d5716841ce6f0f85826a663372494191.ppt
- Количество слайдов: 107
 Microarrays & Expression profiling Matthias E. Futschik Institute for Theoretical Biology Humboldt-University, Berlin, Germany Bioinformatics Summer School, Istanbul-Sile, 2004
Microarrays & Expression profiling Matthias E. Futschik Institute for Theoretical Biology Humboldt-University, Berlin, Germany Bioinformatics Summer School, Istanbul-Sile, 2004
 Yeast c. DNA microarray
Yeast c. DNA microarray
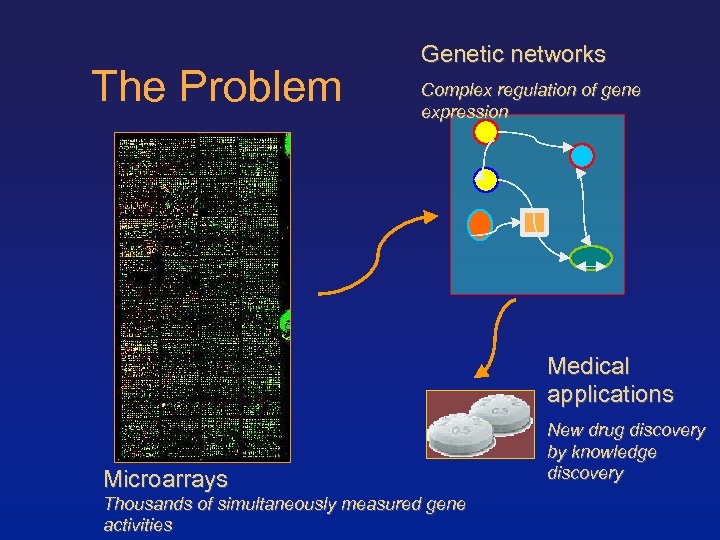 The Problem Genetic networks Complex regulation of gene expression Medical applications Microarrays Thousands of simultaneously measured gene activities New drug discovery by knowledge discovery
The Problem Genetic networks Complex regulation of gene expression Medical applications Microarrays Thousands of simultaneously measured gene activities New drug discovery by knowledge discovery
 Methodology 1. 2. 3. 4. 5. 6. 7. From Image to Numbers: Image-Analysis Well begun is half done: Design of experiments Cleaning up: Preprocessing and Normalisation Go fishing: Significance of Differences in Gene Expression Data Who is with whom: Clustering of samples and genes Gattica becomes alive: Classification and Disease Profiling The whole picture: An integrative approach
Methodology 1. 2. 3. 4. 5. 6. 7. From Image to Numbers: Image-Analysis Well begun is half done: Design of experiments Cleaning up: Preprocessing and Normalisation Go fishing: Significance of Differences in Gene Expression Data Who is with whom: Clustering of samples and genes Gattica becomes alive: Classification and Disease Profiling The whole picture: An integrative approach
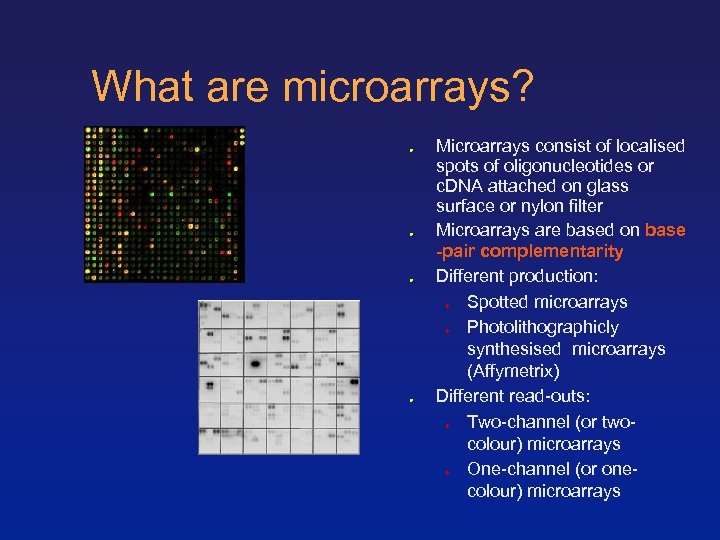 What are microarrays? Microarrays consist of localised spots of oligonucleotides or c. DNA attached on glass surface or nylon filter Microarrays are based on base -pair complementarity Different production: Spotted microarrays Photolithographicly synthesised microarrays (Affymetrix) Different read-outs: Two-channel (or twocolour) microarrays One-channel (or onecolour) microarrays
What are microarrays? Microarrays consist of localised spots of oligonucleotides or c. DNA attached on glass surface or nylon filter Microarrays are based on base -pair complementarity Different production: Spotted microarrays Photolithographicly synthesised microarrays (Affymetrix) Different read-outs: Two-channel (or twocolour) microarrays One-channel (or onecolour) microarrays
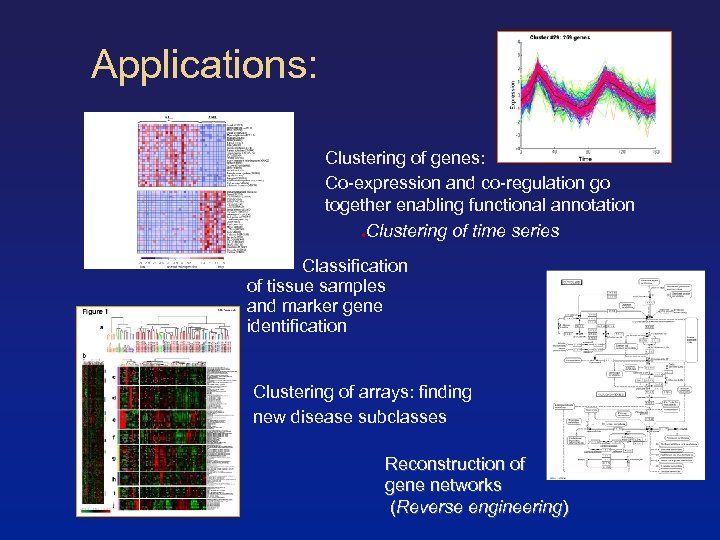 Applications: Clustering of genes: Co-expression and co-regulation go together enabling functional annotation Clustering of time series Classification of tissue samples and marker gene identification Clustering of arrays: finding new disease subclasses Reconstruction of gene networks (Reverse engineering)
Applications: Clustering of genes: Co-expression and co-regulation go together enabling functional annotation Clustering of time series Classification of tissue samples and marker gene identification Clustering of arrays: finding new disease subclasses Reconstruction of gene networks (Reverse engineering)
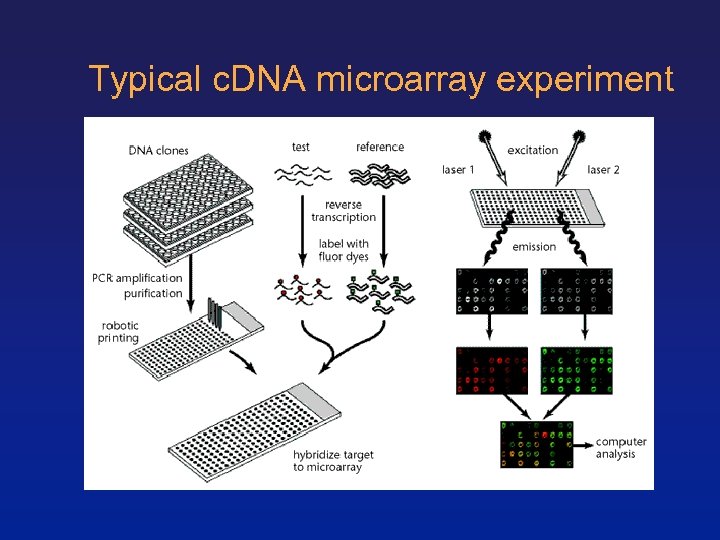 Typical c. DNA microarray experiment
Typical c. DNA microarray experiment
 Microarray technology I Two-colour microarray (c. DNA and spotted oligonucleotide microarrays) Probes are PCR products based on a chosen c. DNA library or synthesized oligonucleotides (length 50 -70) optimized for specifity and binding properties >> probe design Probes are mechanically spotted. To control variation of amount of printed c. DNA/oligos and spot morphology, reference RNA sample is included. Thus, ratios are considered as basic units for analysing gene expression. Absolute intensities should be interpreted with care. MOVIE 1: Array production - Galbraith lab MOVIE 2: Principles – Schreiber lab
Microarray technology I Two-colour microarray (c. DNA and spotted oligonucleotide microarrays) Probes are PCR products based on a chosen c. DNA library or synthesized oligonucleotides (length 50 -70) optimized for specifity and binding properties >> probe design Probes are mechanically spotted. To control variation of amount of printed c. DNA/oligos and spot morphology, reference RNA sample is included. Thus, ratios are considered as basic units for analysing gene expression. Absolute intensities should be interpreted with care. MOVIE 1: Array production - Galbraith lab MOVIE 2: Principles – Schreiber lab
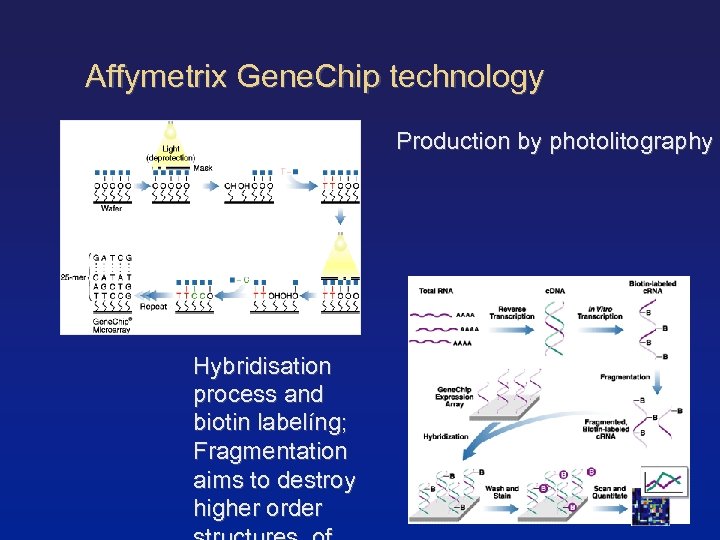 Affymetrix Gene. Chip technology Production by photolitography Hybridisation process and biotin labelíng; Fragmentation aims to destroy higher order
Affymetrix Gene. Chip technology Production by photolitography Hybridisation process and biotin labelíng; Fragmentation aims to destroy higher order
 Microarray technologies II One-colour microarrays (Affymetrix Gene. Chips) Measurement of hybridisation of target RNA to sets of 25 oligonucleotides (probes). Probes are paired: Perfect match (PM) and mis-match (MM). PM are complementary to the gene sequence of interest. MM include a single nucleotide changed in the middle position of the oligonucleotide. MM serve for controlling of experimental variation and non-specific cross-hybridisation. Thus, MMs constitute internal references (on the probe site). Average (PM-MM) delivers measure for gene expression. However, different methods to calculates summary indices exist (e. g. MAS, dchip, RMA. . . )
Microarray technologies II One-colour microarrays (Affymetrix Gene. Chips) Measurement of hybridisation of target RNA to sets of 25 oligonucleotides (probes). Probes are paired: Perfect match (PM) and mis-match (MM). PM are complementary to the gene sequence of interest. MM include a single nucleotide changed in the middle position of the oligonucleotide. MM serve for controlling of experimental variation and non-specific cross-hybridisation. Thus, MMs constitute internal references (on the probe site). Average (PM-MM) delivers measure for gene expression. However, different methods to calculates summary indices exist (e. g. MAS, dchip, RMA. . . )
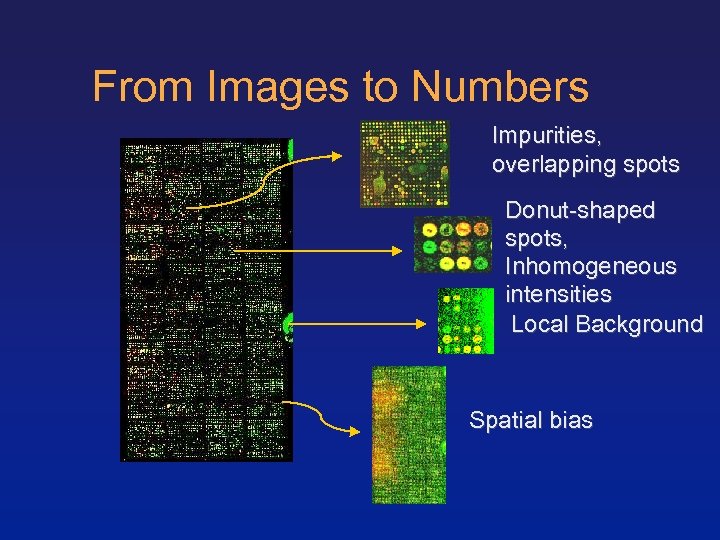 From Images to Numbers Impurities, overlapping spots Donut-shaped spots, Inhomogeneous intensities Local Background Spatial bias
From Images to Numbers Impurities, overlapping spots Donut-shaped spots, Inhomogeneous intensities Local Background Spatial bias
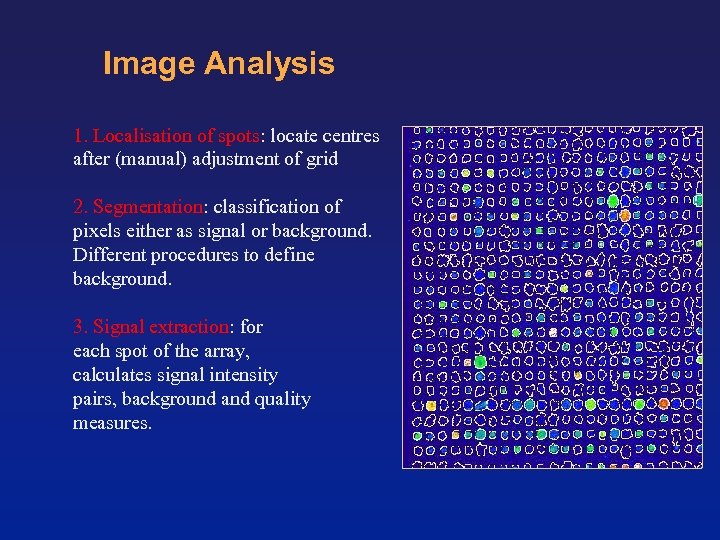 Image Analysis 1. Localisation of spots: locate centres after (manual) adjustment of grid 2. Segmentation: classification of pixels either as signal or background. Different procedures to define background. 3. Signal extraction: for each spot of the array, calculates signal intensity pairs, background and quality measures.
Image Analysis 1. Localisation of spots: locate centres after (manual) adjustment of grid 2. Segmentation: classification of pixels either as signal or background. Different procedures to define background. 3. Signal extraction: for each spot of the array, calculates signal intensity pairs, background and quality measures.
 Data acquisition Scans of slides are usually stored in 16 -bit TIFF files. Thus, scanned intensities vary between 0 and 216. Scanning of separate channels can adjusted by selection of laser power and gain of photo-multiplier. Common aim: balancing of channels. Common problems: avoiding of saturation of high intensity spots while increasing signal to noise ratios.
Data acquisition Scans of slides are usually stored in 16 -bit TIFF files. Thus, scanned intensities vary between 0 and 216. Scanning of separate channels can adjusted by selection of laser power and gain of photo-multiplier. Common aim: balancing of channels. Common problems: avoiding of saturation of high intensity spots while increasing signal to noise ratios.
 Data acquisition Image processing software produces a variety of measures: Spot intensities, local background, spot morphology measures. Software vary in computational approaches of image segmentation and read-out. Open issues: local background correction derivation of ratios for spot intensities flagging of spots, multiple scanning procedures
Data acquisition Image processing software produces a variety of measures: Spot intensities, local background, spot morphology measures. Software vary in computational approaches of image segmentation and read-out. Open issues: local background correction derivation of ratios for spot intensities flagging of spots, multiple scanning procedures
 Design of experiment Two channel microarrays incorporate a reference sample. Choice of reference determines follow-up analysis. A 1 A 2 Reference design: All samples are co-hybridised with common reference sample ● Advantage: Robust and scalable. Length of path of direct comparison equals 2. Disadvantage: Half of the measurements are made on reference sample which is commonly of little or no interest ● ● R A 3
Design of experiment Two channel microarrays incorporate a reference sample. Choice of reference determines follow-up analysis. A 1 A 2 Reference design: All samples are co-hybridised with common reference sample ● Advantage: Robust and scalable. Length of path of direct comparison equals 2. Disadvantage: Half of the measurements are made on reference sample which is commonly of little or no interest ● ● R A 3
 Alternative Designs: Dye-swap design: each comparison includes ● dye-swap to distinguish dye effects from differential expression (important for direct labelling method) Loop-design: No reference sample is involved. Increase of efficiency is, however, accompanied with a decrease of robustness. A 1 A 2 ● Latin-square design: classical design to separate effects of different experimental factors ● A 3
Alternative Designs: Dye-swap design: each comparison includes ● dye-swap to distinguish dye effects from differential expression (important for direct labelling method) Loop-design: No reference sample is involved. Increase of efficiency is, however, accompanied with a decrease of robustness. A 1 A 2 ● Latin-square design: classical design to separate effects of different experimental factors ● A 3
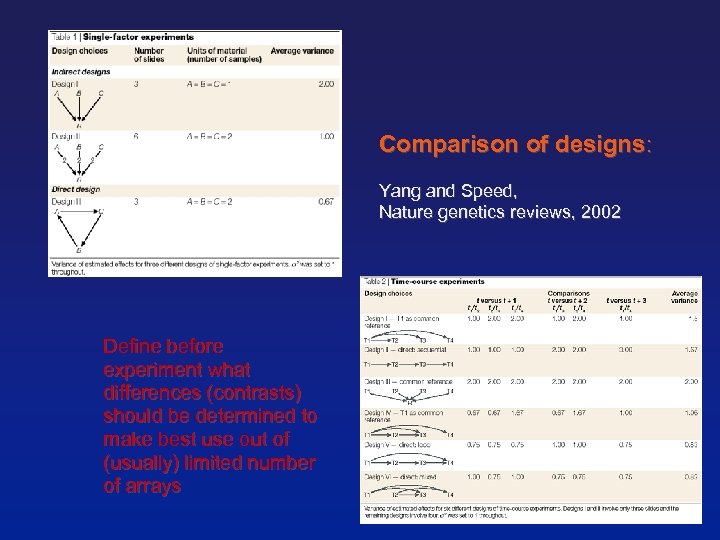 Comparison of designs: Yang and Speed, Nature genetics reviews, 2002 Define before experiment what differences (contrasts) should be determined to make best use out of (usually) limited number of arrays
Comparison of designs: Yang and Speed, Nature genetics reviews, 2002 Define before experiment what differences (contrasts) should be determined to make best use out of (usually) limited number of arrays
 Sources of variation in gene expression measurements using microarrays ● 1. ● Microarray platform Manufacturing or spotting process 1. Manufacturing batch 2. Amplification by PCR and purification 3. Amount of c. DNA spotted, morphology of spot and binding of c. DNA to substrate m. RNA extraction and preparation 1. Protocol of m. RNA extraction and amplification 2. Labelling of m. RNA
Sources of variation in gene expression measurements using microarrays ● 1. ● Microarray platform Manufacturing or spotting process 1. Manufacturing batch 2. Amplification by PCR and purification 3. Amount of c. DNA spotted, morphology of spot and binding of c. DNA to substrate m. RNA extraction and preparation 1. Protocol of m. RNA extraction and amplification 2. Labelling of m. RNA
 Sources of variation in gene expression measurements using microarrays II 1. ● 1. Hybridisation 1. Hybridisation conditions such as temperature, humidity, hyb-buffer Scanning ● scanner ● scanning intensity and PMT settings Imaging 1. software 2. flagging, background correction, . . .
Sources of variation in gene expression measurements using microarrays II 1. ● 1. Hybridisation 1. Hybridisation conditions such as temperature, humidity, hyb-buffer Scanning ● scanner ● scanning intensity and PMT settings Imaging 1. software 2. flagging, background correction, . . .
 Design of experiment Important issues for DOE: Technical replicates assess variability induced by experimental procedures. ● ● Biological replicates (assess generality of results). Number of replicates depends on desired sensitivity and sensibility of measurements and research goal. ● Randomisation to avoid confounding of experimental factors. Blocking to reduce number of experimental factors. ●
Design of experiment Important issues for DOE: Technical replicates assess variability induced by experimental procedures. ● ● Biological replicates (assess generality of results). Number of replicates depends on desired sensitivity and sensibility of measurements and research goal. ● Randomisation to avoid confounding of experimental factors. Blocking to reduce number of experimental factors. ●
 Design of experiment Control spots ● assess reproducibility within and between array, background intensity, cross-hybridisation and/or sensitivity of measurement can consists of empty spots or hybridisation-buffer, genomic DNA, foreign DNA, house-holding genes ● ● Foreign (non-cross-hybridising) c. DNA can be 'spiked in'. Use of dilution series can assess sensitivity of detecting differential expression by 'ratio controls'. ● Validation of results: by other experimental techniques (e. g. Northern, RT-PCR) ● by comparison with independent experiments. ●
Design of experiment Control spots ● assess reproducibility within and between array, background intensity, cross-hybridisation and/or sensitivity of measurement can consists of empty spots or hybridisation-buffer, genomic DNA, foreign DNA, house-holding genes ● ● Foreign (non-cross-hybridising) c. DNA can be 'spiked in'. Use of dilution series can assess sensitivity of detecting differential expression by 'ratio controls'. ● Validation of results: by other experimental techniques (e. g. Northern, RT-PCR) ● by comparison with independent experiments. ●
 Data storage Microarrays experiments produce large amounts of data: data storage and accessibility are of major importance for the follow-up analysis. Not only signal values have to be stored but also: TIFF images and imaging read-out ● Gene annotation Experimental protocol Information about samples Results of pre-processing, normalization and further analysis ● ●
Data storage Microarrays experiments produce large amounts of data: data storage and accessibility are of major importance for the follow-up analysis. Not only signal values have to be stored but also: TIFF images and imaging read-out ● Gene annotation Experimental protocol Information about samples Results of pre-processing, normalization and further analysis ● ●
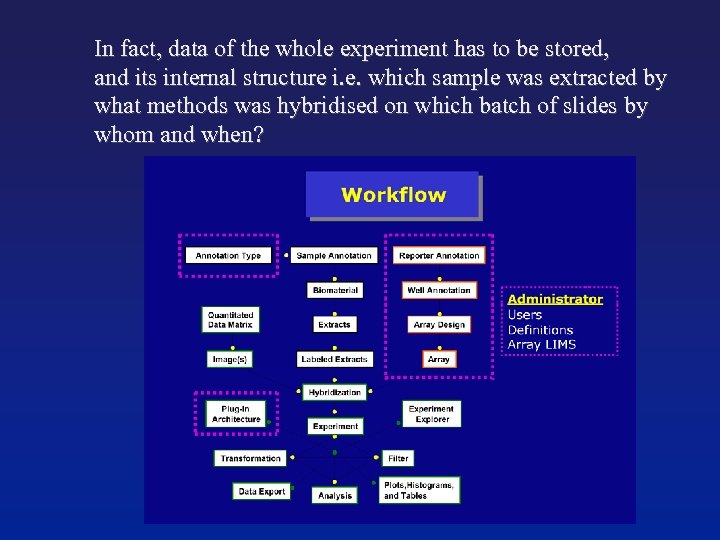 In fact, data of the whole experiment has to be stored, and its internal structure i. e. which sample was extracted by what methods was hybridised on which batch of slides by whom and when?
In fact, data of the whole experiment has to be stored, and its internal structure i. e. which sample was extracted by what methods was hybridised on which batch of slides by whom and when?
 Type of storage ● Flat file: for small-scale experiments and one-off analysis. Example NCBI - Gen. Bank Database: necessary for large scale experiments. ● Microarray DBs typically relational, SQL-based models. Their internal relational structure should reflect the experiment structure. These types of databases will become essiential tools for post-genomic analysis.
Type of storage ● Flat file: for small-scale experiments and one-off analysis. Example NCBI - Gen. Bank Database: necessary for large scale experiments. ● Microarray DBs typically relational, SQL-based models. Their internal relational structure should reflect the experiment structure. These types of databases will become essiential tools for post-genomic analysis.
 Data storage II Sharing microarray data: • NCBI • EBI • Stanford • Journals ie Nature Standardization of information by MGED: ● MIAME (minimal information about a microarray experiment) ● MAGE-ML based on XML for data exchange
Data storage II Sharing microarray data: • NCBI • EBI • Stanford • Journals ie Nature Standardization of information by MGED: ● MIAME (minimal information about a microarray experiment) ● MAGE-ML based on XML for data exchange
 Take-home messages I • Remember: Microarrays are shadows of genetic networks • Watch out for experimental variation • The complexity of microarray experiments should be reflected in the structure used for data storage
Take-home messages I • Remember: Microarrays are shadows of genetic networks • Watch out for experimental variation • The complexity of microarray experiments should be reflected in the structure used for data storage
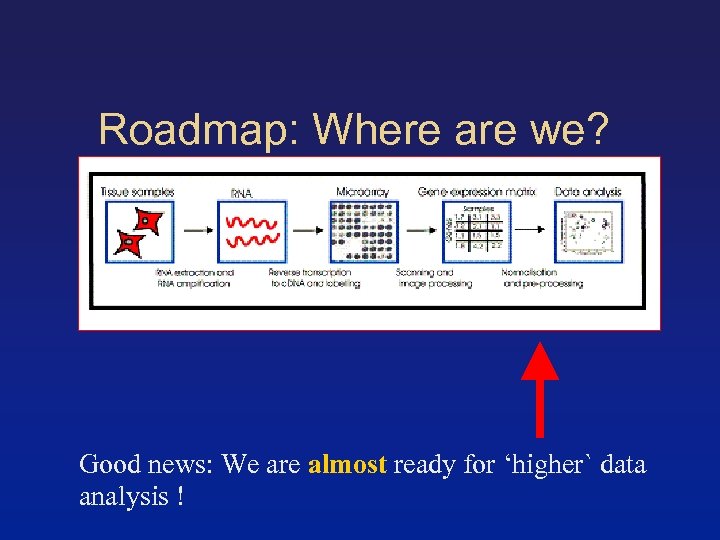 Roadmap: Where are we? Good news: We are almost ready for ‘higher` data analysis !
Roadmap: Where are we? Good news: We are almost ready for ‘higher` data analysis !
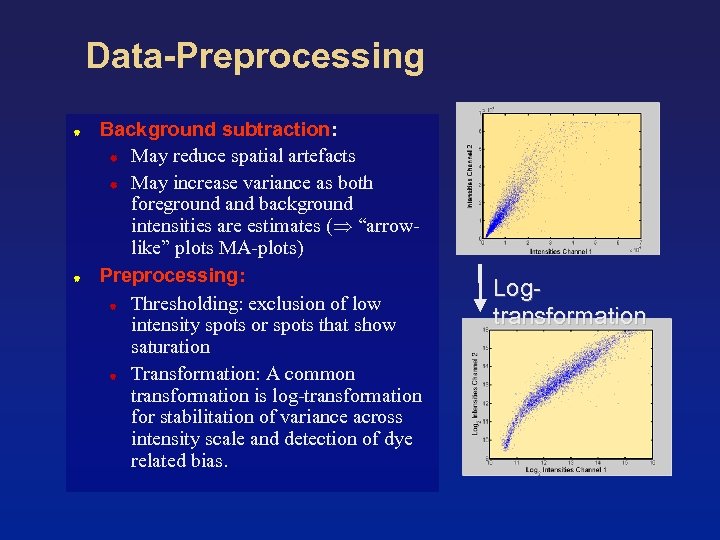 Data-Preprocessing Background subtraction: May reduce spatial artefacts May increase variance as both foreground and background intensities are estimates ( “arrowlike” plots MA-plots) Preprocessing: Thresholding: exclusion of low intensity spots or spots that show saturation Transformation: A common transformation is log-transformation for stabilitation of variance across intensity scale and detection of dye related bias. Logtransformation
Data-Preprocessing Background subtraction: May reduce spatial artefacts May increase variance as both foreground and background intensities are estimates ( “arrowlike” plots MA-plots) Preprocessing: Thresholding: exclusion of low intensity spots or spots that show saturation Transformation: A common transformation is log-transformation for stabilitation of variance across intensity scale and detection of dye related bias. Logtransformation
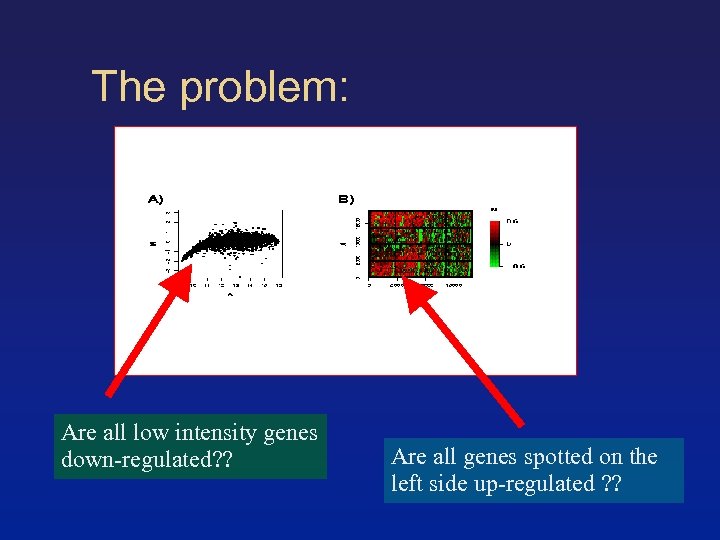 The problem: Are all low intensity genes down-regulated? ? Are all genes spotted on the left side up-regulated ? ?
The problem: Are all low intensity genes down-regulated? ? Are all genes spotted on the left side up-regulated ? ?
 Hybridisation model • Microarrays do not assess gene activities directly, but indirectly by measuring the fluorescence intensities of labelled target c. DNA hybridised to probes on the array. So how do we get what we are interested in? Answer: Find the relation between flourescance spot intensities and m. RNA abundance! • Explicitly modelling the relation between signal intensities and changes in gene expression can separate the measured error into systematic and random errors. • Systematic errors are errors which are reproducible and might be corrected in the normalisation procedure, whereas random errors cannot be corrected, but have to be assessed by replicate experiments.
Hybridisation model • Microarrays do not assess gene activities directly, but indirectly by measuring the fluorescence intensities of labelled target c. DNA hybridised to probes on the array. So how do we get what we are interested in? Answer: Find the relation between flourescance spot intensities and m. RNA abundance! • Explicitly modelling the relation between signal intensities and changes in gene expression can separate the measured error into systematic and random errors. • Systematic errors are errors which are reproducible and might be corrected in the normalisation procedure, whereas random errors cannot be corrected, but have to be assessed by replicate experiments.
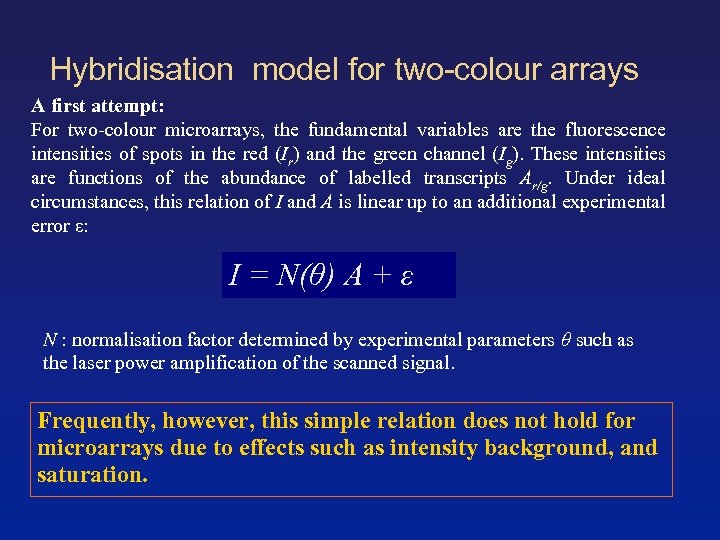 Hybridisation model for two-colour arrays A first attempt: For two-colour microarrays, the fundamental variables are the fluorescence intensities of spots in the red (Ir) and the green channel (Ig). These intensities are functions of the abundance of labelled transcripts Ar/g. Under ideal circumstances, this relation of I and A is linear up to an additional experimental error ε: I = N(θ) A + ε N : normalisation factor determined by experimental parameters θ such as the laser power amplification of the scanned signal. Frequently, however, this simple relation does not hold for microarrays due to effects such as intensity background, and saturation.
Hybridisation model for two-colour arrays A first attempt: For two-colour microarrays, the fundamental variables are the fluorescence intensities of spots in the red (Ir) and the green channel (Ig). These intensities are functions of the abundance of labelled transcripts Ar/g. Under ideal circumstances, this relation of I and A is linear up to an additional experimental error ε: I = N(θ) A + ε N : normalisation factor determined by experimental parameters θ such as the laser power amplification of the scanned signal. Frequently, however, this simple relation does not hold for microarrays due to effects such as intensity background, and saturation.
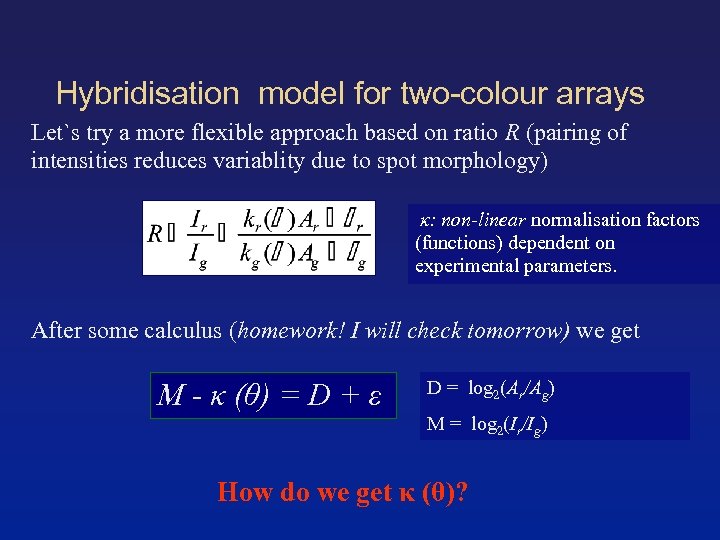 Hybridisation model for two-colour arrays Let`s try a more flexible approach based on ratio R (pairing of intensities reduces variablity due to spot morphology) κ: non-linear normalisation factors (functions) dependent on experimental parameters. After some calculus (homework! I will check tomorrow) we get M - κ (θ) = D + ε D = log 2(Ar/Ag) M = log 2(Ir/Ig) How do we get κ (θ)?
Hybridisation model for two-colour arrays Let`s try a more flexible approach based on ratio R (pairing of intensities reduces variablity due to spot morphology) κ: non-linear normalisation factors (functions) dependent on experimental parameters. After some calculus (homework! I will check tomorrow) we get M - κ (θ) = D + ε D = log 2(Ar/Ag) M = log 2(Ir/Ig) How do we get κ (θ)?
 Normalization – bending data to make it look nicer. . . Normalization describes a variety of data transformations aiming to correct for experimental variation
Normalization – bending data to make it look nicer. . . Normalization describes a variety of data transformations aiming to correct for experimental variation
 Within – array normalization Normalization based on 'householding genes' assumed to be equally expressed in different samples of interest Normalization using 'spiked in' genes: Ajustment of intensities so that control spots show equal intensities across channels and arrays Global linear normalisation assumes that overall expression in samples is constant. Thus, overall intensitiy of both channels is linearly scaled to have value. Non-linear normalisation assumes symmetry of differential expression across intensity scale and spatial dimension of array
Within – array normalization Normalization based on 'householding genes' assumed to be equally expressed in different samples of interest Normalization using 'spiked in' genes: Ajustment of intensities so that control spots show equal intensities across channels and arrays Global linear normalisation assumes that overall expression in samples is constant. Thus, overall intensitiy of both channels is linearly scaled to have value. Non-linear normalisation assumes symmetry of differential expression across intensity scale and spatial dimension of array
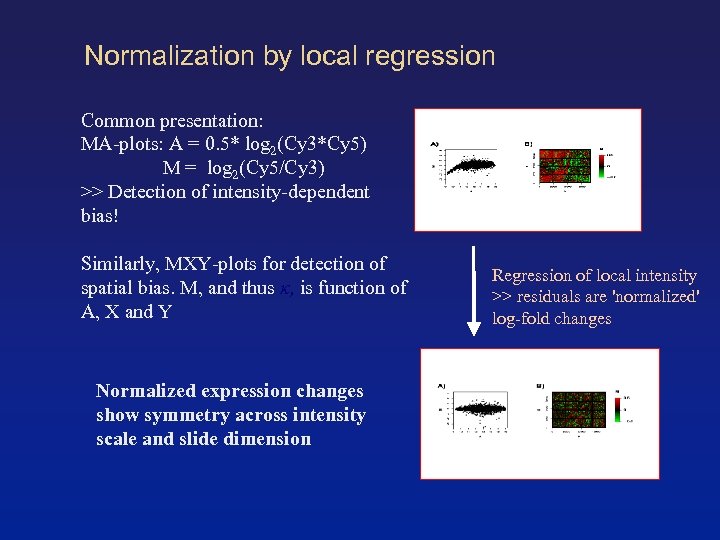 Normalization by local regression Common presentation: MA-plots: A = 0. 5* log 2(Cy 3*Cy 5) M = log 2(Cy 5/Cy 3) >> Detection of intensity-dependent bias! Similarly, MXY-plots for detection of spatial bias. M, and thus κ, is function of A, X and Y Normalized expression changes show symmetry across intensity scale and slide dimension Regression of local intensity >> residuals are 'normalized' log-fold changes
Normalization by local regression Common presentation: MA-plots: A = 0. 5* log 2(Cy 3*Cy 5) M = log 2(Cy 5/Cy 3) >> Detection of intensity-dependent bias! Similarly, MXY-plots for detection of spatial bias. M, and thus κ, is function of A, X and Y Normalized expression changes show symmetry across intensity scale and slide dimension Regression of local intensity >> residuals are 'normalized' log-fold changes
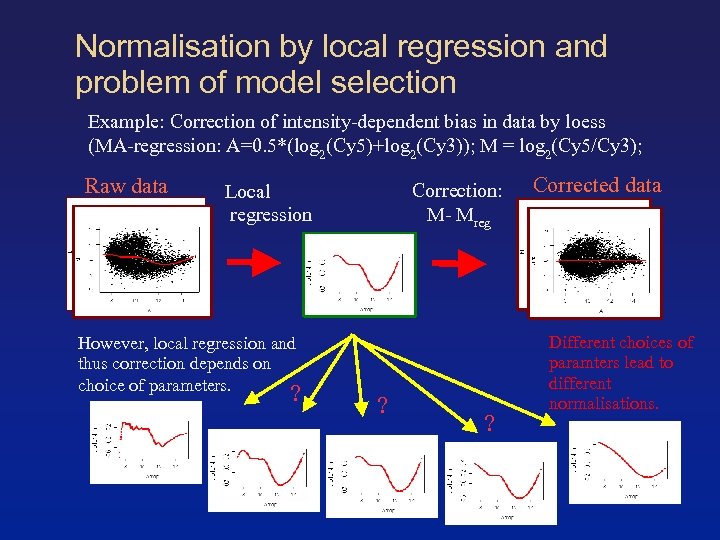 Normalisation by local regression and problem of model selection Example: Correction of intensity-dependent bias in data by loess (MA-regression: A=0. 5*(log 2(Cy 5)+log 2(Cy 3)); M = log 2(Cy 5/Cy 3); Raw data Correction: M- Mreg Local regression However, local regression and thus correction depends on choice of parameters. ? ? ? Corrected data Different choices of paramters lead to different normalisations.
Normalisation by local regression and problem of model selection Example: Correction of intensity-dependent bias in data by loess (MA-regression: A=0. 5*(log 2(Cy 5)+log 2(Cy 3)); M = log 2(Cy 5/Cy 3); Raw data Correction: M- Mreg Local regression However, local regression and thus correction depends on choice of parameters. ? ? ? Corrected data Different choices of paramters lead to different normalisations.
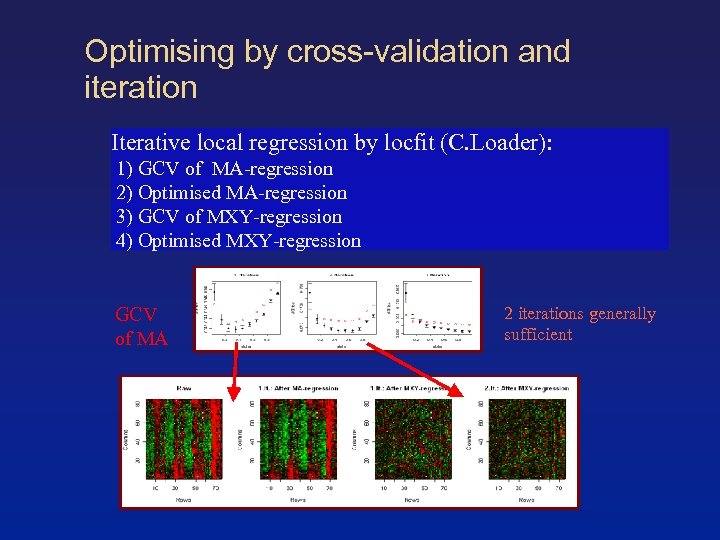 Optimising by cross-validation and iteration Iterative local regression by locfit (C. Loader): 1) GCV of MA-regression 2) Optimised MA-regression 3) GCV of MXY-regression 4) Optimised MXY-regression GCV of MA 2 iterations generally sufficient
Optimising by cross-validation and iteration Iterative local regression by locfit (C. Loader): 1) GCV of MA-regression 2) Optimised MA-regression 3) GCV of MXY-regression 4) Optimised MXY-regression GCV of MA 2 iterations generally sufficient
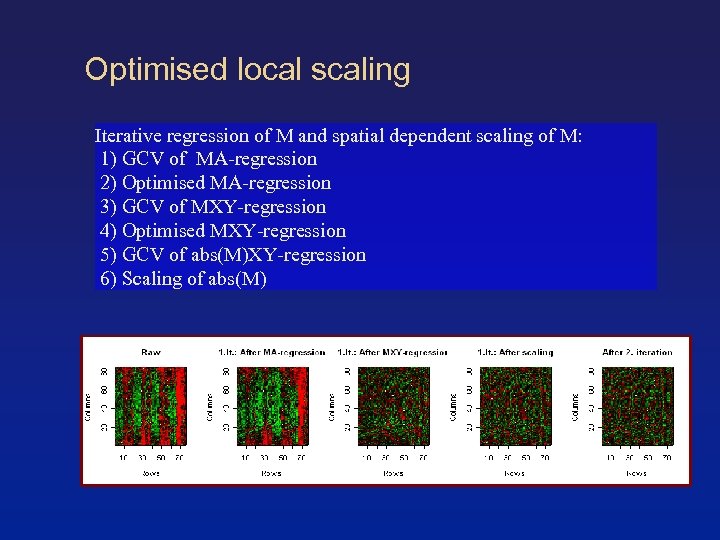 Optimised local scaling Iterative regression of M and spatial dependent scaling of M: 1) GCV of MA-regression 2) Optimised MA-regression 3) GCV of MXY-regression 4) Optimised MXY-regression 5) GCV of abs(M)XY-regression 6) Scaling of abs(M)
Optimised local scaling Iterative regression of M and spatial dependent scaling of M: 1) GCV of MA-regression 2) Optimised MA-regression 3) GCV of MXY-regression 4) Optimised MXY-regression 5) GCV of abs(M)XY-regression 6) Scaling of abs(M)
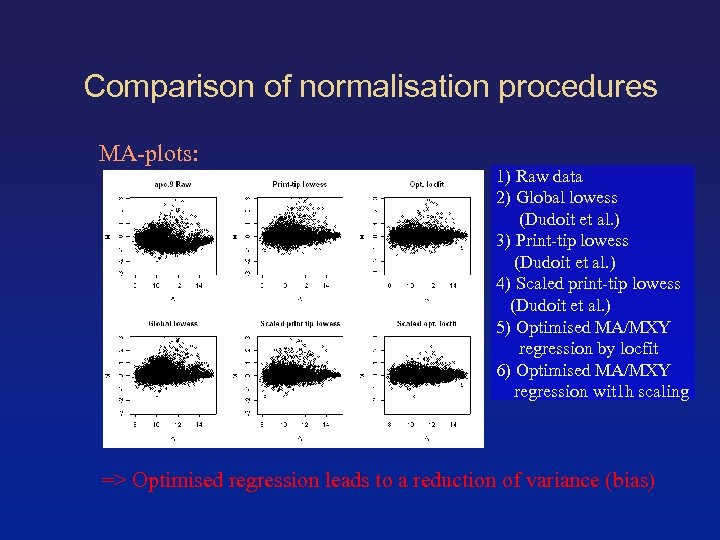 Comparison of normalisation procedures MA-plots: 1) Raw data 2) Global lowess (Dudoit et al. ) 3) Print-tip lowess (Dudoit et al. ) 4) Scaled print-tip lowess (Dudoit et al. ) 5) Optimised MA/MXY regression by locfit 6) Optimised MA/MXY regression wit 1 h scaling => Optimised regression leads to a reduction of variance (bias)
Comparison of normalisation procedures MA-plots: 1) Raw data 2) Global lowess (Dudoit et al. ) 3) Print-tip lowess (Dudoit et al. ) 4) Scaled print-tip lowess (Dudoit et al. ) 5) Optimised MA/MXY regression by locfit 6) Optimised MA/MXY regression wit 1 h scaling => Optimised regression leads to a reduction of variance (bias)
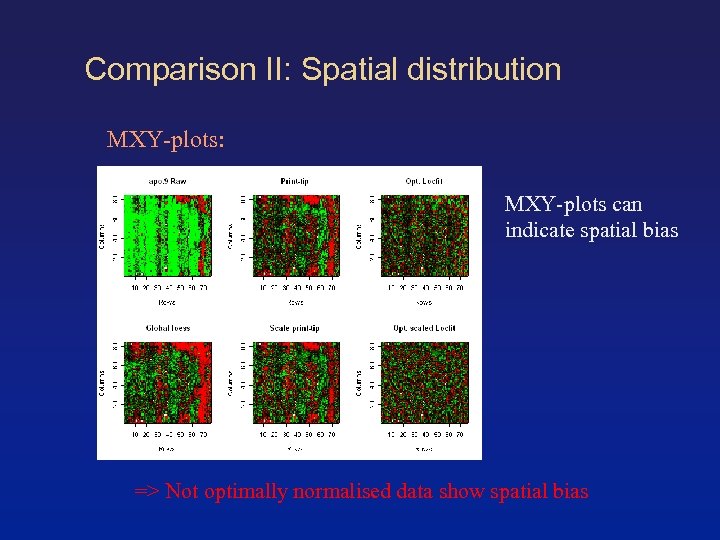 Comparison II: Spatial distribution MXY-plots: MXY-plots can indicate spatial bias => Not optimally normalised data show spatial bias
Comparison II: Spatial distribution MXY-plots: MXY-plots can indicate spatial bias => Not optimally normalised data show spatial bias
 Averaging by sliding window reveals uncorrected bias Distribution of median M within a window of 5 x 5 spots: => Spatial regression requires optimal adjustment to data
Averaging by sliding window reveals uncorrected bias Distribution of median M within a window of 5 x 5 spots: => Spatial regression requires optimal adjustment to data
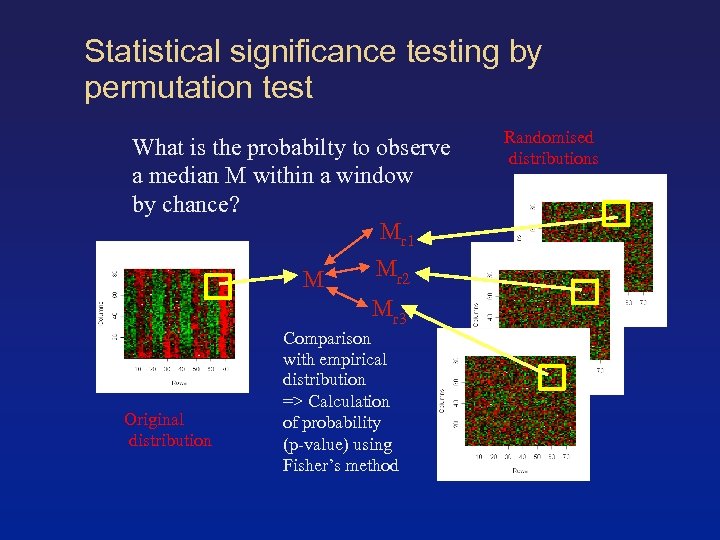 Statistical significance testing by permutation test What is the probabilty to observe a median M within a window by chance? Mr 1 M Mr 2 Mr 3 Original distribution Comparison with empirical distribution => Calculation of probability (p-value) using Fisher’s method Randomised distributions
Statistical significance testing by permutation test What is the probabilty to observe a median M within a window by chance? Mr 1 M Mr 2 Mr 3 Original distribution Comparison with empirical distribution => Calculation of probability (p-value) using Fisher’s method Randomised distributions
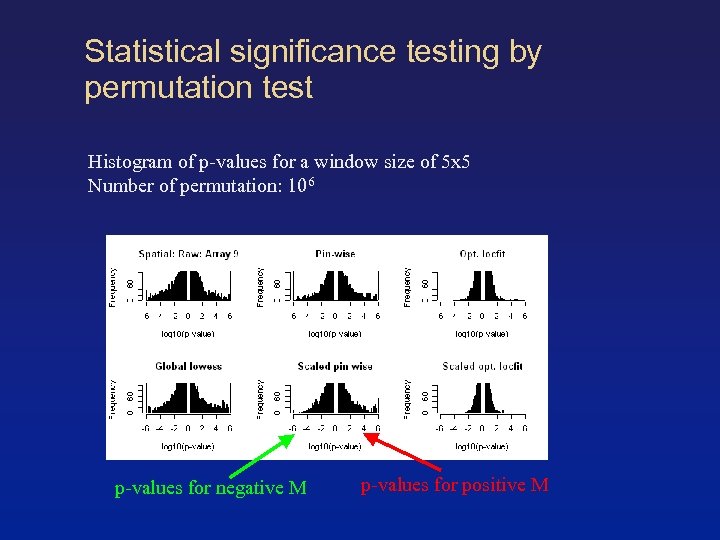 Statistical significance testing by permutation test Histogram of p-values for a window size of 5 x 5 Number of permutation: 106 p-values for negative M p-values for positive M
Statistical significance testing by permutation test Histogram of p-values for a window size of 5 x 5 Number of permutation: 106 p-values for negative M p-values for positive M
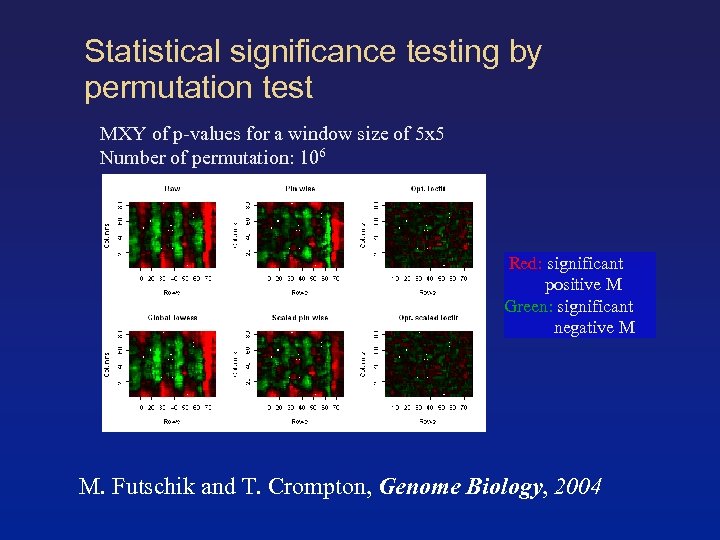 Statistical significance testing by permutation test MXY of p-values for a window size of 5 x 5 Number of permutation: 106 Red: significant positive M Green: significant negative M M. Futschik and T. Crompton, Genome Biology, 2004
Statistical significance testing by permutation test MXY of p-values for a window size of 5 x 5 Number of permutation: 106 Red: significant positive M Green: significant negative M M. Futschik and T. Crompton, Genome Biology, 2004
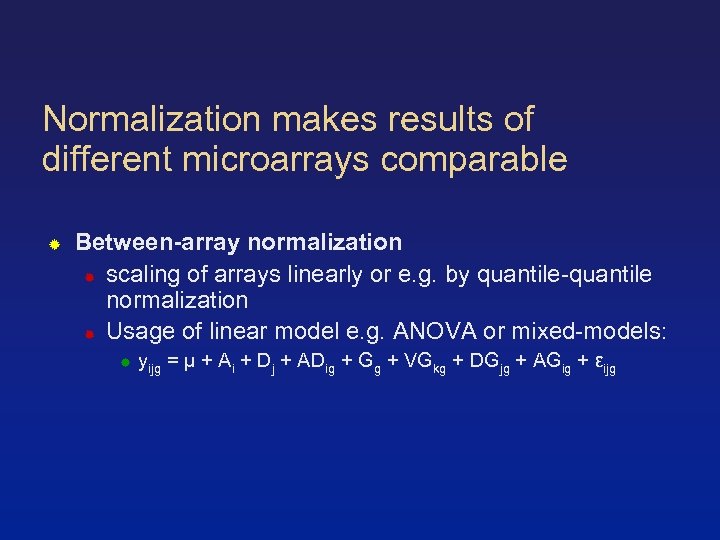 Normalization makes results of different microarrays comparable Between-array normalization scaling of arrays linearly or e. g. by quantile-quantile normalization Usage of linear model e. g. ANOVA or mixed-models: yijg = µ + Ai + Dj + ADig + Gg + VGkg + DGjg + AGig + εijg
Normalization makes results of different microarrays comparable Between-array normalization scaling of arrays linearly or e. g. by quantile-quantile normalization Usage of linear model e. g. ANOVA or mixed-models: yijg = µ + Ai + Dj + ADig + Gg + VGkg + DGjg + AGig + εijg
 Going fishing: What is differentially expressed Classical hypothesis testing: 1) Setting up of null hypothesis H 0(e. g. gene X is not differentially expressed) and alternative hypothesis Ha (e. g. Gene X is differentially expressed) 2) Using a test statistic to compare observed values with values predicted for H 0. 3) Define region for the test statistic for which H 0 is rejected in favour of Ha.
Going fishing: What is differentially expressed Classical hypothesis testing: 1) Setting up of null hypothesis H 0(e. g. gene X is not differentially expressed) and alternative hypothesis Ha (e. g. Gene X is differentially expressed) 2) Using a test statistic to compare observed values with values predicted for H 0. 3) Define region for the test statistic for which H 0 is rejected in favour of Ha.
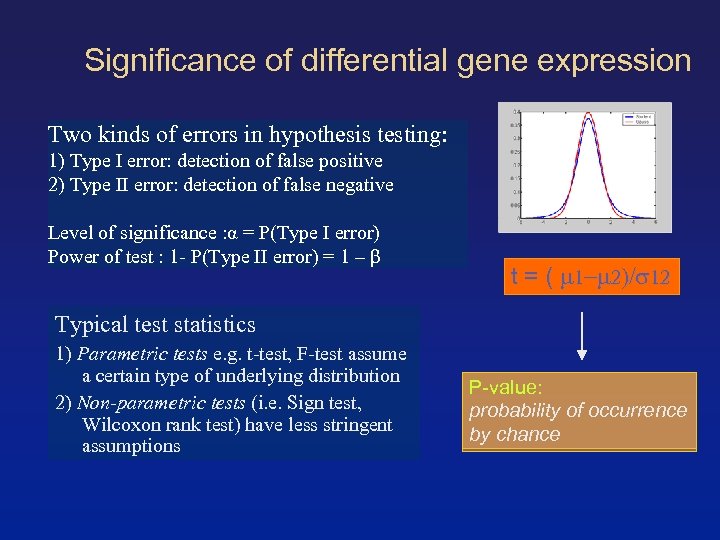 Significance of differential gene expression Two kinds of errors in hypothesis testing: 1) Type I error: detection of false positive 2) Type II error: detection of false negative Level of significance : α = P(Type I error) Power of test : 1 - P(Type II error) = 1 – β t = ( Typical test statistics 1) Parametric tests e. g. t-test, F-test assume a certain type of underlying distribution 2) Non-parametric tests (i. e. Sign test, Wilcoxon rank test) have less stringent assumptions P-value: probability of occurrence by chance
Significance of differential gene expression Two kinds of errors in hypothesis testing: 1) Type I error: detection of false positive 2) Type II error: detection of false negative Level of significance : α = P(Type I error) Power of test : 1 - P(Type II error) = 1 – β t = ( Typical test statistics 1) Parametric tests e. g. t-test, F-test assume a certain type of underlying distribution 2) Non-parametric tests (i. e. Sign test, Wilcoxon rank test) have less stringent assumptions P-value: probability of occurrence by chance
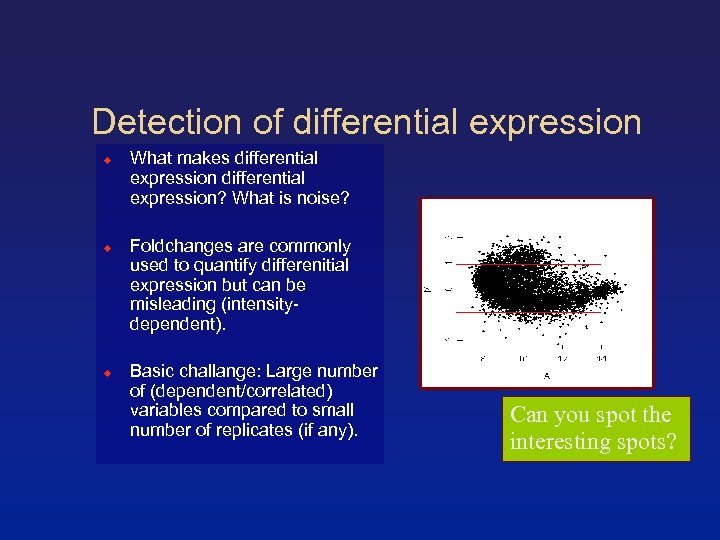 Detection of differential expression What makes differential expression? What is noise? Foldchanges are commonly used to quantify differenitial expression but can be misleading (intensitydependent). Basic challange: Large number of (dependent/correlated) variables compared to small number of replicates (if any). Can you spot the interesting spots?
Detection of differential expression What makes differential expression? What is noise? Foldchanges are commonly used to quantify differenitial expression but can be misleading (intensitydependent). Basic challange: Large number of (dependent/correlated) variables compared to small number of replicates (if any). Can you spot the interesting spots?
 Criteria for gene selection Accuracy: how closely are the results to the true values Precision: how variable are the results compared to the true value Sensitivity: how many true posítive are detected Specificity: how many of the selected genes are true positives.
Criteria for gene selection Accuracy: how closely are the results to the true values Precision: how variable are the results compared to the true value Sensitivity: how many true posítive are detected Specificity: how many of the selected genes are true positives.
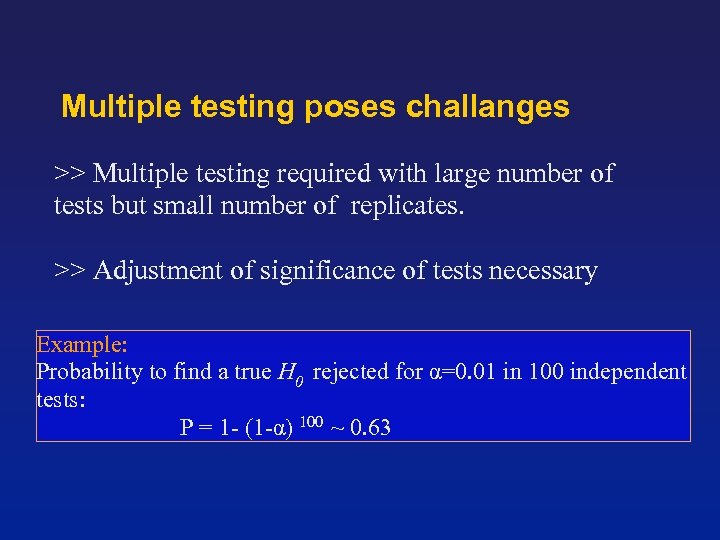 Multiple testing poses challanges >> Multiple testing required with large number of tests but small number of replicates. >> Adjustment of significance of tests necessary Example: Probability to find a true H 0 rejected for α=0. 01 in 100 independent tests: P = 1 - (1 -α) 100 ~ 0. 63
Multiple testing poses challanges >> Multiple testing required with large number of tests but small number of replicates. >> Adjustment of significance of tests necessary Example: Probability to find a true H 0 rejected for α=0. 01 in 100 independent tests: P = 1 - (1 -α) 100 ~ 0. 63
![Compound error measures: Per comparison error rate: PCER= E[V]/N Familiywise error rate: FWER=P(V≥ 1) Compound error measures: Per comparison error rate: PCER= E[V]/N Familiywise error rate: FWER=P(V≥ 1)](https://present5.com/presentation/d5716841ce6f0f85826a663372494191/image-51.jpg) Compound error measures: Per comparison error rate: PCER= E[V]/N Familiywise error rate: FWER=P(V≥ 1) False discovery rate: FDR= E[V/R] N: total number of tests V: number of reject true H 0 (FP) R: number of rejected H (TP+FP) Aim to control the error rate: 1) by p-value adjustment (step-down procedures: Bonferroni, Holm, Westfall-Young, . . . ) 2) by direct comparison with a background distribution (commonly generated by random permuation)
Compound error measures: Per comparison error rate: PCER= E[V]/N Familiywise error rate: FWER=P(V≥ 1) False discovery rate: FDR= E[V/R] N: total number of tests V: number of reject true H 0 (FP) R: number of rejected H (TP+FP) Aim to control the error rate: 1) by p-value adjustment (step-down procedures: Bonferroni, Holm, Westfall-Young, . . . ) 2) by direct comparison with a background distribution (commonly generated by random permuation)
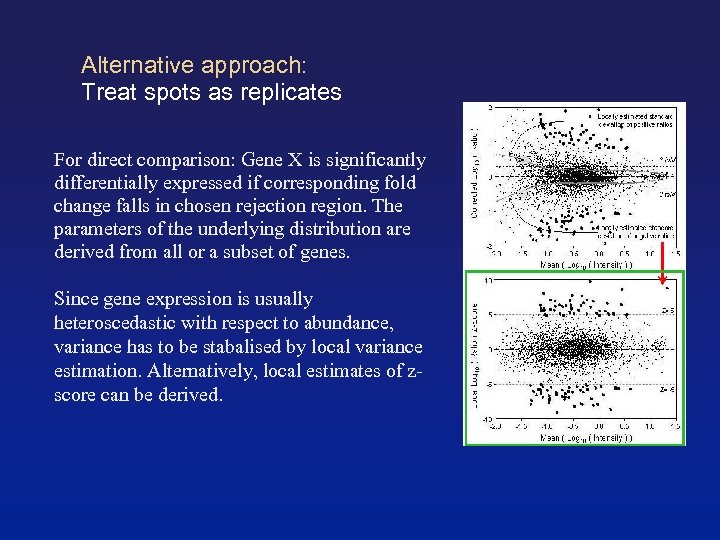 Alternative approach: Treat spots as replicates For direct comparison: Gene X is significantly differentially expressed if corresponding fold change falls in chosen rejection region. The parameters of the underlying distribution are derived from all or a subset of genes. Since gene expression is usually heteroscedastic with respect to abundance, variance has to be stabalised by local variance estimation. Alternatively, local estimates of zscore can be derived.
Alternative approach: Treat spots as replicates For direct comparison: Gene X is significantly differentially expressed if corresponding fold change falls in chosen rejection region. The parameters of the underlying distribution are derived from all or a subset of genes. Since gene expression is usually heteroscedastic with respect to abundance, variance has to be stabalised by local variance estimation. Alternatively, local estimates of zscore can be derived.
 Constistency of replications Case study: SW 480/620 cell line comparison SW 480: derived from primary tumour SW 620: derived from lymphnode metastisis of same patient Model for cancer progression Experimental design: • 4 independent hybridisations, • 4000 genes • c. DNA of SW 620 Cy 5 -labelled, • c. DNA of SW 480 Cy 3 -labelled. This design poses a problem! Can you spot it?
Constistency of replications Case study: SW 480/620 cell line comparison SW 480: derived from primary tumour SW 620: derived from lymphnode metastisis of same patient Model for cancer progression Experimental design: • 4 independent hybridisations, • 4000 genes • c. DNA of SW 620 Cy 5 -labelled, • c. DNA of SW 480 Cy 3 -labelled. This design poses a problem! Can you spot it?
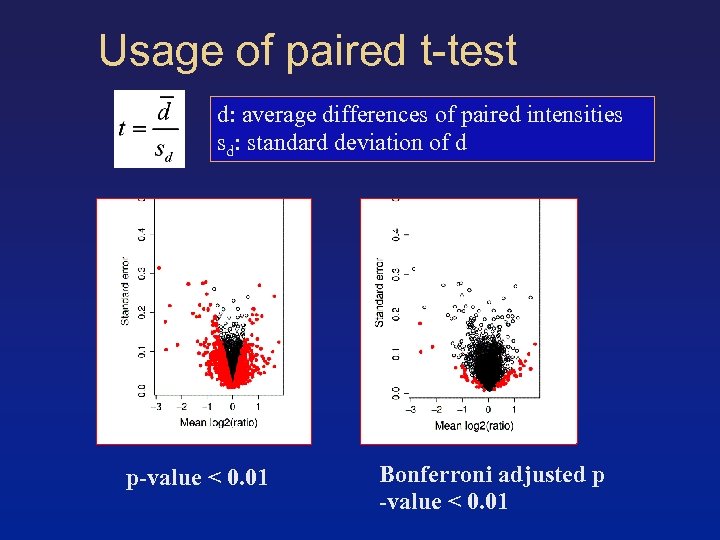 Usage of paired t-test d: average differences of paired intensities sd: standard deviation of d p-value < 0. 01 Bonferroni adjusted p -value < 0. 01
Usage of paired t-test d: average differences of paired intensities sd: standard deviation of d p-value < 0. 01 Bonferroni adjusted p -value < 0. 01
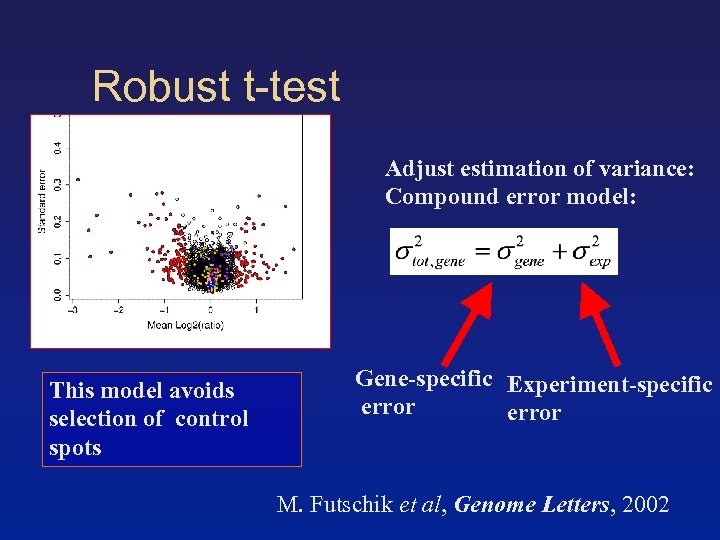 Robust t-test Adjust estimation of variance: Compound error model: This model avoids selection of control spots Gene-specific Experiment-specific error M. Futschik et al, Genome Letters, 2002
Robust t-test Adjust estimation of variance: Compound error model: This model avoids selection of control spots Gene-specific Experiment-specific error M. Futschik et al, Genome Letters, 2002
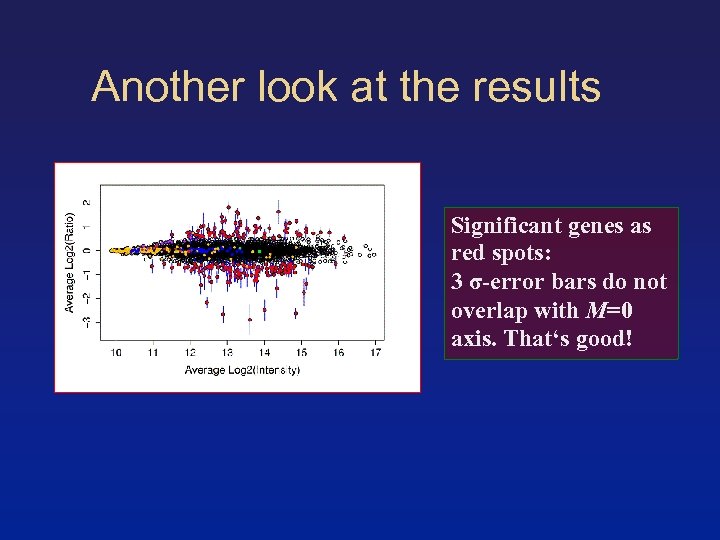 Another look at the results Significant genes as red spots: 3 σ-error bars do not overlap with M=0 axis. That‘s good!
Another look at the results Significant genes as red spots: 3 σ-error bars do not overlap with M=0 axis. That‘s good!
 Take-home messages II Don‘t download analyse array data blindly Visualise distributions: the eye is astonishing good in finding interesting spots Use different statistics and try to understand the differences Remember: Statistical significance is not necessary biological significance!
Take-home messages II Don‘t download analyse array data blindly Visualise distributions: the eye is astonishing good in finding interesting spots Use different statistics and try to understand the differences Remember: Statistical significance is not necessary biological significance!
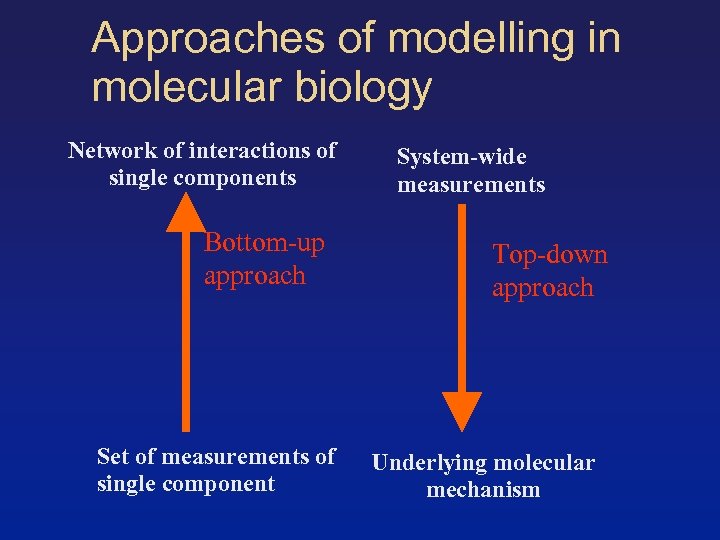 Approaches of modelling in molecular biology Network of interactions of single components Bottom-up approach Set of measurements of single component System-wide measurements Top-down approach Underlying molecular mechanism
Approaches of modelling in molecular biology Network of interactions of single components Bottom-up approach Set of measurements of single component System-wide measurements Top-down approach Underlying molecular mechanism
 Visualisation • Important tool for detection of patterns remains visualization of results. • Examples are MA-plots, dendrograms, Venndiagrams or projection derived fro multi-dimensional scaling. • Do not underestimate the ability of the human eye
Visualisation • Important tool for detection of patterns remains visualization of results. • Examples are MA-plots, dendrograms, Venndiagrams or projection derived fro multi-dimensional scaling. • Do not underestimate the ability of the human eye
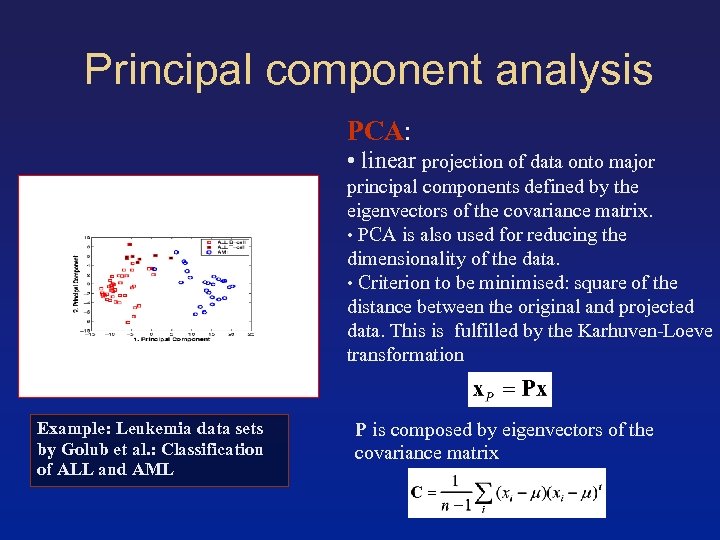 Principal component analysis PCA: • linear projection of data onto major principal components defined by the eigenvectors of the covariance matrix. • PCA is also used for reducing the dimensionality of the data. • Criterion to be minimised: square of the distance between the original and projected data. This is fulfilled by the Karhuven-Loeve transformation Example: Leukemia data sets by Golub et al. : Classification of ALL and AML P is composed by eigenvectors of the covariance matrix
Principal component analysis PCA: • linear projection of data onto major principal components defined by the eigenvectors of the covariance matrix. • PCA is also used for reducing the dimensionality of the data. • Criterion to be minimised: square of the distance between the original and projected data. This is fulfilled by the Karhuven-Loeve transformation Example: Leukemia data sets by Golub et al. : Classification of ALL and AML P is composed by eigenvectors of the covariance matrix
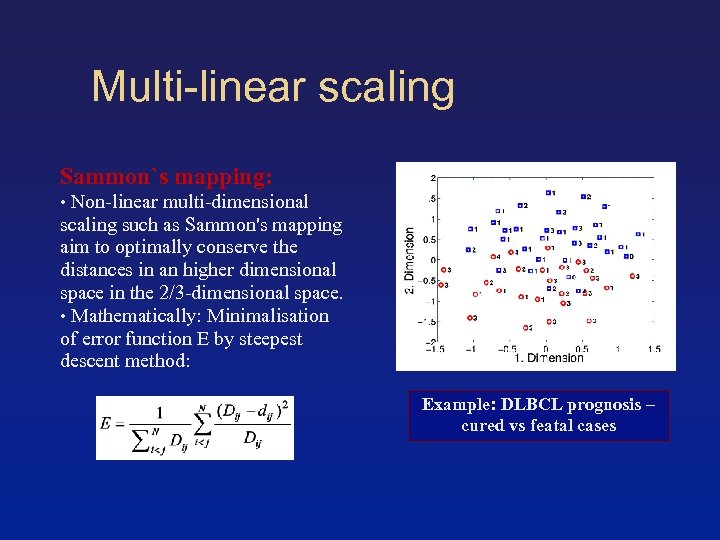 Multi-linear scaling Sammon`s mapping: • Non-linear multi-dimensional scaling such as Sammon's mapping aim to optimally conserve the distances in an higher dimensional space in the 2/3 -dimensional space. • Mathematically: Minimalisation of error function E by steepest descent method: Example: DLBCL prognosis – cured vs featal cases
Multi-linear scaling Sammon`s mapping: • Non-linear multi-dimensional scaling such as Sammon's mapping aim to optimally conserve the distances in an higher dimensional space in the 2/3 -dimensional space. • Mathematically: Minimalisation of error function E by steepest descent method: Example: DLBCL prognosis – cured vs featal cases
 Clustering: Birds of a feather flock together Clustering of genes Clustering of array: Co-expression indicates coregulation: functional annotation Clustering of time series finding new subclasses in samplespace Two-way clustering: Parallel clustering of samples and genes
Clustering: Birds of a feather flock together Clustering of genes Clustering of array: Co-expression indicates coregulation: functional annotation Clustering of time series finding new subclasses in samplespace Two-way clustering: Parallel clustering of samples and genes
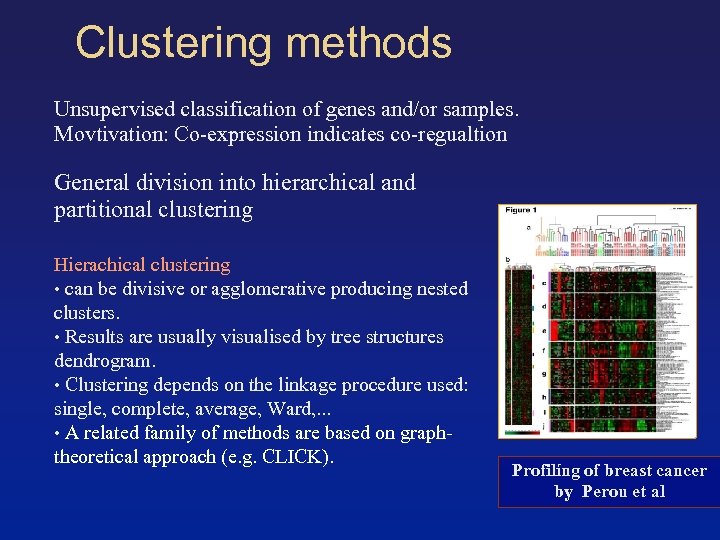 Clustering methods Unsupervised classification of genes and/or samples. Movtivation: Co-expression indicates co-regualtion General division into hierarchical and partitional clustering Hierachical clustering • can be divisive or agglomerative producing nested clusters. • Results are usually visualised by tree structures dendrogram. • Clustering depends on the linkage procedure used: single, complete, average, Ward, . . . • A related family of methods are based on graphtheoretical approach (e. g. CLICK). Profilíng of breast cancer by Perou et al
Clustering methods Unsupervised classification of genes and/or samples. Movtivation: Co-expression indicates co-regualtion General division into hierarchical and partitional clustering Hierachical clustering • can be divisive or agglomerative producing nested clusters. • Results are usually visualised by tree structures dendrogram. • Clustering depends on the linkage procedure used: single, complete, average, Ward, . . . • A related family of methods are based on graphtheoretical approach (e. g. CLICK). Profilíng of breast cancer by Perou et al
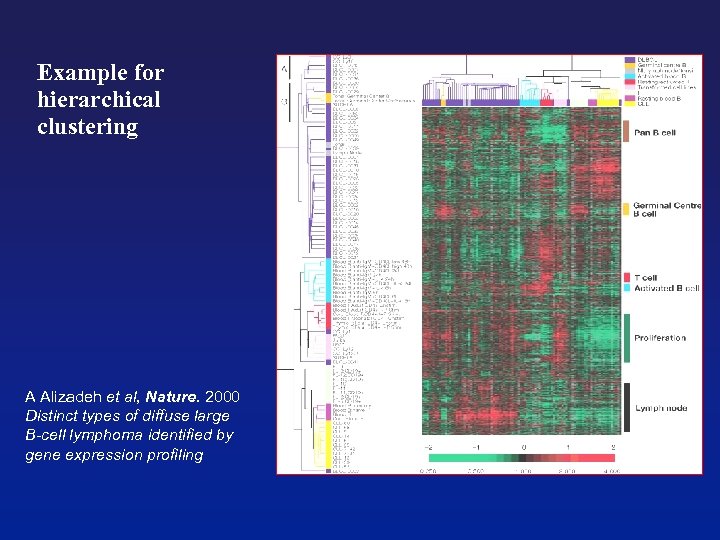 Example for hierarchical clustering A Alizadeh et al, Nature. 2000 Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling
Example for hierarchical clustering A Alizadeh et al, Nature. 2000 Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling
 Clustering methods II Partitional clustering • divides data into a (pre-)chosen number of classes. • Examples: k-means, SOMs, fuzzy c-means, simulated annealing, model -based clustering, HMMs, . . . • Setting the number of clusters is problematic Cluster validity: • Most cluster algorithms always detect clusters, even in random data. • Cluster validation approaches address the number of existing clusters. • Approaches are based on objective functions, figures of merits, resampling, adding noise. .
Clustering methods II Partitional clustering • divides data into a (pre-)chosen number of classes. • Examples: k-means, SOMs, fuzzy c-means, simulated annealing, model -based clustering, HMMs, . . . • Setting the number of clusters is problematic Cluster validity: • Most cluster algorithms always detect clusters, even in random data. • Cluster validation approaches address the number of existing clusters. • Approaches are based on objective functions, figures of merits, resampling, adding noise. .
 Hard clustering vs. soft clustering Hard clustering: Based on classical set theory Assigns a gene to exactly one cluster No differentiation how well gene is represented by cluster centroid Examples: hierachical clustering, k-means, SOMs, . . . Soft clustering: Can assign a gene to several cluster Differentiate grade of representation (cluster membership) Example: Fuzzy c-means, HMMs, . . .
Hard clustering vs. soft clustering Hard clustering: Based on classical set theory Assigns a gene to exactly one cluster No differentiation how well gene is represented by cluster centroid Examples: hierachical clustering, k-means, SOMs, . . . Soft clustering: Can assign a gene to several cluster Differentiate grade of representation (cluster membership) Example: Fuzzy c-means, HMMs, . . .
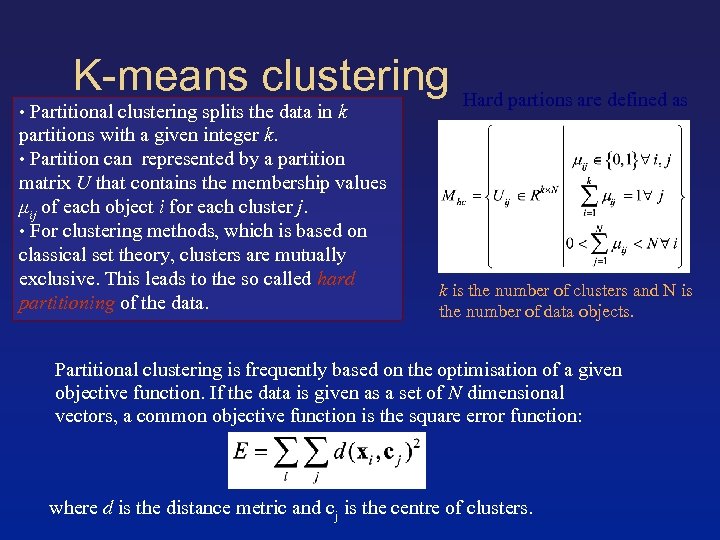 K-means clustering • Partitional clustering splits the data in k partitions with a given integer k. • Partition can represented by a partition matrix U that contains the membership values μij of each object i for each cluster j. • For clustering methods, which is based on classical set theory, clusters are mutually exclusive. This leads to the so called hard partitioning of the data. Hard partions are defined as k is the number of clusters and N is the number of data objects. Partitional clustering is frequently based on the optimisation of a given objective function. If the data is given as a set of N dimensional vectors, a common objective function is the square error function: where d is the distance metric and cj is the centre of clusters.
K-means clustering • Partitional clustering splits the data in k partitions with a given integer k. • Partition can represented by a partition matrix U that contains the membership values μij of each object i for each cluster j. • For clustering methods, which is based on classical set theory, clusters are mutually exclusive. This leads to the so called hard partitioning of the data. Hard partions are defined as k is the number of clusters and N is the number of data objects. Partitional clustering is frequently based on the optimisation of a given objective function. If the data is given as a set of N dimensional vectors, a common objective function is the square error function: where d is the distance metric and cj is the centre of clusters.
 K-means algorithm • • • Initiation: Choose k random vectors as cluster centres cj ; Partitioning: Assign xi to if for all k with k ; Calculation of cluster centres cj based on the partition derived in step 2: The cluster centre cj is defined as the mean value of all vectors within the cluster; Calculation of the square error function E; If the chosen stop criterion is met, stop; otherwise continue with step 2. For the distance metric D, the Euclidean distance is generally chosen.
K-means algorithm • • • Initiation: Choose k random vectors as cluster centres cj ; Partitioning: Assign xi to if for all k with k ; Calculation of cluster centres cj based on the partition derived in step 2: The cluster centre cj is defined as the mean value of all vectors within the cluster; Calculation of the square error function E; If the chosen stop criterion is met, stop; otherwise continue with step 2. For the distance metric D, the Euclidean distance is generally chosen.
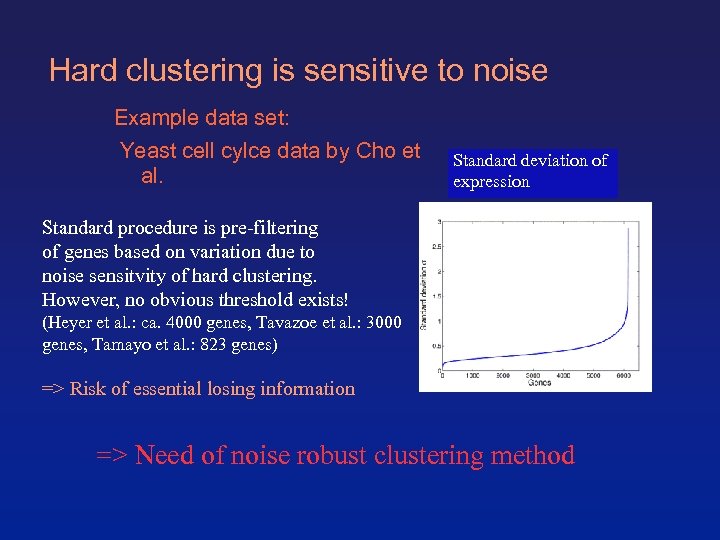 Hard clustering is sensitive to noise Example data set: Yeast cell cylce data by Cho et Standard deviation of al. expression Standard procedure is pre-filtering of genes based on variation due to noise sensitvity of hard clustering. However, no obvious threshold exists! (Heyer et al. : ca. 4000 genes, Tavazoe et al. : 3000 genes, Tamayo et al. : 823 genes) => Risk of essential losing information => Need of noise robust clustering method
Hard clustering is sensitive to noise Example data set: Yeast cell cylce data by Cho et Standard deviation of al. expression Standard procedure is pre-filtering of genes based on variation due to noise sensitvity of hard clustering. However, no obvious threshold exists! (Heyer et al. : ca. 4000 genes, Tavazoe et al. : 3000 genes, Tamayo et al. : 823 genes) => Risk of essential losing information => Need of noise robust clustering method
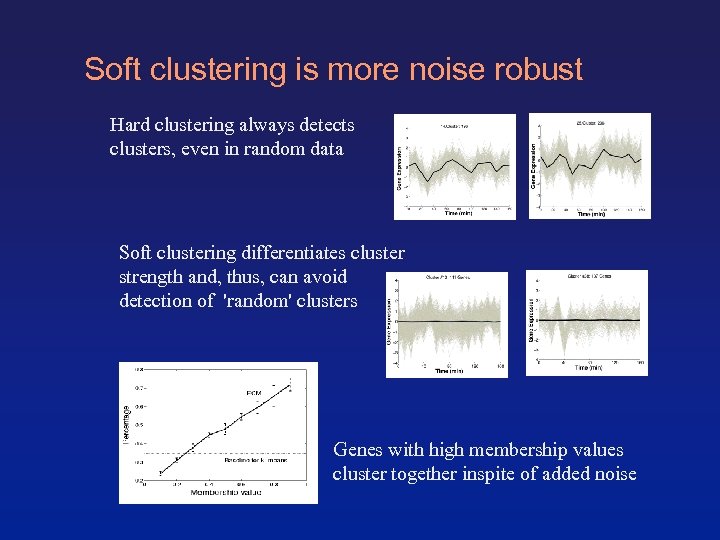 Soft clustering is more noise robust Hard clustering always detects clusters, even in random data Soft clustering differentiates cluster strength and, thus, can avoid detection of 'random' clusters Genes with high membership values cluster together inspite of added noise
Soft clustering is more noise robust Hard clustering always detects clusters, even in random data Soft clustering differentiates cluster strength and, thus, can avoid detection of 'random' clusters Genes with high membership values cluster together inspite of added noise
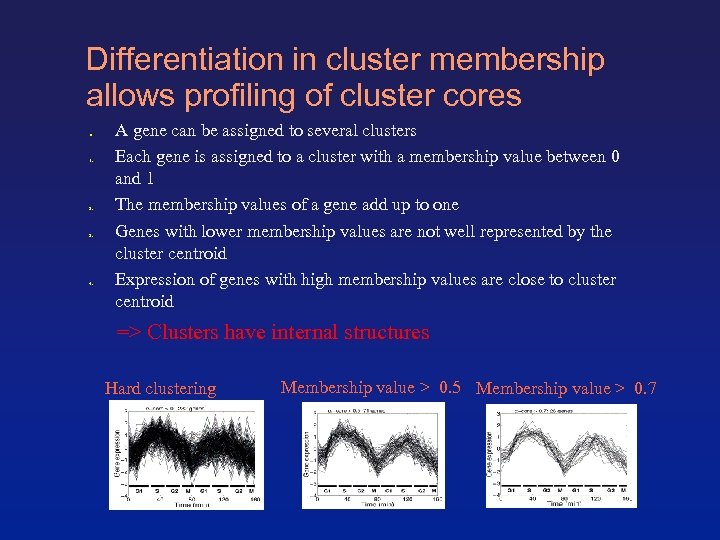 Differentiation in cluster membership allows profiling of cluster cores ● 1. 2. 3. 4. A gene can be assigned to several clusters Each gene is assigned to a cluster with a membership value between 0 and 1 The membership values of a gene add up to one Genes with lower membership values are not well represented by the cluster centroid Expression of genes with high membership values are close to cluster centroid => Clusters have internal structures Hard clustering Membership value > 0. 5 Membership value > 0. 7
Differentiation in cluster membership allows profiling of cluster cores ● 1. 2. 3. 4. A gene can be assigned to several clusters Each gene is assigned to a cluster with a membership value between 0 and 1 The membership values of a gene add up to one Genes with lower membership values are not well represented by the cluster centroid Expression of genes with high membership values are close to cluster centroid => Clusters have internal structures Hard clustering Membership value > 0. 5 Membership value > 0. 7
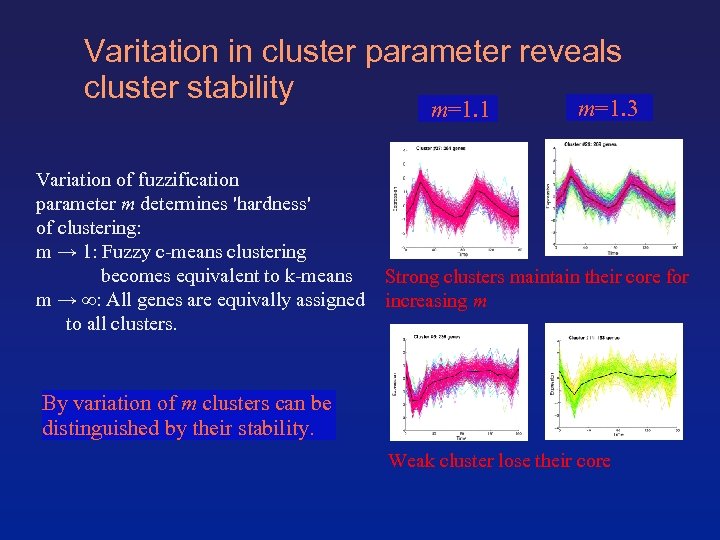 Varitation in cluster parameter reveals cluster stability m=1. 1 Variation of fuzzification parameter m determines 'hardness' of clustering: m → 1: Fuzzy c-means clustering becomes equivalent to k-means m → ∞: All genes are equivally assigned to all clusters. m=1. 3 Strong clusters maintain their core for increasing m By variation of m clusters can be distinguished by their stability. Weak cluster lose their core
Varitation in cluster parameter reveals cluster stability m=1. 1 Variation of fuzzification parameter m determines 'hardness' of clustering: m → 1: Fuzzy c-means clustering becomes equivalent to k-means m → ∞: All genes are equivally assigned to all clusters. m=1. 3 Strong clusters maintain their core for increasing m By variation of m clusters can be distinguished by their stability. Weak cluster lose their core
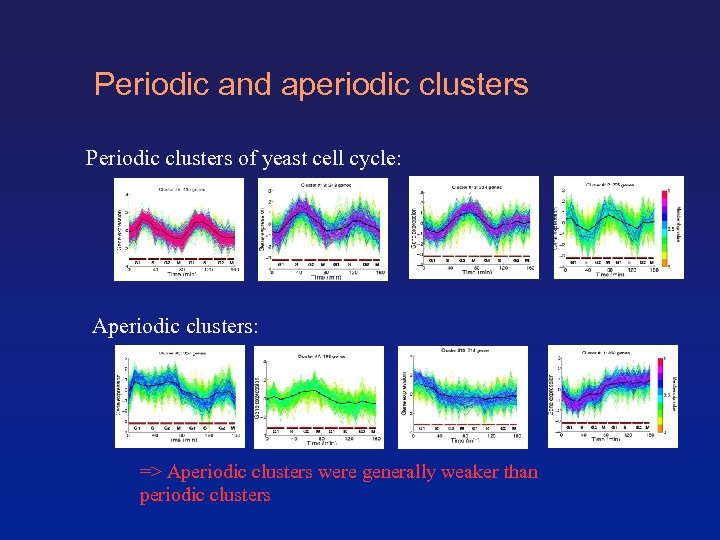 Periodic and aperiodic clusters Periodic clusters of yeast cell cycle: Aperiodic clusters: => Aperiodic clusters were generally weaker than periodic clusters
Periodic and aperiodic clusters Periodic clusters of yeast cell cycle: Aperiodic clusters: => Aperiodic clusters were generally weaker than periodic clusters
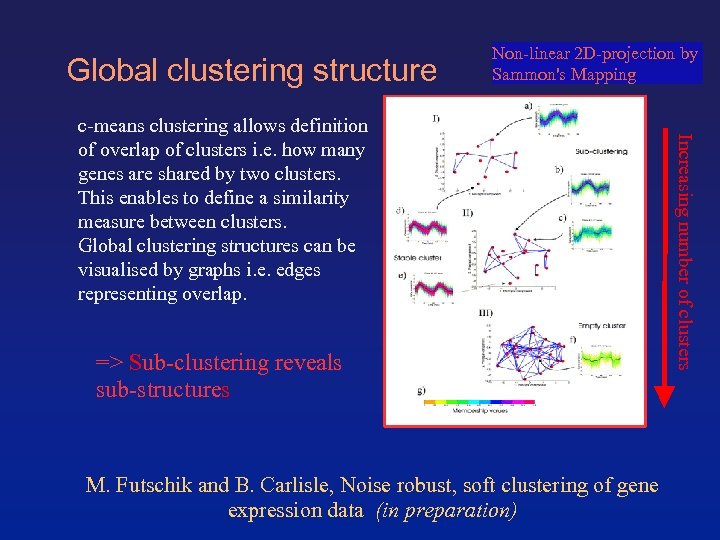 Global clustering structure Non-linear 2 D-projection by Sammon's Mapping => Sub-clustering reveals sub-structures M. Futschik and B. Carlisle, Noise robust, soft clustering of gene expression data (in preparation) Increasing number of clusters c-means clustering allows definition of overlap of clusters i. e. how many genes are shared by two clusters. This enables to define a similarity measure between clusters. Global clustering structures can be visualised by graphs i. e. edges representing overlap.
Global clustering structure Non-linear 2 D-projection by Sammon's Mapping => Sub-clustering reveals sub-structures M. Futschik and B. Carlisle, Noise robust, soft clustering of gene expression data (in preparation) Increasing number of clusters c-means clustering allows definition of overlap of clusters i. e. how many genes are shared by two clusters. This enables to define a similarity measure between clusters. Global clustering structures can be visualised by graphs i. e. edges representing overlap.
 Classifaction of microarray data Many diseases involve (unknown) complex interaction of multiple genes, thus ``single gene approach´´ is limited genome-wide approaches may reveal this interactions To detect this patterns, supervised learníng techniques from pattern recognition, statistics and artificial intelligence can be applied. The medical applications of these “arrays of hope” are various and include the identification of markers for classification, diagnosis, disease outcome prediction, therapeutic responsiveness and target identification.
Classifaction of microarray data Many diseases involve (unknown) complex interaction of multiple genes, thus ``single gene approach´´ is limited genome-wide approaches may reveal this interactions To detect this patterns, supervised learníng techniques from pattern recognition, statistics and artificial intelligence can be applied. The medical applications of these “arrays of hope” are various and include the identification of markers for classification, diagnosis, disease outcome prediction, therapeutic responsiveness and target identification.
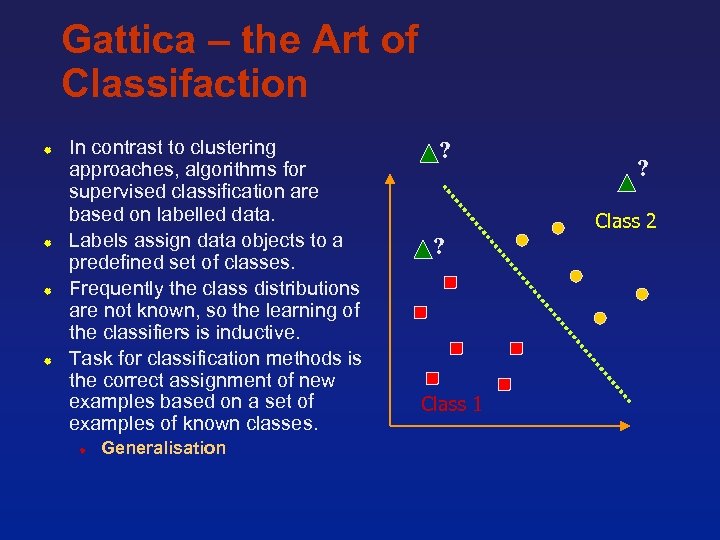 Gattica – the Art of Classifaction In contrast to clustering approaches, algorithms for supervised classification are based on labelled data. Labels assign data objects to a predefined set of classes. Frequently the class distributions are not known, so the learning of the classifiers is inductive. Task for classification methods is the correct assignment of new examples based on a set of examples of known classes. Generalisation ? ? Class 2 ? Class 1
Gattica – the Art of Classifaction In contrast to clustering approaches, algorithms for supervised classification are based on labelled data. Labels assign data objects to a predefined set of classes. Frequently the class distributions are not known, so the learning of the classifiers is inductive. Task for classification methods is the correct assignment of new examples based on a set of examples of known classes. Generalisation ? ? Class 2 ? Class 1
 Challanges in classification of microarray data • Microarray data inherit large experimental and biological variances • experimental bias + tissue hetrogenity • cross-hybridisation • ‘bad design’: confounding effects • Microarray data are sparse • high-dimensionality of gene (feature) space • low number of samples/arrays • Curse of dimensionality • Microarray data are highly redundant • Many genes are co-expressed, thus their expression is strongly correlated.
Challanges in classification of microarray data • Microarray data inherit large experimental and biological variances • experimental bias + tissue hetrogenity • cross-hybridisation • ‘bad design’: confounding effects • Microarray data are sparse • high-dimensionality of gene (feature) space • low number of samples/arrays • Curse of dimensionality • Microarray data are highly redundant • Many genes are co-expressed, thus their expression is strongly correlated.
 Classification I: Models K-nearest neighbour Decision Trees Inclusion of prior knowledge possible Neural Networks Easy to follow the classification process Bayesian classifier Simple and quick method No model assumed Support Vector Machines Based on statistical learning theory; today`s state of art.
Classification I: Models K-nearest neighbour Decision Trees Inclusion of prior knowledge possible Neural Networks Easy to follow the classification process Bayesian classifier Simple and quick method No model assumed Support Vector Machines Based on statistical learning theory; today`s state of art.
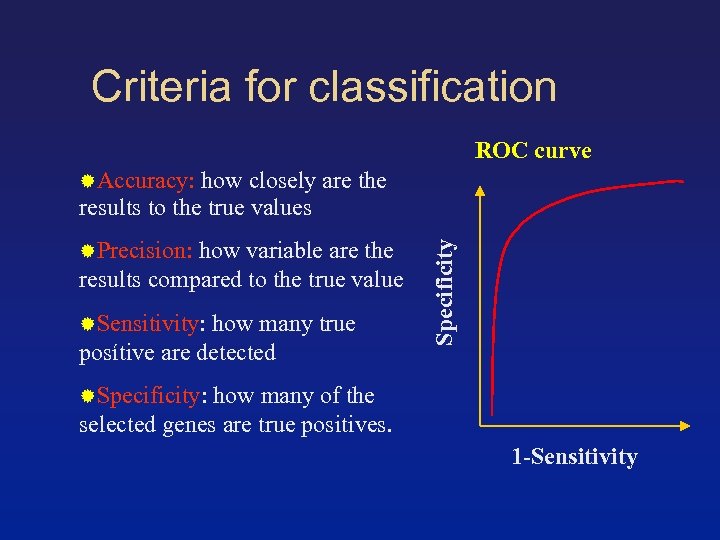 Criteria for classification ROC curve Accuracy: how closely are the Precision: how variable are the results compared to the true value Sensitivity: how many true posítive are detected Specificity results to the true values Specificity: how many of the selected genes are true positives. 1 -Sensitivity
Criteria for classification ROC curve Accuracy: how closely are the Precision: how variable are the results compared to the true value Sensitivity: how many true posítive are detected Specificity results to the true values Specificity: how many of the selected genes are true positives. 1 -Sensitivity
 Getting a good team: Feature Extraction - gene selection Selection of genes based on : Parametric tests e. g. t-test Non-parametric tests eg. Wilcoxon-Rank test But the 11 best players do not necessary form the best team! Selection of groups of genes which act as good: Sensitivity measure Genetic Algorithms Decision tree SVD
Getting a good team: Feature Extraction - gene selection Selection of genes based on : Parametric tests e. g. t-test Non-parametric tests eg. Wilcoxon-Rank test But the 11 best players do not necessary form the best team! Selection of groups of genes which act as good: Sensitivity measure Genetic Algorithms Decision tree SVD
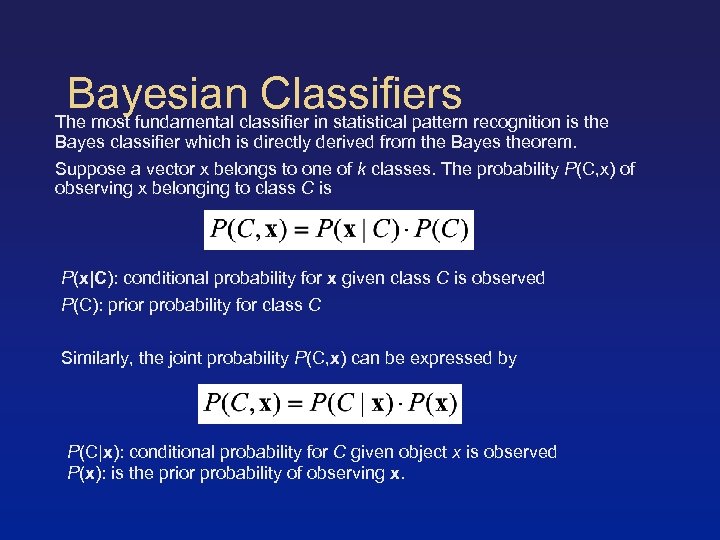 Bayesian Classifiers The most fundamental classifier in statistical pattern recognition is the Bayes classifier which is directly derived from the Bayes theorem. Suppose a vector x belongs to one of k classes. The probability P(C, x) of observing x belonging to class C is P(x|C): conditional probability for x given class C is observed P(C): prior probability for class C Similarly, the joint probability P(C, x) can be expressed by P(C|x): conditional probability for C given object x is observed P(x): is the prior probability of observing x.
Bayesian Classifiers The most fundamental classifier in statistical pattern recognition is the Bayes classifier which is directly derived from the Bayes theorem. Suppose a vector x belongs to one of k classes. The probability P(C, x) of observing x belonging to class C is P(x|C): conditional probability for x given class C is observed P(C): prior probability for class C Similarly, the joint probability P(C, x) can be expressed by P(C|x): conditional probability for C given object x is observed P(x): is the prior probability of observing x.
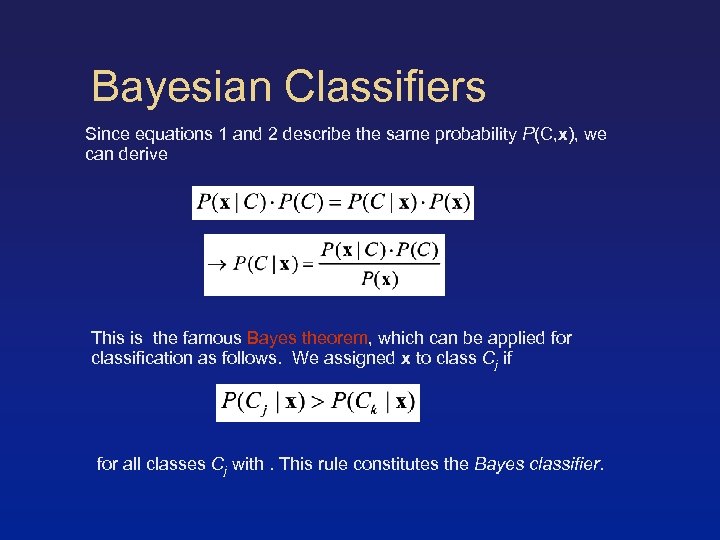 Bayesian Classifiers Since equations 1 and 2 describe the same probability P(C, x), we can derive This is the famous Bayes theorem, which can be applied for classification as follows. We assigned x to class Cj if for all classes Cj with. This rule constitutes the Bayes classifier.
Bayesian Classifiers Since equations 1 and 2 describe the same probability P(C, x), we can derive This is the famous Bayes theorem, which can be applied for classification as follows. We assigned x to class Cj if for all classes Cj with. This rule constitutes the Bayes classifier.
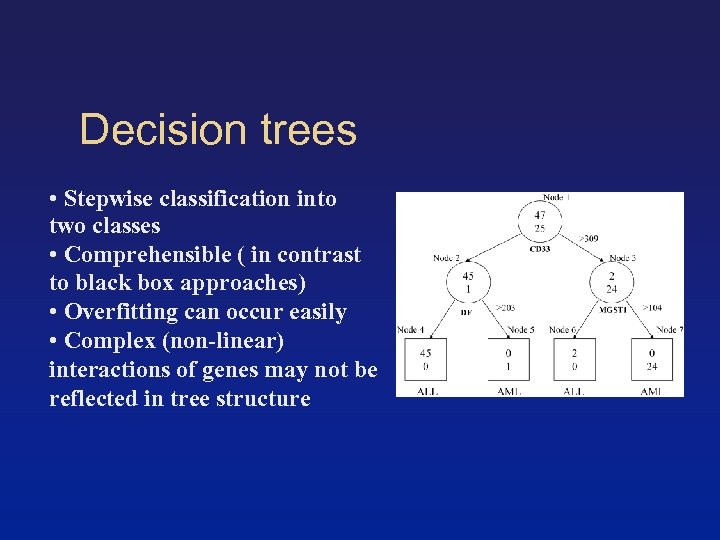 Decision trees • Stepwise classification into two classes • Comprehensible ( in contrast to black box approaches) • Overfitting can occur easily • Complex (non-linear) interactions of genes may not be reflected in tree structure
Decision trees • Stepwise classification into two classes • Comprehensible ( in contrast to black box approaches) • Overfitting can occur easily • Complex (non-linear) interactions of genes may not be reflected in tree structure
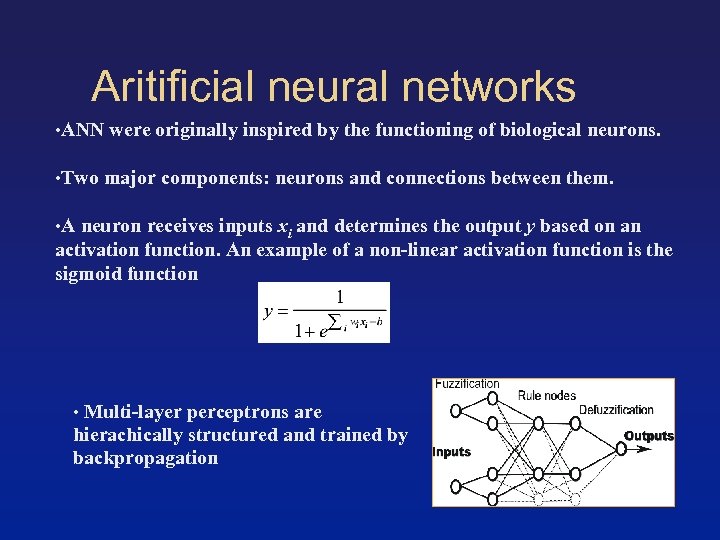 Aritificial neural networks • ANN were originally inspired by the functioning of biological neurons. • Two major components: neurons and connections between them. • A neuron receives inputs xi and determines the output y based on an activation function. An example of a non-linear activation function is the sigmoid function • Multi-layer perceptrons are hierachically structured and trained by backpropagation
Aritificial neural networks • ANN were originally inspired by the functioning of biological neurons. • Two major components: neurons and connections between them. • A neuron receives inputs xi and determines the output y based on an activation function. An example of a non-linear activation function is the sigmoid function • Multi-layer perceptrons are hierachically structured and trained by backpropagation
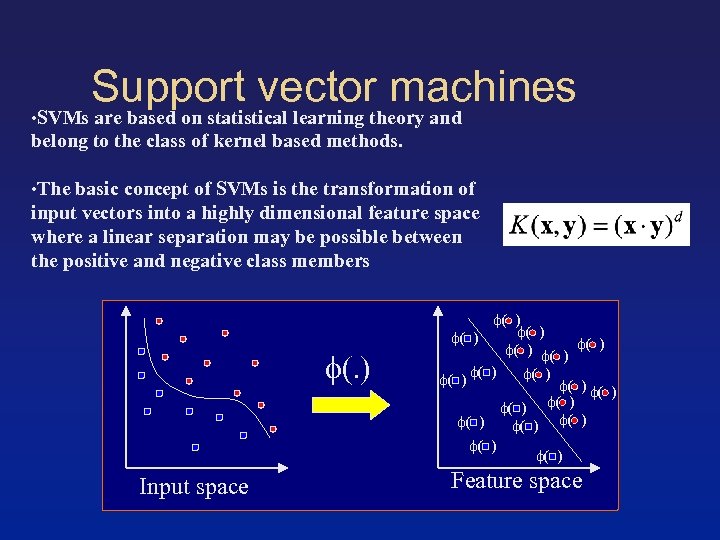 Support vector machines • SVMs are based on statistical learning theory and belong to the class of kernel based methods. • The basic concept of SVMs is the transformation of input vectors into a highly dimensional feature space where a linear separation may be possible between the positive and negative class members (. ) Input space ( ) ( ) ( ) ( ) ( ) Feature space
Support vector machines • SVMs are based on statistical learning theory and belong to the class of kernel based methods. • The basic concept of SVMs is the transformation of input vectors into a highly dimensional feature space where a linear separation may be possible between the positive and negative class members (. ) Input space ( ) ( ) ( ) ( ) ( ) Feature space
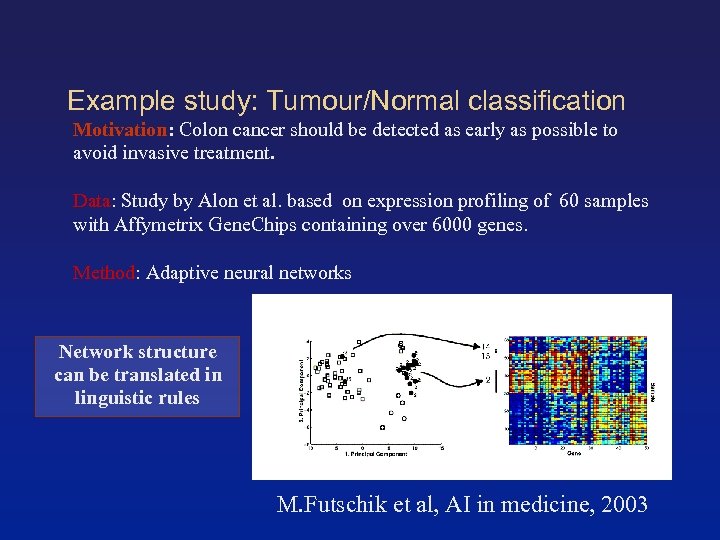 Example study: Tumour/Normal classification Motivation: Colon cancer should be detected as early as possible to avoid invasive treatment. Data: Study by Alon et al. based on expression profiling of 60 samples with Affymetrix Gene. Chips containing over 6000 genes. Method: Adaptive neural networks Network structure can be translated in linguistic rules M. Futschik et al, AI in medicine, 2003
Example study: Tumour/Normal classification Motivation: Colon cancer should be detected as early as possible to avoid invasive treatment. Data: Study by Alon et al. based on expression profiling of 60 samples with Affymetrix Gene. Chips containing over 6000 genes. Method: Adaptive neural networks Network structure can be translated in linguistic rules M. Futschik et al, AI in medicine, 2003
 Take-home messages III 1. Visualisation of results helps their interpretation 2. A lot of tools for classification are around – know their strength and weakness
Take-home messages III 1. Visualisation of results helps their interpretation 2. A lot of tools for classification are around – know their strength and weakness
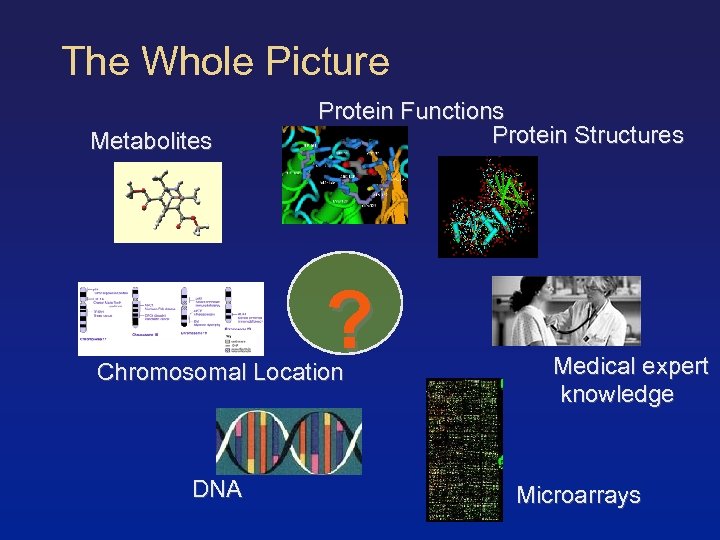 The Whole Picture Metabolites Protein Functions Protein Structures ? Chromosomal Location DNA Medical expert knowledge Microarrays
The Whole Picture Metabolites Protein Functions Protein Structures ? Chromosomal Location DNA Medical expert knowledge Microarrays
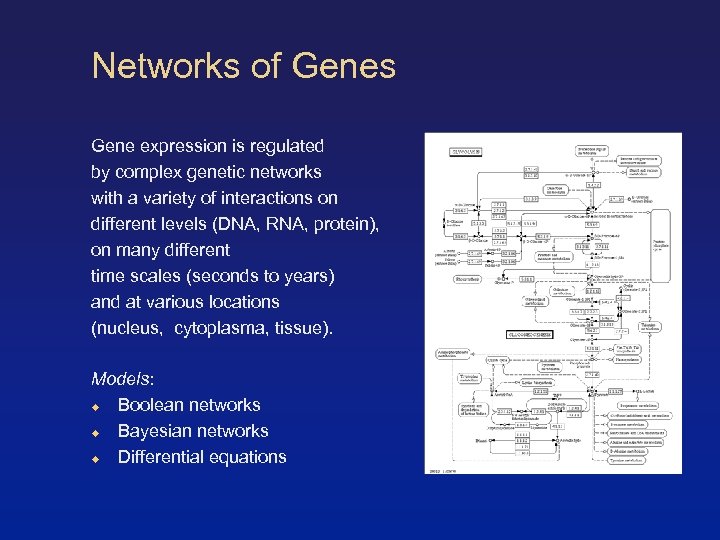 Networks of Genes Gene expression is regulated by complex genetic networks with a variety of interactions on different levels (DNA, RNA, protein), on many different time scales (seconds to years) and at various locations (nucleus, cytoplasma, tissue). Models: Boolean networks Bayesian networks Differential equations
Networks of Genes Gene expression is regulated by complex genetic networks with a variety of interactions on different levels (DNA, RNA, protein), on many different time scales (seconds to years) and at various locations (nucleus, cytoplasma, tissue). Models: Boolean networks Bayesian networks Differential equations
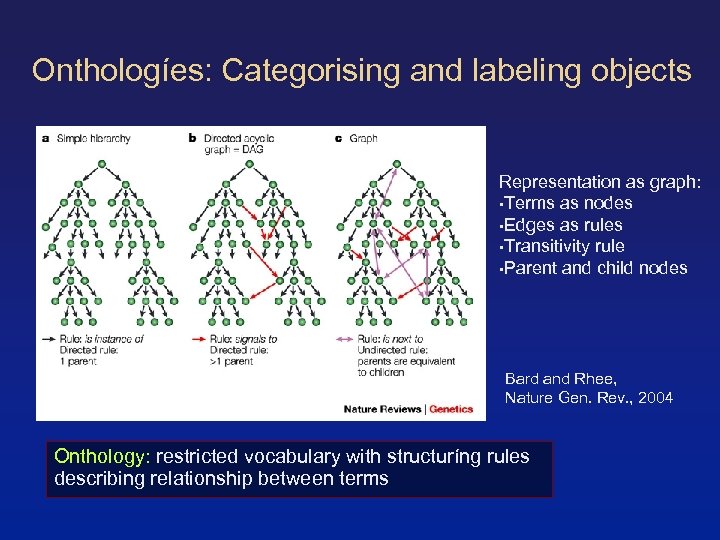 Onthologíes: Categorising and labeling objects Representation as graph: • Terms as nodes • Edges as rules • Transitivity rule • Parent and child nodes Bard and Rhee, Nature Gen. Rev. , 2004 Onthology: restricted vocabulary with structuríng rules describing relationship between terms
Onthologíes: Categorising and labeling objects Representation as graph: • Terms as nodes • Edges as rules • Transitivity rule • Parent and child nodes Bard and Rhee, Nature Gen. Rev. , 2004 Onthology: restricted vocabulary with structuríng rules describing relationship between terms
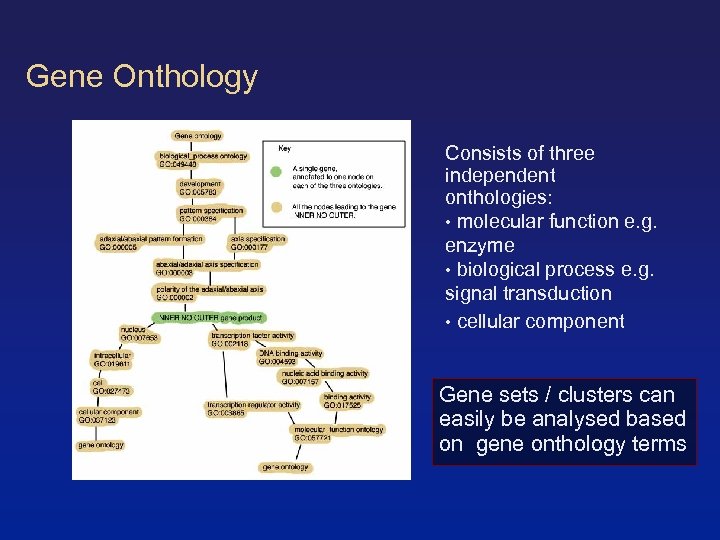 Gene Onthology Consists of three independent onthologies: • molecular function e. g. enzyme • biological process e. g. signal transduction • cellular component Gene sets / clusters can easily be analysed based on gene onthology terms
Gene Onthology Consists of three independent onthologies: • molecular function e. g. enzyme • biological process e. g. signal transduction • cellular component Gene sets / clusters can easily be analysed based on gene onthology terms
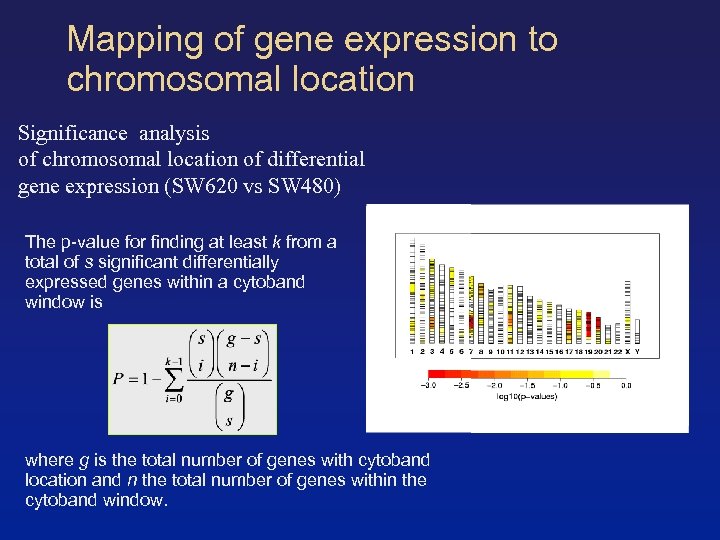 Mapping of gene expression to chromosomal location Significance analysis of chromosomal location of differential gene expression (SW 620 vs SW 480) The p-value for finding at least k from a total of s significant differentially expressed genes within a cytoband window is where g is the total number of genes with cytoband location and n the total number of genes within the cytoband window.
Mapping of gene expression to chromosomal location Significance analysis of chromosomal location of differential gene expression (SW 620 vs SW 480) The p-value for finding at least k from a total of s significant differentially expressed genes within a cytoband window is where g is the total number of genes with cytoband location and n the total number of genes within the cytoband window.
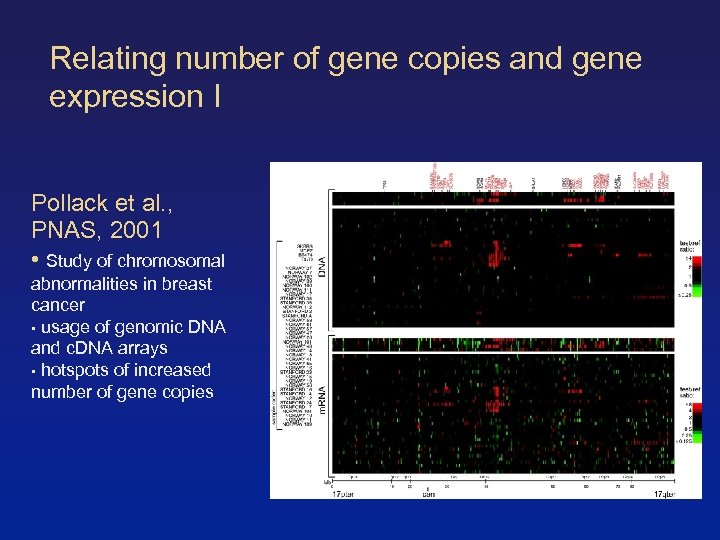 Relating number of gene copies and gene expression I Pollack et al. , PNAS, 2001 • Study of chromosomal abnormalities in breast cancer • usage of genomic DNA and c. DNA arrays • hotspots of increased number of gene copies
Relating number of gene copies and gene expression I Pollack et al. , PNAS, 2001 • Study of chromosomal abnormalities in breast cancer • usage of genomic DNA and c. DNA arrays • hotspots of increased number of gene copies
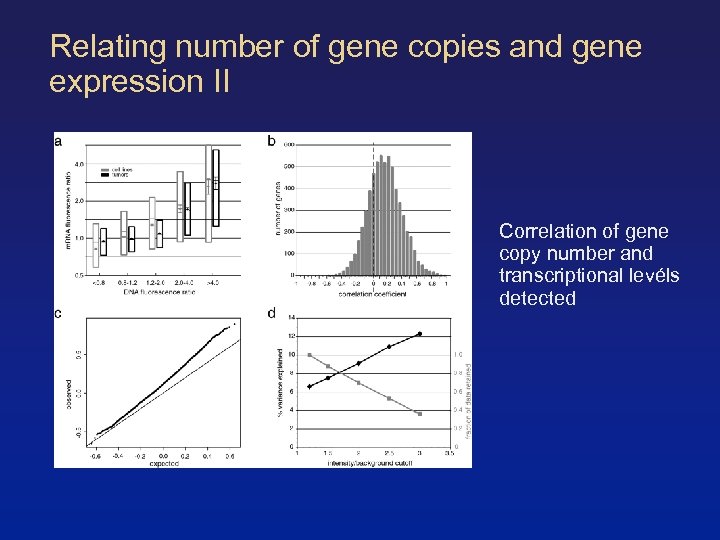 Relating number of gene copies and gene expression II Correlation of gene copy number and transcriptional levéls detected
Relating number of gene copies and gene expression II Correlation of gene copy number and transcriptional levéls detected
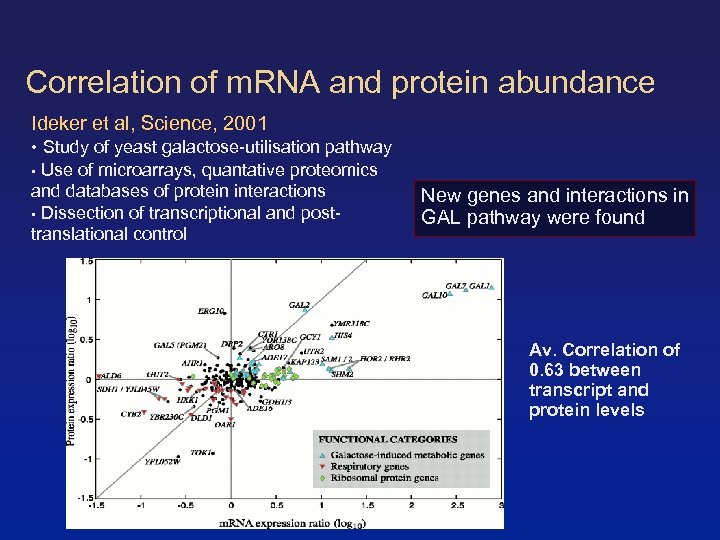 Correlation of m. RNA and protein abundance Ideker et al, Science, 2001 • Study of yeast galactose-utilisation pathway • Use of microarrays, quantative proteomics and databases of protein interactions • Dissection of transcriptional and posttranslational control New genes and interactions in GAL pathway were found Av. Correlation of 0. 63 between transcript and protein levels
Correlation of m. RNA and protein abundance Ideker et al, Science, 2001 • Study of yeast galactose-utilisation pathway • Use of microarrays, quantative proteomics and databases of protein interactions • Dissection of transcriptional and posttranslational control New genes and interactions in GAL pathway were found Av. Correlation of 0. 63 between transcript and protein levels
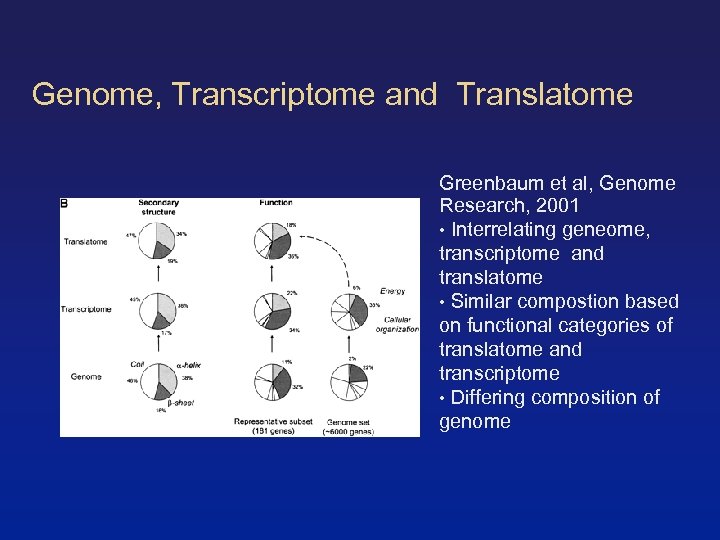 Genome, Transcriptome and Translatome Greenbaum et al, Genome Research, 2001 • Interrelating geneome, transcriptome and translatome • Similar compostion based on functional categories of translatome and transcriptome • Differing composition of genome
Genome, Transcriptome and Translatome Greenbaum et al, Genome Research, 2001 • Interrelating geneome, transcriptome and translatome • Similar compostion based on functional categories of translatome and transcriptome • Differing composition of genome
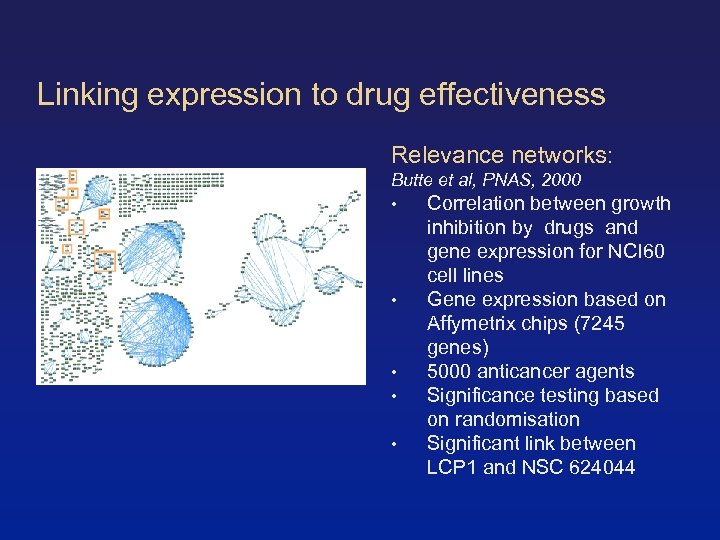 Linking expression to drug effectiveness Relevance networks: Butte et al, PNAS, 2000 • Correlation between growth • • inhibition by drugs and gene expression for NCI 60 cell lines Gene expression based on Affymetrix chips (7245 genes) 5000 anticancer agents Significance testing based on randomisation Significant link between LCP 1 and NSC 624044
Linking expression to drug effectiveness Relevance networks: Butte et al, PNAS, 2000 • Correlation between growth • • inhibition by drugs and gene expression for NCI 60 cell lines Gene expression based on Affymetrix chips (7245 genes) 5000 anticancer agents Significance testing based on randomisation Significant link between LCP 1 and NSC 624044
 Combinining gene expression data with clinical parameters Diffuse large B-cell Lymphoma (DLBCL) • Most common lymphoid malignancy in adults • Treatment by multi-agent chemotherapy • In case of a relapse: bone marrow transplantation • Clinical course of DLBCL is widely variable: Only 40% of treatments successful => Accurate outcome prediction is crucial for stratifying patients for intensified therapy
Combinining gene expression data with clinical parameters Diffuse large B-cell Lymphoma (DLBCL) • Most common lymphoid malignancy in adults • Treatment by multi-agent chemotherapy • In case of a relapse: bone marrow transplantation • Clinical course of DLBCL is widely variable: Only 40% of treatments successful => Accurate outcome prediction is crucial for stratifying patients for intensified therapy
 Case study: DLBCL Current prognostic model: International Prediction Index (IPI) Alternative: Microarray-based prediction of treatment outcome DLBCL study by Shipp et al. (Nature Medicine, 2002, 8(1): 68 -74) • expression profiles of 58 patients using Hu 6800 Affymetrix chips (corresponding to ca. 6800 genes) • Prediction accuracy of outcome using leave-one-out procedure: Knn: 70. 7%; WV: 75. 9%; SVM: 77. 6%
Case study: DLBCL Current prognostic model: International Prediction Index (IPI) Alternative: Microarray-based prediction of treatment outcome DLBCL study by Shipp et al. (Nature Medicine, 2002, 8(1): 68 -74) • expression profiles of 58 patients using Hu 6800 Affymetrix chips (corresponding to ca. 6800 genes) • Prediction accuracy of outcome using leave-one-out procedure: Knn: 70. 7%; WV: 75. 9%; SVM: 77. 6%
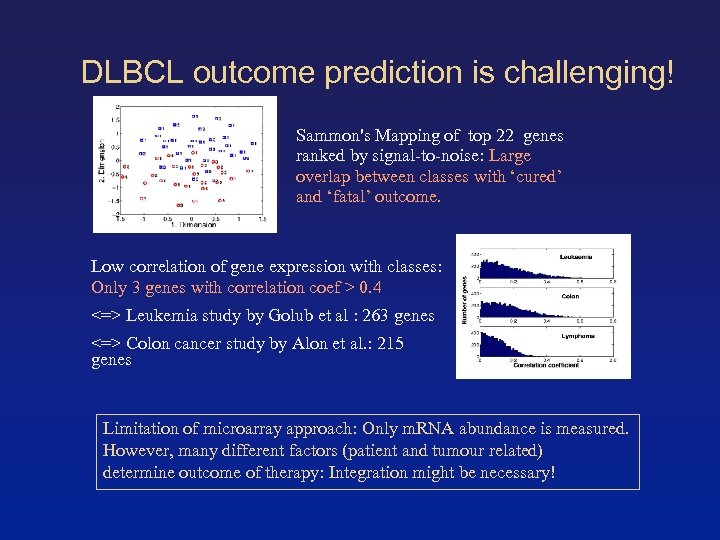 DLBCL outcome prediction is challenging! Sammon's Mapping of top 22 genes ranked by signal-to-noise: Large overlap between classes with ‘cured’ and ‘fatal’ outcome. Low correlation of gene expression with classes: Only 3 genes with correlation coef > 0. 4 <=> Leukemia study by Golub et al : 263 genes <=> Colon cancer study by Alon et al. : 215 genes Limitation of microarray approach: Only m. RNA abundance is measured. However, many different factors (patient and tumour related) determine outcome of therapy: Integration might be necessary!
DLBCL outcome prediction is challenging! Sammon's Mapping of top 22 genes ranked by signal-to-noise: Large overlap between classes with ‘cured’ and ‘fatal’ outcome. Low correlation of gene expression with classes: Only 3 genes with correlation coef > 0. 4 <=> Leukemia study by Golub et al : 263 genes <=> Colon cancer study by Alon et al. : 215 genes Limitation of microarray approach: Only m. RNA abundance is measured. However, many different factors (patient and tumour related) determine outcome of therapy: Integration might be necessary!
 Prognostic models for DLBCL Clinical predictor: • IPI based on five risk factors (age, tumour stage, patient’s performance, number of extranodal sites, LDH concentration) • Survival rate determined in clinical study: Low risk: 73%, low-intermediate: 51%, intermediate-high: 42%, high: 26% • Conversion of IPI into Bayesian classifier using survival rates as conditional probabilities P: e. g. Sample belongs to class ‘cured’ if P(‘cured’|IPI)> P(‘fatal’|IPI) => Overall accuracy of 73. 2%.
Prognostic models for DLBCL Clinical predictor: • IPI based on five risk factors (age, tumour stage, patient’s performance, number of extranodal sites, LDH concentration) • Survival rate determined in clinical study: Low risk: 73%, low-intermediate: 51%, intermediate-high: 42%, high: 26% • Conversion of IPI into Bayesian classifier using survival rates as conditional probabilities P: e. g. Sample belongs to class ‘cured’ if P(‘cured’|IPI)> P(‘fatal’|IPI) => Overall accuracy of 73. 2%.
 Prognostic models for DLBCL Microarray-based predictor: • Identifies clusters by unsupervised learning • Supervised classification • Efu. NN as five layered neural network • Based on 17 genes using signal-to-noise criterion • Accuracy using leave-one-out: 78. 5%
Prognostic models for DLBCL Microarray-based predictor: • Identifies clusters by unsupervised learning • Supervised classification • Efu. NN as five layered neural network • Based on 17 genes using signal-to-noise criterion • Accuracy using leave-one-out: 78. 5%
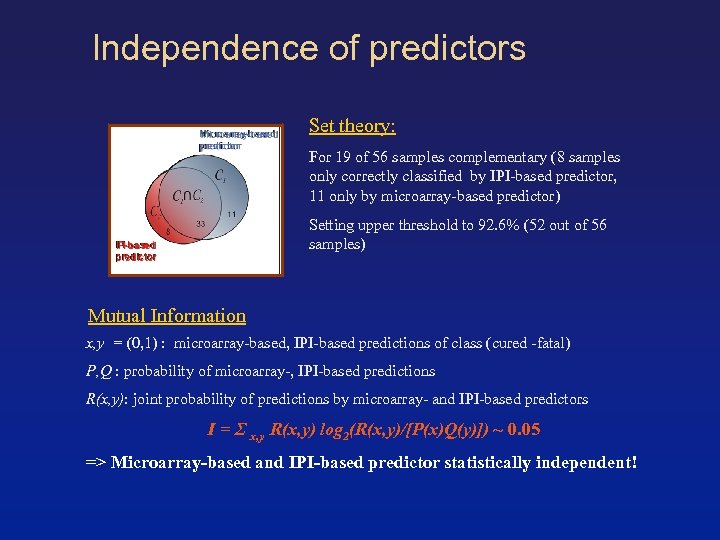 Independence of predictors Set theory: For 19 of 56 samples complementary (8 samples only correctly classified by IPI-based predictor, 11 only by microarray-based predictor) Setting upper threshold to 92. 6% (52 out of 56 samples) Mutual Information x, y = (0, 1) : microarray-based, IPI-based predictions of class (cured -fatal) P, Q : probability of microarray-, IPI-based predictions R(x, y): joint probability of predictions by microarray- and IPI-based predictors I = Σ x, y R(x, y) log 2(R(x, y)/[P(x)Q(y)]) ~ 0. 05 => Microarray-based and IPI-based predictor statistically independent!
Independence of predictors Set theory: For 19 of 56 samples complementary (8 samples only correctly classified by IPI-based predictor, 11 only by microarray-based predictor) Setting upper threshold to 92. 6% (52 out of 56 samples) Mutual Information x, y = (0, 1) : microarray-based, IPI-based predictions of class (cured -fatal) P, Q : probability of microarray-, IPI-based predictions R(x, y): joint probability of predictions by microarray- and IPI-based predictors I = Σ x, y R(x, y) log 2(R(x, y)/[P(x)Q(y)]) ~ 0. 05 => Microarray-based and IPI-based predictor statistically independent!
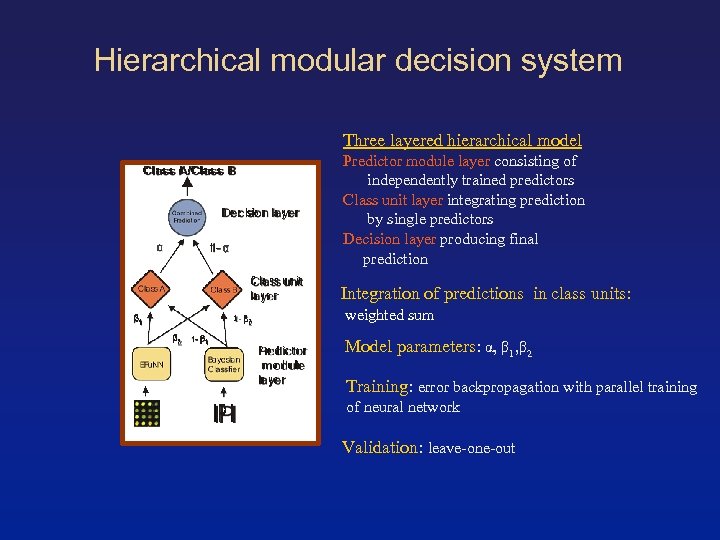 Hierarchical modular decision system Three layered hierarchical model Predictor module layer consisting of independently trained predictors Class unit layer integrating prediction by single predictors Decision layer producing final prediction Integration of predictions in class units: weighted sum Model parameters: α, β 1, β 2 Training: error backpropagation with parallel training of neural network Validation: leave-one-out
Hierarchical modular decision system Three layered hierarchical model Predictor module layer consisting of independently trained predictors Class unit layer integrating prediction by single predictors Decision layer producing final prediction Integration of predictions in class units: weighted sum Model parameters: α, β 1, β 2 Training: error backpropagation with parallel training of neural network Validation: leave-one-out
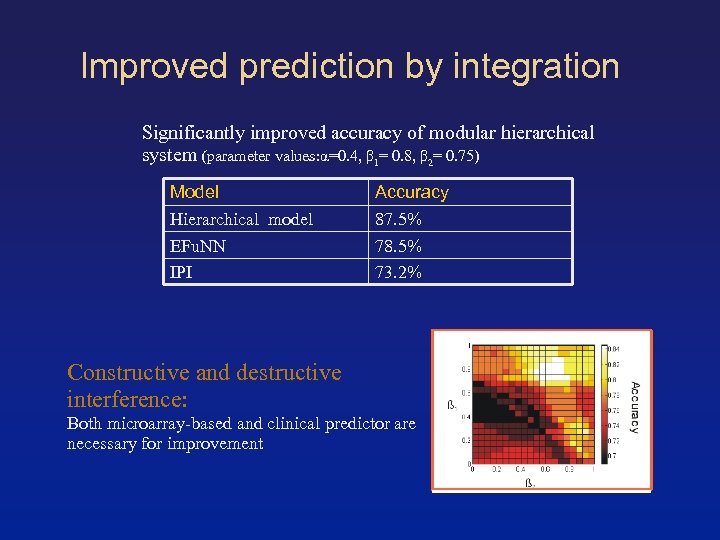 Improved prediction by integration Significantly improved accuracy of modular hierarchical system (parameter values: α=0. 4, β 1= 0. 8, β 2= 0. 75) Model Hierarchical model EFu. NN IPI Accuracy 87. 5% 78. 5% 73. 2% Constructive and destructive interference: Both microarray-based and clinical predictor are necessary for improvement
Improved prediction by integration Significantly improved accuracy of modular hierarchical system (parameter values: α=0. 4, β 1= 0. 8, β 2= 0. 75) Model Hierarchical model EFu. NN IPI Accuracy 87. 5% 78. 5% 73. 2% Constructive and destructive interference: Both microarray-based and clinical predictor are necessary for improvement
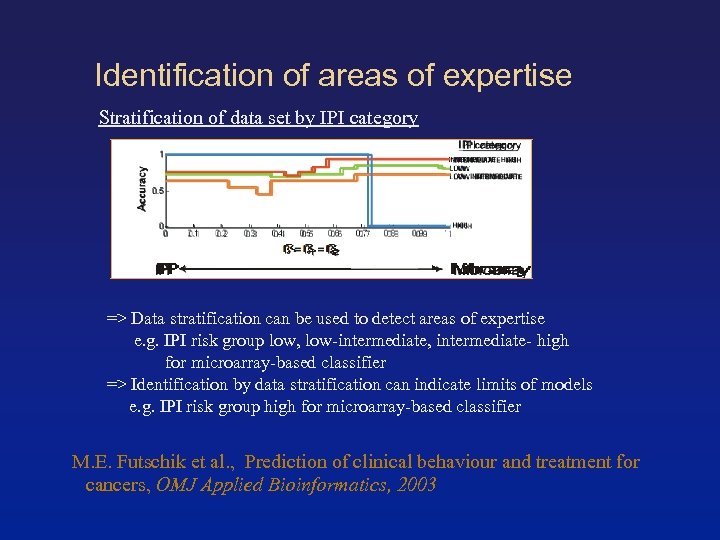 Identification of areas of expertise Stratification of data set by IPI category => Data stratification can be used to detect areas of expertise e. g. IPI risk group low, low-intermediate, intermediate- high for microarray-based classifier => Identification by data stratification can indicate limits of models e. g. IPI risk group high for microarray-based classifier M. E. Futschik et al. , Prediction of clinical behaviour and treatment for cancers, OMJ Applied Bioinformatics, 2003
Identification of areas of expertise Stratification of data set by IPI category => Data stratification can be used to detect areas of expertise e. g. IPI risk group low, low-intermediate, intermediate- high for microarray-based classifier => Identification by data stratification can indicate limits of models e. g. IPI risk group high for microarray-based classifier M. E. Futschik et al. , Prediction of clinical behaviour and treatment for cancers, OMJ Applied Bioinformatics, 2003
 TEŞEKKÜR !
TEŞEKKÜR !


