a3316dc9e94c9c18cf8a13f6ba225af6.ppt
- Количество слайдов: 31

Low-cost, high accuracy, long-DNA synthesis technology 16 -Feb-2005 10 AM NHGRI George Church, Joe Jacobsen et al. Harvard & MIT 0. Killer Applications 1. Chip synthesis, fluidics 2. Multiplex assembly 3. Error correction methods 4. Software CAD-PAM 5. Proteome (in vitro) synthesis 6. Homologous recombination & selection for BACs 7. Integrases 8. Process integration, QA, timeline 9. Safety opportunities
Low-cost, high accuracy, long-DNA synthesis technology 16 -Feb-2005 10 AM NHGRI George Church, Joe Jacobsen et al. Harvard & MIT All stages: Error correction, Software, QA, safety 50 -100 Chip synthesis, fluidics 100 -15 k Pol-Assembly-Multiplex, Proteome synthesis 15 k-100 k Annealing assembly 100 k-5 M. Microbial recombination 100 k-200 M Mammalian recomb, integrases

Synthetic Genomes & Proteomes. Why? • Test or engineer cis-DNA/RNA-elements • Access to any protein (complex) including post-transcriptional modifications • Affinity agents for the above. • Protein design, vaccines, solubility screens • Utility of molecular biology DNA -- RNA -- Protein in vitro "kits" (e. g. PCR -- T 7 -- Roche) Toward these goals design a chassis: • 115 kbp genome. 150 genes. • Nearly all 3 D structures known. • Comprehensive functional data.
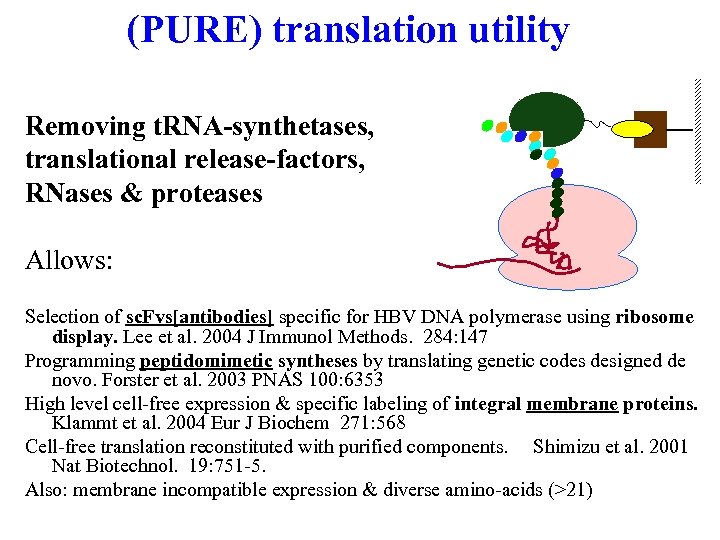

(PURE) translation utility Removing t. RNA-synthetases, translational release-factors, RNases & proteases Allows: Selection of sc. Fvs[antibodies] specific for HBV DNA polymerase using ribosome display. Lee et al. 2004 J Immunol Methods. 284: 147 Programming peptidomimetic syntheses by translating genetic codes designed de novo. Forster et al. 2003 PNAS 100: 6353 High level cell-free expression & specific labeling of integral membrane proteins. Klammt et al. 2004 Eur J Biochem 271: 568 Cell-free translation reconstituted with purified components. Shimizu et al. 2001 Nat Biotechnol. 19: 751 -5. Also: membrane incompatible expression & diverse amino-acids (>21)
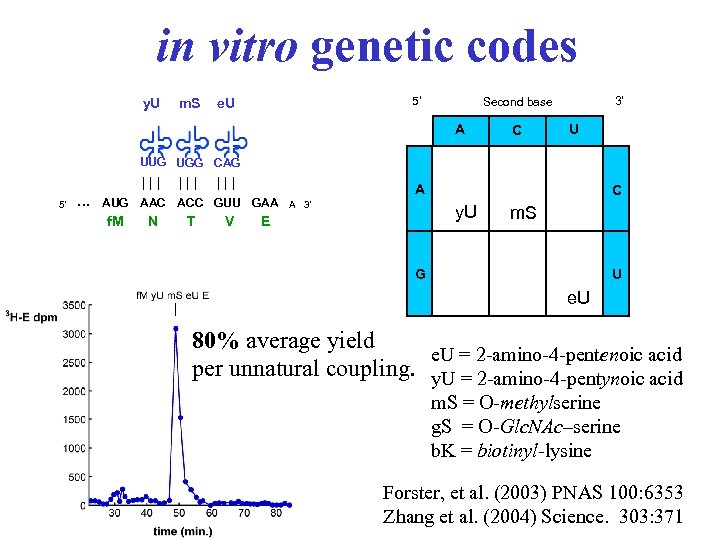
in vitro genetic codes y. U m. S 5' e. U 3' Second base A A C C U U UUG UGG CAG 5' . . . ||| ||| AUG AAC ACC GUU GAA A 3' f. M N T V A C y. U E m. S U G e. U 80% average yield e. U = 2 -amino-4 -pentenoic acid per unnatural coupling. y. U = 2 -amino-4 -pentynoic acid m. S = O-methylserine g. S = O-Glc. NAc–serine b. K = biotinyl-lysine Forster, et al. (2003) PNAS 100: 6353 Zhang et al. (2004) Science. 303: 371
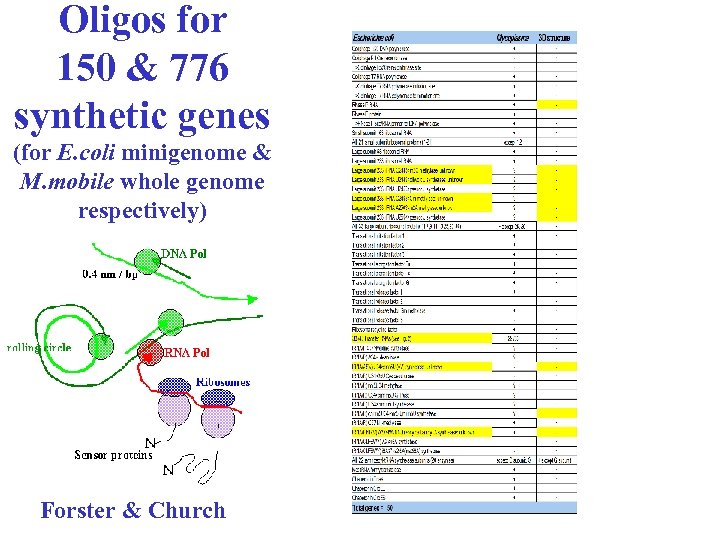

Oligos for 150 & 776 synthetic genes (for E. coli minigenome & M. mobile whole genome respectively) Forster & Church
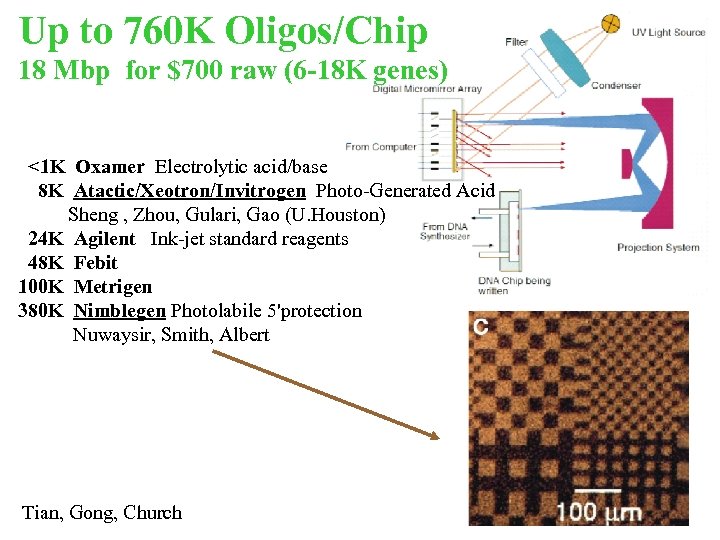
Up to 760 K Oligos/Chip 18 Mbp for $700 raw (6 -18 K genes) <1 K Oxamer Electrolytic acid/base 8 K Atactic/Xeotron/Invitrogen Photo-Generated Acid Sheng , Zhou, Gulari, Gao (U. Houston) 24 K Agilent Ink-jet standard reagents 48 K Febit 100 K Metrigen 380 K Nimblegen Photolabile 5'protection Nuwaysir, Smith, Albert Tian, Gong, Church
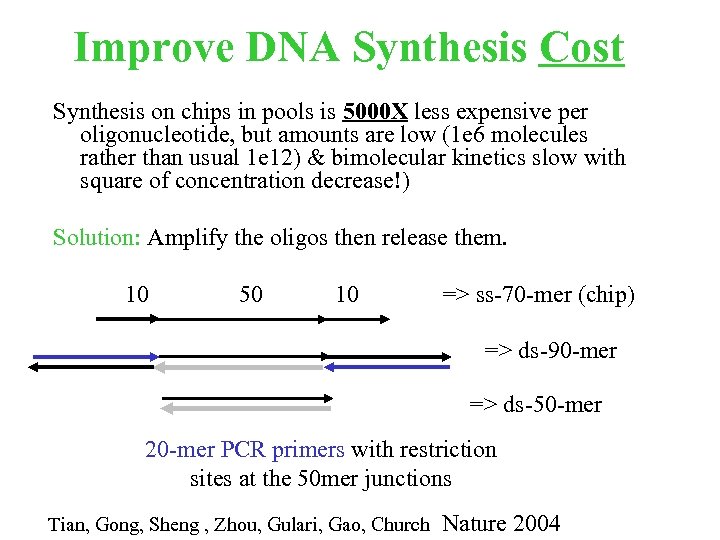

Improve DNA Synthesis Cost Synthesis on chips in pools is 5000 X less expensive per oligonucleotide, but amounts are low (1 e 6 molecules rather than usual 1 e 12) & bimolecular kinetics slow with square of concentration decrease!) Solution: Amplify the oligos then release them. 10 50 10 => ss-70 -mer (chip) => ds-90 -mer => ds-50 -mer 20 -mer PCR primers with restriction sites at the 50 mer junctions Tian, Gong, Sheng , Zhou, Gulari, Gao, Church Nature 2004
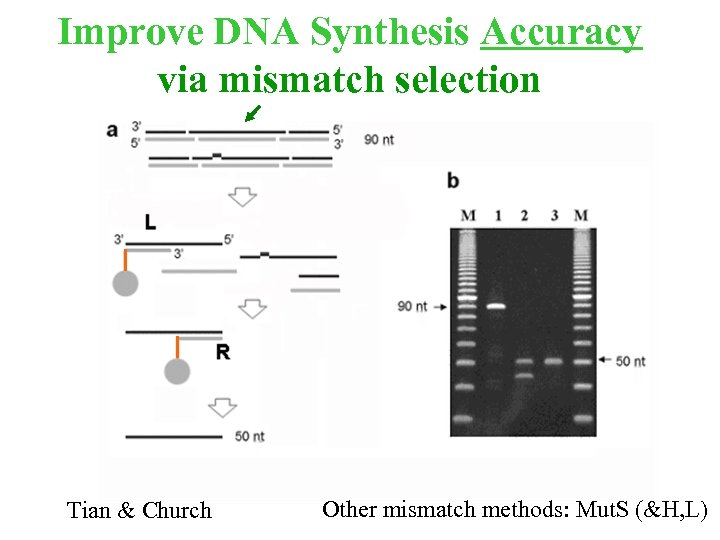
Improve DNA Synthesis Accuracy via mismatch selection Tian & Church Other mismatch methods: Mut. S (&H, L)
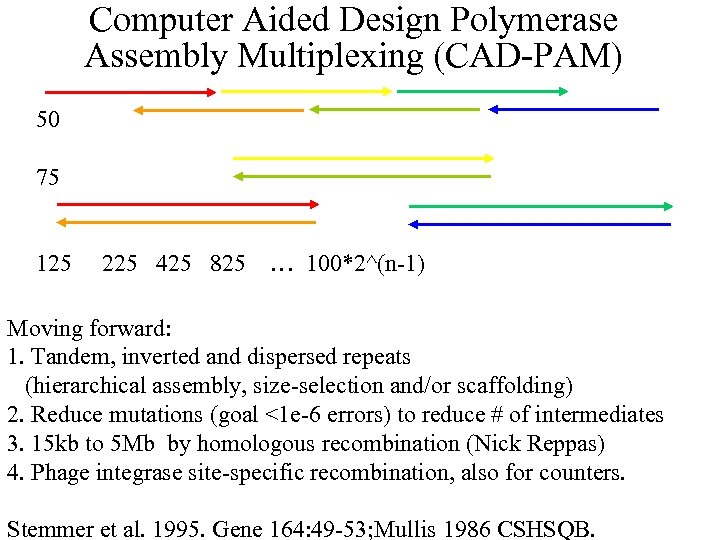
Computer Aided Design Polymerase Assembly Multiplexing (CAD-PAM) 50 75 125 225 425 825 … 100*2^(n-1) Moving forward: 1. Tandem, inverted and dispersed repeats (hierarchical assembly, size-selection and/or scaffolding) 2. Reduce mutations (goal <1 e-6 errors) to reduce # of intermediates 3. 15 kb to 5 Mb by homologous recombination (Nick Reppas) 4. Phage integrase site-specific recombination, also for counters. Stemmer et al. 1995. Gene 164: 49 -53; Mullis 1986 CSHSQB.
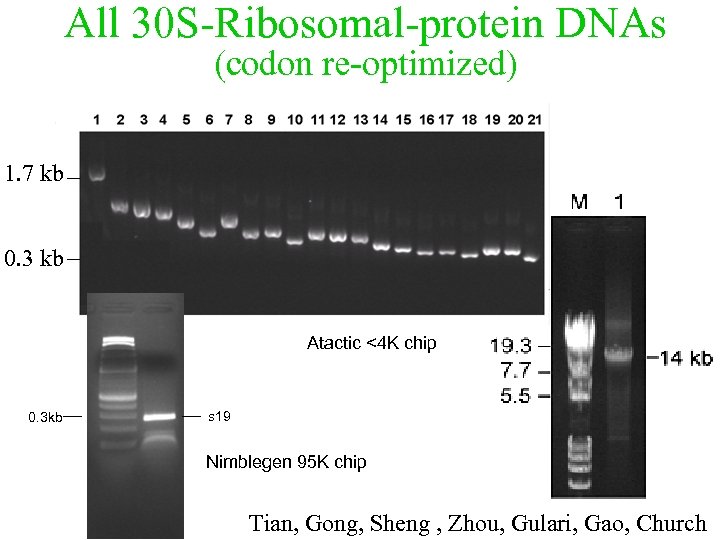
All 30 S-Ribosomal-protein DNAs (codon re-optimized) 1. 7 kb 0. 3 kb Atactic <4 K chip 0. 3 kb s 19 Nimblegen 95 K chip Tian, Gong, Sheng , Zhou, Gulari, Gao, Church
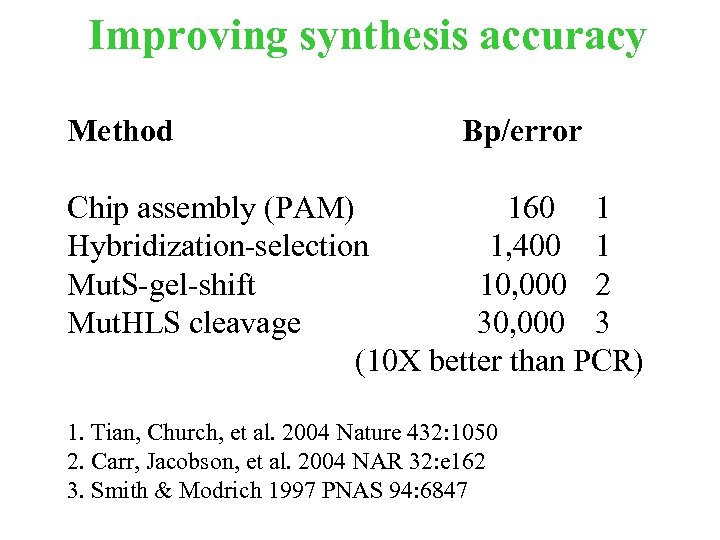

Improving synthesis accuracy Method Bp/error Chip assembly (PAM) 160 1 Hybridization-selection 1, 400 1 Mut. S-gel-shift 10, 000 2 Mut. HLS cleavage 30, 000 3 (10 X better than PCR) 1. Tian, Church, et al. 2004 Nature 432: 1050 2. Carr, Jacobson, et al. 2004 NAR 32: e 162 3. Smith & Modrich 1997 PNAS 94: 6847
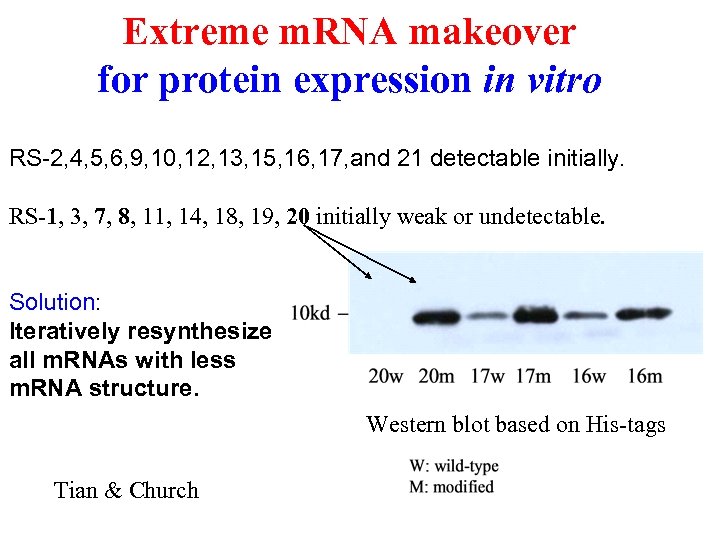

Extreme m. RNA makeover for protein expression in vitro RS-2, 4, 5, 6, 9, 10, 12, 13, 15, 16, 17, and 21 detectable initially. RS-1, 3, 7, 8, 11, 14, 18, 19, 20 initially weak or undetectable. Solution: Iteratively resynthesize all m. RNAs with less m. RNA structure. Western blot based on His-tags Tian & Church
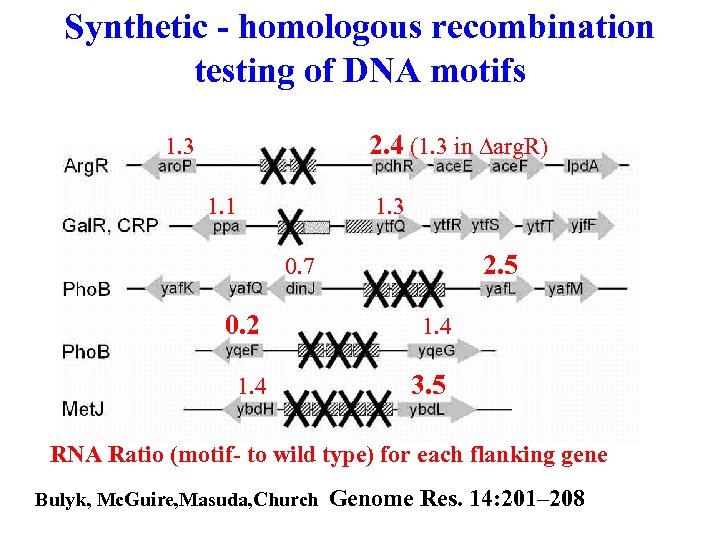

Synthetic - homologous recombination testing of DNA motifs 2. 4 (1. 3 in Darg. R) 1. 3 1. 1 1. 3 2. 5 0. 7 0. 2 1. 4 3. 5 RNA Ratio (motif- to wild type) for each flanking gene Bulyk, Mc. Guire, Masuda, Church Genome Res. 14: 201– 208

Safe Synthetic Biology Church, G. M. (2004) A synthetic biohazard non-proliferation proposal. http: //arep. med. harvard. edu/SBP/Church_Biohazard 04 c. doc 1. Monitor oligo synthesis via expansion of Controlled substances, Select Agents, &/or Recombinant DNA 2. Computational tools are available; very small number of reagent, instrument & synthetic gene suppliers at present. 3. System modeling checks for synthetic biology projects 4. Multi-auxotroph, novel genetic code for the host genome, prevents functional transfer of DNA to other cells.

Public relations & safety Church, G. M. A synthetic biohazard non-proliferation proposal (2004) http: //arep. med. harvard. edu/SBP/Church_Biohazard 04 c. doc • Monitor oligo synthesis via expanding the purview of Controlled substances, Select Agents, &/or Recombinant DNA. • Computational tools for the above (e. g. Craic) • System modeling for all Synthetic Biology Projects • Avoid environmental release uses (at least initially) Beckwith'69, Asilomar'75, AGS-Rifkin'84 -6, Starlink'00, Roundup'04… http: //www. americanscientist. org/template/Book. Review. Type. Detail/assetid/16207 http: //www. social-ecology. org/article. php? story=2003120211014237 Jackson et al. (2001) J Virol. 75: 1205 -10. "immunized genetically resistant mice withthevirusexpressing. IL-4 resulted in significant mortality due to fulminant mousepox. "

Safety via blocking exchange Can we make a cell which is resistant to all viruses and incapable of *functional* DNA exchange in or out? One option is genetic code remapping. Micrococcus luteus is naturally missing 6 codons: UUA(L), CUA(L), AUA(I), GUA(Q), CAA(Q), AGA(R). Kowal, AK, & Oliver, JS NAR 1997, 25: 4685
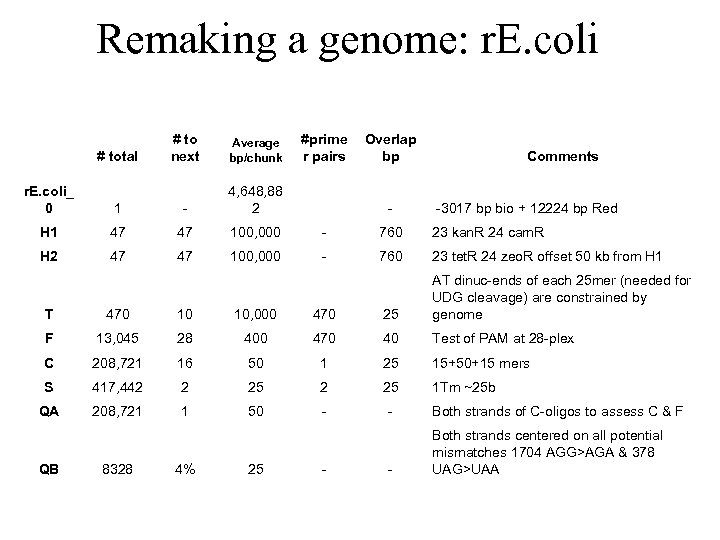
Remaking a genome: r. E. coli # total # to next Average bp/chunk #prime r pairs Overlap bp Comments r. E. coli_ 0 1 - 4, 648, 88 2 H 1 47 47 100, 000 - 760 23 kan. R 24 cam. R H 2 47 47 100, 000 - 760 23 tet. R 24 zeo. R offset 50 kb from H 1 - -3017 bp bio + 12224 bp Red T 470 10 10, 000 470 25 AT dinuc-ends of each 25 mer (needed for UDG cleavage) are constrained by genome F 13, 045 28 400 470 40 Test of PAM at 28 -plex C 208, 721 16 50 1 25 15+50+15 mers S 417, 442 2 25 1 Tm ~25 b QA 208, 721 1 50 - - Both strands of C-oligos to assess C & F - Both strands centered on all potential mismatches 1704 AGG>AGA & 378 UAG>UAA QB 8328 4% 25 -
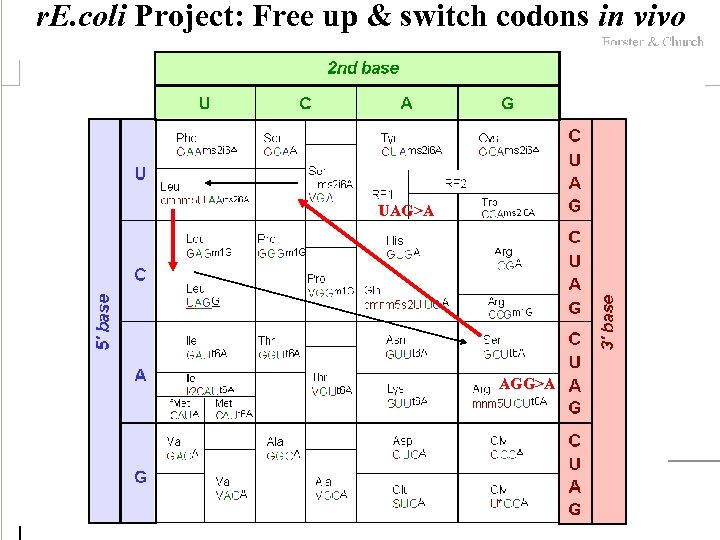
r. E. coli Project: Free up & switch codons in vivo UAG>A AGG>A
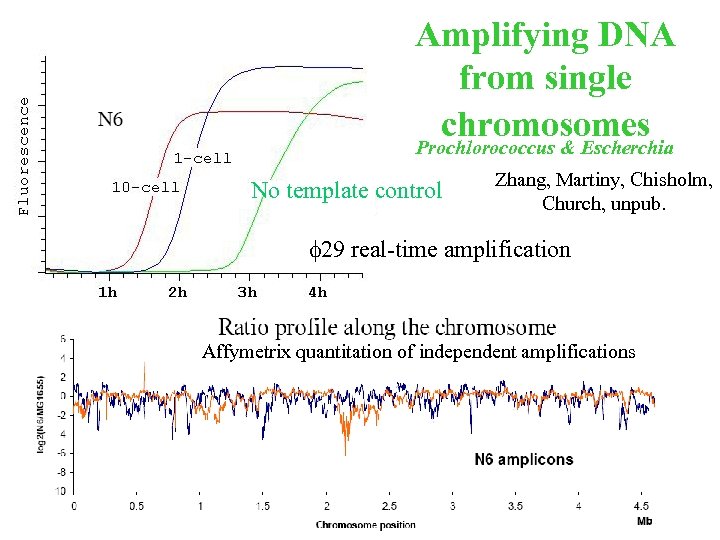
Amplifying DNA from single chromosomes Prochlorococcus & Escherchia No template control Zhang, Martiny, Chisholm, Church, unpub. f 29 real-time amplification Affymetrix quantitation of independent amplifications
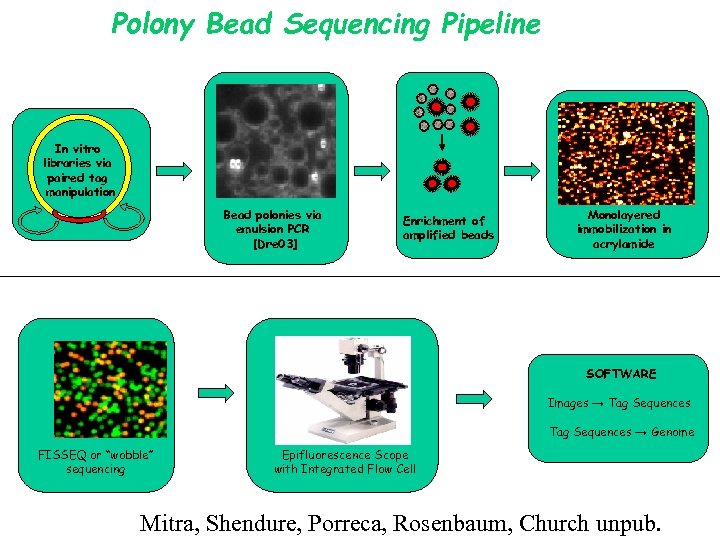
Polony Bead Sequencing Pipeline In vitro libraries via paired tag manipulation Bead polonies via emulsion PCR [Dre 03] Enrichment of amplified beads Monolayered immobilization in acrylamide SOFTWARE Images → Tag Sequences → Genome FISSEQ or “wobble” sequencing Epifluorescence Scope with Integrated Flow Cell Mitra, Shendure, Porreca, Rosenbaum, Church unpub.
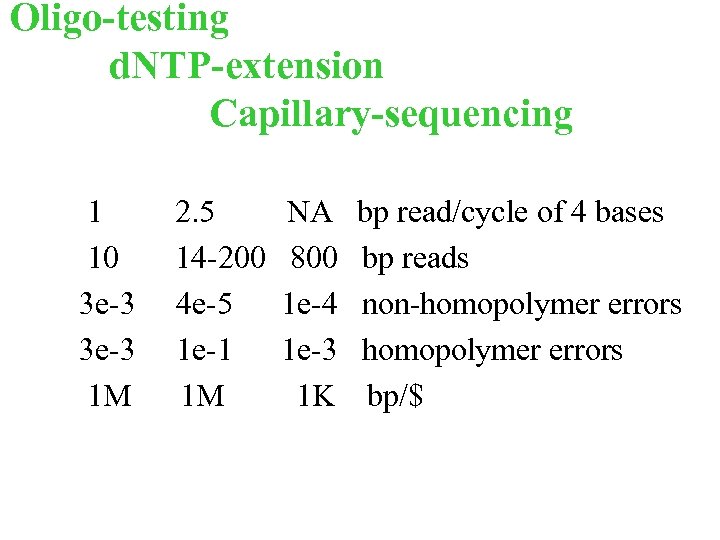

Oligo-testing d. NTP-extension Capillary-sequencing 1 10 3 e-3 1 M 2. 5 14 -200 4 e-5 1 e-1 1 M NA 800 1 e-4 1 e-3 1 K bp read/cycle of 4 bases bp reads non-homopolymer errors bp/$

Integrating with appropriate sequencing strategies Shendure J, Mitra R, Varma C, Church GM (May 2004) Advanced Sequencing Technologies: Methods & Goals. Nature Reviews of Genetics 5, 335 -344. NHGRI Seeks Next Generation of Sequencing Technologies (Jan 2004) http: //www. genome. gov/12513210

Automated homologous recombination • Positive & Negative Selection in same gene: URA 3 (yeast), Thy. A(E. coli), GFP(various) • Electroporation, viral, conjugative delivery 3 ori. T regions: Inc. Pa, F, and R 64(Inc. I) Valenzuela DM, et al. Nat Biotechnol. 2003 Jun; 21(6): 652 -9. High-throughput engineering of the mouse genome coupled with high-resolution expression analysis. up to 25% targeting with BACs. Yang Y, Seed B. Site-specific gene targeting in mouse embryonic stem cells with intact bacterial artificial chromosomes. Nat Biotechnol. 2003 21: 447 -51. Schneckenburger H, et al. J Biomed Opt. 2002 Jul; 7(3): 410 -6. Laser-assisted optoporation of single cells.

Integrase applications (1) In vivo recombination (increase fidelity & efficiency) Nucleofection of muscle-derived stem cells and myoblasts with phi. C 31 integrase. Mol Ther. 2004 10: 679 -87. (2) In vitro plasmid construction (Gateway) (3) In vivo counters allow recording & increased analog I/O through digital reuse of functions. For a 3 -bit (8 state counter) 0 0 0 lac-GFP 0 0 1 ara-GFP 0 1 0 trp-GFP 0 1 1 tet-GFP 1 0 0 etc.
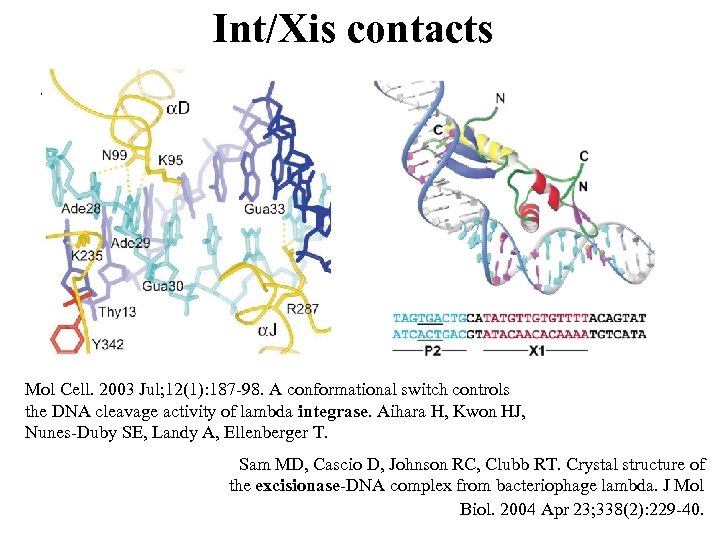
Int/Xis contacts Mol Cell. 2003 Jul; 12(1): 187 -98. A conformational switch controls the DNA cleavage activity of lambda integrase. Aihara H, Kwon HJ, Nunes-Duby SE, Landy A, Ellenberger T. Sam MD, Cascio D, Johnson RC, Clubb RT. Crystal structure of the excisionase-DNA complex from bacteriophage lambda. J Mol Biol. 2004 Apr 23; 338(2): 229 -40.
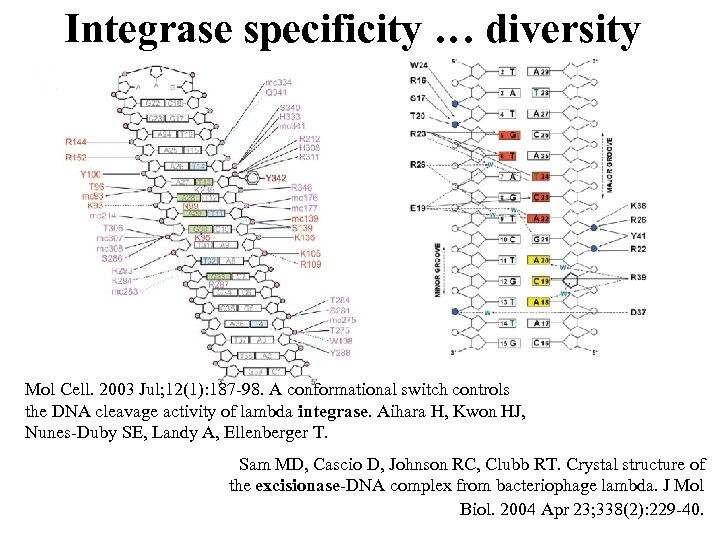

Integrase specificity … diversity Mol Cell. 2003 Jul; 12(1): 187 -98. A conformational switch controls the DNA cleavage activity of lambda integrase. Aihara H, Kwon HJ, Nunes-Duby SE, Landy A, Ellenberger T. Sam MD, Cascio D, Johnson RC, Clubb RT. Crystal structure of the excisionase-DNA complex from bacteriophage lambda. J Mol Biol. 2004 Apr 23; 338(2): 229 -40.
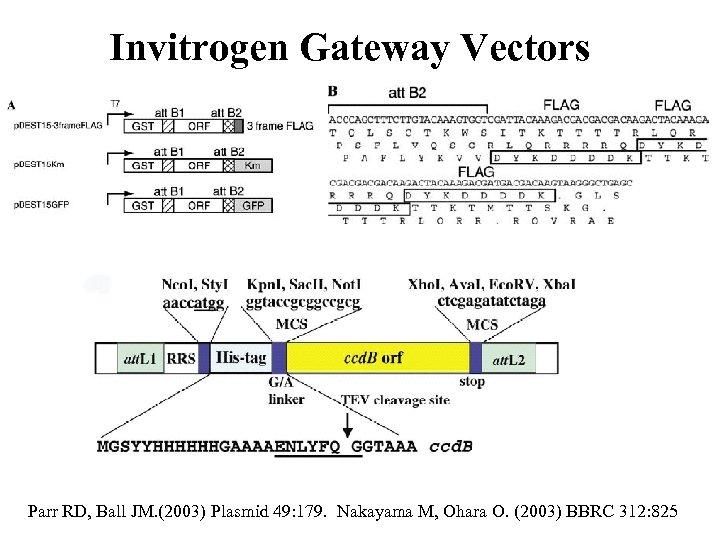
Invitrogen Gateway Vectors Parr RD, Ball JM. (2003) Plasmid 49: 179. Nakayama M, Ohara O. (2003) BBRC 312: 825

Potential Commercial Biology Partners / Competitors Invitrogen Gateway cloning Poetic Genetics Integrases & Gene Therapy Regeneron Mammalian BAC recombination 1% Scarab Genomics Better E. coli strains 20% of genome Avidia/Diversa Shuffling/selection Ensemble DNA catalysts Amyris Terpenoid pathways Kosan Biosciences Polyketide pathways Big & Small Pharma

ibm. com/chips/services/foundry/partners Analog Bits, Artisan Components, Cadence, e. Silicon Corporation, GDA Technologies Inc. , insyte, Jennic Limited, Kisel Microelectronics, Magma, MOSIS, QThink, Qual. Core Logic, Inc. , RF Integration, Sierra Monolithics, SOCLE Technology, Synopsys, Tahoe RF Semiconductors, Tel. ASIC, Tri. CN, Triscend, Virtual Silicon, Mosis. org "50, 000 designs… keep prototype costs low by aggregating many designs onto one mask set, sharing overhead" Fabrication Processes: AMIS, IBM, Austriamicrosystems, OMMIC/PML, Peregrine, TSMC, Vitesse Example 2: linux. org redhat. com Q 2 $46 M up 60% You are here

Low-cost, high accuracy, long-DNA synthesis technology 16 -Feb-2005 10 AM NHGRI George Church, Joe Jacobsen et al. Harvard & MIT 0. Killer Applications 1. Chip synthesis, fluidics 2. Multiplex assembly 3. Error correction methods 4. Software CAD-PAM 5. Proteome (in vitro) synthesis 6. Homologous recombination & selection for BACs 7. Integrases 8. Process integration, QA, timeline 9. Safety opportunities
a3316dc9e94c9c18cf8a13f6ba225af6.ppt