450d8c4986421af8f6753c61bc3cce6a.ppt
- Количество слайдов: 70
 MGH-PGA enomic Analysis of Stress and Inflammatio Pseudomonas aeruginosa Infection Nicole T. Liberati, Dan G. Lee, Jacinto M. Villanueva and Frederick M. Ausubel Department of Molecular Biology Massachusetts General Hospital Carolyn Cannon, Fadie Coleman, Mike Kowalski Jeff Lyczak, Martin Lee, Gloria Meluleni, and Gerald Pier Channing Laboratory Brigham and Women’s Hospital
MGH-PGA enomic Analysis of Stress and Inflammatio Pseudomonas aeruginosa Infection Nicole T. Liberati, Dan G. Lee, Jacinto M. Villanueva and Frederick M. Ausubel Department of Molecular Biology Massachusetts General Hospital Carolyn Cannon, Fadie Coleman, Mike Kowalski Jeff Lyczak, Martin Lee, Gloria Meluleni, and Gerald Pier Channing Laboratory Brigham and Women’s Hospital
 Ausubel/Pier PGA Project
Ausubel/Pier PGA Project
 Contents of Slide Show: Section I: Background Information on Multi Host Pathogenesis System Section II: Background Information on Screening Methodology and Rationale for Constructing the Uni-Gene Library Section III: Progress Report on Uni-Gene Library Construction and Detailed Methodology Section IV: Development of a Linux My. SQL Uni-Gene Library Relational Database Section V: CF Mouse Oropharynx Colonization Model
Contents of Slide Show: Section I: Background Information on Multi Host Pathogenesis System Section II: Background Information on Screening Methodology and Rationale for Constructing the Uni-Gene Library Section III: Progress Report on Uni-Gene Library Construction and Detailed Methodology Section IV: Development of a Linux My. SQL Uni-Gene Library Relational Database Section V: CF Mouse Oropharynx Colonization Model
 Section I Background Information on Multi-Host Pathogenesis System
Section I Background Information on Multi-Host Pathogenesis System
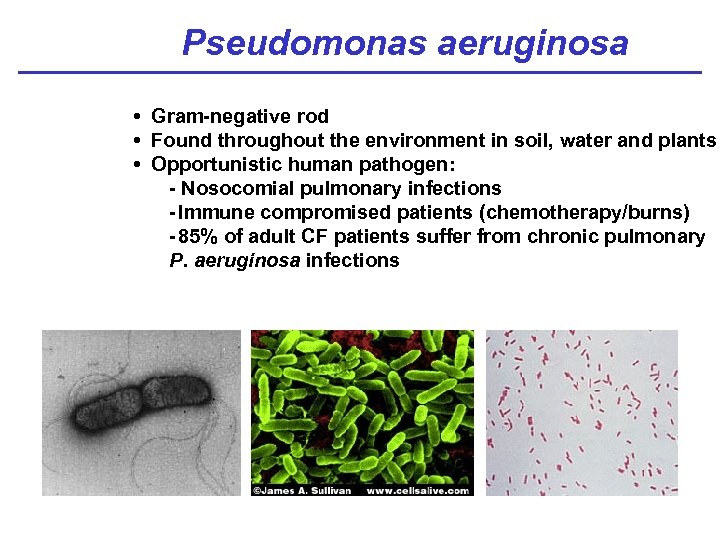 Pseudomonas aeruginosa • Gram-negative rod • Found throughout the environment in soil, water and plants • Opportunistic human pathogen: - Nosocomial pulmonary infections - Immune compromised patients (chemotherapy/burns) - 85% of adult CF patients suffer from chronic pulmonary P. aeruginosa infections
Pseudomonas aeruginosa • Gram-negative rod • Found throughout the environment in soil, water and plants • Opportunistic human pathogen: - Nosocomial pulmonary infections - Immune compromised patients (chemotherapy/burns) - 85% of adult CF patients suffer from chronic pulmonary P. aeruginosa infections
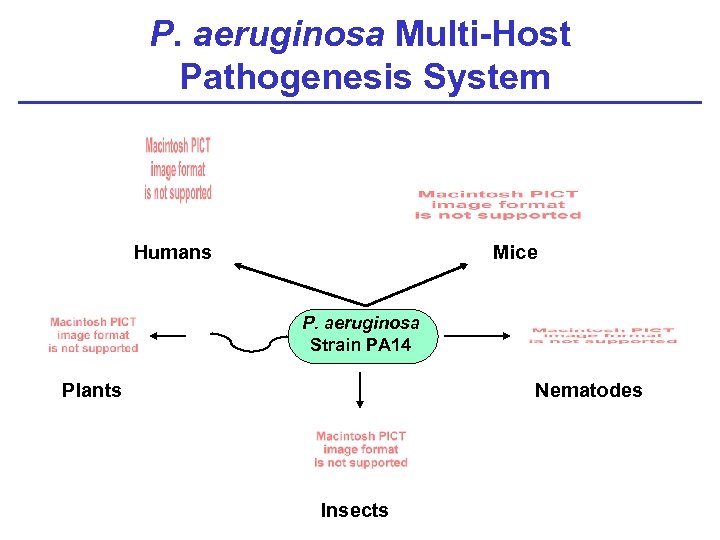 P. aeruginosa Multi-Host Pathogenesis System Humans Mice P. aeruginosa Strain PA 14 Plants Nematodes Insects
P. aeruginosa Multi-Host Pathogenesis System Humans Mice P. aeruginosa Strain PA 14 Plants Nematodes Insects
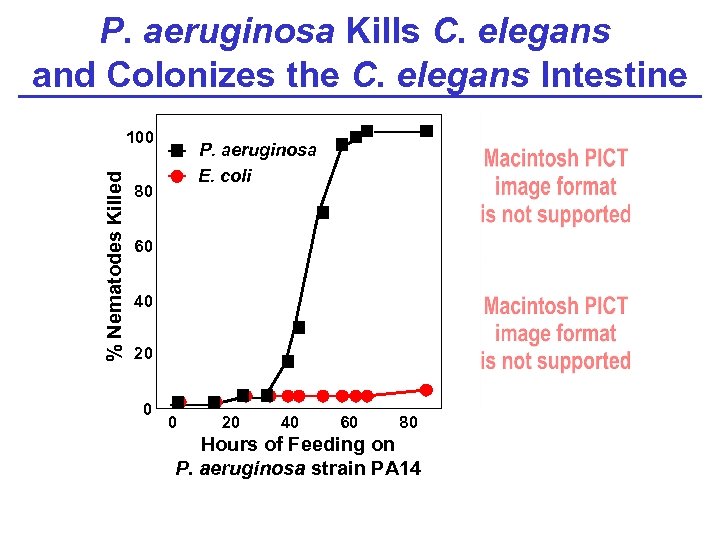 P. aeruginosa Kills C. elegans and Colonizes the C. elegans Intestine % Nematodes Killed 100 P. aeruginosa E. coli 80 60 40 20 0 0 20 40 60 80 Hours of Feeding on P. aeruginosa strain PA 14
P. aeruginosa Kills C. elegans and Colonizes the C. elegans Intestine % Nematodes Killed 100 P. aeruginosa E. coli 80 60 40 20 0 0 20 40 60 80 Hours of Feeding on P. aeruginosa strain PA 14
 P. aeruginosa Kills Galleria mellonella (Wax Moth Caterpillar) Larvae; LD 50 = 1
P. aeruginosa Kills Galleria mellonella (Wax Moth Caterpillar) Larvae; LD 50 = 1
 Section II Background Information on Screening Methodology and Rationale for Constructing the Uni-Gene Library
Section II Background Information on Screening Methodology and Rationale for Constructing the Uni-Gene Library
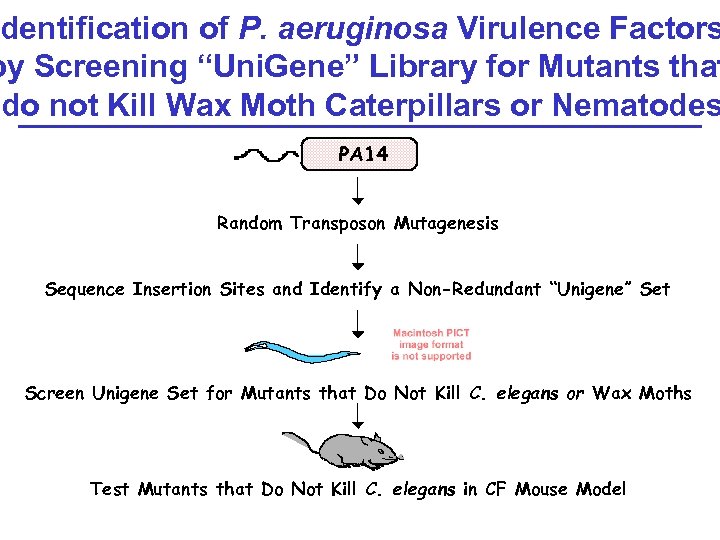 dentification of P. aeruginosa Virulence Factors by Screening “Uni. Gene” Library for Mutants that do not Kill Wax Moth Caterpillars or Nematodes PA 14 Random Transposon Mutagenesis Sequence Insertion Sites and Identify a Non-Redundant “Unigene” Set Screen Unigene Set for Mutants that Do Not Kill C. elegans or Wax Moths Test Mutants that Do Not Kill C. elegans in CF Mouse Model
dentification of P. aeruginosa Virulence Factors by Screening “Uni. Gene” Library for Mutants that do not Kill Wax Moth Caterpillars or Nematodes PA 14 Random Transposon Mutagenesis Sequence Insertion Sites and Identify a Non-Redundant “Unigene” Set Screen Unigene Set for Mutants that Do Not Kill C. elegans or Wax Moths Test Mutants that Do Not Kill C. elegans in CF Mouse Model
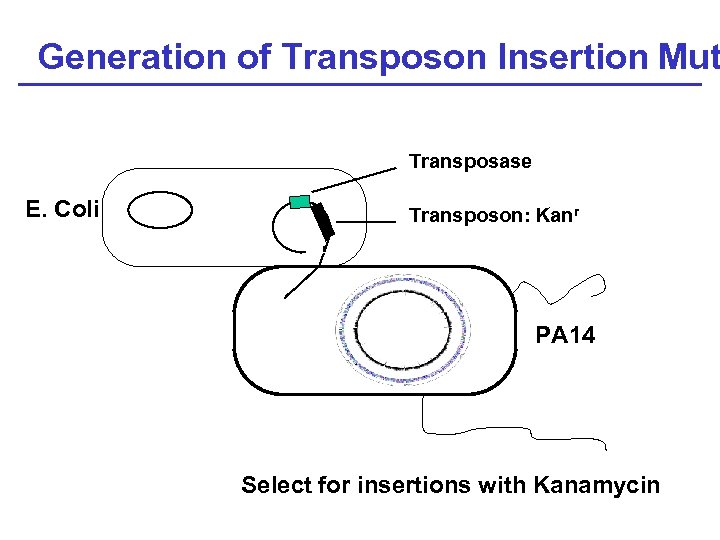 Generation of Transposon Insertion Mut Transposase E. Coli Transposon: Kanr 6 Mb PA 14 Select for insertions with Kanamycin
Generation of Transposon Insertion Mut Transposase E. Coli Transposon: Kanr 6 Mb PA 14 Select for insertions with Kanamycin
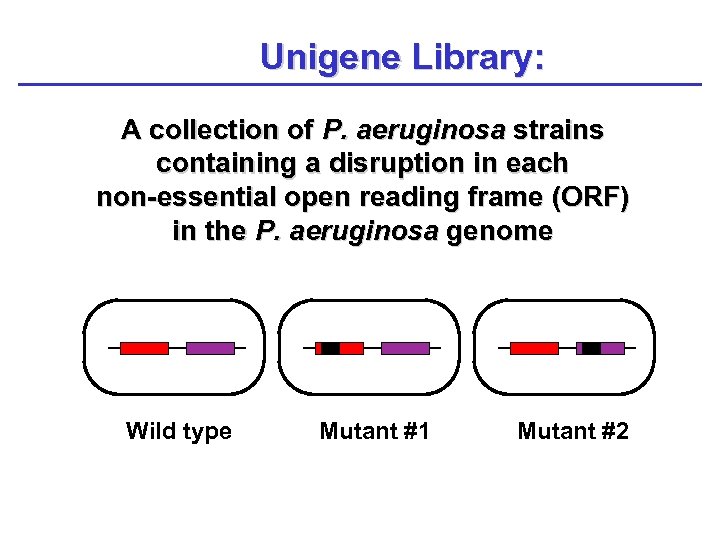 Unigene Library: A collection of P. aeruginosa strains containing a disruption in each non-essential open reading frame (ORF) in the P. aeruginosa genome Wild type Mutant #1 Mutant #2
Unigene Library: A collection of P. aeruginosa strains containing a disruption in each non-essential open reading frame (ORF) in the P. aeruginosa genome Wild type Mutant #1 Mutant #2
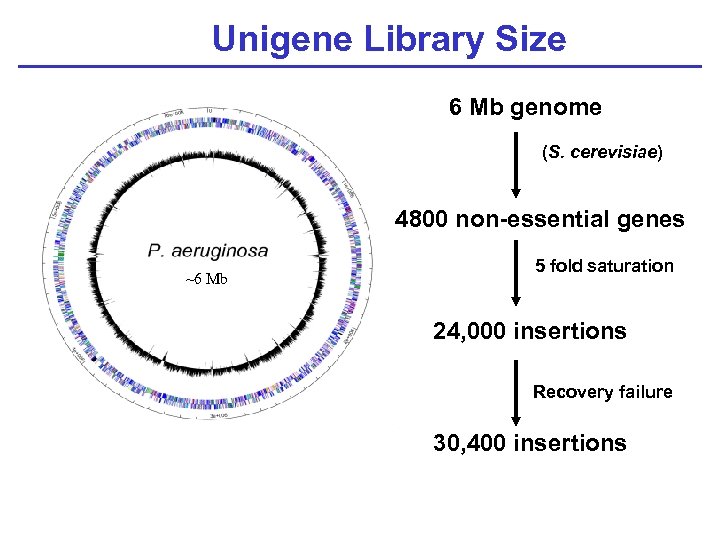 Unigene Library Size 6 Mb genome (S. cerevisiae) 4800 non-essential genes ~6 Mb 5 fold saturation 24, 000 insertions Recovery failure 30, 400 insertions
Unigene Library Size 6 Mb genome (S. cerevisiae) 4800 non-essential genes ~6 Mb 5 fold saturation 24, 000 insertions Recovery failure 30, 400 insertions
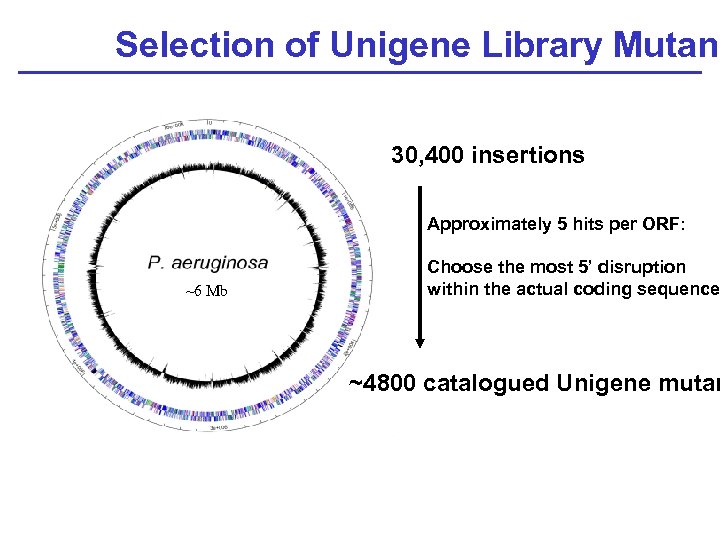 Selection of Unigene Library Mutant 30, 400 insertions Approximately 5 hits per ORF: ~6 Mb Choose the most 5’ disruption within the actual coding sequence ~4800 catalogued Unigene mutan
Selection of Unigene Library Mutant 30, 400 insertions Approximately 5 hits per ORF: ~6 Mb Choose the most 5’ disruption within the actual coding sequence ~4800 catalogued Unigene mutan
 Advantages of Unigene Library Screen 1) Mutation previously identified 2) Limited number of mutants to screen (4800) 3) Non-redundant mutations 4) Built-in confirmation of the involvement of known pathways. 5) Easy to confirm the importance of the mutated gene using other mutant alleles.
Advantages of Unigene Library Screen 1) Mutation previously identified 2) Limited number of mutants to screen (4800) 3) Non-redundant mutations 4) Built-in confirmation of the involvement of known pathways. 5) Easy to confirm the importance of the mutated gene using other mutant alleles.
 Section III Progress Report on Uni-Gene Library Construction and Detailed Methodology
Section III Progress Report on Uni-Gene Library Construction and Detailed Methodology
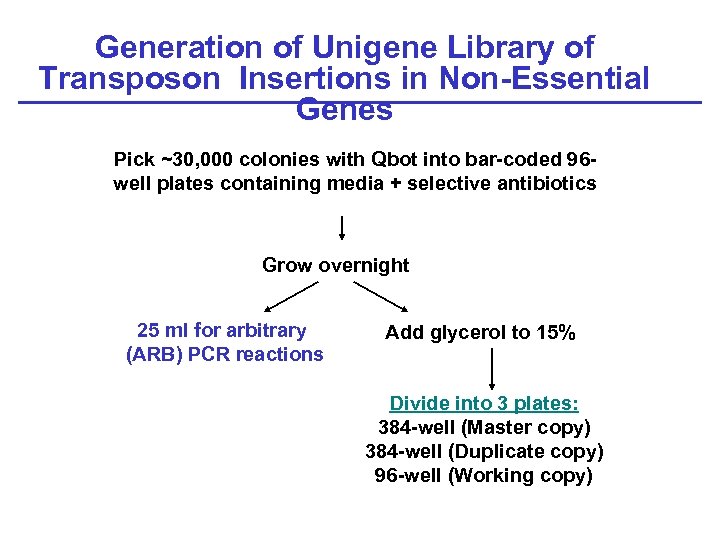 Generation of Unigene Library of Transposon Insertions in Non-Essential Genes Pick ~30, 000 colonies with Qbot into bar-coded 96 well plates containing media + selective antibiotics Grow overnight 25 ml for arbitrary (ARB) PCR reactions Add glycerol to 15% Divide into 3 plates: 384 -well (Master copy) 384 -well (Duplicate copy) 96 -well (Working copy)
Generation of Unigene Library of Transposon Insertions in Non-Essential Genes Pick ~30, 000 colonies with Qbot into bar-coded 96 well plates containing media + selective antibiotics Grow overnight 25 ml for arbitrary (ARB) PCR reactions Add glycerol to 15% Divide into 3 plates: 384 -well (Master copy) 384 -well (Duplicate copy) 96 -well (Working copy)
 Current Status of the Unigene Libra 1) 48 x 96 (4608) mutants created. 2) 60% of the insertion sites identified. 2) Insertion site identification protocol optimized. 3) (1152 mutants created and identified in 2. 5 we 1) 3) Accompanying database is operational. 2) Quality assurance testing is in progress.
Current Status of the Unigene Libra 1) 48 x 96 (4608) mutants created. 2) 60% of the insertion sites identified. 2) Insertion site identification protocol optimized. 3) (1152 mutants created and identified in 2. 5 we 1) 3) Accompanying database is operational. 2) Quality assurance testing is in progress.
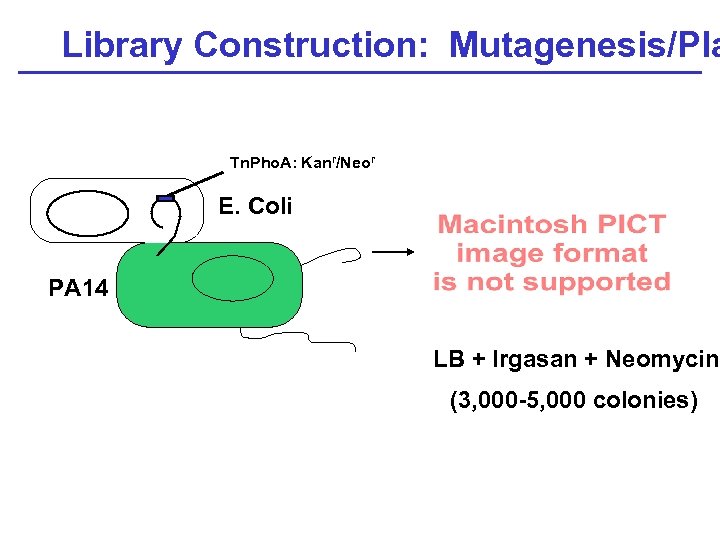 Library Construction: Mutagenesis/Pla Tn. Pho. A: Kanr/Neor E. Coli PA 14 LB + Irgasan + Neomycin (3, 000 -5, 000 colonies)
Library Construction: Mutagenesis/Pla Tn. Pho. A: Kanr/Neor E. Coli PA 14 LB + Irgasan + Neomycin (3, 000 -5, 000 colonies)
![Library Construction: Colony Picking/Cu • Inoculate 250 µL LB + Irgasan [50 µg/m. L] Library Construction: Colony Picking/Cu • Inoculate 250 µL LB + Irgasan [50 µg/m. L]](https://present5.com/presentation/450d8c4986421af8f6753c61bc3cce6a/image-20.jpg) Library Construction: Colony Picking/Cu • Inoculate 250 µL LB + Irgasan [50 µg/m. L] Kanamycin [200 µg/m. L]* • Grow 40 hrs at 37°C (no shaking)
Library Construction: Colony Picking/Cu • Inoculate 250 µL LB + Irgasan [50 µg/m. L] Kanamycin [200 µg/m. L]* • Grow 40 hrs at 37°C (no shaking)
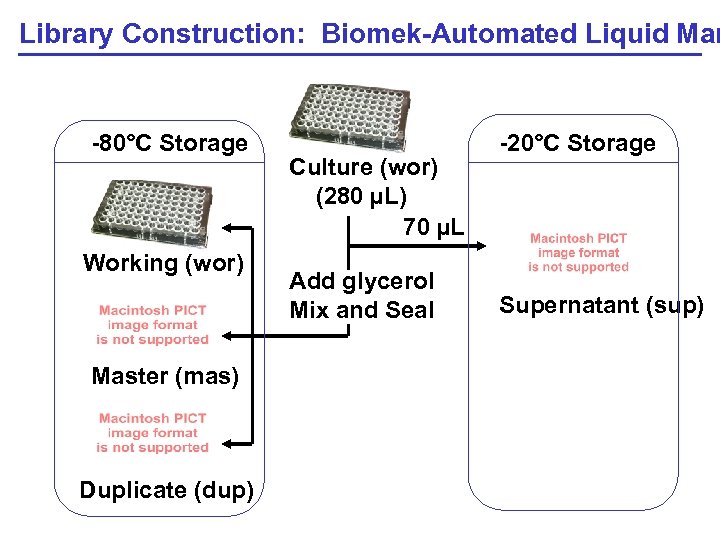 Library Construction: Biomek-Automated Liquid Man -80°C Storage Working (wor) Master (mas) Duplicate (dup) Culture (wor) (280 µL) 70 µL Add glycerol Mix and Seal -20°C Storage Supernatant (sup)
Library Construction: Biomek-Automated Liquid Man -80°C Storage Working (wor) Master (mas) Duplicate (dup) Culture (wor) (280 µL) 70 µL Add glycerol Mix and Seal -20°C Storage Supernatant (sup)
 Library Construction: Bar Coding B Side: Human Readab A Side: Unique ID# PA 14_Pho. A_100_xxx wor sup mas dup ar 1 ar 2 seq
Library Construction: Bar Coding B Side: Human Readab A Side: Unique ID# PA 14_Pho. A_100_xxx wor sup mas dup ar 1 ar 2 seq
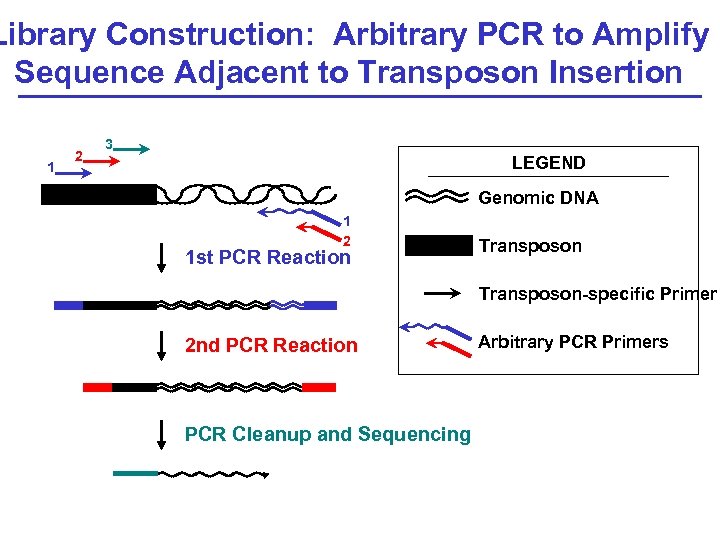 Library Construction: Arbitrary PCR to Amplify Sequence Adjacent to Transposon Insertion 1 2 3 LEGEND Genomic DNA 1 2 1 st PCR Reaction Transposon-specific Primer 2 nd PCR Reaction PCR Cleanup and Sequencing Arbitrary PCR Primers
Library Construction: Arbitrary PCR to Amplify Sequence Adjacent to Transposon Insertion 1 2 3 LEGEND Genomic DNA 1 2 1 st PCR Reaction Transposon-specific Primer 2 nd PCR Reaction PCR Cleanup and Sequencing Arbitrary PCR Primers
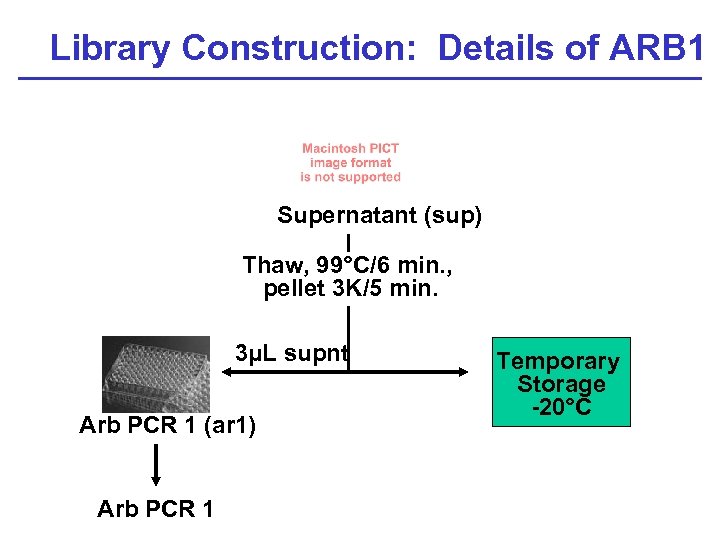 Library Construction: Details of ARB 1 P Supernatant (sup) Thaw, 99°C/6 min. , pellet 3 K/5 min. 3µL supnt Arb PCR 1 (ar 1) Arb PCR 1 Temporary Storage -20°C
Library Construction: Details of ARB 1 P Supernatant (sup) Thaw, 99°C/6 min. , pellet 3 K/5 min. 3µL supnt Arb PCR 1 (ar 1) Arb PCR 1 Temporary Storage -20°C
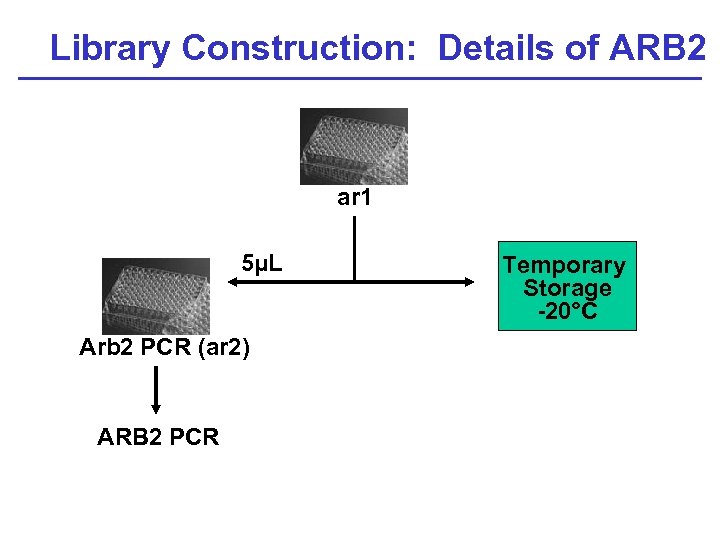 Library Construction: Details of ARB 2 P ar 1 5µL Arb 2 PCR (ar 2) ARB 2 PCR Temporary Storage -20°C
Library Construction: Details of ARB 2 P ar 1 5µL Arb 2 PCR (ar 2) ARB 2 PCR Temporary Storage -20°C
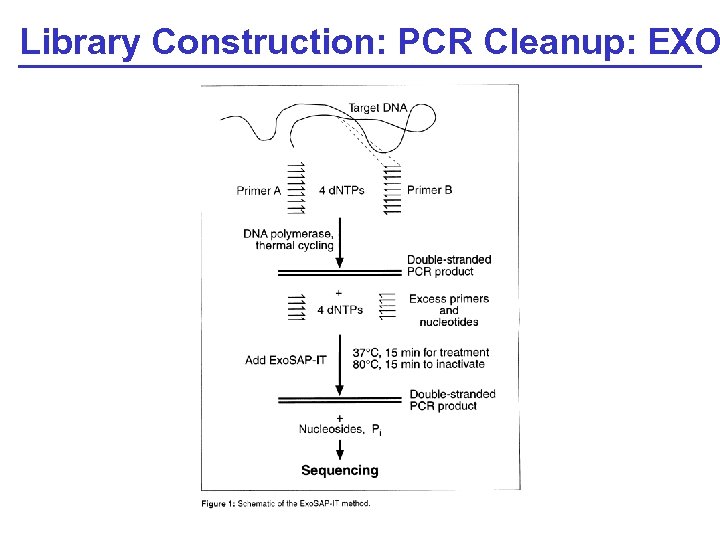 Library Construction: PCR Cleanup: EXO
Library Construction: PCR Cleanup: EXO
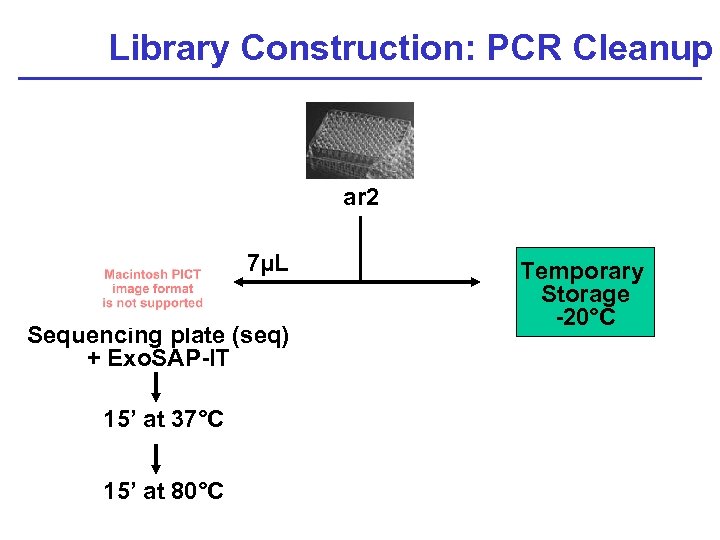 Library Construction: PCR Cleanup ar 2 7µL Sequencing plate (seq) + Exo. SAP-IT 15’ at 37°C 15’ at 80°C Temporary Storage -20°C
Library Construction: PCR Cleanup ar 2 7µL Sequencing plate (seq) + Exo. SAP-IT 15’ at 37°C 15’ at 80°C Temporary Storage -20°C
![Library Construction: Sequencing seq Add Sequencing primer to a [final] of 5 ng/µL and Library Construction: Sequencing seq Add Sequencing primer to a [final] of 5 ng/µL and](https://present5.com/presentation/450d8c4986421af8f6753c61bc3cce6a/image-28.jpg) Library Construction: Sequencing seq Add Sequencing primer to a [final] of 5 ng/µL and Seal Send to DNA Core for Sequencing (Store at 4°C)
Library Construction: Sequencing seq Add Sequencing primer to a [final] of 5 ng/µL and Seal Send to DNA Core for Sequencing (Store at 4°C)
 Example of High Quality Sequence Tn. Pho. A Sequence Length + Mixed + Tn. Pho. A = Sequence Success
Example of High Quality Sequence Tn. Pho. A Sequence Length + Mixed + Tn. Pho. A = Sequence Success
 Example of Low Quality Sequence Tn. Pho. A
Example of Low Quality Sequence Tn. Pho. A
![Sequencing Success Index Optimization of [Taq] in Sequencing Reactio Sequencing Success Index Optimization of [Taq] in Sequencing Reactio](https://present5.com/presentation/450d8c4986421af8f6753c61bc3cce6a/image-31.jpg) Sequencing Success Index Optimization of [Taq] in Sequencing Reactio
Sequencing Success Index Optimization of [Taq] in Sequencing Reactio
 Unigene Library Mutant Identification Optimiz • Taq Manufacturer Roche vs. Promega vs. Prepared Master Mixes • Final Taq Concentration 1. 25 U sufficient • PCR Master Mix Preparations Fresh Master Mixes vs Stored (4°C) Master Mix • Hybaid vs. MJ Research PCR Machines • PCR Cleanup Protocol Exo. SAP-IT vs. Clontech Nucleo. Spin
Unigene Library Mutant Identification Optimiz • Taq Manufacturer Roche vs. Promega vs. Prepared Master Mixes • Final Taq Concentration 1. 25 U sufficient • PCR Master Mix Preparations Fresh Master Mixes vs Stored (4°C) Master Mix • Hybaid vs. MJ Research PCR Machines • PCR Cleanup Protocol Exo. SAP-IT vs. Clontech Nucleo. Spin
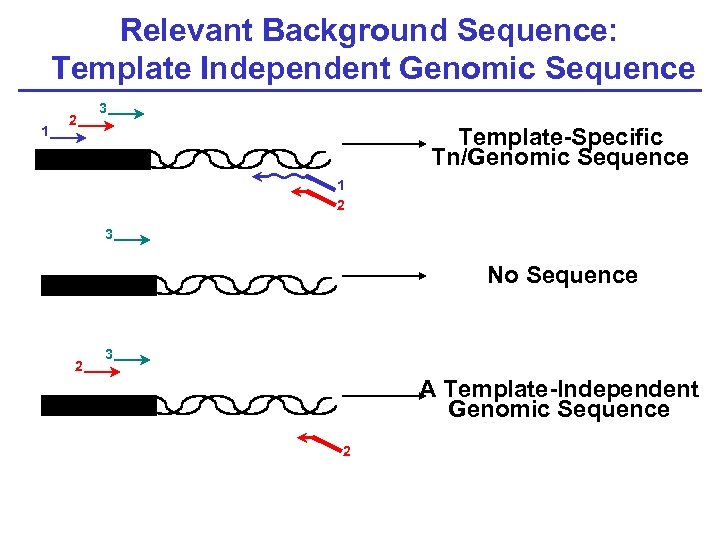 Relevant Background Sequence: Template Independent Genomic Sequence 1 2 3 Template-Specific Tn/Genomic Sequence 1 2 3 No Sequence 2 3 A Template-Independent Genomic Sequence 2
Relevant Background Sequence: Template Independent Genomic Sequence 1 2 3 Template-Specific Tn/Genomic Sequence 1 2 3 No Sequence 2 3 A Template-Independent Genomic Sequence 2
 High Quality Sequence (cont’d) NNNNN ARB PRIMER Sequence
High Quality Sequence (cont’d) NNNNN ARB PRIMER Sequence
 Trouble Shooting: Buried ARB Seque NNNNN ARB PRIMER Sequence High Quality Sequence
Trouble Shooting: Buried ARB Seque NNNNN ARB PRIMER Sequence High Quality Sequence
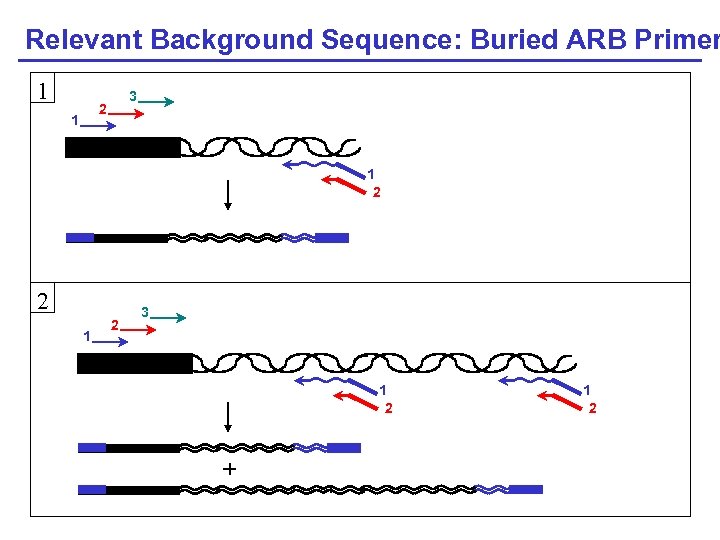 Relevant Background Sequence: Buried ARB Primer 1 3 2 1 1 2 2 1 2 3 1 2 + 1 2
Relevant Background Sequence: Buried ARB Primer 1 3 2 1 1 2 2 1 2 3 1 2 + 1 2
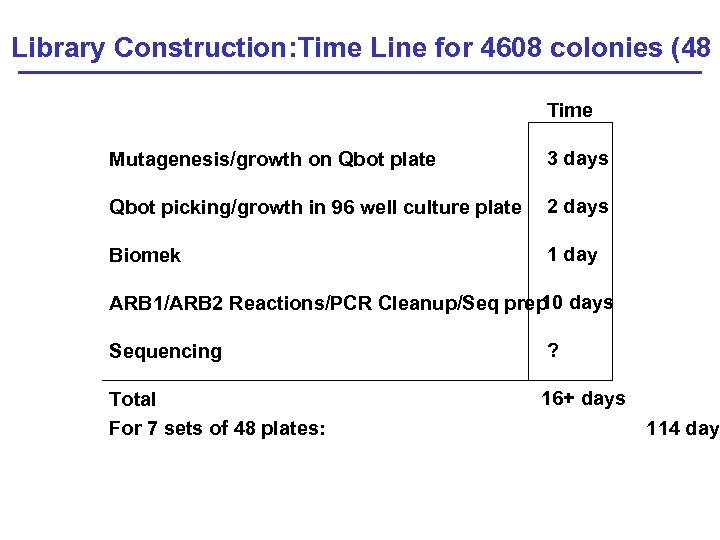 Library Construction: Time Line for 4608 colonies (48 s Time Mutagenesis/growth on Qbot plate 3 days Qbot picking/growth in 96 well culture plate 2 days Biomek 1 day 10 ARB 1/ARB 2 Reactions/PCR Cleanup/Seq prep days Sequencing ? Total For 7 sets of 48 plates: 16+ days 114 days
Library Construction: Time Line for 4608 colonies (48 s Time Mutagenesis/growth on Qbot plate 3 days Qbot picking/growth in 96 well culture plate 2 days Biomek 1 day 10 ARB 1/ARB 2 Reactions/PCR Cleanup/Seq prep days Sequencing ? Total For 7 sets of 48 plates: 16+ days 114 days
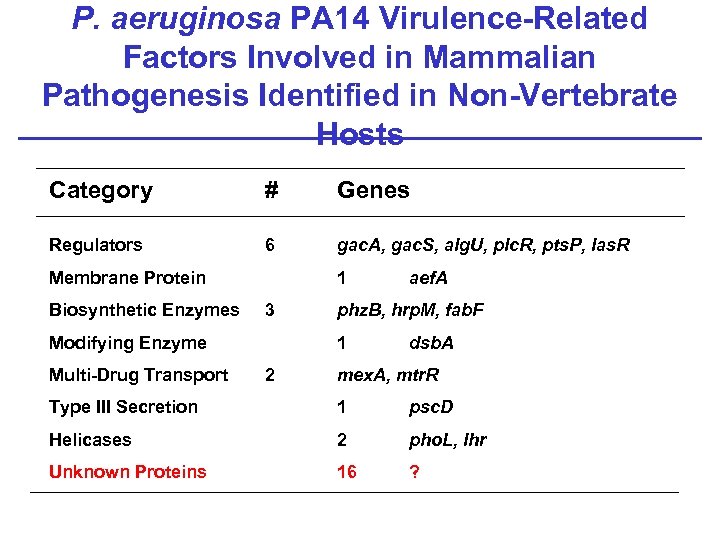 P. aeruginosa PA 14 Virulence-Related Factors Involved in Mammalian Pathogenesis Identified in Non-Vertebrate Hosts Category # Genes Regulators 6 gac. A, gac. S, alg. U, plc. R, pts. P, las. R Membrane Protein Biosynthetic Enzymes 1 3 Modifying Enzyme Multi-Drug Transport phz. B, hrp. M, fab. F 1 2 aef. A dsb. A mex. A, mtr. R Type III Secretion 1 psc. D Helicases 2 pho. L, lhr Unknown Proteins 16 ?
P. aeruginosa PA 14 Virulence-Related Factors Involved in Mammalian Pathogenesis Identified in Non-Vertebrate Hosts Category # Genes Regulators 6 gac. A, gac. S, alg. U, plc. R, pts. P, las. R Membrane Protein Biosynthetic Enzymes 1 3 Modifying Enzyme Multi-Drug Transport phz. B, hrp. M, fab. F 1 2 aef. A dsb. A mex. A, mtr. R Type III Secretion 1 psc. D Helicases 2 pho. L, lhr Unknown Proteins 16 ?
 Section IV Development of a Linux My. SQL Uni-Gene Library Relational Database
Section IV Development of a Linux My. SQL Uni-Gene Library Relational Database
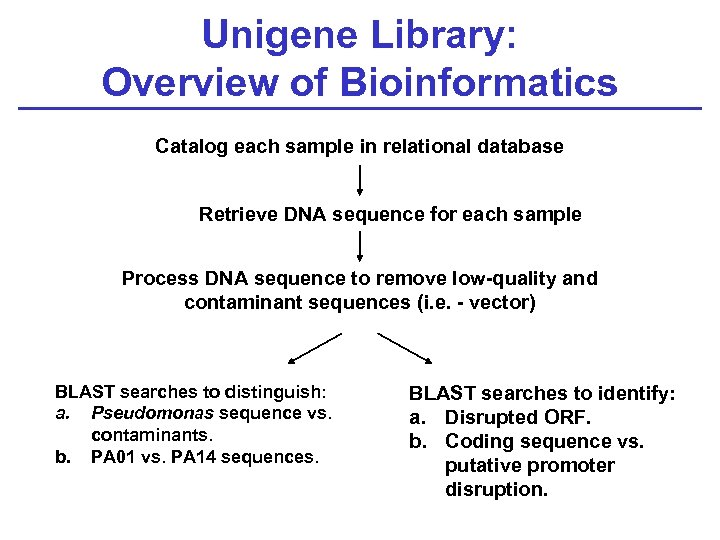 Unigene Library: Overview of Bioinformatics Catalog each sample in relational database Retrieve DNA sequence for each sample Process DNA sequence to remove low-quality and contaminant sequences (i. e. - vector) BLAST searches to distinguish: a. Pseudomonas sequence vs. contaminants. b. PA 01 vs. PA 14 sequences. BLAST searches to identify: a. Disrupted ORF. b. Coding sequence vs. putative promoter disruption.
Unigene Library: Overview of Bioinformatics Catalog each sample in relational database Retrieve DNA sequence for each sample Process DNA sequence to remove low-quality and contaminant sequences (i. e. - vector) BLAST searches to distinguish: a. Pseudomonas sequence vs. contaminants. b. PA 01 vs. PA 14 sequences. BLAST searches to identify: a. Disrupted ORF. b. Coding sequence vs. putative promoter disruption.
 What will the My. SQL Database Do? 1) Store/catalog all of the data. 2) Process DNA sequences and perform BLAST searches. 3) Display the results and allow for user queries.
What will the My. SQL Database Do? 1) Store/catalog all of the data. 2) Process DNA sequences and perform BLAST searches. 3) Display the results and allow for user queries.
 How will the Database Store the Data? • The data will be stored in a relational database. • Individual tables can be thought of as separate Excel spreadsheets with rows and columns. • The tables are connected to each other via specified relationships.
How will the Database Store the Data? • The data will be stored in a relational database. • Individual tables can be thought of as separate Excel spreadsheets with rows and columns. • The tables are connected to each other via specified relationships.
 How will the Database Store the Data? • Tables will be populated (i. e. - individual cells in the table will be filled with entries) as plates, samples, and/or data are generated. • Data entry into the Database will be “restricted” to parallel the creation of the physical library. – Order of different types of inputs is restricted. – Prevent duplicate entries.
How will the Database Store the Data? • Tables will be populated (i. e. - individual cells in the table will be filled with entries) as plates, samples, and/or data are generated. • Data entry into the Database will be “restricted” to parallel the creation of the physical library. – Order of different types of inputs is restricted. – Prevent duplicate entries.
 How will the Database Store the Data? • The Database will store organizational information: – – – Date created. Created by. Storage locations. Bacterial strain. Mutagen/Transposon used. • The Database will store experimental data: – – DNA sequences obtained by PCR. Location of insertion with respect to PAO 1 genome. Identity of PAO 1 ORF disrupted. Phenotypic data?
How will the Database Store the Data? • The Database will store organizational information: – – – Date created. Created by. Storage locations. Bacterial strain. Mutagen/Transposon used. • The Database will store experimental data: – – DNA sequences obtained by PCR. Location of insertion with respect to PAO 1 genome. Identity of PAO 1 ORF disrupted. Phenotypic data?
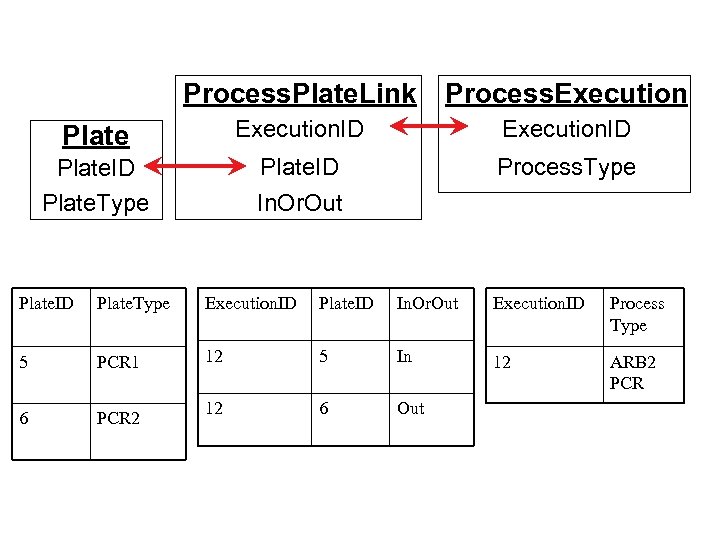 Process. Plate. Link Process. Execution Plate Execution. ID Plate. Type Plate. ID In. Or. Out Process. Type Plate. ID Plate. Type Execution. ID Plate. ID In. Or. Out Execution. ID Process Type 5 PCR 1 12 5 In 12 ARB 2 PCR 6 PCR 2 12 6 Out
Process. Plate. Link Process. Execution Plate Execution. ID Plate. Type Plate. ID In. Or. Out Process. Type Plate. ID Plate. Type Execution. ID Plate. ID In. Or. Out Execution. ID Process Type 5 PCR 1 12 5 In 12 ARB 2 PCR 6 PCR 2 12 6 Out
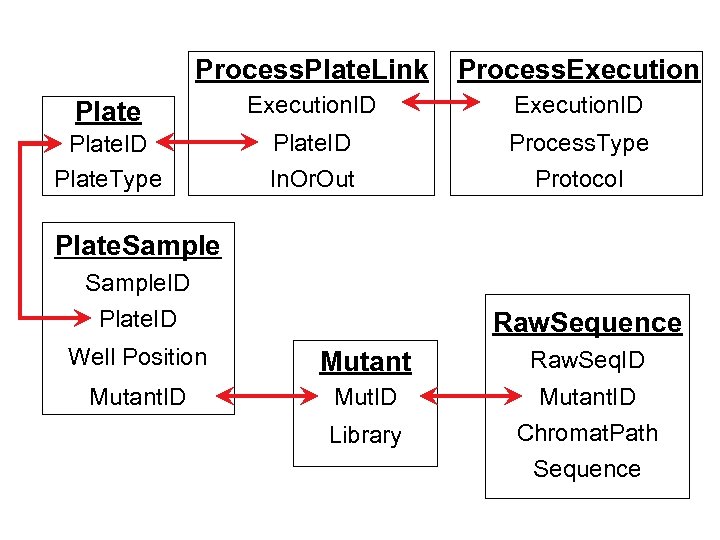 Process. Plate. Link Process. Execution Plate Execution. ID Plate. Type Plate. ID Process. Type In. Or. Out Protocol Plate. Sample. ID Plate. ID Raw. Sequence Well Position Mutant Raw. Seq. ID Mutant. ID Chromat. Path Library Sequence
Process. Plate. Link Process. Execution Plate Execution. ID Plate. Type Plate. ID Process. Type In. Or. Out Protocol Plate. Sample. ID Plate. ID Raw. Sequence Well Position Mutant Raw. Seq. ID Mutant. ID Chromat. Path Library Sequence
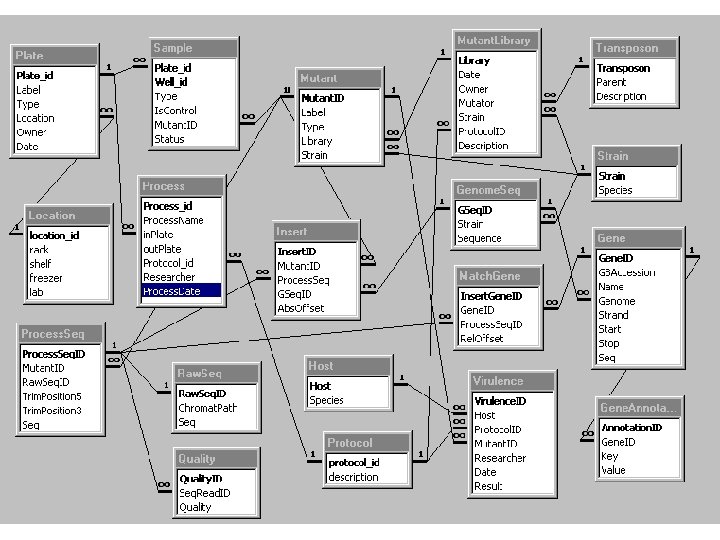
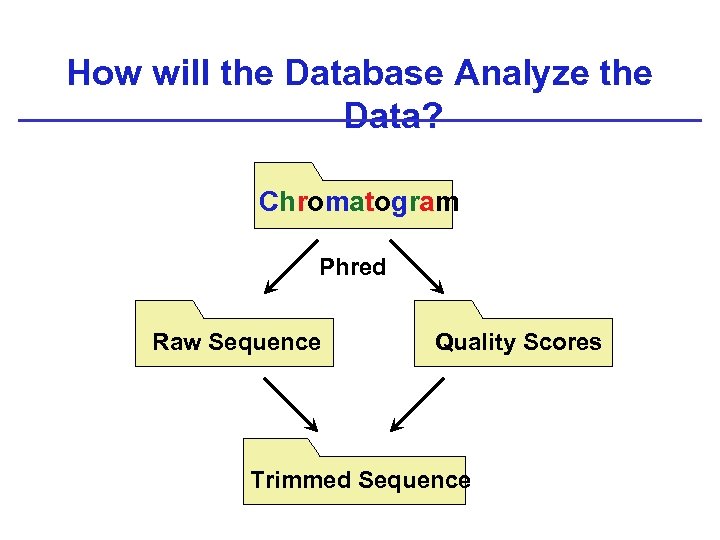 How will the Database Analyze the Data? Chromatogram Phred Raw Sequence Quality Scores Trimmed Sequence
How will the Database Analyze the Data? Chromatogram Phred Raw Sequence Quality Scores Trimmed Sequence
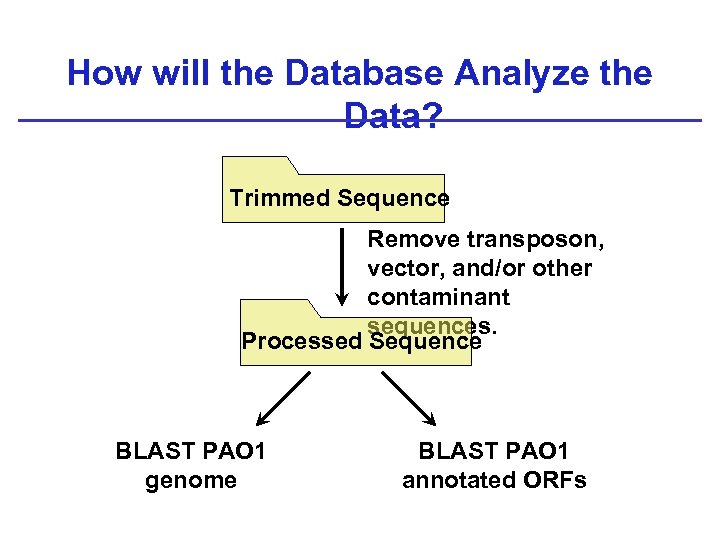 How will the Database Analyze the Data? Trimmed Sequence Remove transposon, vector, and/or other contaminant sequences. Processed Sequence BLAST PAO 1 genome BLAST PAO 1 annotated ORFs
How will the Database Analyze the Data? Trimmed Sequence Remove transposon, vector, and/or other contaminant sequences. Processed Sequence BLAST PAO 1 genome BLAST PAO 1 annotated ORFs
 How will the Database Analyze the Data? • Other BLAST searches that can be performed in the future: – Internal BLAST against the contents of the database to identify siblings vs. adjacent independent insertions. – BLAST against other public databases to determine gene identity of ORFs not found in PAO 1.
How will the Database Analyze the Data? • Other BLAST searches that can be performed in the future: – Internal BLAST against the contents of the database to identify siblings vs. adjacent independent insertions. – BLAST against other public databases to determine gene identity of ORFs not found in PAO 1.
 How will we Retrieve/View the Contents of the Database? • Current status: – A web-accessible table viewer can allow us to examine the contents of each table in the database. – To organize and search the contents, the html file can be opened in Excel and then sorted. • Future goals: – A web-accessible browser with multiple query and view options.
How will we Retrieve/View the Contents of the Database? • Current status: – A web-accessible table viewer can allow us to examine the contents of each table in the database. – To organize and search the contents, the html file can be opened in Excel and then sorted. • Future goals: – A web-accessible browser with multiple query and view options.
 How will we Retrieve/View the Contents of the Database? Types of queries: – – – Insertions in a given gene. Insertions upstream of a given gene. Insertions near a given physical location. Insertions in PAO 1 non-coding sequences. Insertions in sequences NOT found in PAO 1. – – Insertions in genes of a particular pathway/family. Insertions in PAO 1 ORFs of known function. Insertions in putative PAO 1 ORFs of unknown function. Multiple queries.
How will we Retrieve/View the Contents of the Database? Types of queries: – – – Insertions in a given gene. Insertions upstream of a given gene. Insertions near a given physical location. Insertions in PAO 1 non-coding sequences. Insertions in sequences NOT found in PAO 1. – – Insertions in genes of a particular pathway/family. Insertions in PAO 1 ORFs of known function. Insertions in putative PAO 1 ORFs of unknown function. Multiple queries.
 How will we Retrieve/View the Contents of the Database? Options for Viewing Database Contents: – Table view in alphabetical order. – Table view in linear order. – Graphical view with ORF orientation and transposon orientation (zoom in/out/, click on ORF or transposon, etc).
How will we Retrieve/View the Contents of the Database? Options for Viewing Database Contents: – Table view in alphabetical order. – Table view in linear order. – Graphical view with ORF orientation and transposon orientation (zoom in/out/, click on ORF or transposon, etc).
 Future Steps • Select members for unigene (nonredundant) library. • Physically pick members for unigene library. • Store, duplicate and disseminate unigene library. • Incorporate non-PAO 1 sequences into unigene set. • “Completing” the unigene set (targeted
Future Steps • Select members for unigene (nonredundant) library. • Physically pick members for unigene library. • Store, duplicate and disseminate unigene library. • Incorporate non-PAO 1 sequences into unigene set. • “Completing” the unigene set (targeted
 Trouble Shooting • Test cases - designed to test either the entire system (i. e. - start to finish) or a particular module. • Can be designed to test “ideal” inputs or “incorrect” inputs. • Example: – Input 96 (unknown) chromatogram files that contain a few known sequences at defined well positions. – Determine if the expected BLAST hits are associated with the appropriate well position. – This tests: • Ability to process the chromatogram files. • Ability to correctly perform BLAST searches. • Ability to correctly map the resulting BLAST search onto the correct well-position.
Trouble Shooting • Test cases - designed to test either the entire system (i. e. - start to finish) or a particular module. • Can be designed to test “ideal” inputs or “incorrect” inputs. • Example: – Input 96 (unknown) chromatogram files that contain a few known sequences at defined well positions. – Determine if the expected BLAST hits are associated with the appropriate well position. – This tests: • Ability to process the chromatogram files. • Ability to correctly perform BLAST searches. • Ability to correctly map the resulting BLAST search onto the correct well-position.
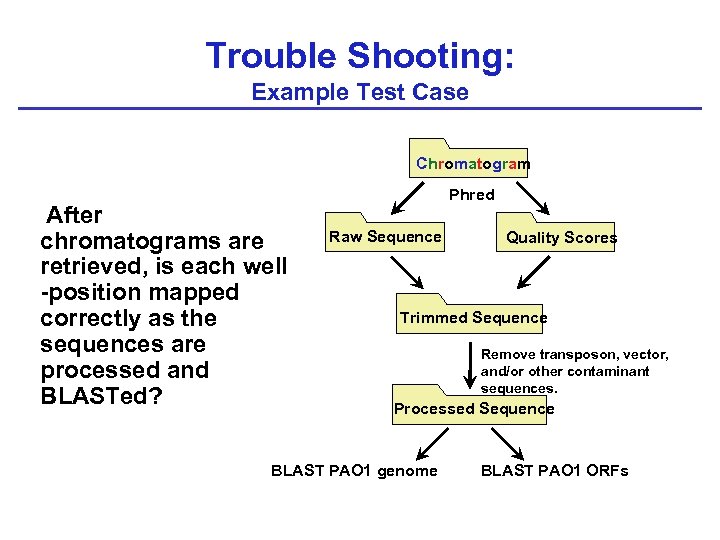 Trouble Shooting: Example Test Case Chromatogram After chromatograms are retrieved, is each well -position mapped correctly as the sequences are processed and BLASTed? Phred Raw Sequence Quality Scores Trimmed Sequence Remove transposon, vector, and/or other contaminant sequences. Processed Sequence BLAST PAO 1 genome BLAST PAO 1 ORFs
Trouble Shooting: Example Test Case Chromatogram After chromatograms are retrieved, is each well -position mapped correctly as the sequences are processed and BLASTed? Phred Raw Sequence Quality Scores Trimmed Sequence Remove transposon, vector, and/or other contaminant sequences. Processed Sequence BLAST PAO 1 genome BLAST PAO 1 ORFs
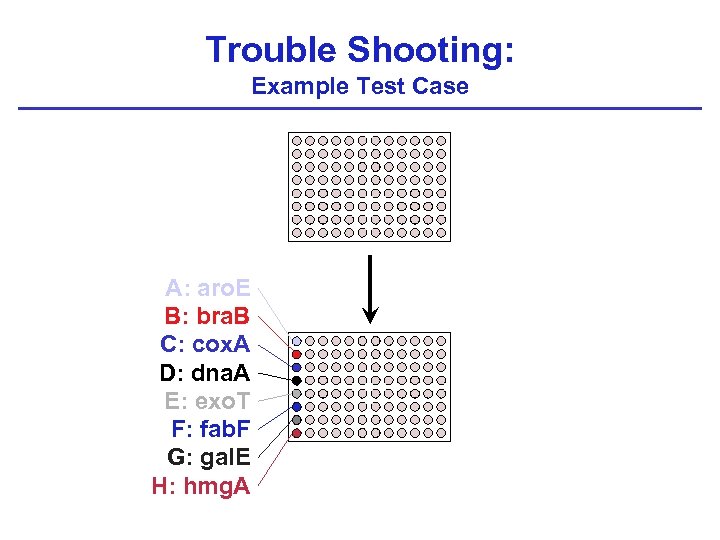 Trouble Shooting: Example Test Case A: aro. E B: bra. B C: cox. A D: dna. A E: exo. T F: fab. F G: gal. E H: hmg. A
Trouble Shooting: Example Test Case A: aro. E B: bra. B C: cox. A D: dna. A E: exo. T F: fab. F G: gal. E H: hmg. A
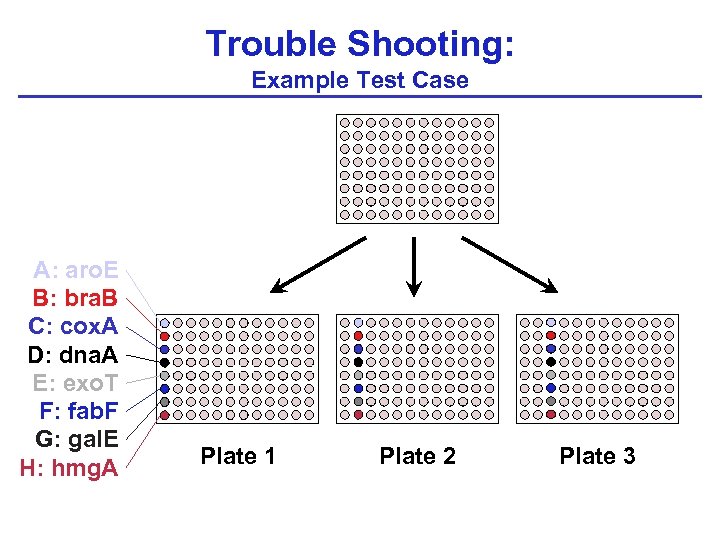 Trouble Shooting: Example Test Case A: aro. E B: bra. B C: cox. A D: dna. A E: exo. T F: fab. F G: gal. E H: hmg. A Plate 1 Plate 2 Plate 3
Trouble Shooting: Example Test Case A: aro. E B: bra. B C: cox. A D: dna. A E: exo. T F: fab. F G: gal. E H: hmg. A Plate 1 Plate 2 Plate 3
 Example Test Case: Expected BLAST Results 1 A 1 : aro. E 1 A 2 : --1 A 3 : --1 A 4 : --1 A 5 : --1 A 6 : --1 A 7 : --1 A 8 : --1 A 9 : --1 A 10 : --1 A 11 : --1 A 12 : --- 1 B 1 : bra. B 1 B 2 : --1 B 3 : --1 B 4 : --1 B 5 : --1 B 6 : --1 B 7 : --1 B 8 : --1 B 9 : --1 B 10 : --1 B 11 : --1 B 12 : --- 1 C 1 : cox. A 1 C 2 : --1 C 3 : --1 C 4 : --1 C 5 : --1 C 6 : --1 C 7 : --1 C 8 : --1 C 9 : --1 C 10 : --1 C 11 : --1 C 12 : --- 1 D 1 : dna. A 1 D 2 : --1 D 3 : --1 D 4 : --1 D 5 : --1 D 6 : --1 D 7 : --1 D 8 : --1 D 9 : --1 D 10 : --1 D 11 : --1 D 12 : ---
Example Test Case: Expected BLAST Results 1 A 1 : aro. E 1 A 2 : --1 A 3 : --1 A 4 : --1 A 5 : --1 A 6 : --1 A 7 : --1 A 8 : --1 A 9 : --1 A 10 : --1 A 11 : --1 A 12 : --- 1 B 1 : bra. B 1 B 2 : --1 B 3 : --1 B 4 : --1 B 5 : --1 B 6 : --1 B 7 : --1 B 8 : --1 B 9 : --1 B 10 : --1 B 11 : --1 B 12 : --- 1 C 1 : cox. A 1 C 2 : --1 C 3 : --1 C 4 : --1 C 5 : --1 C 6 : --1 C 7 : --1 C 8 : --1 C 9 : --1 C 10 : --1 C 11 : --1 C 12 : --- 1 D 1 : dna. A 1 D 2 : --1 D 3 : --1 D 4 : --1 D 5 : --1 D 6 : --1 D 7 : --1 D 8 : --1 D 9 : --1 D 10 : --1 D 11 : --1 D 12 : ---
 “System Requirements” • Programming language / database platform. – Microsoft Access vs. My. SQL. • Backups and database restore. • Storage issues. – Each plate generates ~30 MB of chromatograms (I. e. - 3 X 96 chromatograms on a zip disk). – Each chromatogram spawns several types of data: a raw sequence, a quality score for each nucleotide of the raw score (~2. 67 MB for an average 96 -well plate), a processed sequence, and blast results. • Documentation of database development
“System Requirements” • Programming language / database platform. – Microsoft Access vs. My. SQL. • Backups and database restore. • Storage issues. – Each plate generates ~30 MB of chromatograms (I. e. - 3 X 96 chromatograms on a zip disk). – Each chromatogram spawns several types of data: a raw sequence, a quality score for each nucleotide of the raw score (~2. 67 MB for an average 96 -well plate), a processed sequence, and blast results. • Documentation of database development
 Database Summary • Data storage is mostly complete. Needs some testing. • Sequence analysis is currently being tested. – Once it’s operational, sequence analysis will have to be updated to include more complex scenarios (i. e. - sequences not found in PAO 1). • Data retrieval/viewing is currently undeveloped. • Non-redundant (unigene) library is
Database Summary • Data storage is mostly complete. Needs some testing. • Sequence analysis is currently being tested. – Once it’s operational, sequence analysis will have to be updated to include more complex scenarios (i. e. - sequences not found in PAO 1). • Data retrieval/viewing is currently undeveloped. • Non-redundant (unigene) library is
 Section V CF Mouse Oropharynx Colonization Model
Section V CF Mouse Oropharynx Colonization Model
 Utility of transgenic CF mice for identifying novel P. aeruginosa virulence factors • To date, no apparent phenotype relevant to acquisition and establishment of chronic P. aeruginosa infection has been found in a variety of transgenic CF mouse lines • CF mice given acute P. aeruginosa infections manifest increased inflammation and pathology but do not get chronic infections
Utility of transgenic CF mice for identifying novel P. aeruginosa virulence factors • To date, no apparent phenotype relevant to acquisition and establishment of chronic P. aeruginosa infection has been found in a variety of transgenic CF mouse lines • CF mice given acute P. aeruginosa infections manifest increased inflammation and pathology but do not get chronic infections
 Our approach: try to recapitulate method of natural acquisition of P. aeruginosa by CF patients Aspects of chronic oropharyngeal colonization in mice • Maintain mice on water with antibiotic to prevent P. aeruginosa growth in water-0. 1 mg gentamicin/ml • Treat mice for 5 -7 days with 250 ug levofloxacin/ml in drinking water – Eliminates oropharyngeal colonization by a mucoid Enterobacter that grows on Pseudomonas isolation media and interferes with P. aeruginosa colonization – Remove 48 hrs prior to introduction of P. aeruginosa
Our approach: try to recapitulate method of natural acquisition of P. aeruginosa by CF patients Aspects of chronic oropharyngeal colonization in mice • Maintain mice on water with antibiotic to prevent P. aeruginosa growth in water-0. 1 mg gentamicin/ml • Treat mice for 5 -7 days with 250 ug levofloxacin/ml in drinking water – Eliminates oropharyngeal colonization by a mucoid Enterobacter that grows on Pseudomonas isolation media and interferes with P. aeruginosa colonization – Remove 48 hrs prior to introduction of P. aeruginosa
 Aspects of chronic oropharyngeal colonization in mice • Colonize mice by placing 107 CFU P. aeruginosa/ml drinking water for 5 days • Remove contaminated water, culture mouse throats, give sterile water for 1 week followed by water containing 0. 1 mg gentamicin/ml to keep bacteria from growing in it • Monitor mice by throat culture every 1 -2 weeks.
Aspects of chronic oropharyngeal colonization in mice • Colonize mice by placing 107 CFU P. aeruginosa/ml drinking water for 5 days • Remove contaminated water, culture mouse throats, give sterile water for 1 week followed by water containing 0. 1 mg gentamicin/ml to keep bacteria from growing in it • Monitor mice by throat culture every 1 -2 weeks.
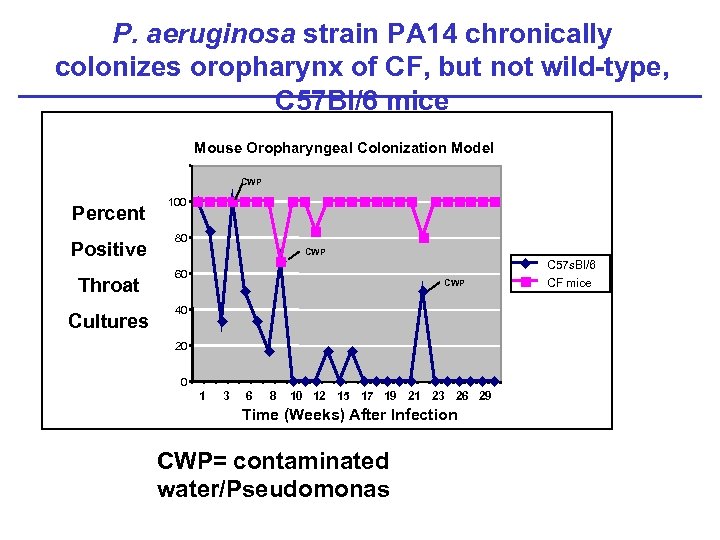 P. aeruginosa strain PA 14 chronically colonizes oropharynx of CF, but not wild-type, C 57 Bl/6 mice Mouse Oropharyngeal Colonization Model CWP Percent Positive Throat Cultures 100 80 CWP 60 CWP 40 20 0 1 3 6 8 10 12 15 17 19 21 23 26 29 Time (Weeks) After Infection CWP= contaminated water/Pseudomonas C 57 s. Bl/6 CF mice
P. aeruginosa strain PA 14 chronically colonizes oropharynx of CF, but not wild-type, C 57 Bl/6 mice Mouse Oropharyngeal Colonization Model CWP Percent Positive Throat Cultures 100 80 CWP 60 CWP 40 20 0 1 3 6 8 10 12 15 17 19 21 23 26 29 Time (Weeks) After Infection CWP= contaminated water/Pseudomonas C 57 s. Bl/6 CF mice
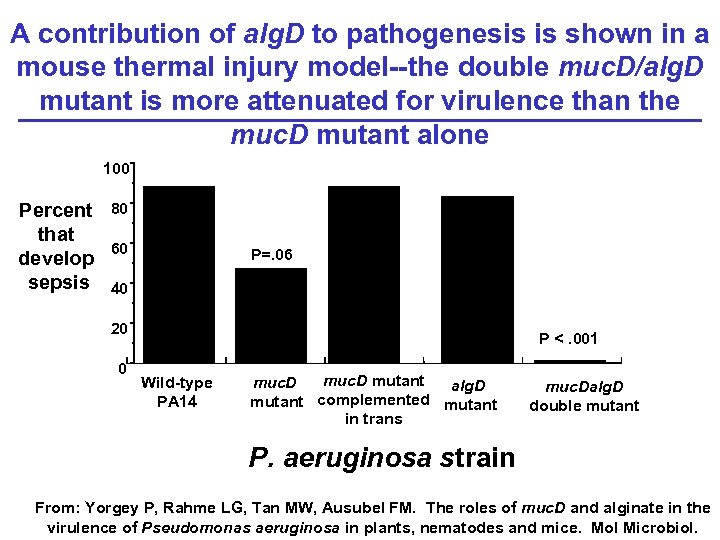 A contribution of alg. D to pathogenesis is shown in a mouse thermal injury model--the double muc. D/alg. D mutant is more attenuated for virulence than the muc. D mutant alone 100 Percent that develop sepsis 80 60 P=. 06 40 20 0 P <. 001 Wild-type PA 14 muc. D mutant muc. D alg. D mutant complemented mutant in trans muc. Dalg. D double mutant P. aeruginosa strain From: Yorgey P, Rahme LG, Tan MW, Ausubel FM. The roles of muc. D and alginate in the virulence of Pseudomonas aeruginosa in plants, nematodes and mice. Mol Microbiol.
A contribution of alg. D to pathogenesis is shown in a mouse thermal injury model--the double muc. D/alg. D mutant is more attenuated for virulence than the muc. D mutant alone 100 Percent that develop sepsis 80 60 P=. 06 40 20 0 P <. 001 Wild-type PA 14 muc. D mutant muc. D alg. D mutant complemented mutant in trans muc. Dalg. D double mutant P. aeruginosa strain From: Yorgey P, Rahme LG, Tan MW, Ausubel FM. The roles of muc. D and alginate in the virulence of Pseudomonas aeruginosa in plants, nematodes and mice. Mol Microbiol.
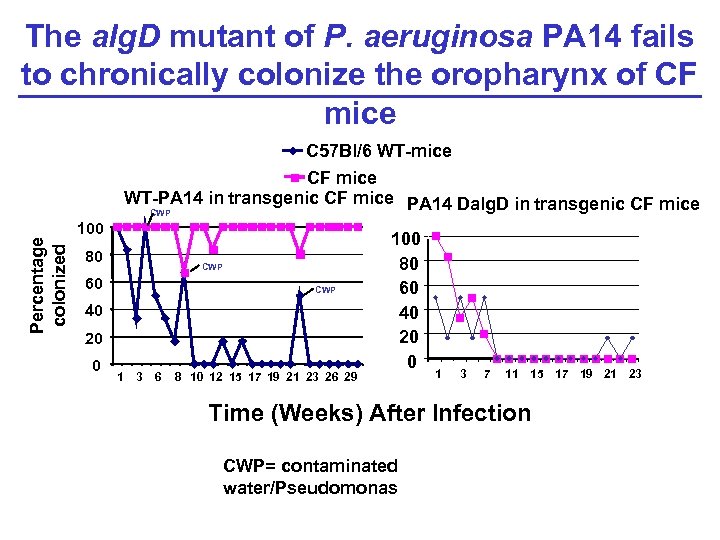 The alg. D mutant of P. aeruginosa PA 14 fails to chronically colonize the oropharynx of CF mice C 57 Bl/6 WT-mice Percentage colonized CF mice WT-PA 14 in transgenic CF mice PA 14 Dalg. D in transgenic CF mice CWP 100 80 CWP 60 CWP 40 20 0 1 3 6 8 10 12 15 17 19 21 23 26 29 100 80 60 40 20 0 1 3 7 11 15 17 19 21 23 Time (Weeks) After Infection CWP= contaminated water/Pseudomonas
The alg. D mutant of P. aeruginosa PA 14 fails to chronically colonize the oropharynx of CF mice C 57 Bl/6 WT-mice Percentage colonized CF mice WT-PA 14 in transgenic CF mice PA 14 Dalg. D in transgenic CF mice CWP 100 80 CWP 60 CWP 40 20 0 1 3 6 8 10 12 15 17 19 21 23 26 29 100 80 60 40 20 0 1 3 7 11 15 17 19 21 23 Time (Weeks) After Infection CWP= contaminated water/Pseudomonas
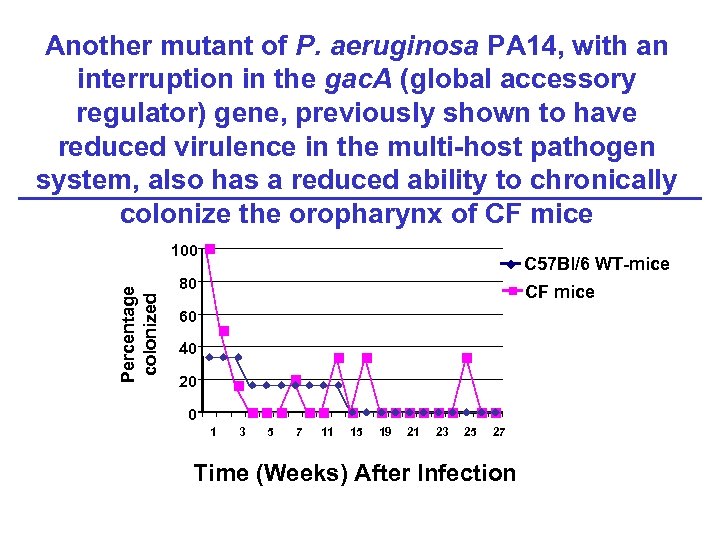 Another mutant of P. aeruginosa PA 14, with an interruption in the gac. A (global accessory regulator) gene, previously shown to have reduced virulence in the multi-host pathogen system, also has a reduced ability to chronically colonize the oropharynx of CF mice Percentage colonized 100 C 57 Bl/6 WT-mice 80 CF mice 60 40 20 0 1 3 5 7 11 15 19 21 23 25 27 Time (Weeks) After Infection
Another mutant of P. aeruginosa PA 14, with an interruption in the gac. A (global accessory regulator) gene, previously shown to have reduced virulence in the multi-host pathogen system, also has a reduced ability to chronically colonize the oropharynx of CF mice Percentage colonized 100 C 57 Bl/6 WT-mice 80 CF mice 60 40 20 0 1 3 5 7 11 15 19 21 23 25 27 Time (Weeks) After Infection
 Summary of CF Mouse Model A model of chronic P. aeruginosa oropharyngeal colonization in CF mice has been developed and tested for applicability for confirming the role of P. aeruginosa multi-host virulence factors.
Summary of CF Mouse Model A model of chronic P. aeruginosa oropharyngeal colonization in CF mice has been developed and tested for applicability for confirming the role of P. aeruginosa multi-host virulence factors.


