Лекция_1.pptx
- Количество слайдов: 34
 Методы изучения регуляторных районов генов Лекция I Меркулова Татьяна Ивановна Институт цитологии и генетики СО РАН
Методы изучения регуляторных районов генов Лекция I Меркулова Татьяна Ивановна Институт цитологии и генетики СО РАН
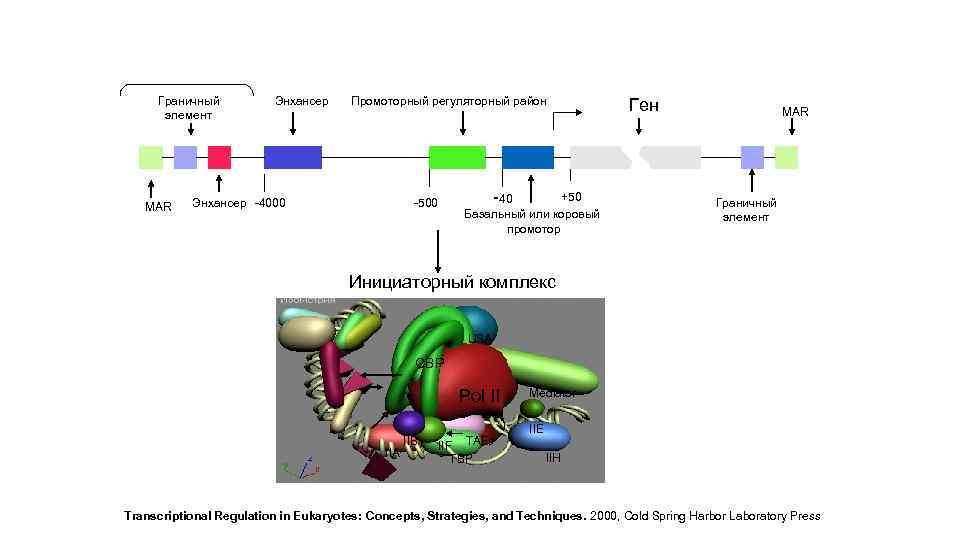 Граничный элемент MAR Энхансер Промоторный регуляторный район Энхансер -4000 -500 -40 +50 Базальный или коровый промотор Ген MAR Граничный элемент Инициаторный комплекс USA CBP Pol II IIA IIB IIF TAFs TBP Mediator IIE IIH Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
Граничный элемент MAR Энхансер Промоторный регуляторный район Энхансер -4000 -500 -40 +50 Базальный или коровый промотор Ген MAR Граничный элемент Инициаторный комплекс USA CBP Pol II IIA IIB IIF TAFs TBP Mediator IIE IIH Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
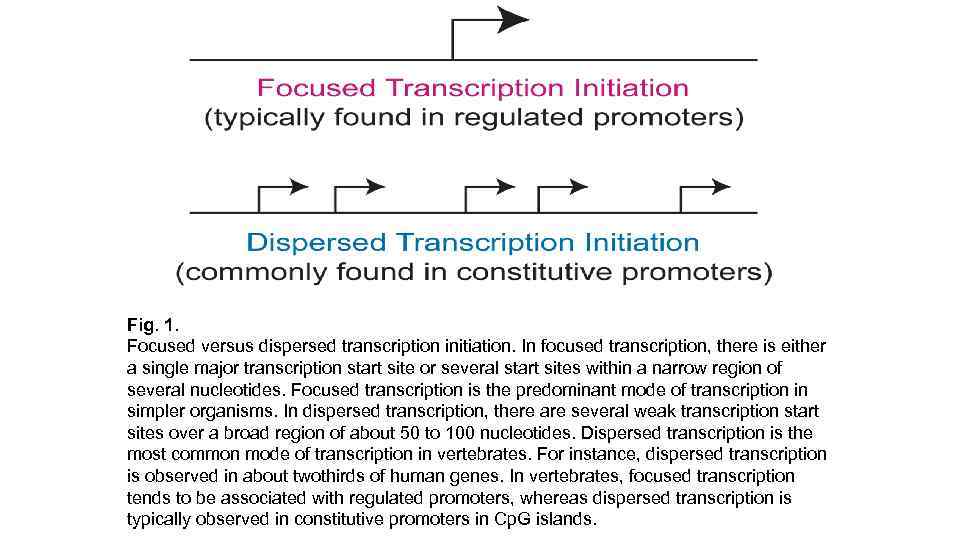 Fig. 1. Focused versus dispersed transcription initiation. In focused transcription, there is either a single major transcription start site or several start sites within a narrow region of several nucleotides. Focused transcription is the predominant mode of transcription in simpler organisms. In dispersed transcription, there are several weak transcription start sites over a broad region of about 50 to 100 nucleotides. Dispersed transcription is the most common mode of transcription in vertebrates. For instance, dispersed transcription is observed in about twothirds of human genes. In vertebrates, focused transcription tends to be associated with regulated promoters, whereas dispersed transcription is typically observed in constitutive promoters in Cp. G islands.
Fig. 1. Focused versus dispersed transcription initiation. In focused transcription, there is either a single major transcription start site or several start sites within a narrow region of several nucleotides. Focused transcription is the predominant mode of transcription in simpler organisms. In dispersed transcription, there are several weak transcription start sites over a broad region of about 50 to 100 nucleotides. Dispersed transcription is the most common mode of transcription in vertebrates. For instance, dispersed transcription is observed in about twothirds of human genes. In vertebrates, focused transcription tends to be associated with regulated promoters, whereas dispersed transcription is typically observed in constitutive promoters in Cp. G islands.
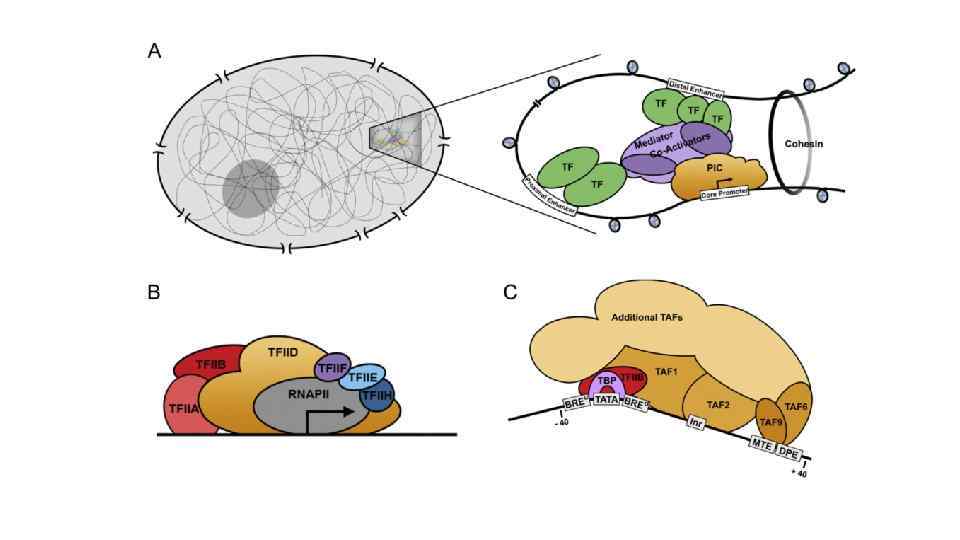
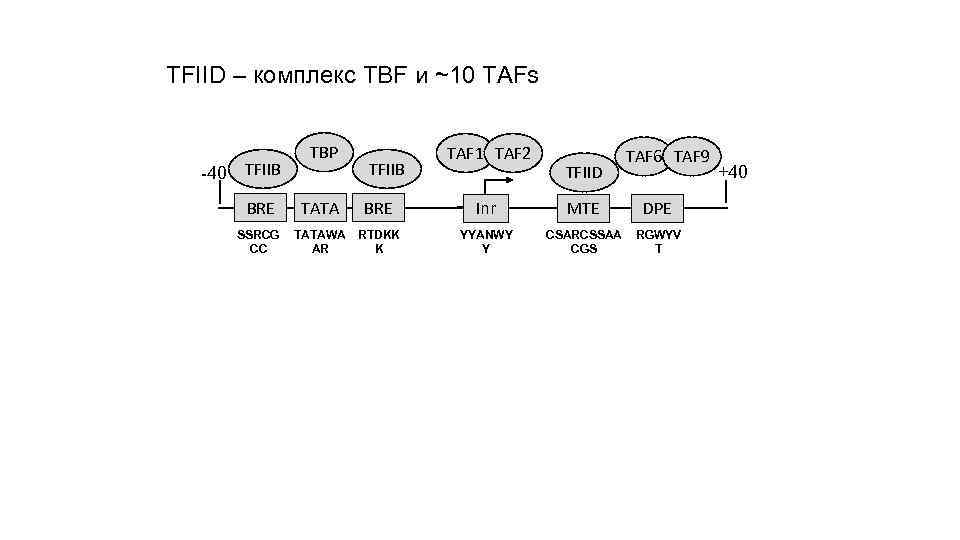 TFIID – комплекс TBF и ~10 TAFs -40 TFIIB BRE SSRCG CC TBP TATA TFIIB BRE TATAWA RTDKK АR K TAF 1 TAF 2 TFIID TAF 6 TAF 9 Inr MTE DPE YYANWY Y CSARCSSAA CGS RGWYV T +40
TFIID – комплекс TBF и ~10 TAFs -40 TFIIB BRE SSRCG CC TBP TATA TFIIB BRE TATAWA RTDKK АR K TAF 1 TAF 2 TFIID TAF 6 TAF 9 Inr MTE DPE YYANWY Y CSARCSSAA CGS RGWYV T +40
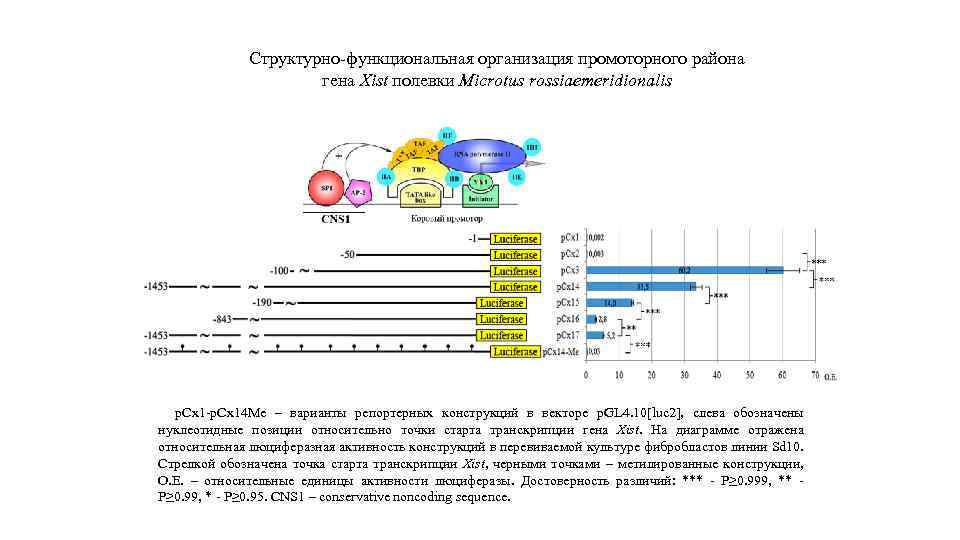
 Структурно-функциональная организация промоторного района гена Xist полевки Microtus rossiaemeridionalis р. Сх1 -р. Сх14 Ме – варианты репортерных конструкций в векторе p. GL 4. 10[luc 2], слева обозначены нуклеотидные позиции относительно точки старта транскрипции гена Xist. На диаграмме отражена относительная люциферазная активность конструкций в перевиваемой культуре фибробластов линии Sd 10. Стрелкой обозначена точка старта транскрипции Xist, черными точками – метилированные конструкции, О. Е. – относительные единицы активности люциферазы. Достоверность различий: *** - P≥ 0. 999, ** P≥ 0. 99, * - P≥ 0. 95. CNS 1 – conservative noncoding sequence.
Структурно-функциональная организация промоторного района гена Xist полевки Microtus rossiaemeridionalis р. Сх1 -р. Сх14 Ме – варианты репортерных конструкций в векторе p. GL 4. 10[luc 2], слева обозначены нуклеотидные позиции относительно точки старта транскрипции гена Xist. На диаграмме отражена относительная люциферазная активность конструкций в перевиваемой культуре фибробластов линии Sd 10. Стрелкой обозначена точка старта транскрипции Xist, черными точками – метилированные конструкции, О. Е. – относительные единицы активности люциферазы. Достоверность различий: *** - P≥ 0. 999, ** P≥ 0. 99, * - P≥ 0. 95. CNS 1 – conservative noncoding sequence.
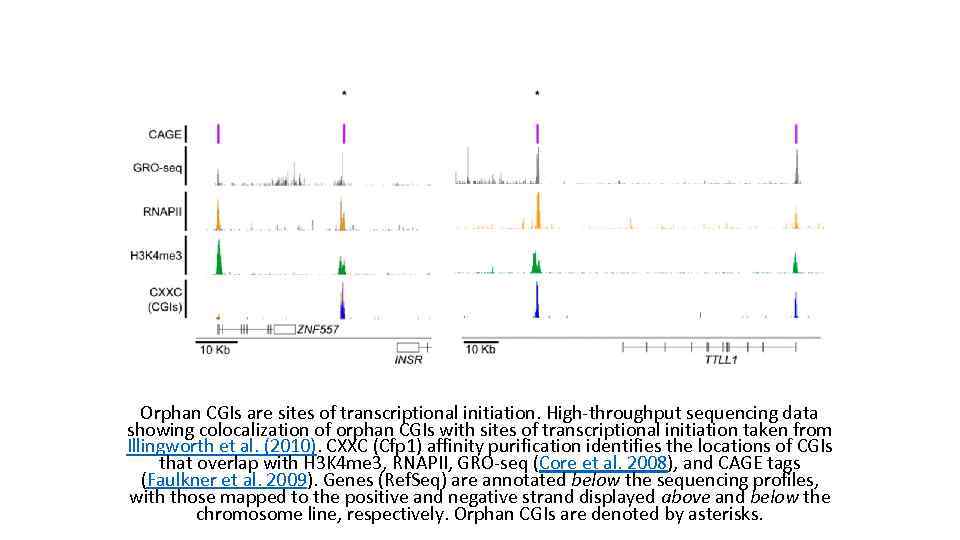 Orphan CGIs are sites of transcriptional initiation. High-throughput sequencing data showing colocalization of orphan CGIs with sites of transcriptional initiation taken from Illingworth et al. (2010). CXXC (Cfp 1) affinity purification identifies the locations of CGIs that overlap with H 3 K 4 me 3, RNAPII, GRO-seq (Core et al. 2008), and CAGE tags (Faulkner et al. 2009). Genes (Ref. Seq) are annotated below the sequencing profiles, with those mapped to the positive and negative strand displayed above and below the chromosome line, respectively. Orphan CGIs are denoted by asterisks.
Orphan CGIs are sites of transcriptional initiation. High-throughput sequencing data showing colocalization of orphan CGIs with sites of transcriptional initiation taken from Illingworth et al. (2010). CXXC (Cfp 1) affinity purification identifies the locations of CGIs that overlap with H 3 K 4 me 3, RNAPII, GRO-seq (Core et al. 2008), and CAGE tags (Faulkner et al. 2009). Genes (Ref. Seq) are annotated below the sequencing profiles, with those mapped to the positive and negative strand displayed above and below the chromosome line, respectively. Orphan CGIs are denoted by asterisks.
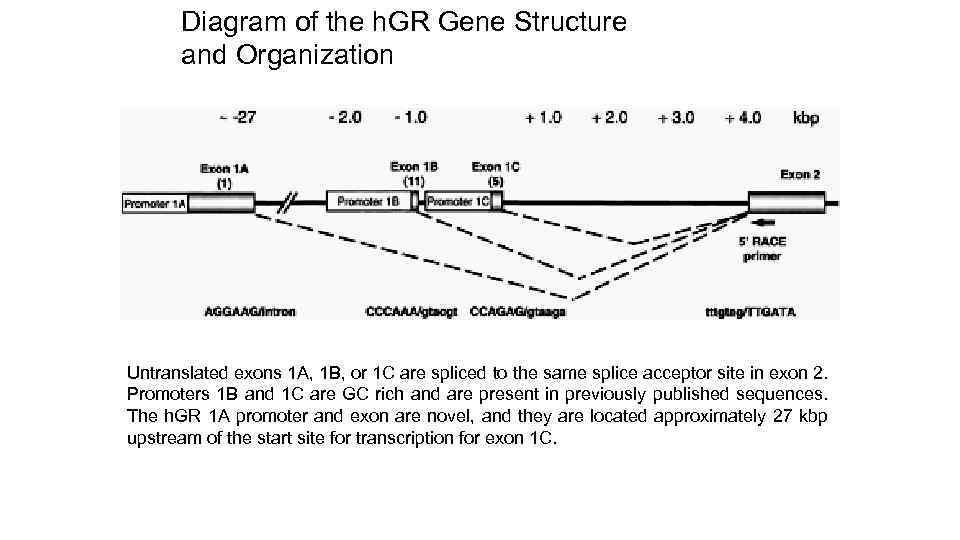 Diagram of the h. GR Gene Structure and Organization Untranslated exons 1 A, 1 B, or 1 C are spliced to the same splice acceptor site in exon 2. Promoters 1 B and 1 C are GC rich and are present in previously published sequences. The h. GR 1 A promoter and exon are novel, and they are located approximately 27 kbp upstream of the start site for transcription for exon 1 C.
Diagram of the h. GR Gene Structure and Organization Untranslated exons 1 A, 1 B, or 1 C are spliced to the same splice acceptor site in exon 2. Promoters 1 B and 1 C are GC rich and are present in previously published sequences. The h. GR 1 A promoter and exon are novel, and they are located approximately 27 kbp upstream of the start site for transcription for exon 1 C.
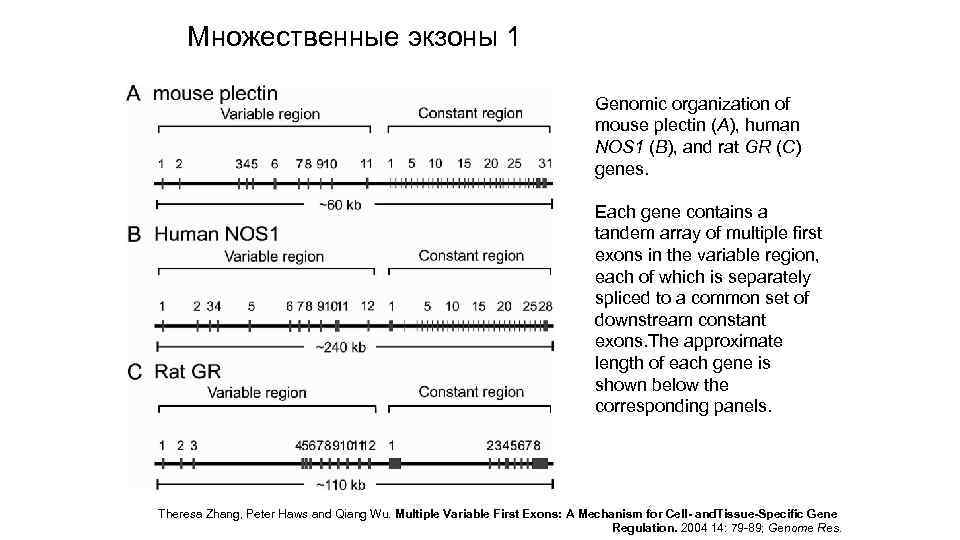 Множественные экзоны 1 Genomic organization of mouse plectin (A), human NOS 1 (B), and rat GR (C) genes. Each gene contains a tandem array of multiple first exons in the variable region, each of which is separately spliced to a common set of downstream constant exons. The approximate length of each gene is shown below the corresponding panels. Theresa Zhang, Peter Haws and Qiang Wu. Multiple Variable First Exons: A Mechanism for Cell- and. Tissue-Specific Gene Regulation. 2004 14: 79 -89; Genome Res.
Множественные экзоны 1 Genomic organization of mouse plectin (A), human NOS 1 (B), and rat GR (C) genes. Each gene contains a tandem array of multiple first exons in the variable region, each of which is separately spliced to a common set of downstream constant exons. The approximate length of each gene is shown below the corresponding panels. Theresa Zhang, Peter Haws and Qiang Wu. Multiple Variable First Exons: A Mechanism for Cell- and. Tissue-Specific Gene Regulation. 2004 14: 79 -89; Genome Res.
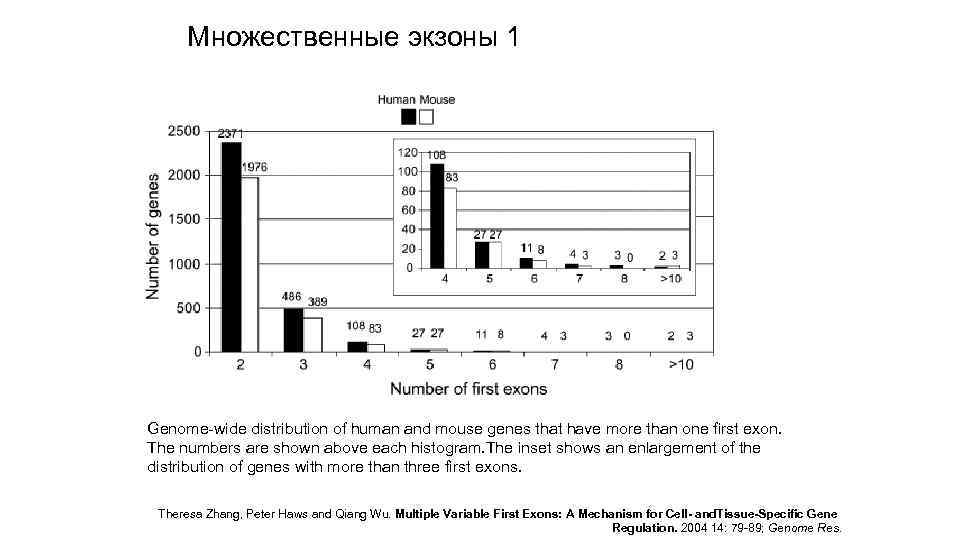 Множественные экзоны 1 Genome-wide distribution of human and mouse genes that have more than one first exon. The numbers are shown above each histogram. The inset shows an enlargement of the distribution of genes with more than three first exons. Theresa Zhang, Peter Haws and Qiang Wu. Multiple Variable First Exons: A Mechanism for Cell- and. Tissue-Specific Gene Regulation. 2004 14: 79 -89; Genome Res.
Множественные экзоны 1 Genome-wide distribution of human and mouse genes that have more than one first exon. The numbers are shown above each histogram. The inset shows an enlargement of the distribution of genes with more than three first exons. Theresa Zhang, Peter Haws and Qiang Wu. Multiple Variable First Exons: A Mechanism for Cell- and. Tissue-Specific Gene Regulation. 2004 14: 79 -89; Genome Res.
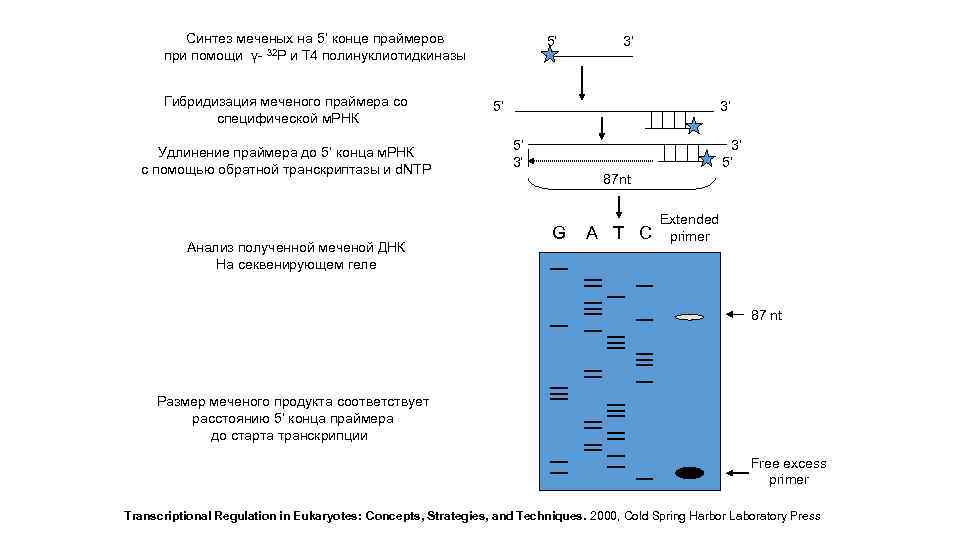 Синтез меченых на 5’ конце праймеров при помощи γ- 32 P и Т 4 полинуклиотидкиназы Гибридизация меченого праймера со специфической м. РНК Удлинение праймера до 5’ конца м. РНК c помощью обратной транскриптазы и d. NTP Анализ полученной меченой ДНК На секвенирующем геле 5’ 3’ 5’ 3’ 3’ 5’ 87 nt G A T C Extended primer 87 nt Размер меченого продукта соответствует расстоянию 5’ конца праймера до старта транскрипции Free excess primer Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
Синтез меченых на 5’ конце праймеров при помощи γ- 32 P и Т 4 полинуклиотидкиназы Гибридизация меченого праймера со специфической м. РНК Удлинение праймера до 5’ конца м. РНК c помощью обратной транскриптазы и d. NTP Анализ полученной меченой ДНК На секвенирующем геле 5’ 3’ 5’ 3’ 3’ 5’ 87 nt G A T C Extended primer 87 nt Размер меченого продукта соответствует расстоянию 5’ конца праймера до старта транскрипции Free excess primer Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
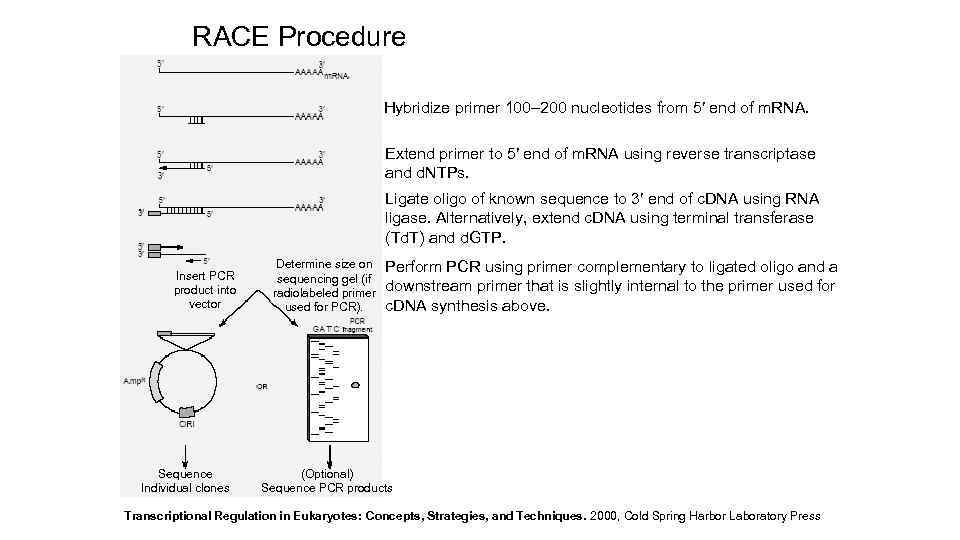 RACE Procedure Hybridize primer 100– 200 nucleotides from 5′ end of m. RNA. Extend primer to 5′ end of m. RNA using reverse transcriptase and d. NTPs. Ligate oligo of known sequence to 3′ end of c. DNA using RNA ligase. Alternatively, extend c. DNA using terminal transferase (Td. T) and d. GTP. Insert PCR product into vector Sequence Individual clones Determine size on sequencing gel (if radiolabeled primer used for PCR). Perform PCR using primer complementary to ligated oligo and a downstream primer that is slightly internal to the primer used for c. DNA synthesis above. (Optional) Sequence PCR products Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
RACE Procedure Hybridize primer 100– 200 nucleotides from 5′ end of m. RNA. Extend primer to 5′ end of m. RNA using reverse transcriptase and d. NTPs. Ligate oligo of known sequence to 3′ end of c. DNA using RNA ligase. Alternatively, extend c. DNA using terminal transferase (Td. T) and d. GTP. Insert PCR product into vector Sequence Individual clones Determine size on sequencing gel (if radiolabeled primer used for PCR). Perform PCR using primer complementary to ligated oligo and a downstream primer that is slightly internal to the primer used for c. DNA synthesis above. (Optional) Sequence PCR products Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
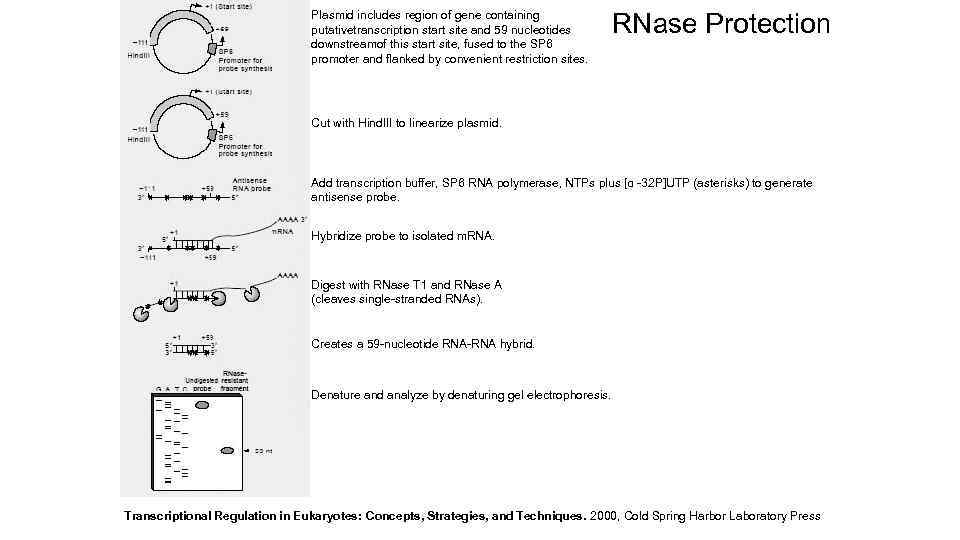 Plasmid includes region of gene containing putativetranscription start site and 59 nucleotides downstreamof this start site, fused to the SP 6 promoter and flanked by convenient restriction sites. RNase Protection Cut with Hind. III to linearize plasmid. Add transcription buffer, SP 6 RNA polymerase, NTPs plus [α -32 P]UTP (asterisks) to generate antisense probe. Hybridize probe to isolated m. RNA. Digest with RNase T 1 and RNase A (cleaves single-stranded RNAs). Creates a 59 -nucleotide RNA-RNA hybrid. Denature and analyze by denaturing gel electrophoresis. Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
Plasmid includes region of gene containing putativetranscription start site and 59 nucleotides downstreamof this start site, fused to the SP 6 promoter and flanked by convenient restriction sites. RNase Protection Cut with Hind. III to linearize plasmid. Add transcription buffer, SP 6 RNA polymerase, NTPs plus [α -32 P]UTP (asterisks) to generate antisense probe. Hybridize probe to isolated m. RNA. Digest with RNase T 1 and RNase A (cleaves single-stranded RNAs). Creates a 59 -nucleotide RNA-RNA hybrid. Denature and analyze by denaturing gel electrophoresis. Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
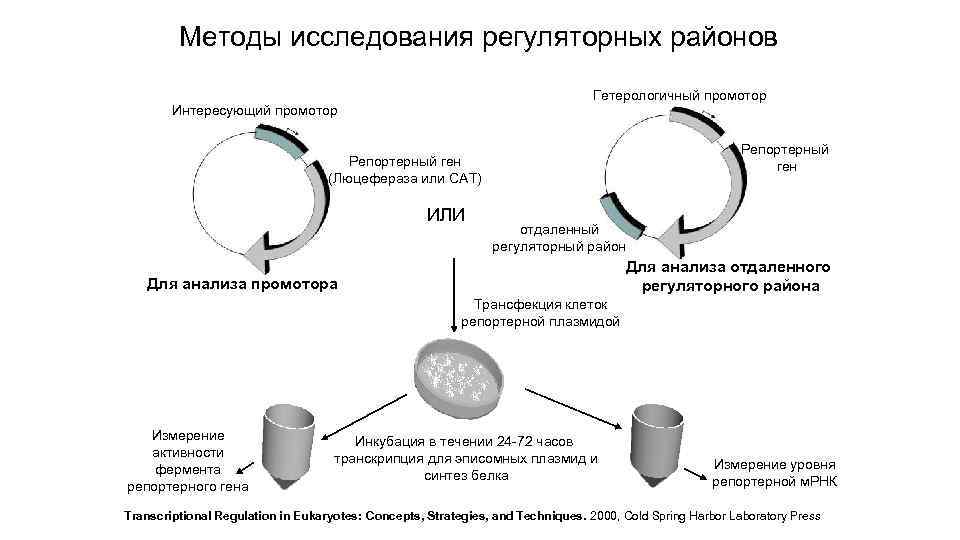 Методы исследования регуляторных районов Гетерологичный промотор Интересующий промотор Репортерный ген (Люцефераза или САТ) ИЛИ отдаленный регуляторный район Для анализа отдаленного регуляторного района Для анализа промотора Трансфекция клеток репортерной плазмидой Измерение активности фермента репортерного гена Инкубация в течении 24 -72 часов транскрипция для эписомных плазмид и синтез белка Измерение уровня репортерной м. РНК Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
Методы исследования регуляторных районов Гетерологичный промотор Интересующий промотор Репортерный ген (Люцефераза или САТ) ИЛИ отдаленный регуляторный район Для анализа отдаленного регуляторного района Для анализа промотора Трансфекция клеток репортерной плазмидой Измерение активности фермента репортерного гена Инкубация в течении 24 -72 часов транскрипция для эписомных плазмид и синтез белка Измерение уровня репортерной м. РНК Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
 Generation and initial analysis of more than 15, 000 full-length human and mouse c. DNA sequences Mammalian Gene Collection (MGC) Program Team*, 2000. Robert L. Strausberg, Elise A. Feingold, Lynette H. Grouse, Jeffery G. Derge, Richard D. Klausner, Francis S. Collins, Lukas Wagner, Carolyn M. Shenmen, Gregory D. Schuler, Stephen F. Altschul, Barry Zeeberg, Kenneth H. Buetow, Carl F. Schaefer, Narayan K. Bhat, Ralph F. Hopkins, Heather Jordan, Troy Moore, Steve I. Max, Jun Wang, Florence Hsieh, Luda Diatchenko, Kate Marusina, Andrew A. Farmer, Gerald M. Rubin, Ling Hong, Mark Stapleton, M. Bento Soares, Maria F. Bonaldo, Tom L. Casavant, Todd E. Scheetz, Michael J. Brownstein, Ted B. Usdin, Shiraki Toshiyuki, Piero Carninci, Christa Prange, Sam S. Raha, Naomi A. Loquellano, Garrick J. Peters, Rick D. Abramson, Sara J. Mullahy, Stephanie A. Bosak, Paul J. Mc. Ewan, Kevin J. Mc. Kernan, Joel A. Malek, Preethi H. Gunaratne, Stephen Richards, Kim C. Worley, Sarah Hale, Angela M. Garcia, Laura J. Gay, Stephen W. Hulyk, Debbie K. Villalon, Donna M. Muzny, Erica J. Sodergren, Xiuhua Lu, Richard A. Gibbs, Jessica Fahey, Erin Helton, Mark Ketteman, Anuradha Madan, Stephanie Rodrigues, Amy Sanchez, Michelle Whiting, Anup Madan, Alice C. Young, Yuriy Shevchenko, Gerard G. Bouffard, Robert W. Blakesley, Jeffrey W. Touchman, Eric D. Green, Mark C. Dickson, Alex C. Rodriguez, Jane Grimwood, Jeremy Schmutz, Richard M. Myers, Yaron S. N. Butterfield, Martin I. Krzywinski, Ursula Skalska, Duane E. Smailus, Angelique Schnerch, Jacqueline E. Schein, Steven J. M. Jones, and Marco A. Marra Contributed by Francis S. Collins, et al.
Generation and initial analysis of more than 15, 000 full-length human and mouse c. DNA sequences Mammalian Gene Collection (MGC) Program Team*, 2000. Robert L. Strausberg, Elise A. Feingold, Lynette H. Grouse, Jeffery G. Derge, Richard D. Klausner, Francis S. Collins, Lukas Wagner, Carolyn M. Shenmen, Gregory D. Schuler, Stephen F. Altschul, Barry Zeeberg, Kenneth H. Buetow, Carl F. Schaefer, Narayan K. Bhat, Ralph F. Hopkins, Heather Jordan, Troy Moore, Steve I. Max, Jun Wang, Florence Hsieh, Luda Diatchenko, Kate Marusina, Andrew A. Farmer, Gerald M. Rubin, Ling Hong, Mark Stapleton, M. Bento Soares, Maria F. Bonaldo, Tom L. Casavant, Todd E. Scheetz, Michael J. Brownstein, Ted B. Usdin, Shiraki Toshiyuki, Piero Carninci, Christa Prange, Sam S. Raha, Naomi A. Loquellano, Garrick J. Peters, Rick D. Abramson, Sara J. Mullahy, Stephanie A. Bosak, Paul J. Mc. Ewan, Kevin J. Mc. Kernan, Joel A. Malek, Preethi H. Gunaratne, Stephen Richards, Kim C. Worley, Sarah Hale, Angela M. Garcia, Laura J. Gay, Stephen W. Hulyk, Debbie K. Villalon, Donna M. Muzny, Erica J. Sodergren, Xiuhua Lu, Richard A. Gibbs, Jessica Fahey, Erin Helton, Mark Ketteman, Anuradha Madan, Stephanie Rodrigues, Amy Sanchez, Michelle Whiting, Anup Madan, Alice C. Young, Yuriy Shevchenko, Gerard G. Bouffard, Robert W. Blakesley, Jeffrey W. Touchman, Eric D. Green, Mark C. Dickson, Alex C. Rodriguez, Jane Grimwood, Jeremy Schmutz, Richard M. Myers, Yaron S. N. Butterfield, Martin I. Krzywinski, Ursula Skalska, Duane E. Smailus, Angelique Schnerch, Jacqueline E. Schein, Steven J. M. Jones, and Marco A. Marra Contributed by Francis S. Collins, et al.
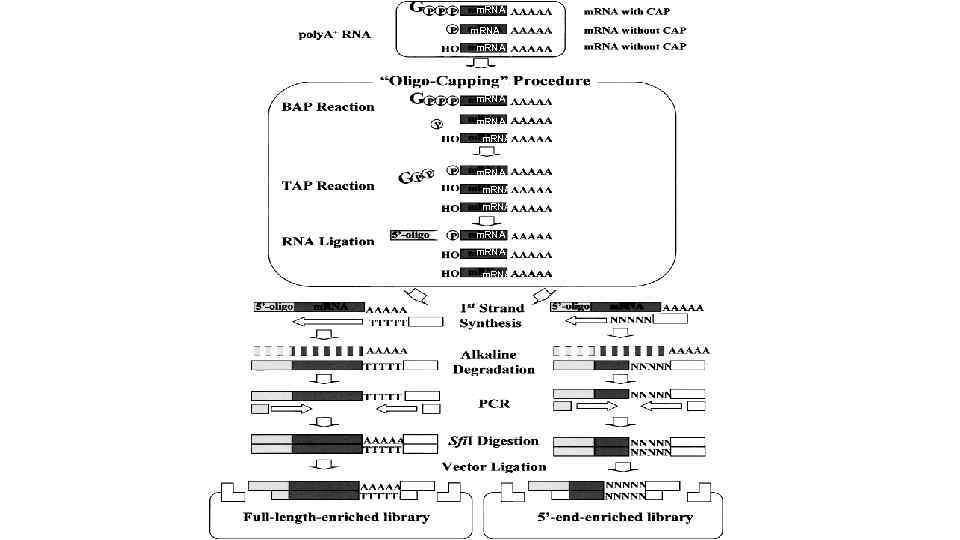 m. RNA m. RNA
m. RNA m. RNA
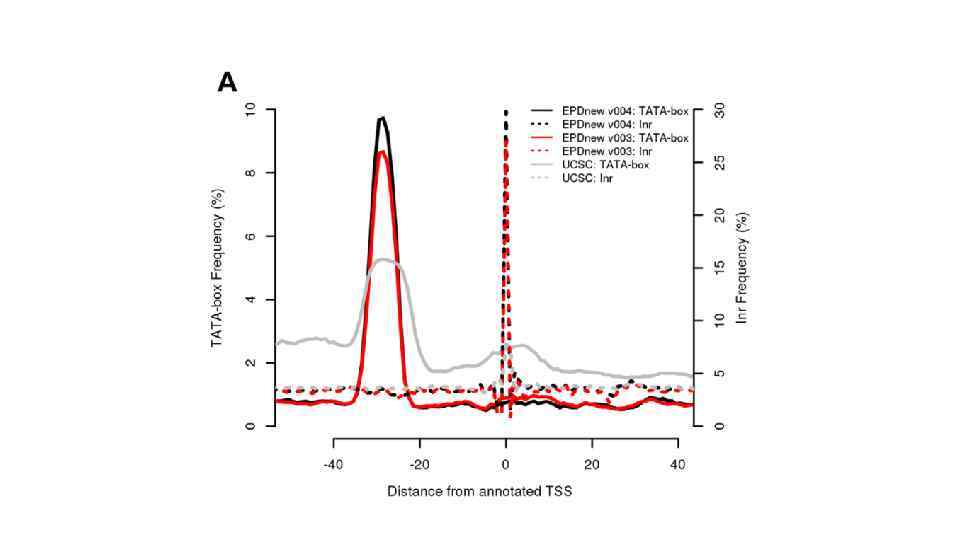
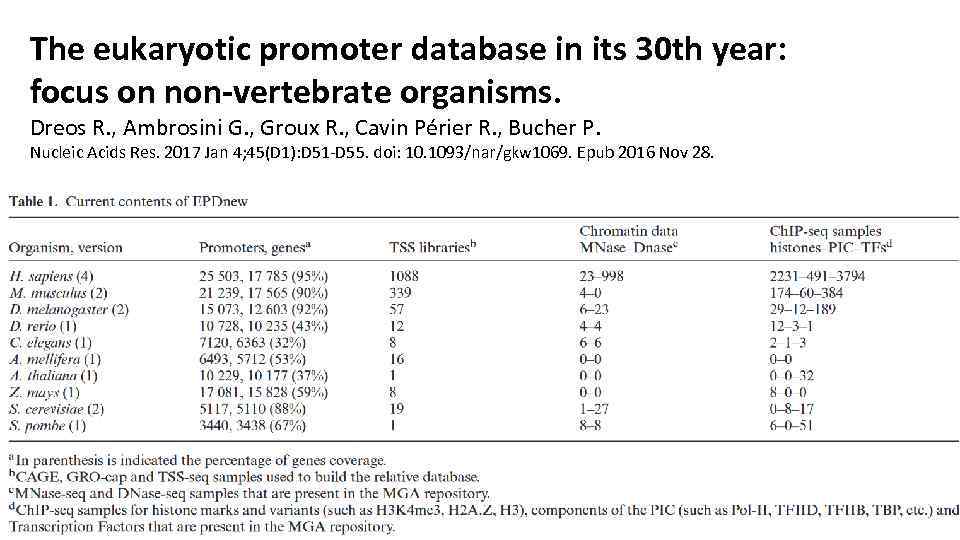 The eukaryotic promoter database in its 30 th year: focus on non-vertebrate organisms. Dreos R. , Ambrosini G. , Groux R. , Cavin Périer R. , Bucher P. Nucleic Acids Res. 2017 Jan 4; 45(D 1): D 51 -D 55. doi: 10. 1093/nar/gkw 1069. Epub 2016 Nov 28.
The eukaryotic promoter database in its 30 th year: focus on non-vertebrate organisms. Dreos R. , Ambrosini G. , Groux R. , Cavin Périer R. , Bucher P. Nucleic Acids Res. 2017 Jan 4; 45(D 1): D 51 -D 55. doi: 10. 1093/nar/gkw 1069. Epub 2016 Nov 28.
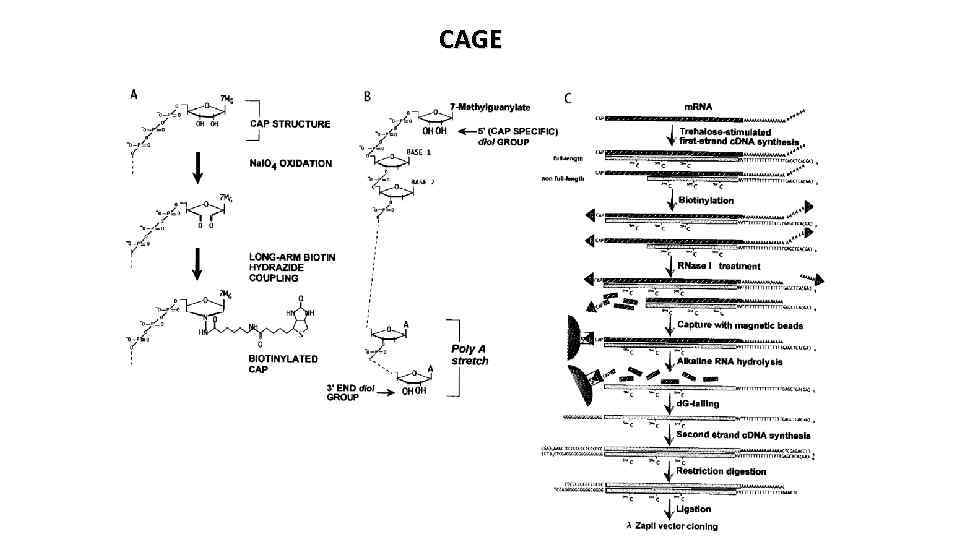 CAGE
CAGE
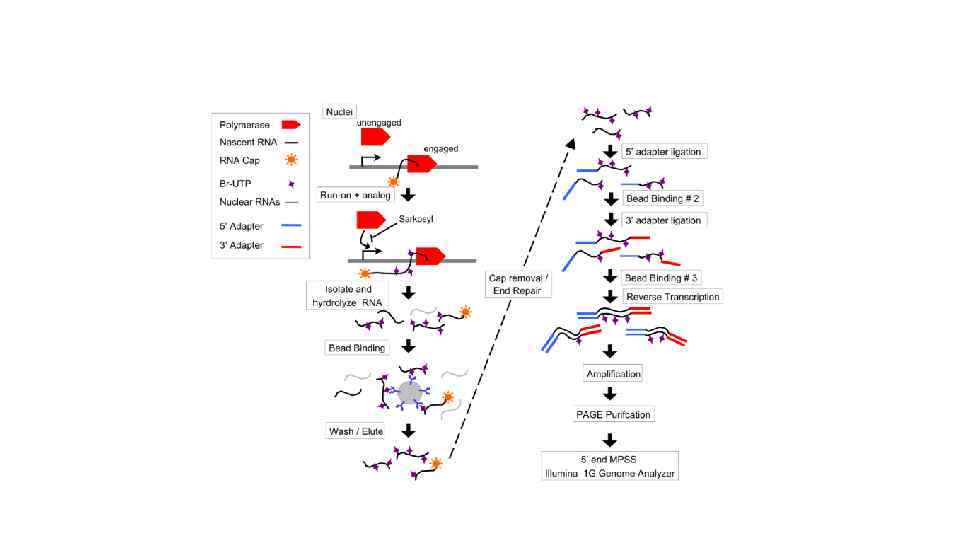
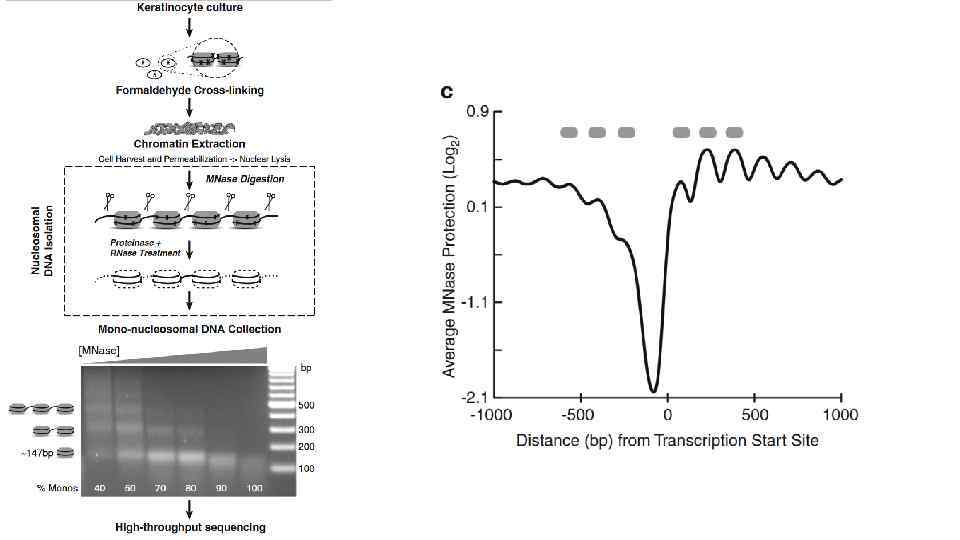
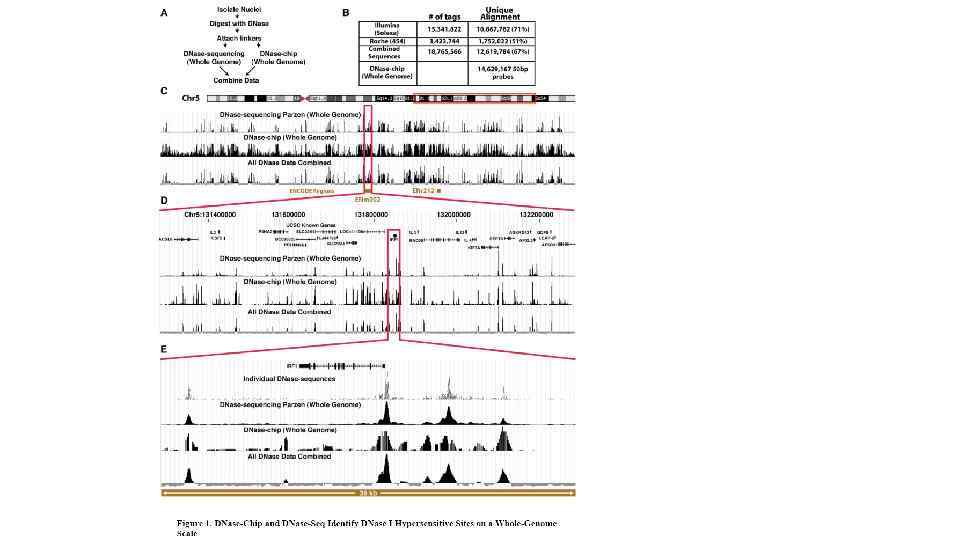
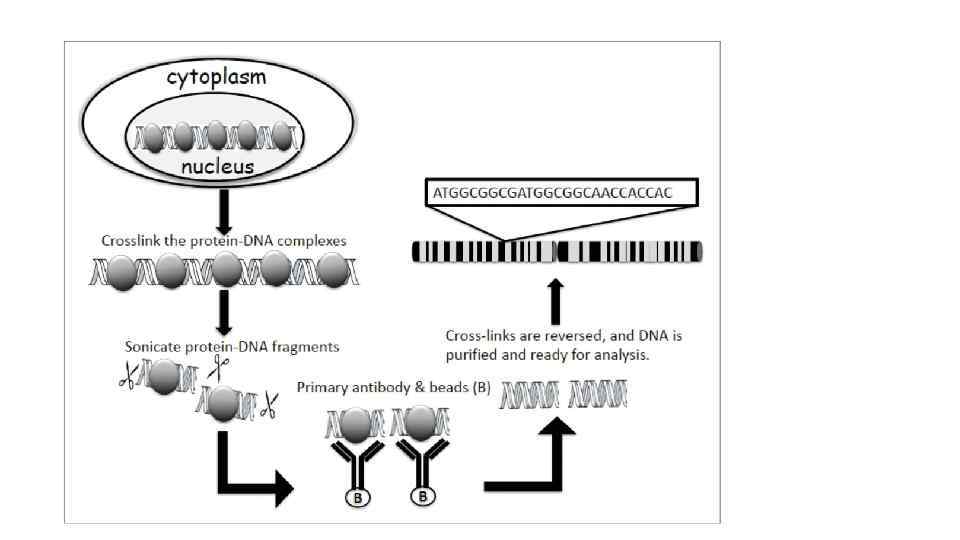
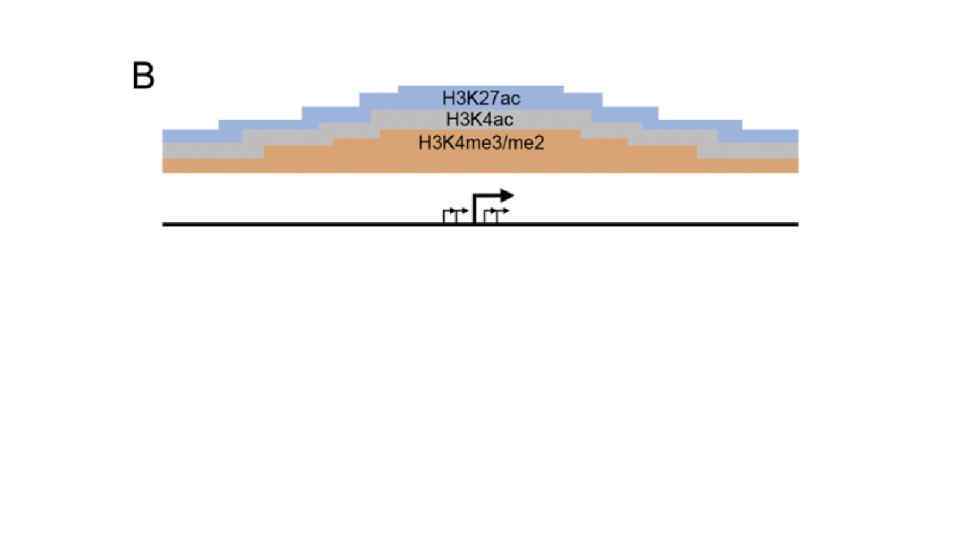
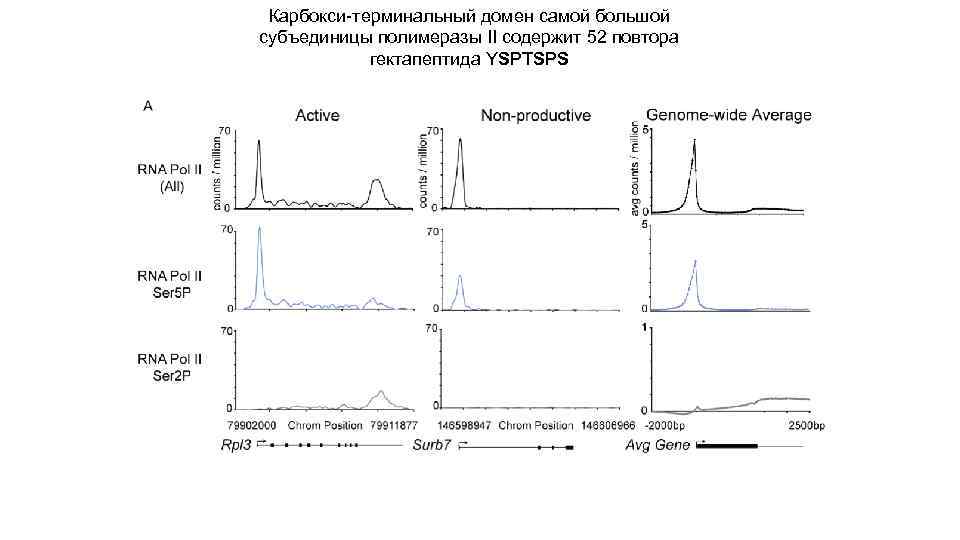 Карбокcи-терминальный домен самой большой субъединицы полимеразы II содержит 52 повтора гектапептида YSPTSPS
Карбокcи-терминальный домен самой большой субъединицы полимеразы II содержит 52 повтора гектапептида YSPTSPS
 The ENCODE Project Consortium ENCODE Project Scientific Management: Hartman, J. Rozowsky, O. Emanuelsson, T. Royce, S. Chung, M. National Human Genome Research Institute (E. A. Gerstein, Z. Lian, J. Lian, Y. Nakayama, S. Weissman, V. Stolc, W. Feingold, P. J. Good, M. S. Guyer, S. Kamholz, L. Liefer, Tongprasit, H. Sethi). K. Wetterstrand, F. S. Collins). Additional ENCODE Pilot Phase Participants: Initial ENCODE Pilot Phase Participants: British Columbia Cancer Agency Genome Sciences Centre (S. Jones, Affymetrix, Inc. (T. R. Gingeras, D. Kampa, E. A. Sekinger, M. Marra, H. Shin, J. Schein); Broad Institute (M. Clamp, K. Lindblad. J. Cheng, H. Hirsch, S. Ghosh, Z. Zhu, S. Patel, A. Piccolboni, Toh, J. Chang, D. B. Jaffe, M. Kamal, E. S. Lander, T. S. Mikkelsen, J. A. Yang, H. Tammana, S. Bekiranov, P. Kapranov, R. Harrison, Vinson, M. C. Zody); Children’s Hospital Oakland Research Institute G. Church, K. Struhl); Ludwig Institute for Cancer Research (B. (P. J. de Jong, K. Osoegawa, M. Nefedov, B. Zhu); National Human Ren, T. H. Kim, L. O. Barrera, C. Qu, S. Van Calcar, R. Luna, C. Genome Research Institute/Computational Genomics Unit (A. D. K. Glass, M. G. Rosenfeld); Municipal Institute of Medical Baxevanis, T. G. Wolfsberg); National Human Genome Research (R. Guigo, S. E. Antonarakis, E. Birney, M. Brent, L. Institute/Molecular Genetics Section (F. S. Collins, G. E. Crawford, J. Pachter, A. Reymond, E. T. Dermitzakis, C. Dewey, D. Keefe, F. Whittle, I. E. Holt, T. J. Vasicek, D. Zhou, S. Luo); NIH Intramural Denoeud, J. Lagarde, J. Ashurst, T. Hubbard, J. J. Wesselink, R. Sequencing Center/National Human Genome Research Institute (E. D. Castelo, E. Eyras); Stanford University (R. M. Myers, A. Sidow, Green, G. G. Bouffard, E. H. Margulies, M. E. Portnoy, N. F. Hansen, P. S. Batzoglou, N. D. Trinklein, S. J. Hartman, S. F. Aldred, E. J. Thomas, J. C. Mc. Dowell, B. Maskeri, A. C. Young, J. R. Idol, R. W. Anton, D. I. Schroeder, S. S. Marticke, L. Nguyen, J. Schmutz, Blakesley); National Library of Medicine (G. Schuler); Pennsylvania J. Grimwood, M. Dickson, G. M. Cooper, E. A. Stone, G. State University (W. Miller, R. Hardison, L. Elnitski, P. Shah); The Asimenos, M. Brudno); University of Virginia (A. Dutta, N. Institute for Genomic Research (S. L. Salzberg, M. Pertea, W. H. Karnani, C. M. Taylor, H. K. Kim, G. Robins); University of Majoros); University of California, Santa Cruz (D. Haussler, D. Washington (G. Stamatoyannopoulos, J. A. Thomas, K. R. Rosenbloom, H. Clawson, A. Siepel, W. J. Kent). Stamatoyannopoulos, M. Dorschner, P. Sabo, M. Hawrylycz, R. ENCODE Technology Development Phase Participants: Humbert, J. Wallace, M. Yu, P. A. Navas, M. Mc. Arthur, W. S. Boston University (Z. Weng, S. Jin, A. Halees, H. Burden, U. Karaoz, Y. Noble); Wellcome Trust Sanger Institute (I. Dunham, C. M. Fu, Y. Yu, C. Ding, C. R. Cantor); Massachusetts General Hospital (R. Koch, R. M. Andrews, G. K. Clelland, S. Wilcox, J. C. Fowler, K. D. E. Kingston, J. Dennis); Nimble. Gen Systems, Inc. (R. D. Green, M. A. James, P. Groth, O. M. Dovey, P. D. Ellis, V. L. Wraight, A. J. Singer, T. A. Richmond, J. E. Norton, P. J Farnham, M. J. Oberley, D. Mungall, P. Dhami, H. Fiegler, C. F. Langford, N. P. Carter, R. Inman); Nimble. Gen Systems, Inc. (M. R. Mc. Cormick, H. Kim, C. L. D. Vetrie); Yale University (M. Snyder, G. Euskirchen, A. E. Middle, M. C. Pirrung); University of California, Urban, U. Nagalakshmi, J. Rinn, G. Popescu, P. Bertone, S. et al! The ENCODE Project Consortium. 2004. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306: 636– 640.
The ENCODE Project Consortium ENCODE Project Scientific Management: Hartman, J. Rozowsky, O. Emanuelsson, T. Royce, S. Chung, M. National Human Genome Research Institute (E. A. Gerstein, Z. Lian, J. Lian, Y. Nakayama, S. Weissman, V. Stolc, W. Feingold, P. J. Good, M. S. Guyer, S. Kamholz, L. Liefer, Tongprasit, H. Sethi). K. Wetterstrand, F. S. Collins). Additional ENCODE Pilot Phase Participants: Initial ENCODE Pilot Phase Participants: British Columbia Cancer Agency Genome Sciences Centre (S. Jones, Affymetrix, Inc. (T. R. Gingeras, D. Kampa, E. A. Sekinger, M. Marra, H. Shin, J. Schein); Broad Institute (M. Clamp, K. Lindblad. J. Cheng, H. Hirsch, S. Ghosh, Z. Zhu, S. Patel, A. Piccolboni, Toh, J. Chang, D. B. Jaffe, M. Kamal, E. S. Lander, T. S. Mikkelsen, J. A. Yang, H. Tammana, S. Bekiranov, P. Kapranov, R. Harrison, Vinson, M. C. Zody); Children’s Hospital Oakland Research Institute G. Church, K. Struhl); Ludwig Institute for Cancer Research (B. (P. J. de Jong, K. Osoegawa, M. Nefedov, B. Zhu); National Human Ren, T. H. Kim, L. O. Barrera, C. Qu, S. Van Calcar, R. Luna, C. Genome Research Institute/Computational Genomics Unit (A. D. K. Glass, M. G. Rosenfeld); Municipal Institute of Medical Baxevanis, T. G. Wolfsberg); National Human Genome Research (R. Guigo, S. E. Antonarakis, E. Birney, M. Brent, L. Institute/Molecular Genetics Section (F. S. Collins, G. E. Crawford, J. Pachter, A. Reymond, E. T. Dermitzakis, C. Dewey, D. Keefe, F. Whittle, I. E. Holt, T. J. Vasicek, D. Zhou, S. Luo); NIH Intramural Denoeud, J. Lagarde, J. Ashurst, T. Hubbard, J. J. Wesselink, R. Sequencing Center/National Human Genome Research Institute (E. D. Castelo, E. Eyras); Stanford University (R. M. Myers, A. Sidow, Green, G. G. Bouffard, E. H. Margulies, M. E. Portnoy, N. F. Hansen, P. S. Batzoglou, N. D. Trinklein, S. J. Hartman, S. F. Aldred, E. J. Thomas, J. C. Mc. Dowell, B. Maskeri, A. C. Young, J. R. Idol, R. W. Anton, D. I. Schroeder, S. S. Marticke, L. Nguyen, J. Schmutz, Blakesley); National Library of Medicine (G. Schuler); Pennsylvania J. Grimwood, M. Dickson, G. M. Cooper, E. A. Stone, G. State University (W. Miller, R. Hardison, L. Elnitski, P. Shah); The Asimenos, M. Brudno); University of Virginia (A. Dutta, N. Institute for Genomic Research (S. L. Salzberg, M. Pertea, W. H. Karnani, C. M. Taylor, H. K. Kim, G. Robins); University of Majoros); University of California, Santa Cruz (D. Haussler, D. Washington (G. Stamatoyannopoulos, J. A. Thomas, K. R. Rosenbloom, H. Clawson, A. Siepel, W. J. Kent). Stamatoyannopoulos, M. Dorschner, P. Sabo, M. Hawrylycz, R. ENCODE Technology Development Phase Participants: Humbert, J. Wallace, M. Yu, P. A. Navas, M. Mc. Arthur, W. S. Boston University (Z. Weng, S. Jin, A. Halees, H. Burden, U. Karaoz, Y. Noble); Wellcome Trust Sanger Institute (I. Dunham, C. M. Fu, Y. Yu, C. Ding, C. R. Cantor); Massachusetts General Hospital (R. Koch, R. M. Andrews, G. K. Clelland, S. Wilcox, J. C. Fowler, K. D. E. Kingston, J. Dennis); Nimble. Gen Systems, Inc. (R. D. Green, M. A. James, P. Groth, O. M. Dovey, P. D. Ellis, V. L. Wraight, A. J. Singer, T. A. Richmond, J. E. Norton, P. J Farnham, M. J. Oberley, D. Mungall, P. Dhami, H. Fiegler, C. F. Langford, N. P. Carter, R. Inman); Nimble. Gen Systems, Inc. (M. R. Mc. Cormick, H. Kim, C. L. D. Vetrie); Yale University (M. Snyder, G. Euskirchen, A. E. Middle, M. C. Pirrung); University of California, Urban, U. Nagalakshmi, J. Rinn, G. Popescu, P. Bertone, S. et al! The ENCODE Project Consortium. 2004. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306: 636– 640.
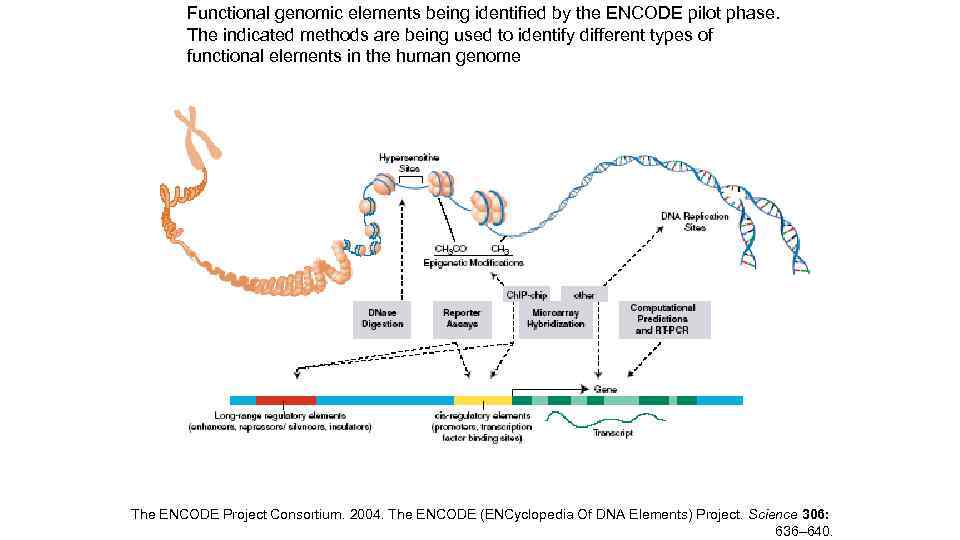 Functional genomic elements being identified by the ENCODE pilot phase. The indicated methods are being used to identify different types of functional elements in the human genome The ENCODE Project Consortium. 2004. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306: 636– 640.
Functional genomic elements being identified by the ENCODE pilot phase. The indicated methods are being used to identify different types of functional elements in the human genome The ENCODE Project Consortium. 2004. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306: 636– 640.
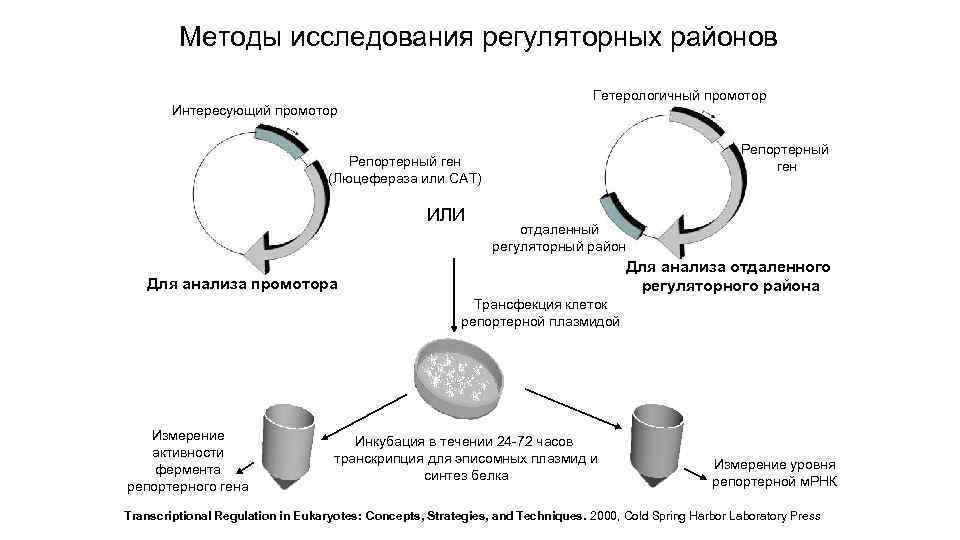 Методы исследования регуляторных районов Гетерологичный промотор Интересующий промотор Репортерный ген (Люцефераза или САТ) ИЛИ отдаленный регуляторный район Для анализа отдаленного регуляторного района Для анализа промотора Трансфекция клеток репортерной плазмидой Измерение активности фермента репортерного гена Инкубация в течении 24 -72 часов транскрипция для эписомных плазмид и синтез белка Измерение уровня репортерной м. РНК Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
Методы исследования регуляторных районов Гетерологичный промотор Интересующий промотор Репортерный ген (Люцефераза или САТ) ИЛИ отдаленный регуляторный район Для анализа отдаленного регуляторного района Для анализа промотора Трансфекция клеток репортерной плазмидой Измерение активности фермента репортерного гена Инкубация в течении 24 -72 часов транскрипция для эписомных плазмид и синтез белка Измерение уровня репортерной м. РНК Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. 2000, Cold Spring Harbor Laboratory Press
 Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome Sara J. Cooper, Nathan D. Trinklein, Elizabeth D. Anton, Loan Nguyen and Richard M. Myers Genome Res. , 2006 16: 1 -10; Clustergram of 642 putative promoter fragments. The clustergram illustrates the hierarchical clustering of promoter activity among 16 diverse cell lines. Each row indicates the promoter activity of a fragment in each of the cell lines, with red indicating the degree of activity and black indicating no activity. Promoter activity has been normalized and log transformed to reflect comparable values between cell lines. Area A represents a cluster of promoter fragments with strong, ubiquitous activity in all cell lines and area B represents a cluster of promoter fragments that exhibit variable function across the 16 cell types. Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome/ Sara J. Cooper, Nathan D. Trinklein, et al. Genome Res. , 2006 16: 1 -10
Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome Sara J. Cooper, Nathan D. Trinklein, Elizabeth D. Anton, Loan Nguyen and Richard M. Myers Genome Res. , 2006 16: 1 -10; Clustergram of 642 putative promoter fragments. The clustergram illustrates the hierarchical clustering of promoter activity among 16 diverse cell lines. Each row indicates the promoter activity of a fragment in each of the cell lines, with red indicating the degree of activity and black indicating no activity. Promoter activity has been normalized and log transformed to reflect comparable values between cell lines. Area A represents a cluster of promoter fragments with strong, ubiquitous activity in all cell lines and area B represents a cluster of promoter fragments that exhibit variable function across the 16 cell types. Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome/ Sara J. Cooper, Nathan D. Trinklein, et al. Genome Res. , 2006 16: 1 -10


