7029379b2bab49f2b8a65f84a5f54198.ppt
- Количество слайдов: 71

National Center for Biote Information (NCB www. ncbi. nlm. nih Bunu databases’in icine koy le Page 24
www. ncbi. nlm. nih. gov Fig. 2. 5 Page 25
Fig. 2. 5 Page 25

Pub. Med is… • National Library of Medicine's search service • 16 million citations in MEDLINE • links to participating online journals • Pub. Med tutorial (via “Education” on side bar) Page 24

Entrez integrates… • the scientific literature; • DNA and protein sequence databases; • 3 D protein structure data; • population study data sets; • assemblies of complete genomes Page 24
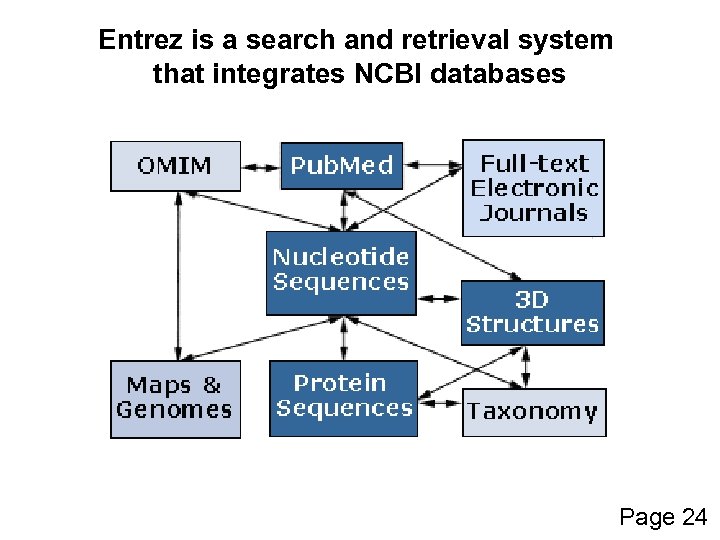
Entrez is a search and retrieval system that integrates NCBI databases Page 24

BLAST is… • Basic Local Alignment Search Tool • NCBI's sequence similarity search tool • supports analysis of DNA and protein databases • 100, 000 searches per day Page 25

OMIM is… • Online Mendelian Inheritance in Man • catalog of human genes and genetic disorders • edited by Dr. Victor Mc. Kusick, others at JHU Page 25

Cancer Chromosomes Contains cytogenetic, clinical, and reference information from integrated information from the NCI Mitelman Database of Chromosome Aberrations in Cancer, the NCI Recurrent Aberrations in Cancer database, and the NCI/NCBI SKY/M-FISH & CGH Database.

CDD Conserved Domain Database, a collection of sequence alignments and profiles representing protein domains conserved in molecular evolution. Select 'Domains' from the Entrez pull down menu.

Core. Nucleotide Contains all nucleotide sequences not included in the EST or GSS subsets. 3 D Domains Contains protein domains from the Entrez Structure database. EST A Nucleotide database subset that contains only Expressed Sequence Tag records. Genes and associated information for a number of organisms in addition to and including human.
Genomes of over 1, 200 organisms can be found in this database, representing both completely sequenced organisms and those for which sequencing is in progress. Genome Project A searchable collection of complete and incomplete (in-progress) large-scale sequencing, assembly, annotation, and mapping projects for cellular organisms. db. Ga. P Associated genotype and phenotype data. GENSAT Gene expression atlas of the mouse central nervous system.

GEO Datasets Curated gene expression and molecular abundance Data. Sets from NCBI's Gene Expression Omnibus, a gene expression and hybridization array repository. GEO Profiles Individual gene expression and molecular abundance profiles assembled from the GEO repository. http: //www. ncbi. nlm. nih. gov/About/tools/restable_mol. html

Books is… • searchable resource of on-line books Page 26

Tax. Browser is… • browser for the major divisions of living organisms (archaea, bacteria, eukaryota, viruses) • taxonomy information such as genetic codes • molecular data on extinct organisms Page 26

Structure site includes… • Molecular Modelling Database (MMDB) • biopolymer structures obtained from the Protein Data Bank (PDB) • Cn 3 D (a 3 D-structure viewer) • vector alignment search tool (VAST) Page 26

Accessing information on molecular sequences Page 26

Accession numbers are labels for sequences NCBI includes databases (such as Gen. Bank) that contain information on DNA, RNA, or protein sequences. You may want to acquire information beginning with a query such as the name of a protein of interest, or the raw nucleotides comprising a DNA sequence of interest. DNA sequences and other molecular data are tagged with accession numbers that are used to identify a sequence or other record relevant to molecular data. Page 26
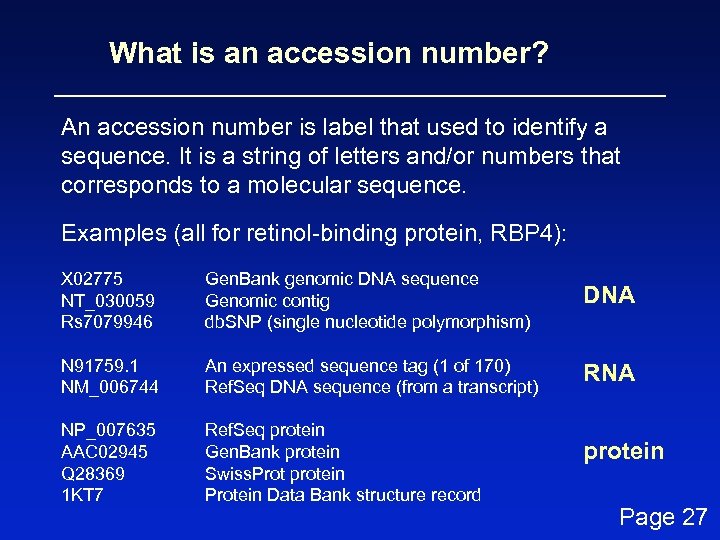

What is an accession number? An accession number is label that used to identify a sequence. It is a string of letters and/or numbers that corresponds to a molecular sequence. Examples (all for retinol-binding protein, RBP 4): X 02775 NT_030059 Rs 7079946 Gen. Bank genomic DNA sequence Genomic contig db. SNP (single nucleotide polymorphism) DNA N 91759. 1 NM_006744 An expressed sequence tag (1 of 170) Ref. Seq DNA sequence (from a transcript) RNA NP_007635 AAC 02945 Q 28369 1 KT 7 Ref. Seq protein Gen. Bank protein Swiss. Prot protein Protein Data Bank structure record protein Page 27
![Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq](https://present5.com/presentation/7029379b2bab49f2b8a65f84a5f54198/image-21.jpg)
Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq [2] Uni. Gene [3] European Bioinformatics Institute (EBI) and Ensembl (separate from NCBI) [4] Ex. PASy Sequence Retrieval System (separate from NCBI) Note: Locus. Link at NCBI was recently retired. The third printing of the book has updated these sections (pages 27 -31). Page 27
![4 ways to access protein and DNA sequences [1] Entrez Gene with Ref. Seq 4 ways to access protein and DNA sequences [1] Entrez Gene with Ref. Seq](https://present5.com/presentation/7029379b2bab49f2b8a65f84a5f54198/image-22.jpg)
4 ways to access protein and DNA sequences [1] Entrez Gene with Ref. Seq Entrez Gene is a great starting point: it collects key information on each gene/protein from major databases. It covers all major organisms. Ref. Seq provides a curated, optimal accession number for each DNA (NM_006744) or protein (NP_007635) Page 27
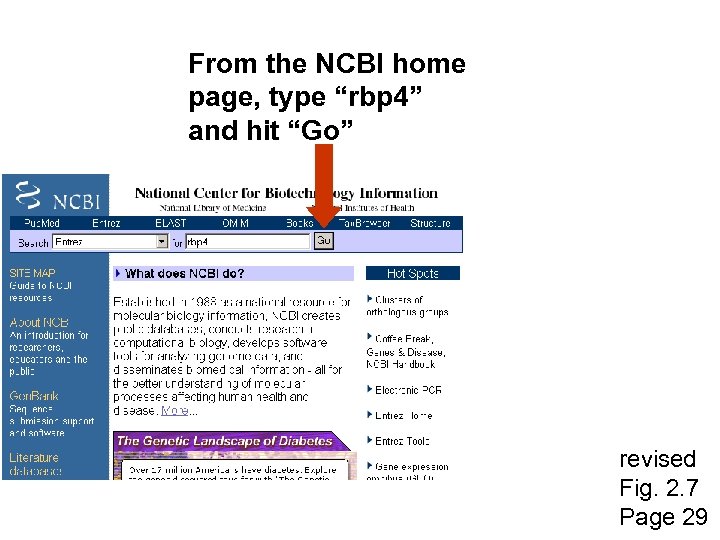

From the NCBI home page, type “rbp 4” and hit “Go” revised Fig. 2. 7 Page 29
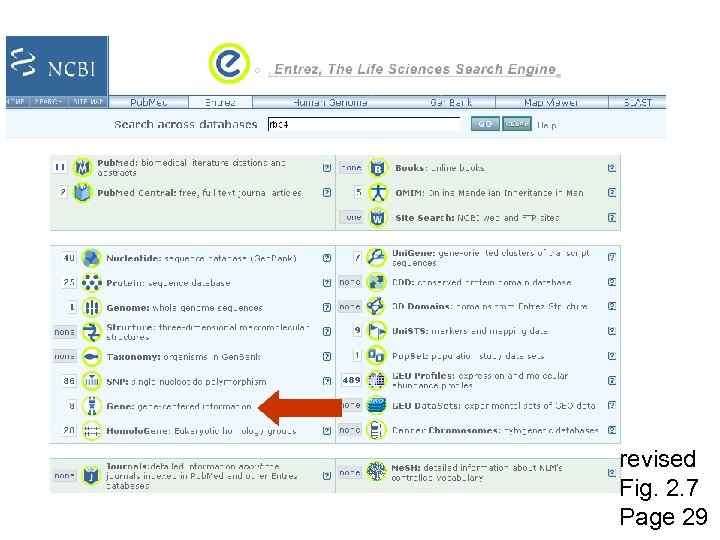
revised Fig. 2. 7 Page 29
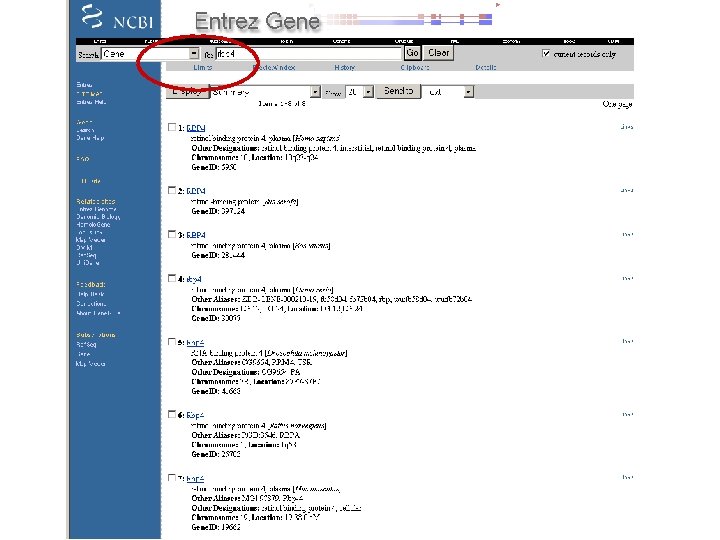
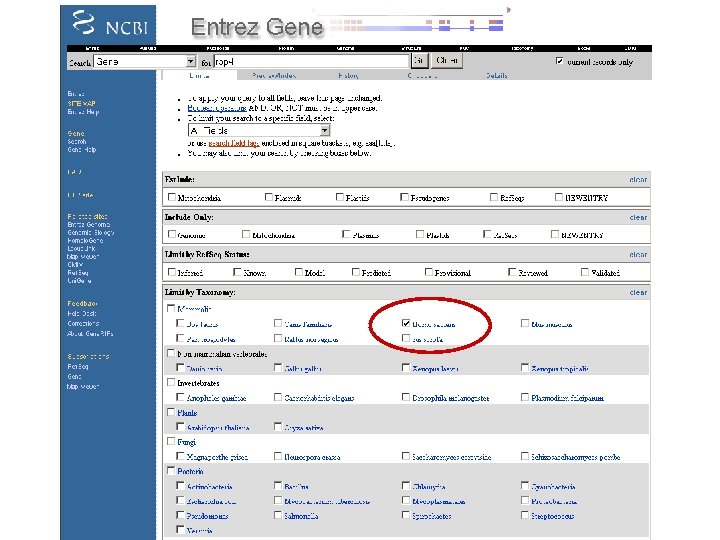

By applying limits, there are now just two entries
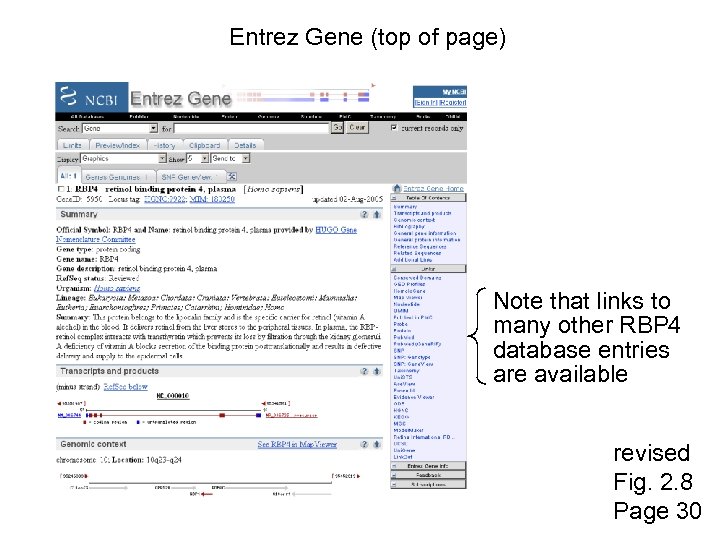
Entrez Gene (top of page) Note that links to many other RBP 4 database entries are available revised Fig. 2. 8 Page 30
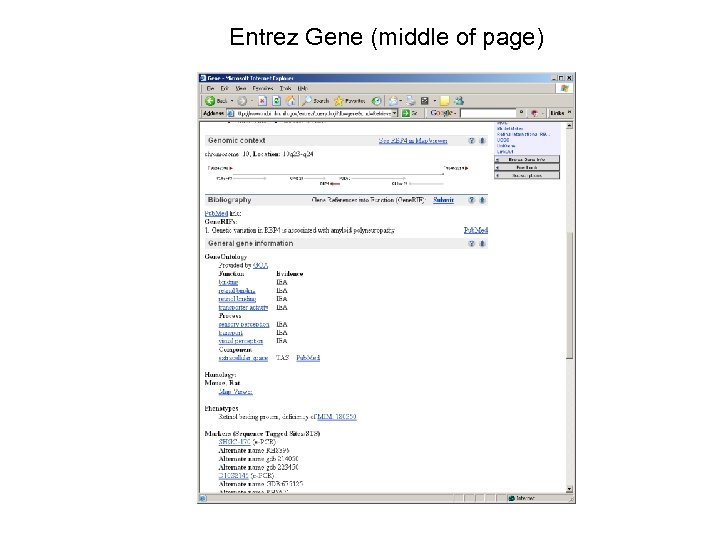
Entrez Gene (middle of page)
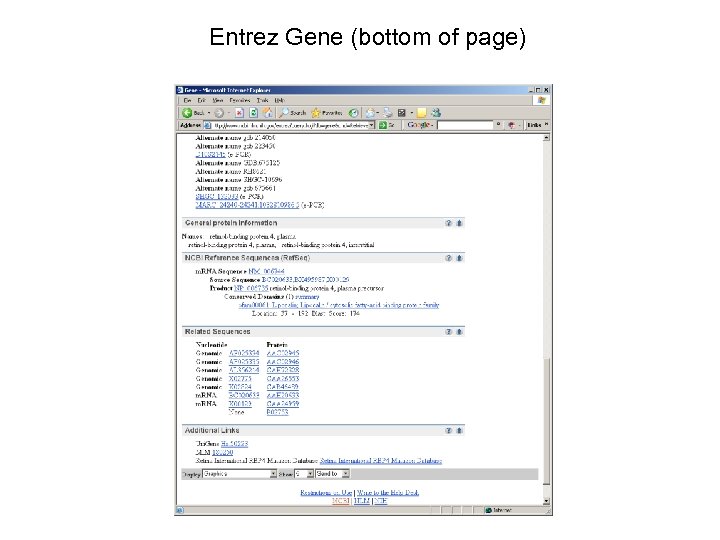
Entrez Gene (bottom of page)
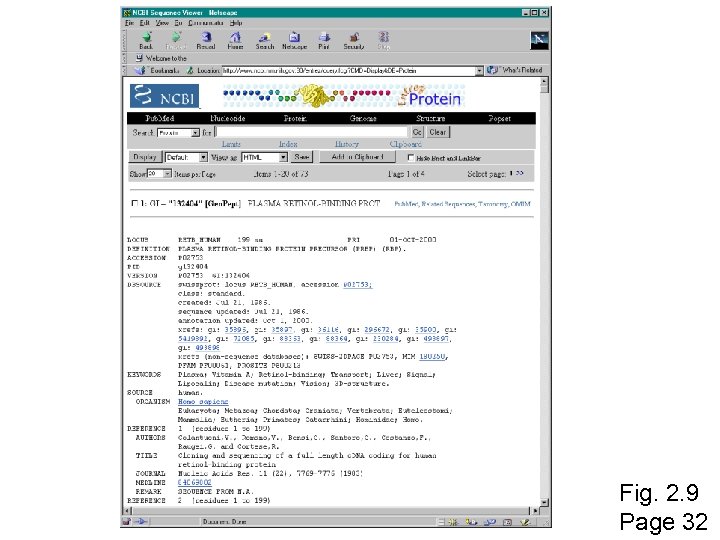
Fig. 2. 9 Page 32
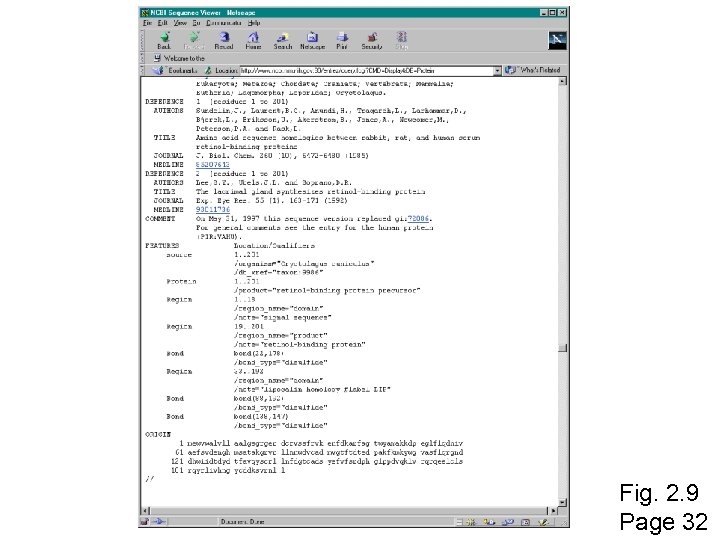
Fig. 2. 9 Page 32
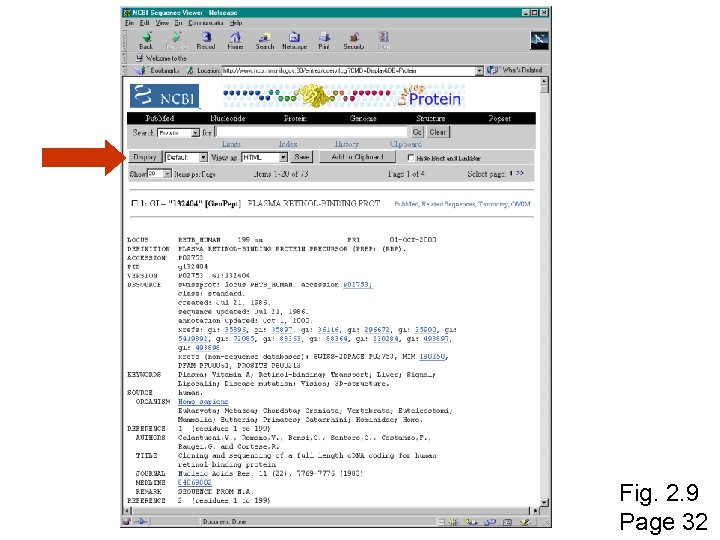
Fig. 2. 9 Page 32
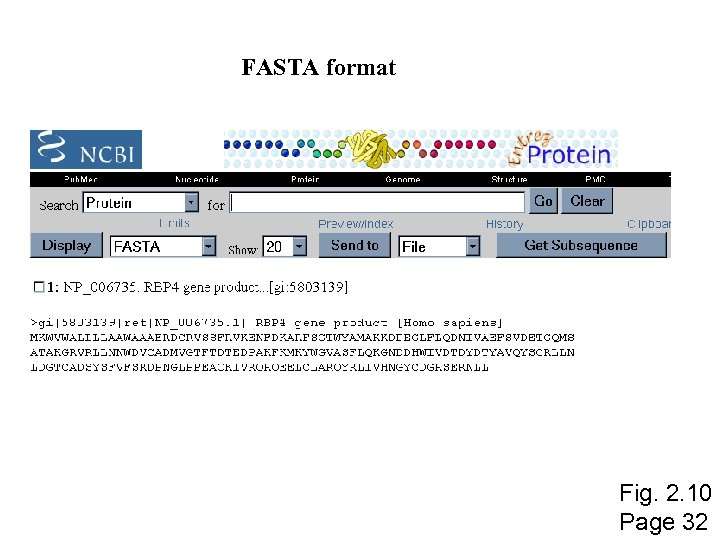
FASTA format Fig. 2. 10 Page 32
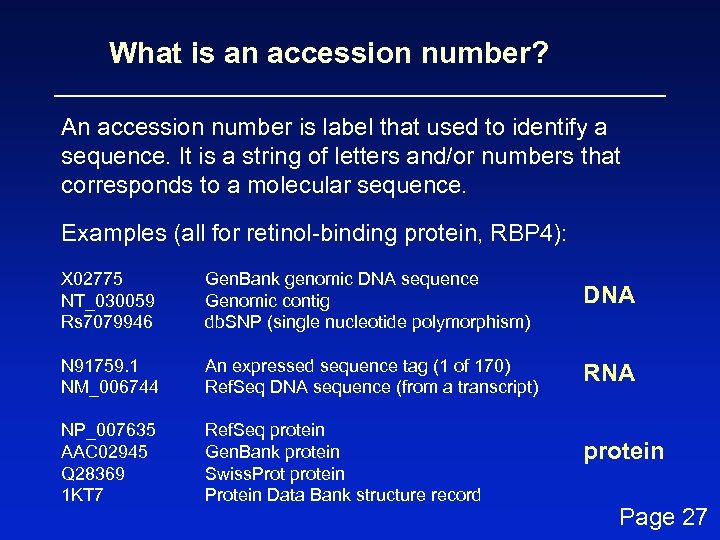

What is an accession number? An accession number is label that used to identify a sequence. It is a string of letters and/or numbers that corresponds to a molecular sequence. Examples (all for retinol-binding protein, RBP 4): X 02775 NT_030059 Rs 7079946 Gen. Bank genomic DNA sequence Genomic contig db. SNP (single nucleotide polymorphism) DNA N 91759. 1 NM_006744 An expressed sequence tag (1 of 170) Ref. Seq DNA sequence (from a transcript) RNA NP_007635 AAC 02945 Q 28369 1 KT 7 Ref. Seq protein Gen. Bank protein Swiss. Prot protein Protein Data Bank structure record protein Page 27
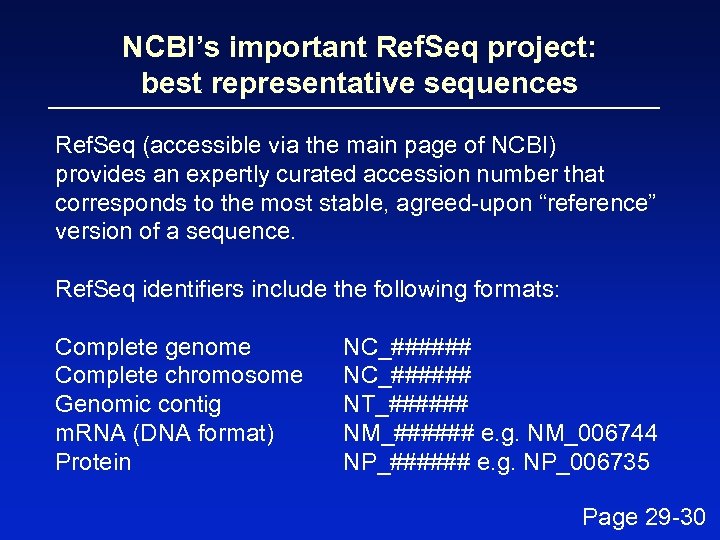

NCBI’s important Ref. Seq project: best representative sequences Ref. Seq (accessible via the main page of NCBI) provides an expertly curated accession number that corresponds to the most stable, agreed-upon “reference” version of a sequence. Ref. Seq identifiers include the following formats: Complete genome Complete chromosome Genomic contig m. RNA (DNA format) Protein NC_###### NT_###### NM_###### e. g. NM_006744 NP_###### e. g. NP_006735 Page 29 -30
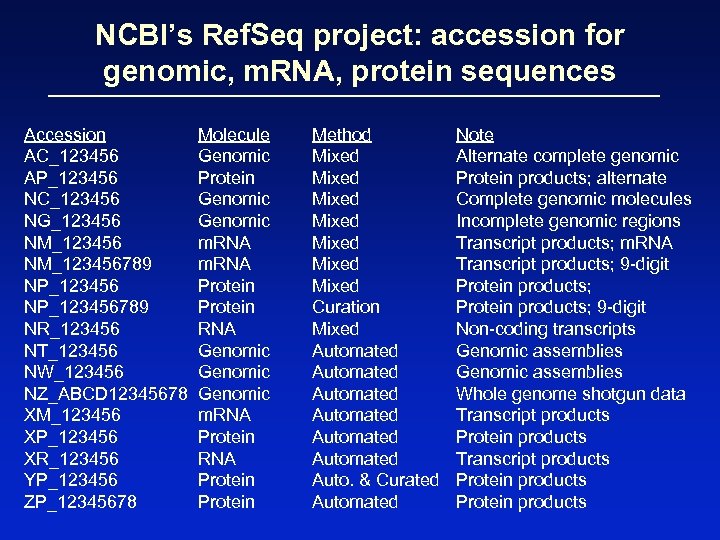

NCBI’s Ref. Seq project: accession for genomic, m. RNA, protein sequences Accession AC_123456 AP_123456 NC_123456 NG_123456 NM_123456789 NP_123456789 NR_123456 NT_123456 NW_123456 NZ_ABCD 12345678 XM_123456 XP_123456 XR_123456 YP_123456 ZP_12345678 Molecule Genomic Protein Genomic m. RNA Protein RNA Genomic m. RNA Protein Method Mixed Mixed Curation Mixed Automated Automated Auto. & Curated Automated Note Alternate complete genomic Protein products; alternate Complete genomic molecules Incomplete genomic regions Transcript products; m. RNA Transcript products; 9 -digit Protein products; 9 -digit Non-coding transcripts Genomic assemblies Whole genome shotgun data Transcript products Protein products
![Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq](https://present5.com/presentation/7029379b2bab49f2b8a65f84a5f54198/image-38.jpg)
Four ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq [2] Uni. Gene [3] European Bioinformatics Institute (EBI) and Ensembl (separate from NCBI) [4] Ex. PASy Sequence Retrieval System (separate from NCBI) Page 31
DNA RNA protein complementary DNA (c. DNA) Uni. Gene Fig. 2. 3 Page 23

Uni. Gene: unique genes via ESTs • Find Uni. Gene at NCBI: www. ncbi. nlm. nih. gov/Uni. Gene • Uni. Gene clusters contain many expressed sequence tags (ESTs), which are DNA sequences (typically 500 base pairs in length) corresponding to the m. RNA from an expressed gene. ESTs are sequenced from a complementary DNA (c. DNA) library. • Uni. Gene data come from many c. DNA libraries. Thus, when you look up a gene in Uni. Gene you get information on its abundance and its regional distribution. Pages 20 -21
Cluster sizes in Uni. Gene This is a gene with 1 EST associated; the cluster size is 1 Fig. 2. 3 Page 23
Cluster sizes in Uni. Gene This is a gene with 10 ESTs associated; the cluster size is 10
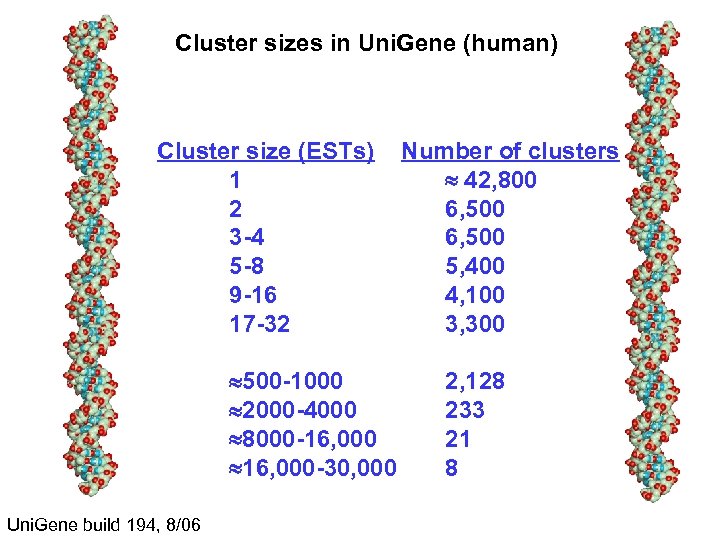

Cluster sizes in Uni. Gene (human) Cluster size (ESTs) 1 2 3 -4 5 -8 9 -16 17 -32 500 -1000 2000 -4000 8000 -16, 000 -30, 000 Uni. Gene build 194, 8/06 Number of clusters 42, 800 6, 500 5, 400 4, 100 3, 300 2, 128 233 21 8

Uni. Gene: unique genes via ESTs Conclusion: Uni. Gene is a useful tool to look up information about expressed genes. Uni. Gene displays information about the abundance of a transcript (expressed gene), as well as its regional distribution of expression (e. g. brain vs. liver). We will discuss Uni. Gene further later (gene expression). Page 31
![Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq](https://present5.com/presentation/7029379b2bab49f2b8a65f84a5f54198/image-45.jpg)
Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq [2] Uni. Gene [3] European Bioinformatics Institute (EBI) and Ensembl (separate from NCBI) [4] Ex. PASy Sequence Retrieval System (separate from NCBI) Page 31

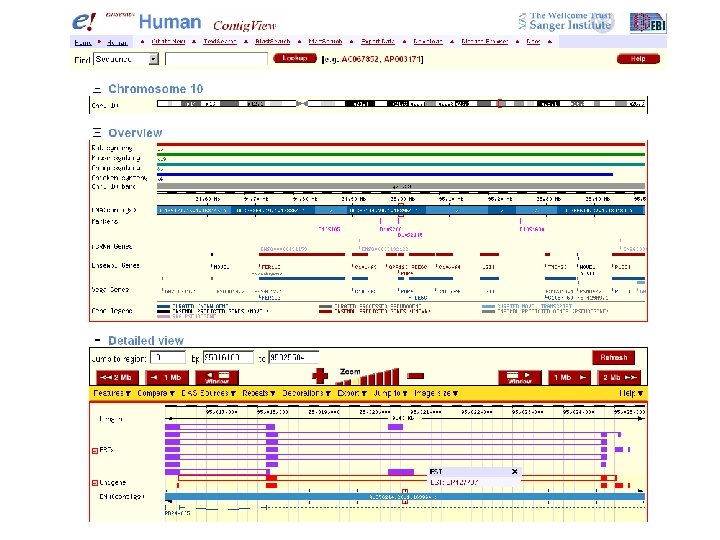
Ensembl to access protein and DNA sequences Try Ensembl at www. ensembl. org for a premier human genome web browser. We will encounter Ensembl as we study the human genome, BLAST, and other topics.
click human
enter RBP 4
![Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq](https://present5.com/presentation/7029379b2bab49f2b8a65f84a5f54198/image-50.jpg)
Five ways to access DNA and protein sequences [1] Entrez Gene with Ref. Seq [2] Uni. Gene [3] European Bioinformatics Institute (EBI) and Ensembl (separate from NCBI) [4] Ex. PASy Sequence Retrieval System (separate from NCBI) Page 33

Ex. PASy to access protein and DNA sequences Ex. PASy sequence retrieval system (Ex. PASy = Expert Protein Analysis System) Visit http: //www. expasy. ch/ Page 33
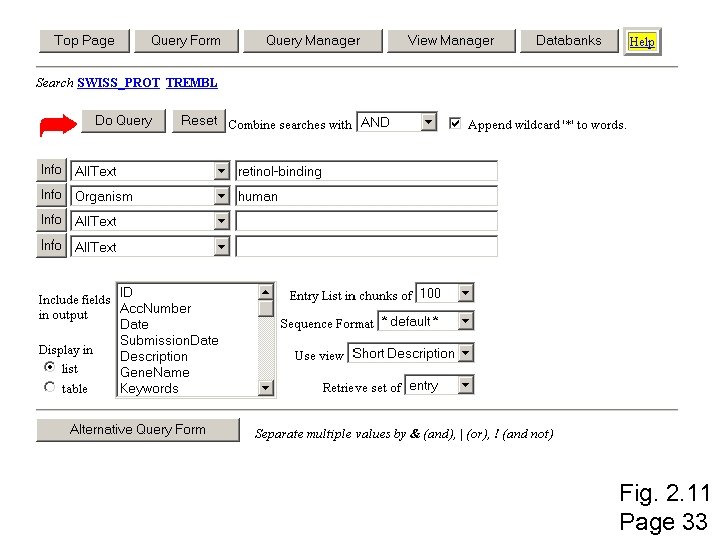
Fig. 2. 11 Page 33

Example of how to access sequence data: HIV-1 pol There are many possible approaches. Begin at the main page of NCBI, and type an Entrez query: hiv-1 pol Page 34
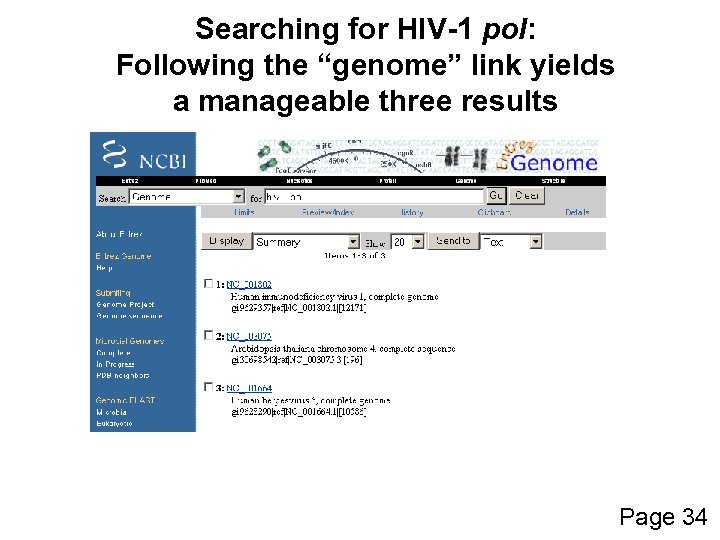
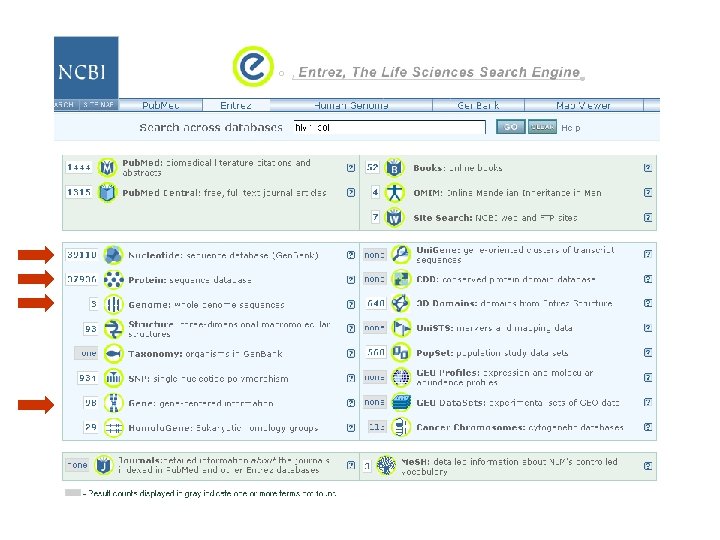

Searching for HIV-1 pol: Following the “genome” link yields a manageable three results Page 34

Example of how to access sequence data: HIV-1 pol For the Entrez query: hiv-1 pol there about 40, 000 nucleotide or protein records (and >100, 000 records for a search for “hiv-1”), but these can easily be reduced in two easy steps: --specify the organism, e. g. hiv-1[organism] --limit the output to Ref. Seq! Page 34
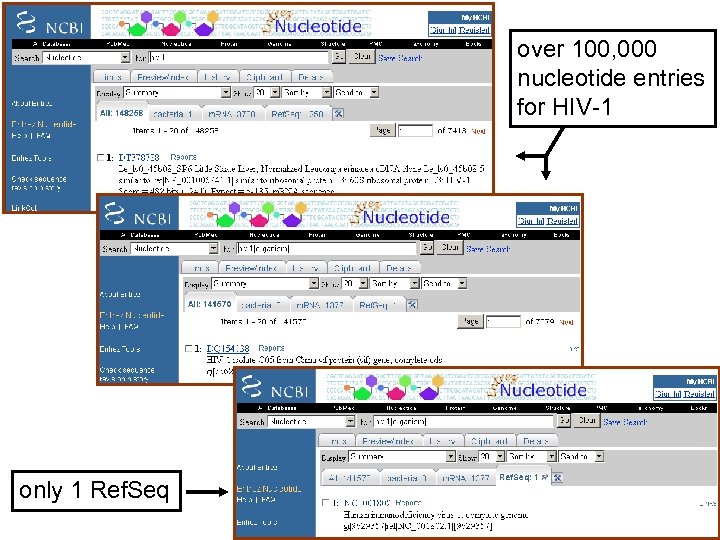
over 100, 000 nucleotide entries for HIV-1 only 1 Ref. Seq
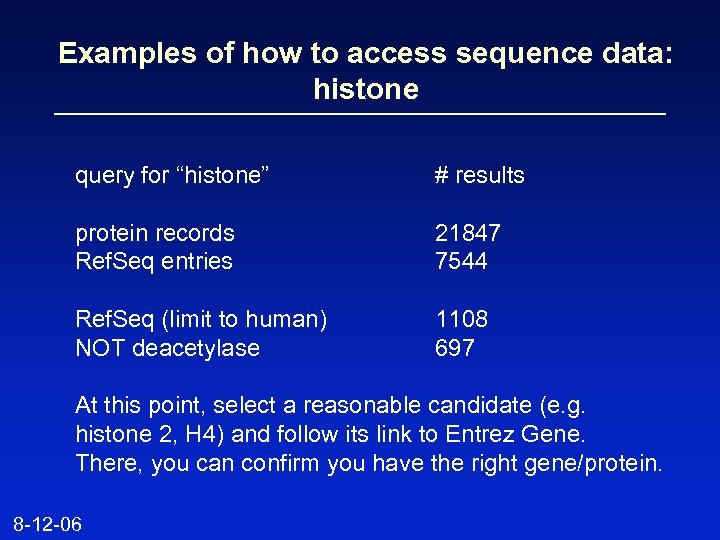

Examples of how to access sequence data: histone query for “histone” # results protein records Ref. Seq entries 21847 7544 Ref. Seq (limit to human) NOT deacetylase 1108 697 At this point, select a reasonable candidate (e. g. histone 2, H 4) and follow its link to Entrez Gene. There, you can confirm you have the right gene/protein. 8 -12 -06

Access to Biomedical Literature Page 35

Pub. Med at NCBI to find literature information

Pub. Med is the NCBI gateway to MEDLINE contains bibliographic citations and author abstracts from over 4, 600 journals published in the United States and in 70 foreign countries. It has >14 million records dating back to 1966. Page 35

Me. SH is the acronym for "Medical Subject Headings. " Me. SH is the list of the vocabulary terms used for subject analysis of biomedical literature at NLM. Me. SH vocabulary is used for indexing journal articles for MEDLINE. The Me. SH controlled vocabulary imposes uniformity and consistency to the indexing of biomedical literature. Page 35
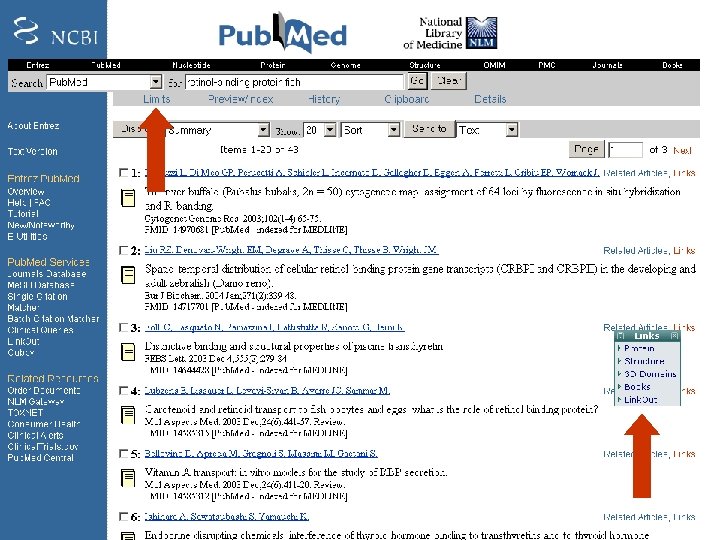

Pub. Med search strategies Try the tutorial (“education” on the left sidebar) Use boolean queries (capitalize AND, OR, NOT) lipocalin AND disease Try using “limits” Try “Links” to find Entrez information and external resources Obtain articles on-line via Welch Medical Library (and download pdf files): http: //www. welch. jhu. edu/ Page 35
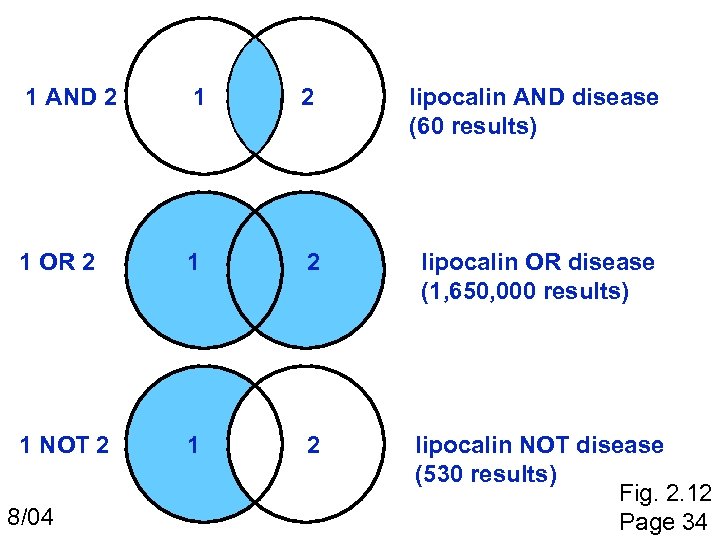
1 AND 2 1 2 lipocalin AND disease (60 results) 1 OR 2 1 2 lipocalin OR disease (1, 650, 000 results) 1 NOT 2 1 2 lipocalin NOT disease (530 results) Fig. 2. 12 Page 34 8/04
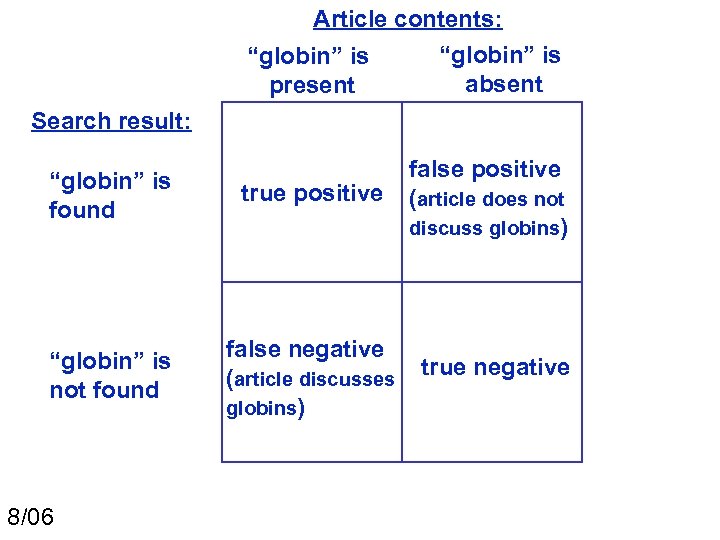

Article contents: “globin” is absent present Search result: “globin” is found true positive false positive (article does not discuss globins) “globin” is not found false negative (article discusses globins) true negative 8/06

Welch. Web is available at http: //www. welch. jhu. edu

http: //www. welch. jhu. edu Brian Brown (bbrown 20@jhmi. edu) and Carrie Iwema (iwema@jhmi. edu) are the Welch Medical Library liasons to the basic sciences
7029379b2bab49f2b8a65f84a5f54198.ppt