31d06aa6aec56ad2eee518f10c7801bf.ppt
- Количество слайдов: 113



Observation * Photos courtesy of www. webshots. com and Peter Smallwood

Observation * Photos courtesy of www. webshots. com and Peter Smallwood

Observation * Photos courtesy of www. webshots. com and Peter Smallwood

Observation * Photos courtesy of www. webshots. com and Peter Smallwood

Experiment * Photos courtesy of www. webshots. com and Peter Smallwood
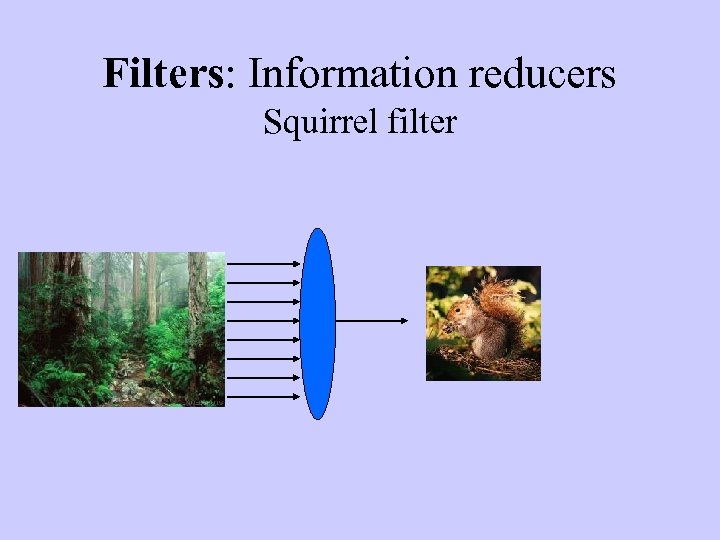

Filters: Information reducers Squirrel filter
Filters: Information reducers Molecule filter
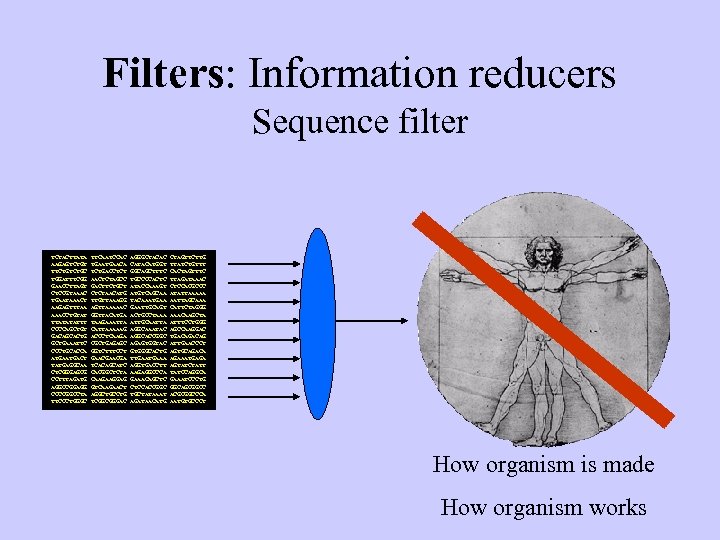
Filters: Information reducers Sequence filter TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TATGAGGCAA CTCGGGAGCG CCTTTAGATG AGGCCGGAGG CCCCGGCCTA TTCCCTGGGC TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA TCACAGCATC CACGGCTCTA CAAGAAGGAG GTCAAGAACT AGGCTGCCTG TCGGCGGGAC AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA AGGTGACCTT AAGAGGCCCA GAAACAGCTC CTCCACCGGC TGCTATAAAT AGATAACATG CTAGTTCTTG TTATCTGTTT CACTAGTTTC TTAGATAAAC CTCCACGCCC ATATTAAAAA AATTAGCAAA CATTCTAGGG AAACAAGCTA ATTTCCTGGG AGCCAAGGAC TGACAG ATTGAACCCT AGTGCAGACA AGAAATGAGA AGTATCTATT TATCCAGGCA GAAATCCCTG GGCAGCGGCC ACGCGGCCCA AATGTGCCCT CTCCGTAAAC CTCTAAC. . . How organism is made How organism works
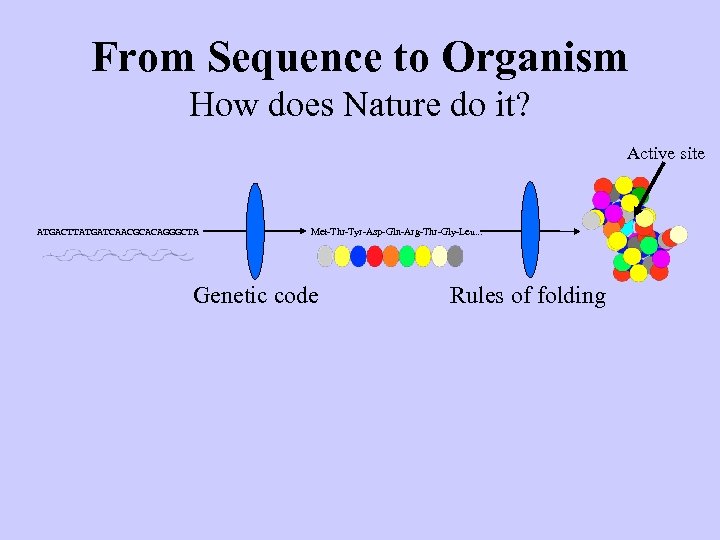
From Sequence to Organism How does Nature do it? Active site ATGACTTATGATCAACGCACAGGGCTA Met-Thr-Tyr-Asp-Gln-Arg-Thr-Gly-Leu. . . Genetic code Rules of folding
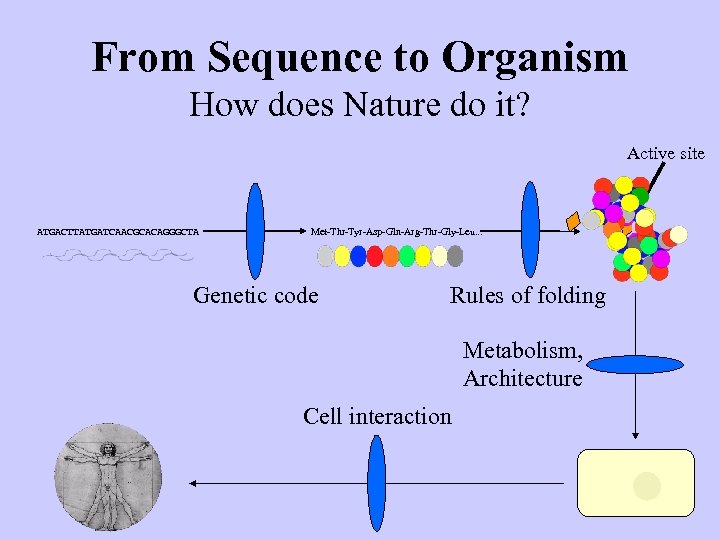
From Sequence to Organism How does Nature do it? Active site ATGACTTATGATCAACGCACAGGGCTA Met-Thr-Tyr-Asp-Gln-Arg-Thr-Gly-Leu. . . Genetic code Rules of folding Metabolism, Architecture Cell interaction
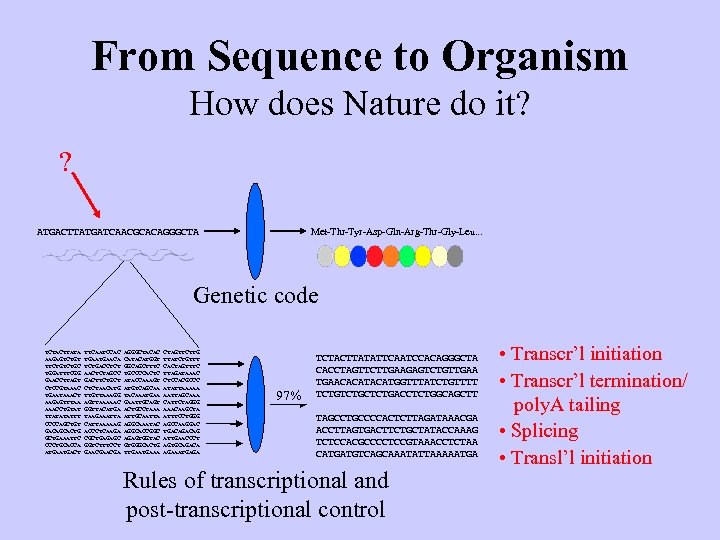
From Sequence to Organism How does Nature do it? ? Met-Thr-Tyr-Asp-Gln-Arg-Thr-Gly-Leu. . . ATGACTTATGATCAACGCACAGGGCTA Genetic code TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA CTAGTTCTTG TTATCTGTTT CACTAGTTTC TTAGATAAAC CTCCACGCCC ATATTAAAAA AATTAGCAAA CATTCTAGGG AAACAAGCTA ATTTCCTGGG AGCCAAGGAC TGACAG ATTGAACCCT AGTGCAGACA AGAAATGAGA 3% 97% TCTACTTATATTCAATCCACAGGGCTA CACCTAGTTCTTGAAGAGTCTGTTGAACACATGGTTTATCTGTTTT TCTGCTCTGACCTCTGGCAGCTT ATGACTTATGATCAACGCACAGGGCTA TAGCCTGCCCCACTCTTAGATAAACGA ACCTTAGTGACTTCTGCTATACCAAAG TCTCCACGCCCCTCCGTAAACCTCTAA CATGATGTCAGCAAATATTAAAAATGA Rules of transcriptional and post-transcriptional control • Transcr’l initiation • Transcr’l termination/ poly. A tailing • Splicing • Transl’l initiation
From Sequence to Organism How does Nature do it? Natural filters/transformations • Selective transcription DNA • Selective processing • Translation • Folding Functional protein
From Sequence to Organism How does Nature do it? Natural filters/transformations DNA Functional protein
From Sequence to Organism How can WE do it? Simulation of Nature “Whether ‘tis nobler in the mind to suffer the slings and arrows of outrageous fortune. . . ” Utterence of Wm Shakespeare Utterence of George W Bush “We must give our military every tool and weapon it needs to prevail. . . ” ? ? ?
From Sequence to Organism How can WE do it? Surrogate Processes “Whether ‘tis nobler in the mind to suffer the slings and arrows of outrageous fortune. . . ” “We must give our military every tool and weapon it needs to prevail. . . ” Utterence of Wm Shakespeare Utterence of George W Bush Words/sentence; Choice of words; Sentence structure; …
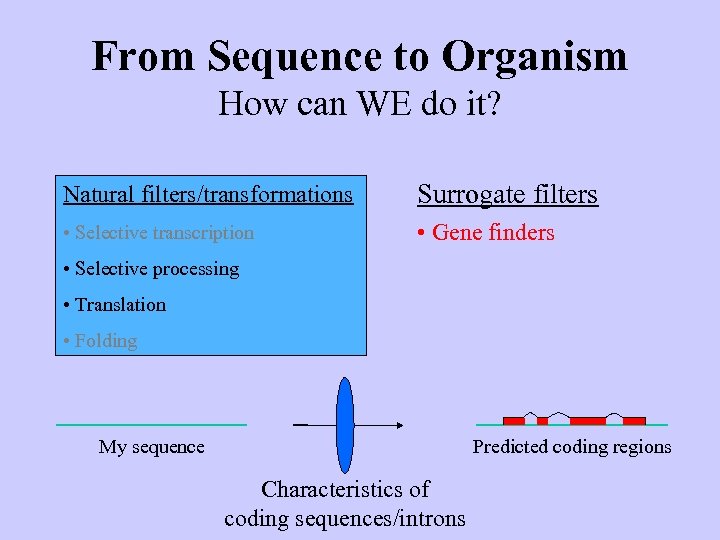
From Sequence to Organism How can WE do it? Natural filters/transformations Surrogate filters • Selective transcription • Gene finders • Selective processing • Translation • Folding Predicted coding regions My sequence Characteristics of coding sequences/introns
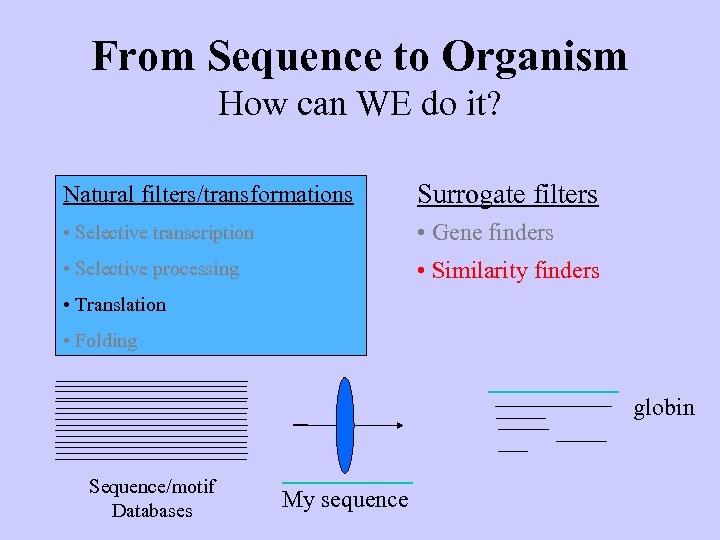
From Sequence to Organism How can WE do it? Natural filters/transformations Surrogate filters • Selective transcription • Gene finders • Selective processing • Similarity finders • Translation • Folding globin Sequence/motif Databases My sequence
From Sequence to Organism How can WE do it? Natural filters/transformations Surrogate filters • Selective transcription • Gene finders • Selective processing • Similarity finders • Translation • Feature finders • Folding Predicted features My sequence Characteristics of features
From Sequence to Organism How can WE do it? Natural filters/transformations Surrogate filters • Selective transcription • Gene finders • Selective processing • Similarity finders • Translation • Feature finders • Folding • Pattern finders My sequences Statistical engine

From Sequence to Organism How can WE do it? Natural filters/transformations Surrogate filters • Selective transcription • Gene finders • Selective processing • Similarity finders • Translation • Feature finders • Folding • Pattern finders 2 nd Most powerful tool

Surrogate Filters How do they work? Case studies • Gene finders • Real problems • Similarity finders • Mixed strategies • Feature finders • Pattern finders You do it
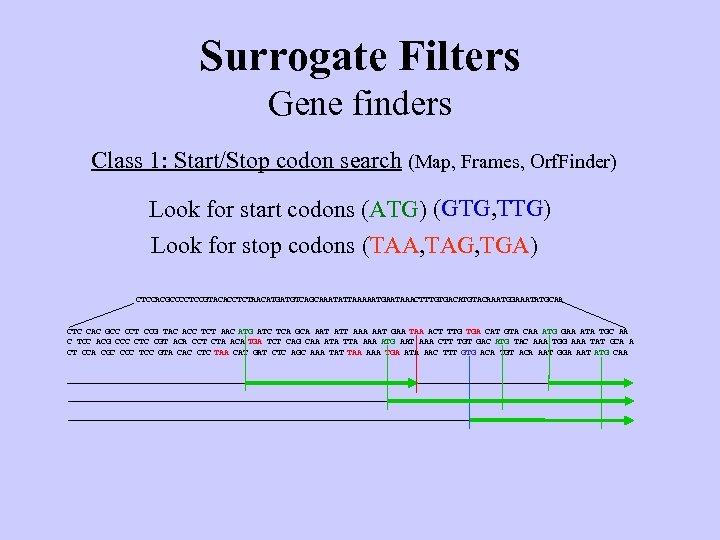

Surrogate Filters Gene finders Class 1: Start/Stop codon search (Map, Frames, Orf. Finder) Look for start codons (ATG) (GTG, TTG) Look for stop codons (TAA, TAG, TGA) CTCCACGCCCCTCCGTACACCTCTAACATGATGTCAGCAAATATTAAAAATGAATAAACTTTGTGACATGTACAAATGGAAATATGCAA CTC CAC GCC CCT CCG TAC ACC TCT AAC ATG ATC TCA GCA AAT ATT AAA AAT GAA TAA ACT TTG TGA CAT GTA CAA ATG GAA ATA TGC AA C TCC ACG CCC CTC CGT ACA CCT CTA ACA TGA TCT CAG CAA ATA TTA AAA ATG AAT AAA CTT TGT GAC ATG TAC AAA TGG AAA TAT GCA A CT CCA CGC CCC TCC GTA CAC CTC TAA CAT GAT CTC AGC AAA TAT TAA AAA TGA ATA AAC TTT GTG ACA TGT ACA AAT GGA AAT ATG CAA
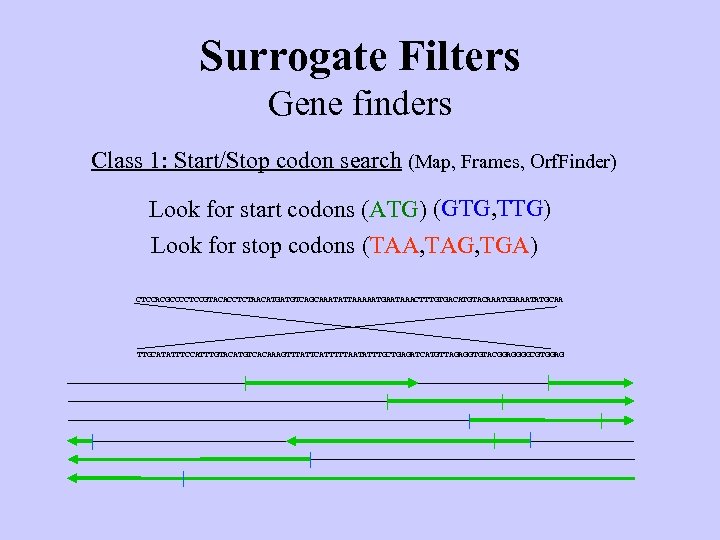

Surrogate Filters Gene finders Class 1: Start/Stop codon search (Map, Frames, Orf. Finder) Look for start codons (ATG) (GTG, TTG) Look for stop codons (TAA, TAG, TGA) CTCCACGCCCCTCCGTACACCTCTAACATGATGTCAGCAAATATTAAAAATGAATAAACTTTGTGACATGTACAAATGGAAATATGCAA TTGCATATTTCCATTTGTACATGTCACAAAGTTTATTCATTTTTAATATTTGCTGAGATCATGTTAGAGGTGTACGGAGGGGCGTGGAG

Surrogate Filters Gene finders Class 1: Start/Stop codon search (Map, Frames, Orf. Finder) Pro: Quick, simple Con: Useless for eukaryotic genomic sequences (introns) Inaccurate (start codon problem) Inaccurate (doubtful short open reading frames)

Surrogate Filters Gene finders Do it Class 1: Start/Stop codon search (Map, Frames, Orf. Finder) 1. Go to http: //www. vcu. edu/~elhaij/Bio. Inf 2. Open 2 nd & 3 rd browsers (Ctrl-N in Netscape) Go to same site (copy and paste URL) 3. In 1 st browser, go to Program List Click on Gene Finders then scroll down Open Orf. Finder 4. In 2 nd browser, open sample sequence

Surrogate Filters Gene finders Do it Class 1: Start/Stop codon search (Map, Frames, Orf. Finder) 5. Paste sample sequence into window 6. Choose Bacterial Code in Genetic codes window 7. Click on Orf. Find
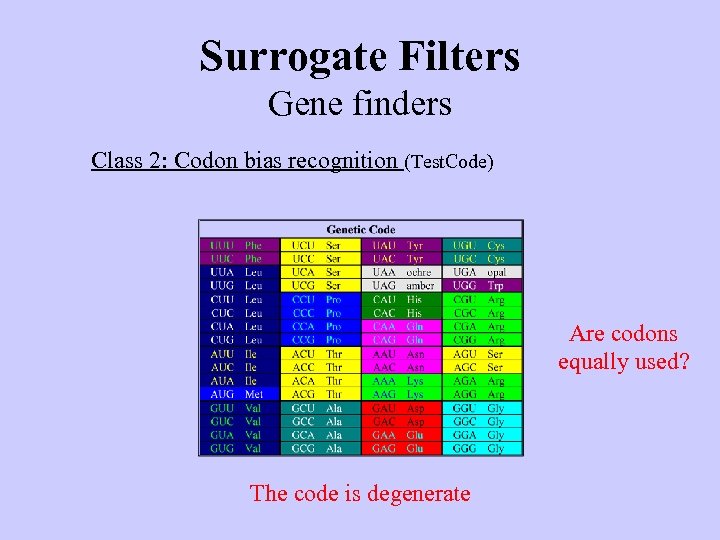


Surrogate Filters Gene finders Class 2: Codon bias recognition (Test. Code) Are codons equally used? The code is degenerate
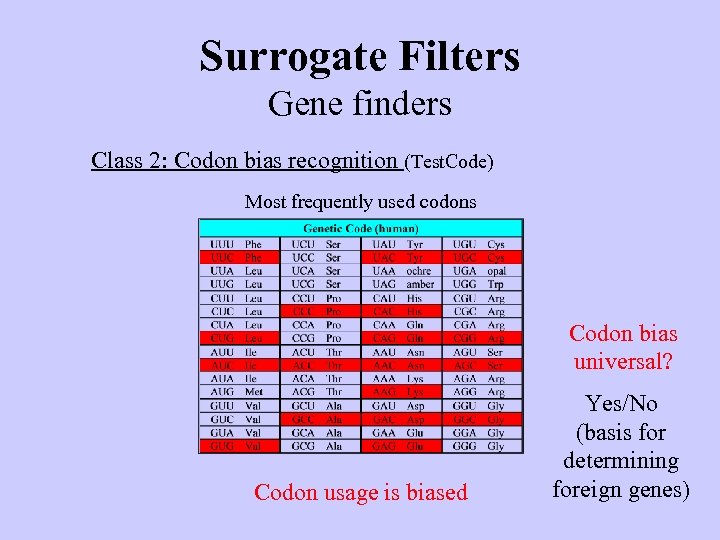
Surrogate Filters Gene finders Class 2: Codon bias recognition (Test. Code) Most frequently used codons Codon bias universal? Codon usage is biased Yes/No (basis for determining foreign genes)

Surrogate Filters Gene finders Class 2: Codon bias recognition (Test. Code) Pro: Quick, simple, available through GCG Better than Class 1 in excluding false open reading frames Con: Useless for eukaryotic genomic sequences (introns) Gives only general areas of open reading frames

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Principle Step 1: Create model through extensive training set * Training set = proven or suspected genes * Organism-specific Step 2: Assess candidate genes through filter of model

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 1: Create model through extensive training set Training Set AAGCTTGACCAAAAAGTTAAAACACTGACGGCAAATAATC AATGACTATCAGAGAATCATCGTGCTGTCAGTAAAA CCTCTGATTTCGATCTTTACCATAATTGTTATGTTGTAAT GACTAACCAGACTATCTTTTACAGAGCTTCTGGTTAACAC TTGTCTAATTAGACATTGATAATGTTTGTGGGGGTTGGTC ATCAGGAATGGTAAATAGCAATTACCCTTCAGACTTTCCT ATGAGACGCTCCGCCAACGAGCAGTGTCTCTTAAAGAACG TTATGAGCGCTCAGTTAACTTCAGAAATTCACGGCGGAAA TCCATAGTTATTATTACTTATGACTAAAACAAAATTACTA TGGCGGCTTGTTTAATATAGATTCTGTGTTCTGAGAAATG ACTTTTAAAGTCCCACTAACTTTTTTCTCATCTATTGCTA TATTTCGACTTTAAAACTTATAGTAGATGGCTTAATTCTC AAATAACAAACTCATTTTTAGTAGATATTTCATGCAAACT GAGGTTTTTAGTGATATTTTCCCCTTATTGAGTACAGCCA CTCCACAAACCTTAGAATGGCTACTCAATATTGCAATTGA TCATGAATATCCCACTGGTAGAGCAGTTTTAATGGAAGAT GCCTGGGGTAATGCAGTTTATTTCGTTGTATCTGGATGGG TAAAAGTTCGGCGCACCTGTGGAGATGATTCGGTAGCTTT AAA AAC AAG AAT ACA. . . TTG TTT
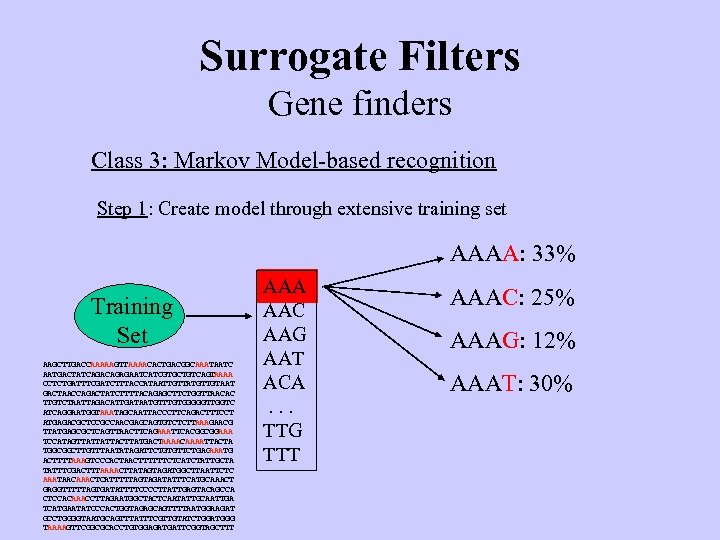

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 1: Create model through extensive training set AAAA: 33% Training Set AAGCTTGACCAAAAAGTTAAAACACTGACGGCAAATAATC AATGACTATCAGAGAATCATCGTGCTGTCAGT AAAA CCTCTGATTTCGATCTTTACCATAATTGTTATGTTGTAAT GACTAACCAGACTATCTTTTACAGAGCTTCTGGTTAACAC TTGTCTAATTAGACATTGATAATGTTTGTGGGGGTTGGTC ATCAGGAATGGTAAATAGCAATTACCCTTCAGACTTTCCT ATGAGACGCTCCGCCAACGAGCAGTGTCTCTT AAAGAACG TTATGAGCGCTCAGTTAACTTCAGAAATTCACGGCGGAAA TCCATAGTTATTATTACTTATGACTAAAACAAAATTACTA TGGCGGCTTGTTTAATATAGATTCTGTGTTCTGAG AAATG ACTTTTAAAGTCCCACTAACTTTTTTCTCATCTATTGCTA TATTTCGACTTTAAAACTTATAGTAGATGGCTTAATTCTC AAATAACAAACTCATTTTTAGTAGATATTTCATGCAAACT GAGGTTTTTAGTGATATTTTCCCCTTATTGAGTACAGCCA CTCCACAAACCTTAGAATGGCTACTCAATATTGCAATTGA TCATGAATATCCCACTGGTAGAGCAGTTTTAATGGAAGAT GCCTGGGGTAATGCAGTTTATTTCGTTGTATCTGGATGGG TAAAAGTTCGGCGCACCTGTGGAGATGATTCGGTAGCTTT AAA AAC AAG AAT ACA. . . TTG TTT AAAC: 25% AAAG: 12% AAAT: 30%
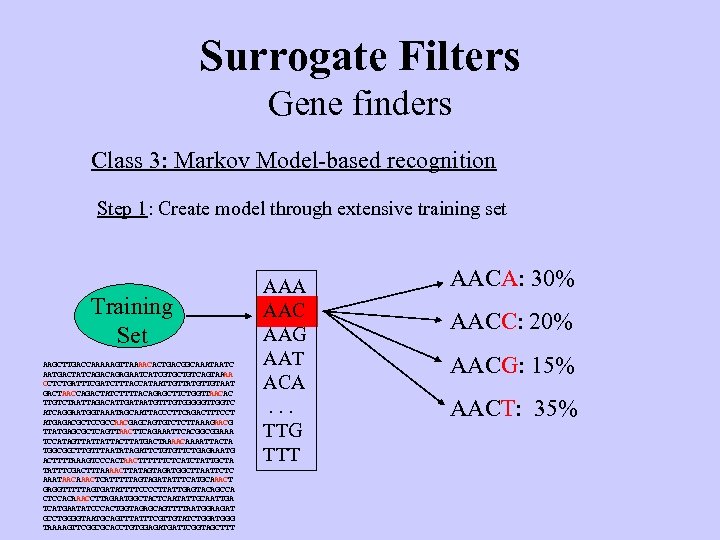

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 1: Create model through extensive training set Training Set AAGCTTGACCAAAAAGTTAAAACACTGACGGCAAATAATC AATGACTATCAGAGAATCATCGTGCTGTCAGTAA AA CCTCTGATTTCGATCTTTACCATAATTGTTATGTTGTAAT GACTAACCAGACTATCTTTTACAGAGCTTCTGGTTAACAC TTGTCTAATTAGACATTGATAATGTTTGTGGGGGTTGGTC ATCAGGAATGGTAAATAGCAATTACCCTTCAGACTTTCCT ATGAGACGCTCCGCCAACGAGCAGTGTCTCTTAAAGAACG TTATGAGCGCTCAGTTAACTTCAGAAATTCACGGCGGAAA TCCATAGTTATTATTACTTATGACTAAAACAAAATTACTA TGGCGGCTTGTTTAATATAGATTCTGTGTTCTGAGAAATG ACTTTTAAAGTCCCACTAACTTTTTTCTCATCTATTGCTA TATTTCGACTTTAAAACTTATAGTAGATGGCTTAATTCTC AAATAACAAACTCATTTTTAGTAGATATTTCATGCAAACT GAGGTTTTTAGTGATATTTTCCCCTTATTGAGTACAGCCA CTCCACAAACCTTAGAATGGCTACTCAATATTGCAATTGA TCATGAATATCCCACTGGTAGAGCAGTTTTAATGGAAGAT GCCTGGGGTAATGCAGTTTATTTCGTTGTATCTGGATGGG TAAAAGTTCGGCGCACCTGTGGAGATGATTCGGTAGCTTT AAA AAC AAG AAT ACA. . . TTG TTT AACA: 30% AACC: 20% AACG: 15% AACT: 35%
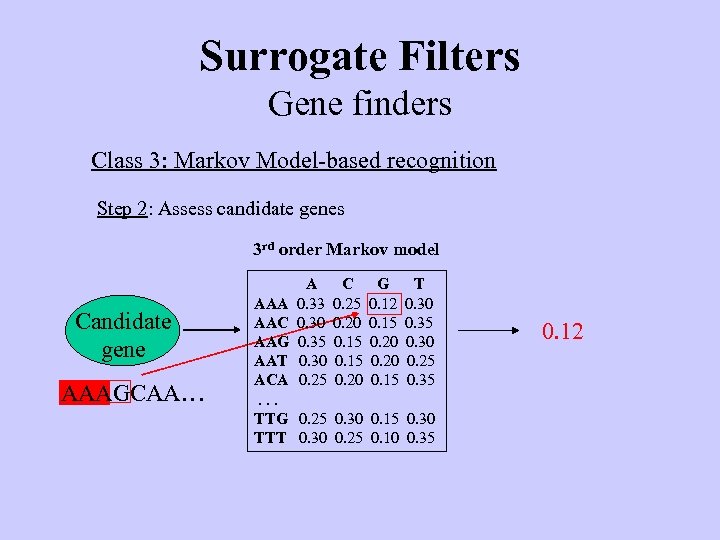

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 2: Assess candidate genes 3 rd order Markov model Candidate gene AAAGCAA… A 0. 33 0. 30 0. 35 0. 30 0. 25 C 0. 25 0. 20 0. 15 0. 20 G 0. 12 0. 15 0. 20 0. 15 T 0. 30 0. 35 0. 30 0. 25 0. 35 AAA AAC AAG AAT ACA. . . TTG 0. 25 0. 30 0. 15 0. 30 TTT 0. 30 0. 25 0. 10 0. 35 0. 12
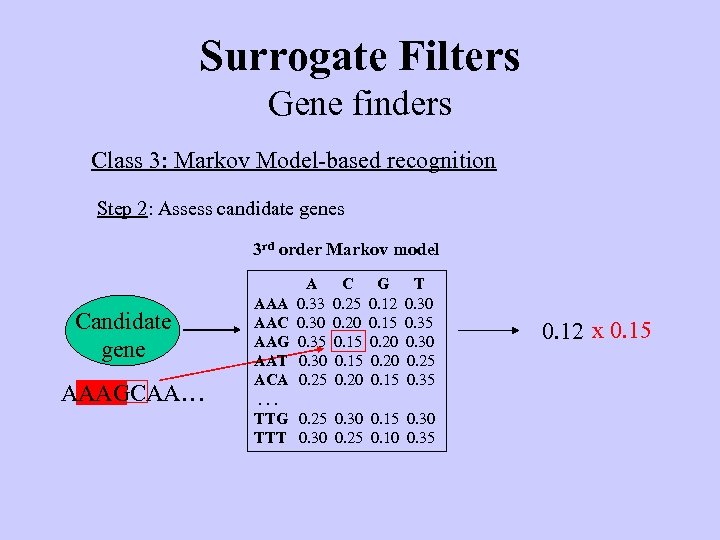

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 2: Assess candidate genes 3 rd order Markov model Candidate gene AAAGCAA… A 0. 33 0. 30 0. 35 0. 30 0. 25 C 0. 25 0. 20 0. 15 0. 20 G 0. 12 0. 15 0. 20 0. 15 T 0. 30 0. 35 0. 30 0. 25 0. 35 AAA AAC AAG AAT ACA. . . TTG 0. 25 0. 30 0. 15 0. 30 TTT 0. 30 0. 25 0. 10 0. 35 0. 12 x 0. 15
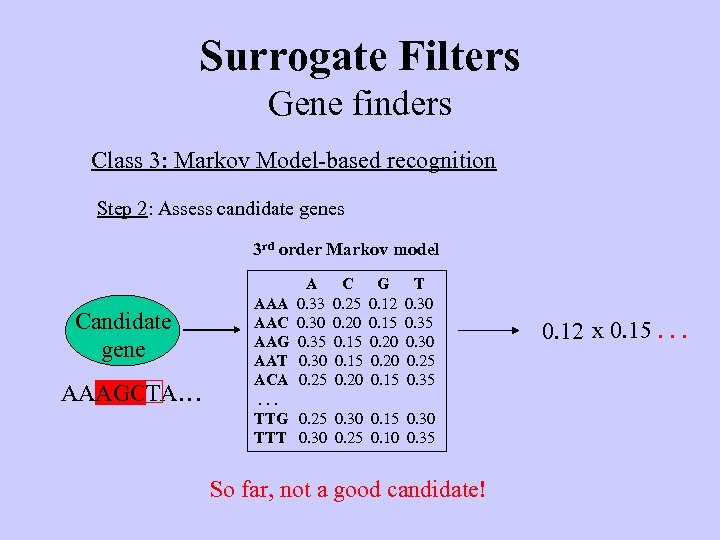

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 2: Assess candidate genes 3 rd order Markov model Candidate gene AAAGCTA… A 0. 33 0. 30 0. 35 0. 30 0. 25 C 0. 25 0. 20 0. 15 0. 20 G 0. 12 0. 15 0. 20 0. 15 T 0. 30 0. 35 0. 30 0. 25 0. 35 AAA AAC AAG AAT ACA. . . TTG 0. 25 0. 30 0. 15 0. 30 TTT 0. 30 0. 25 0. 10 0. 35 So far, not a good candidate! 0. 12 x 0. 15. . .
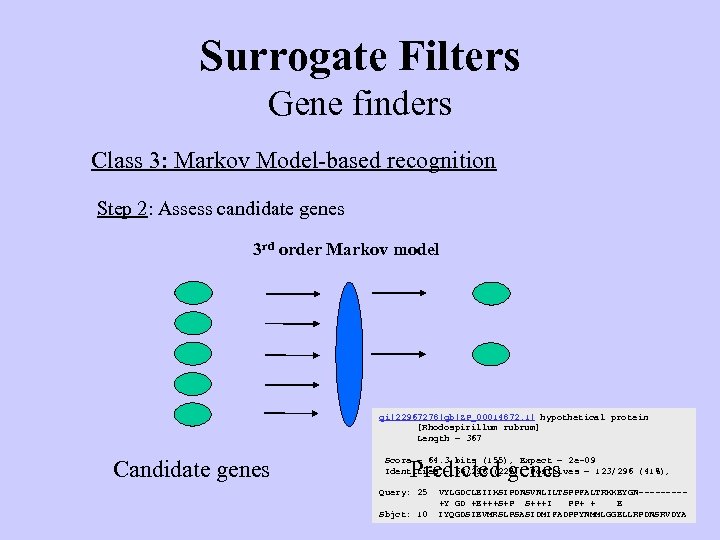

Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 2: Assess candidate genes 3 rd order Markov model gi|22967278|gb|ZP_00014872. 1| hypothetical protein [Rhodospirillum rubrum] Length = 367 Candidate genes Score = 64. 3 bits (155), Expect = 2 e-09 Identities = 68/296 (22%), Positives = 123/296 (41%), Predicted genes Query: 25 Sbjct: 10 VYLGDCLEIIKSIPDNSVNLILTSPPFALTRKKEYGN----+Y GD +E+++S+P S+++I PP+ + E IYQGDSIEVMRSLPSASIDMIFADPPYNMMLGGELLRPDNSRVDYA
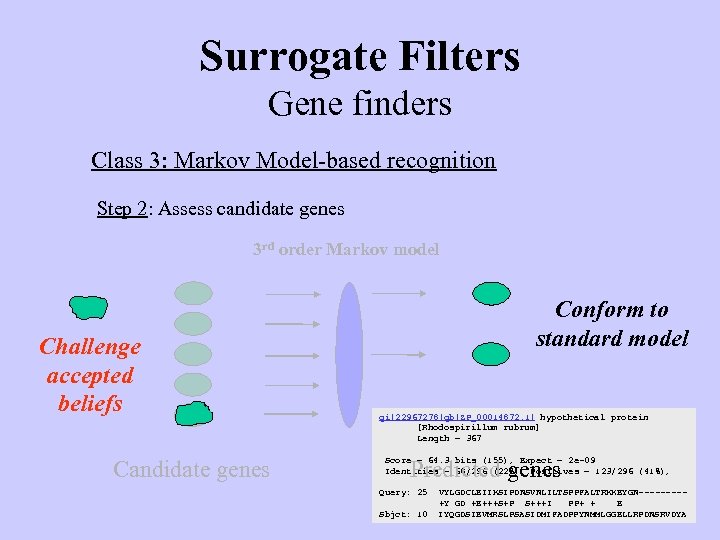
Surrogate Filters Gene finders Class 3: Markov Model-based recognition Step 2: Assess candidate genes 3 rd order Markov model Challenge accepted beliefs Candidate genes Conform to standard model gi|22967278|gb|ZP_00014872. 1| hypothetical protein [Rhodospirillum rubrum] Length = 367 Score = 64. 3 bits (155), Expect = 2 e-09 Identities = 68/296 (22%), Positives = 123/296 (41%), Predicted genes Query: 25 Sbjct: 10 VYLGDCLEIIKSIPDNSVNLILTSPPFALTRKKEYGN----+Y GD +E+++S+P S+++I PP+ + E IYQGDSIEVMRSLPSASIDMIFADPPYNMMLGGELLRPDNSRVDYA

Surrogate Filters Gene finders Class 3: Hidden Markov Model (HMM)-based recognition Pro: Almost accurate method known Con: Needs big training set May miss genes of foreign origin Will miss very small genes

Surrogate Filters Gene finders Do it Class 3: Hidden Markov Model (HMM)-based recognition 1. Go to course web page (3 rd browser) 2. Go to Program List Click on Gene Finders then Gene. Mark 3. Click on “here” in Gene Prediction in Bacteria and Archaea 4. Paste in sample sequence

Surrogate Filters Gene finders Do it Class 3: Hidden Markov Model (HMM)-based recognition 5. Choose Nostoc PCC 7120 as species 6. Check: Generate PDF graphics (screen) Print Gene. Mark 2. 4 predictions 7. Click Start Gene. Mark. hmm


Surrogate Filters Scenario I – Case of the Hidden Heterocyst
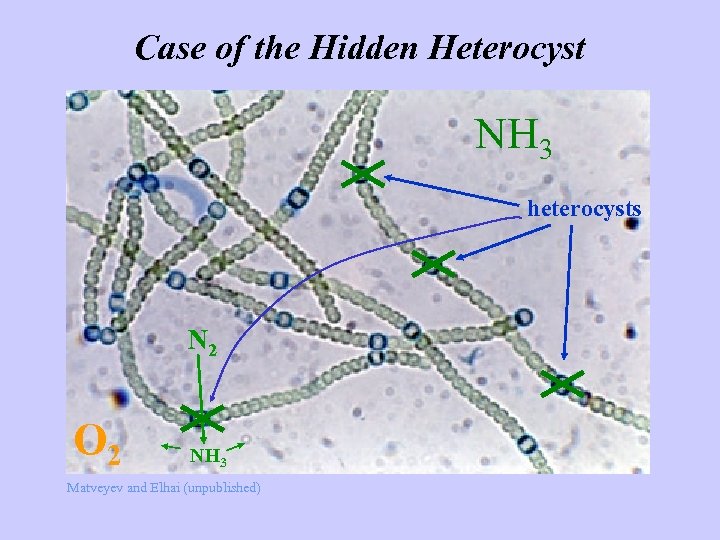
Case of the Hidden Heterocyst NH 3 heterocysts N 2 O 2 NH 3 Matveyev and Elhai (unpublished)
Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Nostoc genome Transposon 1. Use transposon mutagenesis
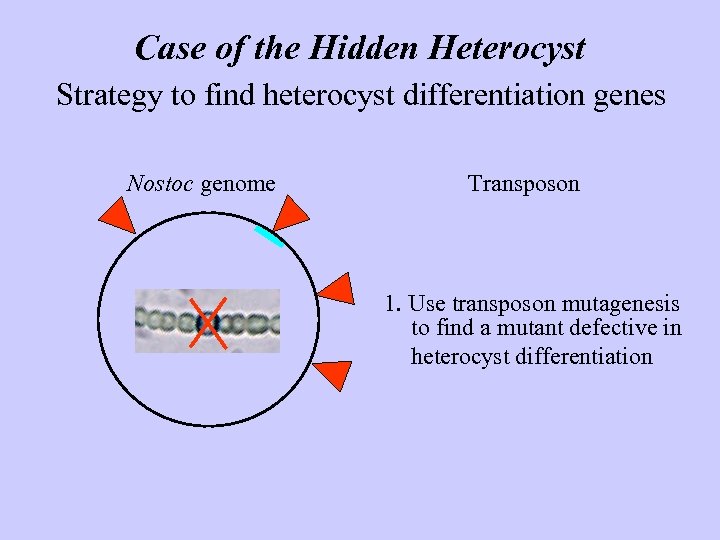
Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Nostoc genome Transposon 1. Use transposon mutagenesis to find a mutant defective in heterocyst differentiation
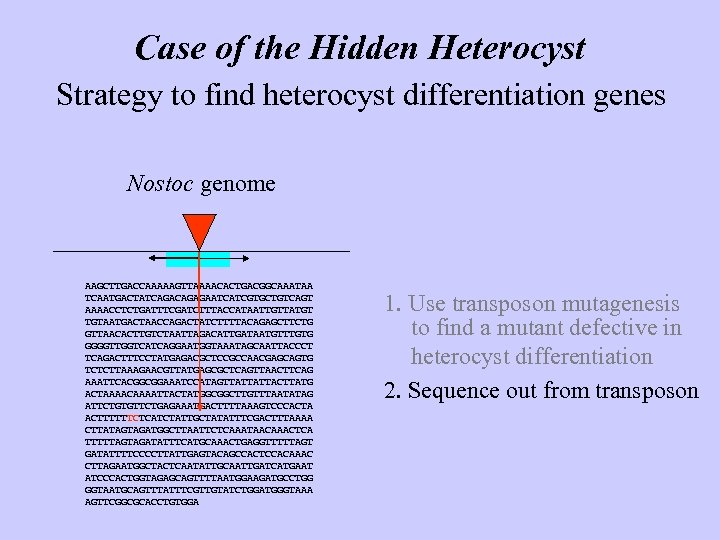

Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Nostoc genome AAGCTTGACCAAAAAGTTAAAACACTGACGGCAAATAA TCAATGACTATCAGAGAATCATCGTGCTGTCAGT AAAACCTCTGATTTCGATCTTTACCATAATTGTTATGT TGTAATGACTAACCAGACTATCTTTTACAGAGCTTCTG GTTAACACTTGTCTAATTAGACATTGATAATGTTTGTG GGGGTTGGTCATCAGGAATGGTAAATAGCAATTACCCT TCAGACTTTCCTATGAGACGCTCCGCCAACGAGCAGTG TCTCTTAAAGAACGTTATGAGCGCTCAGTTAACTTCAG AAATTCACGGCGGAAATCCATAGTTATTATTACTTATG ACTAAAACAAAATTACTATGGCGGCTTGTTTAATATAG ATTCTGTGTTCTGAGAAATGACTTTTAAAGTCCCACTA ACTTTTTTCTCATCTATTGCTATATTTCGACTTTAAAA CTTATAGTAGATGGCTTAATTCTCAAATAACAAACTCA TTTTTAGTAGATATTTCATGCAAACTGAGGTTTTTAGT GATATTTTCCCCTTATTGAGTACAGCCACTCCACAAAC CTTAGAATGGCTACTCAATATTGCAATTGATCATGAAT ATCCCACTGGTAGAGCAGTTTTAATGGAAGATGCCTGG GGTAATGCAGTTTATTTCGTTGTATCTGGATGGGTAAA AGTTCGGCGCACCTGTGGA 1. Use transposon mutagenesis to find a mutant defective in heterocyst differentiation 2. Sequence out from transposon
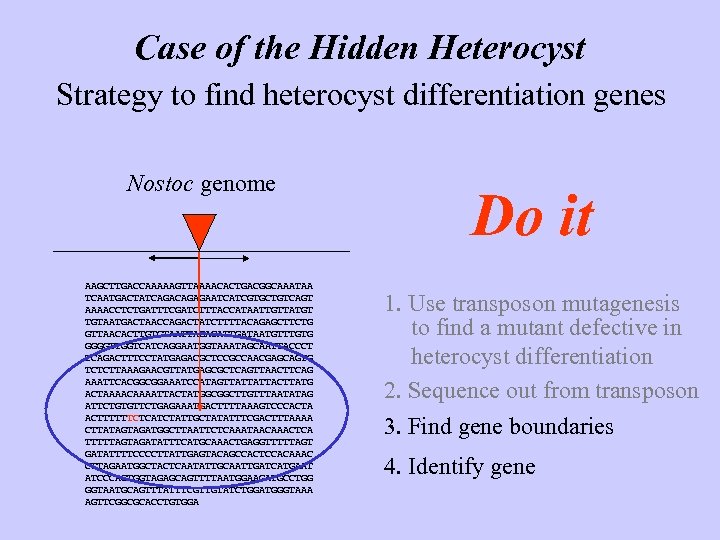
Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Nostoc genome AAGCTTGACCAAAAAGTTAAAACACTGACGGCAAATAA TCAATGACTATCAGAGAATCATCGTGCTGTCAGT AAAACCTCTGATTTCGATCTTTACCATAATTGTTATGT TGTAATGACTAACCAGACTATCTTTTACAGAGCTTCTG GTTAACACTTGTCTAATTAGACATTGATAATGTTTGTG GGGGTTGGTCATCAGGAATGGTAAATAGCAATTACCCT TCAGACTTTCCTATGAGACGCTCCGCCAACGAGCAGTG TCTCTTAAAGAACGTTATGAGCGCTCAGTTAACTTCAG AAATTCACGGCGGAAATCCATAGTTATTATTACTTATG ACTAAAACAAAATTACTATGGCGGCTTGTTTAATATAG ATTCTGTGTTCTGAGAAATGACTTTTAAAGTCCCACTA ACTTTTTTCTCATCTATTGCTATATTTCGACTTTAAAA CTTATAGTAGATGGCTTAATTCTCAAATAACAAACTCA TTTTTAGTAGATATTTCATGCAAACTGAGGTTTTTAGT GATATTTTCCCCTTATTGAGTACAGCCACTCCACAAAC CTTAGAATGGCTACTCAATATTGCAATTGATCATGAAT ATCCCACTGGTAGAGCAGTTTTAATGGAAGATGCCTGG GGTAATGCAGTTTATTTCGTTGTATCTGGATGGGTAAA AGTTCGGCGCACCTGTGGA Do it 1. Use transposon mutagenesis to find a mutant defective in heterocyst differentiation 2. Sequence out from transposon 3. Find gene boundaries 4. Identify gene

Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes 1. Go to course web page (http: //www. vcu. edu/~elhaij/Bio. Inf) 2. Open Nostoc sequence 3. Do what you need to do to find the gene


Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Mission successful: >Translation: 358. . 513 (direct), 51 amino acids VQLAKQAQTAEGTLQIVTNARVTQTVKLVRLEKFLSLQKSVEEALENVK* … or was it? Check predicted protein against databases

Surrogate Filters Similarity finders Do it Blast • Blast. P: Protein sequence to search protein database • Blast. N: Nucleotide sequence to search nucleotide database • Blast. X: Nucleotide sequence (translated) to search protein database • TBlast. N: Protein sequence to search (translated) nucleotide database • Blast 2 Seq: Compare two sequences you specify Fast. A • (Various flavors) Pfam (Protein motif families) Finds conserved motifs similar to protein sequence

*
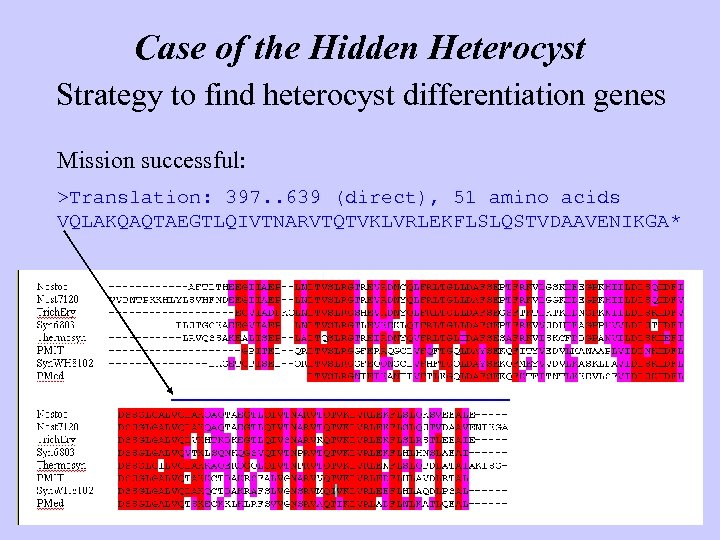
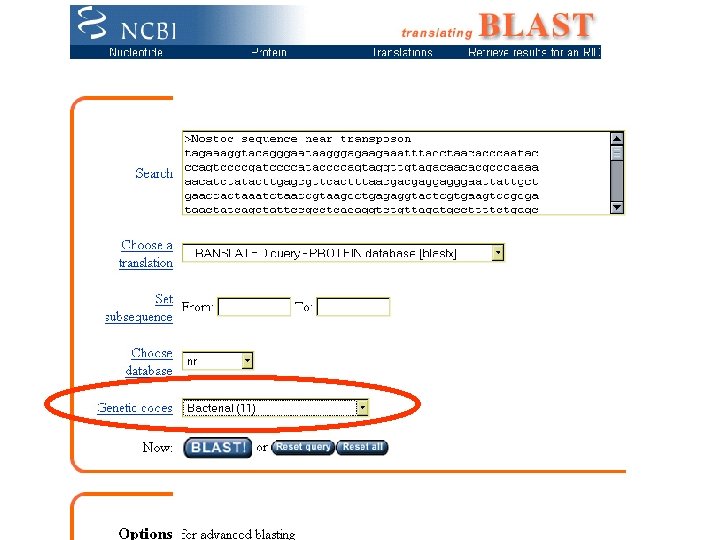

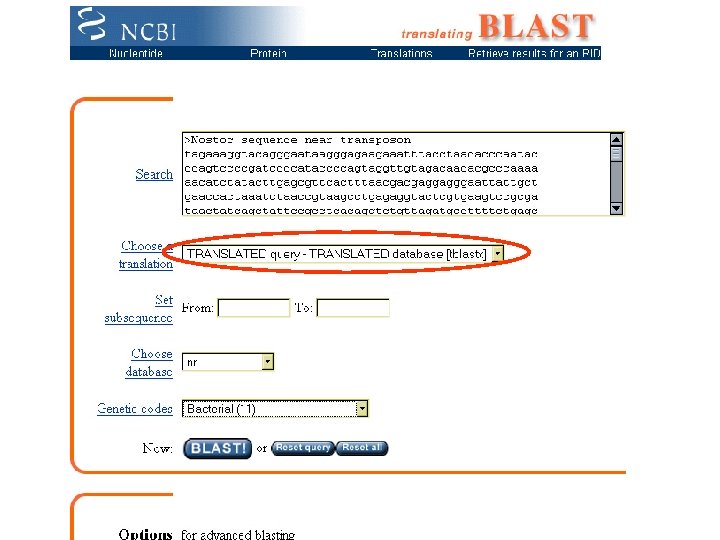

Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes Mission successful: >Translation: 397. . 639 (direct), 51 amino acids VQLAKQAQTAEGTLQIVTNARVTQTVKLVRLEKFLSLQSTVDAAVENIKGA*
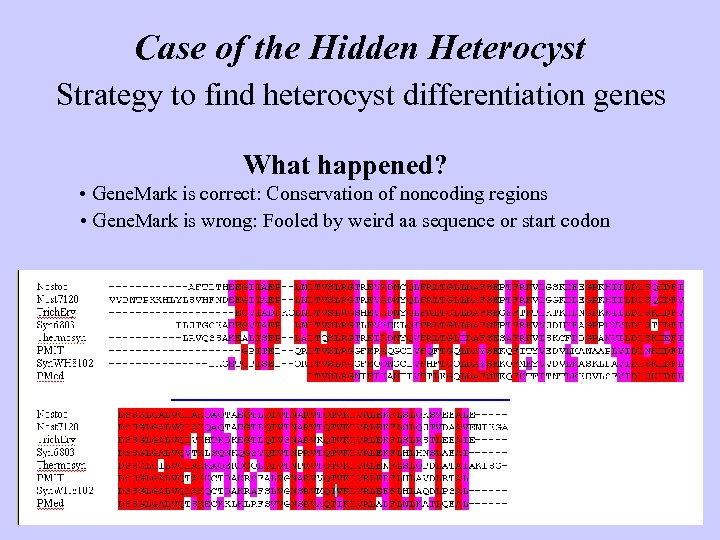
Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes What happened? • Gene. Mark is correct: Conservation of noncoding regions • Gene. Mark is wrong: Fooled by weird aa sequence or start codon

Case of the Hidden Heterocyst Strategy to find heterocyst differentiation genes What happened? • Gene. Mark is correct: Conservation of noncoding regions • Gene. Mark is wrong: Fooled by weird aa sequence or start codon Moral Automated gene finders are wonderful, but common sense is better Don’t trust automated annotation

Surrogate Filters Feature finders Hidden Markov model-based methods • Good for contiguous features (e. g. signal sequences) • Problems with features having gaps (e. g. promoters) Ad hoc methods • Feature-specific rules (e. g. tandem repeats, terminators) Position-dependent frequency tables = Position-specific scoring matrix (PSSM) = Weight table
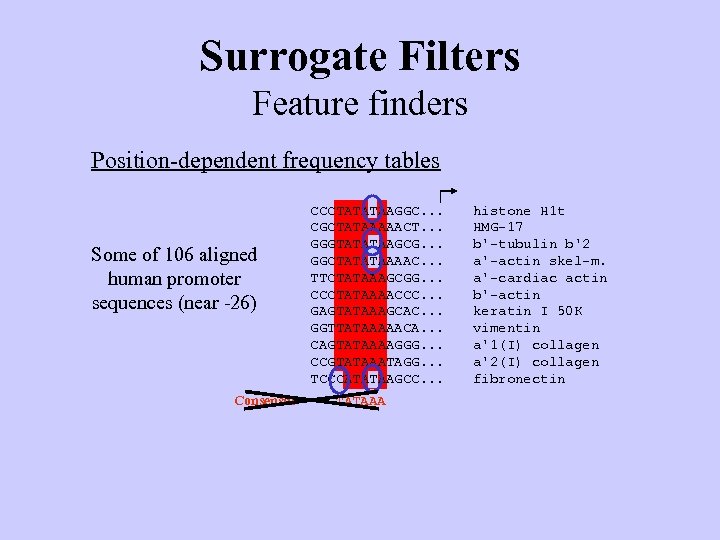

Surrogate Filters Feature finders Position-dependent frequency tables Some of 106 aligned human promoter sequences (near -26) Consensus CCCTATATAAGGC. . . CGCTATAAAAACT. . . GGGTATATAAGCG. . . GGCTATATAAAAC. . . TTCTATAAAGCGG. . . CCCTATAAAACCC. . . GAGTATAAAGCAC. . . GGTTATAAAAACA. . . CAGTATAAAAGGG. . . CCGTATAAATAGG. . . TCCCATATAAGCC. . . TATAAA histone H 1 t HMG-17 b'-tubulin b'2 a'-actin skel-m. a'-cardiac actin b'-actin keratin I 50 K vimentin a'1(I) collagen a'2(I) collagen fibronectin
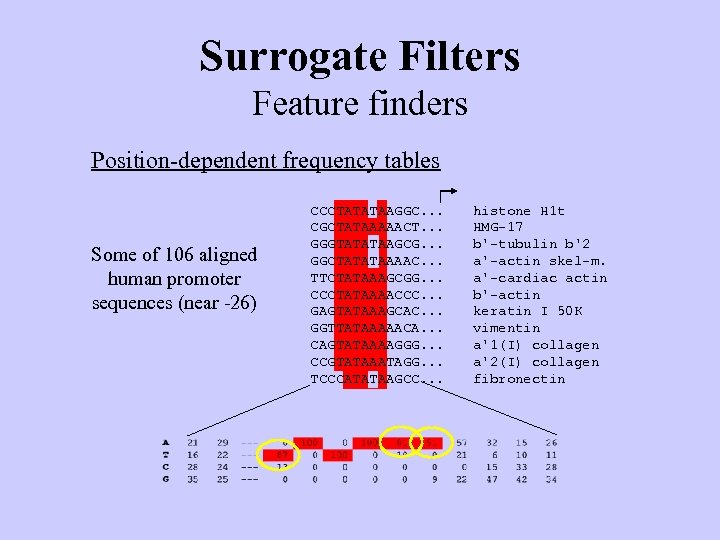
Surrogate Filters Feature finders Position-dependent frequency tables Some of 106 aligned human promoter sequences (near -26) CCCTATATAAGGC. . . CGCTATAAAAACT. . . GGGTATATAAGCG. . . GGCTATATAAAAC. . . TTCTATAAAGCGG. . . CCCTATAAAACCC. . . GAGTATAAAGCAC. . . GGTTATAAAAACA. . . CAGTATAAAAGGG. . . CCGTATAAATAGG. . . TCCCATATAAGCC. . . histone H 1 t HMG-17 b'-tubulin b'2 a'-actin skel-m. a'-cardiac actin b'-actin keratin I 50 K vimentin a'1(I) collagen a'2(I) collagen fibronectin
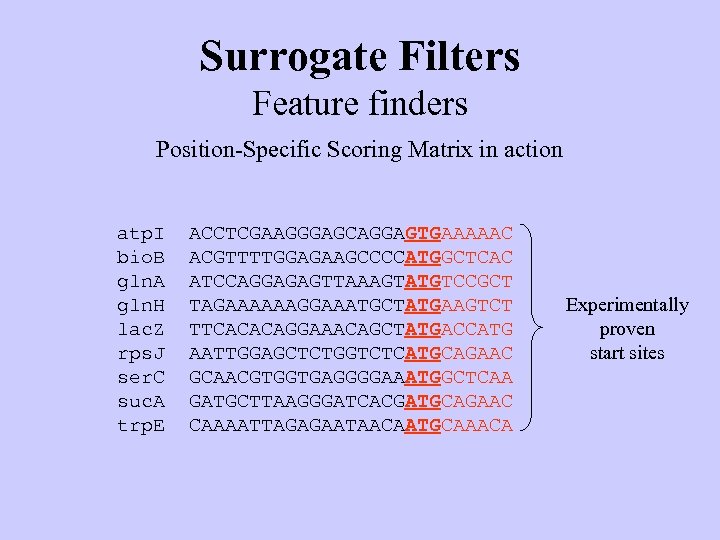

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action atp. I bio. B gln. A gln. H lac. Z rps. J ser. C suc. A trp. E ACCTCGAAGGGAGCAGGAGTGAAAAAC ACGTTTTGGAGAAGCCCCATGGCTCAC ATCCAGGAGAGTTAAAGTATGTCCGCT TAGAAAAAAGGAAATGCTATGAAGTCT TTCACACAGGAAACAGCTATGACCATG AATTGGAGCTCTGGTCTCATGCAGAAC GCAACGTGGTGAGGGGAAATGGCTCAA GATGCTTAAGGGATCACGATGCAGAAC CAAAATTAGAGAATAACAATGCAAACA Experimentally proven start sites
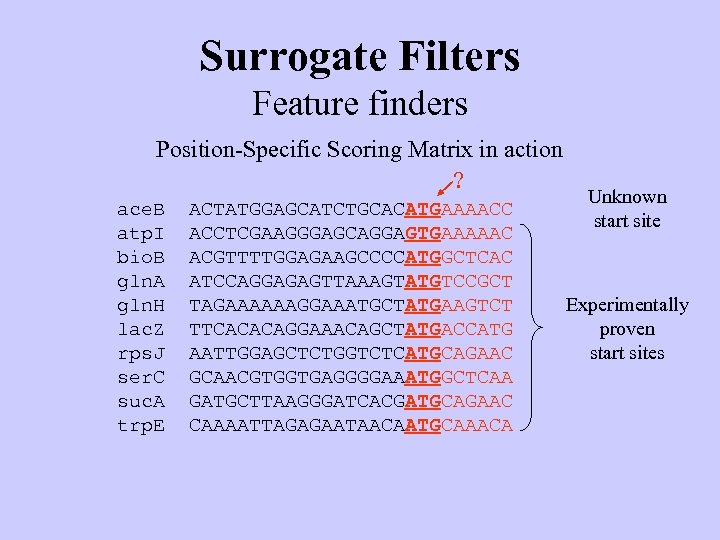

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action ? ace. B atp. I bio. B gln. A gln. H lac. Z rps. J ser. C suc. A trp. E ACTATGGAGCATCTGCACATGAAAACC ACCTCGAAGGGAGCAGGAGTGAAAAAC ACGTTTTGGAGAAGCCCCATGGCTCAC ATCCAGGAGAGTTAAAGTATGTCCGCT TAGAAAAAAGGAAATGCTATGAAGTCT TTCACACAGGAAACAGCTATGACCATG AATTGGAGCTCTGGTCTCATGCAGAAC GCAACGTGGTGAGGGGAAATGGCTCAA GATGCTTAAGGGATCACGATGCAGAAC CAAAATTAGAGAATAACAATGCAAACA Unknown start site Experimentally proven start sites
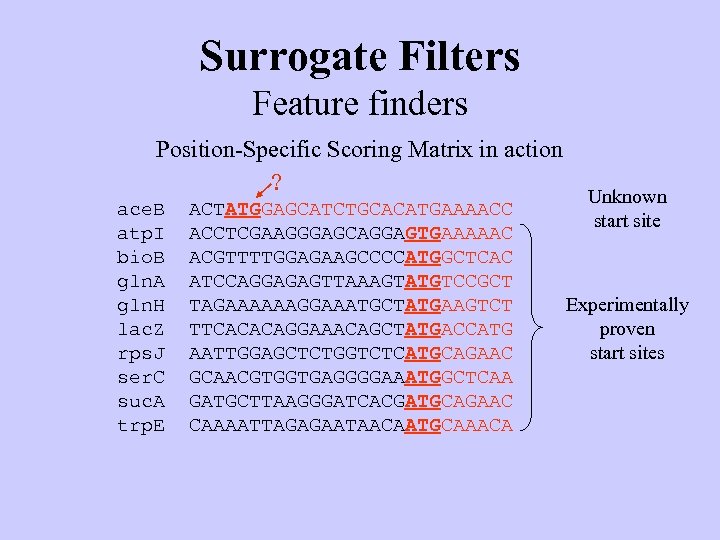

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action ? ace. B atp. I bio. B gln. A gln. H lac. Z rps. J ser. C suc. A trp. E ACTATGGAGCATCTGCACATGAAAACC ACCTCGAAGGGAGCAGGAGTGAAAAAC ACGTTTTGGAGAAGCCCCATGGCTCAC ATCCAGGAGAGTTAAAGTATGTCCGCT TAGAAAAAAGGAAATGCTATGAAGTCT TTCACACAGGAAACAGCTATGACCATG AATTGGAGCTCTGGTCTCATGCAGAAC GCAACGTGGTGAGGGGAAATGGCTCAA GATGCTTAAGGGATCACGATGCAGAAC CAAAATTAGAGAATAACAATGCAAACA Unknown start site Experimentally proven start sites
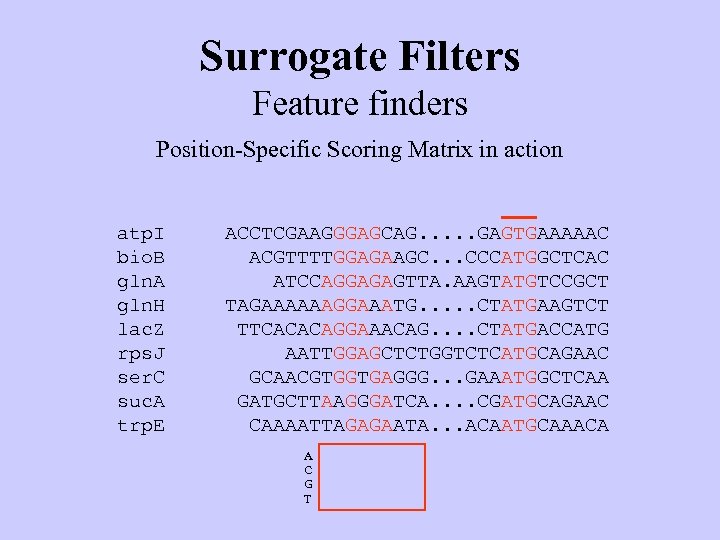

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action atp. I bio. B gln. A gln. H lac. Z rps. J ser. C suc. A trp. E ACCTCGAAGGGAGCAG. . . GAGTGAAAAAC ACGTTTTGGAGAAGC. . . CCCATGGCTCAC ATCCAGGAGAGTTA. AAGTATGTCCGCT TAGAAAAAAGGAAATG. . . CTATGAAGTCT TTCACACAGGAAACAG. . CTATGACCATG AATTGGAGCTCTGGTCTCATGCAGAAC GCAACGTGGTGAGGG. . . GAAATGGCTCAA GATGCTTAAGGGATCA. . CGATGCAGAAC CAAAATTAGAGAATA. . . ACAATGCAAACA A C G T
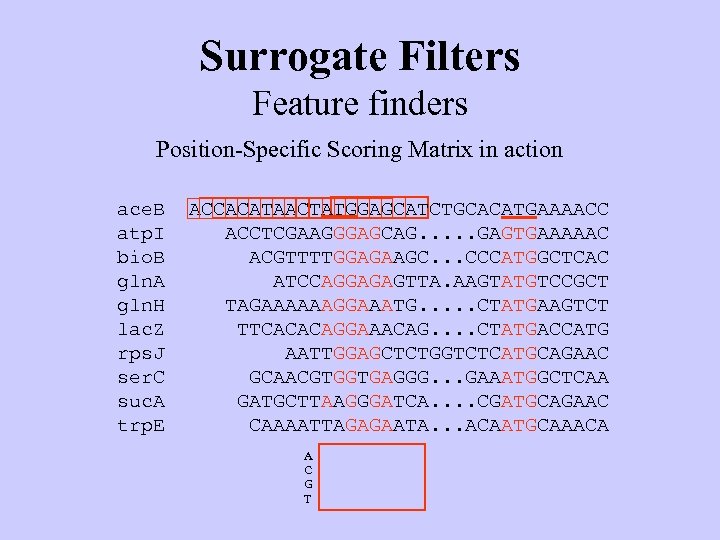

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action ace. B atp. I bio. B gln. A gln. H lac. Z rps. J ser. C suc. A trp. E ACCACATAACTATGGAGCATCTGCACATGAAAACC ACCTCGAAGGGAGCAG. . . GAGTGAAAAAC ACGTTTTGGAGAAGC. . . CCCATGGCTCAC ATCCAGGAGAGTTA. AAGTATGTCCGCT TAGAAAAAAGGAAATG. . . CTATGAAGTCT TTCACACAGGAAACAG. . CTATGACCATG AATTGGAGCTCTGGTCTCATGCAGAAC GCAACGTGGTGAGGG. . . GAAATGGCTCAA GATGCTTAAGGGATCA. . CGATGCAGAAC CAAAATTAGAGAATA. . . ACAATGCAAACA A C G T
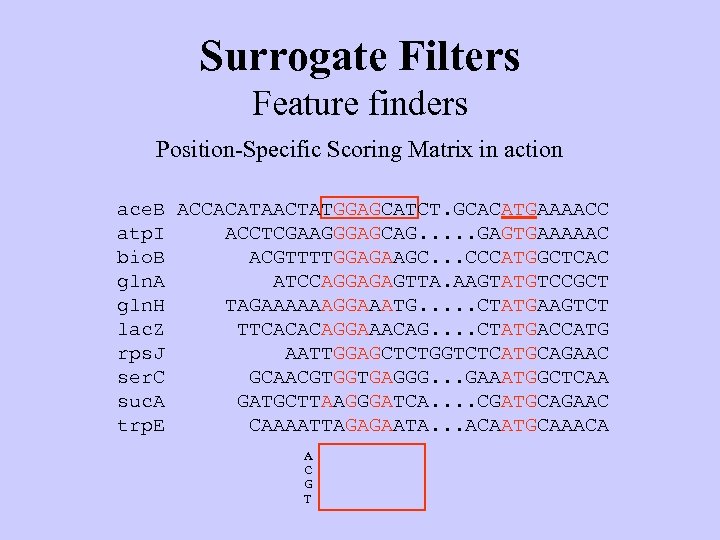

Surrogate Filters Feature finders Position-Specific Scoring Matrix in action ace. B ACCACATAACTATGGAGCATCT. GCACATGAAAACC atp. I ACCTCGAAGGGAGCAG. . . GAGTGAAAAAC bio. B ACGTTTTGGAGAAGC. . . CCCATGGCTCAC gln. A ATCCAGGAGAGTTA. AAGTATGTCCGCT gln. H TAGAAAAAAGGAAATG. . . CTATGAAGTCT lac. Z TTCACACAGGAAACAG. . CTATGACCATG rps. J AATTGGAGCTCTGGTCTCATGCAGAAC ser. C GCAACGTGGTGAGGG. . . GAAATGGCTCAA suc. A GATGCTTAAGGGATCA. . CGATGCAGAAC trp. E CAAAATTAGAGAATA. . . ACAATGCAAACA A C G T
Surrogate Filters Pattern finders New pattern discovery (Meme, Gibbs sampler, Bio. Prospector) Human sequences 5’ to transcriptional start sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 nucleolin sn. RNP E rp S 14 rp S 17 ribosomal p. S 19 a'-tubulin ba'1 b'-tubulin b'2 a'-actin skel-m. a'-cardiac actin b'-actin AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT GCAGGCTCAGTCTTTCGCCTCAGTCTCGAGCTCTCGCTGG TGCCGCCGCGTGACCTTCACACTTCCGGTTCTTT GACACGGAAGTGACCCCCGTCGCTCCGCCCTCTCCCACTC TGGCCTAAGCTTTAACAGGCTTCGCCTGTGCTTCCTGTTT ACCCTACGCCCGACTTGTGCGCCCGGGAAACCCCGTCGTT GGTCTGGGCGTCCCGGCTGGGCCCCGTGTCTGTGCGCACG GGGAGGGTATATAAGCGTTGGCGGACGGTTGTAGCA CCGCGGGCTATATAAAACCTGAGCAGAGGGACAAGCGGCC TCAGCGTTCTATAAAGCGGCCCTCCTGGAGCCACCC CGCGGCGGCGCCCTATAAAACCCAGCGGCGCGACGCGCCA
Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence
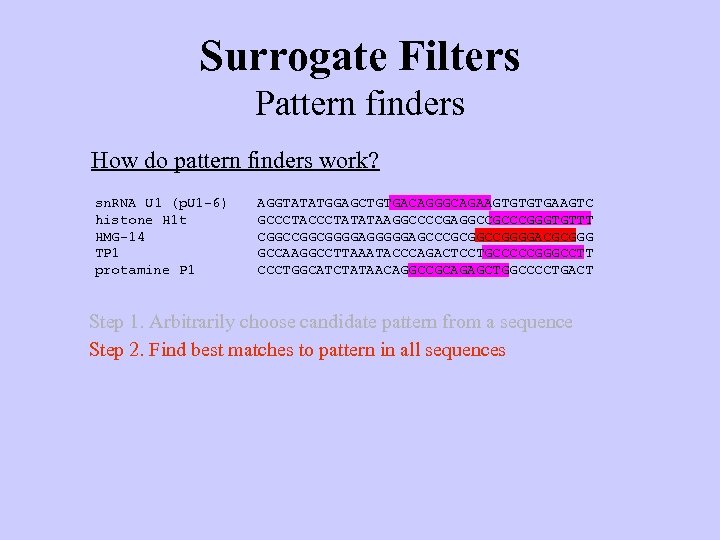

Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence Step 2. Find best matches to pattern in all sequences
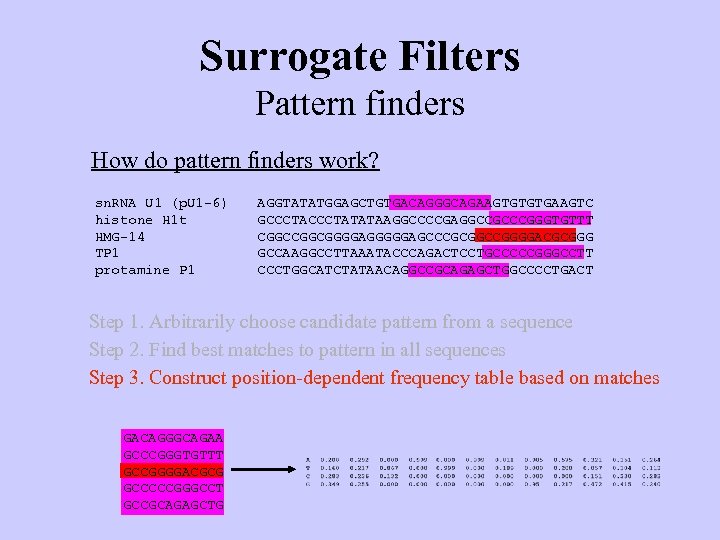

Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence Step 2. Find best matches to pattern in all sequences Step 3. Construct position-dependent frequency table based on matches GACAGGGCAGAA GCCCGGGTGTTT GCCGGGGACGCG GCCCCCGGGCCT GCCGCAGAGCTG
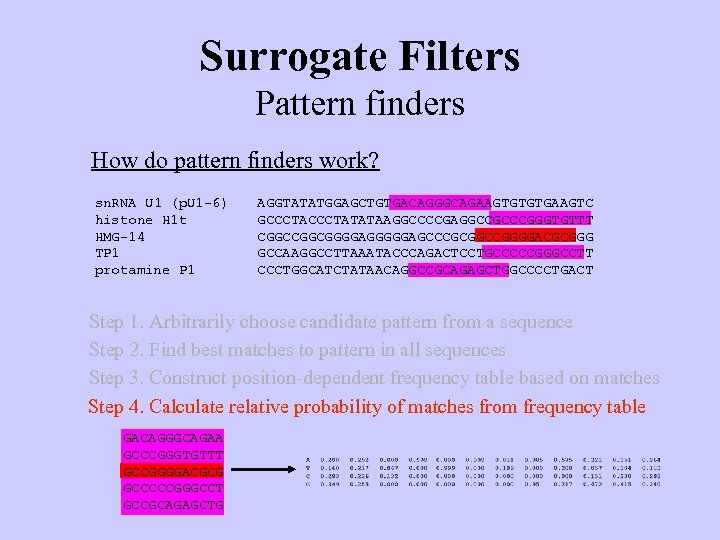

Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence Step 2. Find best matches to pattern in all sequences Step 3. Construct position-dependent frequency table based on matches Step 4. Calculate relative probability of matches from frequency table GACAGGGCAGAA GCCCGGGTGTTT GCCGGGGACGCG GCCCCCGGGCCT GCCGCAGAGCTG
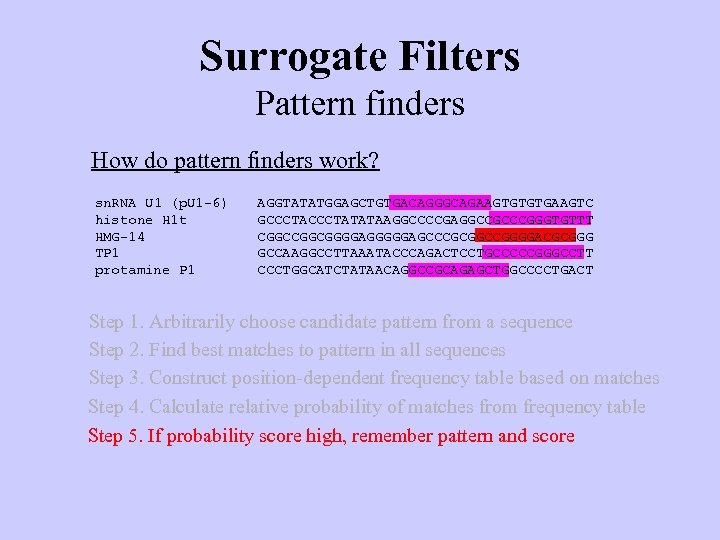

Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence Step 2. Find best matches to pattern in all sequences Step 3. Construct position-dependent frequency table based on matches Step 4. Calculate relative probability of matches from frequency table Step 5. If probability score high, remember pattern and score

Surrogate Filters Pattern finders How do pattern finders work? sn. RNA U 1 (p. U 1 -6) histone H 1 t HMG-14 TP 1 protamine P 1 AGGTATATGGAGCTGTGACAGGGCAGAAGTGTGTGAAGTC GCCCTATATAAGGCCCCGAGGCCGCCCGGGTGTTT CGGCGGGGAGCCCGCGGCCGGGGACGCGGG GCCAAGGCCTTAAATACCCAGACTCCTGCCCCCGGGCCTT CCCTGGCATCTATAACAGGCCGCAGAGCTGGCCCCTGACT Step 1. Arbitrarily choose candidate pattern from a sequence Step 2. Find best matches to pattern in all sequences Step 3. Construct position-dependent frequency table based on matches Step 4. Calculate relative probability of matches from frequency table Step 5. If probability score high, remember pattern and score Step 6. Repeat Steps 1 - 5

Surrogate Filters Scenario II – Case of the Masked Motif • You’ve found a gene related to Purple Tongue Syndrome • Blast. P: Encoded protein related to c. AMP-binding proteins • Are the similarities trivial? Related to c. AMP binding? • Does your protein contain c. AMP-binding site? • What IS a c. AMP-binding site? Task 1. Determine what is a c. AMP-binding site 2. Determine if your protein has one

Surrogate Filters Scenario II – Case of the Masked Motif Strategy 1. Collect sequences of known c. AMP-binding proteins 2. Run Meme, a pattern-finding program Ask it to find any significant motifs Do it 3. Rerun Meme. Demand that every protein has identified motifs 4. Run Pfam over known sequence to check


Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Progressive External Ophthalmoplegia (PEO) • Slow paralysis of voluntary eye muscles • Many other symptoms (e. g. , frequent deafness) • Loss of mitochondrial DNA

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Progressive External Ophthalmoplegia (PEO) • Slow paralysis of voluntary eye muscles • Many other symptoms (e. g. , frequent deafness) • Loss of mitochondrial DNA Inheritance • Mendelian • Autosomal dominant • Linked to chromosome 4 q 34

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Progressive External Ophthalmoplegia (PEO) • Slow paralysis of voluntary eye muscles • Many other symptoms (e. g. , frequent deafness) • Loss of mitochondrial DNA Inheritance • Mendelian • Autosomal dominant • Linked to chromosome 4 q 34 Your task • Examine sequence of 4 q 34 region • Assess likelihood that a gene in the area could cause disease symptoms

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Examining Sequence of 4 q 34 Region tctacttatattcaatccacagggctacacctagttcttggtacacagtacatgctcagcaagagtctgttgaacacatggtttatctgtttgtctcttccgagttcttgacttctgct ctgacctctggcagctttccactagtttctagctttcattctgcttacctggatttcggaactctagcctgccccactcttagataaacgcatgccctctgtggccctggaaccttagtgacttctgctat accaaagtctccacgcccagggtgacacgcagctccgtaaacctctaacatgatgtcagcaaatattaaaaaagtttataaaaacaatgaataaactttgttaaaggtacaaatgaaaat tagcaaacatgggaagataattgagtaaagagtttaaagttaaaaacgaattgcagtcattctaggggaacagttgtatttgaaaacctgtatggttacatgaactgcctaaaaaacaagctaagga aaattaaagctcagatttatatattttaagaaattgcaatttcctgggattaaatagcatttcctcaaccccagctgtcattaaaaagaggcaaatacagccaaggactggatcttctccgga aggctgacagcactgaccctcaagaaggcaccggctgacagaacattctgccctaatatgtgctgaaattccgctgagagcagagtggtacattgaaccctttaggggcttacaaaagaagtgtcct gtgttttagagtcacagagttttgcagaaacaagtatgaattcacctagtggccccctgcaccaggtctttcctgtgggcactgagtgcagacacatcaatatgtaatagcagaatgactgaacgaa cgattgaaaagaaatgagaggcagcaggttgtcagattctatgaggcaatcacagcatcaggtgaccttagtatctatttgagaggactgccatttattctcgggagcgcacggctctaaagaggcc catatccaggcagtgagctctggtggggggcgcctttagatgcaagaaggaggaaacagctcgaaatccctgggcctgagcgcggcccgtgcaggccggagggtcaagaactctccaccggcggcagcggc ccggtgtctgccccggcttcgccccggcctaaggctgcctgtgctataaatacgcggcccacatgccgcggtgacacggtgttccctgggctcggcgggacagataacatgaatgtgccctttaaacgtcc caagttgcagggacagcccccggcccagcctcgctcccggaagcgccttcgcccccgatgccctctgcagctgggaggagggggcgccccgcacctgcccagccaatgcgcgagcgccgcga cccgcctcctctcgcgagagcccggcggggatataagggggagctgcgggccaggcggcggccccctagcgtcgcgcagggtcggggactgcgcgcggtgccaggccgggcgtgggcgagagcacgaacgg gctgcgggctgagagcgtcgagctgtcaccatgggtgatcacgcttggagcttcctaaaggacttcctggccgggggcgtcgccgctgccgtctccaagaccgcggtcgcccccatcgagagggtca aactgctgctgcaggtgaggaccgcgcggtgcaagaggcgcgggcgcggcgggcggggcgcgcgatgcggcgcgagctgcagggcgcggggcgccgcggaaaatctgcgccaggccacaggc ccgggcgcccgcccgcgggggaagaaggtgccctctgcgtagagacaggtccagcgtcagtcgcagattcctggtgtcgggtggcgcccggcgttcgggtgtctatatatggaaacccggagc cggtttacgtgtgccagatcctgcgcccgtgacagcacgggcgtgcactcaggcccggaggcacctagtgattgccagtatttttggcaccgtcttatgcgcacctttacaataaaaacatcaaaat aatcatcacccaagaattcccttatcgtatctcatgcacaatgctgtaggctgacgccttcatctttatgtaacctctgtgagagagttattcttctccattttacagatgaagctgaggttttgaa atattaagaaacaattttcggaataaactcagatcatcctgtctccaaatcttttcctcccctacctggtcgctgaatggtttatcatcctctcgtgttttcctccacctgcccaaaaggtcagggcccct caatgaggaagagcccaatttgggagtcagaattactaacaacaaaacccccacaaattgctcacaacggcagcaaacccttaataattgattacttggattatctgcttgaaaactttggaggcctaatg tttagtggatttattctcctctattagagcatctagtagagatcctcatctccagggtgatcagagtgacactgagaaattgtcattttttggccatcatgtctattaaatccaaagccctttgaag cagggagtgttactcatttctgtcccccagtaagcccctcatacagttctcaaacctagggaaagtgaaataaatggctatagctttatataattcaatcaccttttcagtttatttggggcaatac ctttccctcaaataccctaataattgaagcaacattggattattttggcttgttatccagtaacatggataacagtatccatttacacgtcctcgtatccatttgatttcctcatcctttttttctt caaaaatctaggaagtgcaaacctttttctcctgtcctcttcccttctctctaccctgtcctctgtcaccctcccctccaccaggtccagcatgccagcaaacagatcagtgc tgagaagcagtacaaagggatcattgtgtggtgagaatccctaaggagcagggcttcctctccttctggaggggtaacctggccaacgtgatccgttacttccccaagctctcaacttcgcct tcaaggacaagtacaagcagctcttcttagggggtgtggatcggcataagcagttctggcgctactttgctggtaacctggcgtccggtggggccgctggggccacctccctttgtctacccgctg gactttgctaggaccaggttggctgctgatgtgggcaagggcgccgcccagcgtgagttccatggtctgggcgactgtatcatcaagatcttcaagtctgatggcctgagggggctctaccagggtttcaa cgtctctgtccaaggcatcattatctatagagctgcctacttcggagtctatgatactgccaagggtgagagaggggcatcggggagaaggagggtggtgtggaaagaggatcctatgggatctataactc acaaaggacctgatattgatcttgttttttctagtctctgggataattgaggcttctgaatgaggaggtgatgtgcataagttaatagctgaagcgttccttgtgtcctctactgaaataaactctg gcctttagttattcagagaggaggaggggggagcctgtctccctctagacacagccatagcagttactgagtttaacttgaagccacttccaatgccctgtatacaagctgagcactgcccctccggggtc cggagagggcagcagccacctttgctgtctgcctggtcatatgtgaagcacctgcacaggggcaggttccccgcaaggtcagagcatggagctggaggtgcagtggcctctctccacctgctttctg ctgagaacaggcacttcatagccgttcggcttctgggctctgtccacagggatgctgcctgaccccaagaacgtgcacatttttgtgagctggatgattgcccagagtgtgacggcagtcgcagggctggt gtcctacccctttgacactgttcgtcgtagaatgatgatgcagtccggaaagggggtaagcttgtgctctactcatctaaacttgtttggttttgcccgaggagaacattttacagggctcctttca gtcttccttactggaaattttcaaaattatttgataaggacttagggaagatggtattaattccccctaacgttctcaactatcctattagggaaaagtattttccattttattagagatgat aagaacatgaatagtaagacatttagatgtgaatttaactaggtatccagcattatagagaccctaggccctcttcccttagagcctgggtgcaaaagctagggaaaagaagtagttagctacttcttaca aagaactcttgcttccctcctagttacaggtgttagtgggatggggtgtttagctgggtagagatggcctgaagcaatctgttgtgccagagaaagttttggcttctataggttgaaccatatgaaattgc cactttaaaagtcaaaaacagtccaatgttagcagtttcgtatgtttcaacgaatagttacagccttttagactgcataacctcgtgcaggatcatctgaggctcagcctcagttcggtcctccata aaaaaaggtaaccgcgtagcataatactcctgctccactgcgcccttcttgtttcgcagttgggcagtccatgaattacttggttaattgccccagttcttcactgaccttgaactaatggagtaggaatg acaggagacccagcctgccagtgaagcaaggagatgtccagtgggatgttgcatggagctgggactccatgcccagatgaccctgattttataaaactggtaacagtgtgtacagatatgtttcagg ggaaaagtctctttcctccagcgttacggagccctcaccagcatttgtttccacagccgatattatgtacacggggacagttgactgctggaggaagattgcaaaagacgaaggagccaaggccttcttca aaggtgcctggtccaatgtgctgagaggcatgggcggtgcttttgtattggtgttgtatgatgagatcaaaaaatatgtctaatgtaattaaaacacaagttcacagatttacatgaacttgatctacaag ttcacagatccattgtgtggtttaatagactattcctaggggaagtaaaaagatctgggataaaaccagactgaaggaatacctcagaagagatgcttcattgagtgttcattaaaccacacatgtatttt

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Strategy • Assume that encoded protein is in mitochondria • Protein has function associated with mitochondrial location? – Use Gene finder to identify protein sequence(s) – Use Similarity finder to identify possible function Do it • Protein has structure associated with mitochondrial location? – Use Feature finders to identify pertinent regions – (What ARE pertinent regions? )
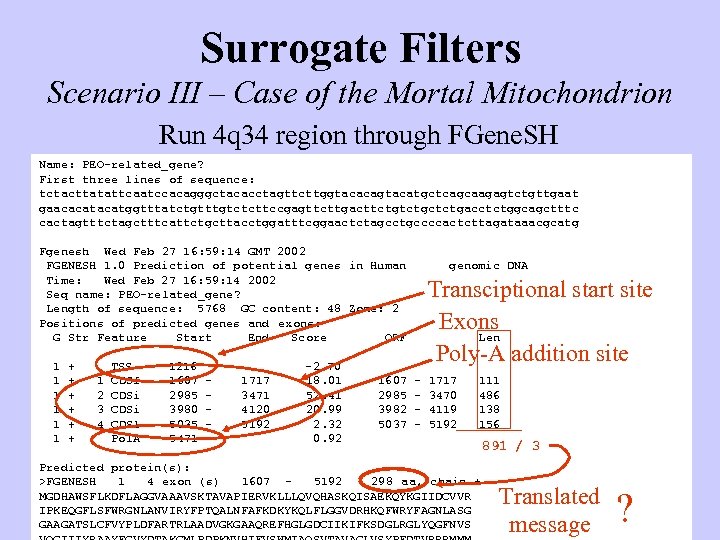


Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Run 4 q 34 region through FGene. SH Name: PEO-related_gene? First three lines of sequence: tctacttatattcaatccacagggctacacctagttcttggtacacagtacatgctcagcaagagtctgttgaat gaacacatggtttatctgtttgtctcttccgagttcttgacttctgctctgacctctggcagctttc cactagtttctagctttcattctgcttacctggatttcggaactctagcctgccccactcttagataaacgcatg Fgenesh Wed Feb 27 16: 59: 14 GMT 2002 FGENESH 1. 0 Prediction of potential genes in Human Time: Wed Feb 27 16: 59: 14 2002 Seq name: PEO-related_gene? Length of sequence: 5768 GC content: 48 Zone: 2 Positions of predicted genes and exons: G Str Feature Start End Score ORF 1 1 1 + + + 1 2 3 4 TSS CDSf CDSi CDSl Pol. A 1216 1607 2985 3980 5035 5471 - 1717 3471 4120 5192 -2. 70 18. 01 52. 41 20. 99 2. 32 0. 92 1607 2985 3982 5037 genomic DNA Transciptional start site Exons Len Poly-A addition site - 1717 3470 4119 5192 111 486 138 156 Predicted protein(s): >FGENESH 1 4 exon (s) 1607 5192 298 aa, chain + MGDHAWSFLKDFLAGGVAAAVSKTAVAPIERVKLLLQVQHASKQISAEKQYKGIIDCVVR IPKEQGFLSFWRGNLANVIRYFPTQALNFAFKDKYKQLFLGGVDRHKQFWRYFAGNLASG GAAGATSLCFVYPLDFARTRLAADVGKGAAQREFHGLGDCIIKIFKSDGLRGLYQGFNVS 891 / 3 Translated message ?
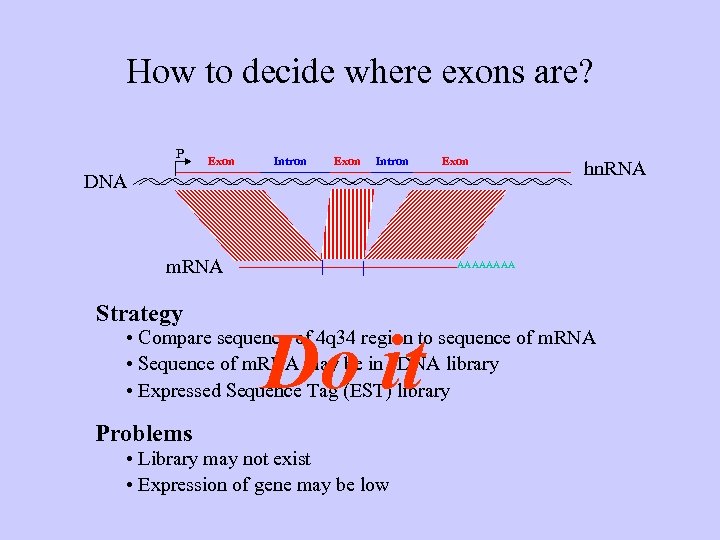
How to decide where exons are? P Exon Intron Exon DNA m. RNA Strategy hn. RNA AAAA Do it • Compare sequence of 4 q 34 region to sequence of m. RNA • Sequence of m. RNA may be in c. DNA library • Expressed Sequence Tag (EST) library Problems • Library may not exist • Expression of gene may be low

*

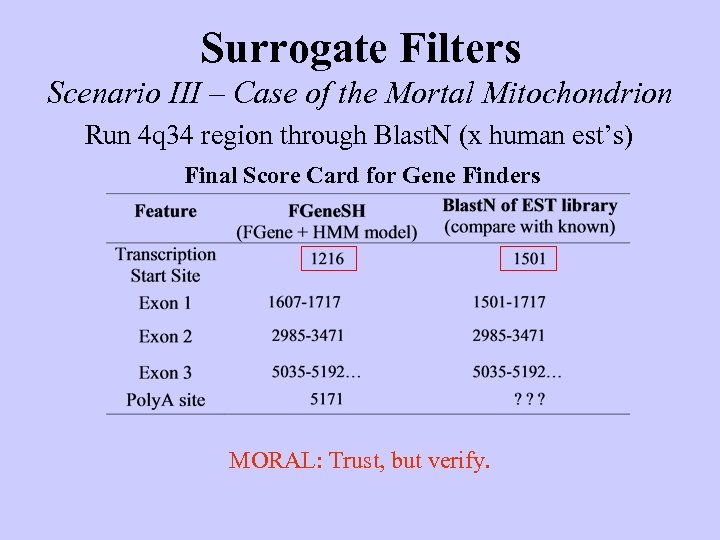
Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Run 4 q 34 region through Blast. N (x human est’s) Final Score Card for Gene Finders MORAL: Trust, but verify.

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Strategy • Assume that encoded protein is in mitochondria • Protein has function associated with mitochondrial location? – Use Gene finder to identify protein sequence(s) – Use Similarity finder to identify possible function • Protein has structure associated with mitochondrial location? – Use Feature finders to identify pertinent structures – (What ARE pertinent structures? )

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Run 4 q 34 region through Blast. P Name: PEO-related_gene? First three lines of sequence: tctacttatattcaatccacagggctacacctagttcttggtacacagtacatgctcagcaagagtctgttgaat gaacacatggtttatctgtttgtctcttccgagttcttgacttctgctctgacctctggcagctttc cactagtttctagctttcattctgcttacctggatttcggaactctagcctgccccactcttagataaacgcatg Fgenesh Wed Feb 27 16: 59: 14 GMT 2002 FGENESH 1. 0 Prediction of potential genes in Human Time: Wed Feb 27 16: 59: 14 2002 Seq name: PEO-related_gene? Length of sequence: 5768 GC content: 48 Zone: 2 Positions of predicted genes and exons: G Str Feature Start End Score ORF 1 1 1 + + + 1 2 3 4 TSS CDSf CDSi CDSl Pol. A 1216 1607 2985 3980 5035 5471 - 1717 3471 4120 5192 -2. 70 18. 01 52. 41 20. 99 2. 32 0. 92 1607 2985 3982 5037 genomic DNA Len - 1717 3470 4119 5192 Predicted protein(s): >FGENESH 1 4 exon (s) 1607 5192 298 aa, chain + MGDHAWSFLKDFLAGGVAAAVSKTAVAPIERVKLLLQVQHASKQISAEKQYKGIIDCVVR IPKEQGFLSFWRGNLANVIRYFPTQALNFAFKDKYKQLFLGGVDRHKQFWRYFAGNLASG GAAGATSLCFVYPLDFARTRLAADVGKGAAQREFHGLGDCIIKIFKSDGLRGLYQGFNVS VQGIIIYRAAYFGVYDTAKGMLPDPKNVHIFVSWMIAQSVTAVAGLVSYPFDTVRRRMMM 111 486 138 156 Translated message

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Strategy • Assume that encoded protein is in mitochondria • Protein has function associated with mitochondrial location? – Use Gene finder to identify protein sequence(s) – Use Similarity finder to identify possible function • Protein has structure associated with mitochondrial location? – Use Feature finders to identify pertinent structures – (What ARE pertinent structures? )

Surrogate Filters Scenario III – Case of the Mortal Mitochondrion Progressive External Ophthalmoplegia (PEO) • Slow paralysis of voluntary eye muscles • Many other symptoms (e. g. , frequent deafness) • Loss of mitochondrial DNA Inheritance • Mendelian • Autosomal dominant • Linked to chromosome 4 q 34

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike
2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike Escherichia coli. . . very small lab rats Courtesy of Kent State University Microbiology

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike Escherichia coli. . . haemorrhagic colitis
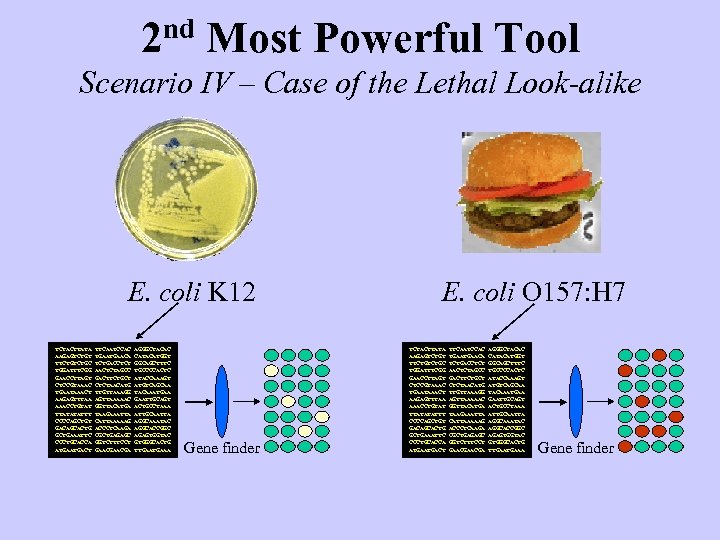

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder
2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder
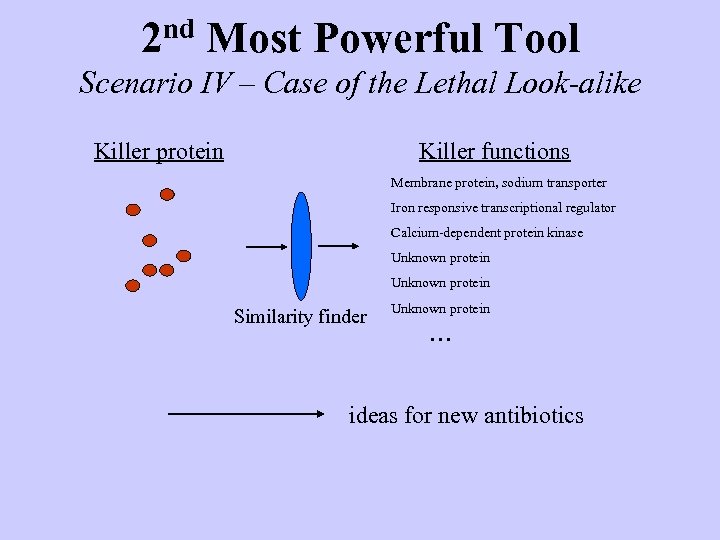
2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike Killer protein Killer functions Membrane protein, sodium transporter Iron responsive transcriptional regulator Calcium-dependent protein kinase Unknown protein Similarity finder Unknown protein . . . ideas for new antibiotics

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder What tool to use? Go to http: //www. vcu. edu/~elhaij/Bio. Inf
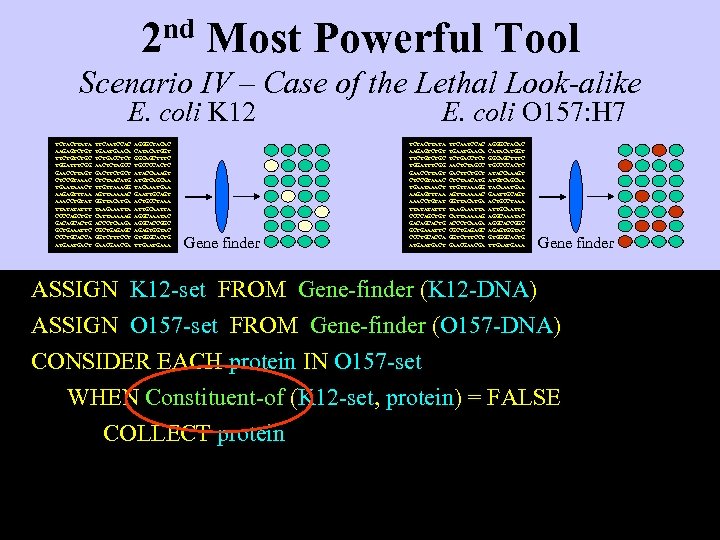

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder ASSIGN K 12 -set FROM Gene-finder (K 12 -DNA) ASSIGN O 157 -set FROM Gene-finder (O 157 -DNA) CONSIDER EACH protein IN O 157 -set WHEN Constituent-of (K 12 -set, protein) = FALSE COLLECT protein
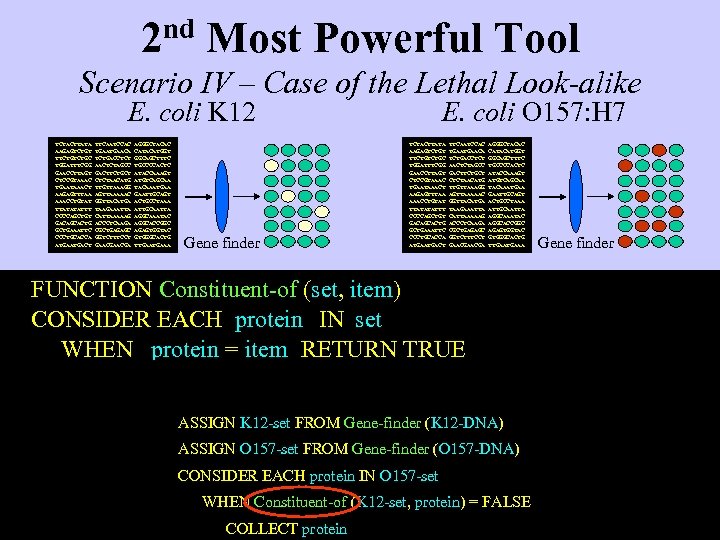

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA FUNCTION Constituent-of (set, item) CONSIDER EACH protein IN set WHEN protein = item RETURN TRUE OTHERWISE RETURN FALSE ASSIGN K 12 -set FROM Gene-finder (K 12 -DNA) ASSIGN O 157 -set FROM Gene-finder (O 157 -DNA) CONSIDER EACH protein IN O 157 -set WHEN Constituent-of (K 12 -set, protein) = FALSE COLLECT protein Gene finder
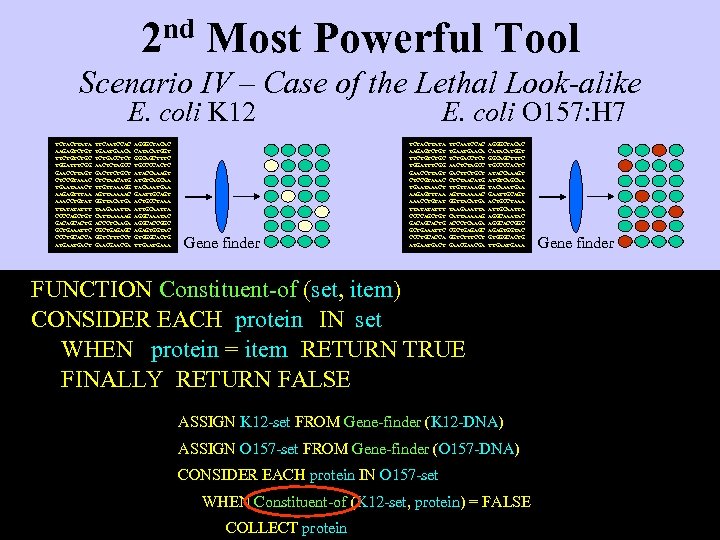

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA FUNCTION Constituent-of (set, item) CONSIDER EACH protein IN set WHEN protein = item RETURN TRUE FINALLY RETURN FALSE ASSIGN K 12 -set FROM Gene-finder (K 12 -DNA) ASSIGN O 157 -set FROM Gene-finder (O 157 -DNA) CONSIDER EACH protein IN O 157 -set WHEN Constituent-of (K 12 -set, protein) = FALSE COLLECT protein Gene finder
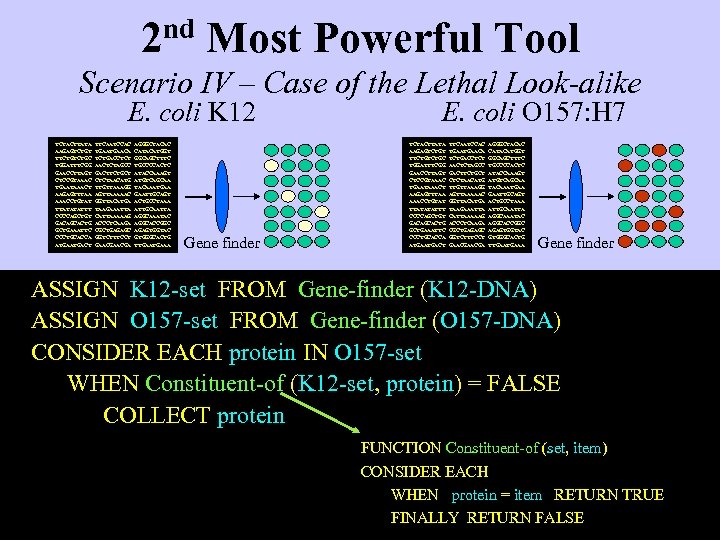

2 nd Most Powerful Tool Scenario IV – Case of the Lethal Look-alike E. coli K 12 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder E. coli O 157: H 7 TCTACTTATA AAGAGTCTGT TTCTGC TGGATTTCGG GAACCTTAGT CTCCGTAAAC TGAATAAACT AAGAGTTTAA AAACCTGTAT TTATATATTT CCCCAGCTGT GACAGCACTG GCTGAAATTC CCCTGCACCA ATGACT TTCAATCCAC TGAACA TCTGACCTCT AACTCTAGCC GACTTCTGCT CTCTAACATG TTGTTAAAGG AGTTAAAAAC GGTTACATGA TAAGAAATTA CATTAAAAAG ACCCTCAAGA CGCTGAGAGC GGTCTTTCCT GAACGA AGGGCTACAC CATACATGGT GGCAGCTTTC TGCCCCACTC ATACCAAAGT ATGTCAGCAA TACAAATGAA GAATTGCAGT ACTGCCTAAA ATTGCAATTA AGGCAAATAC AGGCACCGGC AGAGTGGTAC GTGGGCACTG TTGAAA Gene finder ASSIGN K 12 -set FROM Gene-finder (K 12 -DNA) ASSIGN O 157 -set FROM Gene-finder (O 157 -DNA) CONSIDER EACH protein IN O 157 -set WHEN Constituent-of (K 12 -set, protein) = FALSE COLLECT protein FUNCTION Constituent-of (set, item) CONSIDER EACH WHEN protein = item RETURN TRUE FINALLY RETURN FALSE

2 nd Most Powerful Tool Computer programming • Make your own tools • Change them at will • Use them to teach you Glob in

FIRST Most Powerful Tool Your brain • Keep your nonsense detector on high alert Mission successful: >Translation: 358. . 513 (direct), 51 amino acids VQLAKQAQTAEGTLQIVTNARVTQTVKLVRLEKFLSLQKSVEEALENVK* … or was it?
FIRST Most Powerful Tool Your brain • Keep your nonsense detector on high alert • Appreciate the limitations of bioinformatic tools Challenge accepted beliefs Conform to standard model
FIRST Most Powerful Tool Your brain • Keep your nonsense detector on high alert • Appreciate the limitations of bioinformatic tools • Look out for surprises in the underlying data Challenge accepted beliefs Conform to standard model

31d06aa6aec56ad2eee518f10c7801bf.ppt