824458e57ca9e29b7bef4279c819443f.ppt
- Количество слайдов: 24

Modeling Dependencies in Protein-DNA Binding Sites Yoseph Barash 1 Gal Elidan 1 Nir Friedman 1 Tommy Kaplan 1, 2 1 School of Computer Science & Engineering 2 Hadassah Medical School The Hebrew University, Jerusalem, Israel
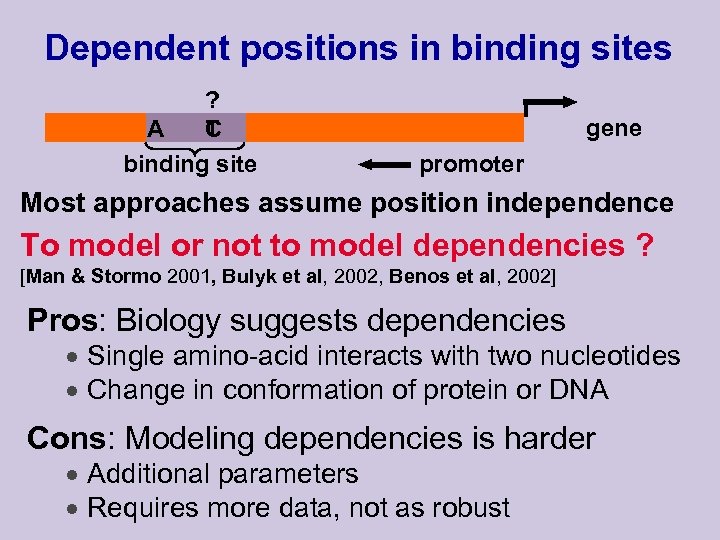
Dependent positions in binding sites ? A T C binding site gene promoter Most approaches assume position independence To model or not to model dependencies ? [Man & Stormo 2001, Bulyk et al, 2002, Benos et al, 2002] Pros: Biology suggests dependencies · Single amino-acid interacts with two nucleotides · Change in conformation of protein or DNA Cons: Modeling dependencies is harder · Additional parameters · Requires more data, not as robust

Data driven approach w w Can we learn dependencies from available genomic data ? Do dependency models perform better ? Outline w Flexible models of dependencies w Learning from (un)aligned sequences w Systematic evaluation Biological insights Yesü
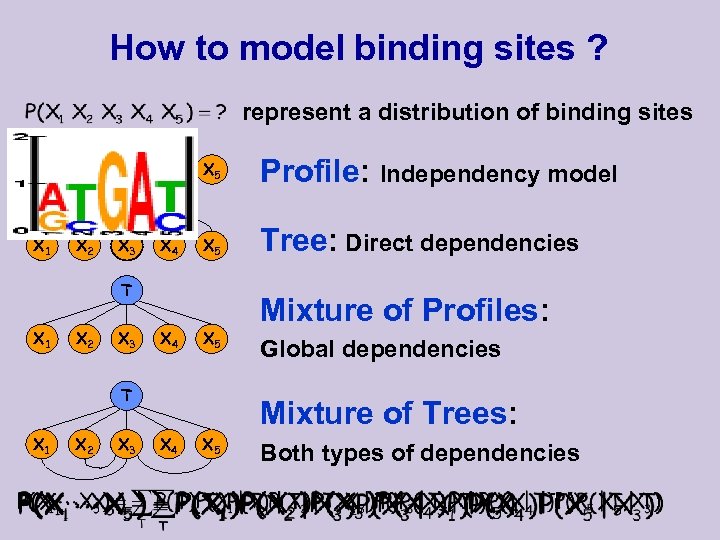
How to model binding sites ? represent a distribution of binding sites X 1 X 2 X 3 X 4 X 5 Profile: Independency model X 1 X 2 X 3 X 4 X 5 Tree: Direct dependencies T X 1 X 2 X 3 X 4 X 5 T X 1 X 2 X 3 Mixture of Profiles: Global dependencies Mixture of Trees: X 4 X 5 Both types of dependencies
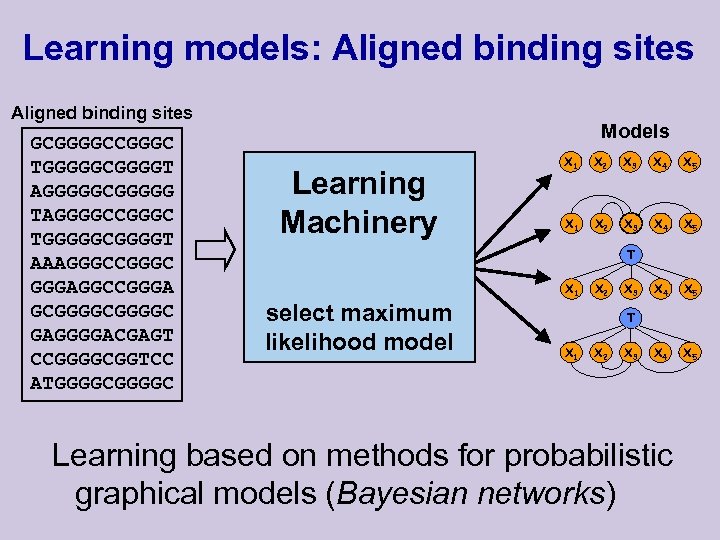
Learning models: Aligned binding sites GCGGGGCCGGGC TGGGGGCGGGGT AGGGGGCGGGGG TAGGGGCCGGGC TGGGGGCGGGGT AAAGGGCCGGGC GGGAGGCCGGGA GCGGGGC GAGGGGACGAGT CCGGGGCGGTCC ATGGGGC Models Learning Machinery X 1 X 2 X 3 X 4 X 5 T X 1 select maximum likelihood model X 2 X 3 T X 1 X 2 X 3 Learning based on methods for probabilistic graphical models (Bayesian networks)
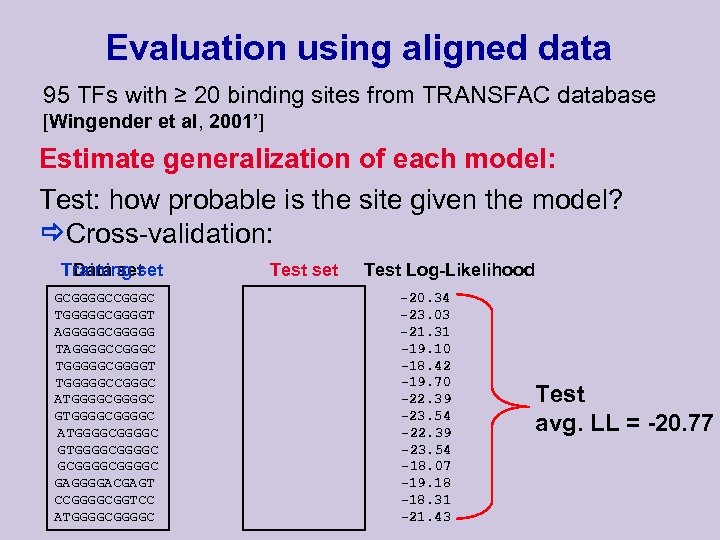

Evaluation using aligned data 95 TFs with ≥ 20 binding sites from TRANSFAC database [Wingender et al, 2001’] Estimate generalization of each model: Test: how probable is the site given the model? Cross-validation: Training set Data set GCGGGGCCGGGC TGGGGGCGGGGT AGGGGGCGGGGG TAGGGGCCGGGC TGGGGGCGGGGT TGGGGGCCGGGC ATGGGGCGGGGC GTGGGGCGGGGC GAGGGGACGAGT CCGGGGCGGTCC ATGGGGC Test set Test Log-Likelihood -20. 34 -23. 03 -21. 31 -19. 10 -18. 42 -19. 70 -22. 39 -23. 54 -18. 07 -19. 18 -18. 31 -21. 43 Test avg. LL = -20. 77
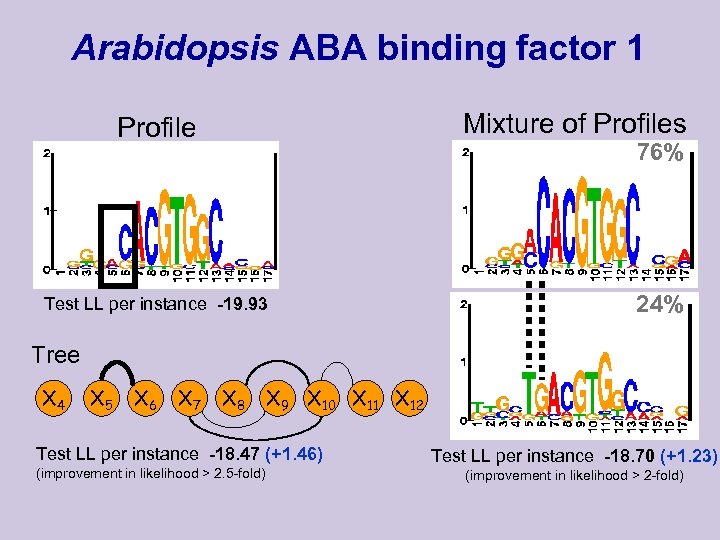
Arabidopsis ABA binding factor 1 Mixture of Profiles Profile 76% Test LL per instance -19. 93 24% Tree X 4 X 5 X 6 X 7 X 8 X 9 X 10 X 11 X 12 Test LL per instance -18. 47 (+1. 46) (improvement in likelihood > 2. 5 -fold) Test LL per instance -18. 70 (+1. 23) (improvement in likelihood > 2 -fold)
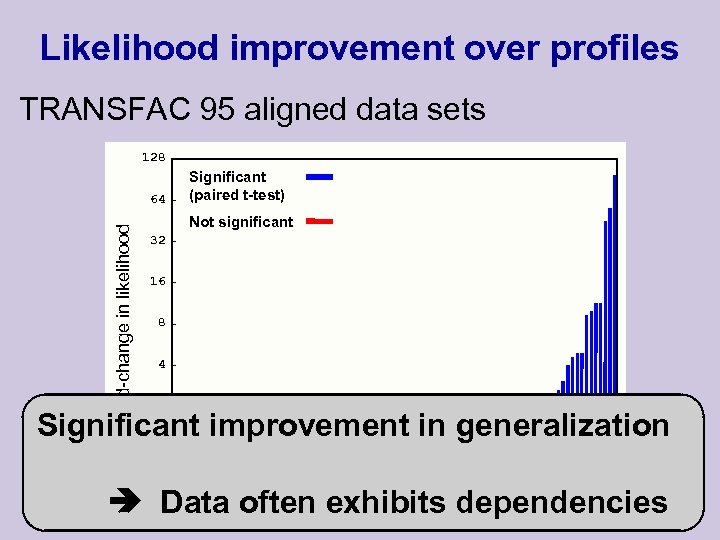
Likelihood improvement over profiles TRANSFAC 95 aligned data sets 128 Fold-change in likelihood 64 Significant (paired t-test) Not significant 32 16 8 4 2 Significant improvement in generalization 1 0. 5 10 20 30 40 50 Data often exhibits 60 70 80 90 dependencies
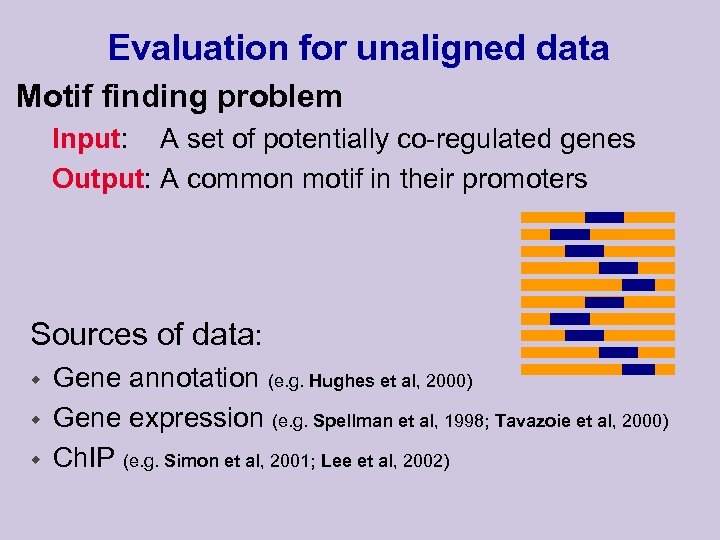

Evaluation for unaligned data Motif finding problem Input: A set of potentially co-regulated genes Output: A common motif in their promoters Sources of data: w w w Gene annotation (e. g. Hughes et al, 2000) Gene expression (e. g. Spellman et al, 1998; Tavazoie et al, 2000) Ch. IP (e. g. Simon et al, 2001; Lee et al, 2002)
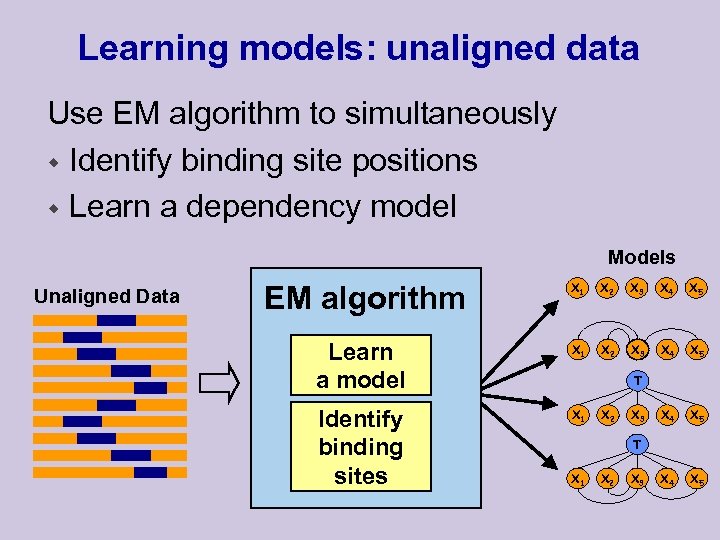
Learning models: unaligned data Use EM algorithm to simultaneously w Identify binding site positions w Learn a dependency model Models Unaligned Data X 1 X 2 X 3 X 4 X 5 Learn a model X 1 X 2 X 3 X 4 X 5 Identify binding sites X 1 X 4 X 5 EM algorithm T X 2 X 3 T X 1 X 2 X 3
![Ch. IP location analysis [Lee et al, 2002] Yeast genome-wide location experiments Target genes Ch. IP location analysis [Lee et al, 2002] Yeast genome-wide location experiments Target genes](https://present5.com/presentation/824458e57ca9e29b7bef4279c819443f/image-11.jpg)
Ch. IP location analysis [Lee et al, 2002] Yeast genome-wide location experiments Target genes for 106 TFs in 146 experiments Gene # genes ~ 6000 YAL 001 C YAL 002 W YAL 003 W YAL 005 C. . . YAL 010 C YAL 012 C YAL 013 W YPR 201 W ABF 1 Targets + –. . . + – – – …. . ZAP 1 Targets – + – –. . . – + + –
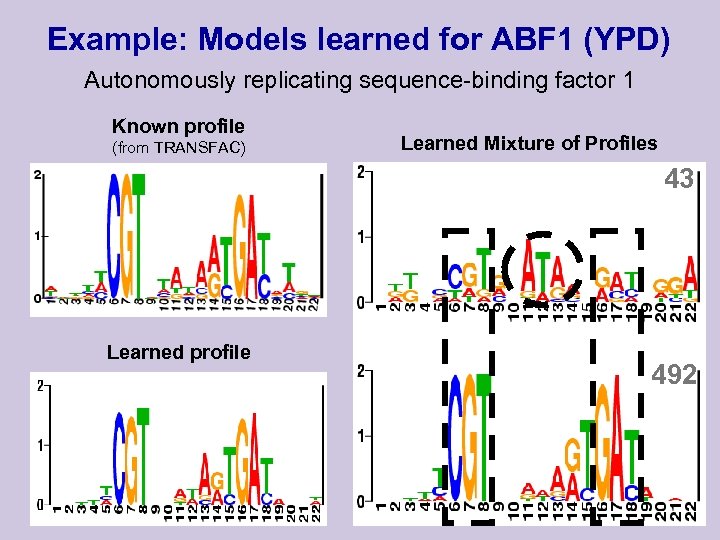
Example: Models learned for ABF 1 (YPD) Autonomously replicating sequence-binding factor 1 Known profile (from TRANSFAC) Learned Mixture of Profiles 43 Learned profile 492
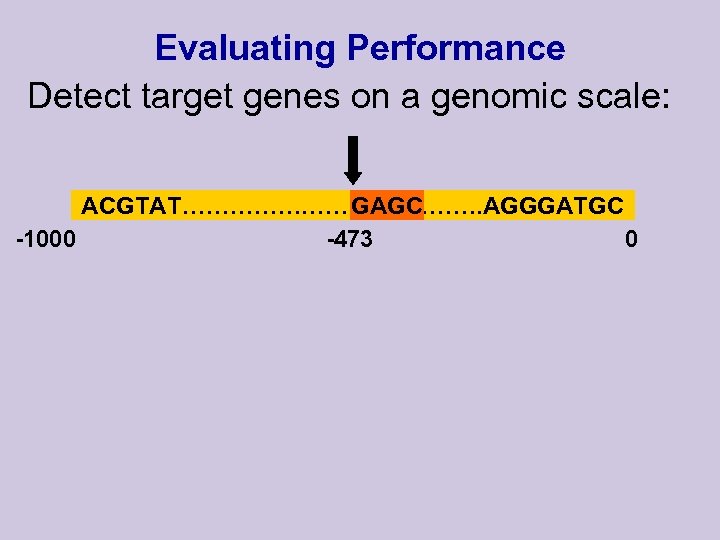
Evaluating Performance Detect target genes on a genomic scale: ACGTAT…………………. AGGGATGC GAGC -473 -1000 0
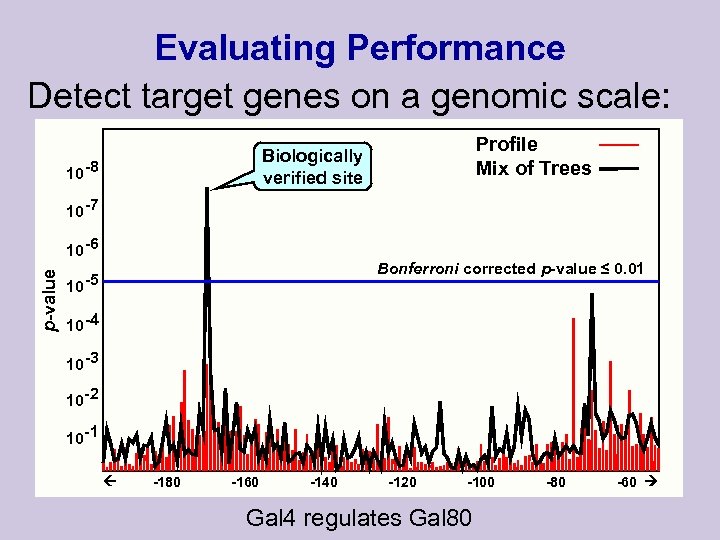
Evaluating Performance Detect target genes on a genomic scale: Profile Mix of Trees Biologically verified site 10 -8 10 -7 p-value 10 -6 Bonferroni corrected p-value ≤ 0. 01 10 -5 10 -4 10 -3 10 -2 10 -1 -180 -160 -140 -120 -100 Gal 4 regulates Gal 80 -60
![Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5 Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5](https://present5.com/presentation/824458e57ca9e29b7bef4279c819443f/image-15.jpg)
Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5 -fold cross-validation test: Data set + – + – – – YAL 001 C YAL 002 W YAL 003 W YAL 005 C YAL 007 C YAL 008 W YAL 009 W YAL 010 C YAL 012 C YAL 013 W YPR 201 W Test set Prediction + – +
![Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5 Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5](https://present5.com/presentation/824458e57ca9e29b7bef4279c819443f/image-16.jpg)
Evaluation using Ch. IP location data [Lee et al, 2002] Evaluate using a 5 -fold cross-validation test: Data set YAL 001 C YAL 002 W YAL 003 W YAL 005 C YAL 007 C YAL 008 W YAL 009 W YAL 010 C YAL 012 C YAL 013 W YPR 201 W Prediction True + – – – – + + – – + – + – – – √ √ FN √ √ √ FP √ √
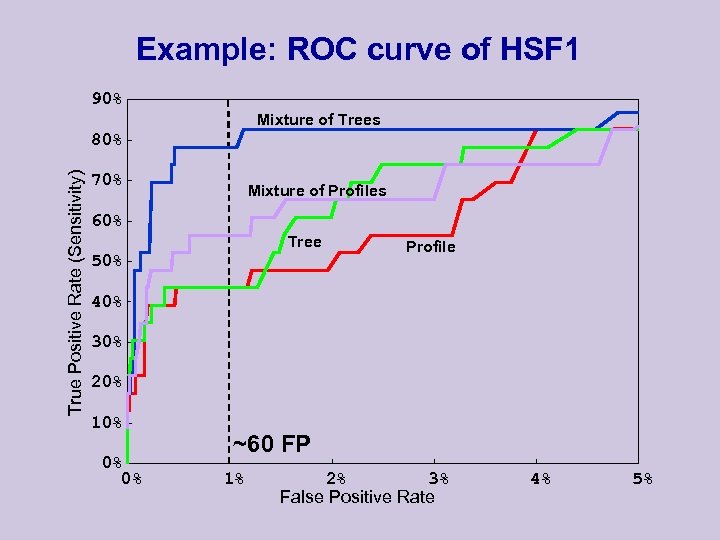
Example: ROC curve of HSF 1 90% Mixture of Trees True Positive Rate (Sensitivity) 80% 70% Mixture of Profiles 60% Tree 50% Profile 40% 30% 20% 10% 0% 0% ~60 FP 1% 2% 3% False Positive Rate 4% 5%
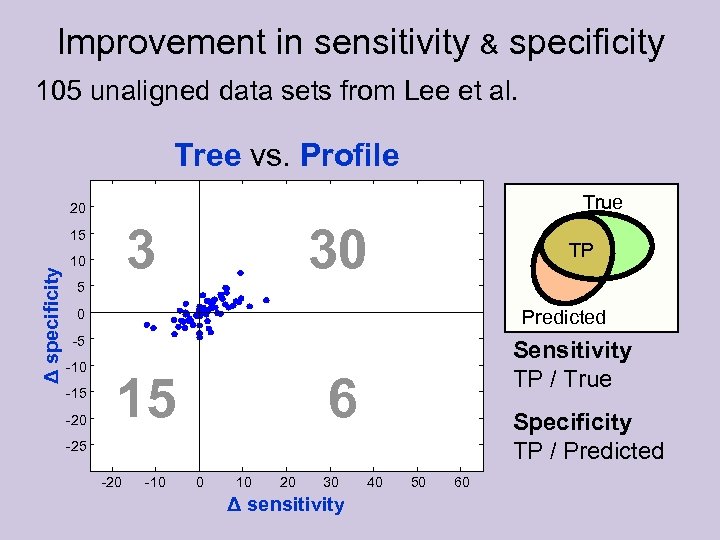
Improvement in sensitivity & specificity 105 unaligned data sets from Lee et al. Tree vs. Profile True 20 3 Δ specificity 15 10 30 TP 5 Predicted 0 -5 -10 -15 -20 15 Sensitivity TP / True 6 Specificity TP / Predicted -25 -20 -10 0 10 20 30 Δ sensitivity 40 50 60
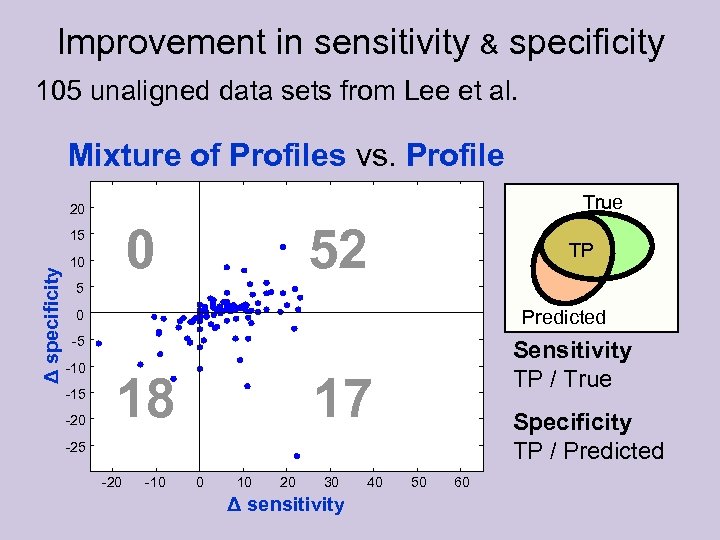
Improvement in sensitivity & specificity 105 unaligned data sets from Lee et al. Mixture of Profiles vs. Profile True 20 0 Δ specificity 15 10 52 TP 5 Predicted 0 -5 -10 -15 -20 18 Sensitivity TP / True 17 Specificity TP / Predicted -25 -20 -10 0 10 20 30 Δ sensitivity 40 50 60
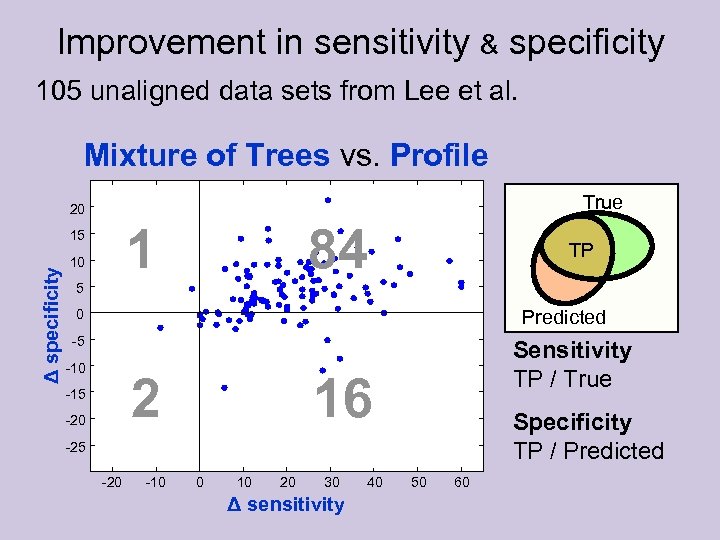
Improvement in sensitivity & specificity 105 unaligned data sets from Lee et al. Mixture of Trees vs. Profile True 20 1 Δ specificity 15 10 84 TP 5 Predicted 0 -5 -10 2 -15 -20 Sensitivity TP / True 16 Specificity TP / Predicted -25 -20 -10 0 10 20 30 Δ sensitivity 40 50 60

“Is it worthwhile to model dependencies? ” Evaluation clearly supports this What about the underlying biology ? (with Prof. Hanah Margalit, Hadassah Medical School)
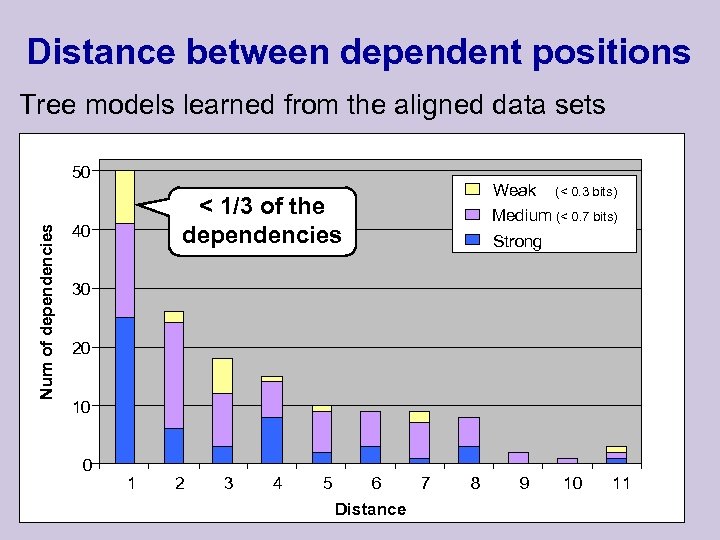
Distance between dependent positions Tree models learned from the aligned data sets Num of dependencies 50 Weak (< 0. 3 bits) Medium (< 0. 7 bits) Strong < 1/3 of the dependencies 40 30 20 10 0 1 2 3 4 5 6 Distance 7 8 9 10 11
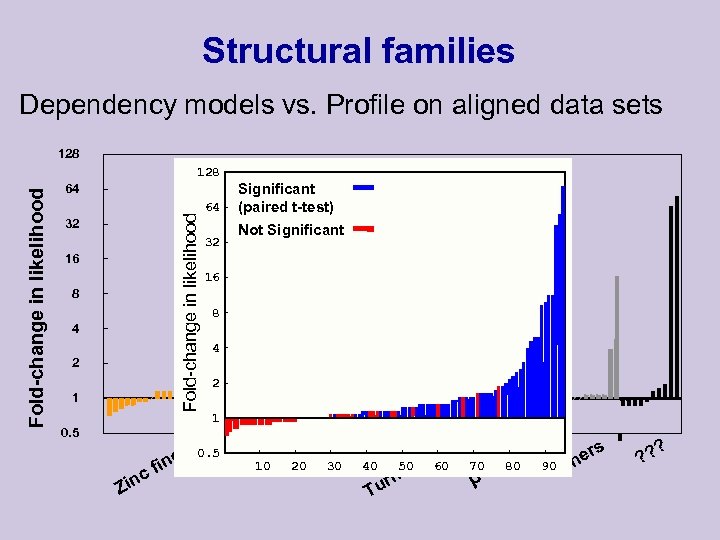
Structural families Dependency models vs. Profile on aligned data sets 128 64 Fold-change in likelihood 128 32 16 8 4 2 1 0. 5 c Zin 64 32 Significant (paired t-test) Not Significant 16 8 4 2 1 r P L ge 0. 5 b. ZI 10 H 20 H b fin 30 ix x el 50 eli 60 H H 40 urn T t 70 βS e 80 he rs e 90 oth ? ? ?

Conclusions Flexible framework for learning dependencies üDependencies are found in many cases üIt is worthwhile to model them Better learning and binding site prediction w Future work w w Link to the underlying structural biology Incorporate as part of other regulatory mechanism models http: //compbio. cs. huji. ac. il/TFBN
824458e57ca9e29b7bef4279c819443f.ppt