6ed7318297b250e5600020cd4067af7a.ppt
- Количество слайдов: 42

László Manczinger, Isidora Radulov, Adina Berbecea, Enikő Sajben-Nagy, Andrea Palágyi, Dorin Tărău, Lucian Dumitru Niţă, Csaba Vágvölgyi Recommendation for working out a new soil ranking system based on the results of the SOILMAP project László Manczinger Department of Microbiology, Faculty of Science and Informatics, University of Szeged, Hungary
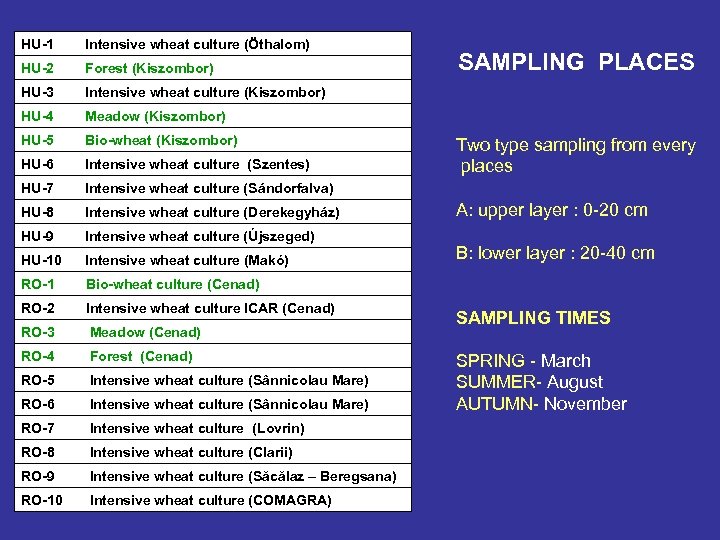

HU-1 Intensive wheat culture (Öthalom) HU-2 Forest (Kiszombor) HU-3 Intensive wheat culture (Kiszombor) HU-4 Meadow (Kiszombor) HU-5 Bio-wheat (Kiszombor) HU-6 Intensive wheat culture (Szentes) HU-7 Intensive wheat culture (Sándorfalva) HU-8 Intensive wheat culture (Derekegyház) HU-9 Intensive wheat culture (Újszeged) HU-10 Intensive wheat culture (Makó) RO-1 Bio-wheat culture (Cenad) RO-2 Intensive wheat culture ICAR (Cenad) RO-3 Meadow (Cenad) RO-4 Forest (Cenad) RO-5 Intensive wheat culture (Sânnicolau Mare) RO-6 Intensive wheat culture (Sânnicolau Mare) RO-7 Intensive wheat culture (Lovrin) RO-8 Intensive wheat culture (Clarii) RO-9 Intensive wheat culture (Săcălaz – Beregsana) RO-10 Intensive wheat culture (COMAGRA) SAMPLING PLACES Two type sampling from every places A: upper layer : 0 -20 cm B: lower layer : 20 -40 cm SAMPLING TIMES SPRING - March SUMMER- August AUTUMN- November
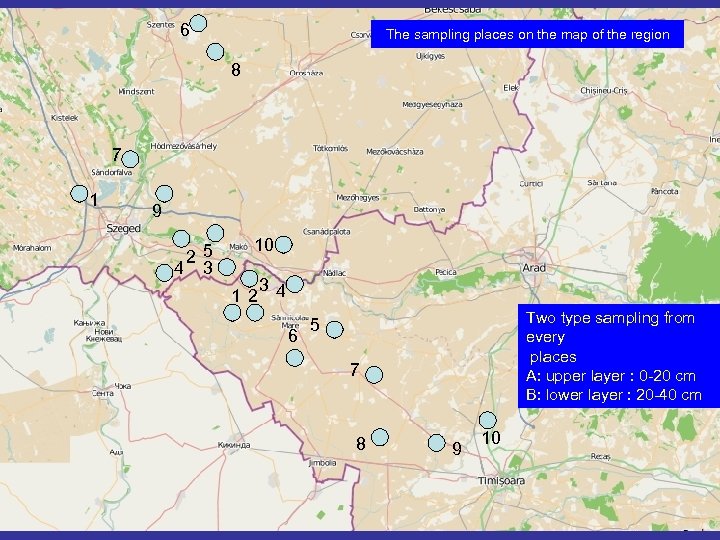
6 The sampling places on the map of the region 8 7 1 9 25 4 3 10 12 34 6 Two type sampling from every places A: upper layer : 0 -20 cm B: lower layer : 20 -40 cm 5 7 8 9 10
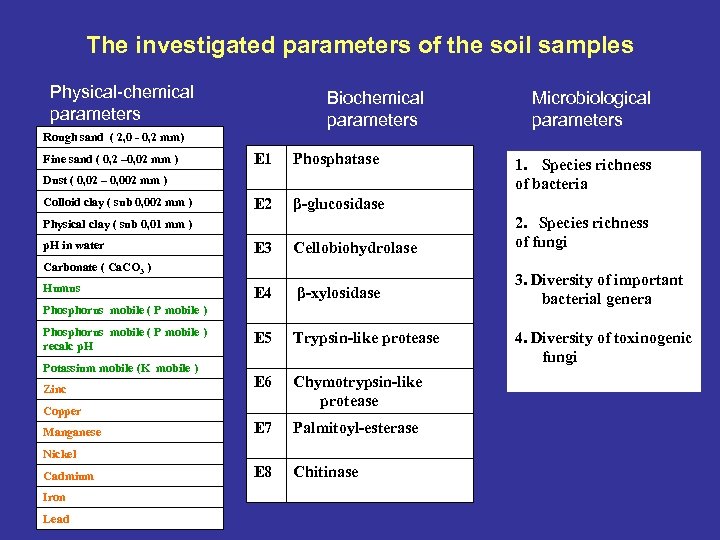

The investigated parameters of the soil samples Physical-chemical parameters Biochemical parameters Microbiological parameters Rough sand ( 2, 0 - 0, 2 mm) Fine sand ( 0, 2 – 0, 02 mm ) E 1 Phosphatase E 2 β-glucosidase Dust ( 0, 02 – 0, 002 mm ) Colloid clay ( sub 0, 002 mm ) E 3 Cellobiohydrolase 2. Species richness of fungi E 4 β-xylosidase 3. Diversity of important bacterial genera E 5 Trypsin-like protease E 6 Chymotrypsin-like protease E 7 Palmitoyl-esterase E 8 Chitinase Physical clay ( sub 0, 01 mm ) p. H in water Carbonate ( Ca. CO 3 ) Humus Phosphorus mobile ( P mobile ) recalc p. H Potassium mobile (K mobile ) Zinc Copper Manganese Nickel Cadmium Iron Lead 1. Species richness of bacteria 4. Diversity of toxinogenic fungi
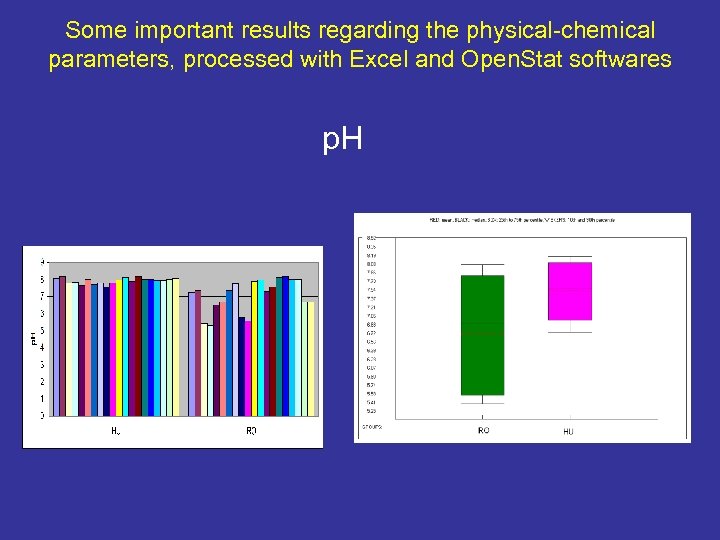
Some important results regarding the physical-chemical parameters, processed with Excel and Open. Stat softwares p. H
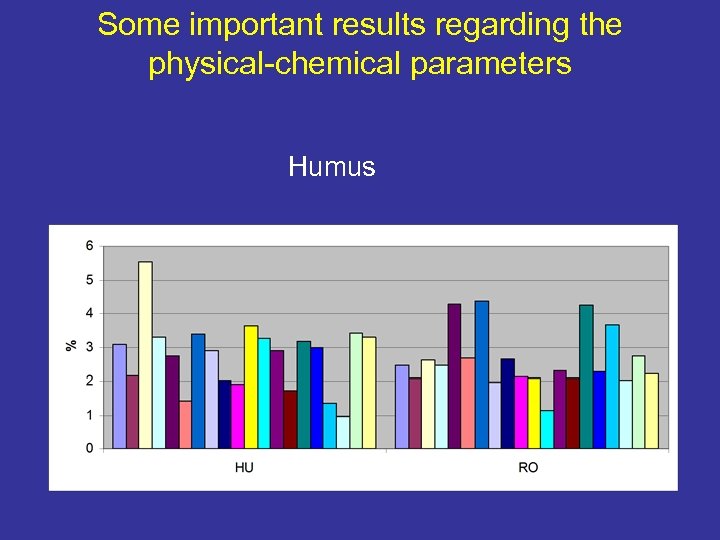
Some important results regarding the physical-chemical parameters Humus
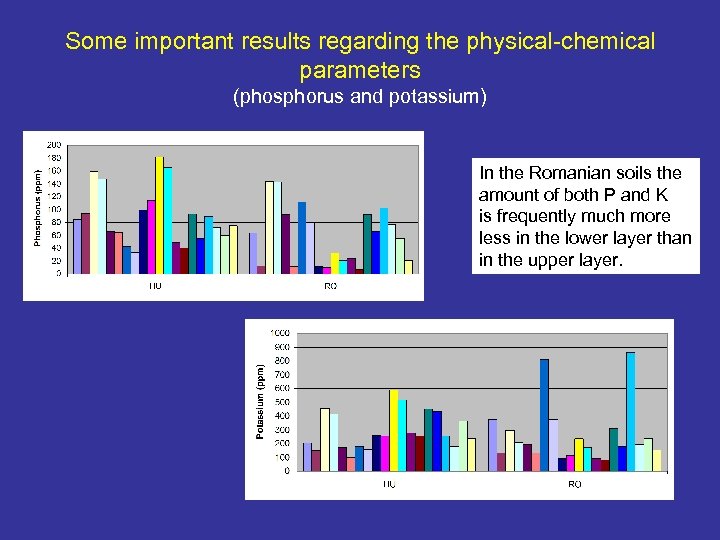
Some important results regarding the physical-chemical parameters (phosphorus and potassium) In the Romanian soils the amount of both P and K is frequently much more less in the lower layer than in the upper layer.
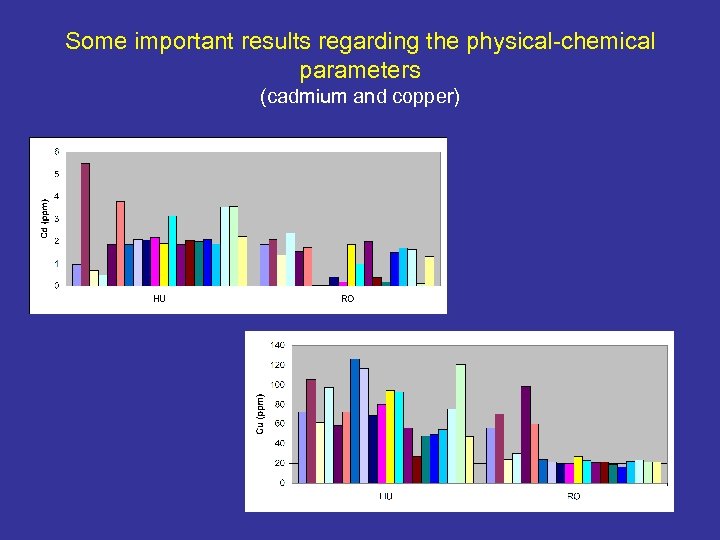
Some important results regarding the physical-chemical parameters (cadmium and copper)
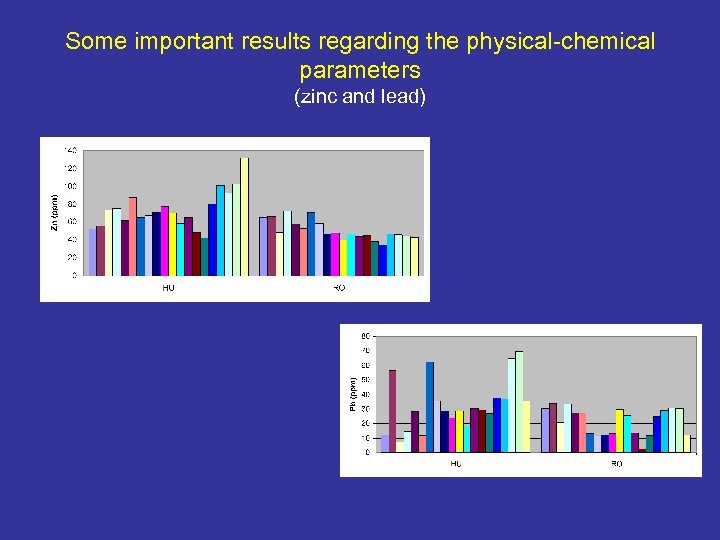
Some important results regarding the physical-chemical parameters (zinc and lead)
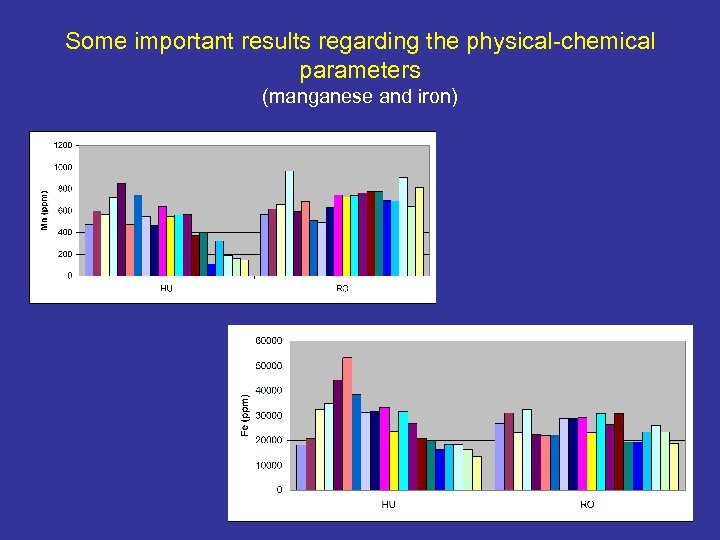
Some important results regarding the physical-chemical parameters (manganese and iron)
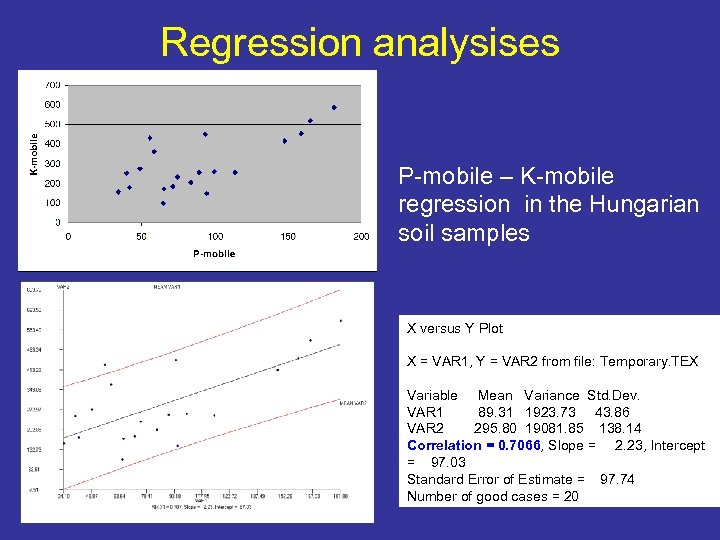
Regression analysises P-mobile – K-mobile regression in the Hungarian soil samples X versus Y Plot X = VAR 1, Y = VAR 2 from file: Temporary. TEX Variable Mean Variance Std. Dev. VAR 1 89. 31 1923. 73 43. 86 VAR 2 295. 80 19081. 85 138. 14 Correlation = 0. 7066, Slope = 2. 23, Intercept = 97. 03 Standard Error of Estimate = 97. 74 Number of good cases = 20
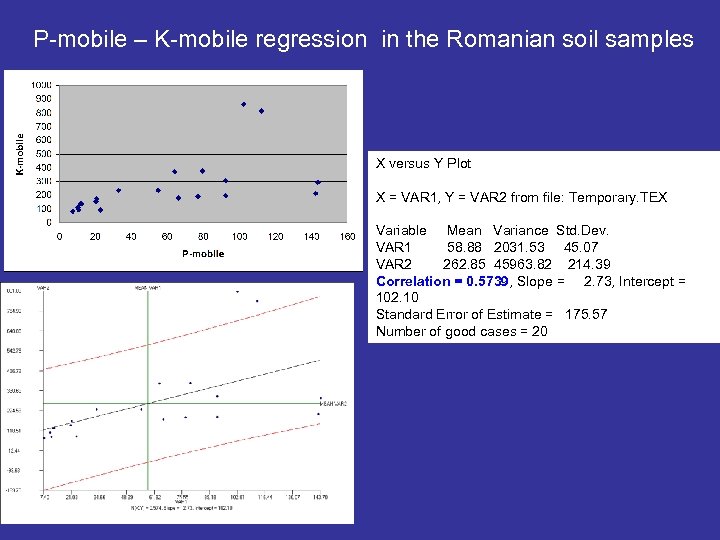
P-mobile – K-mobile regression in the Romanian soil samples X versus Y Plot X = VAR 1, Y = VAR 2 from file: Temporary. TEX Variable Mean Variance Std. Dev. VAR 1 58. 88 2031. 53 45. 07 VAR 2 262. 85 45963. 82 214. 39 Correlation = 0. 5739, Slope = 2. 73, Intercept = 102. 10 Standard Error of Estimate = 175. 57 Number of good cases = 20
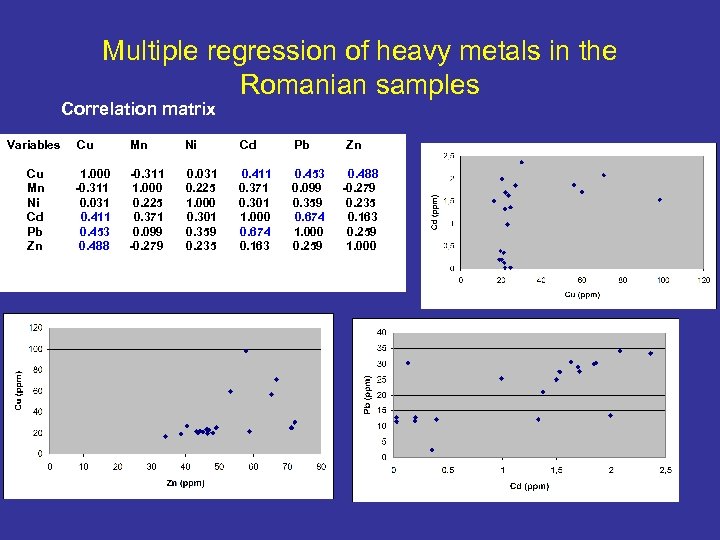
Multiple regression of heavy metals in the Romanian samples Correlation matrix Variables Cu Mn Ni Cd Pb Zn Cu Mn Ni Cd Pb 1. 000 -0. 311 0. 031 0. 411 0. 453 0. 488 -0. 311 1. 000 0. 225 0. 371 0. 099 -0. 279 0. 031 0. 225 1. 000 0. 301 0. 359 0. 235 0. 411 0. 371 0. 301 1. 000 0. 674 0. 163 0. 453 0. 099 0. 359 0. 674 1. 000 0. 259 Zn 0. 488 -0. 279 0. 235 0. 163 0. 259 1. 000
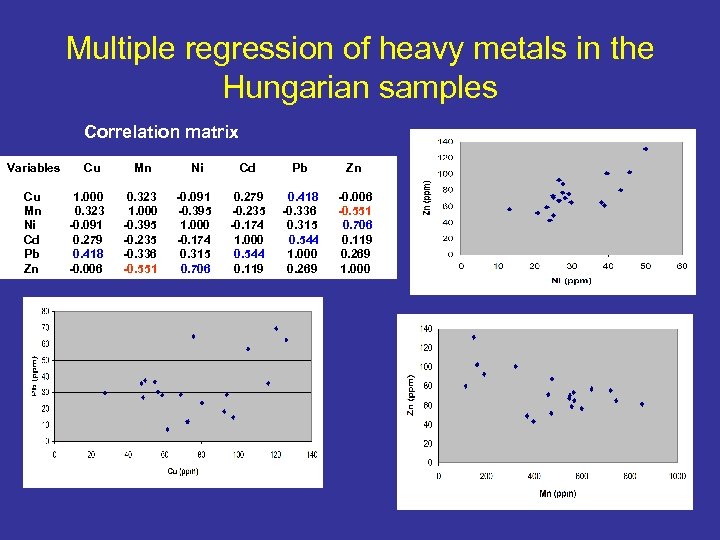
Multiple regression of heavy metals in the Hungarian samples Correlation matrix Variables Cu Mn Ni Cd Pb Zn Cu 1. 000 0. 323 -0. 091 0. 279 0. 418 -0. 006 Mn Ni Cd Pb Zn 0. 323 1. 000 -0. 395 -0. 235 -0. 336 -0. 551 -0. 091 -0. 395 1. 000 -0. 174 0. 315 0. 706 0. 279 -0. 235 -0. 174 1. 000 0. 544 0. 119 0. 418 -0. 336 0. 315 0. 544 1. 000 0. 269 -0. 006 -0. 551 0. 706 0. 119 0. 269 1. 000

Analysis of soil enzyme data
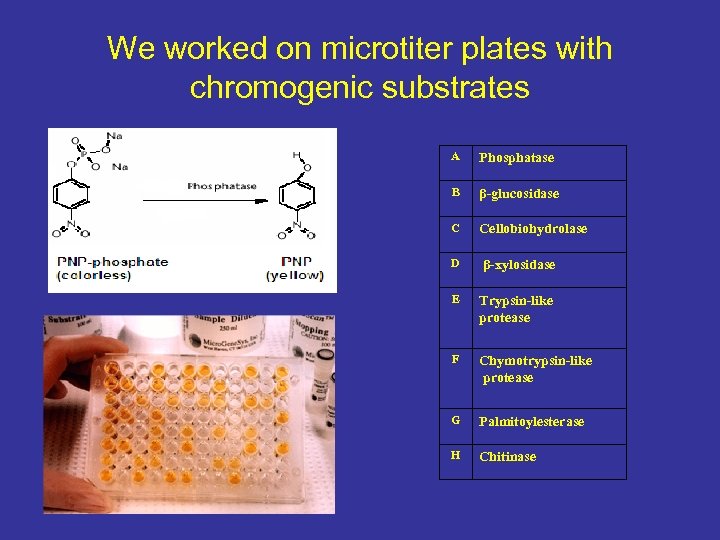

We worked on microtiter plates with chromogenic substrates A Phosphatase B β-glucosidase C Cellobiohydrolase D β-xylosidase E Trypsin-like protease F Chymotrypsin-like protease G Palmitoylesterase H Chitinase
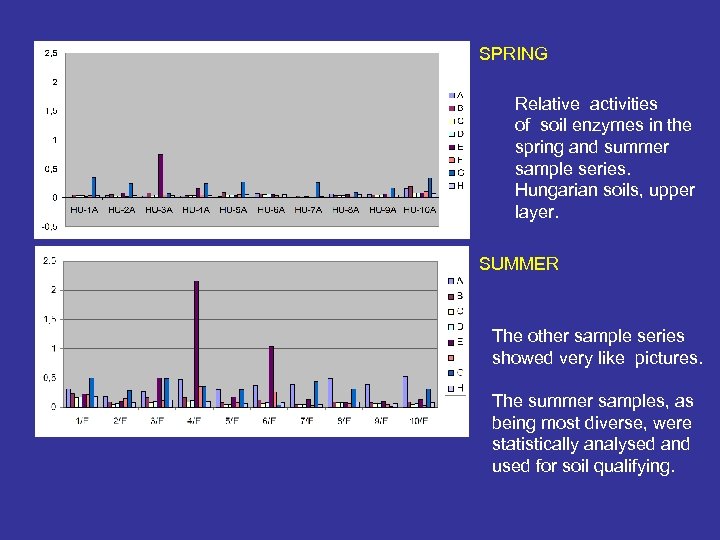
SPRING Relative activities of soil enzymes in the spring and summer sample series. Hungarian soils, upper layer. SUMMER The other sample series showed very like pictures. The summer samples, as being most diverse, were statistically analysed and used for soil qualifying.
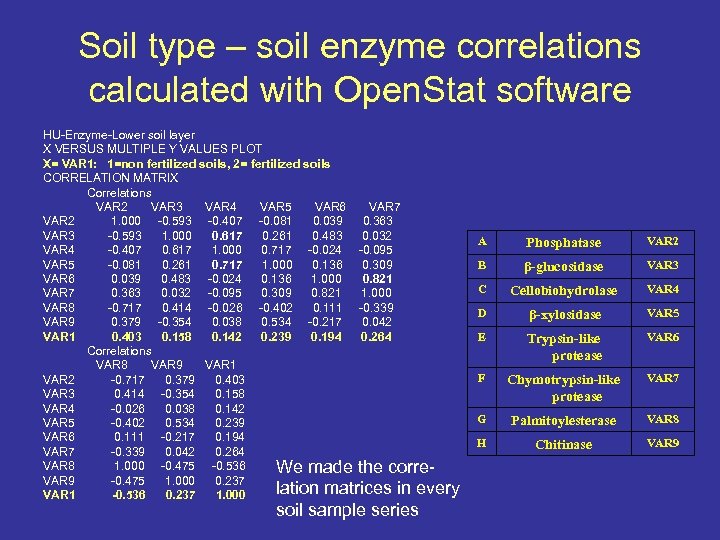
Soil type – soil enzyme correlations calculated with Open. Stat software HU-Enzyme-Lower soil layer X VERSUS MULTIPLE Y VALUES PLOT X= VAR 1: 1=non fertilized soils, 2= fertilized soils CORRELATION MATRIX Correlations VAR 2 VAR 3 VAR 4 VAR 5 VAR 6 VAR 7 VAR 2 1. 000 -0. 593 -0. 407 -0. 081 0. 039 0. 363 VAR 3 -0. 593 1. 000 0. 617 0. 261 0. 483 0. 032 VAR 4 -0. 407 0. 617 1. 000 0. 717 -0. 024 -0. 095 VAR 5 -0. 081 0. 261 0. 717 1. 000 0. 136 0. 309 VAR 6 0. 039 0. 483 -0. 024 0. 136 1. 000 0. 821 VAR 7 0. 363 0. 032 -0. 095 0. 309 0. 821 1. 000 VAR 8 -0. 717 0. 414 -0. 026 -0. 402 0. 111 -0. 339 VAR 9 0. 379 -0. 354 0. 038 0. 534 -0. 217 0. 042 VAR 1 0. 403 0. 158 0. 142 0. 239 0. 194 0. 264 Correlations VAR 8 VAR 9 VAR 1 VAR 2 -0. 717 0. 379 0. 403 VAR 3 0. 414 -0. 354 0. 158 VAR 4 -0. 026 0. 038 0. 142 VAR 5 -0. 402 0. 534 0. 239 VAR 6 0. 111 -0. 217 0. 194 VAR 7 -0. 339 0. 042 0. 264 VAR 8 1. 000 -0. 475 -0. 536 We made the corre. VAR 9 -0. 475 1. 000 0. 237 lation matrices in every VAR 1 -0. 536 0. 237 1. 000 soil sample series A Phosphatase VAR 2 B β-glucosidase VAR 3 C Cellobiohydrolase VAR 4 D β-xylosidase VAR 5 E Trypsin-like protease VAR 6 F Chymotrypsin-like protease VAR 7 G Palmitoylesterase VAR 8 H Chitinase VAR 9
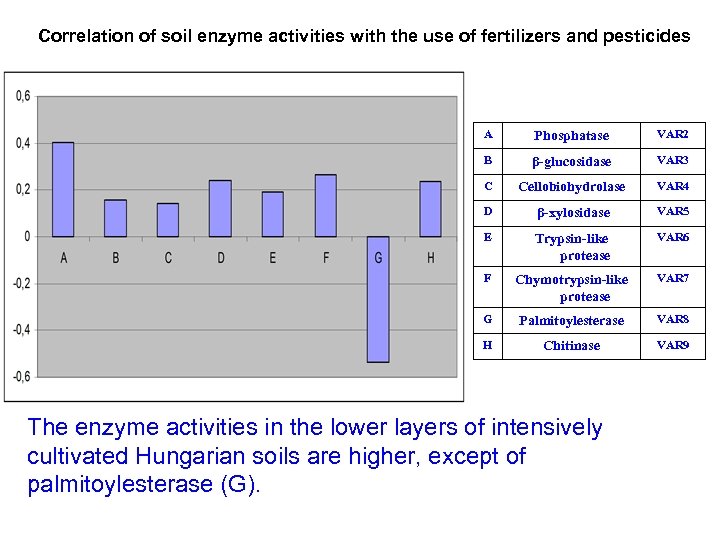
Correlation of soil enzyme activities with the use of fertilizers and pesticides A Phosphatase VAR 2 B β-glucosidase VAR 3 C Cellobiohydrolase VAR 4 D β-xylosidase VAR 5 E Trypsin-like protease VAR 6 F Chymotrypsin-like protease VAR 7 G Palmitoylesterase VAR 8 H Chitinase VAR 9 The enzyme activities in the lower layers of intensively cultivated Hungarian soils are higher, except of palmitoylesterase (G).
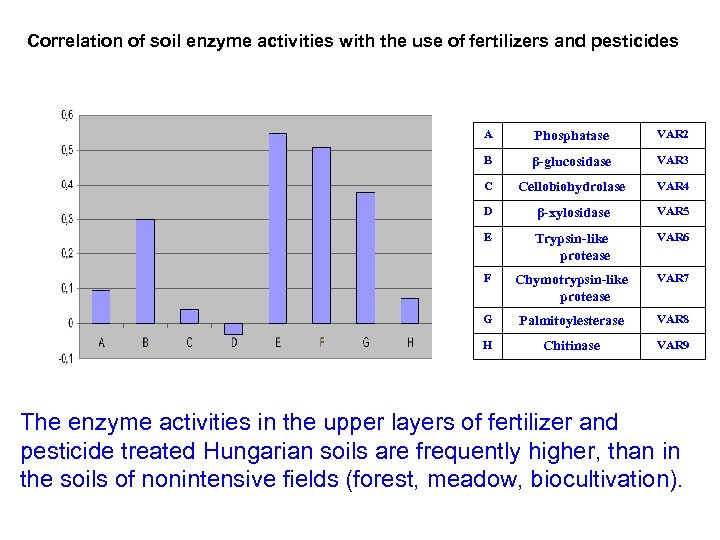

Correlation of soil enzyme activities with the use of fertilizers and pesticides A Phosphatase VAR 2 B β-glucosidase VAR 3 C Cellobiohydrolase VAR 4 D β-xylosidase VAR 5 E Trypsin-like protease VAR 6 F Chymotrypsin-like protease VAR 7 G Palmitoylesterase VAR 8 H Chitinase VAR 9 The enzyme activities in the upper layers of fertilizer and pesticide treated Hungarian soils are frequently higher, than in the soils of nonintensive fields (forest, meadow, biocultivation).
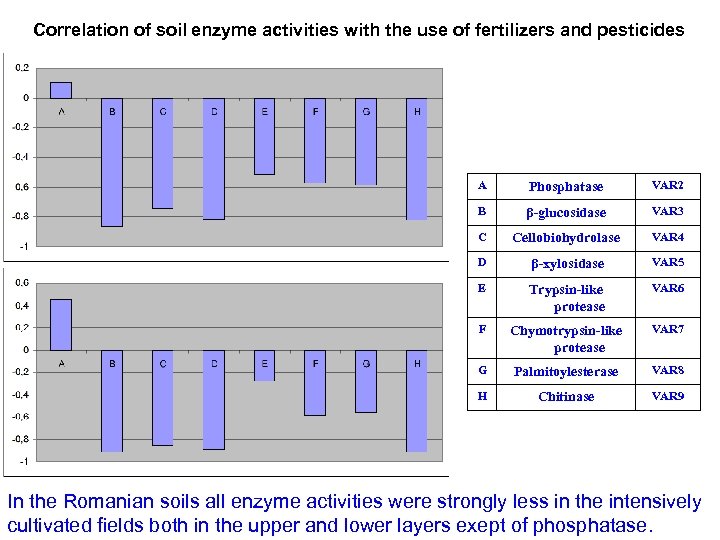
Correlation of soil enzyme activities with the use of fertilizers and pesticides A Phosphatase VAR 2 B β-glucosidase VAR 3 C Cellobiohydrolase VAR 4 D β-xylosidase VAR 5 E Trypsin-like protease VAR 6 F Chymotrypsin-like protease VAR 7 G Palmitoylesterase VAR 8 H Chitinase VAR 9 In the Romanian soils all enzyme activities were strongly less in the intensively cultivated fields both in the upper and lower layers exept of phosphatase.
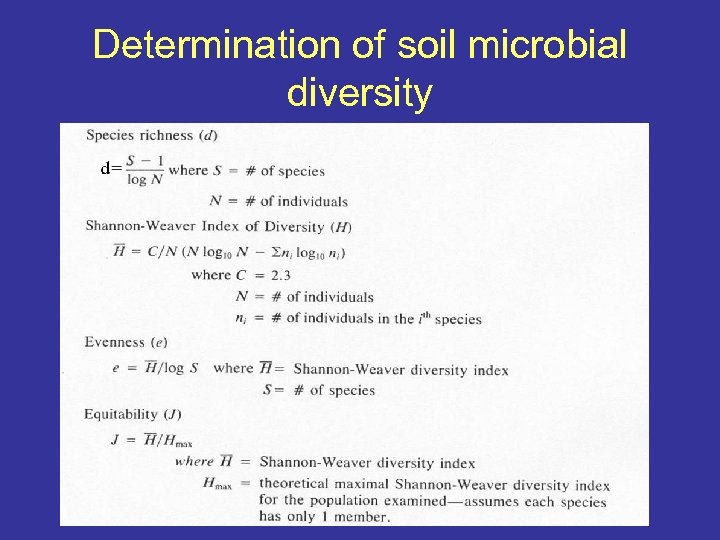

Determination of soil microbial diversity

The new molecular diversity methods - DGGE = Denaturing Gradient Gelelectrophoresis - TGGE = Temperature Gradient Gelelectrophoresis - TTGE= Temporal Temperature Gradient gelelectrophoresis - SSCP= Single Strand Conformational Polymorphism - RISA, ARISA (Automated) Ribosomal Intergenic Spacer Analysis - Community ARDRA, Community ITS RFLP - T-RFLP= Terminal Restriction Fragment Length Polymorphism
RISA Ribosomal Intergenic Spacer Analysis
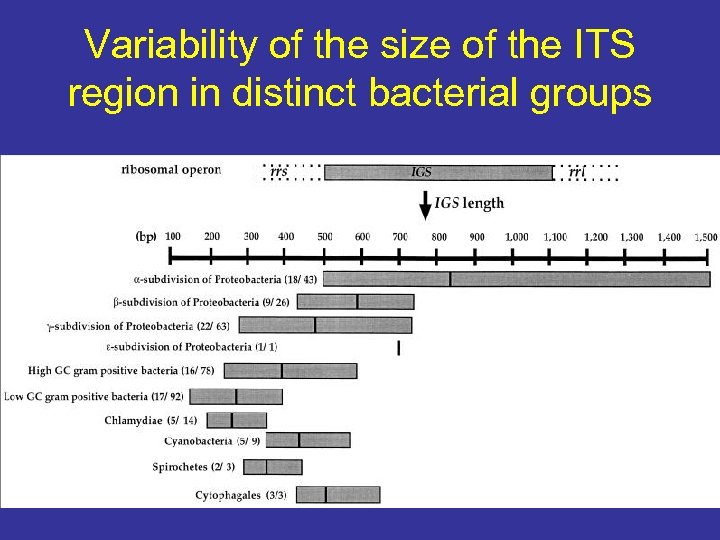
Variability of the size of the ITS region in distinct bacterial groups
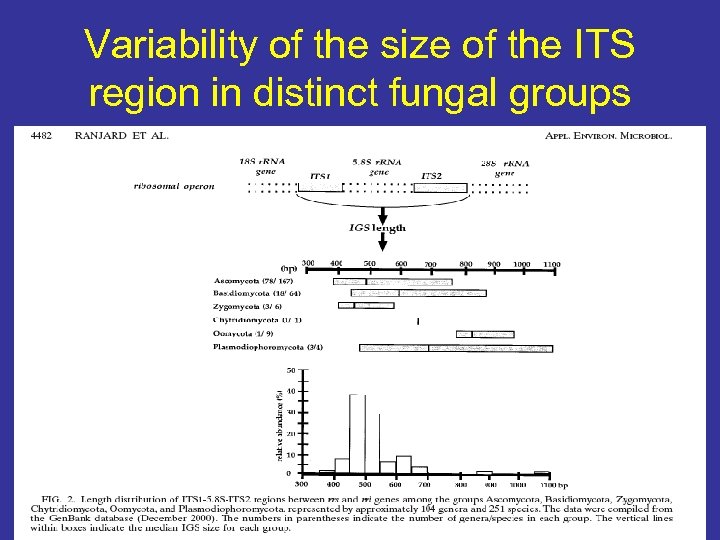

Variability of the size of the ITS region in distinct fungal groups
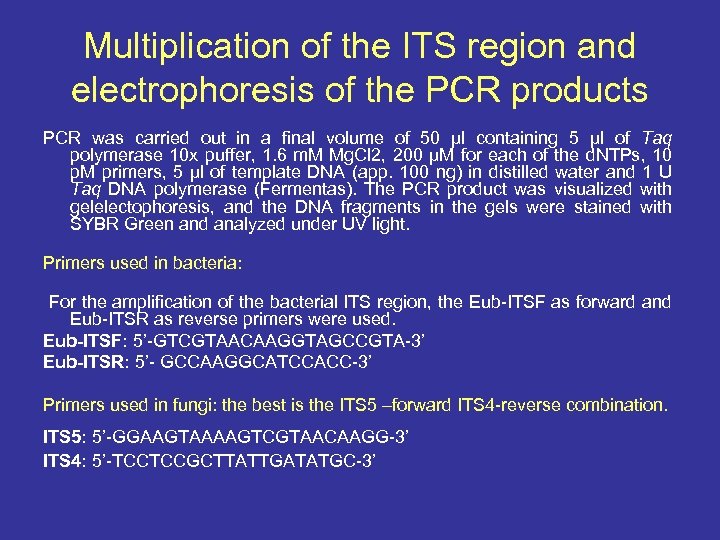

Multiplication of the ITS region and electrophoresis of the PCR products PCR was carried out in a final volume of 50 μl containing 5 μl of Taq polymerase 10 x puffer, 1. 6 m. M Mg. Cl 2, 200 μM for each of the d. NTPs, 10 p. M primers, 5 μl of template DNA (app. 100 ng) in distilled water and 1 U Taq DNA polymerase (Fermentas). The PCR product was visualized with gelelectophoresis, and the DNA fragments in the gels were stained with SYBR Green and analyzed under UV light. Primers used in bacteria: For the amplification of the bacterial ITS region, the Eub-ITSF as forward and Eub-ITSR as reverse primers were used. Eub-ITSF: 5’-GTCGTAACAAGGTAGCCGTA-3’ Eub-ITSR: 5’- GCCAAGGCATCCACC-3’ Primers used in fungi: the best is the ITS 5 –forward ITS 4 -reverse combination. ITS 5: 5’-GGAAGTAAAAGTCGTAACAAGG-3’ ITS 4: 5’-TCCTCCGCTTATTGATATGC-3’
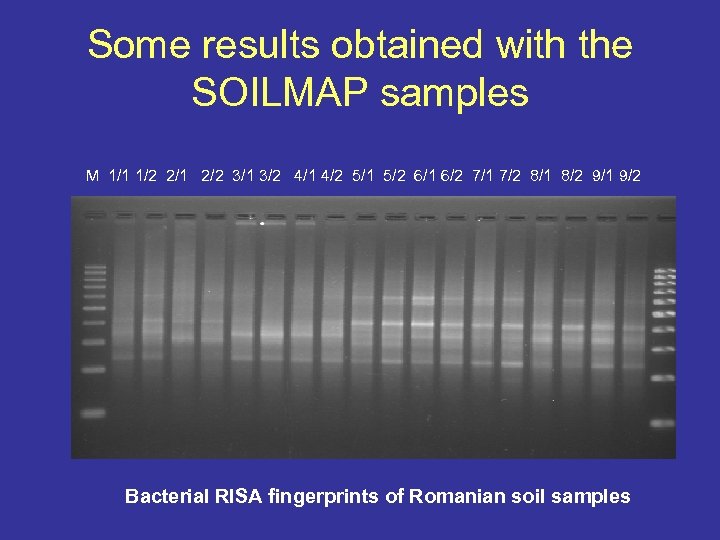
Some results obtained with the SOILMAP samples M 1/1 1/2 2/1 2/2 3/1 3/2 4/1 4/2 5/1 5/2 6/1 6/2 7/1 7/2 8/1 8/2 9/1 9/2 Bacterial RISA fingerprints of Romanian soil samples
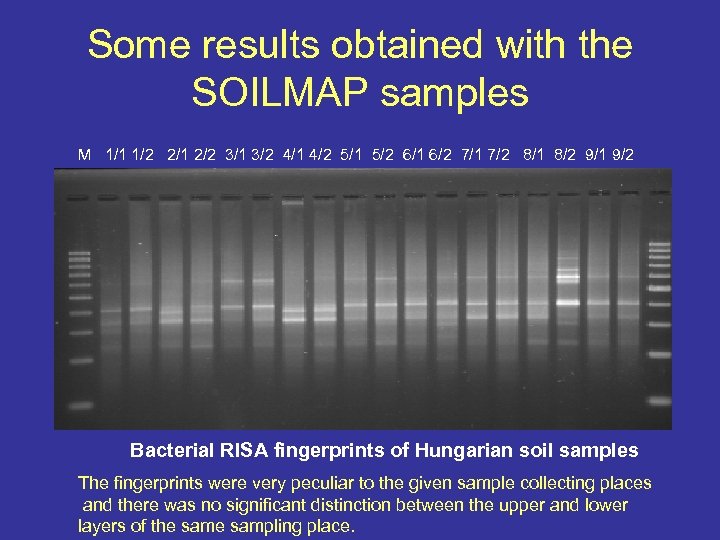

Some results obtained with the SOILMAP samples M 1/1 1/2 2/1 2/2 3/1 3/2 4/1 4/2 5/1 5/2 6/1 6/2 7/1 7/2 8/1 8/2 9/1 9/2 Bacterial RISA fingerprints of Hungarian soil samples The fingerprints were very peculiar to the given sample collecting places and there was no significant distinction between the upper and lower layers of the sampling place.
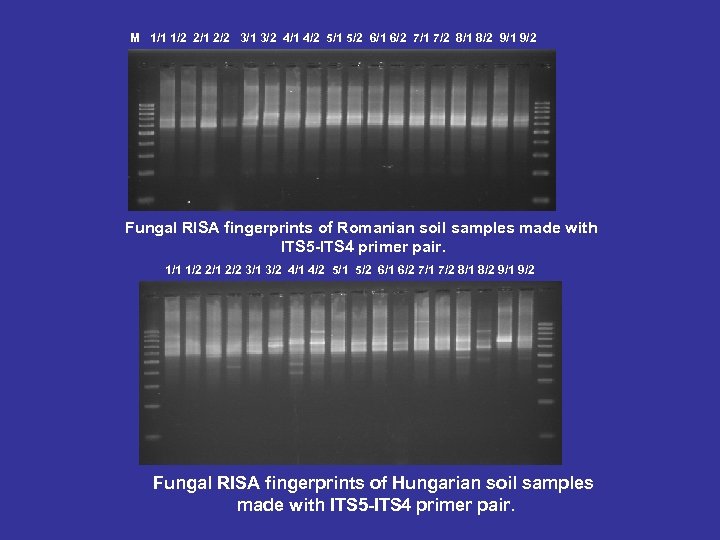
M 1/1 1/2 2/1 2/2 3/1 3/2 4/1 4/2 5/1 5/2 6/1 6/2 7/1 7/2 8/1 8/2 9/1 9/2 Fungal RISA fingerprints of Romanian soil samples made with ITS 5 -ITS 4 primer pair. 1/1 1/2 2/1 2/2 3/1 3/2 4/1 4/2 5/1 5/2 6/1 6/2 7/1 7/2 8/1 8/2 9/1 9/2 Fungal RISA fingerprints of Hungarian soil samples made with ITS 5 -ITS 4 primer pair.

As the fungal fingerprints were not enough diverse we used for soil qualifying the bacterial fingerprints only. Forest Meadow M 1/1 1/2 2/1 2/2 3/1 3/2 4/1 4/2 5/1 5/2 6/1 6/2 7/1 7/2 8/1 8/2 9/1 9/2 Pf Ba Str Bs 200 100 Bacterial RISA fingerprints of Hungarian soil samples
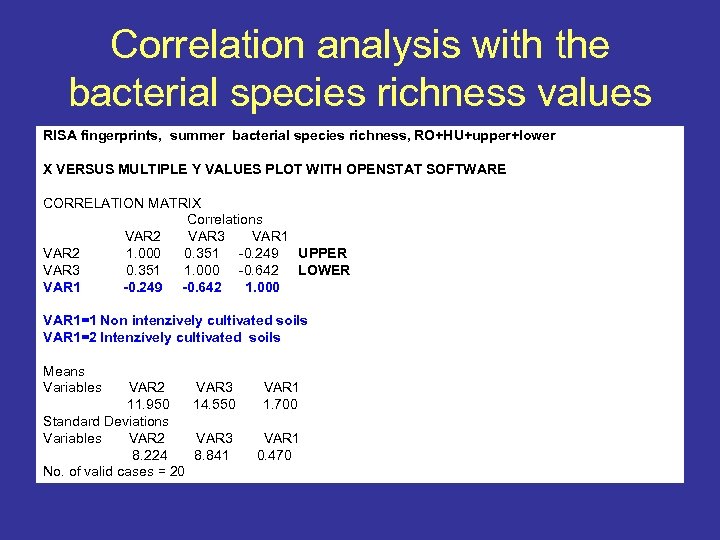

Correlation analysis with the bacterial species richness values RISA fingerprints, summer bacterial species richness, RO+HU+upper+lower X VERSUS MULTIPLE Y VALUES PLOT WITH OPENSTAT SOFTWARE CORRELATION MATRIX Correlations VAR 2 VAR 3 VAR 1 VAR 2 1. 000 0. 351 -0. 249 UPPER VAR 3 0. 351 1. 000 -0. 642 LOWER VAR 1 -0. 249 -0. 642 1. 000 VAR 1=1 Non intenzively cultivated soils VAR 1=2 Intenzívely cultivated soils Means Variables VAR 2 11. 950 Standard Deviations Variables VAR 2 8. 224 No. of valid cases = 20 VAR 3 14. 550 VAR 1 1. 700 VAR 3 8. 841 VAR 1 0. 470
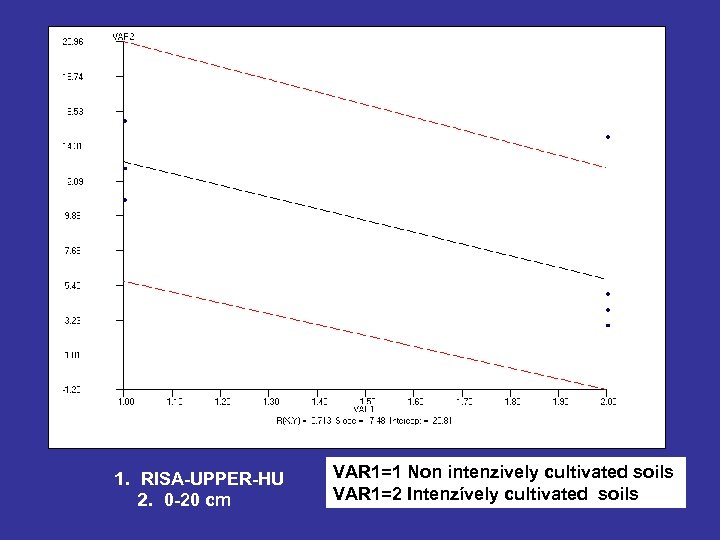
1. RISA-UPPER-HU 2. 0 -20 cm VAR 1=1 Non intenzively cultivated soils VAR 1=2 Intenzívely cultivated soils
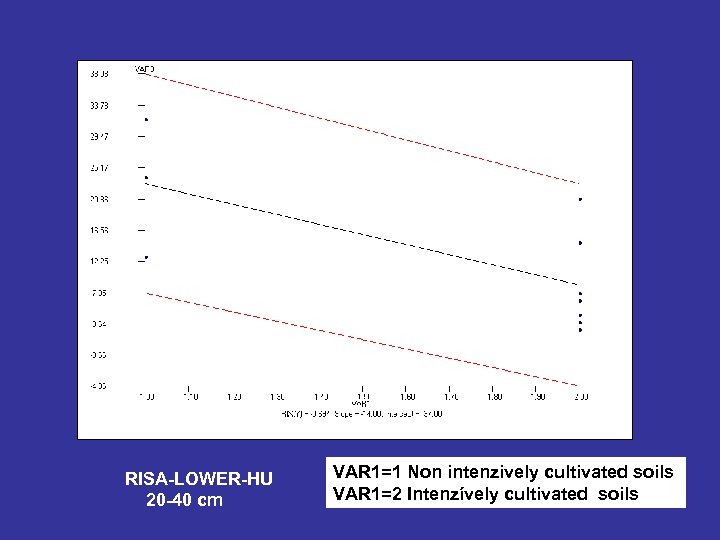
RISA-LOWER-HU 20 -40 cm VAR 1=1 Non intenzively cultivated soils VAR 1=2 Intenzívely cultivated soils
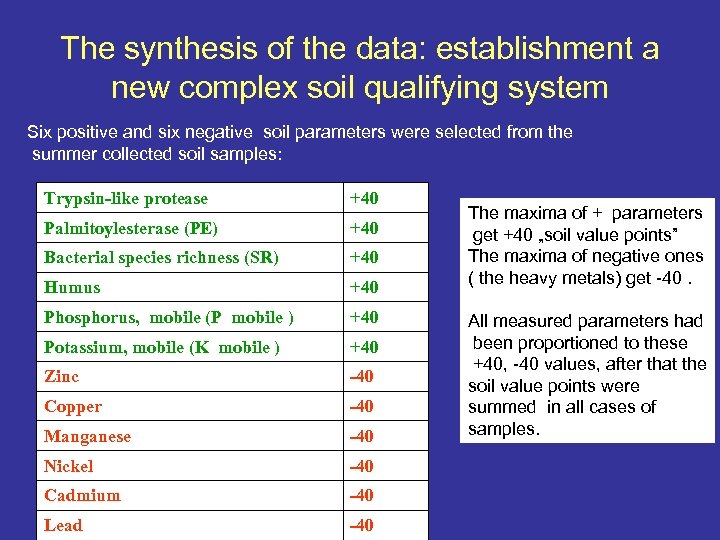

The synthesis of the data: establishment a new complex soil qualifying system Six positive and six negative soil parameters were selected from the summer collected soil samples: Trypsin-like protease +40 Palmitoylesterase (PE) +40 Bacterial species richness (SR) +40 Humus +40 Phosphorus, mobile (P mobile ) +40 Potassium, mobile (K mobile ) +40 Zinc -40 Copper -40 Manganese -40 Nickel -40 Cadmium -40 Lead -40 The maxima of + parameters get +40 „soil value points” The maxima of negative ones ( the heavy metals) get -40. All measured parameters had been proportioned to these +40, -40 values, after that the soil value points were summed in all cases of samples.
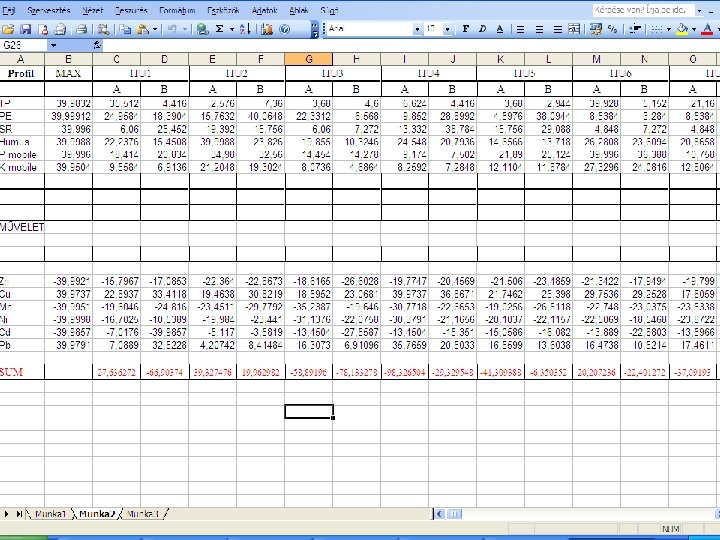

The quality values of Hungarian and Romanian soils A: 0 -20 cm , B. 20 -40 cm Forest soils: HU 2 and RO 4
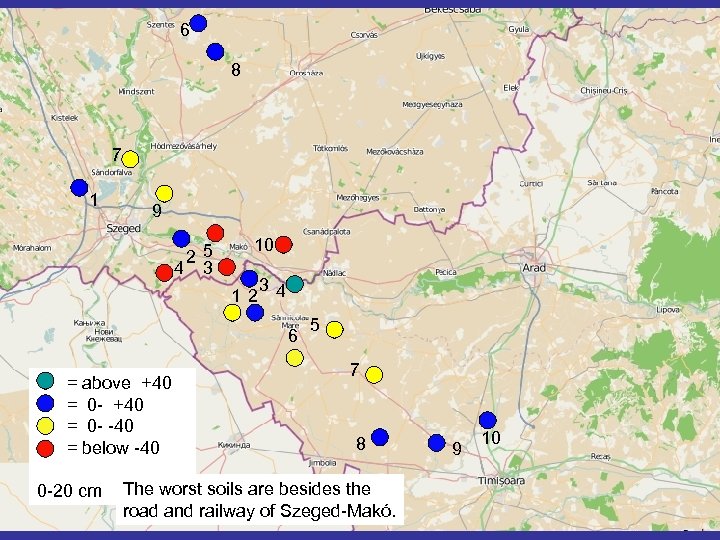
6 8 7 1 9 25 4 3 10 12 34 6 = above +40 = 0 - -40 = below -40 0 -20 cm 5 7 8 The worst soils are besides the road and railway of Szeged-Makó. 9 10
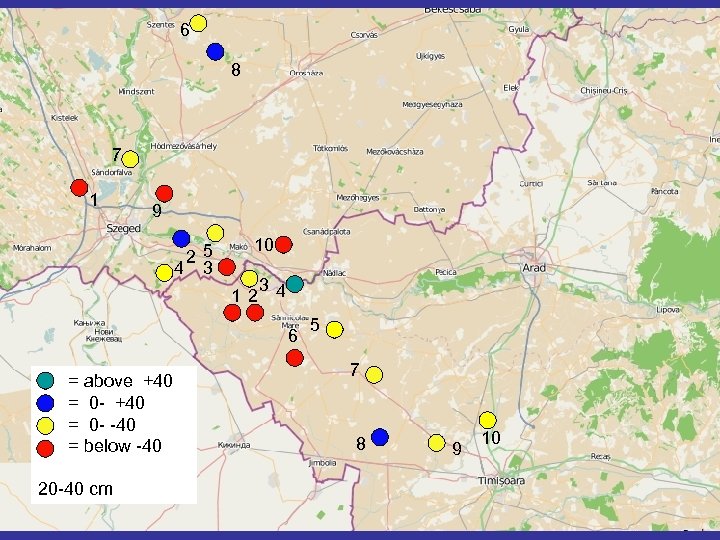
6 8 7 1 9 25 4 3 10 12 34 6 = above +40 = 0 - -40 = below -40 20 -40 cm 5 7 8 9 10
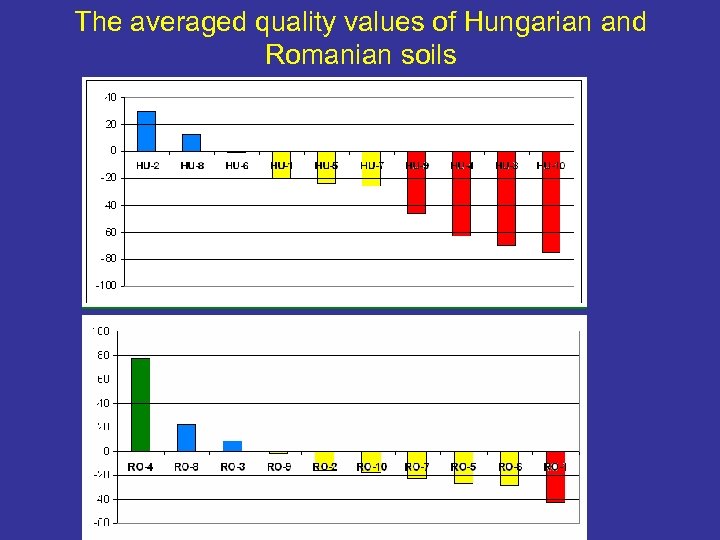
The averaged quality values of Hungarian and Romanian soils
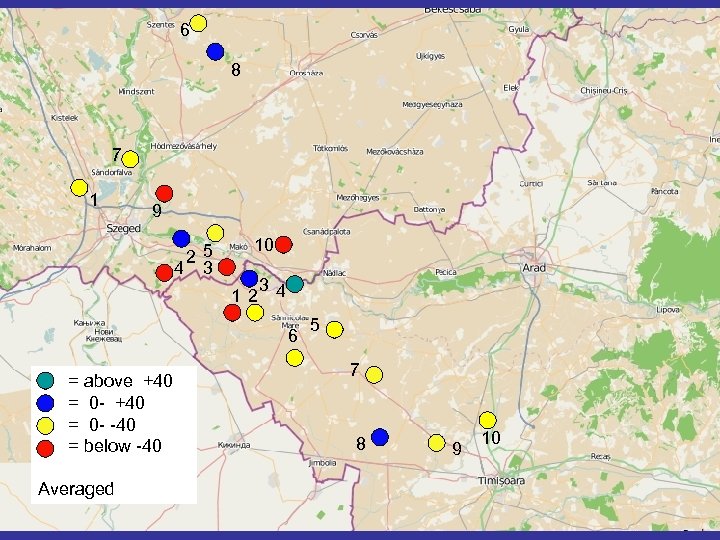
6 8 7 1 9 25 4 3 10 12 34 6 = above +40 = 0 - -40 = below -40 Averaged 5 7 8 9 10
Thank you for your attention!
6ed7318297b250e5600020cd4067af7a.ppt