8db9ca3f1ab9dbbdb0a9cb8cc7dbc5f7.ppt
- Количество слайдов: 11
 Manual Annotation of the human and mouse gene index: www. allgenes. org
Manual Annotation of the human and mouse gene index: www. allgenes. org
 Allgenes. org A web interface providing access to the assembled EST and m. RNA sequences, or Do. TS RNA transcripts, contained within GUS (Genomics Unified Schema), a relational database. Computed & manual annotation has been applied to the human and mouse Do. TS RNA transcripts to associate them with their corresponding genes, creating a human and mouse gene index, Allgenes.
Allgenes. org A web interface providing access to the assembled EST and m. RNA sequences, or Do. TS RNA transcripts, contained within GUS (Genomics Unified Schema), a relational database. Computed & manual annotation has been applied to the human and mouse Do. TS RNA transcripts to associate them with their corresponding genes, creating a human and mouse gene index, Allgenes.
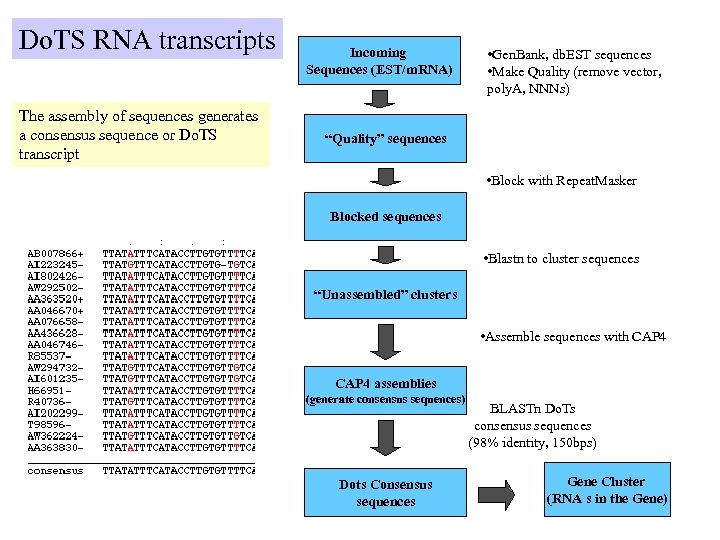 Do. TS RNA transcripts The assembly of sequences generates a consensus sequence or Do. TS transcript Incoming Sequences (EST/m. RNA) • Gen. Bank, db. EST sequences • Make Quality (remove vector, poly. A, NNNs) “Quality” sequences • Block with Repeat. Masker Blocked sequences • Blastn to cluster sequences “Unassembled” clusters • Assemble sequences with CAP 4 assemblies (generate consensus sequences) Dots Consensus sequences BLASTn Do. Ts consensus sequences (98% identity, 150 bps) Gene Cluster (RNA s in the Gene)
Do. TS RNA transcripts The assembly of sequences generates a consensus sequence or Do. TS transcript Incoming Sequences (EST/m. RNA) • Gen. Bank, db. EST sequences • Make Quality (remove vector, poly. A, NNNs) “Quality” sequences • Block with Repeat. Masker Blocked sequences • Blastn to cluster sequences “Unassembled” clusters • Assemble sequences with CAP 4 assemblies (generate consensus sequences) Dots Consensus sequences BLASTn Do. Ts consensus sequences (98% identity, 150 bps) Gene Cluster (RNA s in the Gene)
 Manual Annotation efforts utilize: A web-based tool to update GUS
Manual Annotation efforts utilize: A web-based tool to update GUS
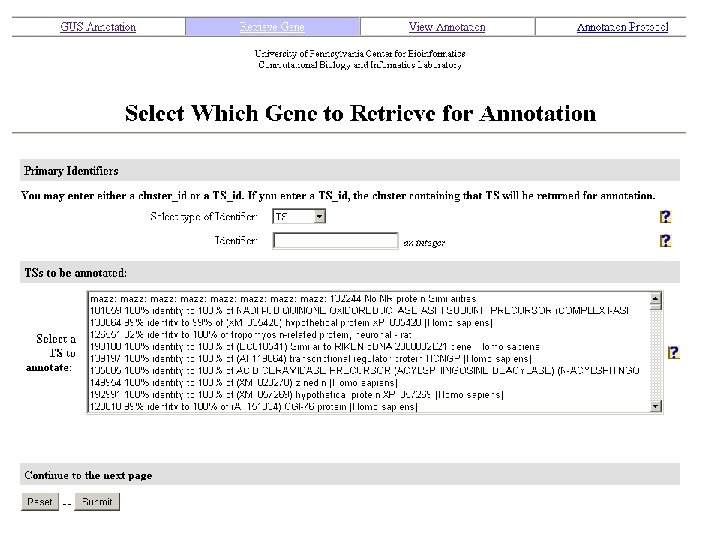
 Annotation Tasks include: On Gene page: 1. Assigning reference RNA sequence. 2. Determining members of Gene cluster (RNA transcripts) – removing or adding members - validating RNA transcripts assigned to gene using genomic alignment, BLAST similarity and/or c. DNA clone linkages 3. Adding approved abbreviated gene name or symbol (if known) and evidence Gene description field: 4. Adding approved full gene name, aliases and evidence for them 5. Adding gene synonyms and evidence for them Gene Annotation is displayed on Gene page of allgenes. org
Annotation Tasks include: On Gene page: 1. Assigning reference RNA sequence. 2. Determining members of Gene cluster (RNA transcripts) – removing or adding members - validating RNA transcripts assigned to gene using genomic alignment, BLAST similarity and/or c. DNA clone linkages 3. Adding approved abbreviated gene name or symbol (if known) and evidence Gene description field: 4. Adding approved full gene name, aliases and evidence for them 5. Adding gene synonyms and evidence for them Gene Annotation is displayed on Gene page of allgenes. org
 On RNAPage: 6. Modifying TS (RNA) description of reference sequence to reflect HUGO or MGI approved full gene name, if assigned. 7. GO Function assignment/verification – GO Function manually assigned; predicted GO Functions are verified RNA Annotation is displayed on RNA page of allgenes. org Evidence is Retrieved from GUS (e. g. , a protein domain similarity) to confirm an assignment or Evidence is added manually for the assignment (comment e. g. , source).
On RNAPage: 6. Modifying TS (RNA) description of reference sequence to reflect HUGO or MGI approved full gene name, if assigned. 7. GO Function assignment/verification – GO Function manually assigned; predicted GO Functions are verified RNA Annotation is displayed on RNA page of allgenes. org Evidence is Retrieved from GUS (e. g. , a protein domain similarity) to confirm an assignment or Evidence is added manually for the assignment (comment e. g. , source).
 Proposed New Annotation Tasks: Gene level annotation - New & Old annotator tasks; underlined are tasks that are surface level annotation 1. ) Creation of gene defined exons and introns of gene – use genomic sequence definition of gene boundaries 5’ exon boundary (transcription start site) 3’ exon boundary (poly adenylation signal) 2. ) assign gene name/abbrievated gene symbol 3. ) assign full gene name (MGI or HUGO full gene name) 4. ) assign abbrievated gene synonymns 5. ) assign full gene name aliases 6. ) assign gene category (e. g. non-coding) 7. ) confirm/assign gene chromosomal location 8. ) OMIM Link assignment (verification if computationally determined)
Proposed New Annotation Tasks: Gene level annotation - New & Old annotator tasks; underlined are tasks that are surface level annotation 1. ) Creation of gene defined exons and introns of gene – use genomic sequence definition of gene boundaries 5’ exon boundary (transcription start site) 3’ exon boundary (poly adenylation signal) 2. ) assign gene name/abbrievated gene symbol 3. ) assign full gene name (MGI or HUGO full gene name) 4. ) assign abbrievated gene synonymns 5. ) assign full gene name aliases 6. ) assign gene category (e. g. non-coding) 7. ) confirm/assign gene chromosomal location 8. ) OMIM Link assignment (verification if computationally determined)
 RNA level annotation – 1. ) Define RNA transcripts from gene (create RNAs – stable sequences) ss Using exons defined by curated gene 5’ and 3’ UTRs 2. ) Assign RNA categories to created RNAs (e. g. alternative form) 3. ) Assign/confirm RNA description 4. ) Anatomy expression assignment(s) 5. ) Assign GO terms to curated RNA (non-coding RNAs, e. g. , small RNA involved in splicing)
RNA level annotation – 1. ) Define RNA transcripts from gene (create RNAs – stable sequences) ss Using exons defined by curated gene 5’ and 3’ UTRs 2. ) Assign RNA categories to created RNAs (e. g. alternative form) 3. ) Assign/confirm RNA description 4. ) Anatomy expression assignment(s) 5. ) Assign GO terms to curated RNA (non-coding RNAs, e. g. , small RNA involved in splicing)
 Computational analysis on curated RNAs will be performed: Protein level annotation – 1. ) confirm/assign GO Function 2. ) confirm/assign GO Biological Process 3. ) confirm/assign GO Cellular Component 4. ) Assign protein name 5. ) Assign protein name synonyms 6. ) Assign Protein category (post translational modifications) 7. ) Protein-protein interactions assigned 8. ) Protein pathway assignments
Computational analysis on curated RNAs will be performed: Protein level annotation – 1. ) confirm/assign GO Function 2. ) confirm/assign GO Biological Process 3. ) confirm/assign GO Cellular Component 4. ) Assign protein name 5. ) Assign protein name synonyms 6. ) Assign Protein category (post translational modifications) 7. ) Protein-protein interactions assigned 8. ) Protein pathway assignments
 Mouse DT. 491900 No NR protein Similarities Aligns mouse chr 5 (other cluster members with fly protein similarity) (RNA_id = 229048) Predicted protein sequence (framefinder translation) MKRKASEVKEAEANAALEEEKRRQQAELEAFENRLKGRRKKSRKRDEVAVELSPWQKYKSYLLPVCAVVV AVLMWYIFHGVD (query. Seq. ) Human DT. 426371 39% identity to 44% of (AE 003480) CG 15011 gene product [Drosophila melanogaster] (other cluster member) Aligns human chr 4 Predicted Protein Sequence (framefinder translation) MLRIKCHCKITSLYVECRKITTADVNEKNLLSCCKNQCPKELPCGHRCKEMCHPGECPFNCNQKVKLRCP CKRIKKELQCNKVRENQVSIECDTTCKEMKRKASEIKEAEAKAALEEEKRRQQAELEAFENRLKGRRKKN RKRDEVAVELSLWQKHKYYLISVCGVVVVVFAWYITHDVN GO Function Prediction for human protein is /DNA binding/transcription factor Due to Zn finger domain similarity present. This type of Zn finger is found in Drosophila shuttle craft protein, a transcription factor, which has a role in the late stages of embryonic neurogenesis. Human gene may overlap with adjacent corin gene. Score = 337 (123. 7 bits), Expect = 3. 9 e-36, P = 3. 9 e-36 Identities = 67/82 (81%), Positives = 72/82 (87%) Query: Sbjct: 1 MKRKASEVKEAEANAALEEEKRRQQAELEAFENRLKGRRKKSRKRDEVAVELSPWQKYKS 60 MKRKASE+KEAEA AALEEEKRRQQAELEAFENRLKGRRKK+RKRDEVAVELS WQK+K 99 MKRKASEIKEAEAKAALEEEKRRQQAELEAFENRLKGRRKKNRKRDEVAVELSLWQKHKY 158 61 YLLPVCAVVVAVLMWYIFHGVD 82 YL+ VC VVV V WYI H V+ 159 YLISVCGVVVVVFAWYITHDVN 180 Additional DT. Mouse DT. 60100860 100% identity to 100% of (AK 005913) putative [Mus musculus] maybe 5’ end of gene (1700012 H 24 Rik & AW 538212) sequence reversed, gap in alignment – also recent update has modified the assembly 2 sequences removed (RNA_id = 10404576)
Mouse DT. 491900 No NR protein Similarities Aligns mouse chr 5 (other cluster members with fly protein similarity) (RNA_id = 229048) Predicted protein sequence (framefinder translation) MKRKASEVKEAEANAALEEEKRRQQAELEAFENRLKGRRKKSRKRDEVAVELSPWQKYKSYLLPVCAVVV AVLMWYIFHGVD (query. Seq. ) Human DT. 426371 39% identity to 44% of (AE 003480) CG 15011 gene product [Drosophila melanogaster] (other cluster member) Aligns human chr 4 Predicted Protein Sequence (framefinder translation) MLRIKCHCKITSLYVECRKITTADVNEKNLLSCCKNQCPKELPCGHRCKEMCHPGECPFNCNQKVKLRCP CKRIKKELQCNKVRENQVSIECDTTCKEMKRKASEIKEAEAKAALEEEKRRQQAELEAFENRLKGRRKKN RKRDEVAVELSLWQKHKYYLISVCGVVVVVFAWYITHDVN GO Function Prediction for human protein is /DNA binding/transcription factor Due to Zn finger domain similarity present. This type of Zn finger is found in Drosophila shuttle craft protein, a transcription factor, which has a role in the late stages of embryonic neurogenesis. Human gene may overlap with adjacent corin gene. Score = 337 (123. 7 bits), Expect = 3. 9 e-36, P = 3. 9 e-36 Identities = 67/82 (81%), Positives = 72/82 (87%) Query: Sbjct: 1 MKRKASEVKEAEANAALEEEKRRQQAELEAFENRLKGRRKKSRKRDEVAVELSPWQKYKS 60 MKRKASE+KEAEA AALEEEKRRQQAELEAFENRLKGRRKK+RKRDEVAVELS WQK+K 99 MKRKASEIKEAEAKAALEEEKRRQQAELEAFENRLKGRRKKNRKRDEVAVELSLWQKHKY 158 61 YLLPVCAVVVAVLMWYIFHGVD 82 YL+ VC VVV V WYI H V+ 159 YLISVCGVVVVVFAWYITHDVN 180 Additional DT. Mouse DT. 60100860 100% identity to 100% of (AK 005913) putative [Mus musculus] maybe 5’ end of gene (1700012 H 24 Rik & AW 538212) sequence reversed, gap in alignment – also recent update has modified the assembly 2 sequences removed (RNA_id = 10404576)


