2cdc7d26fd7983eeb72ff0c10e114dc1.ppt
- Количество слайдов: 53
 Mechanistic Study and Identification of Essential residues of Family 3 b-Glucosidase
Mechanistic Study and Identification of Essential residues of Family 3 b-Glucosidase
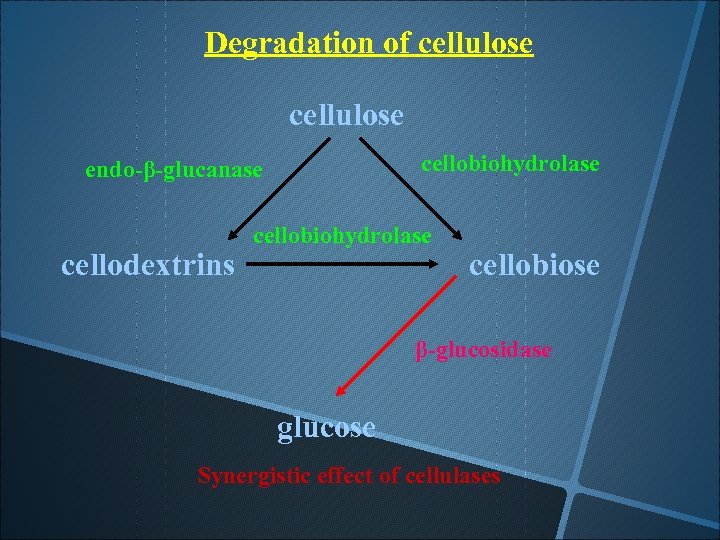 Degradation of cellulose cellobiohydrolase endo-β-glucanase cellodextrins cellobiohydrolase cellobiose β-glucosidase glucose Synergistic effect of cellulases
Degradation of cellulose cellobiohydrolase endo-β-glucanase cellodextrins cellobiohydrolase cellobiose β-glucosidase glucose Synergistic effect of cellulases
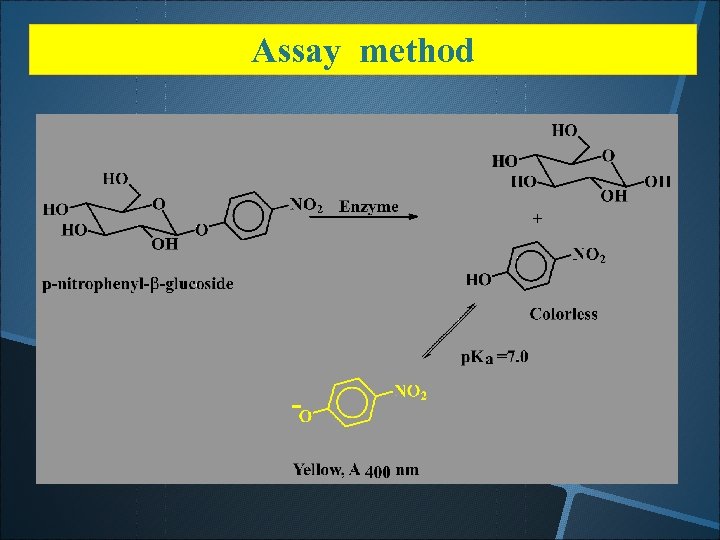 Assay method
Assay method
 Screening for b-glucosidase Acetobacter pasteurianus Neisseria subflava Micrococcus luteus Nocardia brevicatena Thermoactinomyces candidus Azospirillum brasilense Rhodococcus sp. Enterococcus faecalis Flavobacterium meningosepticum
Screening for b-glucosidase Acetobacter pasteurianus Neisseria subflava Micrococcus luteus Nocardia brevicatena Thermoactinomyces candidus Azospirillum brasilense Rhodococcus sp. Enterococcus faecalis Flavobacterium meningosepticum
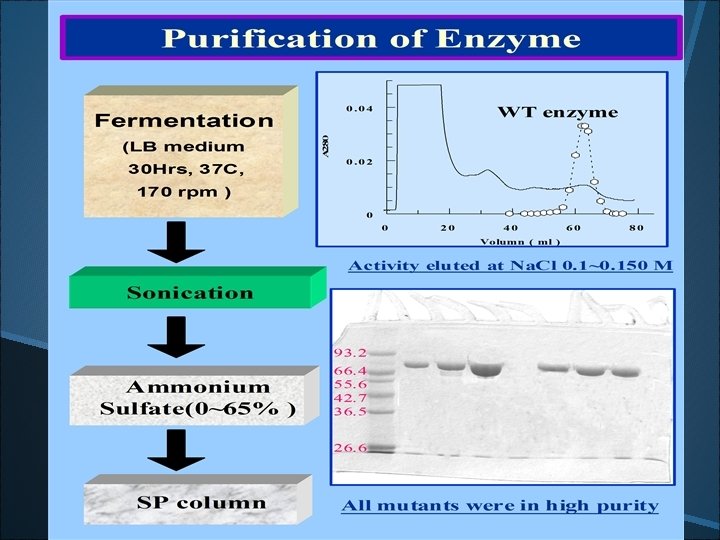
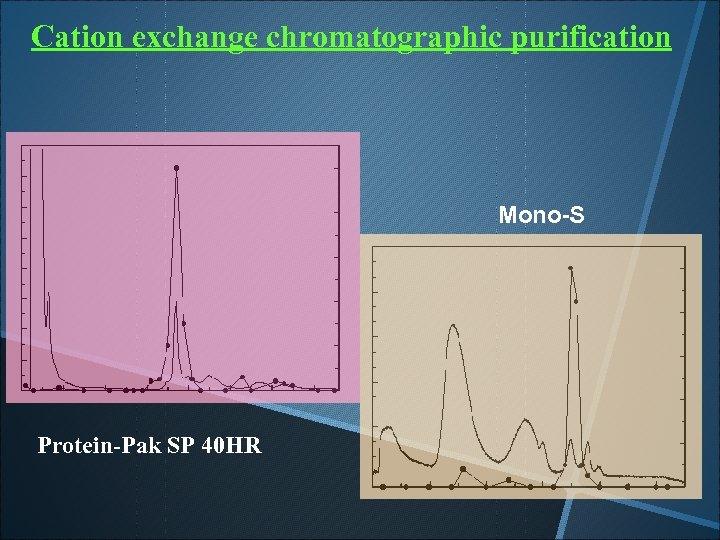 Cation exchange chromatographic purification Mono-S Protein-Pak SP 40 HR
Cation exchange chromatographic purification Mono-S Protein-Pak SP 40 HR
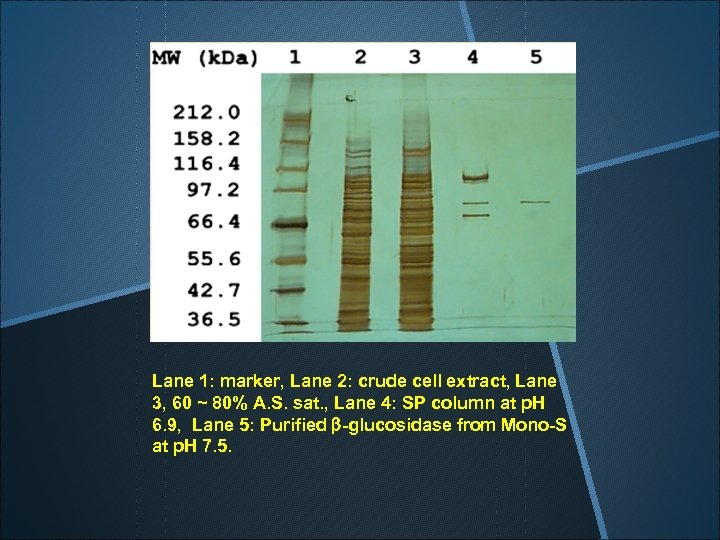 Lane 1: marker, Lane 2: crude cell extract, Lane 3, 60 ~ 80% A. S. sat. , Lane 4: SP column at p. H 6. 9, Lane 5: Purified b-glucosidase from Mono-S at p. H 7. 5.
Lane 1: marker, Lane 2: crude cell extract, Lane 3, 60 ~ 80% A. S. sat. , Lane 4: SP column at p. H 6. 9, Lane 5: Purified b-glucosidase from Mono-S at p. H 7. 5.
 Characteristics of b-glucosidase u No significant activity on hydrolysis of other glycosides. u The estimated Mr. of the enzyme is 150 k. Da by gel filtration and 78 k. Da by SDS-PAGE. u This dimeric enzyme has a p. I= 9. 0 and an optimal activity at p. H 5. 0 and temperature of 50 ℃.
Characteristics of b-glucosidase u No significant activity on hydrolysis of other glycosides. u The estimated Mr. of the enzyme is 150 k. Da by gel filtration and 78 k. Da by SDS-PAGE. u This dimeric enzyme has a p. I= 9. 0 and an optimal activity at p. H 5. 0 and temperature of 50 ℃.
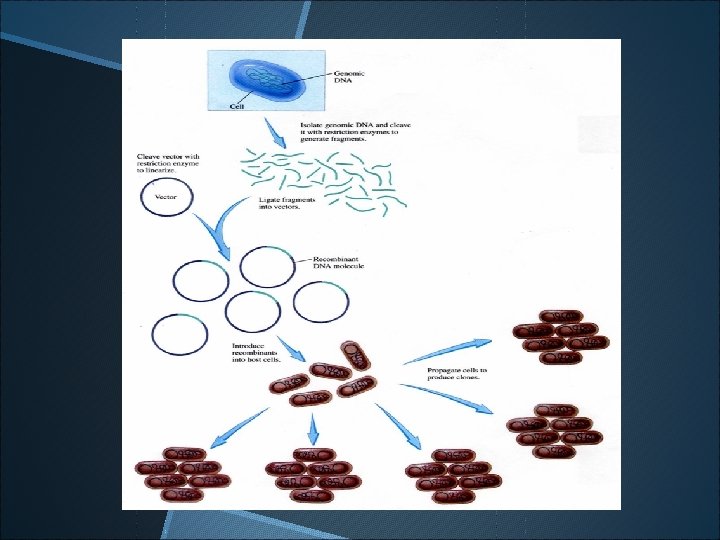
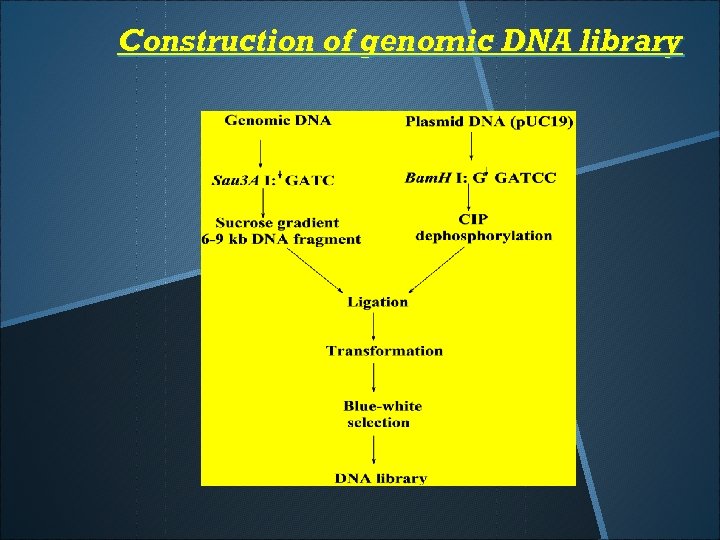 Construction of genomic DNA library
Construction of genomic DNA library
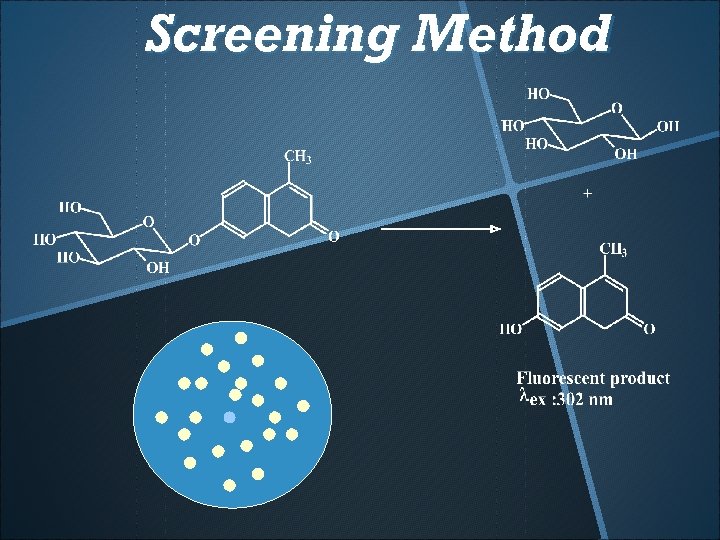 Screening Method
Screening Method
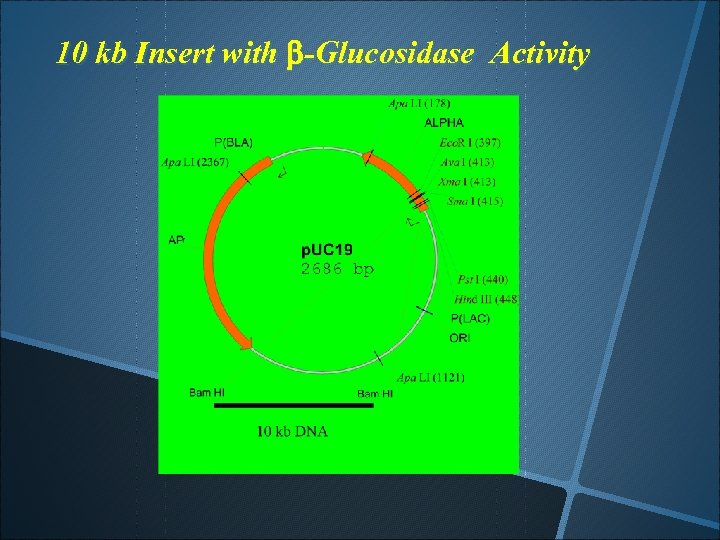 10 kb Insert with b-Glucosidase Activity
10 kb Insert with b-Glucosidase Activity
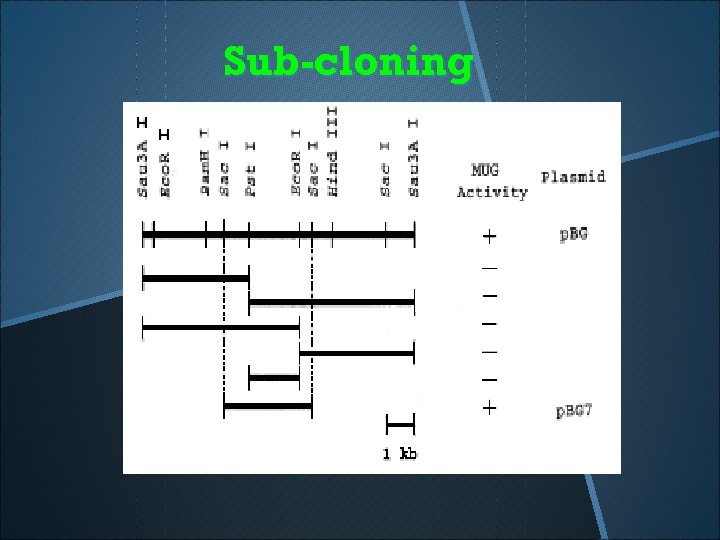 Sub-cloning
Sub-cloning
 Li, Y-K. * and Lee, J-A. 1999 “Cloning and expression of β-glucosidase from Flavobacterium meningosepticum: a new member of family B β-glucosidase” Enzym. Microb. Technol. 24, 144 -150.
Li, Y-K. * and Lee, J-A. 1999 “Cloning and expression of β-glucosidase from Flavobacterium meningosepticum: a new member of family B β-glucosidase” Enzym. Microb. Technol. 24, 144 -150.
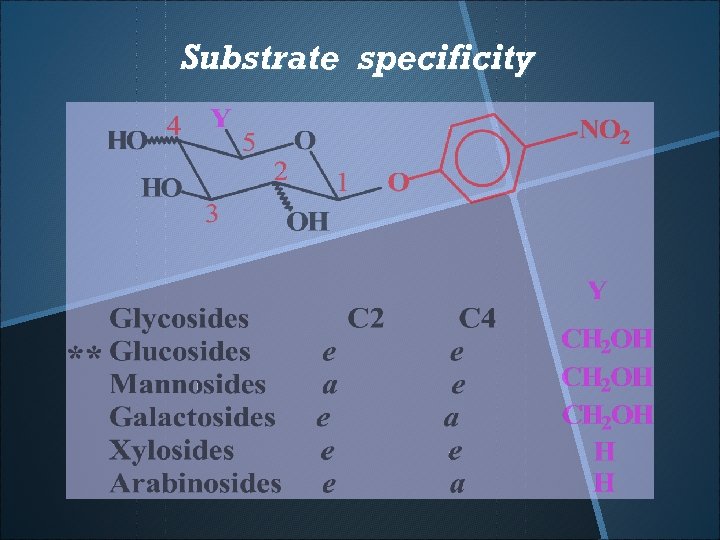 Substrate specificity
Substrate specificity
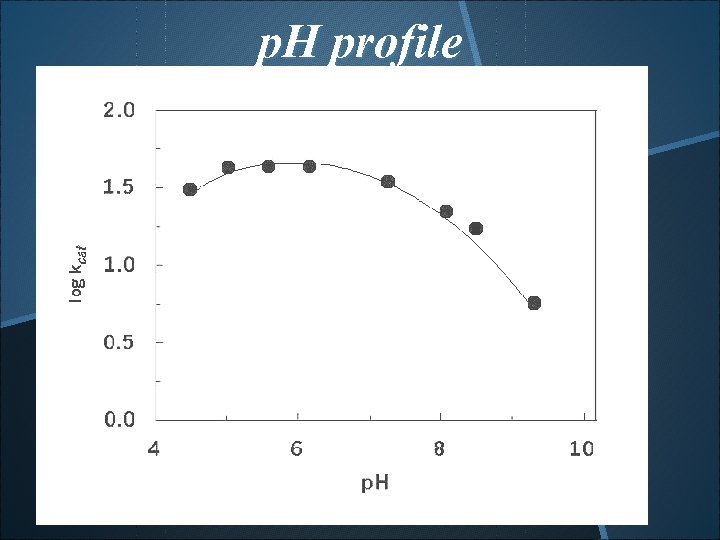 p. H profile p. K 1=3. 8~4. 0, p. K 2=6. 6
p. H profile p. K 1=3. 8~4. 0, p. K 2=6. 6
 p. K 1 p. K 2
p. K 1 p. K 2
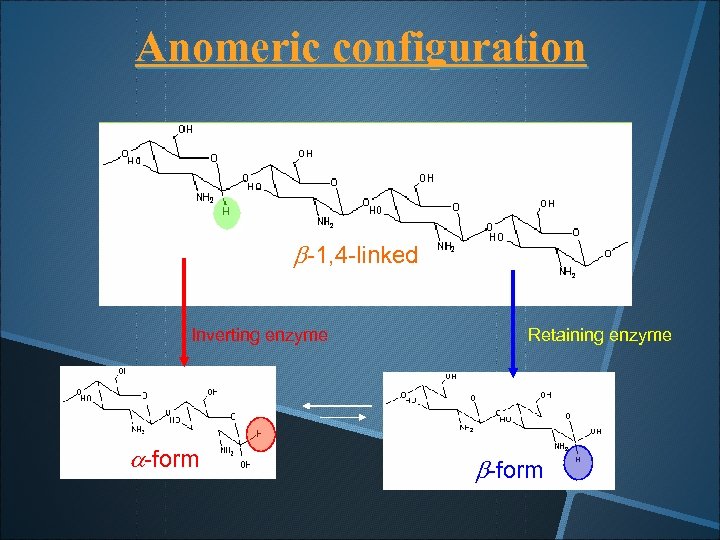 Anomeric configuration b-1, 4 -linked Inverting enzyme a-form Retaining enzyme b-form
Anomeric configuration b-1, 4 -linked Inverting enzyme a-form Retaining enzyme b-form
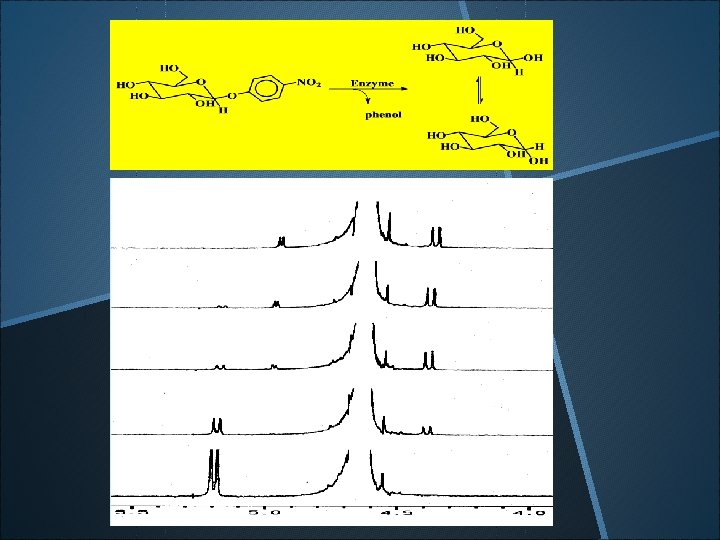 a-H 35 min 25 min 15 min 0 min b-H
a-H 35 min 25 min 15 min 0 min b-H
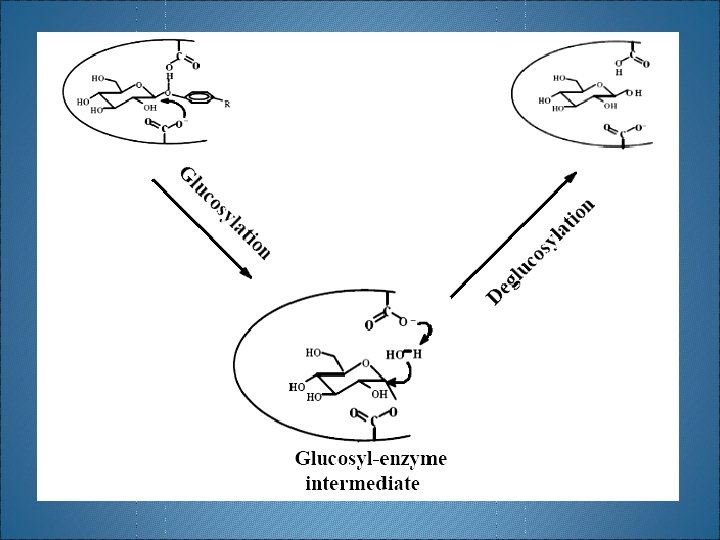 H 2 O Phenols
H 2 O Phenols
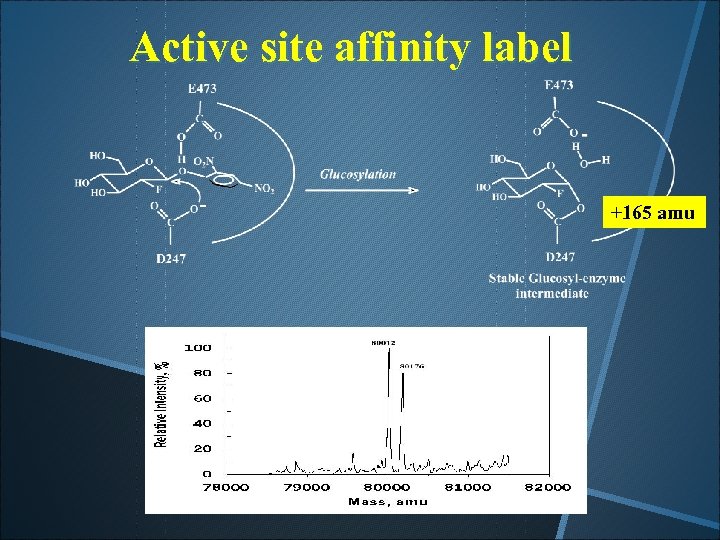 Active site affinity label +165 amu
Active site affinity label +165 amu
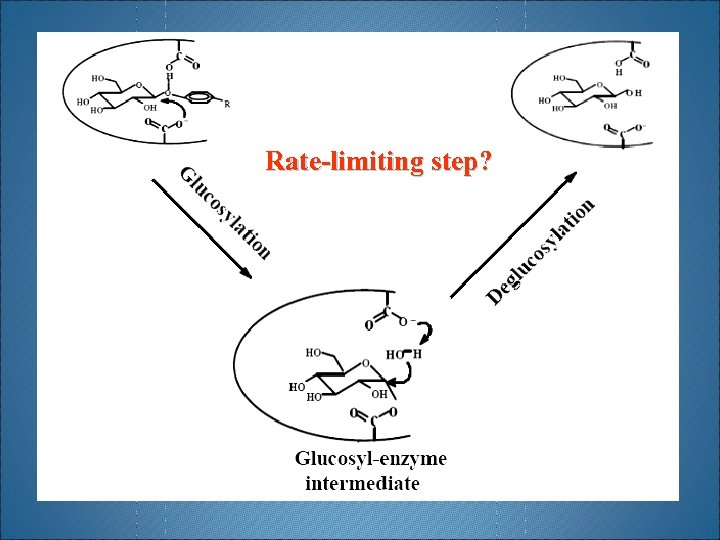 Rate-limiting step? H 2 O Phenols
Rate-limiting step? H 2 O Phenols
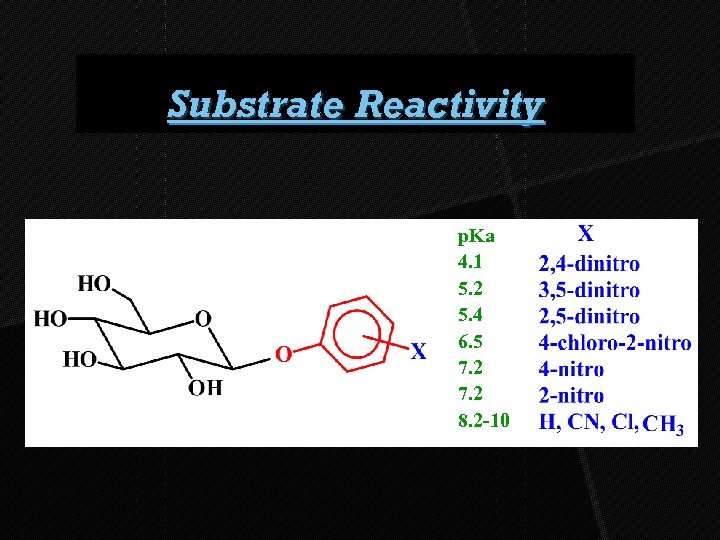 Substrate Reactivity p. Ka 4. 1 5. 2 5. 4 6. 5 7. 2 8. 2 -10
Substrate Reactivity p. Ka 4. 1 5. 2 5. 4 6. 5 7. 2 8. 2 -10
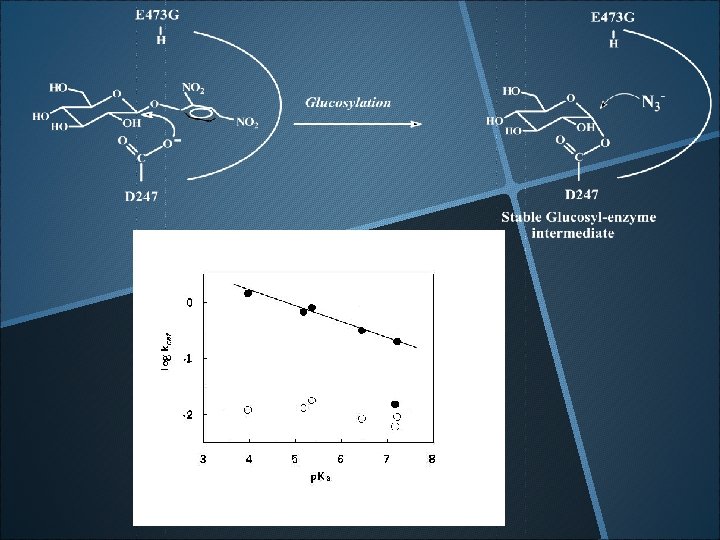
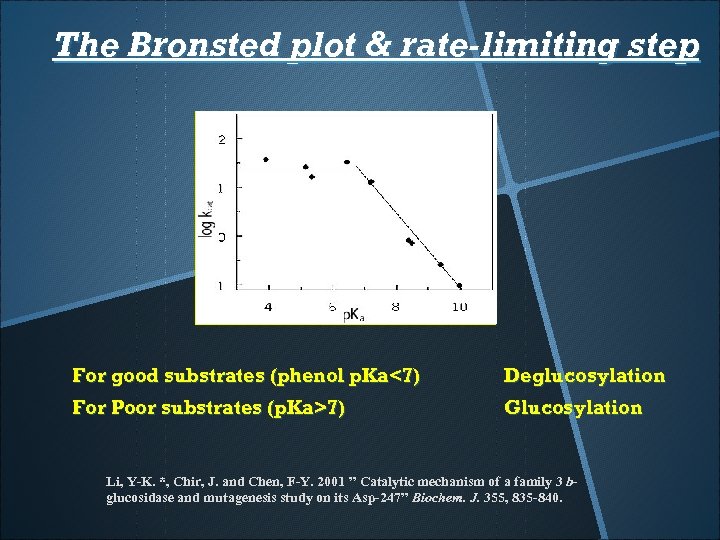 The Bronsted plot & rate-limiting step For good substrates (phenol p. Ka<7) Deglucosylation For Poor substrates (p. Ka>7) Glucosylation Li, Y-K. *, Chir, J. and Chen, F-Y. 2001 ” Catalytic mechanism of a family 3 bglucosidase and mutagenesis study on its Asp-247” Biochem. J. 355, 835 -840.
The Bronsted plot & rate-limiting step For good substrates (phenol p. Ka<7) Deglucosylation For Poor substrates (p. Ka>7) Glucosylation Li, Y-K. *, Chir, J. and Chen, F-Y. 2001 ” Catalytic mechanism of a family 3 bglucosidase and mutagenesis study on its Asp-247” Biochem. J. 355, 835 -840.
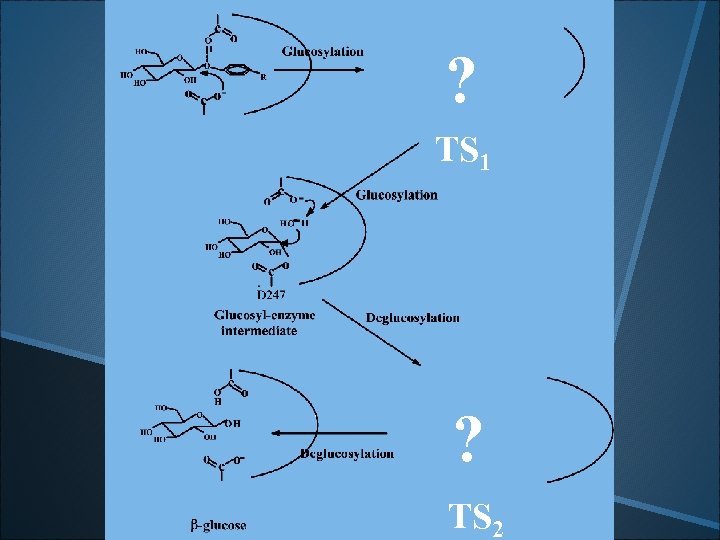 . ? TS 1 ? . TS 2
. ? TS 1 ? . TS 2
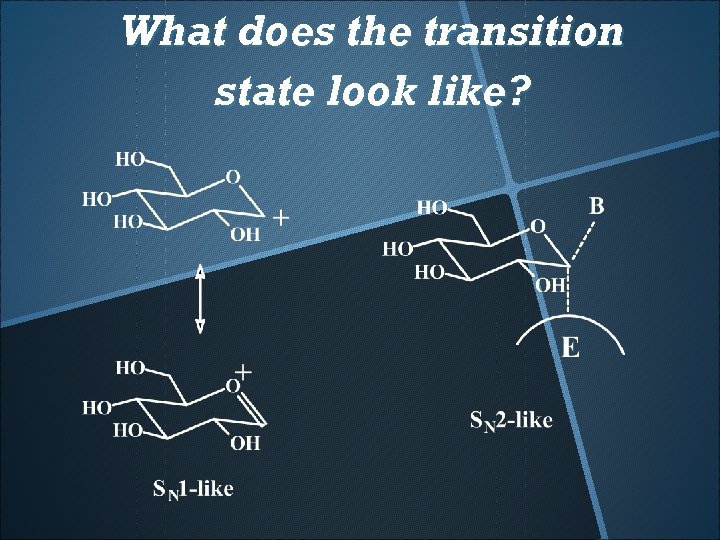 What does the transition state look like?
What does the transition state look like?
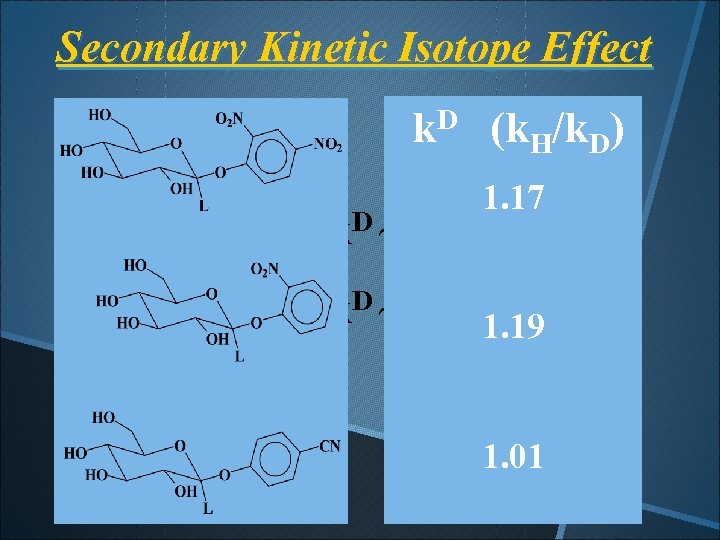 Secondary Kinetic Isotope Effect k. D (k. H/k. D) For SN D ~ 1 -like: k 1. 17 1. 15 -1. 20 For SN 2 -like: k. D ~ 1. 0 1. 19 1. 01
Secondary Kinetic Isotope Effect k. D (k. H/k. D) For SN D ~ 1 -like: k 1. 17 1. 15 -1. 20 For SN 2 -like: k. D ~ 1. 0 1. 19 1. 01
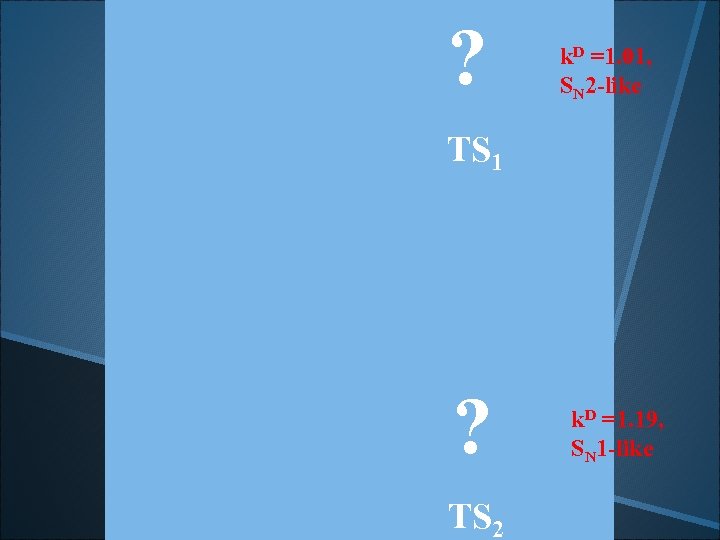 ? . k. D =1. 01, SN 2 -like TS 1 ? . TS 2 k. D =1. 19, SN 1 -like
? . k. D =1. 01, SN 2 -like TS 1 ? . TS 2 k. D =1. 19, SN 1 -like
 Which are the essential residues of b-Glucosidase?
Which are the essential residues of b-Glucosidase?
 AF 015915 69 GMDVIHG 127 WGRVSEGSGEDPY 167 VKHFALYGAPEG 241 NGFIV TDY 454 ANKADVVVLAIGETAELSGESSS AF 005277 46 LSDGPTG 114 GGRLFEAYSEDPL 148 LKHLVANES-ET 222 TGLVM SDW 567 AAQADVAVVVVGLTEEEETESVD AL 355920 43 LSDGPNG 111 NGRGFESFSEDST 145 IKHFVCNDM-ED 219 KGTII SDW 559 AKSVDCVILCVGLTAEWETEGED X 05918 53 VSDGPNG 111 GGRGFESFSEDPY 145 VKHFVCNDL-ED 219 DGMLM SDW 571 AAKHDKAVLIIGLNGEWETEGYD M 59852 39 VTDGPNG 108 NGRNFECYSEDPA 142 IKHFVANES-EI 216 DGVVM SDW 542 ARKSDIVLLLVGREGEWDTEGLD X 15644 42 MTDGPHG 117 CGRNFEYFPEDPY 151 LKHFAANNQ-EH 226 DGFVV SDW 401 ASSSDVAVVFAGLPDEYESEGFD Z 94045 42 VSDGPHG 117 SGRNFEYFSEDPY 151 LKHFAANNQ-EH 225 EGIVV SDW 401 ALKADVAVIFAGLPEHYECEGYD U 92808 52 VSDGPHG 125 CGRNFEYFSEDPY 159 LKHFAANNQ-EH 233 DGLVM SDW 404 AMNADKVVVFAGLPDSFESEGFD D 14068 97 ETDAGQG 177 NGRNFEYAGEDPL 211 LKHFVLNDQ-ET 285 RGYVM SDW 475 AAGADVALVFAN---QWIGEAND AB 003689 94 ISDAGLG 163 GGRNFEYAGEDPL 197 LKHYAMNDL-ET 271 PGFVM SDW 461 ARAADVVVVYAT---QFTFEGMD AF 090429 97 ETDASLG 166 NGRNFEYLGEDPL 200 VKHFSLNGQ-ET 274 KGWVM SDW 466 ARQSDIVILFAN---QWMSEGMD Y 14327 81 GTDGPAG 144 AGRNFETFSEDPL 178 AKHYAANTQ-ET 251 KGWVM SDW 544 ARDSDVAVVFAY---DDGAETAD D 86507 133 AYDVVHG 167 WGRASEGFGEDTY 207 VKHFAAYGAVEG 281 KGITV SDH 499 AKQADVVVAVVGESQGMAHEASS U 00007 133 AYDVLHG 191 WGRASEGFGEDTY 231 VKHFAAYGAVEG 305 KGITV SDH 523 AKQSDVVVAVVGEAQGMAHEASS AF 006658 110 GMDVIHG 168 WGRVSEGNGEDPF 208 VKHFALYGASEA 282 DGFVV TDY 495 AAGADVIVAALGESSEMSGESSS AF 015915 Flavobacterium meningosepticum (This study) D 14068 Cellvibrio gilvus ATCC 13127 AF 005277 Cellulomonas biazotea AB 003689 Acetobacter xylinus BPR 2001 AL 355920 Schizosaccharomyces pombe AF 090429 Azospirillum irakense KBC 1 X 05918 Kluyveromyces fragilis Y 14327 Saccharopolyspora erythraea M 59852 Agrobacterium tumefaciens D 86507 Salmonella typhimurium LT 2 X 15644 Clostridium thermocellum ATCC 27405 U 00007 Escherichia coli K 12/BHB 2600 Z 94045 Clostridium stercorarium AF 006658 Bacteroides fragilis 638 R U 92808 Ruminococcus albus (15 sequences are aligned. )
AF 015915 69 GMDVIHG 127 WGRVSEGSGEDPY 167 VKHFALYGAPEG 241 NGFIV TDY 454 ANKADVVVLAIGETAELSGESSS AF 005277 46 LSDGPTG 114 GGRLFEAYSEDPL 148 LKHLVANES-ET 222 TGLVM SDW 567 AAQADVAVVVVGLTEEEETESVD AL 355920 43 LSDGPNG 111 NGRGFESFSEDST 145 IKHFVCNDM-ED 219 KGTII SDW 559 AKSVDCVILCVGLTAEWETEGED X 05918 53 VSDGPNG 111 GGRGFESFSEDPY 145 VKHFVCNDL-ED 219 DGMLM SDW 571 AAKHDKAVLIIGLNGEWETEGYD M 59852 39 VTDGPNG 108 NGRNFECYSEDPA 142 IKHFVANES-EI 216 DGVVM SDW 542 ARKSDIVLLLVGREGEWDTEGLD X 15644 42 MTDGPHG 117 CGRNFEYFPEDPY 151 LKHFAANNQ-EH 226 DGFVV SDW 401 ASSSDVAVVFAGLPDEYESEGFD Z 94045 42 VSDGPHG 117 SGRNFEYFSEDPY 151 LKHFAANNQ-EH 225 EGIVV SDW 401 ALKADVAVIFAGLPEHYECEGYD U 92808 52 VSDGPHG 125 CGRNFEYFSEDPY 159 LKHFAANNQ-EH 233 DGLVM SDW 404 AMNADKVVVFAGLPDSFESEGFD D 14068 97 ETDAGQG 177 NGRNFEYAGEDPL 211 LKHFVLNDQ-ET 285 RGYVM SDW 475 AAGADVALVFAN---QWIGEAND AB 003689 94 ISDAGLG 163 GGRNFEYAGEDPL 197 LKHYAMNDL-ET 271 PGFVM SDW 461 ARAADVVVVYAT---QFTFEGMD AF 090429 97 ETDASLG 166 NGRNFEYLGEDPL 200 VKHFSLNGQ-ET 274 KGWVM SDW 466 ARQSDIVILFAN---QWMSEGMD Y 14327 81 GTDGPAG 144 AGRNFETFSEDPL 178 AKHYAANTQ-ET 251 KGWVM SDW 544 ARDSDVAVVFAY---DDGAETAD D 86507 133 AYDVVHG 167 WGRASEGFGEDTY 207 VKHFAAYGAVEG 281 KGITV SDH 499 AKQADVVVAVVGESQGMAHEASS U 00007 133 AYDVLHG 191 WGRASEGFGEDTY 231 VKHFAAYGAVEG 305 KGITV SDH 523 AKQSDVVVAVVGEAQGMAHEASS AF 006658 110 GMDVIHG 168 WGRVSEGNGEDPF 208 VKHFALYGASEA 282 DGFVV TDY 495 AAGADVIVAALGESSEMSGESSS AF 015915 Flavobacterium meningosepticum (This study) D 14068 Cellvibrio gilvus ATCC 13127 AF 005277 Cellulomonas biazotea AB 003689 Acetobacter xylinus BPR 2001 AL 355920 Schizosaccharomyces pombe AF 090429 Azospirillum irakense KBC 1 X 05918 Kluyveromyces fragilis Y 14327 Saccharopolyspora erythraea M 59852 Agrobacterium tumefaciens D 86507 Salmonella typhimurium LT 2 X 15644 Clostridium thermocellum ATCC 27405 U 00007 Escherichia coli K 12/BHB 2600 Z 94045 Clostridium stercorarium AF 006658 Bacteroides fragilis 638 R U 92808 Ruminococcus albus (15 sequences are aligned. )
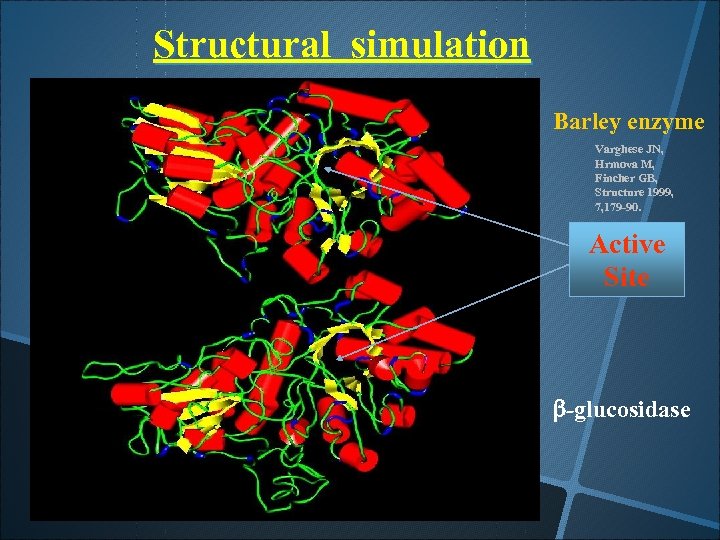 Structural simulation Barley enzyme Varghese JN, Hrmova M, Fincher GB, Structure 1999, 7, 179 -90. Active Site b-glucosidase
Structural simulation Barley enzyme Varghese JN, Hrmova M, Fincher GB, Structure 1999, 7, 179 -90. Active Site b-glucosidase
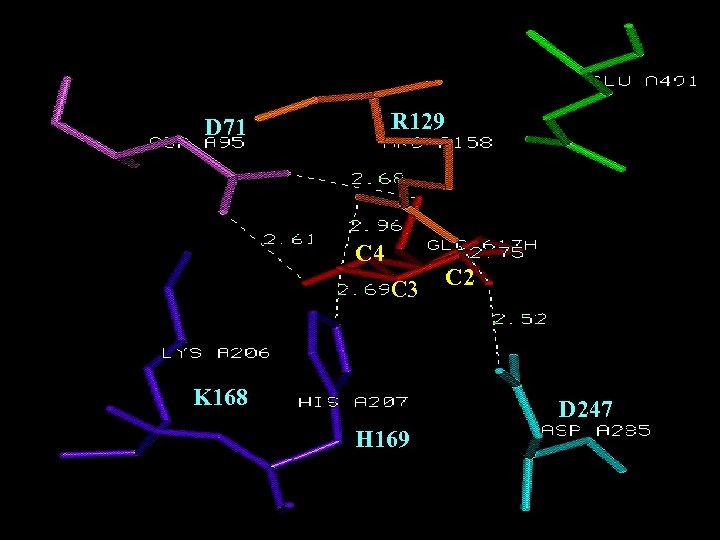 Conserved Sequences R 129 D 71 C 4 C 3 K 168 C 2 D 247 H 169
Conserved Sequences R 129 D 71 C 4 C 3 K 168 C 2 D 247 H 169
 Quick change Mutagenesis
Quick change Mutagenesis
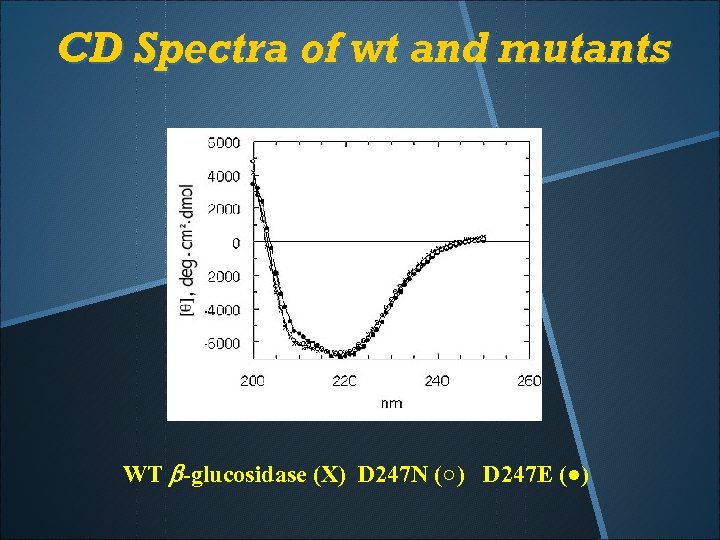 CD Spectra of wt and mutants WT b-glucosidase (X) D 247 N (○) D 247 E (●)
CD Spectra of wt and mutants WT b-glucosidase (X) D 247 N (○) D 247 E (●)
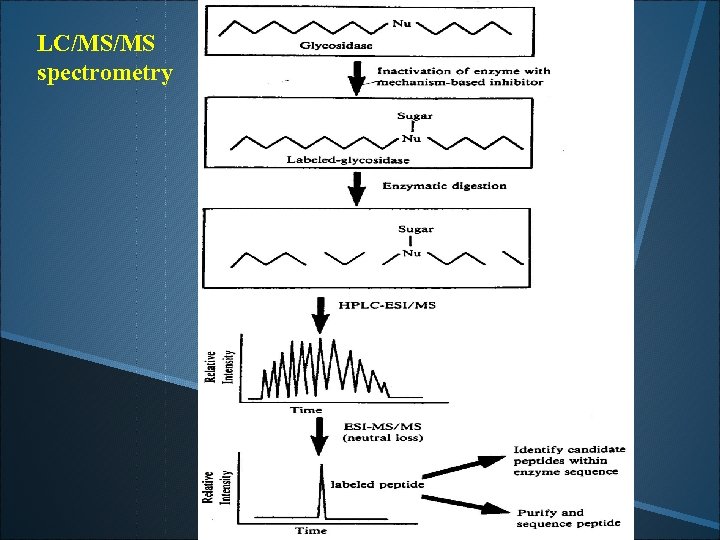 LC/MS/MS spectrometry
LC/MS/MS spectrometry
 Active site affinity label
Active site affinity label
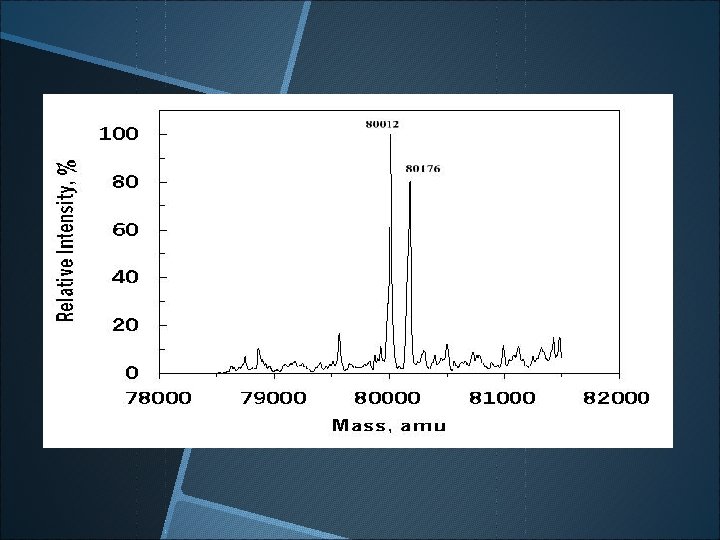
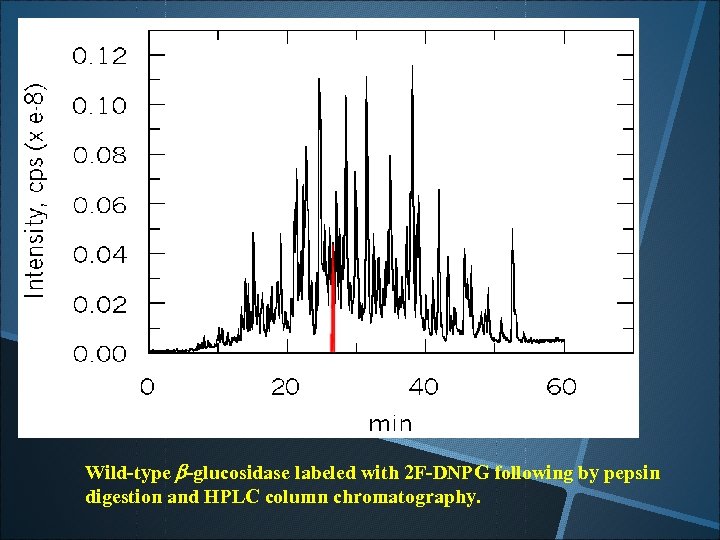 Wild-type b-glucosidase labeled with 2 F-DNPG following by pepsin digestion and HPLC column chromatography.
Wild-type b-glucosidase labeled with 2 F-DNPG following by pepsin digestion and HPLC column chromatography.
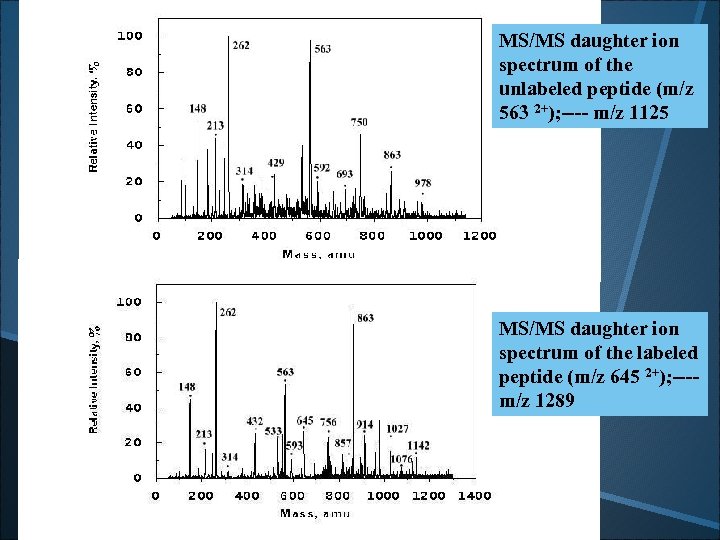 MS/MS daughter ion spectrum of the unlabeled peptide (m/z 563 2+); ---- m/z 1125 MS/MS daughter ion spectrum of the labeled peptide (m/z 645 2+); ---m/z 1289
MS/MS daughter ion spectrum of the unlabeled peptide (m/z 563 2+); ---- m/z 1125 MS/MS daughter ion spectrum of the labeled peptide (m/z 645 2+); ---m/z 1289
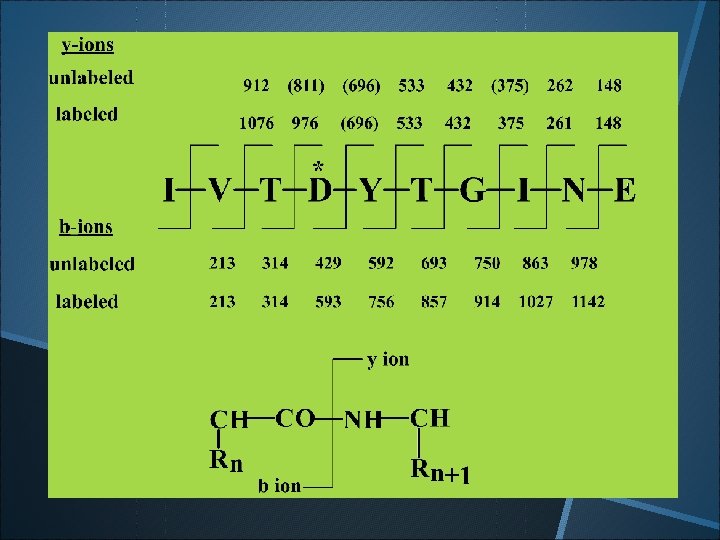
 D 247 in TDY sequence is the catalytic nucleophile!
D 247 in TDY sequence is the catalytic nucleophile!
 Does E 473 serve as general acid/base? Li, Y-K. *, Chir, J. , Tanaka, S. and Chen, F-Y. (2002) Biochemistry, 41, 2751 -2759.
Does E 473 serve as general acid/base? Li, Y-K. *, Chir, J. , Tanaka, S. and Chen, F-Y. (2002) Biochemistry, 41, 2751 -2759.
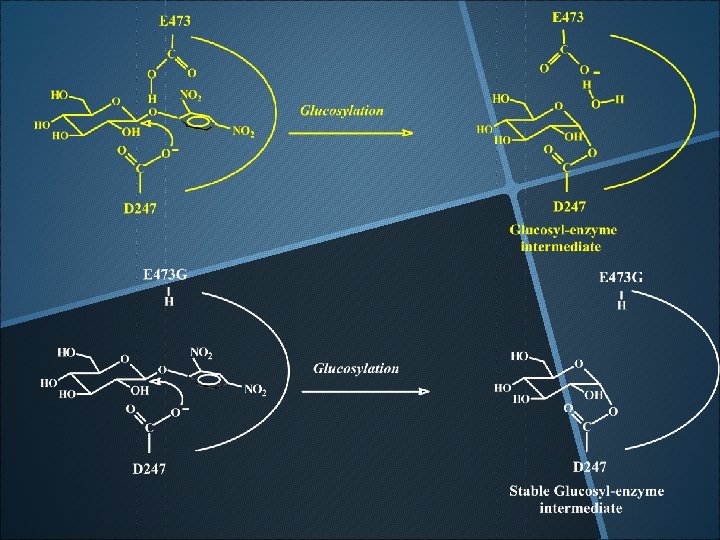
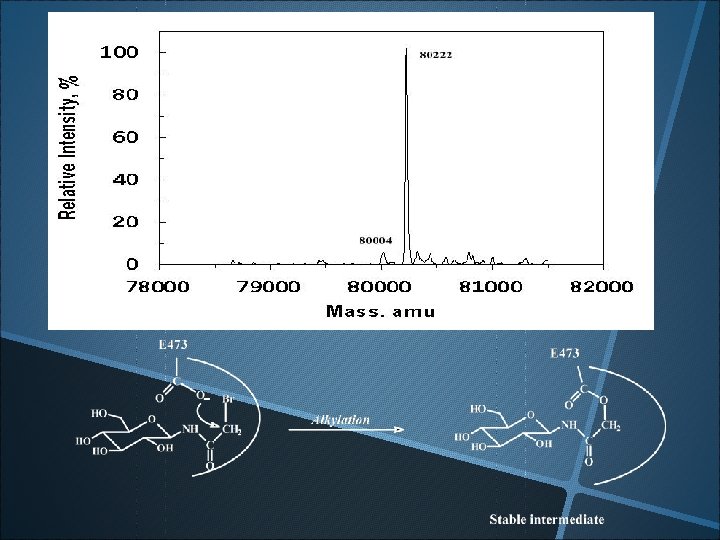
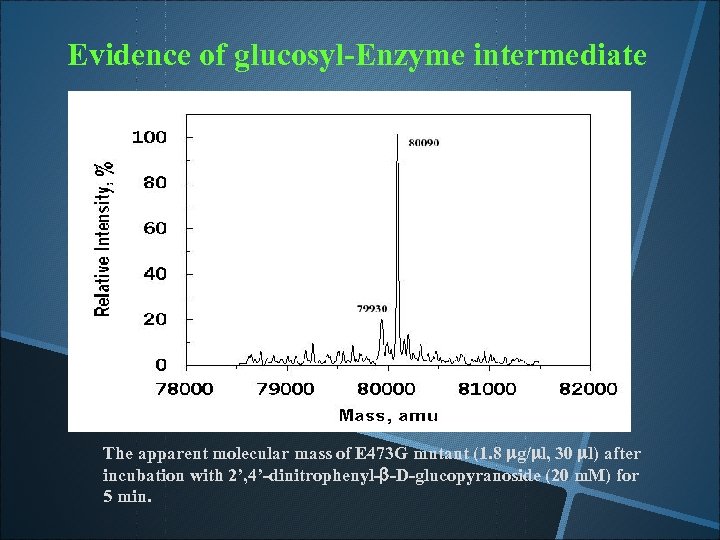 Evidence of glucosyl-Enzyme intermediate The apparent molecular mass of E 473 G mutant (1. 8 mg/ml, 30 ml) after incubation with 2’, 4’-dinitrophenyl-b-D-glucopyranoside (20 m. M) for 5 min.
Evidence of glucosyl-Enzyme intermediate The apparent molecular mass of E 473 G mutant (1. 8 mg/ml, 30 ml) after incubation with 2’, 4’-dinitrophenyl-b-D-glucopyranoside (20 m. M) for 5 min.
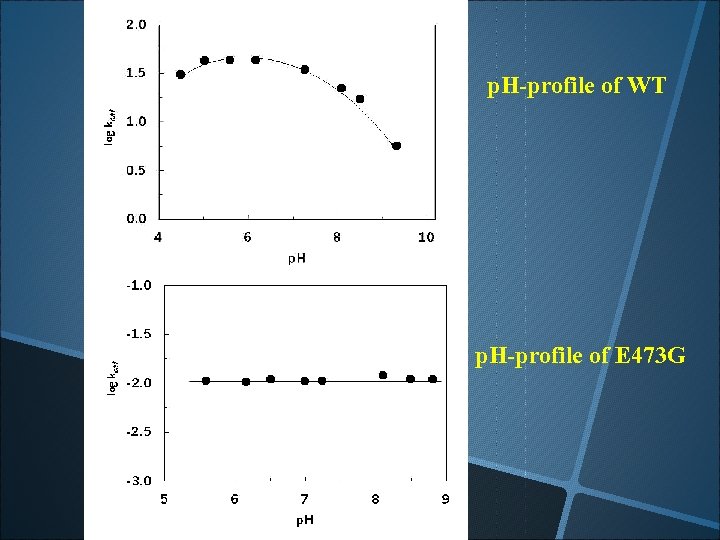 p. H-profile of WT p. H-profile of E 473 G
p. H-profile of WT p. H-profile of E 473 G
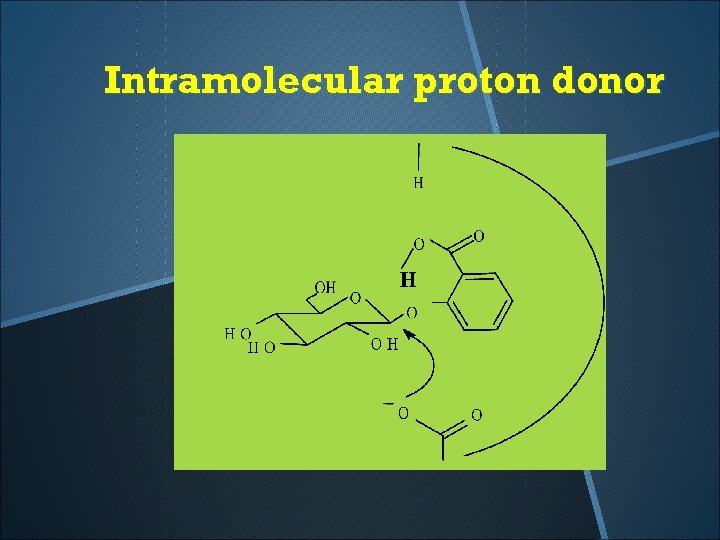 Intramolecular proton donor
Intramolecular proton donor
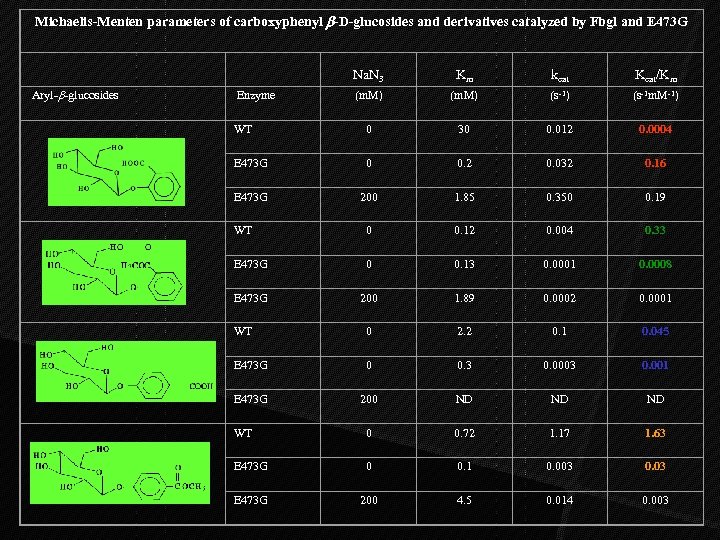 Michaelis-Menten parameters of carboxyphenyl b-D-glucosides and derivatives catalyzed by Fbgl and E 473 G Na. N 3 Km kcat Kcat/Km (m. M) (s-1) (s-1 m. M-1) Aryl-b-glucosides Enzyme WT 0 30 0. 012 0. 0004 E 473 G 0 0. 2 0. 032 0. 16 E 473 G 200 1. 85 0. 350 0. 19 WT 0 0. 12 0. 004 0. 33 E 473 G 0 0. 13 0. 0001 0. 0008 E 473 G 200 1. 89 0. 0002 0. 0001 WT 0 2. 2 0. 1 0. 045 E 473 G 0 0. 3 0. 0003 0. 001 E 473 G 200 ND ND ND WT 0 0. 72 1. 17 1. 63 E 473 G 0 0. 1 0. 003 0. 03 E 473 G 200 4. 5 0. 014 0. 003
Michaelis-Menten parameters of carboxyphenyl b-D-glucosides and derivatives catalyzed by Fbgl and E 473 G Na. N 3 Km kcat Kcat/Km (m. M) (s-1) (s-1 m. M-1) Aryl-b-glucosides Enzyme WT 0 30 0. 012 0. 0004 E 473 G 0 0. 2 0. 032 0. 16 E 473 G 200 1. 85 0. 350 0. 19 WT 0 0. 12 0. 004 0. 33 E 473 G 0 0. 13 0. 0001 0. 0008 E 473 G 200 1. 89 0. 0002 0. 0001 WT 0 2. 2 0. 1 0. 045 E 473 G 0 0. 3 0. 0003 0. 001 E 473 G 200 ND ND ND WT 0 0. 72 1. 17 1. 63 E 473 G 0 0. 1 0. 003 0. 03 E 473 G 200 4. 5 0. 014 0. 003
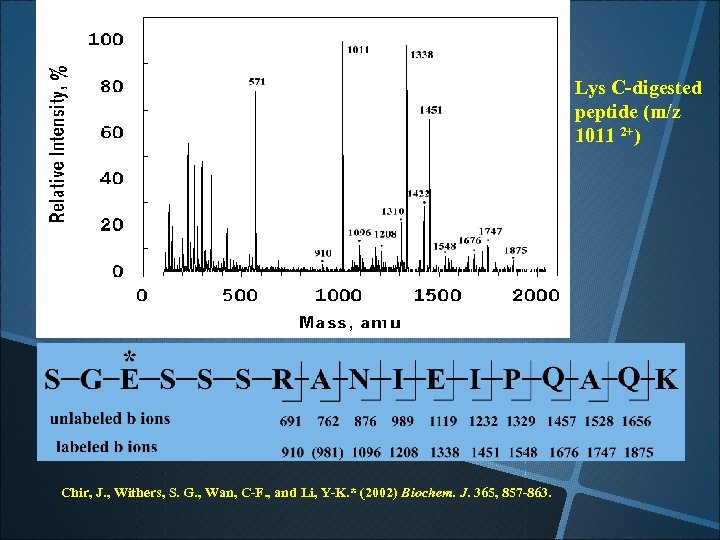 Lys C-digested peptide (m/z 1011 2+) Chir, J. , Withers, S. G. , Wan, C-F. , and Li, Y-K. * (2002) Biochem. J. 365, 857 -863.
Lys C-digested peptide (m/z 1011 2+) Chir, J. , Withers, S. G. , Wan, C-F. , and Li, Y-K. * (2002) Biochem. J. 365, 857 -863.
 E 473 is the acid/base catalyst of the enzyme!
E 473 is the acid/base catalyst of the enzyme!
 Summary A b-glucosidase containing 726 amino acid from F. meningosepticum was cloned and identified as a new member of GH-3 enzyme GH-3 b-glucosidase involves a two-step, double displacement mechanism. D 247 and E 473 function as the nucleophile and the general acid/base catalyst, respectively.
Summary A b-glucosidase containing 726 amino acid from F. meningosepticum was cloned and identified as a new member of GH-3 enzyme GH-3 b-glucosidase involves a two-step, double displacement mechanism. D 247 and E 473 function as the nucleophile and the general acid/base catalyst, respectively.


