and Ed Hovy Biomedical Knowledge Engineering Group, Information Sciences Institute, University of Southern California

Where’s all the knowledge? The primary research literature. . . … is the end-product of all scientific research … forms the basis for human understanding of the subject. . . is written in natural language … is structured … is interpretable Image taken from U. S. Geological Survey Energy Resource Surveys Program … is expensive … is terse
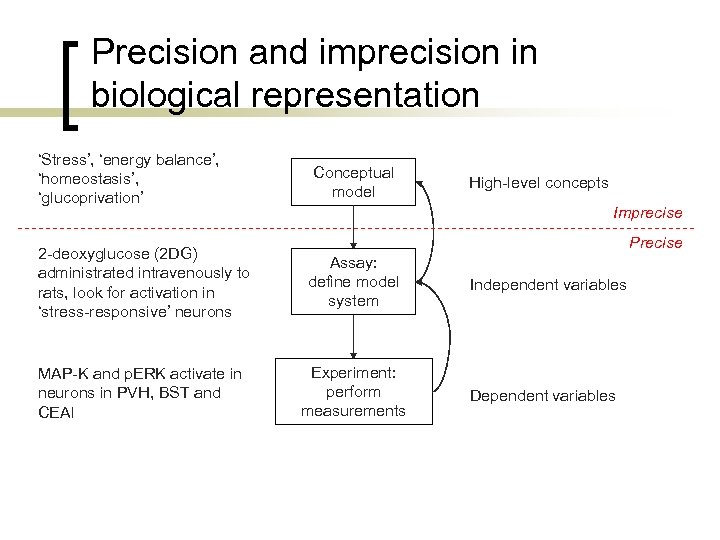
Precision and imprecision in biological representation ‘Stress’, ‘energy balance’, ‘homeostasis’, ‘glucoprivation’ Conceptual model High-level concepts Imprecise Precise 2 -deoxyglucose (2 DG) administrated intravenously to rats, look for activation in ‘stress-responsive’ neurons Assay: define model system MAP-K and p. ERK activate in neurons in PVH, BST and CEAl Experiment: perform measurements Independent variables Dependent variables
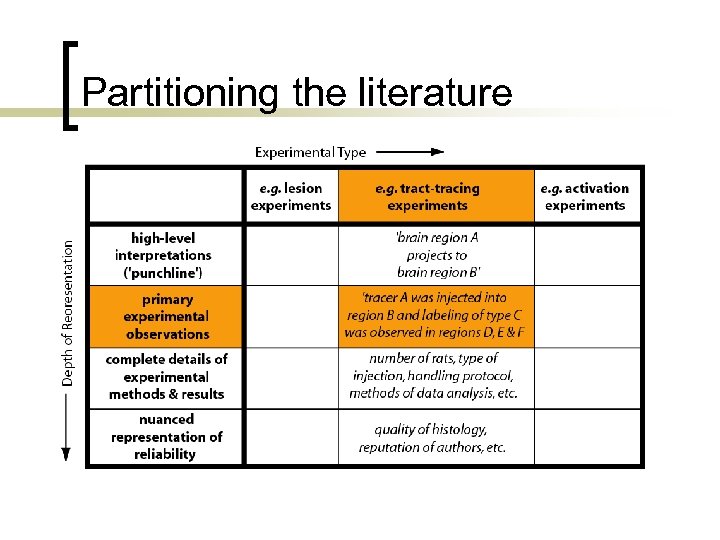
Partitioning the literature
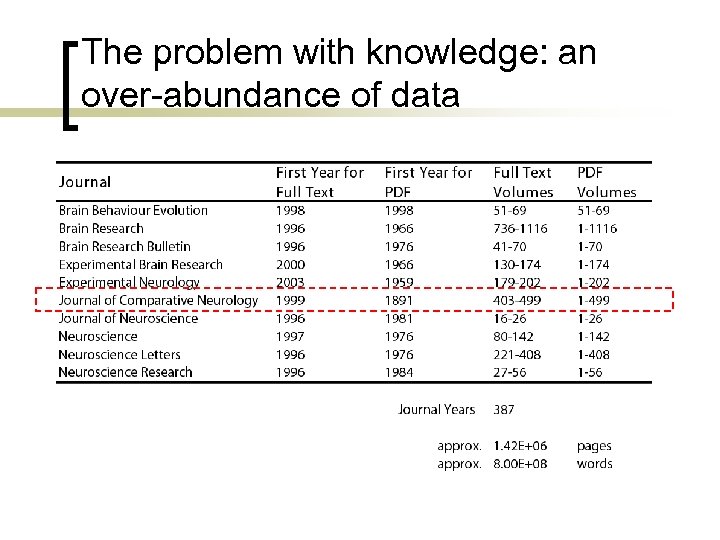
The problem with knowledge: an over-abundance of data
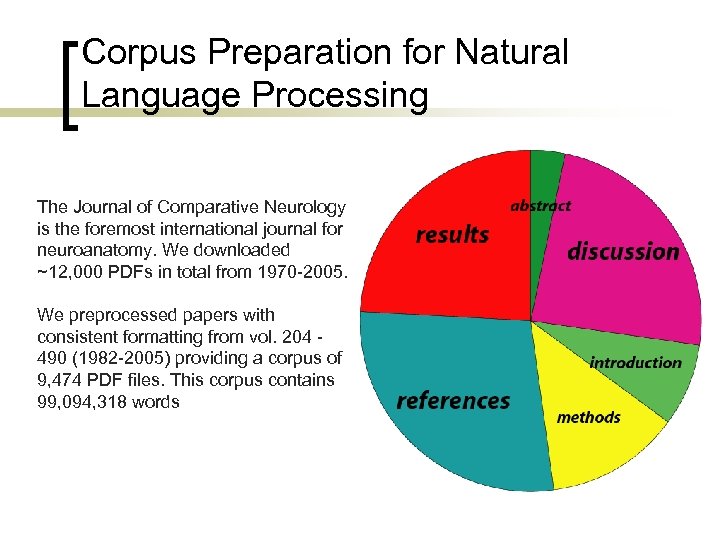

Corpus Preparation for Natural Language Processing The Journal of Comparative Neurology is the foremost international journal for neuroanatomy. We downloaded ~12, 000 PDFs in total from 1970 -2005. We preprocessed papers with consistent formatting from vol. 204 490 (1982 -2005) providing a corpus of 9, 474 PDF files. This corpus contains 99, 094, 318 words
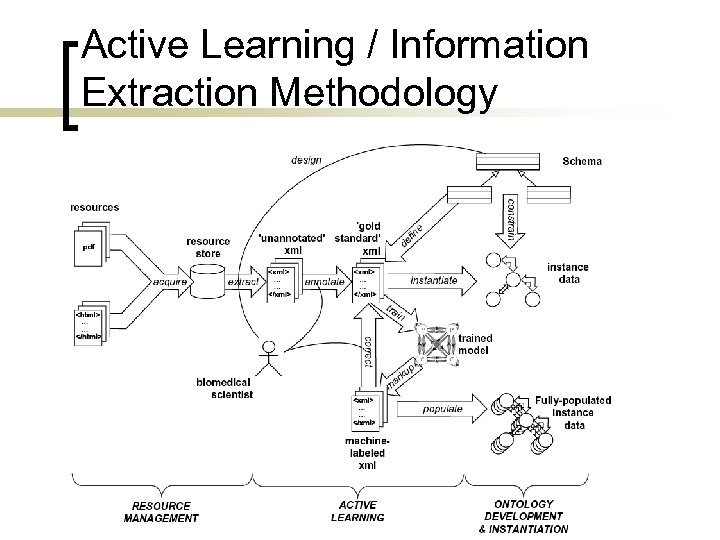
Active Learning / Information Extraction Methodology
![The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site](https://present5.com/presentation/66bbe799c9a9bfb291d6b0117c7369d0/image-8.jpg)
The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site [1] ¡ Location n n brain structure topography side ‘anterograde’ Labeled region [1. . . *] ¡ Location n ¡ ¡ brain structure topography ipsi-contra relative to injection site? Label type Label density ‘retrograde’
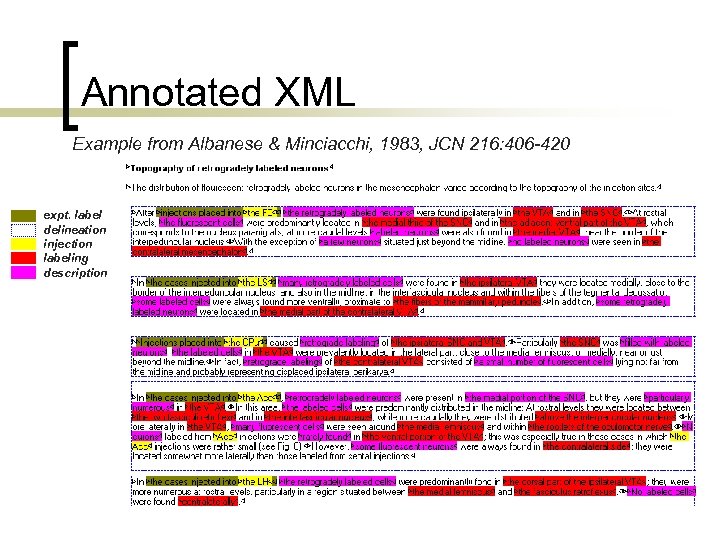

Annotated XML Example from Albanese & Minciacchi, 1983, JCN 216: 406 -420 expt. label delineation injection labeling description
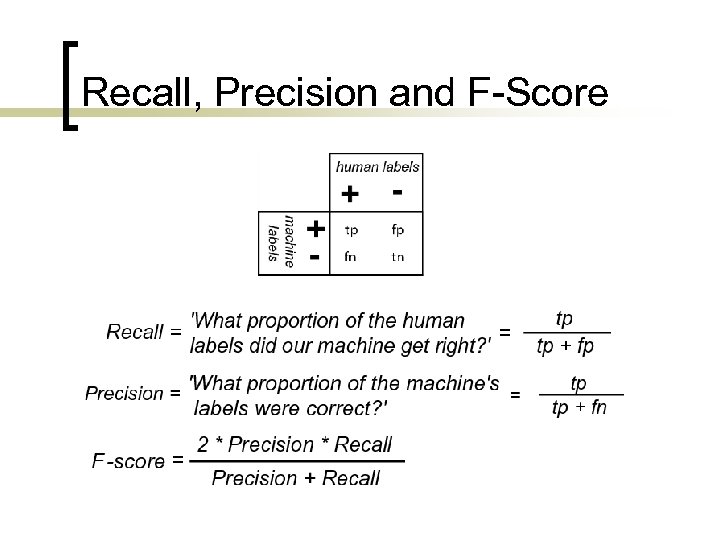

Recall, Precision and F-Score
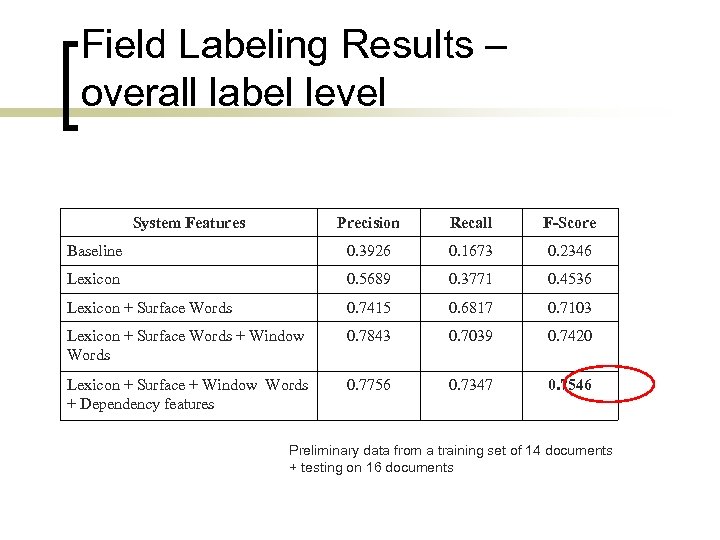
Field Labeling Results – overall label level System Features Precision Recall F-Score Baseline 0. 3926 0. 1673 0. 2346 Lexicon 0. 5689 0. 3771 0. 4536 Lexicon + Surface Words 0. 7415 0. 6817 0. 7103 Lexicon + Surface Words + Window Words 0. 7843 0. 7039 0. 7420 Lexicon + Surface + Window Words + Dependency features 0. 7756 0. 7347 0. 7546 Preliminary data from a training set of 14 documents + testing on 16 documents
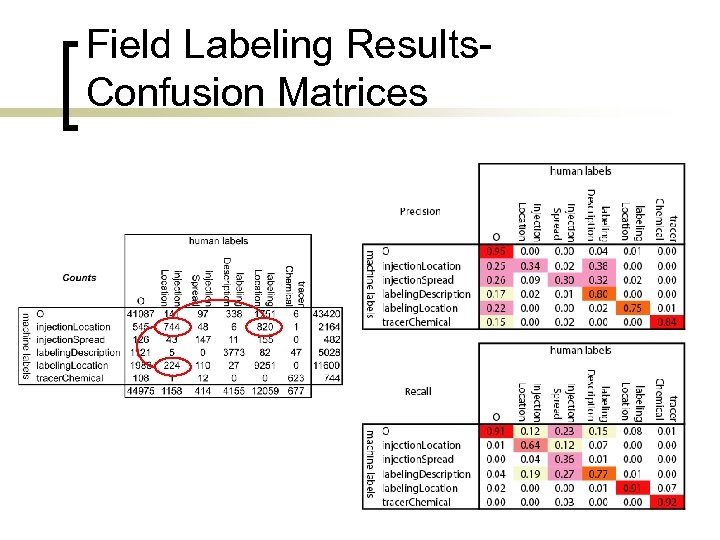

Field Labeling Results. Confusion Matrices
![Generalizing the methodology: ‘Histology’ [from Gonzalo-Ruiz et al 1992, JCN 321: 300 -311] Generalizing the methodology: ‘Histology’ [from Gonzalo-Ruiz et al 1992, JCN 321: 300 -311]](https://present5.com/presentation/66bbe799c9a9bfb291d6b0117c7369d0/image-13.jpg)
Generalizing the methodology: ‘Histology’ [from Gonzalo-Ruiz et al 1992, JCN 321: 300 -311]
![The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site](https://present5.com/presentation/66bbe799c9a9bfb291d6b0117c7369d0/image-14.jpg)
The logical structure of a tracttracing experiment n n Tracer Chemical [1] Injection Site [1] ¡ Location n n brain structure topography side ‘anterograde’ Labeled region [1. . . *] ¡ Location n ¡ ¡ brain structure topography ipsi-contra relative to injection site? Label type Label density ‘retrograde’

Time and effort n n n Current performance achieved by annotating 40 documents Each document contains 97 sentences (in results section) on average Annotation rate ¡ ¡ n ~ 40 Sent/hr (no support) ~115 Sent/hr (after 20 documents) Time taken to annotate document to train system to perform at this standard ¡ ¡ ~65 hours with no support Estimate ~2 months for a 50% RA (20 hours / week)

Can we discover the schema from the text? n n n Given a large review or a grant proposal specific to a single laboratory Annotate independent and dependent variables in papers. Can we learn and extract these patterns?
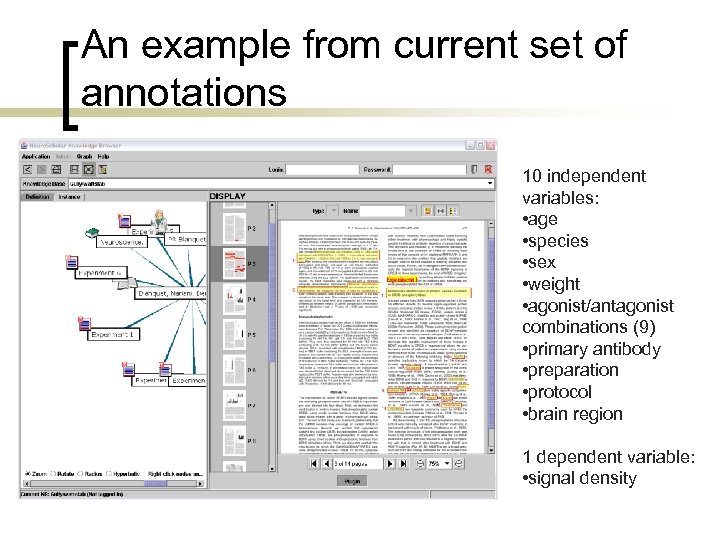

An example from current set of annotations 10 independent variables: • age • species • sex • weight • agonist/antagonist combinations (9) • primary antibody • preparation • protocol • brain region 1 dependent variable: • signal density

Acknowledgements Funding ¡ ¡ Information Sciences Institute, seed funding * National Library of Medicine (RO 1 -LM 07061) * NSF (LONI MAP project) HBP (USCBP) Neuroscience consultants ¡ ¡ ¡ ¡ Alan Watts * Larry Swanson * Arshad Khan * Rick Thompson * Joel Hahn * Lori Gorton * Kim Rapp * Computer Scientists ¡ Eduard Hovy * ¡ Donghui Feng * ¡ Patrick Pantel * Developers ¡ Tommy Ingulfsen * ¡ Wei-Cheng
66bbe799c9a9bfb291d6b0117c7369d0.ppt