dcd4cabe2cde8d02744bbfb5a0c5562c.ppt
- Количество слайдов: 31
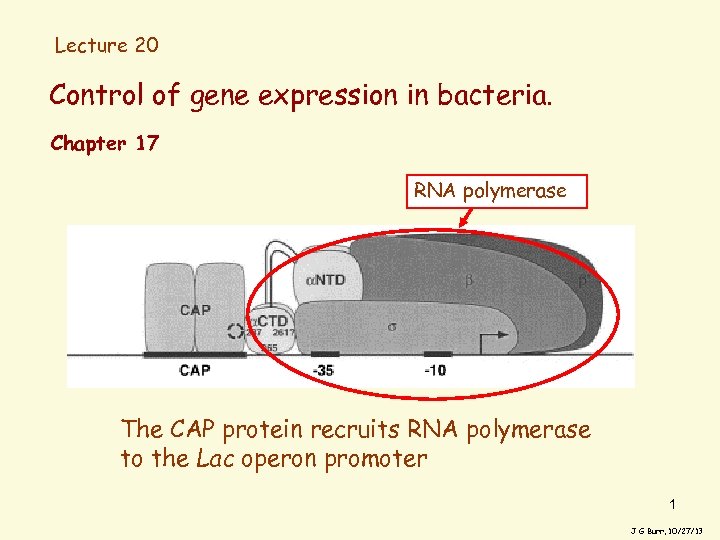
Lecture 20 Control of gene expression in bacteria. Chapter 17 RNA polymerase The CAP protein recruits RNA polymerase to the Lac operon promoter 1 J G Burr, 10/27/13
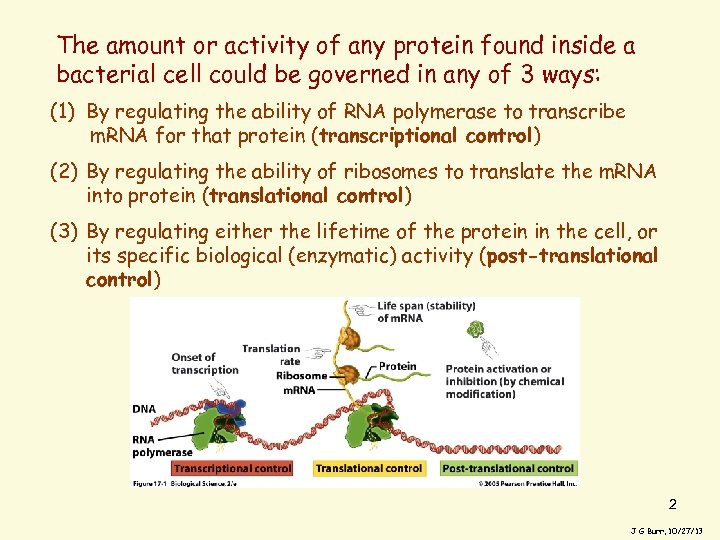
The amount or activity of any protein found inside a bacterial cell could be governed in any of 3 ways: (1) By regulating the ability of RNA polymerase to transcribe m. RNA for that protein (transcriptional control) (2) By regulating the ability of ribosomes to translate the m. RNA into protein (translational control) (3) By regulating either the lifetime of the protein in the cell, or its specific biological (enzymatic) activity (post-translational control) 2 J G Burr, 10/27/13
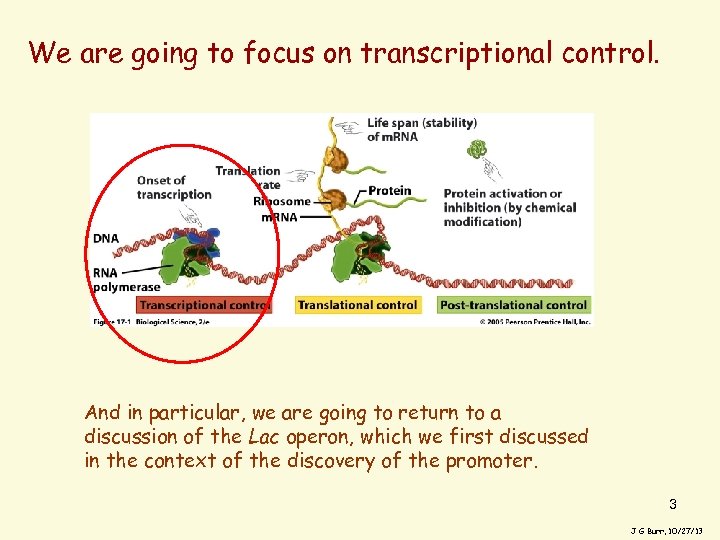
We are going to focus on transcriptional control. And in particular, we are going to return to a discussion of the Lac operon, which we first discussed in the context of the discovery of the promoter. 3 J G Burr, 10/27/13
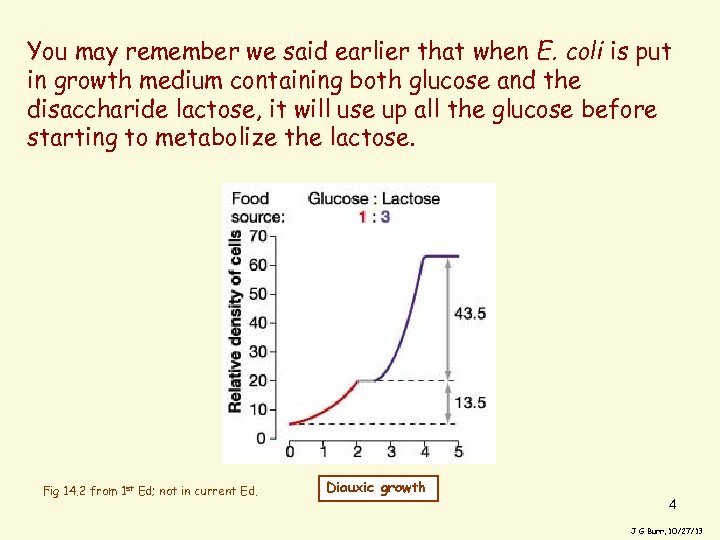
You may remember we said earlier that when E. coli is put in growth medium containing both glucose and the disaccharide lactose, it will use up all the glucose before starting to metabolize the lactose. Fig 14. 2 from 1 st Ed; not in current Ed. Diauxic growth 4 J G Burr, 10/27/13
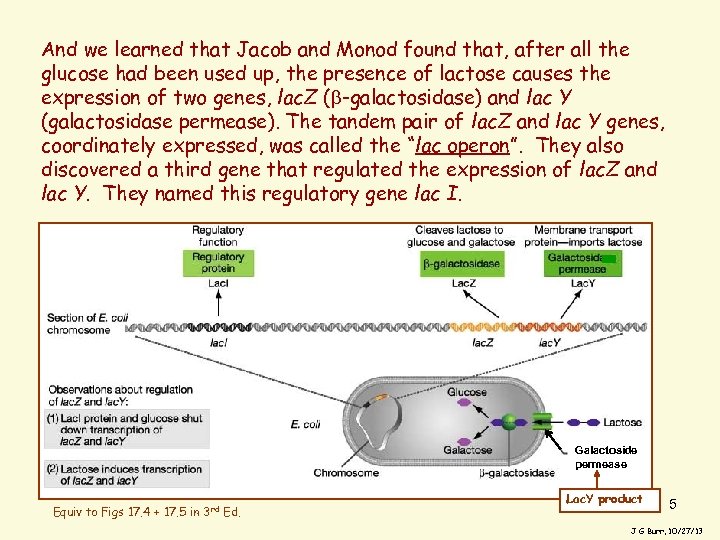
And we learned that Jacob and Monod found that, after all the glucose had been used up, the presence of lactose causes the expression of two genes, lac. Z ( -galactosidase) and lac Y (galactosidase permease). The tandem pair of lac. Z and lac Y genes, coordinately expressed, was called the “lac operon”. They also discovered a third gene that regulated the expression of lac. Z and lac Y. They named this regulatory gene lac I. Galactoside permease Equiv to Figs 17. 4 + 17. 5 in 3 rd Ed. Lac. Y product 5 J G Burr, 10/27/13
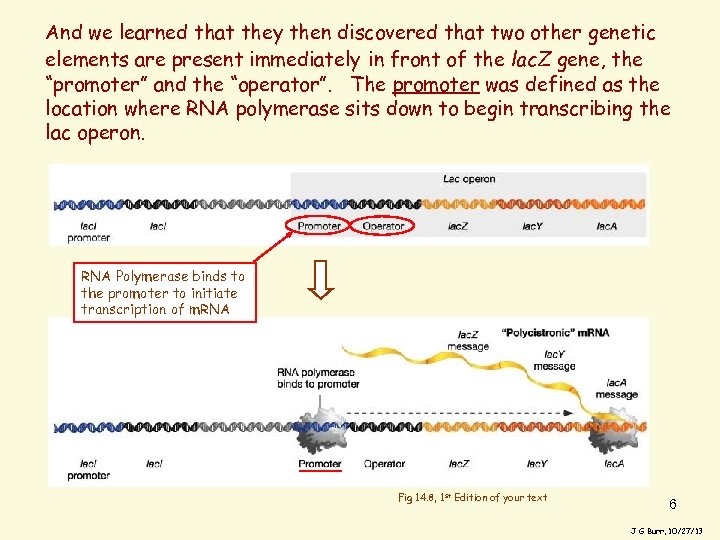
And we learned that they then discovered that two other genetic elements are present immediately in front of the lac. Z gene, the “promoter” and the “operator”. The promoter was defined as the location where RNA polymerase sits down to begin transcribing the lac operon. RNA Polymerase binds to the promoter to initiate transcription of m. RNA Fig 14. 8, 1 st Edition of your text 6 J G Burr, 10/27/13
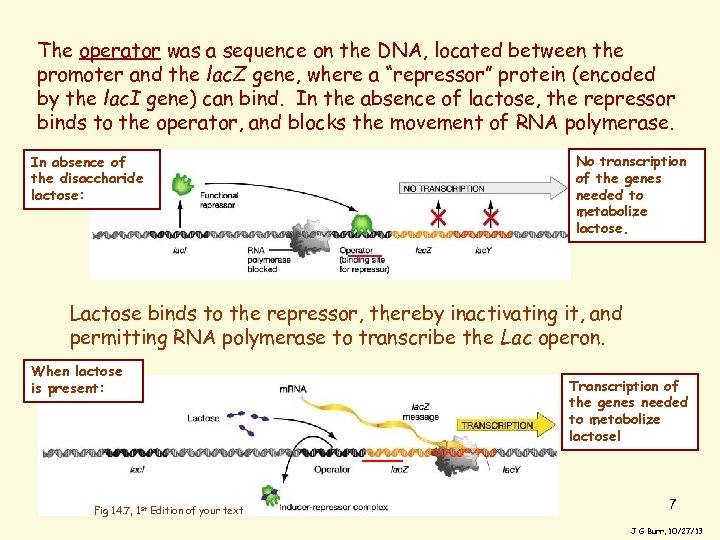
The operator was a sequence on the DNA, located between the promoter and the lac. Z gene, where a “repressor” protein (encoded by the lac. I gene) can bind. In the absence of lactose, the repressor binds to the operator, and blocks the movement of RNA polymerase. In absence of the disaccharide lactose: No transcription of the genes needed to metabolize lactose. Lactose binds to the repressor, thereby inactivating it, and permitting RNA polymerase to transcribe the Lac operon. When lactose is present: Fig 14. 7, 1 st Edition of your text Transcription of the genes needed to metabolize lactose! 7 J G Burr, 10/27/13
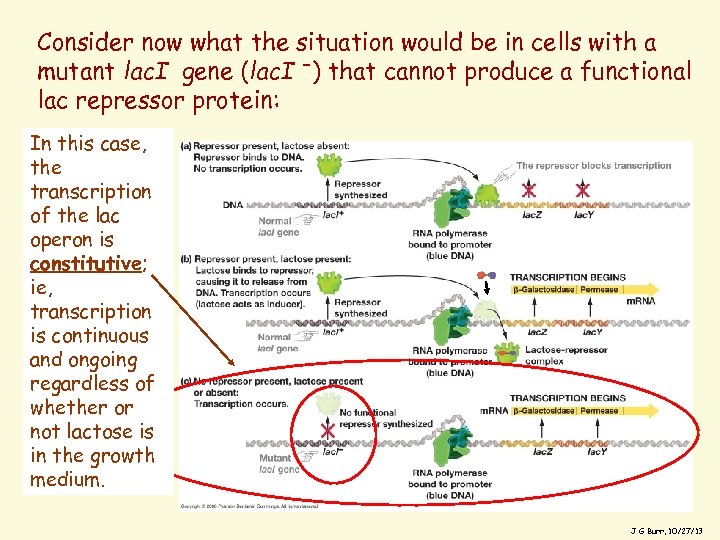
Consider now what the situation would be in cells with a mutant lac. I gene (lac. I -) that cannot produce a functional lac repressor protein: In this case, the transcription of the lac operon is constitutive; ie, transcription is continuous and ongoing regardless of whether or not lactose is in the growth medium. 8 J G Burr, 10/27/13
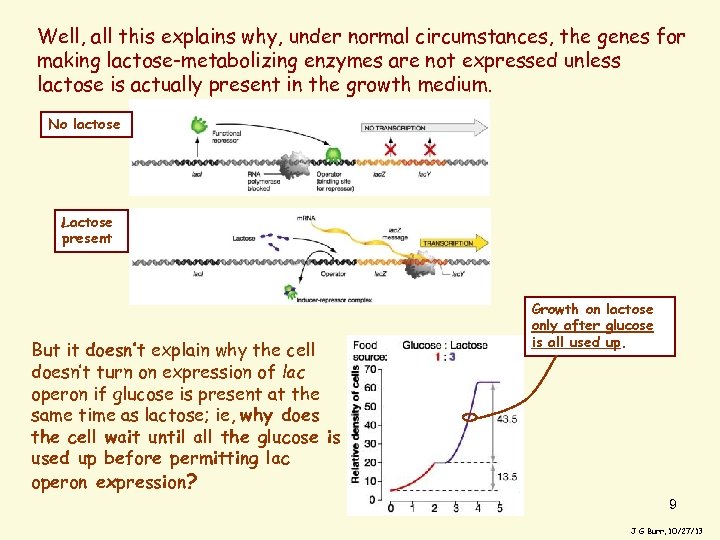
Well, all this explains why, under normal circumstances, the genes for making lactose-metabolizing enzymes are not expressed unless lactose is actually present in the growth medium. No lactose Lactose present But it doesn’t explain why the cell doesn’t turn on expression of lac operon if glucose is present at the same time as lactose; ie, why does the cell wait until all the glucose is used up before permitting lac operon expression? Growth on lactose only after glucose is all used up. 9 J G Burr, 10/27/13
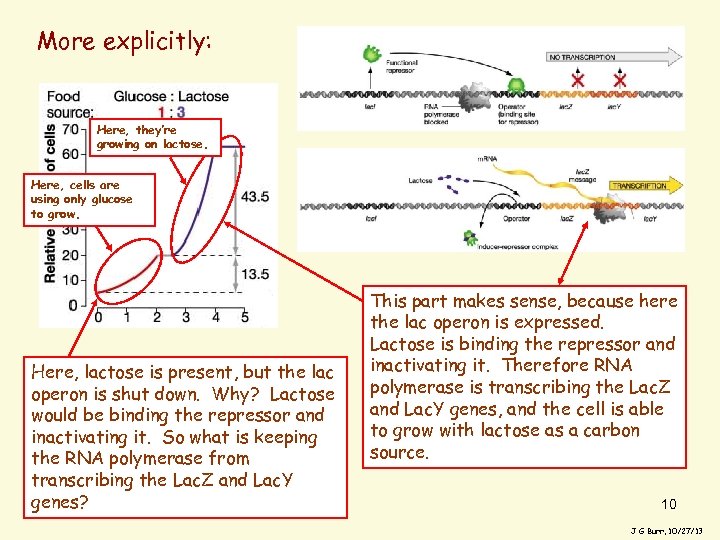
More explicitly: Here, they’re growing on lactose. Here, cells are using only glucose to grow. Here, lactose is present, but the lac operon is shut down. Why? Lactose would be binding the repressor and inactivating it. So what is keeping the RNA polymerase from transcribing the Lac. Z and Lac. Y genes? This part makes sense, because here the lac operon is expressed. Lactose is binding the repressor and inactivating it. Therefore RNA polymerase is transcribing the Lac. Z and Lac. Y genes, and the cell is able to grow with lactose as a carbon source. 10 J G Burr, 10/27/13
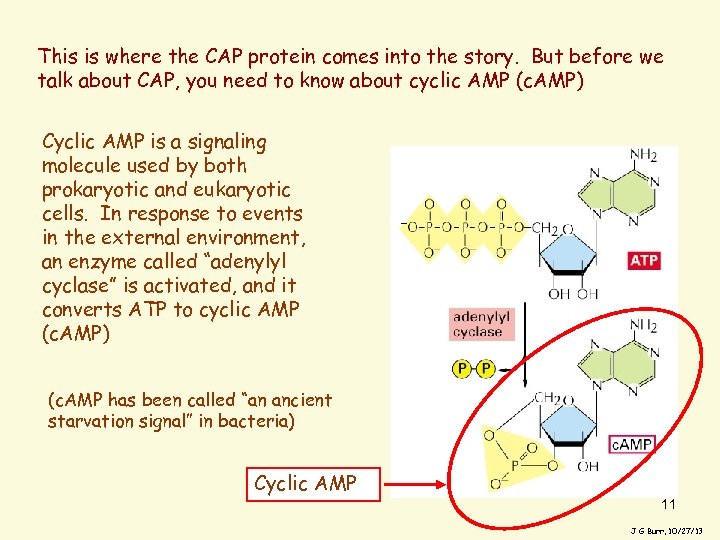
This is where the CAP protein comes into the story. But before we talk about CAP, you need to know about cyclic AMP (c. AMP) Cyclic AMP is a signaling molecule used by both prokaryotic and eukaryotic cells. In response to events in the external environment, an enzyme called “adenylyl cyclase” is activated, and it converts ATP to cyclic AMP (c. AMP) (c. AMP has been called “an ancient starvation signal” in bacteria) Cyclic AMP 11 J G Burr, 10/27/13
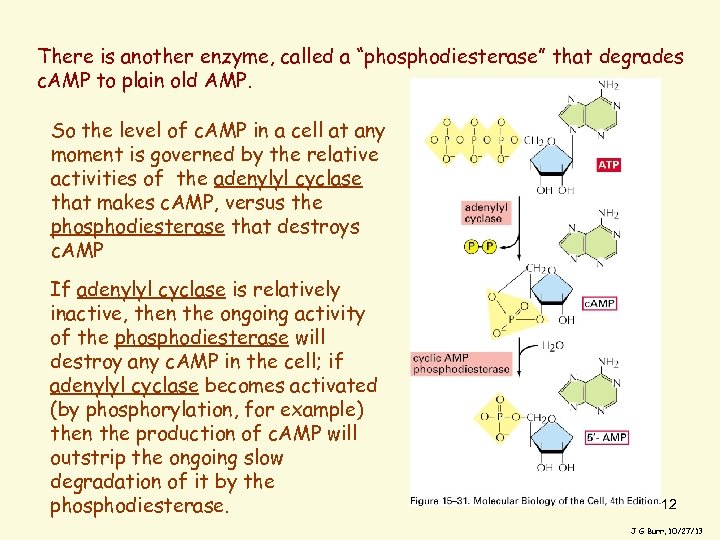
There is another enzyme, called a “phosphodiesterase” that degrades c. AMP to plain old AMP. So the level of c. AMP in a cell at any moment is governed by the relative activities of the adenylyl cyclase that makes c. AMP, versus the phosphodiesterase that destroys c. AMP If adenylyl cyclase is relatively inactive, then the ongoing activity of the phosphodiesterase will destroy any c. AMP in the cell; if adenylyl cyclase becomes activated (by phosphorylation, for example) then the production of c. AMP will outstrip the ongoing slow degradation of it by the phosphodiesterase. 12 J G Burr, 10/27/13
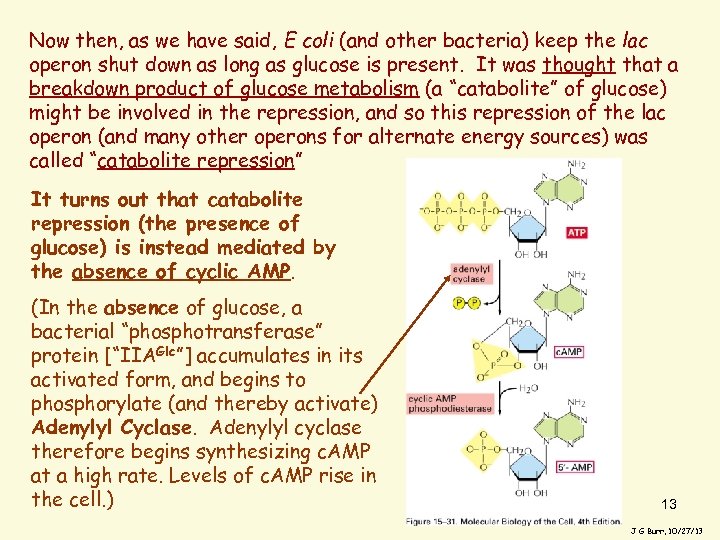
Now then, as we have said, E coli (and other bacteria) keep the lac operon shut down as long as glucose is present. It was thought that a breakdown product of glucose metabolism (a “catabolite” of glucose) might be involved in the repression, and so this repression of the lac operon (and many other operons for alternate energy sources) was called “catabolite repression” It turns out that catabolite repression (the presence of glucose) is instead mediated by the absence of cyclic AMP. (In the absence of glucose, a bacterial “phosphotransferase” protein [“IIAGlc”] accumulates in its activated form, and begins to phosphorylate (and thereby activate) Adenylyl Cyclase. Adenylyl cyclase therefore begins synthesizing c. AMP at a high rate. Levels of c. AMP rise in the cell. ) 13 J G Burr, 10/27/13
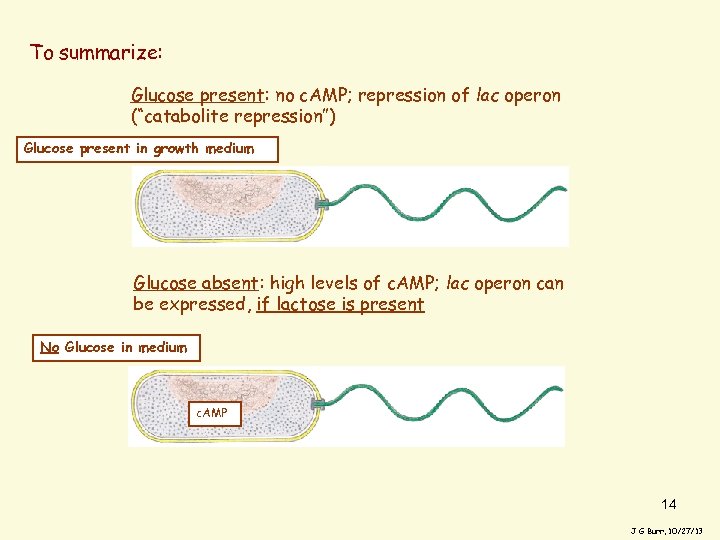
To summarize: Glucose present: no c. AMP; repression of lac operon (“catabolite repression”) Glucose present in growth medium Glucose absent: high levels of c. AMP; lac operon can be expressed, if lactose is present No Glucose in medium c. AMP 14 J G Burr, 10/27/13
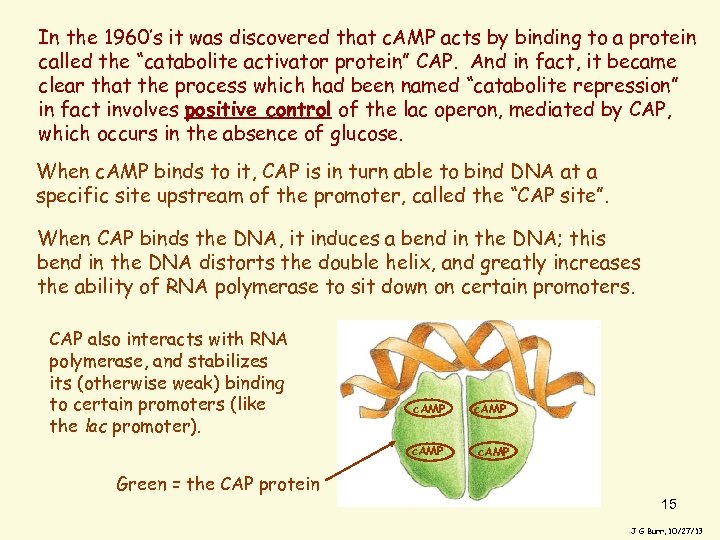
In the 1960’s it was discovered that c. AMP acts by binding to a protein called the “catabolite activator protein” CAP. And in fact, it became clear that the process which had been named “catabolite repression” in fact involves positive control of the lac operon, mediated by CAP, which occurs in the absence of glucose. When c. AMP binds to it, CAP is in turn able to bind DNA at a specific site upstream of the promoter, called the “CAP site”. When CAP binds the DNA, it induces a bend in the DNA; this bend in the DNA distorts the double helix, and greatly increases the ability of RNA polymerase to sit down on certain promoters. CAP also interacts with RNA polymerase, and stabilizes its (otherwise weak) binding to certain promoters (like the lac promoter). c. AMP Green = the CAP protein 15 J G Burr, 10/27/13

Sidebar: There is plenty of CAP protein around in the cell at all times, but if the cell is not starving for glucose, all this CAP protein is inactive (ie, unable to bind to DNA); as we have been saying, it is only when there is no glucose available that the cell makes c. AMP, and this c. AMP then binds to pre-existing CAP, thereby activating CAP. So CAP protein activity is not regulated at the transcriptional or translational level; ie, the cell is constantly making plenty of m. RNA for CAP, and translating CAP m. RNA into CAP protein. Rather, this is an example of a protein whose activity (we would say) is “regulated at the post-translational level. ” c. AMP Green = the CAP protein 16 J G Burr, 10/27/13
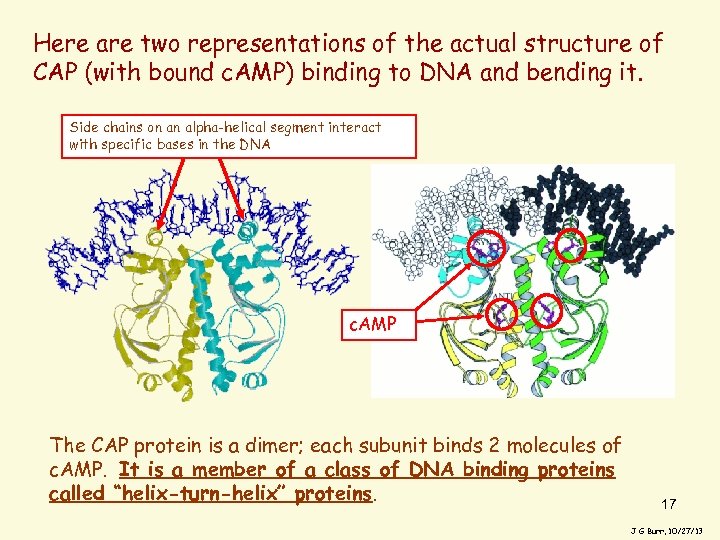
Here are two representations of the actual structure of CAP (with bound c. AMP) binding to DNA and bending it. Side chains on an alpha-helical segment interact with specific bases in the DNA c. AMP The CAP protein is a dimer; each subunit binds 2 molecules of c. AMP. It is a member of a class of DNA binding proteins called “helix-turn-helix” proteins. 17 J G Burr, 10/27/13
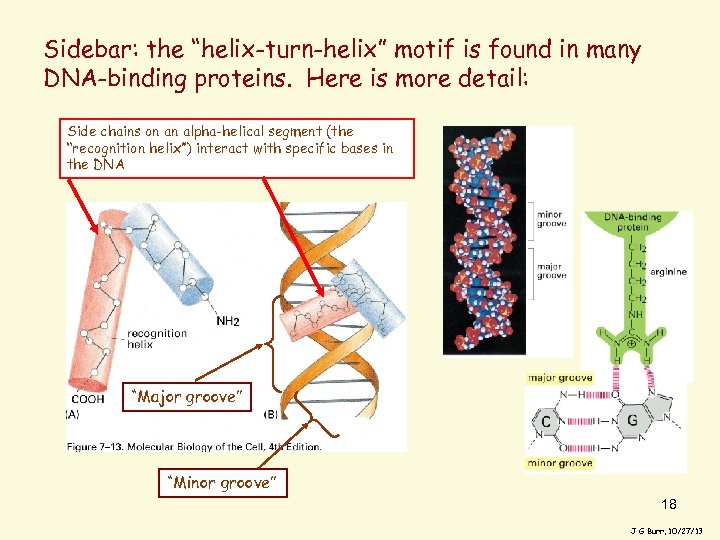
Sidebar: the “helix-turn-helix” motif is found in many DNA-binding proteins. Here is more detail: Side chains on an alpha-helical segment (the “recognition helix”) interact with specific bases in the DNA “Major groove” “Minor groove” 18 J G Burr, 10/27/13
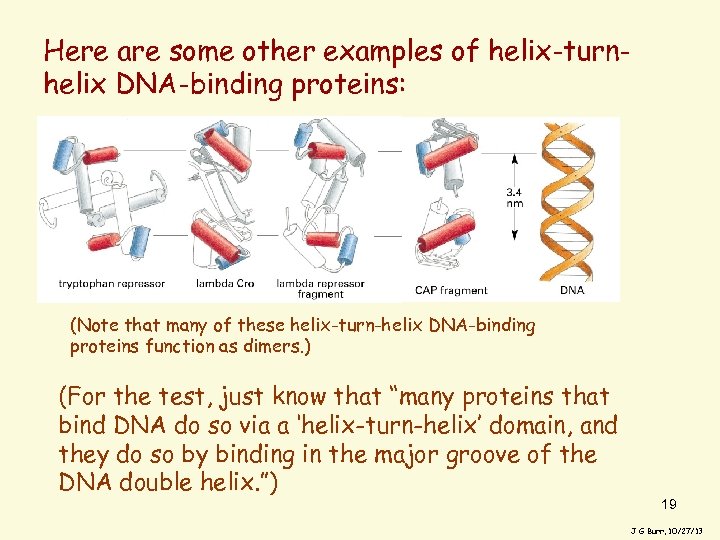
Here are some other examples of helix-turnhelix DNA-binding proteins: (Note that many of these helix-turn-helix DNA-binding proteins function as dimers. ) (For the test, just know that “many proteins that bind DNA do so via a ‘helix-turn-helix’ domain, and they do so by binding in the major groove of the DNA double helix. ”) 19 J G Burr, 10/27/13
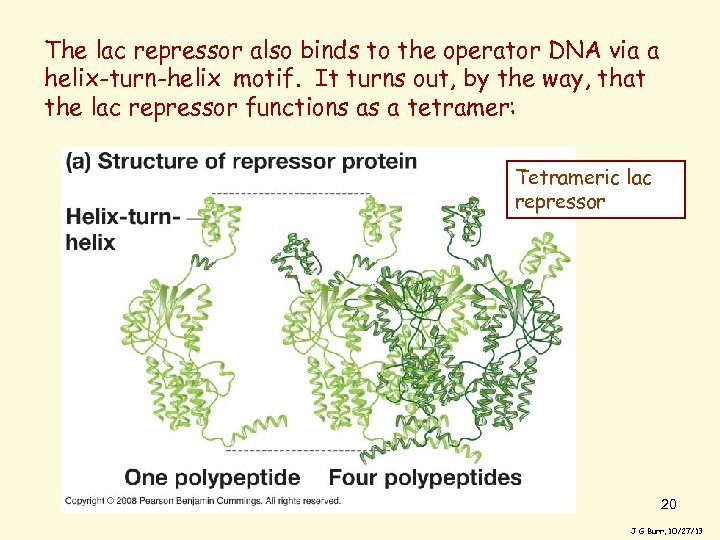
The lac repressor also binds to the operator DNA via a helix-turn-helix motif. It turns out, by the way, that the lac repressor functions as a tetramer: Tetrameric lac repressor 20 J G Burr, 10/27/13
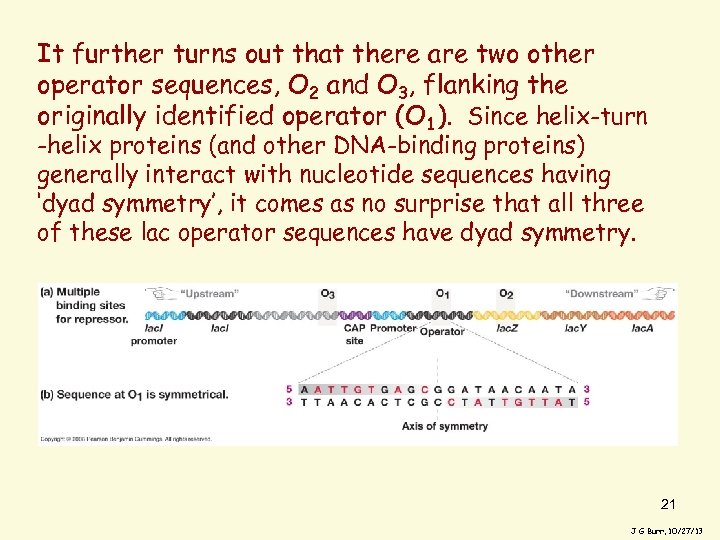

It further turns out that there are two other operator sequences, O 2 and O 3, flanking the originally identified operator (O 1). Since helix-turn -helix proteins (and other DNA-binding proteins) generally interact with nucleotide sequences having ‘dyad symmetry’, it comes as no surprise that all three of these lac operator sequences have dyad symmetry. 21 J G Burr, 10/27/13
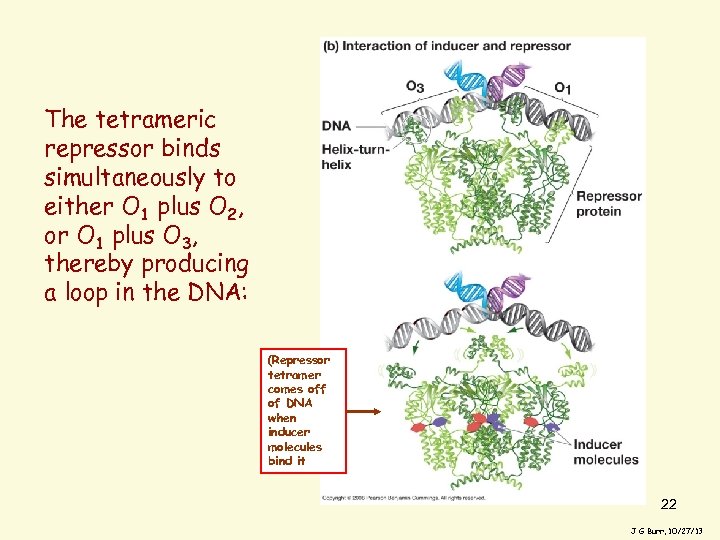
The tetrameric repressor binds simultaneously to either O 1 plus O 2, or O 1 plus O 3, thereby producing a loop in the DNA: (Repressor tetramer comes off of DNA when inducer molecules bind it 22 J G Burr, 10/27/13
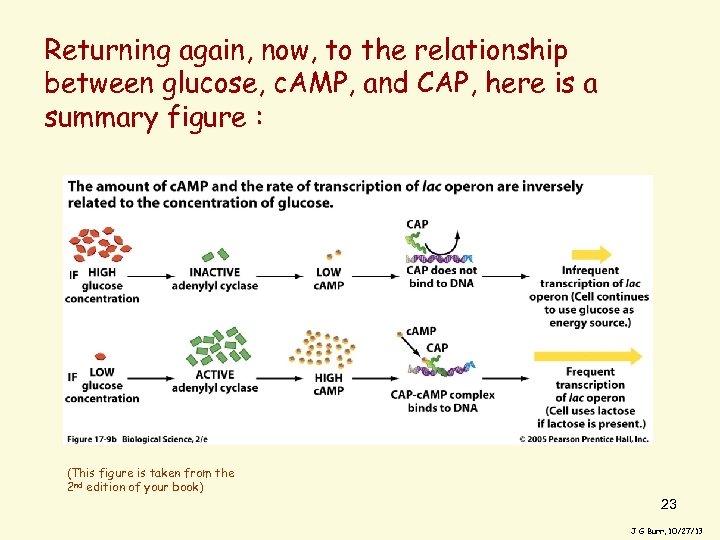
Returning again, now, to the relationship between glucose, c. AMP, and CAP, here is a summary figure : (This figure is taken from the 2 nd edition of your book) 23 J G Burr, 10/27/13
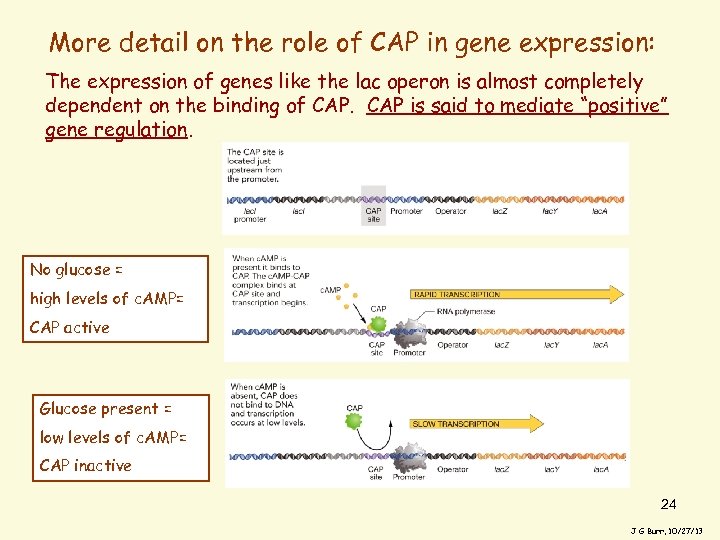
More detail on the role of CAP in gene expression: The expression of genes like the lac operon is almost completely dependent on the binding of CAP is said to mediate “positive” gene regulation. No glucose = high levels of c. AMP= CAP active Glucose present = low levels of c. AMP= CAP inactive 24 J G Burr, 10/27/13
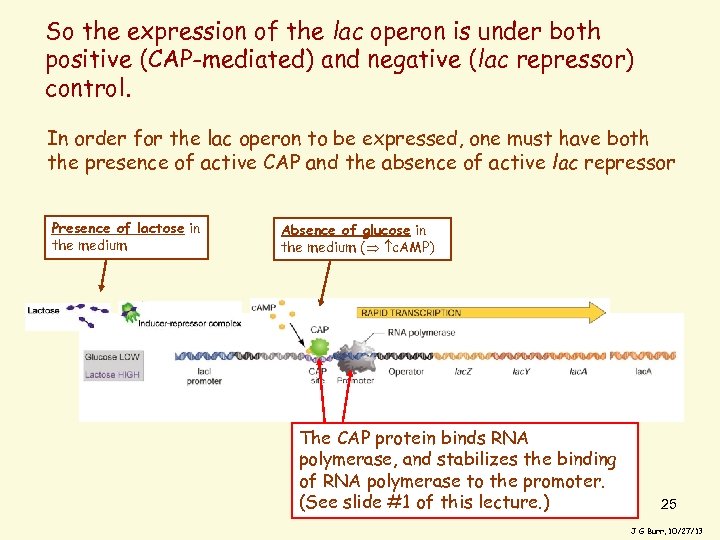
So the expression of the lac operon is under both positive (CAP-mediated) and negative (lac repressor) control. In order for the lac operon to be expressed, one must have both the presence of active CAP and the absence of active lac repressor Presence of lactose in the medium Absence of glucose in the medium ( c. AMP) The CAP protein binds RNA polymerase, and stabilizes the binding of RNA polymerase to the promoter. (See slide #1 of this lecture. ) 25 J G Burr, 10/27/13
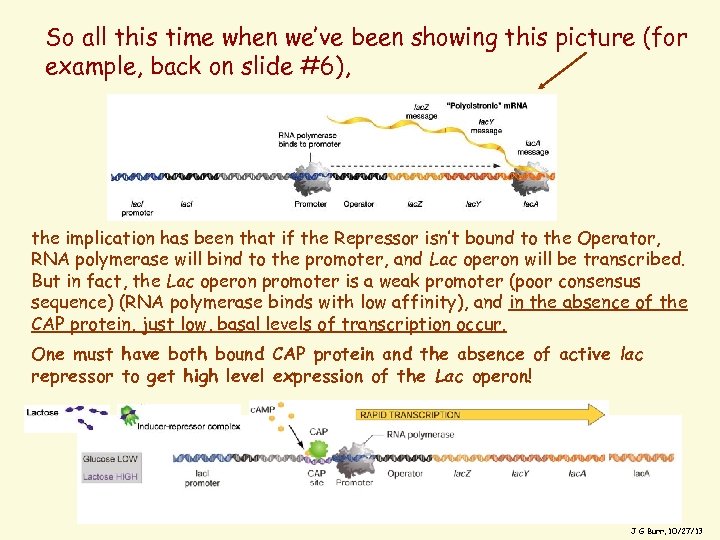
So all this time when we’ve been showing this picture (for example, back on slide #6), the implication has been that if the Repressor isn’t bound to the Operator, RNA polymerase will bind to the promoter, and Lac operon will be transcribed. But in fact, the Lac operon promoter is a weak promoter (poor consensus sequence) (RNA polymerase binds with low affinity), and in the absence of the CAP protein, just low, basal levels of transcription occur. One must have both bound CAP protein and the absence of active lac repressor to get high level expression of the Lac operon! 26 J G Burr, 10/27/13
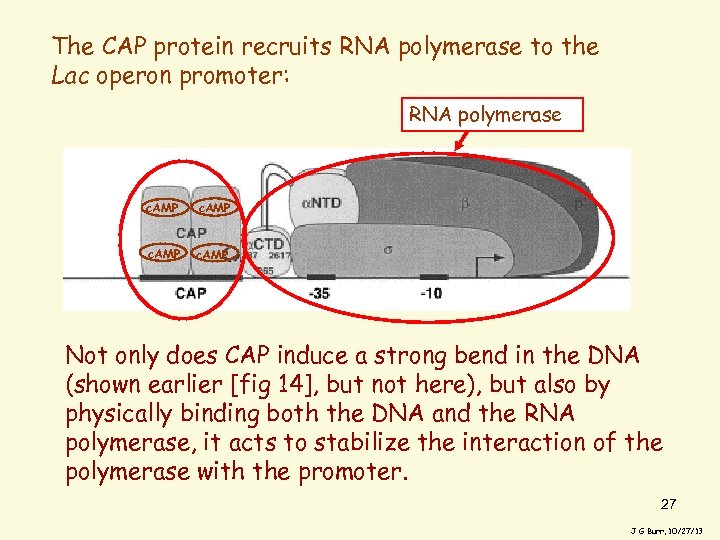
The CAP protein recruits RNA polymerase to the Lac operon promoter: RNA polymerase c. AMP Not only does CAP induce a strong bend in the DNA (shown earlier [fig 14], but not here), but also by physically binding both the DNA and the RNA polymerase, it acts to stabilize the interaction of the polymerase with the promoter. 27 J G Burr, 10/27/13
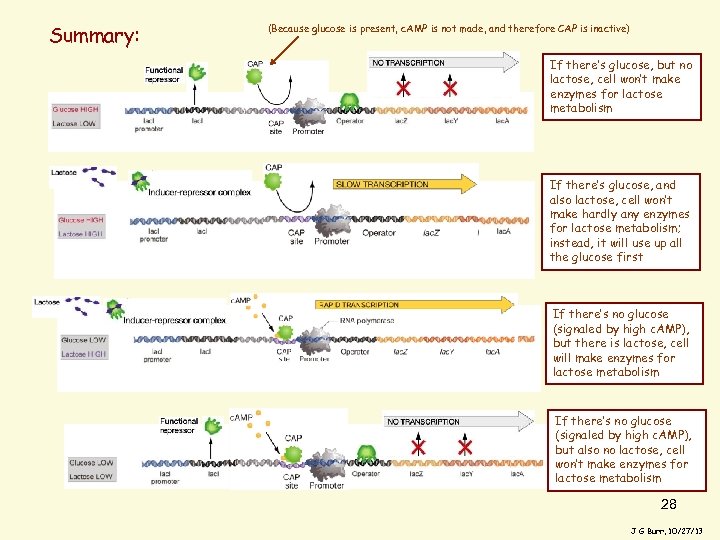
Summary: (Because glucose is present, c. AMP is not made, and therefore CAP is inactive) If there’s glucose, but no lactose, cell won’t make enzymes for lactose metabolism If there’s glucose, and also lactose, cell won’t make hardly any enzymes for lactose metabolism; instead, it will use up all the glucose first If there’s no glucose (signaled by high c. AMP), but there is lactose, cell will make enzymes for lactose metabolism If there’s no glucose (signaled by high c. AMP), but also no lactose, cell won’t make enzymes for lactose metabolism 28 J G Burr, 10/27/13
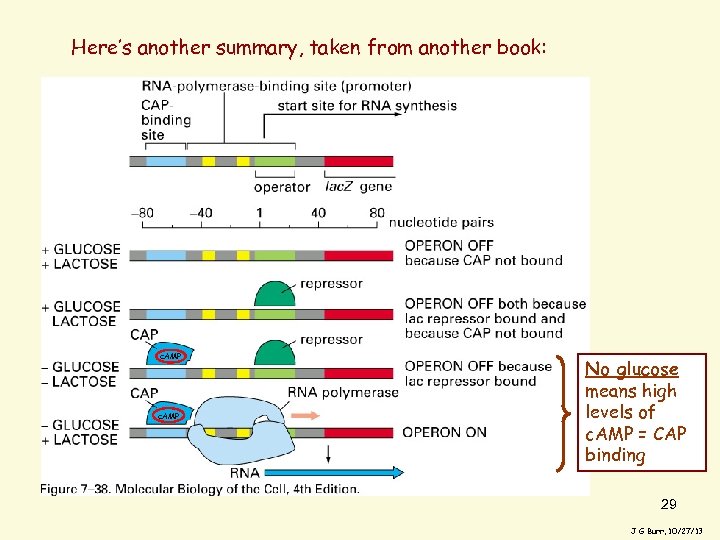
Here’s another summary, taken from another book: c. AMP Alberts, fig 7 -38 No glucose means high levels of c. AMP = CAP binding 29 J G Burr, 10/27/13
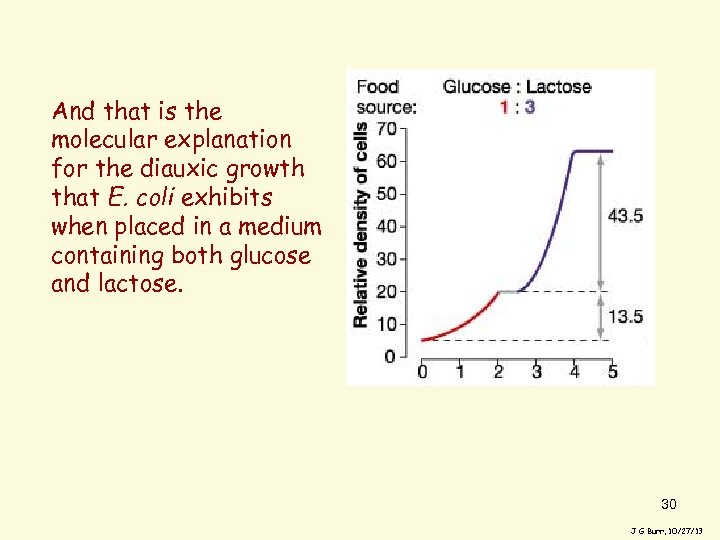
And that is the molecular explanation for the diauxic growth that E. coli exhibits when placed in a medium containing both glucose and lactose. 30 J G Burr, 10/27/13

Jacob, Monod, and André Lwoff won the 1965 Nobel Prize for their contribution to working out this story. The Nobel Prize in Physiology or Medicine 1965 "for their discoveries concerning genetic control of enzyme and virus synthesis" François Jacob (For the test, know the names Jacob and Monod as the “Lac operon guys. ’ Jaques Monod André Lwoff For personal anecdotes about these guys, go to http: //www. dnaftb. org/dnaftb/33/concept/ 31 J G Burr, 10/27/13
dcd4cabe2cde8d02744bbfb5a0c5562c.ppt